GDMT (Golgi-derived microtubule) asymmetry is required for polarized cell motility, but its origin is elusive. Combining experimental and computational approaches, we find that GDMTs arise from spatially restricted hotspots that rely on γ-TuNA (γ-TuRC nucleation activator) activity. The nonrandom nucleation pattern underlies GDMT array asymmetry.
Abstract
Noncentrosomal microtubule (MT) nucleation at the Golgi generates MT network asymmetry in motile vertebrate cells. Investigating the Golgi-derived MT (GDMT) distribution, we find that MT asymmetry arises from nonrandom nucleation sites at the Golgi (hotspots). Using computational simulations, we propose two plausible mechanistic models of GDMT nucleation leading to this phenotype. In the “cooperativity” model, formation of a single GDMT promotes further nucleation at the same site. In the “heterogeneous Golgi” model, MT nucleation is dramatically up-regulated at discrete and sparse locations within the Golgi. While MT clustering in hotspots is equally well described by both models, simulating MT length distributions within the cooperativity model fits the data better. Investigating the molecular mechanism underlying hotspot formation, we have found that hotspots are significantly smaller than a Golgi subdomain positive for scaffolding protein AKAP450, which is thought to recruit GDMT nucleation factors. We have further probed potential roles of known GDMT-promoting molecules, including γ-TuRC-mediated nucleation activator (γ-TuNA) domain-containing proteins and MT stabilizer CLASPs. While both γ-TuNA inhibition and lack of CLASPs resulted in drastically decreased GDMT nucleation, computational modeling revealed that only γ-TuNA inhibition suppressed hotspot formation. We conclude that hotspots require γ-TuNA activity, which facilitates clustered GDMT nucleation at distinct Golgi sites.
INTRODUCTION
While the centrosome is traditionally referred to as the main microtubule (MT) organizing center (MTOC) in vertebrate cells, noncentrosomal MT nucleation plays an equally important role in MT array formation (Sanders and Kaverina, 2015; Dyachuk et al., 2016). A common location of interphase MT nucleation is the Golgi (Chabin-Brion et al., 2001; Efimov et al., 2007). Golgi-derived MTs (GDMTs) are found in cultured cells (Efimov et al., 2007; Rivero et al., 2009) and in multiple differentiated cell types (Chabin-Brion et al., 2001; Zaal et al., 2011; Ori-McKenney et al., 2012; Oddoux et al., 2013; Yalgin et al., 2015; Zhu et al., 2015). The first and best-characterized function of GDMTs is providing polarity to the MT array in migrating cells (Efimov et al., 2007). Unlike the radial centrosomal aster (Salaycik et al., 2005), GDMTs arise as a front-oriented asymmetric array (Efimov et al., 2007), which supports polarized post-Golgi trafficking and directional cell motility (Miller et al., 2009; Hurtado et al., 2011; Vinogradova et al., 2012). Although GDMT asymmetry is functionally important, its mechanistic basis is unknown. One possibility is that asymmetry arises from distribution of essential molecules to one side of Golgi stacks (specifically, cis-Golgi compartments). However, the complexity of Golgi ribbon folding and positioning makes this simple explanation unlikely. Here, we propose that geometry of the GDMT array is based on tight concentration of active nucleation factors in a small specialized domain of the Golgi membrane.
The factors required for GDMT formation include molecules responsible for 1) MT nucleation, 2) MT stabilization, and 3) scaffolding of both activities to the Golgi (reviewed in Sanders and Kaverina, 2015). GDMT nucleation requires a MT nucleation template, the γ-tubulin ring complex (γ-TuRC) (Efimov et al 2007), and MT nucleation activators containing γ-TuRC nucleation–activating (γ-TuNA, aa59-80 in CDK5RAP2; Choi et al., 2010) domains. γ-TuNA activates MT nucleation by binding to γ-TuRC and causing a change in the ring structure (Choi et al., 2010; Kollman et al., 2015). Two such proteins, CDK5RAP2 and myomegalin-8 (MMG8), partially redundantly support GDMT nucleation (Fong et al., 2008; Choi et al., 2010; Wang et al., 2010; Roubin et al., 2013; Wang et al., 2014). However, double knockout of these two factors decreases but does not eliminate GDMT nucleation (Wu et al., 2016), suggesting redundant support of GDMT nucleation by additional γ-TuNA-containing proteins (e.g., TPX2; Alfaro-Aco et al., 2017). γ-TuRC, CDK5RAP2, and MMG8 are thought to be recruited to the cis-Golgi membrane by a scaffolding protein, AKAP450 (AKAP-9, AKAP350, CG-NAP; Takahashi et al., 1999, 2002; Shanks et al., 2002; Rivero et al., 2009; Hurtado et al., 2011). AKAP450 has been reproducibly shown to be essential for GDMT formation (Rivero et al., 2009; Hurtado et al., 2011; Maia et al., 2013; Wu et al., 2016), possibly because it scaffolds multiple nucleation factors.
Another class of molecules important for GDMT formation is MT plus-tip interacting proteins (+TIPs), CLASPs (in mammals, CLASP1 and CLASP2), which localize to the trans-Golgi network (TGN) (Efimov et al., 2007). CLASPs are capable of promoting MT rescues and MT stability (Galjart, 2010). CLASP depletion or removal from the Golgi results in a severe reduction of GDMT levels, possibly through destabilization of newly nucleated MTs (Efimov et al., 2007).
Previously, we have shown that in human retinal pigment epithelial (RPE1) cells, GDMTs account for roughly half the entire MT population (Efimov et al., 2007). Yet, paradoxically, the essential nucleation template γ-TuRC is not enriched at the Golgi under physiological conditions, although it is found at isolated Golgi membranes (Chabin-Brion et al., 2001; Ori-McKenney et al., 2012) and in acentrosomal cells, where it is no longer recruited to the pericentriolar material (PCM; O’Rourke et al., 2014; Wu et al., 2016). Other major proteins implicated in GDMT nucleation concentrate at the centrosome either to a similar (CLASPs, AKAP450) or to a significantly greater (CDK5RAP2) extent than to the Golgi. The only exception is MMG8, which has been shown to localize to the Golgi rather than the centrosome (Wang et al., 2014). However, MMG8 knockout by itself is not sufficient to decrease GDMT nucleation (Wu et al., 2016).
How is the Golgi able to support nucleation of such a large MT subset without highly concentrated nucleation and stabilization factors? Using super-resolution microscopy in conjunction with quantitative modeling, we show that GDMT nucleation is restricted to a small number of hotspots at the Golgi, and that these hotspots require γ-TuRC activation by γ-TuNA. We propose that the hotspots represent small groups of highly concentrated active molecules at the Golgi where efficient MT nucleation can occur without recruitment of a large easily detectable molecule pool, and that uneven distribution of hotspots at the Golgi ribbon underlies GDMT array asymmetry in motile cells.
RESULTS AND DISCUSSION
To determine the distribution of microtubule nucleation on the Golgi, we expressed Emerald-EB3 or EB3-mCherry (MT +TIP marker) and a Golgi marker (TGN-RFP or GTN-GFP) in RPE1 and MRC-5 cells and imaged them by spinning disk confocal microscopy. Newly formed GDMTs were detected by precise location of newly formed EB3 comets in 3D space in close proximity to the Golgi. This mapping of microtubule nucleation sites indicated that GDMTs form close together (Figure 1, A and B). In these cells, more than half of newly formed GDMTs were formed within 0.6 µm of each other, suggesting that nucleation sites are clustered at specific hotspots at the Golgi (Figure 1C). Moreover, each group of GDMTs formed within a short amount of time (within ∼25 s), after which there was often no further nucleation observed at that site (Figure 1, B, E, and F; Supplemental Movie S1). These observations suggest that these specific sites exist temporarily to support MT nucleation. Thus, the MT nucleation-supporting hotspots at the Golgi are restricted in both space and time.
