ABSTRACT
Each influenza season, a set of wild-type viruses, representing one H1N1, one H3N2, and one to two influenza B isolates, are selected for inclusion in the annual seasonal influenza vaccine. In order to develop broadly reactive subtype-specific influenza vaccines, a methodology called computationally optimized broadly reactive antigens (COBRA) was used to design novel hemagglutinin (HA) vaccine immunogens. COBRA technology was effectively used to design HA immunogens that elicited antibodies that neutralized H5N1 and H1N1 isolates. In this report, the development and characterization of 17 prototype H3N2 COBRA HA proteins were screened in mice and ferrets for the elicitation of antibodies with HA inhibition (HAI) activity against human seasonal H3N2 viruses that were isolated over the last 48 years. The most effective COBRA HA vaccine regimens elicited antibodies with broader HAI activity against a panel of H3N2 viruses than wild-type H3 HA vaccines. The top leading COBRA HA candidates were tested against cocirculating variants. These variants were not efficiently detected by antibodies elicited by the wild-type HA from viruses selected as the vaccine candidates. The T-11 COBRA HA vaccine elicited antibodies with HAI and neutralization activity against all cocirculating variants from 2004 to 2007. This is the first report demonstrating broader breadth of vaccine-induced antibodies against cocirculating H3N2 strains compared to the wild-type HA antigens that were represented in commercial influenza vaccines.
IMPORTANCE There is a need for an improved influenza vaccine that elicits immune responses that recognize a broader number of influenza virus strains to prevent infection and transmission. Using the COBRA approach, a set of vaccines against influenza viruses in the H3N2 subtype was tested for the ability to elicit antibodies that neutralize virus infection against not only historical vaccine strains of H3N2 but also a set of cocirculating variants that circulated between 2004 and 2007. Three of the H3N2 COBRA vaccines recognized all of the cocirculating strains during this era, but the chosen wild-type vaccine strains were not able to elicit antibodies with HAI activity against these cocirculating strains. Therefore, the COBRA vaccines have the ability to elicit protective antibodies against not only the dominant vaccine strains but also minor circulating strains that can evolve into the dominant vaccine strains in the future.
KEYWORDS: COBRA, hemagglutination-inhibition, mice, influenza, H3N2
INTRODUCTION
Current influenza vaccination strategies elicit antibodies that bind to influenza HA (hemagglutinin) and neuraminidase (NA) glycoproteins on the surface of the virus (1, 2). Antigenic variation in circulating strains due to evolution of these two proteins results in immune evasion by preventing antibody binding and subsequent neutralization of infection. There is a need to develop influenza vaccine strategies that will elicit more broadly reactive or more universal influenza vaccine responses to recognize a larger number of circulating influenza virus variants within and across subtypes in any given season and over multiple seasons (1, 2).
For influenza viruses with a wide diversity of HA proteins, such as those in the H3N2 subtype, developing a broadly reactive influenza vaccine is a challenge. The viral HA protein from H3N2 influenza viruses rapidly evolves via antigenic drift, resulting in frequent emergence of antigenic variant strains (3–5) that requires updating of the annual influenza vaccine. Antigenic mismatches between the selected strain in the vaccine and cocirculating H3N2 viruses often contribute to reduced vaccine efficacy (6–8). An estimated 20 antigenic clusters (9, 10) have been detected since the H3N2 subtype was introduced into the human population in 1968 (10, 11). To address the need for more broadly reactive influenza vaccines, our group has previously reported on the methodology of antigen design, termed computationally optimized broadly reactive antigen (COBRA), using multiple rounds of layered consensus building to generate influenza vaccine HA immunogens. COBRA HA antigens are able to elicit potent, broadly reactive HA-specific antibody responses that protect against both seasonal and novel pandemic influenza virus strains that have undergone genetic drift. These vaccines induce immunity targeting both the globular head and stem regions of HA. This method has been used to address the diversity of H5N1 highly pathogenic avian influenza (1, 12–15) and seasonal and pandemic H1N1 subtypes (16). COBRA H5 HA designs elicited antibodies with HA inhibition (HAI) activity against 25 H5N1 strains that were isolated over a 12-year span. COBRA H1 HA antigens were able to elicit antibody responses to a panel of 17 H1N1 viruses isolated over an almost 100-year span of time (16). In this study, a set of 17 COBRA HA antigens was designed for the human seasonal H3N2 influenza subtype. Each of the COBRA HA antigens was expressed on virus-like particles (VLPs) and tested with adjuvant in mice for elicited antibody responses. These H3 COBRA vaccine candidates were used individually, in cocktails, or in prime-boost regimens to evaluate the effect on elicited antibodies compared to wild-type (WT) H3N2 HA antigens. For the first time, COBRA HA immunogens were tested against a set of viral variants that cocirculated between 2004 and 2007 that were not all neutralized using the Advisory Committee on Immunization Practices (ACIP) vaccine-selected candidates that were used during this time period. Three of the lead COBRA H3 HA antigens not only were able to elicit HAI activity against historical and future vaccine viruses but also neutralized the variant cocirculating strains. This is the first report demonstrating broader breadth of vaccine-induced antibodies against cocirculating H3N2 strains compared to the wild-type HA antigens represented in commercial influenza vaccines.
RESULTS
Characterization of H3 COBRA HA by phylogeny and predictive modeling.
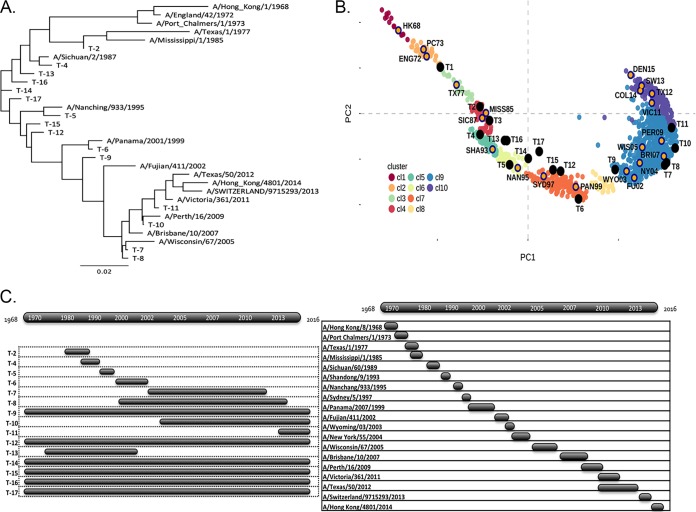
A multilayered consensus building approach was applied to 6,340 human H3N2 HA amino acid sequences to generate 17 unique COBRA HA sequences, of which 15 were further examined in this study. COBRA constructs were designed to cover multiple time periods and antigenic spaces based on date of isolation, with sequences extracted from as early as 1968 (Table 1). Consensus sequences were designed from HA sequences extracted from 3- to 12-year intervals and yielded COBRA constructs T-1 through T-11, except T-9. Six of the COBRA constructs were assembled using different strategies against the full known history of human H3N2, from 1968 to the first 3 months of 2013, including T-9, T-12, T-14, T-15, T-16, and T-17. All COBRA HA sequences were unique and did not have complete sequence identity to any of the vaccine strains. Constructs designed against a narrow period of time, such as 1998 to 2001 for T-6, share an ancestor node, such as Pan/99, with only 4 dissimilar amino acids (Fig. 1A). In contrast, the five COBRA constructs designed to span a large antigenic space between 1968 to 2013, including T-9, T-12, T-14, T-15, T-16, and T-17, all had a minimum of 16 unique residues from any of the wild-type strains, and subsequently they appear on separate branches distant from all other vaccines strains. When visualized as a principal-component analysis (PCA) graph, COBRA constructs were often arranged alongside the boundaries of the antigenic clusters, in contrast to WT vaccines that appear burrowed within the color-coded antigenic cluster (Fig. 1B). To illustrate the frequency of changes to the H3N2 component of the annual influenza vaccine, a timeline was constructed with 19 wild-type vaccines and the 15 COBRA constructs examined in this study (Fig. 1C). For the majority of seasons, a vaccine component was effective for a maximum of 3 to 4 years, such as Pan/99, but often a new recommendation was made each consecutive year.
TABLE 1.
