Abstract
Fresh-cut luffa (Luffa cylindrica) fruits commonly undergo browning. However, little is known about the molecular mechanisms regulating this process. We used the RNA-seq technique to analyze the transcriptomic changes occurring during the browning of fresh-cut fruits from luffa cultivar ‘Fusi-3’. Over 90 million high-quality reads were assembled into 58,073 Unigenes, and 60.86% of these were annotated based on sequences in four public databases. We detected 35,282 Unigenes with significant hits to sequences in the NCBInr database, and 24,427 Unigenes encoded proteins with sequences that were similar to those of known proteins in the Swiss-Prot database. Additionally, 20,546 and 13,021 Unigenes were similar to existing sequences in the Eukaryotic Orthologous Groups of proteins and Kyoto Encyclopedia of Genes and Genomes databases, respectively. Furthermore, 27,301 Unigenes were differentially expressed during the browning of fresh-cut luffa fruits (i.e., after 1–6 h). Moreover, 11 genes from five gene families (i.e., PPO, PAL, POD, CAT, and SOD) identified as potentially associated with enzymatic browning as well as four WRKY transcription factors were observed to be differentially regulated in fresh-cut luffa fruits. With the assistance of rapid amplification of cDNA ends technology, we obtained the full-length sequences of the 15 Unigenes. We also confirmed these Unigenes were expressed by quantitative real-time polymerase chain reaction analysis. This study provides a comprehensive transcriptome sequence resource, and may facilitate further studies aimed at identifying genes affecting luffa fruit browning for the exploitation of the underlying mechanism.
Introduction
Luffa cylindrica (i.e., luffa) of the family Cucurbitaceae is one of the most important vegetables and widely used medicinal plants in China. People are increasingly starting to pay more attention to diet, nutrition, and health. Because luffa is nutritious, delicious, and has beneficial effects on human health, the land area on which it is cultivated continues to increase [1–3]. There are currently two luffa species, namely towel gourd (Luffa cylindrica Roem.) and sinkwa towelsponge gourd (Luffa acutangula Roxb.). The former is affected by flesh browning, which considerably influences its flavor, odor, nutritional value, and shelf life [4,5]. Consequently, luffa browning has gradually become a prominent topic of interest among researchers. The mechanism regulating the browning of fruits and vegetables has been investigated for several decades, with differences detected in the enzymatic browning of diverse crops. Researchers have been unable to inhibit the enzymatic browning mechanism, and the luffa browning mechanism remains uncharacterized [6,7].
The browning of fruits and vegetables results from non-enzymatic and enzymatic processes [8,9]. Non-enzymatic browning, which involves many types of chemical processes, such as the Maillard reaction, caramelization, vitamin C oxidation, and polyphenol polymerization, is a major cause of browning [10,11]. Enzymatic browning in fresh-cut products is generally considered a multi-factorial process [12,13]. Polyphenol oxidase (EC 1.10.3.1, PPO) encoded by PPO multigene family members is a major inducer of the enzymatic browning of fruits, vegetables, and fresh horticultural products, following bruising, cutting, or other cellular damages [14,15]. Many other enzymes, including peroxidase (E1.11.1.7, POD), phenylalanine ammonia lyase (EC 4.3.1.24, PAL), superoxide dismutase (EC1.15.1.1, SOD), and catalase (EC 1.11.1.6, CAT), are also involved in the complex processes that induce or inhibit enzymatic browning of fresh-cut foods [16–18]. The expression levels of the genes encoding these enzymes likely change during the enzymatic browning of fresh-cut fruits and vegetables. Furthermore, the transcriptome profiles from recent studies revealed that the expression levels of genes associated with ethylene, abscisic acid, and gibberellin metabolism were affected by the browning of pears and apples [19,20]. However, little information is available concerning the mechanisms underlying the browning of fresh-cut luffa fruits and how to prevent it.
High-throughput transcriptome sequencing technology has recently been widely used to analyze the gene expression levels of whole organisms. The application of this technology enables the comprehensive study of gene expression during specific states or conditions [21,22]. Additionally, it can be used to identify important functional genes and characterize the molecular mechanisms regulating biological traits. In this study, we used high-throughput sequencing technology to examine gene expression profiles in fresh-cut luffa fruits. We annotated and classified the functions of Unigenes, and analyzed the browning-associated metabolic pathways. Our objective was to clarify the molecular basis of the browning of fresh-cut luffa fruits.
Materials and methods
Plant materials
Luffa ‘Fusi-3’ plants were grown in a greenhouse at the Vegetable Research Institute of the Fujian Academy of Agricultural Sciences. Undamaged fruits of a uniform size were collected at 18 days after pollination. Twenty representative luffa ‘Fusi-3’ fruits were harvested from each tree. The fruit samples were cut into 1-cm-thick slices with a stainless steel knife, and incubated at 25 ± 1°C for 1, 3, and 6 h to induce browning. Samples immediately after slicing were used as controls. The control and treated samples were collected at the same time points. The sliced samples were combined and immediately frozen in liquid nitrogen and peeled before being stored at −80°C.
RNA preparation, library construction, and RNA sequencing
Total RNA was extracted, mRNA was purified, and cDNA libraries were constructed by Guangzhou Gene Denovo Biological Technology Co., Ltd. (Guangzhou, China). The cDNA libraries were constructed as previously described [23] and then sequenced using the HiSeq™ 2500 system (Illumina Inc., San Diego, CA, USA).
The total RNA used for a quantitative real-time polymerase chain reaction (qRT-PCR) assay was extracted from each sample using the EZNA Plant RNA Kit (Bio-tek, Beijing, China). The quantity and quality of the RNA samples were assessed using the NanoDrop ND-1000 spectrophotometer (Thermo Scientific, CA, USA) and a 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). The libraries were then sequenced using the HiSeq™ 2000 system (Illumina Inc.). Equal amounts of RNA from three samples collected at the same browning time point were combined.
Determination of total phenol content
The total phenol content was determined with the Folin–Ciocalteu assay, using a modified version of the method described by Dewanto et al. (2002) [24]. The absorbance (750 nm) of a 25-μL aliquot of extract was recorded, and the total phenol concentration was expressed as gallic acid equivalents (mg GAE g−1 fresh weight) based on a calibration curve (1–6 μg mL−1 gallic acid). Each biological replicate was measured three times.
De novo transcriptome assembly and functional annotation
The RNA sequencing (RNA-seq) raw data were filtered by removing adapter sequences with in-house Perl scripts to obtain high-quality reads. Reads with low-quality regions (reads with a base quality < 20) were excluded using the sliding window trimming approach. The remaining reads were de novo assembled using Trinity software (http://trinityrnaseq.github.io/) [25], with min_kmer_cov set to 2 by default and all other parameters also set to their default values.
We used the luffa Unigene sequences to search the National Center for Biotechnology Information nonredundant (NCBInr) protein, Gene Ontology (GO), Eukaryotic Orthologous Groups of proteins (KOG), Kyoto Encyclopedia of Genes and Genomes (KEGG), and Swiss-Prot databases. Protein functional annotations were based on a BLASTX search (E < 10−5), which was used to identify the most similar proteins. The GO terms were assigned to Unigenes using Blast2GO [26]. The distribution of the GO functional classifications for the Unigenes was plotted using WEGO software [27]. We identified the genes potentially important for luffa browning using the aforementioned databases.
Analysis of Unigene expression levels
Reads from four cDNA libraries were mapped to the assembled Unigenes using Bowtie [28]. A fragments per kilobase of exon model per million mapped reads (FPKM) fold-change > 2 and a false discovery rate (FDR) < 0.05 were used as the thresholds to identify significant differences in gene expression [29]. Expression-level differences between two samples were examined using the DEGseq R package [30]. A fold-change ≥ 2 (in both directions; log2 ratio ≥ 1) and FDR ≤ 0.05 were used as standards to determine the significance of gene expression level differences among samples. A time series cluster analysis, which is based on the short time-series expression miner (STEM) method (http://www.cs.cmu.edu/~jernst/stem/) [31], was used to identify global trends and similar temporal model expression patterns among all differentially expressed genes (DEGs).