FIGURE 1:
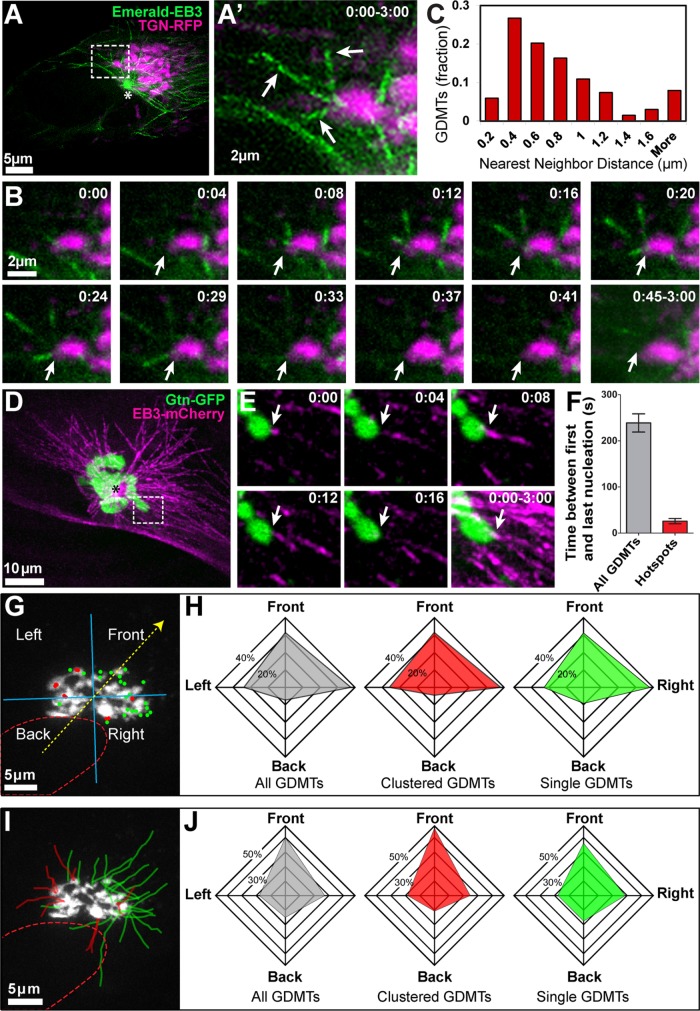
Microtubules are nucleated at specific sites within the Golgi ribbon. (A) An RPE1 cell expressing Emerald-EB3 (green, MT +TIP marker) and TGN-RFP (magenta, Golgi marker). A maximum-intensity projection of a confocal spinning disk microscopy sequence over a 3-min period and Z thickness 3 μm is shown (A, A′). Inset in A is enlarged in A′, showing newly formed GDMTs formed at the same site (arrows). (B) Single–time point maximum-intensity Z-projections from A′ show that clustered GDMTs (arrows) form within a short period of time. Arrows: clustered GDMTs; asterisks: centrosomal MTs. Time, minutes:seconds. Emerald-EB3 (green), TGN-RFP (magenta). (C) Quantification of nearest-neighbor distance between GDMT nucleation sites, based on 3D live-time imaging as in A and B. (D) MRC-5 cells expressing EB3-mCherry (magenta, MT +TIP marker) and GTN-GFP (green, Golgi marker). Maximum-intensity projection as described in A is shown. (E) Single–time point maximum-intensity projections are shown as described in B. Arrows: clustered GDMTs. Time, minutes:seconds. EB3-mCherry (magenta), GTN-GFP (green). (F) Time between GDMT nucleation events. Average time between first and last GDMT nucleation events was calculated over a 5-min period and within hotspots (GDMT nucleation events within 0.4 µm of each other). Error bars; SD. (p < 0.001, Student’s t test, n = 10 cells and 30 hotspots). (G) Distribution of GDMT nucleation sites on the Golgi, depicted over a maximum-intensity Z-projection of the TGN-RFP signal as a Golgi marker (white). The nucleus–Golgi axis was used to determine cell polarity (yellow dotted arrow) and four quadrants were generated to categorize nucleation site placement into four quadrants according to this axis (blue cross). Dots indicate GDMT nucleation events observed over a period of 3 min. Red dots: clustered GDMT nucleation events <0.4 µm apart. Green dots: single GDMT nucleation events. Red dotted line: nucleus. (H) Distribution of GDMT nucleation sites on the Golgi relative to cell polarity axis. Polarity quadrants were determined as in G. Front- or side-oriented nucleation sites occur more often than back-oriented nucleation sites (p < 0.001, χ2 test, n = 10 cells). (I, J) Distribution of GDMT directionality. (I) GDMT tracks were generated using the MTrackJ plugin for Image. Red tracks denote clustered GDMTs (nucleation sites <0.4 µm apart); green tracks are single GDMTs. (J) Relative distribution of GDMT directionality. For each GDMT track (as in I), the blue cross denoting the four quadrants (generated as in G) was centered at the nucleation site and MT directionality was determined. Front- or side-oriented directionality was more prevalent than back-oriented directionality (p < 0.05, χ2 test, n = 10 cells).
Our previous work showed that in motile cells the GDMT array extends asymmetrically toward the cell front (Efimov et al., 2007). To determine whether the distribution of MT nucleation sites at the Golgi ribbon is involved in this asymmetry, we analyzed GDMT nucleation positions and growth directionality as compared with the nucleus/Golgi polarity axis (Uetrecht and Bear, 2009). MT nucleation sites at the Golgi were located predominantly at the periphery of the Golgi complex (Figure 1G) and were distributed asymmetrically in the front three-quarters of the Golgi (Figure 1H). Each GDMT extended away from the Golgi center of mass, and, in correlation with their nucleation positions, most GDMTs were directed toward the leading edges and sides of polarized cells (Figure 1, I and J), possibly providing new tracks for post-Golgi vesicles (Miller et al., 2009; Hurtado et al., 2011). Interestingly, the directionality of hotspot-nucleated GDMTs (clustered within 0.4 μm from each other; see below) and their positioning at the Golgi ribbon were indistinguishable from those of single GDMTs as well as of the whole GDMT population, indicating that MT nucleation sites where single MTs were detected likely have the same nature as MT nucleation hotspots.
The results summarized above are based on time-lapse videos that follow GDMT nucleation in the steady state at 37°C, which is an important setup close to physiological conditions. However, MT growth rates are too high (166 ± 43 nm/s) to allow the first appearance of MTs to be detected precisely, given the time resolution of 3D time-lapse imaging (∼4.5 s/frame), which might result in seemingly greater distances between the nucleation sites. To analyze GDMT nucleation without the background of preexisting MTs and with greater time resolution, we depolymerized all MTs with nocodazole and followed MT regrowth in real time (Supplemental Movies S2 and S3). In this assay, nocodazole was washed out with ice-cold media while samples were on the microscope stage. By slowly warming up the media, initial MT nucleation was slowed, providing better temporal resolution in tracking MT nucleation events. At the same time, this procedure increases the number of simultaneously formed MTs, probably due to high levels of free tubulin dimers in the cytosol, and allows us to better evaluate the MT-nucleating capacity of the Golgi membrane. Figure 2, A, C, and D, shows several examples of multiple MTs formed at tightly restricted hotspots at several Golgi fragments (arrows). In comparison with the steady-state conditions, MTs appear to nucleate closer to each other (Figure 2B), probably due to higher temporal resolution.
FIGURE 2:
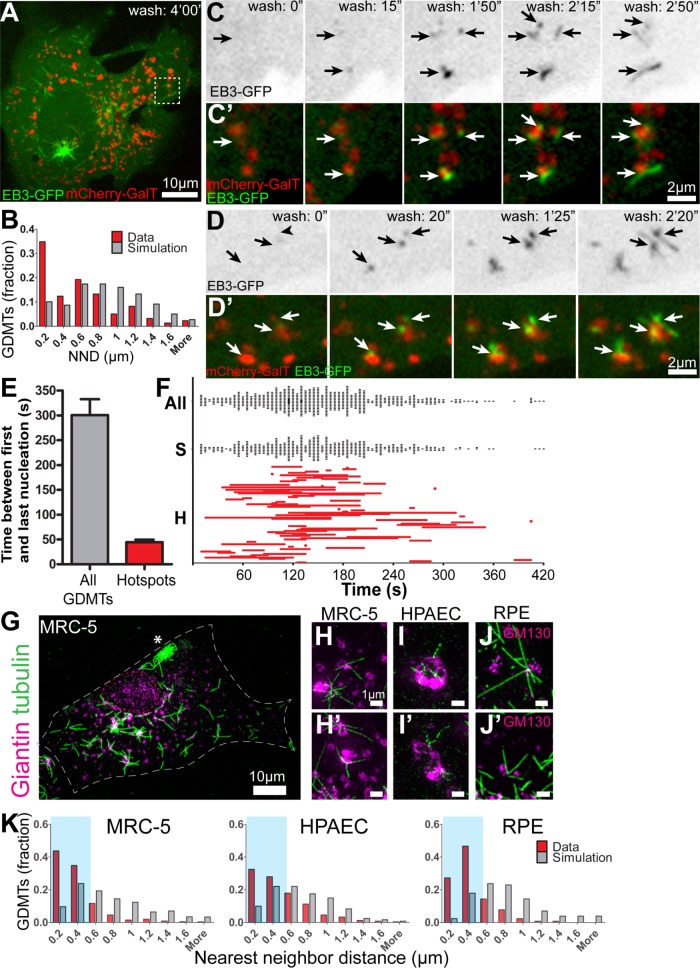
GDMT nucleation is spatially restricted to distinct hotspots. (A) An RPE1 cell expressing EB3-GFP (green) and mCherry-GalT (red, Golgi marker) 4 min after nocodazole washout. Single-plane confocal spinning disk microscopy. Insets are shown over time in C, C′, D, and D′. Asterisk: centrosomal MTs. (B) Newly formed GDMTs are distributed nonrandomly on Golgi fragments following nocodazole washout. Nearest-neighbor distances of GDMT minus ends were calculated for each fragment associated with multiple GDMTs. A paired random data set was generated using Matlab (p < 0.001, Student’s t test, n = 9 cells). Based on data as in A, C, and D. (C, D) Examples of simultaneous multiple GDMT nucleation events (arrows) at Golgi fragments following nocodazole washout. Frames from a time-lapse image sequence. (C, D) EB3-GFP, inverted grayscale image. (C′, D′) EB3-GFP (green) and mCherry-GalT (red, Golgi marker). Time from the start of the movie, minutes:seconds. (E) Time between GDMT nucleation events. Average time between first and last GDMT nucleation event was calculated over a 7-min period and within hotspots (GDMT nucleation events within 0.4 µm of each other). Error bars: SD. (p < 0.001, Student’s t test, n = 9 cells and 76 hotspots.) (F) Distribution of GDMT nucleation events and hotspot duration over time. GDMT nucleation events are plotted over a 7-min period, based on data from E. All GDMTs (All) and single GDMT nucleation events are plotted as single data points. Duration of hotspots (H) is plotted from first to last nucleation event within each hotspot. All, all GDMTs; S, single GDMT nucleation events; H, hotspots. (G–J) Examples of GDMT clustering in different cell types 40 s after nocodazole washout. Immunofluorescence. (G) An MRC-5 cell laser scanning confocal microscopy overview image (maximum-intensity Z-projection; see also Supplementary Figure S1, A–C, for other cell types). Tubulin, green. Giantin, magenta. (H–J′) SIM image maximum-intensity projections. Tubulin, green. GM130, magenta. Note GDMTs clusters extending from a Golgi fragment in each cell type. (K) Distribution of nearest-neighbor distances between GDMT minus end positions on Golgi fragments producing multiple GDMTs. A paired random data set was generated using Matlab. (p < 0.001, Student’s t test, n = 8-–10 cells per cell type.)