Designations and descriptions of HA antigens used for vaccination and strain or antigenic space protein derived
| H3 COBRA designation | Description | Full name of virus | Abbreviation |
|---|---|---|---|
| T-2 | H3 antigenic space from 1980–1986 | A/Hong Kong/1/1968 | HK/68 |
| T-4 | H3 antigenic space from 1987–1992 | A/Port Chalmers/1/1973 | PC/73 |
| T-5 | H3 antigenic space from 1993–1997 | A/Mississippi/1/1985 × PR8 | Miss/85 |
| T-6 | H3 antigenic space from 1998–2001 | A/Sichuan/60/1989 × PR8 | Sich/89 |
| T-7 | H3 antigenic space from 2002–2010 | A/Nanchang/933/1995 | Nan/95 |
| T-8 | H3 antigenic space from 1999–2012 based on site B similarity | A/Panama/2007/1999 | Pan/99 |
| T-9 | H3 antigenic space from 1968–2013 (2nd order) | A/Fujian/411/2002 | Fuj/02 |
| T-10 | H3 antigenic space from 2002–2013 | A/Wisconsin/67/2005 | Wisc/05 |
| T-11 | H3 antigenic space from 2011–2013 | A/Brisbane/10/2007 | Bris/07 |
| T-12 | H3 antigenic space from 1968–2013 (3rd order) | A/Perth/16/2009 | Per/09 |
| T-13 | H3 antigenic space from 1975–2001 (1st order) | A/Victoria/361/2011 | Vic/11 |
| T-14 | H3 antigenic space from 1968–2013 | A/Texas/50/2012 | TX/12 |
| T-15 | H3 antigenic space from 1968–2013 | A/Switzerland/971593/2013 | Switz/13 |
| T-16 | H3 antigenic space from 1968–2013 | A/Hong Kong/4801/2014 | HK/14 |
| T-17 | H3 antigenic space from 1968–2013 | A/Denmark/32/2015 | Denm/15 |
FIG 1.
Characterization of the H3N2 influenza COBRA HA vaccines. (A) The unrooted phylogenetic tree was inferred from HA amino acid sequences derived from 17 representative H3N2 isolates and also the COBRA HA using the maximum likelihood method. Sequences were aligned with MUSCLE 3.7 software, and the alignment was refined by Gblocks 0.91b software. Phylogeny was determined using the maximum likelihood method with PhyML software. Trees were rendered using TreeDyn 198.3 software (46). (B) Genetic map of the HA of H3N2 viruses isolated between 1968 and 2015 that was generated from the numbers of amino acid substitutions between strains in the antigenic map. The two principal components, principal component 1 (PC1) and principal component 2 (PC2), are shown on the x and y axes. The clusters are colored according to the era of isolation. The spheres that represent each COBRA HA are shown in gold circles, and wild-type HA proteins are depicted as black dots. (C) Schematic of the timeline of H3N2 influenza infections in humans. Each bubble represents a time period. (Top) The time period that H3N2 HA sequences were used to develop each COBRA HA sequence. (Bottom) Time interval that each H3N2 vaccine component was used in the seasonal influenza inactivated vaccine over the past 48 years, from 1968 to 2016.
To assess whether sequence differences have the potential to alter the three-dimensional conformation of the HA molecule, predictive protein structure homology modeling was performed through the SWISS-MODEL web server (17–19). Utilizing the crystal structure template and diffraction data from a relatively contemporary H3N2 from 2004 (PDB entry 2YP2), predictive structures were assembled and further manipulated with MacPyMol (20), and several evident differences between three WT vaccines and COBRA T-7 were assessed (11, 21, 22). Comparison of Pan/99, Bris/07, and Vic/11 to COBRA T-7 suggests COBRA antigenic site A contains protein configurations that resemble portions of all three WT HA structures, while site B of T-7 COBRA appears most similar to site B of Bris/07 exclusively (data not shown). Antigenic site D of T-7, which includes the 220-loop and receptor-binding sites (RBS) conferring affinity to either avian or mammalian receptors (11), shares structural characteristics of Bris/07 and Vic/11 but not Pan/99 site D (data not shown). Additional comparisons were performed between COBRA candidates and the panel of H3N2 HA molecules listed in Fig. 1C, and multiple-sequence alignment demonstrates all COBRA sequences have at least one dissimilar residue within the antigenic sites A through E. Structure homology modeling and multiple-sequence alignment demonstrates COBRA constructs share some characteristics with WT vaccines, but not all five antigenic sites are identical to any one WT H3N2 HA; subsequently, the subtle but apparent differences in the antigenic sites have the potential to broaden recognition to a greater number of antigenically different H3N2 strains.
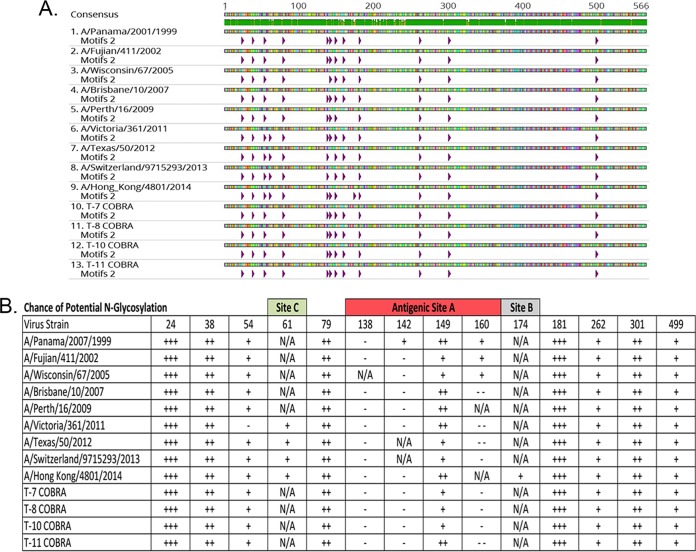
The putative glycosylation sequence motifs were compared between several H3 wild-type HA and H3 COBRA HA sequences (Fig. 2). COBRA HA antigens had numbers and patterns of putative glycosylation motifs similar to those of wild-type HA antigens from strains isolated between 1999 and 2014. All HA proteins analyzed had the same 11 glycosylation motifs out of 14 identified (Fig. 2A). The last four H3N2 vaccine strains had an additional glycosylation motif at amino acid 61 in antigenic site C. HA from Per/09 and HK/14 do not contain a glycosylation motif at amino acid 160 in antigenic site A, and HK/14 has an additional glycosylation site in antigenic site B (Fig. 2B).
FIG 2.
Schematics of putative glycosylation sites in selected H3 HA vaccines. (A) Nine wild-type H3 HA and four COBRA HA amino acids are depicted as putative N-linked glycosylation sites based upon the N{P}[ST]{P} search parameters, and these potential N-glycosylation sites were analyzed by using the NetNGlyc 1.0 server (37). Each putative N-glycosylation site is depicted with a purple arrow underneath the Arg amino acid in the HA sequence. (B) The N-glycosylation site predicted with a ++ or +++ score would be identified as a strong potential one with asparagine N-glycosylated. Low or no N-glycosylation sites, with +, −, or −− scores for N-glycosylation results, indicated low likelihood of an N-glycosylation site.
H3N2 VLP vaccines elicit distinct HAI antibody responses.
Each VLP vaccine had one of the HA proteins as well as a mismatched neuraminidase (N3) protein from the A/mallard/Alberta/24/2001 viral strain and the viral core HIV p55 Gag (23). All VLP vaccines were detectable by enzyme-linked immunosorbent assay (ELISA) for quantification using a broadly reactive, group 2 stalk-specific antibody, CR8020 (24). Each VLP agglutinated red blood cells, indicating functional HA conformation. All H3 VLPs produced moderate to high VLP yields, with endpoint HA titers ranging between 6,400 and 10,240 when expressed with mismatched neuraminidase from N3 (data not shown).