Quantitative real-time polymerase chain reaction analysis
The RNA-seq data were validated by qRT-PCR using gene-specific primers. Total RNA was extracted from fruits, and a 2-μg sample was used to prepare cDNA with the PrimeScript™ II 1st Strand cDNA Synthesis kit (Takara). The resulting cDNA was used as the template for qRT-PCR, which was conducted using a real-time PCR plate (Applied Biosystems, Foster City, CA, USA), SYBR green as the fluorescent reagent, and an ABI 7500 Fast Real-time PCR system (Applied Biosystems). A standard curve for each target gene was constructed based on qRT-PCR data from a series of diluted cDNA samples. The 20-μL qRT-PCR solutions consisted of 2 μl cDNA, 10 μL SYBR Premix Ex Taq™ II, 6.8 μL EASY Dilution buffer, and 0.6 μL 10 μM forward and reverse primers. The PCR was conducted with the following program: 95°C for 30 s; 40 cycles of 95°C for 10 s and 60°C for 34 s. The 18S rRNA gene (GenBank accession: KM656452), which was stably expressed according to the RNA-seq data, was used to normalize expression levels [32]. The 2−ΔΔCt method was used to calculate relative gene expression levels with Microsoft Excel software. The qRT-PCR experiment consisted of three independent biological replicates, and the expression levels calculated for each sample were based on three technical replicates.
Results
Determination of total phenol content
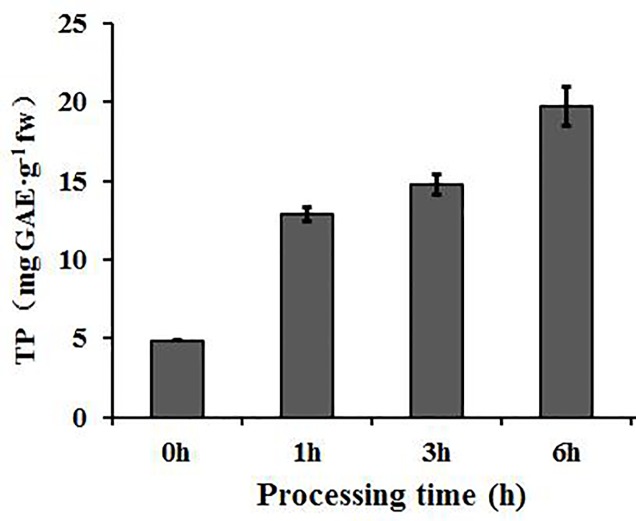
Previous studies indicated that among Luffa cylindrica cultivars, ‘Fusi-3’ produces fruits that are the most susceptible to browning [33]. Additionally, the total phenol content of browning fresh-cut vegetables under constant temperatures tends to increase over time [13,34]. For a comprehensive transcriptome analysis of luffa fruits, total RNA was extracted from peeled ‘Fusi-3’ fruit slices. The total phenol content exhibited an increasing trend during the browning treatment period (i.e., 0–6 h) (Fig 1).
Fig 1. Total phenol content in luffa fruit slices at different browning time points.
The vertical bars represent the standard error of triplicate experiments.
RNA sequencing and assembly of reads
To analyze the transcription profiles of luffa fruits undergoing browning, four cDNA libraries were created from ‘Fusi-3’ fruits at 18 days after pollination, and analyzed using the Illumina HiSeq™ 2000 system. We generated about 92 million sequence reads. After removing adapter sequences and low-quality reads, we obtained 91 million valid sequencing reads, with an average length of 90.2 bp. After a de novo assembly analysis, 58,073 Unigenes were obtained, with an average length of 896.93 bp and an N50 of 1,510 bp. Additionally, 29.86% of the Unigenes were > 1,000 bp long (Table 1).
Table 1. Summary of the transcriptome during the browning of fresh-cut luffa fruits.
| Sequences | 0h | 1h | 3h | 6h |
|---|---|---|---|---|
| BEFORE TRIMMING | ||||
| Total nucleotides (bp) | 2,998,625,250 | 2,760,038,000 | 3,010,284,000 | 2,732,343,750 |
| Number of raw reads | 23,989,002 | 22,080,304 | 24,082,272 | 21,858,750 |
| AFTER TREMMING | ||||
| Number of clean reads | 23,505,162 | 21,668,306 | 23,622,892 | 21,429,916 |
| GC content (%) | 45.81 | 46.29 | 46.08 | 47.16 |
| Q30 percentage (%) | 94.74 | 94.77 | 94.83 | 94.73 |
| AFTER ASSEMBLY | ||||
| Number of Unigenes of combined data | 58,073 | |||
| Total nucleotides (nt) of Unigenes (bp) | 52,087,451 | |||
| Mean length of Unigenes (bp) | 896.93 | |||
| Sequence length > 1000 bp (%) | 29.86 | |||
| N50 of Unigenes (bp) | 1510 | |||
Sequence annotation
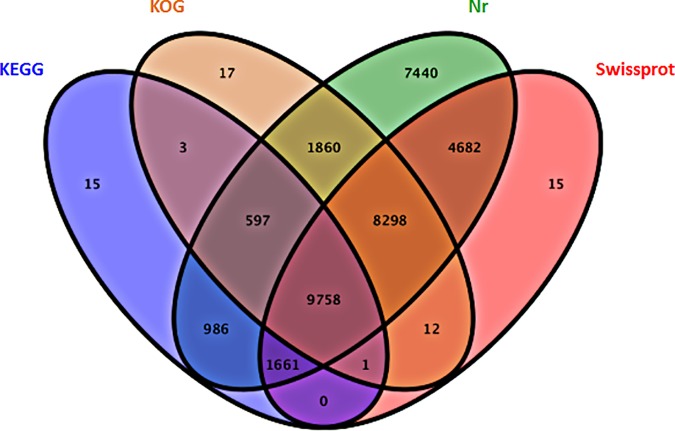
The annotation of Unigenes provided information regarding functions, protein sequence similarities, and KOG, GO, and KEGG pathway details. The data were screened against the following four public databases: NCBInr, Swiss-Prot, KOG, and KEGG. Unigenes were annotated according to the best BLASTX matches (E < 10−5) and sequence identities > 30%. A total of 35,345 (60.86%) Unigenes were annotated. The assembled sequences included 37,321 (60.75%), 24,427 (30.87%), 20,546 (30.52%), and 13,021 (18.83%) Unigenes with matches in the NCBInr, Swiss-Prot, KOG, and KEGG databases, respectively (S1 Table and Fig 2).
Fig 2. Unigenes annotated by a BLASTX search of databases (E < 10−5).
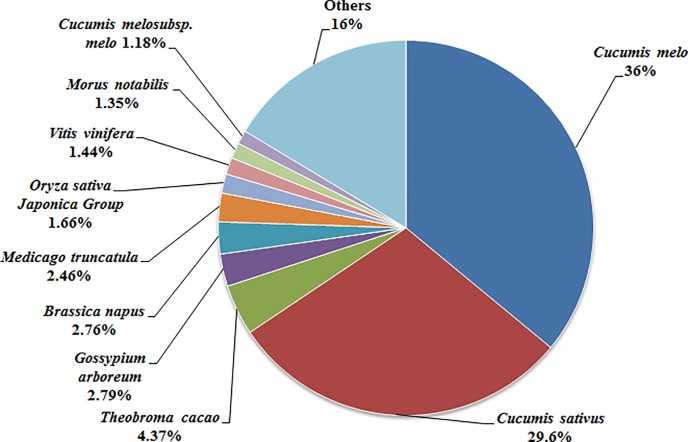
The most abundant BLAST hits were associated with plant species, and 35,282 (60.75%) of the Unigenes with matches in the NCBInr database had homologs in 10 plant species. The five species with the most hits were Cucumis melo [12,702 (36%)], Cucumis sativus [10,445 (29.6%)], Theobroma cacao [1,543 (4.37%)], Gossypium arboreum [983 (2.79%)], and Brassica napus [973 (2.76%)] (Fig 3). The fact that the luffa Unigenes had the most matches with C. melo sequences suggested that C. melo may be a useful reference material for further analyses of luffa functional genes. A total of 27,858 Unigenes were annotated by one or more databases (i.e., 78.81% of all assembled Unigenes), implying they have relatively well conserved functions.
Fig 3. Unigenes with matches in the NCBInr database and homologs in 10 plant species.