To better understand the dynamics of MT nucleation at the hotspots, we next analyzed the timing of GDMT nucleation within them. GDMT formation increases while the medium temperature rises within the first minute after washout, and the nucleation rate starts to decrease 3 min later as the free tubulin pool is depleted. We found that MTs within hotspots form at significantly shorter intervals than the whole GDMT population (Figure 2, C– F; Supplemental Movies S2 and S3), which is consistent with our findings in the steady state (Figure 1F). This behavior indicates that molecular complexes acting as functional hotspots are rapidly formed and inactivated, either through dissolution or through saturation.
To investigate the organization of GDMTs on the Golgi with more precision, we then turned to structured illumination microscopy (SIM) of fixed, immunostained cells. MT regrowth in these experiments was timed after introduction of an acute warm medium as a trigger for immediate nucleation, and cells were fixed after 40 s, which is within the active nucleation period and significantly before the nucleation rates start to decay. GDMTs were clustered on Golgi fragments in epithelial cells (RPE1), primary lung fibroblasts (MRC-5), and primary pulmonary artery endothelial cells (HPAEC), showing that this phenomenon is conserved in different lineages (Figures 2G– 2J). Further analysis of the MT minus-ends on Golgi fragments showed that newly formed GDMTs are nonrandomly distributed across the Golgi (Figure 2K), with a significant fraction of GDMT minus ends clustered within 0.4 µm from each other. To standardize our analyses based on these observations, we define GDMT nucleation hotspots as spheres with a diameter of 0.4 μm containing both Golgi membranes and multiple MT minus ends.
We speculate that GDMT nucleation hotspots could be formed through several mechanisms. In one scenario, when a single GDMT is formed, it can subsequently recruit MT nucleation and stabilizing factors to the same site at the Golgi, promoting nucleation of additional GDMTs (cooperativity hypothesis). Such recruitment can occur, for example, via dynein or other minus end–directed MT motors and would suggest sequential nucleation within hotspots over time. Interestingly, the important MT nucleation activator CDK5RAP2 is collected at the centrosome by dynein-dependent transport (Jia et al., 2013). Alternatively, specific sites at the Golgi membrane can become enriched with MT nucleation and stabilization factors independent of MT presence (heterogeneous Golgi hypothesis). This mechanism would be able to support simultaneous clustered nucleation events.
To test whether our data are consistent with these hypotheses, we turned to computational simulations and fitted four models (summarized in Figure 3A) of GDMT nucleation to quantitative imaging data. As a null model to compare against, the “no cooperativity” model assumes that all nucleation is random in space and that hotspots arise through random coincidence as MTs nucleate near each other. The “heterogeneous Golgi” model assumes similarly that nucleation is random in space, but that there are small (0.4 μm diameter) circular regions of the Golgi where nucleation occurs at a higher rate. To test the cooperativity hypothesis, we considered two variants. In both, the entire membrane has the capacity to randomly nucleate single microtubules at a fixed rate. In the “constrained cooperativity” model, MTs can nucleate in one of two ways: 1) random nucleation anywhere on the membrane at a low rate or 2) nucleation of a new MT from an existing hotspot at a higher rate. This model is “constrained” in the sense that all hotspots are assumed to nucleate new MTs at the same rate. To account for potential size-dependent effects, we also include a more flexible “cooperativity” model where hotspots of different sizes can nucleate new MTs at different rates. Each of these models is fitted to quantitative data of hotspot size, both to assess the reasonableness of that model and to make mechanistic inferences based on the values of best-fit parameters (see Materials and Methods and Supplemental Materials and Methods for further details).
FIGURE 3:
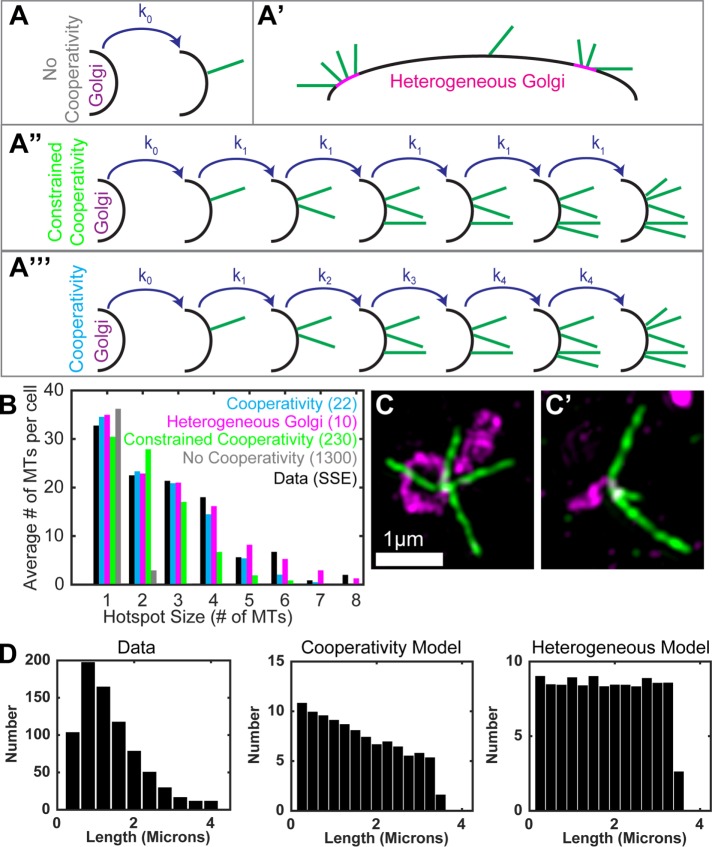
Two potential mechanistic models underlying GDMT nucleation hotspot formation. (A) Schematic representation of the four models to be compared against data. (A) No cooperativity: GDMTs are randomly nucleated. (A′) Heterogeneous Golgi: Small, focal subdomains of the Golgi membrane have increased MT nucleation capability. (A″) Constrained cooperativity: GDMT hotspots promote nucleation of additional GDMTs, independent of number of GDMTs at the hotspot (hotspot size-independent). (A′″) Cooperativity: GDMT hotspots promote nucleation of additional GDMTs at the same site, dependent on number of MTs at the hotspot (hotspot size-dependent). (B) Comparison of best-fit results for each model against data. The vertical axis quantifies the total number of GDMTs contained within hotspots of each size, averaged over the eight cells imaged (100 cells simulated in the case of model results). Sum of squared errors (SSE) between the data and model distributions indicates the quality of fit for each model. (C) Examples of hotspots with GDMTs of similar (C) or varying (C′) length in MRC-5 cells 40 s after nocodazole washout. SIM image maximum intensity projections. Tubulin, green; Giantin, magenta. (D) Comparison of simulated microtubule length distributions against data.