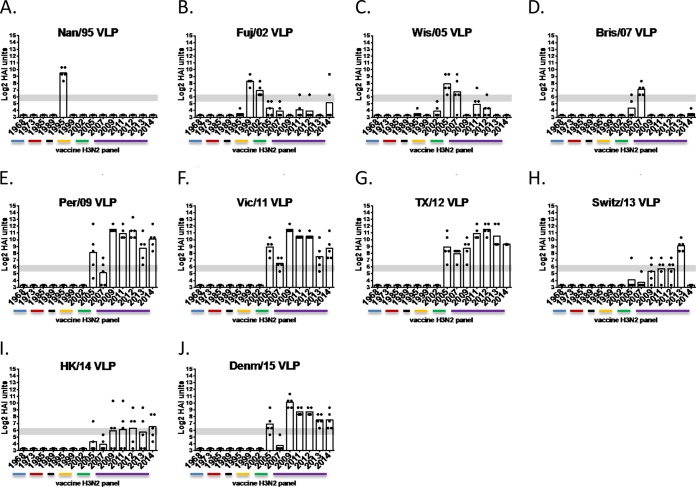
BALB/c mice were vaccinated with VLP vaccines or VLP vaccines expressing wild-type HA or COBRA HA proteins plus adjuvant, and the elicited antibodies were analyzed for HAI activity against a panel of H3N2 viral isolates representing vaccine strains from the past 48 years. Mice vaccinated with one of these 11 VLP vaccines that were produced using wild-type HA proteins had HAI activity primarily against the homologous strain with some cross-reactivity to a few other strains at an HAI titer of 1:80 (Fig. 3A). Mice vaccinated with VLP vaccines expressing the HA protein from either Per/09, Vic/11, or TX/12 elicited antibodies with HAI activity against a subset of vaccine viruses representing Wis/05 to TX/12 (Fig. 3E to G). Mice vaccinated with VLPs expressing the HA from Switz/13, a clade 3c3.a virus, had antisera with HAI activity homologous to that of Switz/13 virus, as well as Per/09, Vic/11, and TX/12 (Fig. 3H). However, the antisera failed to recognize the clade 3c2.a virus, HK/14. In addition, between 40 and 60% of the mice vaccinated with the HK/14 VLP vaccine had antibodies to Per/09, Vic/11, TX/12, and Switz/13 (Fig. 3I). In contrast, 80% of mice had antisera with HAI activity against the homologous HK/14 virus. Denma/15, a recent isolate of the subclade 3c3.b, elicited antibodies that recognized antigens spanning Wis/05 to HK/14 but failed to recognize vaccine strain Bris/07 (Fig. 3J). Similar results were observed in collected ferret sera following vaccination with a subset of wild-type and COBRA HA VLP vaccines using a similar vaccine regimen (data not shown).
FIG 3.
HAI serum antibody titers induced by vaccination of mice with wild-type H3N2 VLP vaccines. HAI titers were determined for each group of mice (n = 5) vaccinated three times (days 0, 28, and 56) with 1 of the 10 H3N2 VLP vaccines expressing surface wild-type HA proteins against a panel of 14 H3N2 influenza viruses. Values are the geometric mean titers plus standard errors of the means (SEM) (error bars) from antisera collected on day 70. The gray bar indicates the 1:40 to 1:80 HAI titer range. The strains are grouped by antigenic eras as depicted by colored bars on the x axis.
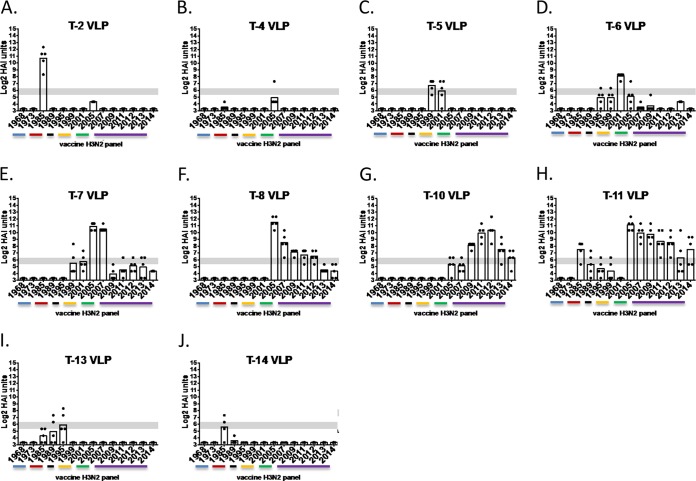
In contrast to animals vaccinated with VLPs expressing wild-type HA proteins, a second set of mice was vaccinated with VLPs expressing 1 of the 15 COBRA HA proteins; the HAI results of 10 of them are displayed (Fig. 4). Sera collected from mice vaccinated with T-2, T-4, T-5, or T-6 COBRA VLP vaccine had HAI activity against fewer than 3 H3N2 vaccine viruses in the 14-member vaccine virus panel. All animals vaccinated with T-6 seroconverted to Nan/95, Pan/99, and Fuj/02, except for one animal that had an HAI titer of 1:20 against Nan/95 (Fig. 4D). T-7 VLP elicited antibodies with HAI activity against Fuj/02, Wis/05, Bris/07, Per/09, Vic/11, and TX/12 (Fig. 4E). Both T-7 and T-8 VLP vaccines elicited HAI activity against the broadest number of H3N2 viruses in our panel. Mice vaccinated with T-10 COBRA HA VLP vaccine, for which the HA antigen was designed using HA sequences from H3N2 viruses isolated between 2002 and early 2013, had HAI activity against Per/09, Vic/11, and TX/12, with lower HAI activity (4 out of 5 mice with HAI titer of 1:40 or higher) against Wis/05, Bris/07, and Switz/13 (Fig. 4G). T-11 HA COBRA sequence was designed using HA sequences isolated between 2011 and early 2013 (Fig. 4H). Mice vaccinated with T-11 VLP vaccines had antibodies with HAI activity against Wis/05, Per/09, Vic/11, TX/12, and HK/14, but fewer mice had HAI activity against Switz/13 (Fig. 4H). Interestingly, COBRA HA antigens (T-9, T-12, T-15, T-16, and T-17) designed against the entire H3N2 virus era from 1968 to present failed to elicit robust HAI antibodies against any of the vaccine viruses in the panel (data not shown).
FIG 4.
HAI serum antibody titers induced by vaccination of mice with COBRA H3N2 VLP vaccines. HAI titers were determined for each group of mice (n = 5) vaccinated three times (days 0, 28, and 56) with 1 of the 17 H3N2 COBRA VLP vaccines against a panel of 14 H3N2 influenza viruses. Values are the geometric mean titers plus standard errors of the means (SEM) (error bars) from antisera collected on day 70. The gray bar indicates the 1:40 to 1:80 HAI titer range. The strains are grouped by antigenic eras as depicted by colored bars on the x axis.
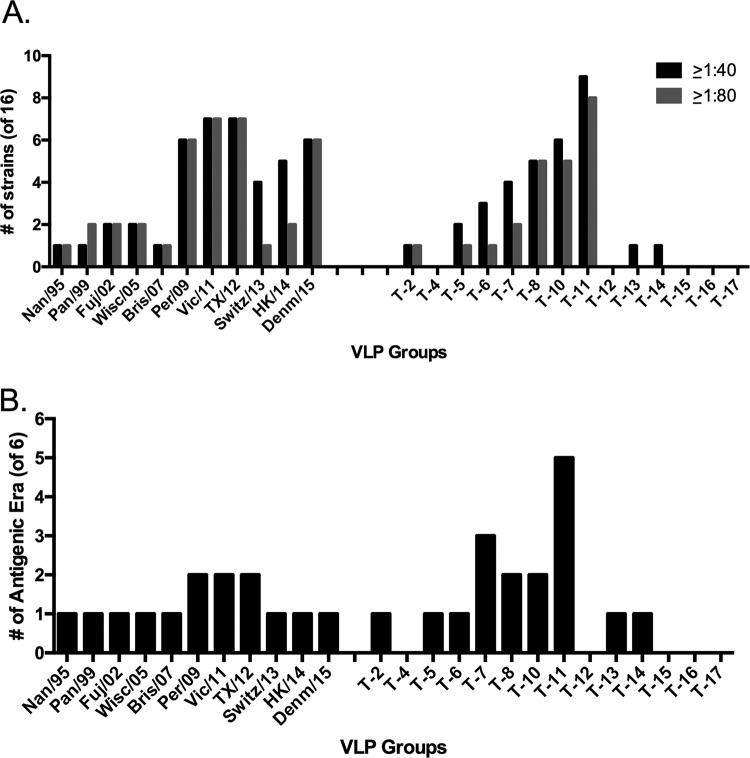
The number of viruses detected by antibodies elicited by each vaccine (geometric mean titer [GMT] of 1:40 or >1:80) in each group was determined (Fig. 5A). Mice vaccinated with Per/09, Vic/11, TX/12, or Denma/15 had antisera with HAI activity against the highest number of H3N2 vaccine strains in the panel. These vaccines elicited antibodies that recognized 6 to 7 viruses from the panel. In contrast, mice vaccinated with the T-11 COBRA vaccine had antibodies with HAI activity against 9 of the 12 representative vaccine strains since 1985 (last 30 years) in the panel, and T-8 or T-10 elicited HAI activity against 5 to 6 viruses in the panel. All of these COBRA vaccines elicited high HAI titers against themselves (≥1:640); therefore, any lack of response against the panel was not due to lack of antibody production (data not shown).
FIG 5.