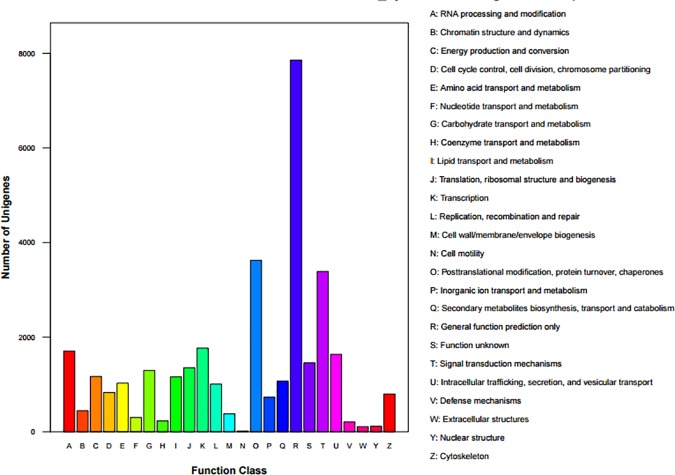
The luffa Unigenes were searched against the KOG database to predict and classify their possible functions. Of 37,321 NCBInr matches, 31,567 sequences had KOG classifications among 25 KOG categories (Fig 4). The most common categories were “general function prediction only”, with 7,857 Unigenes, and “posttranslational modification, protein turnover, and chaperones” (3,626 Unigenes). “Cell motility” had the fewest Unigenes, with only 17. Our data revealed that annotated Unigenes were associated with most of the activities related to growth and development.
Fig 4. Histogram of the KOG classifications.
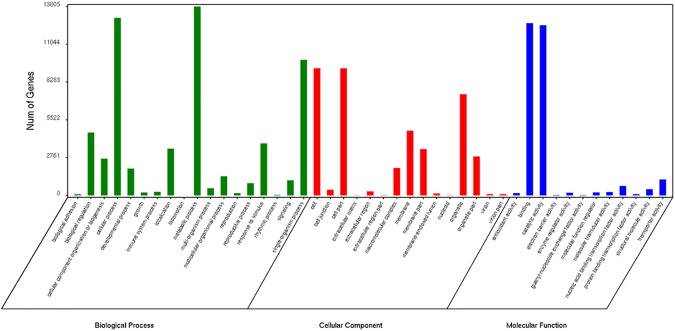
The GO assignments were used to classify the functions of the luffa transcripts. Based on sequence homologies, the 24,021 annotated sequences with BLAST hits to PlantGDB proteins were categorized into 45 functional groups (Fig 5). “Metabolic process”, “cellular process”, and “binding” were the dominant terms in the “biological process”, “cellular component”, and “molecular function” main categories, respectively. We also identified a relatively large number of genes associated with “catalytic activity”, “single-organism process”, and “cell”, with only a few genes related to “extracellular region part”, “nucleoid”, “extracellular matrix”, and “locomotion”.
Fig 5. Summary of the GO classification of Unigenes.
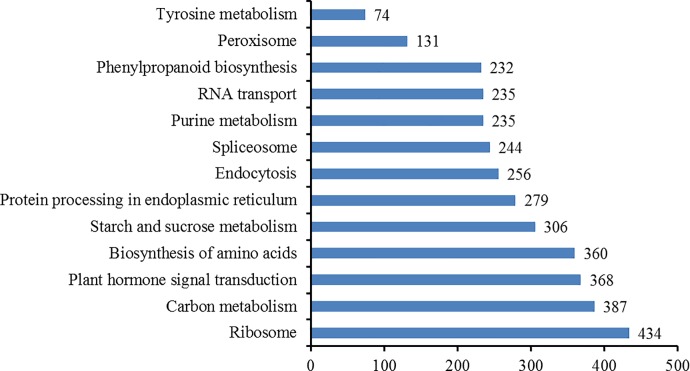
Pathway-based analyses may be useful for clarifying the biological functions of genes. The KEGG pathway database contains information regarding networks of intracellular molecular interactions, as well as their organism-specific variations [35]. To identify the biological pathways that are active during the browning of fresh-cut luffa fruits, we mapped the 58,073 Unigenes to the reference canonical pathways in the KEGG database. A total of 13,021 Unigenes were annotated (S2 Table). The top 10 pathways (i.e., “ribosome”, “carbon metabolism”, “plant hormone signal transduction” “biosynthesis of amino acids”, “starch and sucrose metabolism”, “protein processing in endoplasmic reticulum”, “endocytosis”, “spliceosome”, “purine metabolism”, and “RNA transport”) and three potential browning-related pathways (i.e., “phenylpropanoid biosynthesis”, “peroxisome”, “tyrosine metabolism”) are presented in Fig 6. Moreover, there were 47, 131, and 232 Unigenes associated with the “tyrosine metabolism”, “peroxisome” and “phenylpropanoid biosynthesis” pathways, respectively. These results indicated that diverse metabolic processes are active in luffa fruit flesh, resulting in the synthesis of diverse metabolites. These annotations may serve as a valuable resource for investigating specific processes, functions, and pathways, and facilitate the identification of novel genes involved in the browning of luffa fruit flesh.
Fig 6. Pathway assignments based on the KEGG database.
Analysis of differentially expressed genes
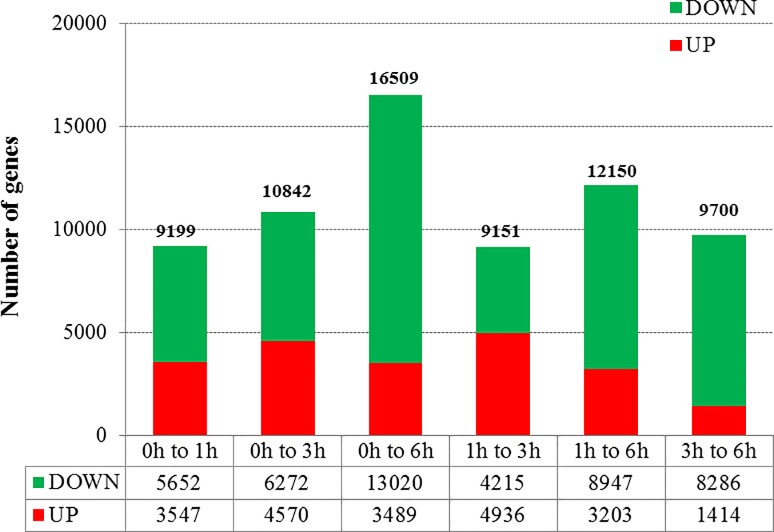
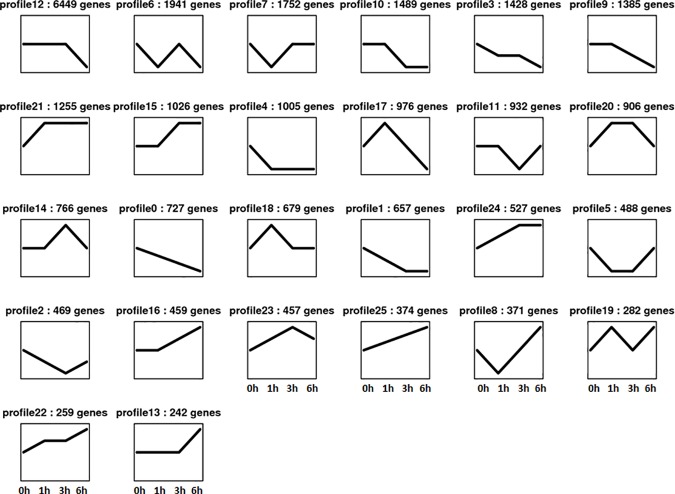
Based on a published protocol, we developed an algorithm to identify genes that were differentially expressed between two samples [36]. The FDR was used to determine the P-value threshold in multiple analyses for calculating the expression-level differences between two samples [37]. We calculated the number of expressed sequence tags corresponding to each gene in each library to estimate gene expression levels. A total of 27,301 Unigenes were differentially expressed during the browning of luffa fruits (S3 Table). The highest number of DEGs was detected during the comparison between the 0 and 6 h time points [i.e., 16,509 (3,489 up- and 13,020 down-regulated at 6 h) (log2 ratio ≥ 1; FDR ≤ 0.05; P ≤ 0.01)] (Fig 7). We used the STEM method to refine the sets of genes that were differently expressed at a minimum of four time points (i.e., 0, 1, 3, and 6 h). This method is commonly used for gene clustering in transcriptomic studies [31]. A total of 27,301 DEGs were clustered into 26 possible model profiles based on expression patterns (Fig 8). The DEGs were then subjected to GO and KEGG pathway enrichment analyses.
Fig 7. Differentially expressed genes in fresh-cut luffa fruits at various time points.
Fig 8. Clustering and classification of 27,301 differentially expressed genes.