Based on these simulations, there is no clear preference for either the cooperativity or heterogeneous Golgi model (though the constrained cooperativity and no cooperativity models are rejected). To provide potentially distinguishing evidence, we have analyzed the length distribution of hotspot MTs (Figure 3D) and found that it resembles the simulated results of the cooperativity model more closely than that of the heterogeneous Golgi model. We thus lean toward the cooperativity model as a working hypothesis. However, we cannot exclude the possibility that additional regulation exists that leads to such a distribution through noncooperative dynamics of heterogeneous Golgi membranes. It is also possible that both heterogeneous and cooperative mechanisms are combined to a certain degree in regulating molecular complexes serving as GDMT nucleation hotspots in real cells.
Regardless of the mechanism that restricts nucleation to the hotspots, these sites should have a specific molecular composition allowing rapid and efficient MT nucleation. It is not clear so far what constitutes a hotspot. While a number of molecules have been shown essential for GDMT formation, most of them do not robustly localize to the Golgi. For example, cytosolic concentration of γ-tubulin is high (>80%, Moudjou et al., 1996, or even 99%, Bauer et al., 2016, of cellular γ-tubulin content), but it is not enriched at the Golgi unless extensive experimental perturbation is introduced (Ríos et al., 2004; Wang et al., 2014). The best characterized molecule that is absolutely essential for GDMT nucleation is AKAP450, which is referred to as a cis-Golgi protein (Rivero et al., 2009; Hurtado et al., 2011). Here, we find that AKAP450 localizes to a subdomain of cis-Golgi (Figure 4, B and E). Consistent with its role in MT nucleation, GDMT’s nucleation sites are concentrated at AKAP450-positive fragments (Figure 4, A and C, and Supplemental Figure S1I). However, the AKAP450-positive domain is significantly larger than the average area covered by hotspots, which does not exceed 7.3 ± 2.8 µm2 per cell (given our quantification of the hotspot number and estimated size). This indicates that while AKAP450 is required, it is insufficient for formation of GDMT nucleation hotspots.
FIGURE 4:
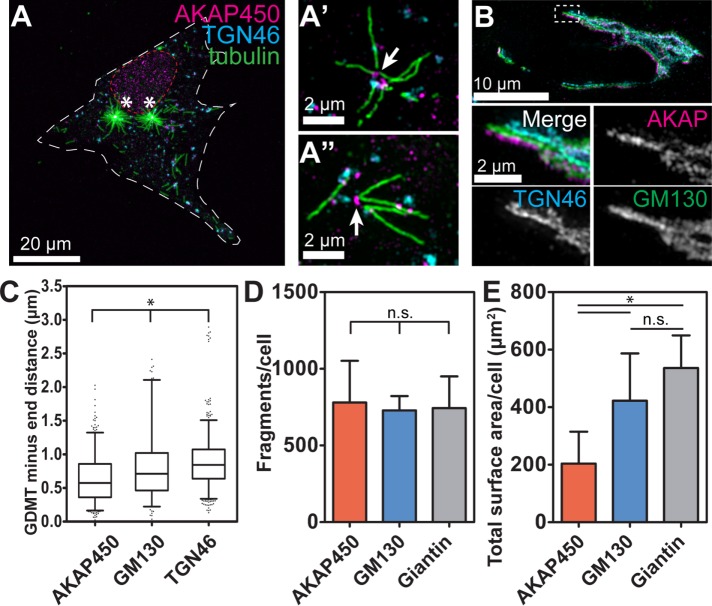
AKAP450 is not sufficient for restriction of GDMT nucleation to hotspots. (A–A″) Localization of GDMT nucleation sites and AKAP450 in an MRC-5 cell 40 s after nocodazole washout. (A) A laser scanning confocal microscopy overview image (maximum-intensity Z-projection). Immunostaining. Tubulin, green. TGN46, cyan. AKAP450, magenta. (A′, A″) SIM image maximum intensity projections. Tubulin, green. TGN46, cyan. AKAP450, magenta. Note that GDMT ends always colocalize with AKAP450. (B) MRC-5 cell expressing GM130-GFP (green) and immunostained for AKAP450 (magenta) and TGN46 (cyan). Note that AKAP450 localizes to a subdomain of the Golgi. SIM, maximum-intensity Z-projection. (C) GDMT minus ends are concentrated closer to AKAP450 positive structures than to GM130-positive or TGN46-positive structures in MRC-5 cells. Distance between GDMT minus ends and centers of mass of AKAP450, GM130, and TGN46 was measured and plotted in 5-95% boxplot. (p < 0.001, one-way ANOVA and Tukey’s multiple comparison test; n = 8–16 cells for each protein.) (D) Total numbers of Golgi fragments positive for AKAP450, GM130, or Giantin are comparable following nocodazole washout in MRC-5 cells. (p = 0.87, one-way ANOVA, n = 8 cells for each protein.) (E) Surface area of AKAP450-positive structures following nocodazole washout in MRC-5 cells is significantly smaller than that of GM130- or Giantin-positive Golgi structures. (p < 0.001 one-way ANOVA, with Tukey’s multiple comparison test showing significant differences between AKAP450- and GM130- or Giantin-positive structures, but not between GM130- and Giantin-positive structures [p = 0.13]; n = 8 cells for each protein.)
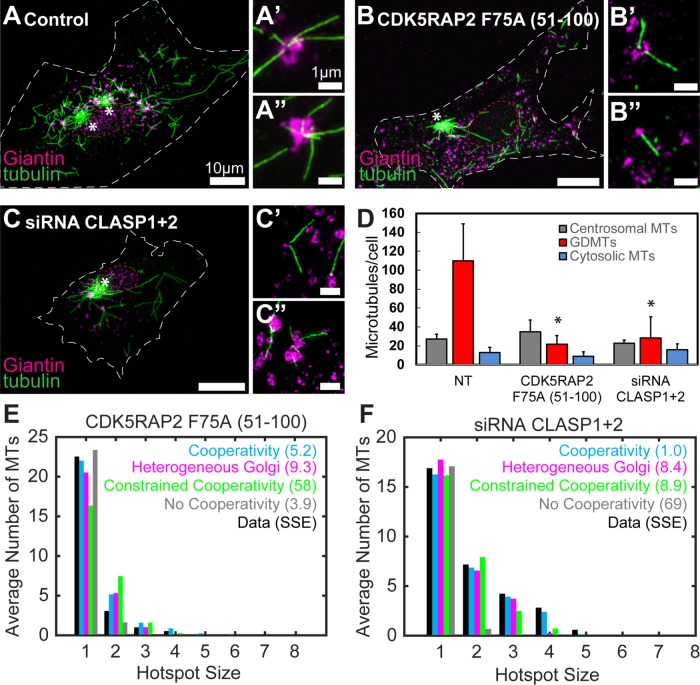
Besides AKAP450, the best evidence for protein involvement in noncentrosomal MT nucleation points to γ-TuRC activators, such as CDK5RAP2, and MT stabilizer CLASPs. Thus, we next assessed whether these proteins contribute to GDMT nucleation hotspots. First, we examined the role of γ-TuNA-dependent γ-TuRC activation (Fong et al., 2008; Choi et al., 2010). Interestingly, active γ-TuNA expression causes MT nucleation throughout the cytoplasm (Choi et al., 2010), indicating that local activation of γ-TuRC, rather than its local accumulation, defines noncentrosomal MT nucleation sites. Expression of a CDK5RAP2 fragment containing a functionally dead γ-TuNA domain (aa51-100 F75A; Choi et al., 2010) was shown to suppress MT nucleation by a dominant negative mechanism. We find that expression of the dead γ-TuNA does not suppress centrosomal MT levels (Figure 5, A, B, and D), possibly because the centrosomal machinery includes other redundant mechanisms to activate γ-TuRC, or is protected from the penetration of this polypeptide. At the same time, the dead γ-TuNA strongly reduces noncentrosomal MT nucleation (Figure 5, A, B, and D). This reduction is due to loss of GDMTs rather than of other noncentrosomal MTs (Figure 5D). Moreover, the decrease of GDMT levels resembles the reported effect of double knockout of CDK5RAP2 and MMG8, rather than of single CDK5RAP2 knockout, probably because this dominant-negative effect extends to at least these two γ-TuNA-containing proteins. Importantly, our computational simulation indicates that the pattern of remaining GDMTs fits random distribution of nucleation sites. The no cooperativity, cooperativity, and heterogeneous Golgi models provide roughly equivalent fits to the data (Figure 5E). Thus, neither cooperativity nor Golgi heterogeneity mechanisms are needed to account for the resulting data pattern; rather, this pattern is indicative of random nucleation. This indicates that besides decrease of MT nucleation per se, inhibition of γ-TuNA activity suppresses clustered nucleation. Accordingly, we conclude that MT nucleation hotspots at the Golgi require local activation of γ-TuRCs by γ-TuNA.