Total number of strains and antigenic eras from vaccine panel recognized by the vaccine-elicited antibodies. Antisera collected from mice vaccinated with wild-type or COBRA VLP vaccines were tested for HAI activity. (A and B) Viral strain (A) or antigenic era of strains (B) detected by the antisera from each mouse in a vaccine group was considered positive only if all mice had antisera with a GMT greater than 1:40 or 1:80 and, for the antigenic era, a titer of 1:80. The strains detected were assessed as positive, and the number of strains in the panel inhibited at the indicated titers was recorded.
Viruses in the panel were categorized by antigenic eras based upon cross-reactive HAI activity elicited by the wild-type vaccine strains (Fig. 4). Each colored line underneath the x axis shows groups of viruses from similar antigenic eras. For example, the vaccine strains from 2009 to 2014 are antigenically similar. For some antigenic eras, the viral panel only contains one or two representative strains, with 6 different antigenic eras represented in this study (Fig. 4). The HAI activity elicited by the wild-type HA VLP and COBRA VLP vaccines was assessed by the number of antigenic eras in which the elicited antisera detected at least one virus. Mice vaccinated with wild-type HA VLP vaccines elicited antibodies with HAI activity that detected viruses in 1 to 2 antigenic eras (Fig. 5B). The COBRA HA vaccines T-7 and T-11 elicited antibodies that detected 3 or more antigenic eras, with T-11-elicited antibodies detecting viruses in 5 of the 6 antigenic eras (Fig. 5B).
COBRA HA antigens elicit antibodies with HAI activity against a broader number of antigenic drift variants than wild-type HA antigens.
For each influenza season, a set of wild-type viruses are selected representing one H1N1, one H3N2, and one to two influenza B isolates. These wild-type viruses and, subsequently, the HA associated with each virus are used by vaccine manufacturers to generate the seasonal influenza vaccine. However, the selected strain in the seasonal vaccine is similar to a percentage of virus variants in any given season. The ability of seasonal influenza vaccine to elicit antibodies that recognize all of the viral variants during an influenza season can vary widely in any given season. To address the ability of COBRA HA-elicited antisera to recognize not only viruses from the predominant cluster of viruses represented by the HA in the selected vaccine candidate but also the cocirculating viral variants, we tested antisera elicited by wild-type HA and COBRA HA VLP vaccines in mice.
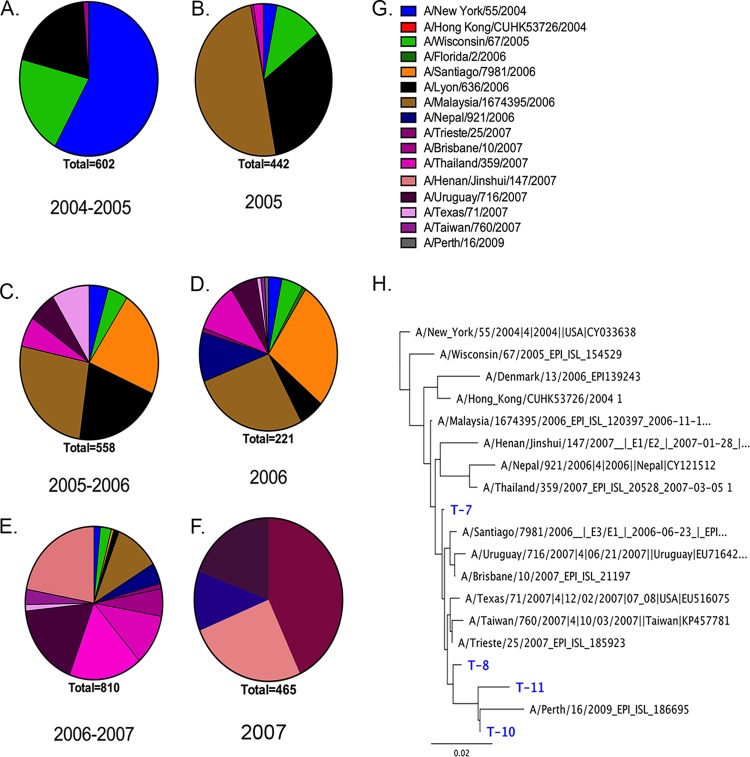
As a proof of concept, we selected the H3N2 era from 2004 to 2007 and identified HA sequences from five consecutive influenza seasons (three Northern and two Southern Hemisphere influenza seasons). A retrospective sequence analysis determined the frequency of related strains per season (Fig. 6), and representative influenza strains were selected for each antigenic cluster. During the 2004-2005 Northern Hemisphere influenza season, the majority of HA sequences collected were phylogenetically related to the 2005-2006 vaccine selected strain, NY/04 (Fig. 6A). However, the subsequent Southern Hemisphere season was dominated by strains related to the Malay/06 HA sequence and not the NY/04 HA sequence (Fig. 6B). Concurrently, a subdominant cluster emerged in 2005 that was phylogenetically similar to Denma/06 (Fig. 6C). Strains genetically related to Malay/06 and Denma/06 were present in the following 2005-2006 season; concurrently, a third cluster emerged consisting of strains similar to San/06 HA. A minor proportion of HA sequences in 2005 to 2006 contained HA sequences that were genetically similar to future 2007-like strains, such as Trie/07 and Thai/07. In 2006, strains related to San/06-like and Malay/06-like HA proteins cocirculated with other minor populations of strains that were genetically related to Nep/06 and Thai/07 (Fig. 6D). The 2006-2007 influenza season was composed of HA sequences that were phylogenetically related to the 2007 HA sequences in the fifteen-member cocirculating panel, including Thai/07, Hen/07, Uru/07, TX/07, and the 2008-2009 vaccine antigen, Bris/07 (Fig. 6E). A minor population of strains that were genetically related to Malay/06 and Nep/06 were present from 2006 to 2007, but other HA sequences from 2004 to 2006 were not represented in the sequences. Nep/06, Joh/08, Tiaw/07, and Uru/07 were the representative strains from the 4 dominant clusters in the 2007 influenza season (Fig. 6F).
FIG 6.
Frequencies of influenza HA clusters in consecutive seasons between 2004 and 2007. Influenza HA sequences posted in GISAID databases were aligned and clustered into family per season. Three Northern Hemisphere seasons (A, C, and E) and two Southern Hemisphere seasons (B, D, and F) that differ by 2% in amino acids in the HA sequence are depicted in clusters in each pie chart. (G) Representative influenza viruses from each cluster in the 5 pie charts are listed and match the color of the pie charts. (H) The unrooted phylogenetic tree was inferred from COBRA HA and HA amino acid sequences derived from the representative H3N2 variant viruses from seasonal clusters between 2004 and 2007 using the maximum likelihood method. Sequences were aligned with MUSCLE 3.7 software, and the alignment was refined by Gblocks 0.91b software. Phylogeny was determined using the maximum likelihood method with PhyML software. Trees were rendered using Geneious software (46).
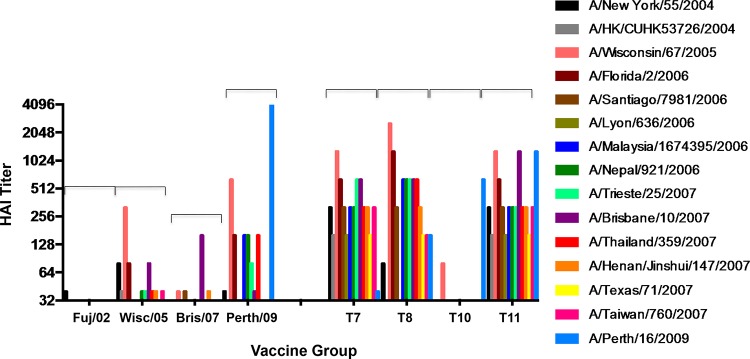
These comprehensive lists of representative sequences (Fig. 6G) were assessed for genetic relatedness to reference antigens, including the 12 drift viruses and the four recommended vaccine strains of those seasons, such as NY/04, Wis/05, Bris/07, and Per/09 (Fig. 6H). Mice were vaccinated three times with four wild-type HA VLP vaccines, Fuj/02, Wis/05, Bris/07, and Per/09, and the antibody-elicited HAI activity against a panel of vaccines and variant viruses that represent the 2004-2007 era was compared to that of antisera elicited from mice vaccinated with 1 of 4 COBRA HA VLP vaccines, T-7, T-8, T-10, and T-11 (Fig. 7). Wis/05 or Bris/07 vaccine elicited antibodies with HAI activity against a portion of the viruses in the panel, with the highest HAI titers against each homologous virus. Wis/05 VLP-vaccinated mice had HAI titers greater than 1:80 against 4 of the 16 viruses in the panel, and Bris/07-elicited antisera recognized 2 of the viruses in the panel. Mice vaccinated with Per/09 had HAI activity against 7 of 16 viruses. In contrast, mice vaccinated with T-11 had antisera with HAI activity against all 16 viruses in the panel, with the lowest HAI titer being 1:160. Mice vaccinated with T-7 or T-8 had antisera with HAI activity against 14 to 15 of the viruses in the panel. Interestingly, mice vaccinated with T-10 had antisera that was highly restrictive and only recognized 2 of the vaccine strains in the 16-member panel.