The numbers in the top corner of each panel represent the identification number of the 26 profiles and the number of identified genes in the cluster, respectively.
Identification of potential browning-related genes
We selected DEGs with an FDR ≤ 0.05 and an expression ratio > 2 for GO functional and KEGG pathway analyses (S4 Table). We used the MapMan tool to visualize the annotated processes of interest or metabolic pathways. We conducted pathway enrichment analyses to identify significantly enriched metabolic or signal transduction pathways associated with the DEGs [38]. According to GO functional annotations and the screening of 58,073 Unigenes, we identified 11 Unigenes with potential roles in the browning of fresh-cut luffa fruits, including three PPO genes (Unigene0043875, Unigene0050171, and Unigene0015096), two PAL genes (Unigene0044748 and Unigene0024978), one POD gene (Unigene0011770), two CAT genes (Unigene 0036262 and Unigene0033876), and three SOD genes (Unigene0021782, Unigene0008835, and Unigene0015506), as well as four WRKY transcription factors (S1 Table). The Unigenes were annotated based on the NCBInr protein database and divided into the following categories: oxidoreductase, ammonia lyase, antioxidant, catalytic activity, and DNA-binding function related. Furthermore, a KEGG analysis revealed that 15 genes may affect tyrosine metabolism, phenylpropanoid biosynthesis, or the peroxisome pathway (S4 Table). Additionally, with the aid of rapid amplification of cDNA ends technology, we obtained the full-length CAT (Unigene0036262) sequence, which has been deposited in the GenBank database (Table 2).
Table 2. Unigenes potentially associated with the browning of luffa fruits.
| Unigene ID |
Nuleotide length (bp) |
Full length |
Protein Length (aa) |
Homologous function in nr |
Homology species & Accession number |
E-value |
GenBank accession numbers |
|---|---|---|---|---|---|---|---|
| Unigene0043875 | 2,189 | Yes | 589 | Polyphenol oxidase, chloroplastic-like |
Cucumis melo XP_008442274.1 |
0.0 | KM506756 |
| Unigene0050171 | 1,990 | Yes | 574 | Polyphenol oxidase, chloroplastic-like |
Cucumis melo XP_008442274.1 |
0.0 | KR819890 |
| Unigene0015096 | 2,189 | Yes | 593 | PREDICTED: polyphenol oxidase, chloroplastic-like |
Cucumis melo XP_011047843.1 |
0.0 | KX092429.1 |
| Unigene0044748 | 2,365 | Yes | 715 | Phenylalanine ammonia-lyase 1 |
Cucumis sativus XP_004143255.1 |
0.0 | KP341758 |
| Unigene0024978 | 2,274 | Yes | 713 | Phenylalanine ammonia-lyase-like |
Cucumis sativus XP_004145752.1 |
0.0 | KR491944 |
| Unigene0011770 | 1,319 | Yes | 331 | Peroxidase 2-like |
Cucumis melo XP_008449774.1 |
0.0 | KM506755 |
| Unigene0036262 | 1,259 | 5′ | 492 | Catalase isozyme 1 |
Cucumis melo XP_008452956.1 |
0.0 | KP222260 |
| Unigene0033876 | 1,709 | Yes | 492 | Catalase |
Cucurbita moschata AHF27430.1 |
0.0 | KR184674 |
| Unigene0021782 | 1,775 | Yes | 152 | Superoxide dismutase [Cu-Zn]-like |
Cucumis sativus NP_001267697.1 |
5E-89 | KP178922 |
| Unigene0008835 | 799 | Yes | 157 | PREDICTED: superoxide dismutase [Cu-Zn] 2 |
Cucumis sativus XP_004147539.1 |
2E-98 | KX092445.1 |
| Unigene0015506 | 1,011 | Yes | 221 | PREDICTED: superoxide dismutase [Cu-Zn] |
Cucumis sativus XP_008465422.1 |
7E-131 | KX092446.1 |
| Unigene0018509 | 2,189 | Yes | 299 | probable WRKY transcription factor 69-like |
Cucumis sativus NM_001280699.1 |
5E-172 | KY621843 |
| Unigene0021412 | 1,990 | Yes | 616 | PREDICTED: probable WRKY transcription factor 31 |
Cucumis sativus XM_004134727.2 |
0.0 | KY621844 |
| Unigene0025291 | 2,365 | Yes | 435 | PREDICTED: probable WRKY transcription factor 31 |
Cucumis sativus XM_011660396.1 |
0.0 | KY621845 |
| Unigene0034271 | 2,274 | Yes | 388 | PREDICTED: probable WRKY transcription factor 7 isoform X2 |
Cucumis melo XM_008454358.2 |
0.0 | KY621846 |
| Unigene0019122 | 1,273 | Yes | 178 | PREDICTED: ethylene-responsive transcription factor ERF109-like |
Cucumis melo XP_008454945.1 |
2E-93 | MF678591 |
| Unigene0003760 | 1,012 | Yes | 119 | PREDICTED: abscisic acid receptor PYL8-like |
Cucumis melo XP_008461524.1 |
6E-120 | MF678592 |
| Unigene0031055 | 1,111 | Yes | 205 | Gibberellin 2-oxidase |
Cucurbita maxima XP_008439633.1 |
0.0 | MF678593 |
Validation of differentially expressed genes based on RNA-seq data by quantitative real-time PCR
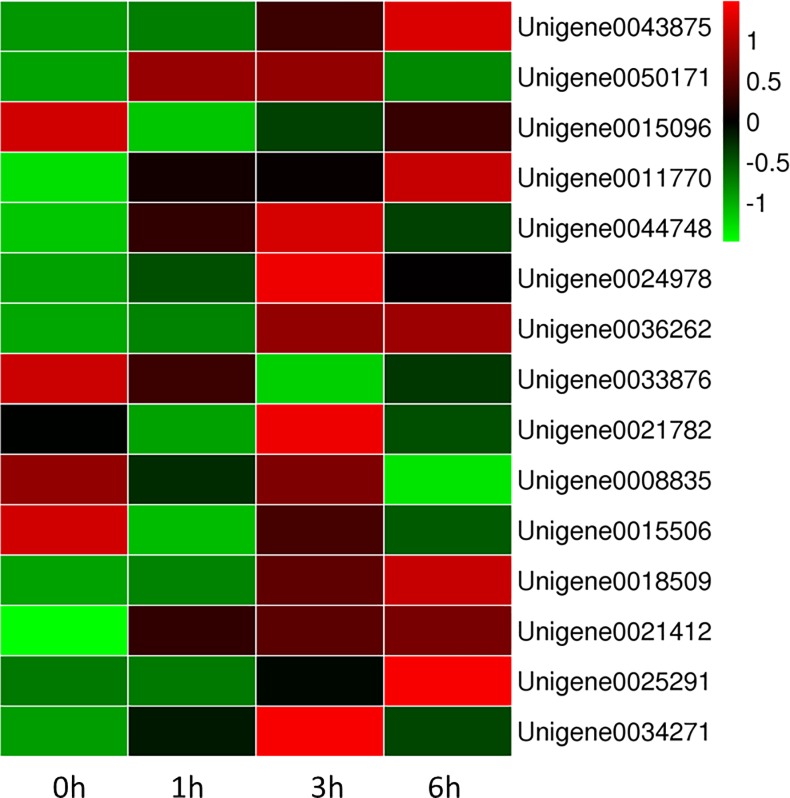
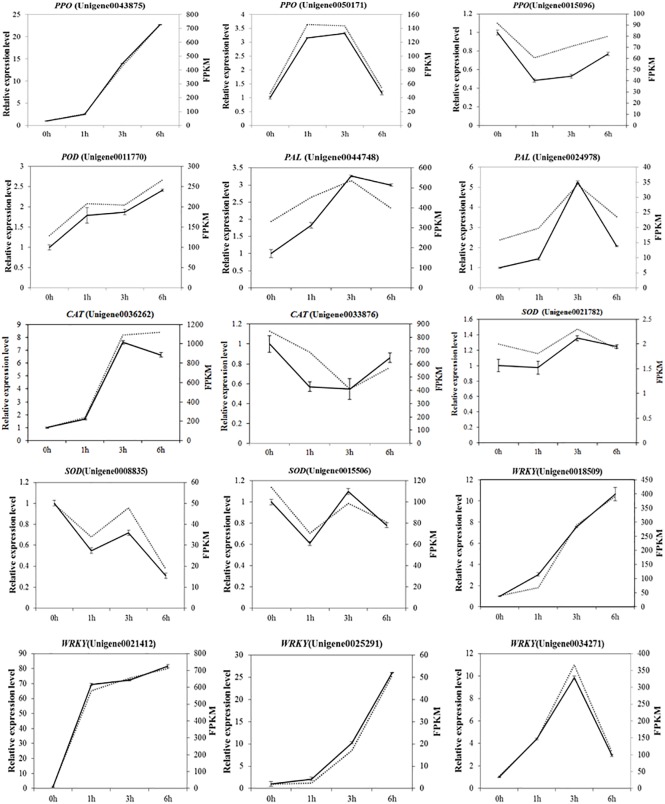
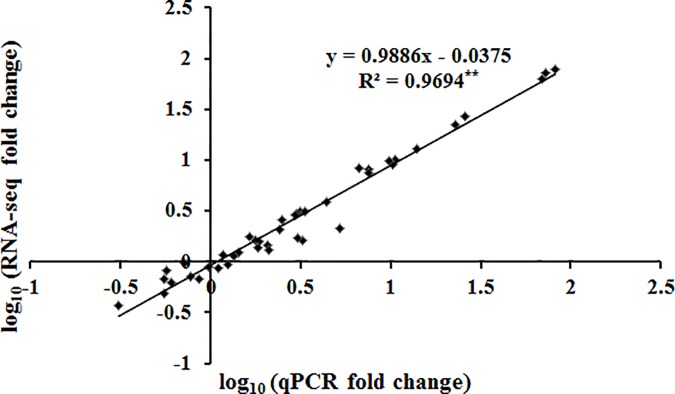
The heat maps for the 11 browning-related genes and four WRKY transcription factors indicated that the expression levels varied at different time points (i.e., 0, 1, 3, and 6 h) (Fig 9). We used qRT-PCR with gene-specific primers (Table 3) to analyze the expression profiles of genes likely involved in the browning of fresh-cut luffa fruits. The expression levels were mainly up-regulated relative to the 0 h expression levels, except for SOD (Unigene0021782), which was down-regulated at the 1 h time point, and PPO (Unigene0015096), CAT (Unigene0033876), and SOD (Unigene0008835 and Unigene0015506), which were down-regulated at the 1, 3, and 6 h time points (Fig 10). A linear regression analysis revealed a strong positive correlation (R2 = 0.9694; P ≤ 0.01) between the fold-change values based on qRT-PCR and RNA-seq data (Fig 11). These results confirm the reliability of the RNA-seq data generated in this study.