FIGURE 5:
γ-TuNA inhibition, but not depletion of CLASPs, leads to loss of GDMT nucleation hotspots. (A–C) Control (A), expressing CDK5RAP2 F75A (51–100, B), or CLASP1+2-depleted (C) MRC-5 cells 40 s after nocodazole washout. Immunofluorescence. (A, B, C) Laser scanning confocal microscopy overview images (maximum intensity Z-projection). Immunostaining. Tubulin, green. Giantin, magenta. (A′, A″, B′, B″, C′, C″) SIM image maximum intensity projections. Tubulin, green. Giantin, magenta. Note decrease of GDMTs in B and C as compared with A. (D) GDMT levels, but not centrosomal or cytosolic MTs, were significantly reduced in MRC-5 cells expressing CDK5RAP2 F75A (51–100) or CLASP1+2 depleted. (p < 0.001, one-way ANOVA, with Tukey’s multiple comparison test showing significant differences between nontreated and CDK5RAP2 F75A-expressing cells or CLASP1+2–depleted cells, but not between CDK5RAP2 F75A-expressing cells and CLASP1+2–depleted cells [p = 0.32], n = 8–16 cells per condition.) (E) Comparison of best-fit results for each model against data (averaged over 15 cells) from MRC-5 cells expressing CDK5RAP2 F75A (51–100). (F) Comparison of best-fit results for each model against data (averaged over 16 cells) from MRC-5 cells depleted of CLASP1+2.
Next, we investigated the role of CLASPs in GDMT hotspot formation. Confirming previous work from our lab and others (Efimov et al., 2007; Miller et al., 2009; Wu et al., 2016), we find that CLASPs are essential for GDMT but not centrosomal MT nucleation. Interestingly, while depletion of CLASP1 and CLASP2 resulted in severe reduction of GDMT levels (Figure 5, C and E), hotspots were still observed (Figure 5C). Computational modeling showed that both cooperativity and heterogeneous Golgi models still similarly matched the data (Figure 5F). The best-fit parameters for the heterogeneous Golgi model show a roughly threefold reduction in the base nucleation rate when compared with controls (Supplemental Figure 2C). Similarly, in the cooperativity model, all coefficients describing the nucleation rates (k) are significantly lower than in controls (Supplemental Figure 2D). Thus, in contrast to γ-TuNA inhibition, CLASPs depletion influences the overall nucleation rate rather than the hotspot nucleation rate specifically. This suggests that the mechanism underlying hotspot formation is not abolished when overall noncentrosomal MT nucleation rates are low (for example, in the absence of CLASPs), but many of them do not produce MTs under these conditions.
Taking these results together, this work identifies a nonrandom distribution pattern of newly formed GDMTs that is conserved in different cell lineages and that exists both in the steady state and upon MT regrowth after full MT network depolymerization. GDMT nucleation hotspots could be formed through Golgi membrane heterogeneity or a saturating cooperativity mechanism of MT nucleation. We cautiously favor the cooperativity model, supported by the hotspot GDMT length distribution.
We propose that due to low concentration and/or inactivity of nucleation and stabilization factors outside the pericentrosomal region, it is beneficial to concentrate and/or activate them at specific Golgi subdomains for efficient MT nucleation. This is manifested through clustered GDMT formation at those nucleation hotspots. Our data indicate that γ-TuNA-containing proteins likely act as such factors, either concentrated or locally activated at the hotspots to define their MT-nucleating capacity. The cooperative mode of MT nucleation at hotspots might mean that γ-TuNA-containing proteins or their activating factors are concentrated at the Golgi membrane in a MT- or a MT motor–dependent manner. While our data indicate that γ-TuNA-containing proteins are essential for hotspot formation, it is still unclear whether additional factors restrict GDMT nucleation to these precise loci. For example, AKAP450-dependent recruitment of γ-TuNA-containing proteins to the Golgi could be locally enhanced by an as yet unidentified mechanism. Alternatively, specific accumulation of one of the less characterized factors implicated in GDMT nucleation and stabilization (e.g., tubulin cofactor TBCE, Bellouze et al., 2014, or MTCL1, Sato et al., 2014) are involved in hotspot formation. It is clear from our data that MT nucleation hotspots are very dynamic, but whether the short lifetimes of hotspots are due to the saturation or dissolution of active molecular complexes is as yet an open question.
Functionally, clustered MT do not appear to be any different from nonclustered, because they are distributed equally asymmetrically and directed toward the cell front with the same degree of directionality. Thus, it is likely that single GDMTs are produced at the same hotspot sites with high potential of nucleation where only one MT succeeds in forming for stochastic reasons. Overall, our data suggest that nonrandom nucleation of GDMTs at hotspots with high nucleating capacity underlies intrinsic asymmetry of GDMT arrays in motile cells, and might contribute to other nonrandom features of MT network geometry in a variety of cell types.
MATERIALS AND METHODS
Cell culture, transfections, and depletions
RPE1 (Clontech), MRC-5 (American Type Culture Collection), and HPAEC (Lonza) cells were maintained in DMEM/F12 with 10% FBS, MEM with 10% FBS, and EGM-2 (Lonza) media, respectively, and grown at 37°C with 5% CO2. RPE1 cells were transfected using Nucleofection (Amaxa). MRC-5 cells were transfected using Nucleofection (Amaxa) for GM130-GFP expression, Fugene6 (Promega, Madison, WI) for γ-TuNA activity inhibition and live-cell experiments, or Transit-X2 (MirusBio, Madison, WI) for siRNA-mediated depletions.
DNA and siRNA constructs
Constructs used for live cell imaging: EB3-GFP (kind gift from J. V. Small, Vienna), mCherry-GalT (modified from Clontech; Efimov et al., 2007), EB3-mCherry (Efimov et al., 2008), GTN-GFP (kind gift from A. D. Lindstedt, Carnegie Mellon University), Emerald-EB3 (kind gift from M. Davidson; Addgene plasmid # 54076), and TGN-RFP (Deora et al., 2007). GM130-GFP (kind gift from C. Sütterlin, University of California, Irvine) was used in immunofluorescence experiments. To inhibit γ-TuNA activity, cells were transfected with pCMV2-FLAG-CDK5RAP2 F75A (51-100) (kind gift from R. Qi, Hong Kong University of Science and Technology; Choi et al., 2010). siRNA constructs (Dharmacon) used to deplete CLASPs 1 and 2 and nontargeting siRNA constructs were previously described (CLASP1: 5′-GGATGATTTACAAGACTGG-3′; CLASP2: 5′-GACATACATGGGTCTTAGA-3′; Efimov et al., 2007). RPE1 and MRC-5 cells were transfected 24 h prior to experiments. MRC-5 cells were treated with siRNA for 72 h to ensure sufficient depletion (Efimov et al., 2007; Miller et al., 2009). Depletion efficiency was determined by Western blotting (Supplemental Figure S1I). Cells expressing the CDK5RAP2 fragments were identified by FLAG immunostaining (Supplemental Figure S1F).
Reagents and antibodies
Nocodazole (Sigma-Aldrich, St. Louis, MO, M1404) was prepared at 16.6 mM in DMSO (cell culture grade). A concentration of 8.3 µM was used to depolymerize MTs. Antibodies used for immunofluorescence experiments were mouse anti-α-tubulin (1:1000, DM1A; Sigma-Aldrich, St. Louis, MO; T6199), mouse anti-GM130 (1:300; BD Transduction Laboratories, Franklin Lake, NJ; 610823), mouse anti-AKAP450 (1:250; kind gift from J. Goldenring; and BD Transduction Laboratories, Franklin Lake, NJ, 611518), mouse anti-FLAG (1:250; Sigma, St. Louis, MO; M2), rabbit anti-Giantin (1:1000; Abcam, Cambridge, MA; ab24586), rabbit anti-β-tubulin (1:1000, Abcam; Cambridge, MA; ab18251), sheep anti-TGN46 (1:5000; AbD Serotec; now BioRad, Hercules, CA; AHP500G). Alexa488-, Alexa568-, and Alexa647-conjugated highly cross-absorbed secondary antibodies were from Invitrogen (Grand Island, NY), and CF405M-conjugated highly cross-absorbed secondary antibodies were from Biotium (Fremont, CA). Coverslips were mounted in Vectashield Mounting Medium (Vector Labs, Burlingame, CA). Primary and secondary antibodies used for Western blot were mouse-anti-actin (1:5000; ThermoScientific; Fremont, CA; MS-1295), rabbit-anti-CLASP1 (1:1000; Abcam, Cambridge, MA; 108620), rabbit-anti-CLASP2 (1:1000; VU-83; Efimov et al., 2007), anti-mouse-IR680 and anti-rabbit-IR800 (1:10,000, Rockland, Gilbertsville, PA).