FIG 7.
H3N2 influenza drift variants circulating between 2004 and 2007. Pooled mouse sera from mice vaccinated with one of four wild-type HA VLP vaccines (Fuj/02, Wisc/05, Bris/07, and Per/09) and four COBRA HA VLP vaccines (T-7, T-8, T-10, and T-11) were tested for HAI activity against variant viruses representing different clusters of influenza H3N2 isolates, as depicted in Fig. 6. Values are the average titers from antisera collected on day 70 against the panel of 15 vaccine and variant H3N2 viruses, representing four vaccine and 11 variant strains isolated from 2006 and 2007.
H3 COBRA HA antigens elicit antibodies that are more effective at neutralizing viral infection against a broader number of antigenic drift variants than wild-type H3 HA antigens.
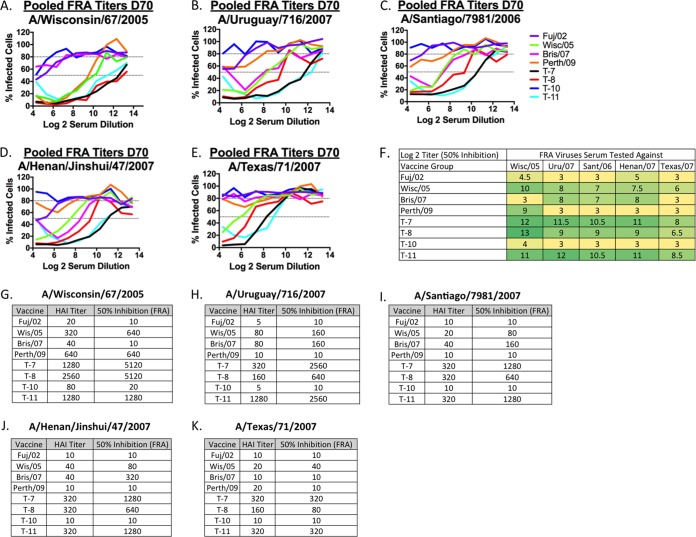
To evaluate the ability of the elicited antibody response to block live virus infection, serum was tested in a focal reduction assay (FRA). Similar to the HAI results, mice vaccinated with T-7 and T-11 COBRA VLP vaccines had high levels of neutralizing antibodies at 50% inhibition compared to mice receiving mock vaccination (Fig. 8). For example, T-7 and T-11 COBRA VLP vaccines elicited antibodies that neutralized viral infection at a log2 serum dilution of 11 to 12 (Fig. 6B and F). This neutralization titer was ∼3 logs higher than that of the antibodies elicited by the wild-type HA antigens from Fuj/02 or Per/09 (log2 serum dilution titer of 3). Consistently, the T-7 and T-11 COBRA vaccines outperformed wild-type HA VLP vaccines, and in most instances T-8 COBRA was superior to the wild-type HA VLP vaccines. There was consistency in the HAI and FRA results against two vaccine strains, Wisc/05 and Uru/07, and three of the variant viruses, Sant/06, Hen/07, and TX/07 (Fig. 6G to K).
FIG 8.
Focal reduction assay (FRA). BALB/c mice were vaccinated at days 0, 28, and 56 with VLPs expressing WT HA antigens from Fuj/02, Wisc/05, Bris/07, or Per/09 or COBRA HA T-7, T-8, T-10, and T-11 antigens. At day 70, sera were collected and tested in an HAI assay and the FRA. (A to E) For each virus, the virus concentration was standardized to 1.2 × 104 FFU/ml (corresponding to 600 FFU/50 μl, which is the volume of virus added to each plate). A monolayer of MDCK SIAT cells (2.5 × 105 to 3 × 105 cells/ml) (100 μl/well in a 96-well plate) was added the day before the assay is run. Cells were 95 to 100% confluent at the time of the assay to determine the number of foci detected as percent infected cells normalized to 100%. Pooled sera from each group of mice were tested against two vaccine strains, Wisc/05 (A) and Uru/07 (B), and three deep-dive viruses, Sant/06 (C), Henan/06 (D), and TX/06 (E). The dotted lines represent 50% and 80% inhibition by sera compared to virus-only control wells. (F) A heat map of the log2 serum dilution titer for 50% inhibition per virus for each vaccine. (G to K) The GMT HAI and FRA (50% inhibition) titers for each vaccine listed for Wisc/05 (G), Uru/07 (H), Sant/06 (I), Henan/06 (J), and TX/06 (K) viruses.
DISCUSSION
There is an urgent need to develop an influenza vaccine that elicits protective immune responses against a broader range of influenza subtypes and variant isolates compared to currently licensed influenza vaccines. Several vaccine approaches are in development to elicit a broadly protective immune response with enhanced potency and durability against both seasonal and prepandemic influenza subtypes (25, 26). The elicitation of HAI antibodies is currently a surrogate marker for protection against influenza in humans vaccinated with seasonal influenza vaccines. However, the vaccine formulation has to be updated annually to address the evolution of influenza viruses each season. It is currently impossible to predict which antigenic variants may evolve; therefore, novel vaccine candidates are needed that will elicit immune responses to potential variants and cocirculating strains. Recently, we designed a new technology, termed COBRA, as a more broadly reactive vaccine approach for both emerging/prepandemic influenza viruses and currently circulating drifting influenza subtypes. The COBRA method uses multiple rounds of consensus building based upon not only the phylogenetic sequence of each isolate but also the outbreak and specific time that each isolate was collected, thereby eliminating the bias in the number of sequences uploaded to online databases. We have published a series of studies demonstrating the effectiveness of the COBRA HA antigens for H5N1 (13–15). We also demonstrated that the COBRA approach was an effective strategy for developing an H1N1 seasonal influenza vaccine (27). These H1N1 COBRA HA proteins were tested in mice using a virus-like particle (VLP) format that elicited antibodies with HAI activity against a range of H1N1 viruses spanning the last 100 years. In addition, these vaccinated mice were protected against H1N1 influenza virus challenge.
The development of a broadly reactive influenza vaccine that can protect and prevent transmission against all influenza A and influenza B viruses would be an ultimate achievement of the current universal influenza vaccine initiatives and will have a great impact on human health (for reviews, see references 25, 26, and 28). However, this may be an unrealistic goal for the foreseeable future. Therefore, various research groups are examining approaches to develop an influenza A-only vaccine or subtype-specific influenza vaccines. Both approaches offer challenges. Our group has recently demonstrated that a subtype-specific vaccine that elicits antibodies with HAI activity against both historical and future influenza strains, covering many HA antigenic types, is possible (16). The COBRA methodology refocuses the HA immunogen in order to elicit functional antibodies that are broad in coverage within a subtype. COBRA HA-elicited antibodies have HAI activity against the receptor binding site, which has benefits over other universal vaccine candidates. Induction of HAI antibodies is accepted as a surrogate/correlate for protective efficacy in humans (29). Therefore, COBRA-based vaccine candidates have a clear path forward to clinical proof of concept, unlike many other universal vaccine candidates.
In this report, we applied COBRA methodology to develop 17 COBRA HA prototype antigens against the H3N2 influenza subtype. Mice vaccinated with some of these COBRA vaccines had antibodies that recognized a broad range of human seasonal H3N2 viruses (T-7, T-8, T-10, and T-11), whereas other COBRA vaccines elicited antibodies that recognized a set of viruses from a narrow era (T-2, T-5, T-6, and T-7). The five COBRA constructs (T-9, T-12, T-13, T-14, and T-15) designed with sequences from the entirety of H3N2 influenza viruses isolated from 1968 to 2013 failed to elicit HAI-specific antibodies to more than one virus in the panel.