Fig 9. Clustering of 15 browning-related genes.
The color scale represents FPKM-normalized log2-transformed gene expression levels. Different columns represent different samples, and different lines represent different genes.
Table 3. Details regarding the qRT-PCR primers used to analyze 15 genes.
| Unigene ID | Directions | Primers | Product size |
TM (°C) | Relative Expression Level |
|---|---|---|---|---|---|
| Unigene0043875 | Forward | CGAATGTTCACTGTGCCTAT | 191 | 60 | up |
| Reverse | CTTCGGAGCGTCGTAGT | ||||
| Unigene0050171 | Forward | CTAGCCGTGGAAACCGA | 218 | 62 | up |
| Reverse | TGATTGGCTCACAGTGGA | ||||
| Unigene0015096 | Forward | AGTTCCCGATCACTCTTAAT | 216 | 58 | down |
| Reverse | TCACGAAGCTGCCG | ||||
| Unigene0044748 | Forward | CGCTGGACCCAACGTACAGG | 195 | 66 | up |
| Reverse | GGTTGACAGTGTCGACTGTGTCC | ||||
| Unigene0024978 | Forward | GCTCAATTTTCTGAGCTTGT | 169 | 59 | up |
| Reverse | GGACGTGGCTTGTTAC | ||||
| Unigene0011770 | Forward | GCCAATGTACTTGCTCTTCT | 199 | 60 | up |
| Reverse | GGTCCAACTCATGTAACTTCT | ||||
| Unigene0036262 | Forward | CTAGTGGAGAAACTTGCTAACT | 189 | 59 | up |
| Reverse | GGATAACAGTGGAGAAACGT | ||||
| Unigene0033876 | Forward | TCACCATAACAACCACCATGAAG | 152 | 62 | down |
| Reverse | CACACACCTTTCTCTCTTTCCG | ||||
| Unigene0021782 | Forward | CACAGGAAAGATGGTGAAGG | 210 | 60 | up |
| Reverse | CCAGCAGGGTTGAAATGT | ||||
| Unigene0018509 | Forward | CATATCCACCGTCCGATTC | 214 | 59 | up |
| Reverse | TGGTGGTGAGATTTGGTG | ||||
| Unigene0021412 | Forward | CTAATGAACCCAAATTTATTGGCT | 165 | 60 | up |
| Reverse | GAAGGGTACGTGGAACG | ||||
| Unigene0025291 | Forward | CAGTTTATGGATCTTGGATTGG | 200 | 59 | up |
| Reverse | GAAGAAGAAGACGAAGAACTG | ||||
| Unigene0034271 | Forward | CATCACTCCGAAGACCATTC | 172 | 60 | up |
| Reverse | GGGAAGTAGTAGTAACAACGG | ||||
| Unigene0008835 | Forward | CCACGCTCTTGGCGATACA | 75 | 61 | down |
| Reverse | CCATGGTCCTTCTTCAATGGA | ||||
| Unigene0015506 | Forward | CACTTCTCCATAGCAAATGC | 127 | 59 | down |
| Reverse | GGTCAGGGAAGGCG | ||||
| 18SrRNA | Forward | GTGTTCTTCGGAATGACTGG | 271 | 60 | - |
| Reverse | ATCGTTTACGGCATGGACTA |
Fig 10. Quantitative real-time polymerase chain reaction analyses of the expression levels of selected Unigenes during the browning of luffa fruits.
18S rRNA was used as the internal control. The error bars represent the standard error of three biological replicates.
Fig 11. Correlation analysis of fold-change values based on RNA-seq and qRT-PCR data.
The RNA-seq fold-change refers to the ratio of FPKM values from 0 to 6 h. The qRT-PCR fold-change refers to the expression levels at 1, 3, and 6 h normalized against the expression level at 0 h. **significant correlation (P < 0.01).
Discussion
Luffa is an important vegetable crop worldwide. It is rich in nutrients and medicinal compounds, including carotenoids, β-carotene, and α-carotene, which maintain the immune system [39]. To identify genes involved in the browning of luffa fruits, total RNA was extracted from ‘Fusi-3’ fruits and used for mRNA preparation, fragmentation, and cDNA synthesis. The cDNA was sequenced using the Illumina Genome Analyzer IIx platform, and the resulting sequencing data were subjected to bioinformatic analysis. We generated 92,010,328 raw reads from a 200-bp insert (mean size) cDNA library. After trimming adapter sequences and removing low quality reads, 90,226,276 clean reads were obtained. Of these clean reads, 94.19% were considered high-quality reads, which were analyzed further. We annotated 35,345 Unigenes (i.e., 60.86% of the assembled Unigenes), suggesting their functions were relatively conserved. In addition to identifying genes or proteins, we also analyzed the associated GO terms and potential roles in metabolic pathways. Combining our data regarding Unigenes expressed in fresh-cut luffa fruits with the results of a previous transcriptomic study of the same luffa variety reveals that luffa fruits may be a useful source of abundant genes [40]. The current study represents the first attempt at applying Illumina paired-end sequencing technology to investigate the changes in the transcriptome during the browning of fresh-cut luffa fruits.