Live cell microscopy
Cells were cultured on MatTek dishes coated with 10 µg/µl fibronectin and transfected 24 h before experiment. For live-cell imaging of steady state MT dynamics, cells were transfected with Emerald-EB3 and TGN-RFP (RPE1) or EB3-mCherry and GTN-GFP (MRC-5) and imaged using a Nikon TE2000 inverted microscope equipped with 488- and 568-nm lasers, a Yokogawa CSU-10 spinning disk head, a PLAN APO VC 100x NA1.4 oil lens, intermediate magnification 1.5×, and EM-CCD camera (Andor Technology), controlled by Nikon Elements software. Image stacks with 0.3 μm between slices were recorded covering the entire Golgi (over 3-4 μm) for 5 min with maximal frame rate (∼4–5 s per stack). For live-cell imaging of nocodazole washouts, single-plane images with 5-s intervals were taken using the spinning disk confocal microscope setup described in (Efimov et al., 2007).
Nocodazole washout assay
Cells were cultured on coverslips (No. 1.5, 12 mm circular) coated in 10 µg/µl fibronectin for 24 h before experiments. Cells were incubated with 8.3 µM nocodazole full media for 2 h at 37°C with 5% CO2 to completely disassemble MTs (Supplemental Figure S1, A–G). To remove nocodazole, cells were placed on ice and washed eight times with ice-cold media. Then coverslips were placed in prewarmed media for 40 s, followed by 10 s in prewarmed preextraction in extraction buffer (60 mM PIPES, 25 mM HEPES, 10 mM EGTA, 2 mM MgCl2, 0.1% Saponin, pH 6.9, supplemented with 0.25 nM nocodazole, 0.25 nM taxol), before fixation in 4% paraformaldehyde, 0.025% glutaraldehyde in cytoskeleton buffer (150 mM NaCl, 5 mM MgCl2, 5 mM EGTA, 5 mM glucose, 10 mM MES) for 10 min at room temperature. Autofluorescence was quenched with 0.2% NaBH4 in PBS, followed by permeabilization in 0.25% Triton X-100 in PBS and blocking in 1% bovine serum albumin (Fisher) and 5% donor horse serum (Atlanta Biologicals) in PBS. Live-cell imaging microtubule regrowth assays were performed on the microscope stage by washing out nocodazole with ice-cold media.
Immunofluorescence microscopy of fixed cells
Whole cell images were acquired using a laser scanning confocal microscope: Nikon A1r with advanced photo-kinetic capacity, based on a TiE Motorized Inverted Microscope with 20×, 40×, 60×, and 100× lenses and DIC module, equipped with a high-speed resonant scanner (30 fps), run by NIS Elements C software. For quantitative analysis, images were acquired using either DeltaVision OMX (GE technology) or N-SIM (Nikon) systems. DeltaVision OMX used 405-, 488-, 568-, and 642-nm lasers, a 60× Plan-Apo NA1.42 oil lens, and a sCMOS camera. Nikon N-SIM used 405-, 488-, 561-, and 647-nm lasers, an SR Apo TIRF 100× NA1.49 oil lens, and an EMCCD camera (DU-987, Andor Technology).
Western blotting
Cells were cultured on 60-mm circular tissue culture dishes (Sarstedt) with fibronectin-coated coverslips (five per dish maximum). Cells were siRNA depleted for 72 h prior to MT regrowth assay (coverslips; see above) or cell harvest for Western blot. Cells were lysed using CHAPS lysis buffer (1% CHAPS, 30 mM Tris, pH 8.5, 30 mM NaCl, 1 mM Na3VO4) supplemented with protease and phosphatase inhibitors (Boston BioProducts, Ashland, MA; PI-215 and PI-270). Nuclei were removed by centrifugation (13,200 × g at 4°C, 15 min), and protein concentration was measured by Bradford assay (Pierce Coomassie Plus Assay kit; ThermoScientific, Fremont, CA; 23236). A sample of 25bµg protein per condition was resolved on 8% SDS–PAGE gels and transferred to nitrocellulose membranes (GE Healthcare Life Sciences, Pittsburgh, PA; 10600002) overnight at 4°C. Prestained SDS–PAGE standards were used as a marker. Membranes were blocked with 5% nonfat powdered milk (Boston BioProducts, Ashland, MA; P-1400) in Tris-buffered saline and probed for CLASP1, CLASP2, and actin (Supplemental Figure S1I). Membranes were imaged on a Licor Odyssey infrared imaging system.
Details of all quantitative analysis methods
GDMT nucleation sites positioning analysis in steady state.
Image stack sequences covering the entire Golgi during 3 min were used (Figure 1). The ImageJ plug-in MtrackJ was used to track newly formed Emerald-EB3 comets (growing MT plus-ends) emanating from the Golgi in 3D (Meijering et al., 2012). To quantify only newly nucleated MTs at the Golgi, tracks starting in the bottom or top slices of a stack or in the first time-frame were excluded. A custom MatLab script was generated to measure all possible distances between MT track origination sites and determine the closest site pairs (using the nearest-neighbor distance algorithm). The histogram of nearest-neighbor distances between GDMT track origination sites is shown in Figure 1C.
Analysis for 2D nucleation site distribution and MT directionality.
Centers of area were measured for the nucleus and the Golgi in maximum-intensity Z-projections (Figure 1, G and I), and the line connecting these centers was considered the nucleus–Golgi axis (Uetrecht and Bear 2009). Then directionality quadrants were generated by rotating the axis 45° in either direction. For Figure 1G, the resulting cross was centered on the center of the Golgi area, and nucleation sites were categorized into four quadrants. For Figure 1I, the cross was moved to each nucleation site to determine MT directionality, and MT were categorized into four direction groups relative to the cell polarity.
GDMT nucleation sites positioning analysis in live-cell MT regrowth.
Single plane time-lapse movies at 5 s intervals were used. ImageJ plugin MtrackJ was used to track newly formed EB3-GFP labeled MTs (Figure 2B). Golgi fragment size in 2D was estimated using Ferret’s diameter, resulting in center of mass coordinates and radius. A custom MatLab script was generated to measure all possible distances between MTs originating from the same Golgi fragment determine the closest site pairs (using the nearest-neighbor distance algorithm). Paired random data points for nearest- neighbor distances were generated using a custom script in Matlab that accounts for Golgi fragment size.
Analysis of hotspot duration.
GDMT nucleation events were determined in both steady state and live-cell MT regrowth (see above). Duration of hotspots was determined by calculating the time between the first and last GDMT nucleation event within these hotspots (Figures 1F and 2, E and F).
GDMT nucleation site positioning analysis in fixed cell after nocodazole washout.
GDMT minus end positions were determined using line scans in 3D SIM image stacks (Figure 2, H– K). Golgi fragment size was determined by Ferret’s diameter (XY) and minimum and maximum range (Z), resulting in center of mass coordinates (XYZ) and the radius of the fragment.
A custom MatLab script was generated to measure all possible distances between MTs originating from the same Golgi fragment to determine the closest site pairs (similar to the above but in 3D). A custom script in Matlab generated paired random data points for nearest-neighbor distances, which accounts for Golgi fragment size.
Distance between MT minus ends and Golgi subdomains.
GDMT minus end coordinates and Golgi fragment centers of mass (determined as above) were used to calculate distances for each Golgi marker (Figure 4C).
The total number of Golgi fragments and total Golgi surface area per cell (Figure 4, D and E) were measured using the Surface mapping function in Imaris (Bitplane).
MT number per cell (Figure 5D) was counted in 3D renderings of image stacks using Imaris.
Statistics
All quantitative data were collected from experiments performed in at least duplicate (Figure 2K) or triplicate (all other work). Categorical data (Figure 1, G and I) were analyzed using a χ2 test. Other data sets were analyzed using a one-way ANOVA F-test, Tukey’s multiple comparison test, and/or Student’s t test, as described in the figure legends.
Image processing
Image analysis was performed using ImageJ, Matlab, and Imaris. For all fluorescence images presented here, adjustments were made to brightness, contrast, and gamma settings to make small structures visible.
Specific image panels were processed as follows:
Figure 1, A, A′, and D. Maximum intensity projections over time and Z are shown of 3D time lapse movies.
Figure 1, B and E. Maximum intensity projections over Z are shown of 3D timelapse movies.
Figure 1, G and I. Maximum intensity projections over time and Z of TGN-RFP are shown.
Figure 2, A, C, and D. Single slices are shown of time-lapse movies.
Figures 2, G– J, 3C, 4, A and B, and 5, A– C. Maximum-intensity Z projections are shown from laser scanning confocal microscope (2E, 4A, 5A–5C) or SIM (2F–2H, 3C, 3C′, 4A′. 4A″, B, 5A′, 5A″, 5B′, 5B″, 5C′, 5C″) images.