Overall, designing a broadly reactive influenza vaccine for H3 influenza viruses provided different challenges and opportunities compared to those of our previous COBRA vaccines design and assessments. First, there have been more antigenic changes in human H3N2 influenza viruses over the last 48 years than in H1N1 influenza viruses. This resulted in the selection of more H3N2 viral vaccine candidates that were used for fewer seasons than H1N1 viruses (Fig. 1C). This is an indication of the more rapid evolution and immune escape of H3N2 influenza viruses compared to those of other influenza A or influenza B viruses. Therefore, a COBRA vaccine design needed to take into account the development of vaccines that would recognize viruses representing multiple antigenic changes over a shorter period of time than H1N1 viruses. For example, the H3N2 vaccine component for the Northern Hemisphere changed 8 times between the 2005-2006 and 2016-2017 seasons. In contrast, the H1N1 vaccine component was changed 4 times over this same 12-year period.
We hypothesize that COBRA HA antigens retain immunodominant epitopes that are represented on viruses from different antigenic eras. When these are used as immunogens, COBRA HA antigens elicit a pool of polyclonal antibodies that can neutralize a broader number of viral strains, most likely by binding to a combination of HA epitopes. While any two wild-type HA antigens may have different epitopes, the COBRA antigens elicit antibodies to all of the potential epitopes and therefore can neutralize many H3N2 viruses from different antigenic eras. The pool of antibodies elicited by any given wild-type HA antigen is restricted to a narrow set of HA epitopes found only on a subset of circulating viruses. Therefore, COBRA vaccines can neutralize more variant wild-type H3N2 influenza viruses than the antibodies elicited by wild-type HA antigens. We cannot rule out that both head-based antibodies, which interfere with HA receptor binding (30, 31), and stalk-directed antibodies (32–34) are enhanced by COBRA designs. However, since there was a consistency between the FRA-elicited titers and the HAI-elicited titers, we can conclude that the antibodies elicited by COBRA vaccines are directed predominantly against the HA head, and that neutralization of these diverse strains is due to interference of receptor binding. Further assessment of monoclonal antibodies induced by COBRA HA antigens will be needed to map specifically which combinations of epitopes are bound by antibodies elicited by COBRA vaccines.
Three lead candidates, T-8, T-10, and T-11, were able to elicit antibodies that recognized a set panel of H3N2 viruses representing vaccine strains from 2004 to 2016 with similar efficiency (Fig. 4). Therefore, the various cocirculating strains during the three Northern and three Southern Hemisphere seasons between 2004 and 2007 were examined, and wild-type H3N2 viruses were selected as representative viruses for each cluster of H3N2 viruses cocirculating at that time. Since some clusters carried over from season to season, we were able to identify a panel of 13 cocirculating H3N2 viruses to represent this 3-year period, along with 4 vaccine strains selected for inclusion in the commercial vaccine (Fig. 7). One vaccine strain, A/Perth/16/2009, is also listed as the next vaccine strain that is not included in the cluster charts. There were clear differences in HAI activity against these cocirculating strains during this period. The antigenic match between the vaccine and cocirculating strains during the 2004-2005, 2005-2006, and 2006-2007 Northern Hemisphere seasons ranged between 5%, 5%, and 91% for the H3N2 component of the vaccine (7). In general, we confirmed that the three wild-type H3N2 strains that were used in the vaccine during this time, Fuj/02, Wis/05, and Bris/07, elicited antibodies with HAI activity against the homologous strain and low titers to most of the other vaccine and cocirculating H3N2 viral strains during this time period (Table 2). In contrast, the T-7 and T-11 COBRA vaccines were highly effective at eliciting antibodies with high-titer HAI activity, as well as FRA activity, against all of these cocirculating strains, with T-8 recognizing ∼85% of the H3N2 viruses in the panel. Interestingly, T-10 and T-11 elicited antibodies with HAI activity against the vaccine strains during that time period. However, T-10 failed to recognize any of the cocirculating variant viruses, whereas T-11-elicited antibodies recognized all of them. T-10 and T-11 COBRA HA antigens differed by only 4 amino acids, which demonstrates that minor sequence variations can have major consequences in HA structure and vaccine design. Future studies will be needed to determine the exact structural differences between T-10 and T-11 and where antibodies bind on the wild-type HA molecules that are elicited by these two COBRA HA antigens.
TABLE 2.
Hemagglutination inhibition titers from WTor COBRA-vaccinated mice against a panel of variants from 2004 to 2007

Various clusters of H3N2 viruses carry over from season to season, and the number of viruses within each of these clusters increases or decreases over this 3-year period. For example, the cluster of H3N2 viruses represented by A/Denmark/13/2006 represented 30% of the cocirculating viruses in the 2004-2005 Northern Hemisphere season and began to decrease 3 to 4 seasons later (Fig. 7). In contrast, the cluster represented by A/Thailand/359/2007 or A/Uruguay/716/2007 was circulating at low levels in the 2004-2005 Northern Hemisphere season, but viruses from these two clusters steadily increased over time and became ∼25% of the cocirculating strains, representing ∼50% of all H3N2 viruses circulating in the 2006-2007 Northern Hemisphere season. In fact, A/Uruguay/716/2007 was selected as one of the potential vaccine strains for the 2006-2007 season. Therefore, while the commercial vaccine had to change the H3N2 component 3 times over those six seasons, the T-11 COBRA HA antigen, for example, would have been able to elicit antibodies with HAI activity to protect humans from infection during this same time period. In most seasons, the virus that the World Health Organization recommended for the H3N2 vaccine strain did not come from a cluster that included the majority of the circulating viruses (Fig. 7). In most cases, that cluster had a plurality of the cocirculating strain, at best. Since both T-7 and T-11 recognized all of the viruses in the panel, if people had been vaccinated with either of these COBRA vaccines, then they already could have possessed antibodies that would have protected them against the emergence of these future dominant H3N2 viruses. In reality, three new commercial vaccines were produced. Therefore, by protecting against all of the minor cocirculating drift variants in one season, the COBRA vaccines are able to protect against future dominant clusters.
Ideally, influenza vaccines should elicit antibodies to both commonly shared as well as new epitopes, thereby neutralizing more new variant viruses in circulation. Currently, most universal influenza vaccines in development afford protection by disease modulation with potential for impacting transmission, but these may not prevent influenza virus infection and therefore are unlikely to replace current licensed vaccine products that elicit neutralizing antibody responses. The generation of subtype-specific universal or broadly protective influenza vaccines (BPIV) capable of eliciting potent, broadly cross-reactive HA-specific functional antibodies (HAI) can broadly protect against both seasonal and potentially novel pandemic influenza strains that have undergone genetic drift. Ultimately, establishing long-lived COBRA-induced immune responses that protect against cocirculating strains for multiple seasons would be a paradigm shift. Establishing memory B cells that secret antibodies in response to COBRA vaccination initially may take multiple doses or adjuvants when first used in humans. However, since humans have preexisting memory B cells, the COBRA vaccines may also recall more memory B cells secreting antibodies with a broader recognition of influenza HA epitopes on multiple HA proteins. Future studies to test these hypotheses in humans will be necessary to answer many of these questions. This is the first report to demonstrate a broadly reactive influenza vaccine that elicits antibodies with broad activity against cocirculating variants within a season and over several seasons.
MATERIALS AND METHODS
Antigen construction and synthesis.
Influenza A HA protein sequences from 6,430 human H3N2 infections collected from 1 January 1968 to 15 December 2013 were downloaded from the Global Initiative on Sharing Avian Influenza Data (GISAID) database (see the supplemental material for a complete list of accession numbers and isolate descriptions) and organized by the year of collection. For each round of consensus generation, multiple-sequence alignment was performed using the Geneious MUSCLE alignment method, and phylogenetic neighbor-joining trees were constructed such that trees were rooted to the oldest sequences collected in 1968. Full-length HA sequences were aligned, and the most common residue found among a designated set of viruses was used to yield the primary consensus sequence. Ambiguities were avoided through alignment of an odd number of sequences.
Multiple rounds of consensus assembly were layered to yield secondary, tertiary, and quaternary consensus sequences that were designed to represent different antigenic spaces that overlapped periods of time for which wild-type vaccine strains were recommended from 1968 to 2013. The final amino acid sequences then were renamed T-series computationally optimized broadly reactive antigens (COBRA). Fifteen H3N2 HA constructs were synthesized and inserted into the pTR600 expression vector, as previously described (35). We designed COBRA HA antigens to represent different antigenic spaces. For example, constructs T-2 through T-7 were designed to represent nonoverlapping periods of time, spanning 4 to 6 years, using H3 sequences from human isolates collected between 1980 and 2010, and T-11 was generated using HA sequences isolated from 2011 to 2013. The COBRA HA constructs T-9, T-12, T-13, T-14, T-15, T-16, and T-17 represent an antigenic space from 1968 to 2013 using various multiconsensus layering approaches.
Cartography and predictive structure-homology modeling methods.