Enzymatic processes are generally recognized as the main determinants of browning. The accumulation of transcripts for genes associated with the oxidation of phenolic compounds is a hallmark of enzymatic browning. Many studies have confirmed that browning-related genes are differentially expressed in fresh-cut vegetables [41–43]. The PPO and POD genes have been associated with the browning of fruit and vegetable tissues, and their encoded enzymes function together to induce browning [44,45]. The PPO enzyme encoded by members of the PPO multigene family is considered to be a major factor responsible for enzymatic browning of several fresh-cut vegetables [17,46]. Zhou et al. (2015) [47] reported that the enzymatic browning of fresh-cut Chinese chestnut can be inhibited by applying salicylic acid to decrease PPO activity. Their findings provide indirect evidence that PPO influences the browning of fresh-cut Chinese chestnut. Moreover, a previous study revealed that a 3-day treatment with 1 mg L−1 ozone decreased POD activity in fresh-cut lettuce, resulting in obviously inhibited browning [48]. Additionally, Chisari et al. (2007) [46] reported that the POD enzyme also affected the browning of fresh-cut melon. In this study, the observed total phenol contents and the results of a KEGG analysis revealed that three PPO genes (Unigene0015096, Unigene0043875, and Unigene0015096) and one POD gene (Unigene0011770) related to the tyrosine metabolism pathway were differentially expressed while the phenolic acid (i.e., gallic acid) content increased in fresh-cut luffa fruit slices (S4 Table) [49,50]. The production of phenolic acids, including gallic acid, L-tyrosine, chlorogenic acid, and gentisic acid, catalyzed by browning-related enzymes (i.e., PPO and POD) can be affected by the tyrosine metabolism pathway [51–54]. However, the exact roles of PPO and POD during responses to the browning of fresh-cut luffa fruits remains to be determined.
In addition to PPO and POD, the PAL, CAT, and SOD genes are also considered to have important roles in the enzymatic browning of fruits and vegetables [55,56]. The PAL enzyme is reportedly involved in the browning of fresh-cut potatoes [13]. Additionally, a lettuce PAL homolog was observed to be responsible for regulating the browning of fresh-cut edges [57]. Our results indicated that the relative expression levels of two luffa PAL genes (Unigene0044748 and Unigene0024978) involved in the phenylpropanoid biosynthesis pathway were up-regulated (S4 Table). It is widely accepted that phenylpropanoid biosynthesis pathways are important for controlling the browning of cut vegetables and fruits (S4 Table) [58–61]. Additionally, the CAT enzyme has a pivotal role in the browning of fresh-cut fruits and vegetables [62–64]. Dong et al. (2015) [44] proposed that the activity of CAT can be enhanced by a short-term carbon dioxide treatment, leading to inhibited browning of fresh-cut burdock during storage at low temperatures. Meanwhile, Fan et al. (2016) [65] reported that significantly higher activities of antioxidant enzymes (i.e., SOD and CAT) were maintained in browning ‘Laiyang’ pear fruits, although PPO activity was inhibited. Another important enzyme related to the browning of fresh-cut fruits and vegetables is SOD, which is essential for enhancing antioxidant activities in fresh-cut Chinese water chestnut [66]. In this study, two CAT genes (Unigene0036262 and Unigene0033876) and three SOD genes (Unigene0021782, Unigene0008835, and Unigene0015506) related to the peroxisome pathway were also differentially expressed during the browning of fresh-cut luffa fruits (S4 Table).
The browning of fruits and vegetables is caused by the overproduction of reactive oxygen species [67–69]. Members of the WRKY transcription factor superfamily have been detected exclusively in plants, and affect diverse physiological responses and metabolic pathways [70,71]. Previous studies confirmed that WRKY transcription factors are important inducers of the expression of genes associated with the reactive oxygen species pathway during plant responses to biotic, abiotic, and oxidative stresses [72,73]. In this study, we observed that the expression levels of four Unigenes (Unigene0018509, Unigene0021412, Unigene0025291, and Unigene0034271) annotated as WRKY transcription factors were significantly up-regulated at various time points (i.e., 0, 1, 3 and 6 h) during the browning of fresh-cut luffa fruits. Analyses revealed that these four transcription factors (GenBank ID: KY621843–KY621846) were members of the Group II WRKY superfamily, and contained a conserved WRKY domain and C2H2-type zinc fingers [74]. Furthermore, the current study is the first to reveal a relationship between these four WRKY transcription factors and the browning of luffa fruits. Further studies involving structural and functional characterizations are required to determine whether the identified candidate genes influence the browning of fresh-cut luffa fruits.
Transcriptome profiles of different luffa cultivars showed that the expression levels of PPO POD and PAL genes related with browning in XTR05 (browning sensitive) were significantly higher than that in YLB05 (browning resistant) [40]. In this study, we also identified three PPO genes, one POD gene and two PAL genes were differentially expressed during the browning of fresh-cut luffa fruits (i.e., after 1–6 h). Moreover, we identified two CAT genes, three SOD genes and four WRKY genes, which were differentially expressed and may play an important role in browning process of fresh-cut luffa fruits. We also observed the differential expression of other Unigenes, such as Unigene0019122, Unigene0003760, and Unigene0031055, which were predicted to encode an ethylene-responsive transcription factor ERF109-like protein, an abscisic acid receptor PYL8-like protein, and gibberellin 2-oxidase, respectively (Table 2). These Unigenes may contribute to the browning of fruits and vegetables [40,75,76]. However, their potential effects on the browning of fresh-cut luffa fruits will need to be confirmed in future studies.
Conclusions
We herein describe the first use of Illumina sequencing technology to investigate the changes in the transcriptome during the browning of fresh-cut luffa fruit slices. A total of 27,301 differentially expressed Unigenes were detected in four sequenced libraries. Additionally, 11 browning-related genes from five gene families (i.e., PPO, PAL, POD, CAT, and SOD) as well as four WRKY transcription factors were observed to be differentially regulated in fresh-cut luffa fruits. These genetic resources and putative signaling pathways related to luffa defense responses against browning may be useful for future molecular studies of L. cylindrica.
Supporting information
(XLSX)
(XLSX)
(XLSX)
(XLSX)
Acknowledgments
The authors thank Zhi Zou and two anonymous reviewers for their helpful suggestions, and Liwen Bianji, Edanz Editing China (www.liwenbianji.cn/ac) for editing the English text of a draft of this manuscript.
Data Availability
All relevant data are within the paper and its Supporting Information files.
Funding Statement
This work was supported by Natural Science fundation of Fujian Province (2015J01118), Project of Innovative Team in Fujian Academy of Agricultural Sciences (2016PI-40), The Fujian Provincial Public Research Institute of Fundamental Research (2017R1026-6), and Fujian Academy of Agricultural Sciences Vegetable Science and Technology Innovation Team (STIT2017-1-2). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Nath N, Deka K. Effect of GA3 and KNO3 on seedling establishment of Luffa acutangula (L.) Roxb. Int J Pure App Biosci. 2015; 3(6): 99–103. [Google Scholar]
- 2.Partap S, Kumar A, Sharma NK, Jha KK. Luffa cylindrica: an important medicinal plant. J Nat Prod Plant Res. 2012; 2: 127–134. [Google Scholar]
- 3.Sutharshana V. Protective role of Luffa cylindrical. J Pham Sci Res. 2013; 5: 184–186. [Google Scholar]