COMPUTATIONAL MODEL DESCRIPTIONS
To compare the different hypotheses described in the text, we fitted four spatial, stochastic, and dynamic models of MT nucleation (see Figure 3A) to imaging data. These models are spatial, in the sense that MTs are associated with a nucleation location on the Golgi, a requirement for investigating the spatial phenomenon of hotspot formation. They are dynamic in that we track MT nucleation across a 40 s time window, mimicking the nucleation time frame between Nocodazole washout and fixation in experiments. Temporal dynamics is required to account for potential effects of cooperativity. Finally, they are stochastic in the sense that nucleation events are probabilistic in space and in time.
To describe each model, we specify the spatial geometry of the Golgi (which is the same for all models) and the assumptions about how microtubules (MTs) nucleate (which is specific to each model). For simplicity, we consider the in silico Golgi geometry to be a sphere. The radius of this sphere is chosen so that its surface area (SA) is the average (over all cells imaged) SA calculated from super-resolution images of the Golgi. The four models considered differ in their MT nucleation properties.
No cooperativity model
This model assumes that all MTs are nucleated randomly. It is described by a single rate parameter (k0), describing the rate of microtubule nucleation in units of #/(μm2 s). In this model, hotspots only form due to coincidental nucleation of MTs close to each other (within 0.4 µm).
Constrained cooperativity model
Here it is assumed that MTs can nucleate in one of two ways: 1) random nucleation anywhere on the membrane with rate parameter k0 (units of #/[μm2 s]) or 2) nucleation of a new MT from an existing hotspot with constrained rate parameter k1 (units of #/[hotspot s]). Because the largest hotspot size observed in data contained eight MTs, we assume the maximal number of MTs that can be contained in a hotspot is eight, after which hotspots no longer nucleate new MTs. This model is fully described by two parameters, k0 and k1.
Cooperativity model
This is an extension of the constrained cooperativity model where it is assumed that hotspots of different size can nucleate new MTs at different rates (k1–k4). Given the small numbers of hotspots of size larger than 4 in the data and to simplify this model, we assume that nucleation of hotspots of size 4–7 is the same. This model is fully described by five parameters, k0–k1.
Heterogeneous Golgi model
This model is fundamentally distinct from the cooperativity models. Rather than assuming that hotspots form due to feedback, we assume that focal locations on the Golgi have a higher propensity to nucleate new MTs than the remainder of the membrane. For this model, we assume there are two types of Golgi: low-nucleation-rate membrane and high-nucleation-rate membrane. In this model, all nucleation is random in time and space. However, there are now two nucleation rates associated with Golgi regions of different propensities. The geometric structure of these membrane regions could in principle take any form. For simplicity, we assume the high nucleation rate region comprises N small focal circles 40 nm in diameter, while the remainder of the membrane exhibits the low nucleation rate. The nucleation rate outside of the circles is k0 (units of #/[μm2 s]) and the nucleation rate in the focal regions is increased by a multiplicative factor f (fold increase, f > 1, no units). Thus, the nucleation rate within these focal regions is fk0. This model is fully described by three parameters, k0, f, and N. Parameter estimates (see Supplemental Materials and Methods for parameter estimates and sensitivity analysis) of the heterogeneous Golgi model predict that ∼50 regions of increased nucleation, comprising <1% of total Golgi surface area, would be responsible for these hotspots, and that the MT nucleation capacity there must be significantly higher (∼300 times) than for the rest of the Golgi membrane.
Fitting methodologies and length distribution comparisons
To compare each of these models against data, we perform a broad search over the parameter spaces of the models to determine the parameter set(s) that best fit the hotspot distribution data. For this comparison, in the data we calculate an average frequency distribution quantifying the average number of microtubules per cell associated with hotspots of different sizes. Results of every model simulation performed are quantified similarly: The resulting MTs are classified as hotspots of size 1–8 (based on the same 40-nm proximity condition as in the data) and a frequency distribution is calculated. To determine how well that model/parameter set combination accounts for data, the sum of squared errors (SSE) between the data and simulated frequency distributions is computed. This SSE statistic is reported in Figures 3B and 5, E and F.
To determine the best-fit parameter set for each model, a standard log spaced grid search was performed for the “no cooperativity” and “constrained cooperativity” models with 100 (in one dimension) and 10,000 (in two dimensions) parameters, respectively. Due to the increased number of parameters, in the remaining models, Latin hypercube sampling (in log coordinates for each parameter) with 100,000 parameters was performed. In all cases, the best-fit parameter set is chosen according to the minimal SSE and those results are reported in the main text. See the Supplemental Material for analysis of further details and sensitivity of results with respect to parameters.
To further distinguish between the length of the “cooperativity” and “heterogeneous Golgi” models, we simulated the length distributions from these models and compared them with observed MT length distributions from fixed images (Figure 3). To generate model length distributions, the best-fit parameters for each model from model fits were considered and the model was rerun with the nucleation time of each MT stored. MTs were assumed to grow at 10 μm per minute based on time lapse imaging of these cells and were assumed to grow consistently with this fixed velocity after nucleation (no dynamic instability). Because this growth rate would be expected to generate lengths considerably greater than those observed, and because it was found that nucleation is likely suppressed until some time after the washout due to temperature effects, we instead modeled length distributions based on a 20-s window of nucleation and growth instead of 40-s (the time between washout and fixation).
Supplementary Material
Acknowledgments
This work was supported by American Heart Association fellowship 17POST32650000 (to A.A.W.M.S.), National Institutes of Health (NIH) grants R01-GM078373 and R01-DK106228 (to I.K.), an award from NIH program grant UL1TR000445 (to A.A.W.M.S.), and National Science Foundation grant DMS1562078 (to W.R.H). We utilized the Vanderbilt Cell Imaging Shared Resource (funded by NIH grants CA68485, DK20593, DK58404, DK59637, EY08126, NIH SEI10, 1sS10, 1S10OD018075, and S10OD012324) and the core(s) of the Vanderbilt Diabetes Research and Training Center (funded by NIH grant DK020593). We thank Brian S. Domin for technical help.
Abbreviations used:
- GDMT
Golgi-derived microtubule
- MT
microtubule
- SIM
structured illumination microscopy
- γ-TuNA
γ-tubulin nucleation activator
- γ-TuRC
γ-tubulin ring complex.
Footnotes
This article was published online ahead of print in MBoC in Press (http://www.molbiolcell.org/cgi/doi/10.1091/mbc.E17-06-0425) on September 20, 2017.