Human influenza A (subtype H3N2) hemagglutinin protein sequences were obtained from the NCBI Influenza Virus Resource database and trimmed to remove signal peptides, transmembrane regions, and cytoplasmic tails, and the resulting ectodomain sequences were aligned using MAFFT (35). The pairwise dissimilarity matrix was calculated from the multiple-sequence alignment based on the Hamming distance between pairs of sequences with no prior assumptions regarding the function or structure of the sequences. Principal-component analysis (PCA) was applied to the dissimilarity matrix for the purpose of dimension reduction and to facilitate visualization of the relative distances between HA proteins. The first two or three principal components were retained for visualizing protein relationships in sequence space and represent a reasonable approximation of the general structure of the phylogenetic tree. Calculations were performed using custom scripts written in python and R.
Each COBRA HA predictive structure was generated using the SWISS-MODEL algorithm (17–19), implementing PDB entry 2yp2.1.a, the crystal structure of a human H3N2 2004 virus (11), as the HA template, and renderings were performed using MacPyMol. Antigenic sites A through E were color coded, and residues were based on references that have identified and characterized antigenic residues based on monoclonal antibody binding and/or mass spectrometry (11, 21, 22).
Vaccine preparation.
Mammalian 293T cells were transfected with each of three plasmids expressing the influenza neuraminidase (A/mallard/Alberta/24/01, H7N3), the HIV p55 Gag sequences, and one of the various H3N2 wild-type or COBRA HA-expressing plasmids on previously described mammalian expression vectors (36). Following 72 h of incubation at 37°C, supernatants from transiently transfected cells were collected, centrifuged to remove cellular debris, and filtered through a 0.22-μm pore membrane. Mammalian virus-like particles (VLPs) were purified and sedimented by ultracentrifugation on a 20% glycerol cushion at 135,000 × g for 4 h at 4°C. VLPs were resuspended in phosphate-buffered saline (PBS), and total protein concentration was assessed by conventional bicinchoninic acid assay (BCA). Hemagglutination activity of each preparation of VLPs was determined by adding equal volumes of turkey red blood cells (RBCs) to a V-bottom 96-well plate and incubating with serially diluted volumes of VLPs for a 30-min incubation at room temperature. The highest dilution of VLP with full agglutination of RBCs was considered the endpoint HA titer.
Determination of HA content.
A high-affinity, 96-well flat-bottom ELISA plate was coated with 5 to 10 μg of total protein of VLP and serial dilutions of a recombinant H3 antigen (3006_H3_Vc; Protein Sciences, Meriden, CT) in ELISA carbonate buffer (50 mM carbonate buffer, pH 9.5), and the plate was incubated overnight at 4°C on a rocker. The next morning, plates were washed in PBS with 0.05% Tween 20 (PBST), and then nonspecific epitopes were blocked with 1% bovine serum albumin (BSA) in PBST solution for 1 h at room temperature. Buffer was removed, and then stalk-specific group 2 antibody CR8020 (24) was added to the plate and incubated for 1 h at 37°C. Plates were washed and then probed with goat anti-human IgG horseradish-peroxidase-conjugated secondary antibody (2040-05; Southern Biotech, Birmingham, AL) for 1 h at 37°C. Plates were washed, and then freshly prepared o-phenylenediamine dihydrochloride (OPD) (P8287; Sigma) substrate in citrate buffer (P4922; Sigma) was added to wells, followed by 1N H2SO4 stopping reagent. Plates were read at 492-nm absorbance using a microplate reader (Powerwave XS; Biotek, Winooski, VT), and background was subtracted from negative-control wells. Linear regression standard curve analysis was performed using the known concentrations of recombinant standard antigen to estimate HA content in VLP lots.
NGSs and antigenic epitope analysis.
The potential N-glycosylation sites (NGSs) (Asn-X-Ser/Thr, where X is any amino acid except Pro) were analyzed using the NetNGlyc 1.0 server (http://www.cbs.dtu.dk/services/NetNGlyc/). The scores of two pluses (++) or three pluses (+++) in the NetNGlyc results were classified as strong potential NGSs (having strong potentiality for N-glycosylation). Based on the server-based analysis, ∼76% of positively scored loci are modified by N-glycans that contain Thr (37, 38). The HA amino acid sequences from 1 to 566 were annotated by the predict function in Geneious 10, searched for the N(6)[ST]{P} motif, and Geneious 10 populates these putative glycosylation sites onto the schematic, depicted with purple arrows.
Mouse and ferret studies.
BALB/c mice (Mus musculus, females, 6 to 8 weeks old) were purchased from Jackson Laboratory (Bar Harbor, ME, USA), housed in microisolator units, and allowed free access to food and water. They were cared for under University of Georgia Research Animal Resources guidelines for laboratory animals. All procedures were reviewed and approved by the Institutional Animal Care and Use Committee (IACUC). Mice (5 mice per group) were vaccinated with purified VLPs (3.0 μg/mouse) based upon HA content from the ELISA quantification, and vaccines were delivered via intramuscular injection at week 0 and then boosted with the same vaccine at the same dose at weeks 4 and 8. Vaccines at each dose were formulated with an emulsified squalene-in-water adjuvant (Sanofi Pasteur, Lyon, France). The final concentration after mixing 1:1 with VLPs is 2.5% squalene. Twenty-eight days after each vaccination, blood samples were collected via the submandibular cheek and transferred to a microcentrifuge tube. The tubes were centrifuged at 10,000 rpm for 10 min, and serum samples were removed and frozen at −20°C ± 5°C.
Fitch ferrets (Mustela putorius furo, female, 6 to 12 months of age), negative for antibodies to circulating influenza A (H1N1 and H3N2) and influenza B viruses, were descented and purchased from Triple F Farms (Sayre, PA). Ferrets were pair housed in stainless steel cages (Shor-line, Kansas City, KS) containing Sani-Chips laboratory animal bedding (P. J. Murphy Forest Products, Montville, NJ). Ferrets were provided with Teklad global ferret diet (Harlan Teklad, Madison, WI) and fresh water ad libitum. Ferrets (n = 4) were vaccinated with the H3 COBRA VLP or wild-type H3 HA VLP vaccines or phosphate-buffered saline alone as a mock vaccination. Vaccines (15-μg dose based upon HA content) were formulated with an emulsified squalene-in-water AF03 adjuvant (Sanofi Pasteur, Lyon, France) in a final 1:1 mixture with VLPs. Ferrets were boosted 28 days later. Blood was harvested from all anesthetized ferrets via the anterior vena cava at days 14, 28, and 56. Serum was transferred to a centrifuge tube and centrifuged at 6,000 rpm. Clarified serum was removed and frozen at −20 ± 5°C. Animals were monitored daily during the infection for adverse events, including weight loss, loss of activity, nasal discharge, sneezing, and diarrhea.
HAI assay.
The hemagglutination inhibition (HAI) assay was used to assess functional antibodies to the HA able to inhibit agglutination of guinea pig and turkey erythrocytes. The protocols were adapted from the WHO laboratory influenza surveillance manual (39) and used the host species that is frequently used to characterize contemporary H3N2 strains that have preferential binding to alpha-(2,6)-linked sialic acid receptors (40, 41). We also used turkey erythrocytes to compare whether there was a differential in HAI depending on erythrocyte used. To inactivate nonspecific inhibitors, sera were treated with receptor-destroying enzyme (RDE) (Denka Seiken, Co., Japan) prior to being tested. Briefly, three parts RDE were added to one part serum and incubated overnight at 37°C. RDE was inactivated by incubation at 56°C for ∼30 min. RDE-treated sera were diluted in a series of 2-fold serial dilutions in V-bottom microtiter plates. An equal volume of each H3N2 virus, adjusted to approximately 8 hemagglutination units (HAU)/50 μl, was added to each well. The plates were covered and incubated at room temperature for 20 min, and then 0.8% guinea pig erythrocytes (Lampire Biologicals, Pipersville, PA, USA) in PBS were added. Red blood cells were stored at 4°C and used within 72 h of preparation. The plates were mixed by agitation and covered, and the RBCs were allowed to settle for 1 h at room temperature. The HAI titer was determined by the reciprocal dilution of the last well that contained nonagglutinated RBCs. Positive and negative serum controls were included for each plate. All mice were negative (HAI of ≤1:10) for preexisting antibodies to currently circulating human influenza viruses prior to vaccination, and seroprotection was defined as an HAI titer of >1:40 and seroconversion as a 4-fold increase in titer compared to the baseline per the WHO and European Committee for Medicinal Products to evaluate influenza vaccines (42); however, we often examined a more stringent threshold of >1:80. Mice are naive and seronegative at the time of vaccination; thus, seroconversion and seroprotection rates are interchangeable in this study.