- 4.Wang C, Wang H, Lou L, Su X. The identification method of Luffa cylindrical flesh browning. Jiangsu Agric Sci. 2012; 40: 137–138. [Google Scholar]
- 5.Bustos MC, Mazzobre MF, Buera MP. Stabilization of refrigerated avocado pulp: Effect of Allium and Brassica extracts on enzymatic browning. LWT-Food Sci Technol. 2015; 61: 89–97. [Google Scholar]
- 6.Deuchande T, Larrigaudiere C, Gine-Bordonaba J, Carvalho PSM, Vasconcelos MW. Biochemical basis of CO2-related internal browning disorders in pears (Pyrus communis L. cv. Rocha) during long term storage. J Agr Food Chem. 2016; 64: 4336–4345. [DOI] [PubMed] [Google Scholar]
- 7.Jiang J, Jiang L, Luo H, Yu Z. Establishment of a statistical model for browning of fresh-cut lotus root during storage. Postharvest Biol Tec. 2014; 92: 164–171. [Google Scholar]
- 8.Du Y, Dou S, Wu S. Efficacy of phytic acid as an inhibitor of enzymatic and non-enzymatic browning in apple juice. Food Chem. 2012; 135: 580–582. doi: 10.1016/j.foodchem.2012.04.131 [DOI] [PubMed] [Google Scholar]
- 9.Wu S. Glutathione suppresses the enzymatic and non-enzymatic browning in grape juice. Food Chem. 2014; 160: 8–10. doi: 10.1016/j.foodchem.2014.03.088 [DOI] [PubMed] [Google Scholar]
- 10.Wei W, Xu C, Wu L, Ren J, Qu X. Non-enzymatic-browning-reaction: a versatile route for production of nitrogen-doped carbon dots with tunable multicolor luminescent display. Sci Rep. 2014; 4: 3564 doi: 10.1038/srep03564 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hong PK, Betti M. Non-enzymatic browning reaction of glucosamine at mild conditions: Relationship between colour formation, radical scavenging activity and α-dicarbonyl compounds production. Food Chem. 2016; 212: 234–243. doi: 10.1016/j.foodchem.2016.05.170 [DOI] [PubMed] [Google Scholar]
- 12.Friedman M. Food browning and its prevention: an overview. J Agric Food Chem. 1996; 44: 631–653. [Google Scholar]
- 13.Cantos E, Tudela JA, Gil MI, Espín JC. Phenolic compounds and related enzymes are not rate-limiting in browning development of fresh-cut potatoes. J Agric Food Chem. 2002; 50: 3015–3023. [DOI] [PubMed] [Google Scholar]
- 14.Martinez MV, Whitaker JR. The biochemistry and control of enzymatic browning. Trends Food Sci Tech. 1995; 6: 195–200. [Google Scholar]
- 15.Lante A, Tinello F, Nicoletto M. UV-A light treatment for controlling enzymatic browning of fresh-cut fruits. Innov Food Sci Emerg. 2016; 34: 141–147. [Google Scholar]
- 16.Lamikanra O, Watson MA. Effects of ascorbic acid on peroxidase and polyphenoloxidase activities in fresh-cut cantaloupe melon. J Food Sci. 2001; 66: 1283–1286. [Google Scholar]
- 17.Degl’Innocenti E, Pardossi A, Tognoni F, Guidi L. Physiological basis of sensitivity to enzymatic browning in ‘lettuce’, ‘escarole’ and ‘rocket salad’when stored as fresh-cut products. Food Chem. 2007; 104: 209–215. [Google Scholar]
- 18.Droillard MJ, Paulin A, Massot JC. Free radical production, catalase and superoxide dismutase activities and membrane integrity during senescence of petals of cut carnations (Dianthus caryophyllus). Physiol Plantarum. 1987; 71: 197–202. [Google Scholar]
- 19.Liu P, Xue C, Wu TT, Heng W, Jia B, Ye Z, Liu L, et al. Molecular analysis of the processes of surface brown spot (SBS) formation in pear fruit (Pyrus bretschneideri Rehd. Cv. Dangshansuli) by de novo transcriptome assembly. PLoS One. 2013; 8: e74217 doi: 10.1371/journal.pone.0074217 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Mellidou I, Buts K, Hatoum D, Ho QT, Johnston JW, Watkins CB, et al. Transcriptomic events associated with internal browning of apple during postharvest storage. BMC Plant Biol. 2014; 14: 328–344. doi: 10.1186/s12870-014-0328-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Dassanayake M, Haas JS, Bohnert HJ, Cheeseman JM. Shedding light on an extremophile life style through transcriptomics. New Phytol. 2010; 183: 764–775. [DOI] [PubMed] [Google Scholar]
- 22.Sangwan RS, Tripathi S, Singh J, Narnoliya LK, Sangwan NS. De novo sequencing and assembly of Centella asiatica leaf transcriptome for mapping of structural, functional and regulatory genes with special reference to secondary metabolism. Gene. 2013; 525: 58–76. doi: 10.1016/j.gene.2013.04.057 [DOI] [PubMed] [Google Scholar]
- 23.Han XJ, Wang YD, Chen YC, Lin LY, Wu QK. Transcriptome sequencing and expression analysis of terpenoid biosynthesis genes in Litsea cubeba. PLoS One. 2013; 8: e76890 doi: 10.1371/journal.pone.0076890 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Dewanto V, Wu X, Adom KK, Liu RH. Thermal processing enhances the nutritional value of tomatoes by increasing total antioxidant activity. J Agric Food Chem. 2002; 50: 3010–3014. [DOI] [PubMed] [Google Scholar]
- 25.Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol. 2011; 29: 644–652. doi: 10.1038/nbt.1883 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics. 2005; 21: 3674–3676. doi: 10.1093/bioinformatics/bti610 [DOI] [PubMed] [Google Scholar]
- 27.Ye J, Fang L, Zheng H, Zhang Y, Chen J, Zhang Z, et al. WEGO: a web tool for plotting GO annotations. Nucleic Acids Res. 2006; 34(Web Server issue): W293–297. doi: 10.1093/nar/gkl031 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Langmead B, Trapnell C, Pop M, Salzberg SL. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009; 10: R25 doi: 10.1186/gb-2009-10-3-r25 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Li B, Dewey C. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics. 2011; 12: 323 doi: 10.1186/1471-2105-12-323 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Wang L, Feng Z, Wang X, Wang X, Zhang X. DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics. 2010; 26: 136–138. doi: 10.1093/bioinformatics/btp612 [DOI] [PubMed] [Google Scholar]
- 31.Ernst J, Bar-Joseph Z. STEM: a tool for the analysis of short time series gene expression data. BMC Bioinformatics. 2006; 7(1): 191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zhu HS, Chen MD, Wen QF, Lan XL, Li YP, Wang B, et al. Cloning of 18S rRNA gene from Luffa cylindrical and its application as an internal standard. Acta Agric Nucl Sin J. 2016; 30: 0035–0041. [Google Scholar]
- 33.Wen QF, Liu JT, Zhu HS, Chen MD, Wang B, Zhang QR. Cloning and expression analysis of catalase CAT1 gene from Luffa cylindrical. Acta Hortic Sin. 2016; 43: 2039–2048. [Google Scholar]
- 34.Sun Y, Zhang W, Zeng T, Nie Q, Zhang F, Zhu L. Hydrogen sulfide inhibits enzymatic browning of fresh-cut lotus root slices by regulating phenolic metabolism. Food Chem. 2015; 177: 376–381. doi: 10.1016/j.foodchem.2015.01.065 [DOI] [PubMed] [Google Scholar]
- 35.Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 2008; 36(Database issue): D480–484. doi: 10.1093/nar/gkm882 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Audic S, Claverie JM. The significance of digital gene expression profiles. Genome Res. 1997; 7: 986–95. [DOI] [PubMed] [Google Scholar]
- 37.Benjamini Y, Yekutieli D. The control of the false discovery rate in multiple testing under dependency. Ann Stat. 2001; 29: 1165–1188. [Google Scholar]
- 38.Li XY, Sun H, Pei J, Dong Y, Wang F, Chen H, et al. De novo sequencing and comparative analysis of the blueberry transcriptome to discover putative genes related to antioxidants. Gene. 2012; 511: 54–61. doi: 10.1016/j.gene.2012.09.021 [DOI] [PubMed] [Google Scholar]
- 39.Isutsa DK, Mallowa SO. Increasing leaf harvest intensity enhances edible leaf vegetable yields and decreases mature fruit yields in multi-purpose pumpkin. J Agric Biol Sci. 2013; 8: 610–615. [Google Scholar]
- 40.Chen X, Tan T, Xu C, Huang S, Tan J, Zhang M, et al. Genome-wide transcriptome profiling reveals novel insights into Luffa cylindrica browning. Biochem Bioph Res Co. 2015; 463: 1243–1249. [DOI] [PubMed] [Google Scholar]
- 41.Docimo T, Francese G, De Palma M, Mennella D, Toppino L, Lo Scalzo R, et al. Insights in the fruit flesh browning mechanisms in Solanum melongena genetic lines with opposite post-cut behavior. J Agr Food Chem. 2016; 64: 4675–4685. [DOI] [PubMed] [Google Scholar]