REFERENCES
- Alfaro-Aco R, Thawani A, Petry S. Structural analysis of the role of TPX2 in branching microtubule nucleation. J Cell Biol. 2017;216:983–997. doi: 10.1083/jcb.201607060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bauer M, Cubizolles F, Schmidt A, Nigg EA. Quantitative analysis of human centrosome architecture by targeted proteomics and fluorescence imaging. EMBO J. 2016;35:2152–2166. doi: 10.15252/embj.201694462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bellouze S, Schäfer MK, Buttigieg D, Baillat G, Rabouille C, Haase G. Golgi fragmentation in pmn mice is due to a defective ARF1/TBCE cross-talk that coordinates COPI vesicle formation and tubulin polymerization. Hum Mol Genet. 2014;23:5961–5975. doi: 10.1093/hmg/ddu320. [DOI] [PubMed] [Google Scholar]
- Chabin-Brion K, Marceiller J, Perez F, Settegrana C, Drechou A, Durand G, Poüs C. The Golgi complex is a microtubule-organizing organelle. Mol Biol Cell. 2001;12:2047–2060. doi: 10.1091/mbc.12.7.2047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi Y-KK, Liu P, Sze SK, Dai C, Qi RZ. CDK5RAP2 stimulates microtubule nucleation by the gamma-tubulin ring complex. J Cell Biol. 2010;191:1089–1095. doi: 10.1083/jcb.201007030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deora AA, Diaz F, Schreiner R, Rodriguez-Boulan E. Efficient electroporation of DNA and protein into confluent and differentiated epithelial cells in culture. Traffic. 2007;8:1304–1312. doi: 10.1111/j.1600-0854.2007.00617.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dyachuk V, Bierkamp C, Merdes A. Non-centrosomal microtubule organization in differentiated cells. In: Lüders J, editor. The Microtubule Cytoskeleton: Organisation, Function and Role in Disease. Vienna: Springer; 2016. pp. 27–41. [Google Scholar]
- Efimov A, Kharitonov A, Efimova N, Loncarek J, Miller PM, Andreyeva N, Gleeson P, Galjart N, Maia A, McLeod IX, et al. Asymmetric CLASP-dependent nucleation of noncentrosomal microtubules at the trans-Golgi network. Dev Cell. 2007;12:917–930. doi: 10.1016/j.devcel.2007.04.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Efimov A, Schiefermeier N, Grigoriev I, Ohi R, Brown MC, Turner CE, Small JV, Kaverina I. Paxillin-dependent stimulation of microtubule catastrophes at focal adhesion sites. J Cell Sci. 2008;121:196–204. doi: 10.1242/jcs.012666. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fong K-WW, Choi Y-KK, Rattner JB, Qi RZ. CDK5RAP2 is a pericentriolar protein that functions in centrosomal attachment of the gamma-tubulin ring complex. Mol Biol Cell. 2008;19:115–125. doi: 10.1091/mbc.E07-04-0371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Galjart N. Plus-end-tracking proteins and their interactions at microtubule ends. Curr Biol. 2010;20:R528–R537. doi: 10.1016/j.cub.2010.05.022. [DOI] [PubMed] [Google Scholar]
- Hurtado L, Caballero C, Gavilan MP, Cardenas J, Bornens M, Rios RM. Disconnecting the Golgi ribbon from the centrosome prevents directional cell migration and ciliogenesis. J Cell Biol. 2011;193:917–933. doi: 10.1083/jcb.201011014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jia Y, Fong K-WW, Choi Y-KK, See S-SS, Qi RZ. Dynamic recruitment of CDK5RAP2 to centrosomes requires its association with dynein. PLoS One. 2013;8 doi: 10.1371/journal.pone.0068523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kollman JM, Greenberg CH, Li S, Moritz M, Zelter A, Fong KK, Fernandez J-JJ, Sali A, Kilmartin J, Davis TN, Agard DA. Ring closure activates yeast γTuRC for species-specific microtubule nucleation. Nat Struct Mol Biol. 2015;22:132–137. doi: 10.1038/nsmb.2953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maia A, Zhu X, Miller P, Gu G, Maiato H, Kaverina I. Modulation of Golgi-associated microtubule nucleation throughout the cell cycle. Cytoskeleton. 2013;70:32–43. doi: 10.1002/cm.21079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meijering E, Dzyubachyk O, Smal I. Methods for cell and particle tracking. Methods Enzymol. 2012;504:183–200. doi: 10.1016/B978-0-12-391857-4.00009-4. [DOI] [PubMed] [Google Scholar]
- Miller PM, Folkmann AW, Maia ARR, Efimova N, Efimov A, Kaverina I. Golgi-derived CLASP-dependent microtubules control Golgi organization and polarized trafficking in motile cells. Nat Cell Biol. 2009;11:1069–1080. doi: 10.1038/ncb1920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moudjou M, Bordes N, Paintrand M, Bornens M. gamma-Tubulin in mammalian cells: the centrosomal and the cytosolic forms. J Cell Sci. 1996;109:875–887. doi: 10.1242/jcs.109.4.875. [DOI] [PubMed] [Google Scholar]
- O’Rourke BP, Gomez-Ferreria MA, Berk RH, Hackl AM, Nicholas MP, O’Rourke SC, Pelletier L, Sharp DJ. Cep192 controls the balance of centrosome and non-centrosomal microtubules during interphase. PLoS One. 2014;9 doi: 10.1371/journal.pone.0101001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oddoux S, Zaal KJ, Tate V, Kenea A, Nandkeolyar SA, Reid E, Liu W, Ralston E. Microtubules that form the stationary lattice of muscle fibers are dynamic and nucleated at Golgi elements. J Cell Biol. 2013;203:205–213. doi: 10.1083/jcb.201304063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ori-McKenney KM, Jan L, Jan Y-N. Golgi outposts shape dendrite morphology by functioning as sites of acentrosomal microtubule nucleation in neurons. Neuron. 2012;76:921–930. doi: 10.1016/j.neuron.2012.10.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ríos RM, Sanchís A, Tassin A, Fedriani C, Bornens M. GMAP-210 recruits gamma-tubulin complexes to cis-Golgi membranes and is required for Golgi ribbon formation. Cell. 2004;118:323–335. doi: 10.1016/j.cell.2004.07.012. [DOI] [PubMed] [Google Scholar]
- Rivero S, Cardenas J, Bornens M, Rios RM. Microtubule nucleation at the cis-side of the Golgi apparatus requires AKAP450 and GM130. EMBO J. 2009;28:1016–1028. doi: 10.1038/emboj.2009.47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roubin R, Acquaviva C, Chevrier V, Sedjaï F, Zyss D, Birnbaum D, Rosnet O. Myomegalin is necessary for the formation of centrosomal and Golgi-derived microtubules. Biol Open. 2013;2:238–250. doi: 10.1242/bio.20123392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salaycik KJ, Fagerstrom CJ, Murthy K, Tulu US, Wadsworth P. Quantification of microtubule nucleation, growth and dynamics in wound-edge cells. J Cell Sci. 2005;118:4113–4122. doi: 10.1242/jcs.02531. [DOI] [PubMed] [Google Scholar]
- Sanders AA, Kaverina I. Nucleation and dynamics of Golgi-derived microtubules. Front Neurosci. 2015;9:431. doi: 10.3389/fnins.2015.00431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sato Y, Hayashi K, Amano Y, Takahashi M, Yonemura S, Hayashi I, Hirose H, Ohno S, Suzuki A. MTCL1 crosslinks and stabilizes non-centrosomal microtubules on the Golgi membrane. Nat Commun. 2014;5:5266. doi: 10.1038/ncomms6266. [DOI] [PubMed] [Google Scholar]
- Shanks RA, Steadman BT, Schmidt PH, Goldenring JR. AKAP350 at the Golgi apparatus. I. Identification of a distinct Golgi apparatus targeting motif in AKAP350. J Biol Chem. 2002;277:40967–40972. doi: 10.1074/jbc.M203307200. [DOI] [PubMed] [Google Scholar]
- Takahashi M, Shibata H, Shimakawa M, Miyamoto M, Mukai H, Ono Y. Characterization of a novel giant scaffolding protein, CG-NAP, that anchors multiple signaling enzymes to centrosome and the Golgi apparatus. J Biol Chem. 1999;274:17267–17274. doi: 10.1074/jbc.274.24.17267. [DOI] [PubMed] [Google Scholar]
- Takahashi M, Yamagiwa A, Nishimura T, Mukai H, Ono Y. Centrosomal proteins CG-NAP and kendrin provide microtubule nucleation sites by anchoring gamma-tubulin ring complex. Mol Biol Cell. 2002;13:3235–3245. doi: 10.1091/mbc.E02-02-0112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uetrecht AC, Bear JE. Golgi polarity does not correlate with speed or persistence of freely migrating fibroblasts. Eur J Cell Biol. 2009;88:711–717. doi: 10.1016/j.ejcb.2009.08.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vinogradova T, Paul R, Grimaldi AD, Loncarek J, Miller PM, Yampolsky D, Magidson V, Khodjakov A, Mogilner A, Kaverina I. Concerted effort of centrosomal and Golgi-derived microtubules is required for proper Golgi complex assembly but not for maintenance. Mol Biol Cell. 2012;23:820–833. doi: 10.1091/mbc.E11-06-0550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Z, Wu T, Shi L, Zhang L, Zheng W, Qu JY, Niu R, Qi RZ. Conserved motif of CDK5RAP2 mediates its localization to centrosomes and the Golgi complex. J Biol Chem. 2010;285:22658–22665. doi: 10.1074/jbc.M110.105965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Z, Zhang C, Qi RZ. A newly identified myomegalin isoform functions in Golgi microtubule organization and ER-Golgi transport. J Cell Sci. 2014;127:4904–4917. doi: 10.1242/jcs.155408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu J, de Heus C, Liu Q, Bouchet BP, Noordstra I, Jiang K, Hua S, Martin M, Yang C, Grigoriev I, et al. Molecular pathway of microtubule organization at the Golgi apparatus. Dev Cell. 2016;39:44–60. doi: 10.1016/j.devcel.2016.08.009. [DOI] [PubMed] [Google Scholar]
- Yalgin C, Ebrahimi S, Delandre C, Yoong LF, Akimoto S, Tran H, Amikura R, Spokony R, Torben-Nielsen B, White KP, Moore AW. Centrosomin represses dendrite branching by orienting microtubule nucleation. Nat Neurosci. 2015;18:1437–1445. doi: 10.1038/nn.4099. [DOI] [PubMed] [Google Scholar]
- Zaal KJ, Reid E, Mousavi K, Zhang T, Mehta A, Bugnard E, Sartorelli V, Ralston E. Who needs microtubules? Myogenic reorganization of MTOC, Golgi complex and ER exit sites persists despite lack of normal microtubule tracks. PLoS One. 2011;6:e29057. doi: 10.1371/journal.pone.0029057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu X, Hu R, Brissova M, Stein RW, Powers AC, Gu G, Kaverina I. Microtubules negatively regulate insulin secretion in pancreatic β cells. Dev Cell. 2015;34:656–668. doi: 10.1016/j.devcel.2015.08.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.