- 42.Xu C, Zeng B, Huang J, Huang W, Liu Y. Genome-wide transcriptome and expression profile analysis of phalaenopsis during explant browning. PloS One. 2015; 10: e0123356 doi: 10.1371/journal.pone.0123356 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Min T, Xie J, Zheng M, Yi Y, Hou W, Wang L, et al. The effect of different temperatures on browning incidence and phenol compound metabolism in fresh-cut lotus (Nelumbo nucifera G.) root. Postharvest Biol Tec. 2017; 123: 69–76. [Google Scholar]
- 44.Pen LT, Jiang YM. Effects of chitosan coating on shelf life and quality of fresh-cut Chinese water chestnut. LWT-Food Sci Technol. 2003; 36: 359–364. [Google Scholar]
- 45.Dong T, Shi J, Jiang CZ, Feng Y, Cao Y, Wang Q. A short-term carbon dioxide treatment inhibits the browning of fresh-cut burdock. Postharvest Biol Tec. 2015; 110: 96–102. [Google Scholar]
- 46.Chisari M, Barbagallo RN, Spagna G. Characterization and role of polyphenol oxidase and peroxidase in browning of fresh-cut melon. J Agr Food Chem. 2007; 56: 132–138. [DOI] [PubMed] [Google Scholar]
- 47.Zhou D, Li L, Wu Y, Fan J, Ouyang J. Salicylic acid inhibits enzymatic browning of fresh-cut Chinese chestnut (Castanea mollissima) by competitively inhibiting polyphenol oxidase. Food Chem. 2015; 171: 19–25. doi: 10.1016/j.foodchem.2014.08.115 [DOI] [PubMed] [Google Scholar]
- 48.Rico D, Martín-Diana AB, Frías JM, Henehan G, Barry-Ryan C. Effect of ozone and calcium lactate treatments on browning and texture properties of fresh-cut lettuce. J Sci Food Agr. 2006; 86(13): 2179–2188. [Google Scholar]
- 49.Lee HS. Tyrosinase inhibitors of Pulsatilla cernua root-derived materials. J Agr Food Chem. 2002; 50(6): 1400–1403. [DOI] [PubMed] [Google Scholar]
- 50.Carreira A, Dillinge K, Eliskases-Lechner F, Loureiro V, Ginzinger W, Rohm H. Influence of selected factors on browning of Camembert cheese. J Dairy Res. 2002; 69(2): 281–292. [DOI] [PubMed] [Google Scholar]
- 51.Richard FC, Goupy PM, Nicolas JJ, Lacombe JM, Pavia AA. Cysteine as an inhibitor of enzymic browning. 1. Isolation and characterization of addition compounds formed during oxidation of phenolics by apple polyphenol oxidase. J Agr Food Chem. 1991; 39(5): 841–847. [Google Scholar]
- 52.Fukumoto LR, Toivonen PM, Delaquis PJ. Effect of wash water temperature and chlorination on phenolic metabolism and browning of stored iceberg lettuce photosynthetic and vascular tissues. J Agr Food Chem. 2002; 50(16): 4503–4511. [DOI] [PubMed] [Google Scholar]
- 53.Sullivan ML. Beyond brown: polyphenol oxidases as enzymes of plant specialized metabolism. Front Plant Sci. 2014; 5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Zhan L, Hu J, Pang L, Li Y, Shao J. Light exposure reduced browning enzyme activity and accumulated total phenols in cauliflower heads during cool storage. Postharvest Biol Tec. 2014; 88: 17–20. [Google Scholar]
- 55.Chi M, Bhagwat B, Lane WD, Tang G, Su Y, Sun R, et al. Reduced polyphenol oxidase gene expression and enzymatic browning in potato (Solanum tuberosum L.) with artificial microRNAs. BMC Plant Biol. 2014; 14(1): 62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Dokhanieh AY, Aghdam MS, Sarcheshmeh MAA. Impact of postharvest hot salicylic acid treatment on aril browning and nutritional quality in fresh-cut pomegranate. Hortic Environ Biote. 2016; 57: 378–384. [Google Scholar]
- 57.Luna MC, Tudela JA, Tomás-Barberán FA, Gil MI. Modified atmosphere (MA) prevents browning of fresh-cut romaine lettuce through multi-target effects related to phenolic metabolism. Postharvest Biol Tec. 2016; 119: 84–93. [Google Scholar]
- 58.Saltveit ME. Wound induced changes in phenolic metabolism and tissue browning are altered by heat shock. Postharvest Biol Tec. 2000; 21(1): 61–69. [Google Scholar]
- 59.Hisaminato H, Murata M, Homma S. Relationship between the enzymatic browning and phenylalanine ammonia-lyase activity of cut lettuce, and the prevention of browning by inhibitors of polyphenol biosynthesis. Biosci Biotech Bioch. 2001; 65(5): 1016–1021. [DOI] [PubMed] [Google Scholar]
- 60.Saltveit ME. Effect of 1-methylcyclopropene on phenylpropanoid metabolism, the accumulation of phenolic compounds, and browning of whole and fresh-cut ‘iceberg’ lettuce. Postharvest Biol Tec. 2004; 34(1): 75–80. [Google Scholar]
- 61.Chuanjun X, Zhiwei R, Ling L, Biyu Z, Junmei H, Wen H, et al. The effects of polyphenol oxidase and cycloheximide on the early stage of browning in Phalaenopsis explants. Hortic Plant J. 2015; 1(3): 172–180. [Google Scholar]
- 62.Soliva-Fortuny RC, Martín-Belloso O. New advances in extending the shelf-life of fresh-cut fruits: a review. Trends Food Sci Tech. 2003; 14: 341–353. [Google Scholar]
- 63.Artés-Hernández F, Robles PA, Gómez PA, Tomás-Callejas A, Artés F. Low UV-C illumination for keeping overall quality of fresh-cut watermelon. Postharvest Biol Tec. 2010; 55: 114–120. [Google Scholar]
- 64.Maghoumi M, Gómez PA, Mostofi Y, Zamani Z, Artés-Hernández F, Artés F. Combined effect of heat treatment, UV-C and superatmospheric oxygen packing on phenolics and browning related enzymes of fresh-cut pomegranate arils. LWT-Food Sci Technol. 2013; 54: 389–396. [DOI] [PubMed] [Google Scholar]
- 65.Fan M, Li W, Hu X, Sun YN, Yu G, Zhang X. Effect of micro-vacuum storage on active oxygen metabolism, internal browning and related enzyme activities in Laiyang pear (Pyrus bretschneideri Reld). LWT-Food Sci Technol. 2016; 72: 467–474. [Google Scholar]
- 66.You Y, Jiang Y, Sun J, Liu H, Song L, Duan X. Effects of short-term anoxia treatment on browning of fresh-cut Chinese water chestnut in relation to antioxidant activity. Food Chem. 2012; 132: 1191–1196. [DOI] [PubMed] [Google Scholar]
- 67.Yang E, Lu W, Qu H, Lin H, Wu F, Yang S, et al. Altered energy status in pericarp browning of litchi fruit during storage. Pak J Bot. 2009; 41(5): 2271–2279. [Google Scholar]
- 68.Yi C, Jiang Y, Shi J, Qu H, Xue S, Duan X, et al. ATP-regulation of antioxidant properties and phenolics in litchi fruit during browning and pathogen infection process. Food Chem. 2010; 118(1): 42–47. [Google Scholar]
- 69.Lin Y, Lin H, Zhang S, Chen Y, Chen M, Lin Y. The role of active oxygen metabolism in hydrogen peroxide-induced pericarp browning of harvested longan fruit. Postharvest Biol Tec. 2014; 96: 42–48. [Google Scholar]
- 70.Phukan UJ, Jeena GS, Shukla RK. WRKY transcription factors: molecular regulation and stress responses in plants. Front Plant Sci. 2016; 7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Kayıhan C, Öz MT, Eyidoğan F, Yücel M, Öktem HA. Physiological, biochemical, and transcriptomic responses to boron toxicity in leaf and root tissues of contrasting wheat cultivars. Plant Mol Biol Rep. 2017; 35(1): 97–109. [Google Scholar]
- 72.Eulgem T, Somssich IE. Networks of WRKY transcription factors in defense signalling. Curr Opin Plant Biol. 2007; 10: 366–371. doi: 10.1016/j.pbi.2007.04.020 [DOI] [PubMed] [Google Scholar]
- 73.Dan Y, Zhang S, Matherly A. Regulation of hydrogen peroxide accumulation and death of Agrobacterium-transformed cells in tomato transformation. Plant Cell Tiss Org. 2016; 127(1): 229–236. [Google Scholar]
- 74.Eulgem T, Rushton PJ, Robatzek S, Somssich IE. The WRKY superfamily of plant transcription factors. Trends Plant Sci. 2000; 5(5): 199–206. [DOI] [PubMed] [Google Scholar]
- 75.Lu X, Sun D, Li Y, Shi W, Sun G Pre- and post-harvest salicylic acid treatments alleviate internal browning and maintain quality of winter pineapple fruit. Sci Hortic. 2011; 130: 97–101. [Google Scholar]
- 76.Pusittigul I, Kondo S, Siriphanich J. Internal browning of pineapple (Ananas comosus L.) fruit and endogenous concentrations of abscisic acid and gibberellins during low temperature storage. Sci Hortic. 2012; 146: 45–51. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
(XLSX)
(XLSX)
(XLSX)
(XLSX)
Data Availability Statement
All relevant data are within the paper and its Supporting Information files.