FRA.
The focus reduction assay (FRA) used in this study was initially developed by the World Health Organization Collaborating Centre in London, United Kingdom (43, 44), and modified by the U.S. Centers for Disease Control and Prevention (CDC) (Thomas Rowe, personal communication). MDCK-SIAT1 cells were cells plated at 2.5 × 105 to 3 × 105 cells/ml (100 μl/well in a 96-well plate) the day before the assay was run. Cells need to be 95 to 100% confluent at the time of the assay in 96-well plates overnight to form a confluent monolayer in Dulbecco's modified Eagle medium (DMEM) containing 5% heat-inactivated fetal bovine serum and antibiotics. The following day, the cell monolayers were rinsed with 0.01 M PBS, pH 7.2 (Gibco), followed by the addition of 2-fold serially diluted RDE-treated serum at 50 μl per well, starting with a 1:20 dilution in virus growth medium containing 1 μg/ml tosylsulfonyl phenylalanyl chloromethyl ketone (TPCK)-treated trypsin, termed VGM-T (DMEM containing 0.1% BSA, penicillin-streptomycin, and 1 μg/ml TPCK-treated trypsin [Sigma, St. Louis, MO, USA]). Afterwards, 50 μl of virus for all FRAs shown here was standardized to 1.2 × 104 focus-forming units (FFU)/ml (corresponds to 600 FFU/50 μl), and VGM-T was added to each plate or cell control wells. The virus stocks were standardized by previous titration in the FRA. Following 2 h of incubation at 37°C with 5% CO2, the cells in each well were overlaid with 100 μl of equal volumes of 1.2% Avicel RC/CL (43) (type RC581 NF; FMC Health and Nutrition, Philadelphia, PA, USA) in 2× modified Eagle medium containing 1 μg/ml TPCK-treated trypsin, 0.1% BSA, and antibiotics. Plates were incubated for 18 to 22 h at 37°C, 5% CO2. The overlays then were removed from each well and the monolayer was washed once with PBS to remove any residual Avicel. The plates were fixed with ice-cold 4% formalin in PBS for 30 min at 4°C, followed by a PBS wash and permeabilization using 0.5% Triton X-100 in PBS-glycine at room temperature for 20 min. Plates were washed three times with wash buffer (PBST) and incubated for 1 h with a monoclonal antibody against influenza A nucleoprotein (45) (IRR) in ELISA buffer (PBS, 10% horse serum, 0.1% Tween 80). Following washing three times with PBST, the cells were incubated with goat anti-mouse peroxidase-labeled IgG (SeraCare, Inc., Milford, MA) in ELISA buffer for 1 h at RT. Plates were washed three times with PBST, and infectious foci (spots) were visualized using TrueBlue substrate (SeraCare, Inc., Milford, MA USA) containing 0.03% H2O2 and incubated at room temperature for 10 to 15 min. The reaction was stopped by washing five times with distilled water. Plates were dried and foci enumerated using a CTL BioSpot analyzer with ImmunoCapture 6.4.87 software (CTL, Shaker Heights, OH). The FRA titer was reported as the reciprocal of the highest dilution of serum corresponding to 50% focus reduction compared to the virus control minus the cell control.
In order for a plate to pass quality control, both the average of the octuplet virus control wells (VC) and the average of the octuplet cell control wells (CC) must pass. The virus controls initially were between 150 and 650 foci (spots), and the cell controls must be less than 21 foci. The virus control wells subsequently were expanded to between 200 and 1,600 spots. Additionally, the reference vaccine strain virus was run in triplicate plates in each individual assay, and at least two out of three plates must pass VC and CC criteria and homologous ferret antisera must have the same titer. Each assay plate (one virus per plate) contained a panel of reference antisera as well as human vaccine serum control to assess overall assay consistency.
Viruses and HA antigens.
H3N2 viruses were obtained through the Influenza Reagents Resource (IRR), BEI Resources, the Centers for Disease Control (CDC), or were provided by Sanofi-Pasteur. Viruses were passaged once under the same growth conditions as those under which they were received, in either embryonated chicken eggs or semiconfluent Madin-Darby canine kidney (MDCK) cell culture per the instructions provided by the WHO (39). Titers of virus lots were determined with both guinea pig and turkey erythrocytes, and they were made into aliquots for single-use applications. The H3N2 vaccine panel includes the following strains: A/Hong Kong/1/1968 (HK/68) (NR-28620), A/Port Chalmers/1/1973 (PC/73), A/Mississippi/1/1985 × PR/8 (Miss/85) (NR-3502), A/Sichuan/60/1989 × PR/8 (Sic/89) (NR-3492), A/Nanching/933/1995 (Nan/95), A/Sydney/05/1997 (Syd/97), A/Panama/2007/1999 (Pan/99), A/Fujian/411/2002 (Fuj/02), A/New York/55/2004 (NY/04) (FR-462), A/Wisonsin/67/2005 (Wis/05) (FR-397), A/Brisbane/10/2007 (Bris/07), A/Perth/16/2009 (Per/09), A/Victoria/361/2011 (Vic/11), A/Texas/50/2012 (TX/12), and A/Switzerland/9715293/2013 (J1506B) (Switz/13), A/Hong Kong/4801/2014 (HK/14), and A/Denmark/32/2015 (Denma/15). All viruses were characterized for the ability to agglutinate turkey and guinea pig erythrocytes. The 14-member panel included all antigens except for Syd/97 and NY/04, which are only included in the abbreviated vaccine panel spanning 1995 to 2013. A panel of 12 cocirculating H3N2 variants from the period of 2004 to 2007 were obtained through BEI and IRR and included A/Nepal/921/2006 (Nep/06), A/Florida/2/2006 (FL/06), A/Henan/Jinshui/147/2007 (Hen/07), A/Santiago/7981/2006 (San/06), A/Texas/71/2007 (TX/07), A/Uruguay/716/2007 (Uru/07), and A/Taiwan/760/2007 (Tai/07). When viruses could not be obtained, H3-A/Thailand/1/2004 N1-Gag VLPs were synthesized using codon-optimized HA sequences. This included the following antigens for the 2004-2007 drift panel: A/Hong Kong/CUHK53726/2004 (HK/04), A/Denmark/13/2006 (Denm/06), A/Malaysia/1674395/2006 (Malay/06), A/Thailand/359/2007 (Thai/07), and A/Trieste/25/2007 (Trie/07).
Phylogenetic comparison of cocirculating variants within an influenza season (2004 to 2007).
Multiple-sequence alignments were performed on extracted HA sequences that retained the majority of the HA1 domain; sequences were organized based on collection during the Northern Hemisphere season or the Southern Hemisphere season. Neighbor-joining Juke-Cantor phylogenetic tree models were assembled using Geneious, and branches were compared for their sequence similarity to a panel of 15 drift viruses described above in “Viruses and HA antigens.” The number of sequences that branched within 98.9% HA1 sequence identity of 1 of the 15 antigens were counted, and frequencies were calculated based on the total number of sequences available per season.
ACKNOWLEDGMENTS
T.A., J.D., and H.K. are full-time employees and shareholders of Sanofi Pasteur.
We thank Maryann Giel-Moloney, Raul Gomila, Kirill Kalnin, Tod Strugnell, and Thortsen Vogel for helpful discussions and reviews. We also thank Maryann Giel-Moloney, Arnel Sicam, and Benjamin Park for technical assistance. The H3N2 influenza A viruses were obtained through the Influenza Reagent Resource, Influenza Division, WHO Collaborating Center for Surveillance, Epidemiology, and Control of Influenza and the Centers for Disease Control and Prevention, Atlanta, GA, USA. We thank the University of Georgia Animal Resource staff, technicians, and veterinarians for the excellent animal care.
This work was funded by the University of Georgia (MRA-001) and by Sanofi Pasteur (CRA UGA 001). In addition, T.M.R. was funded in part by the Georgia Research Alliance.
REFERENCES
- 1.Monto AS. 2010. Seasonal influenza and vaccination coverage. Vaccine 28(Suppl 4):D33–D44. doi: 10.1016/j.vaccine.2010.08.027. [DOI] [PubMed] [Google Scholar]
- 2.Ellebedy AH, Webby RJ. 2009. Influenza vaccines. Vaccine 27(Suppl 4):D65–D68. doi: 10.1016/j.vaccine.2009.08.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Epperson S, Jhung M, Richards S, Quinlisk P, Ball L, Moll M, Boulton R, Haddy L, Biggerstaff M, Brammer L, Trock S, Burns E, Gomez T, Wong KK, Katz J, Lindstrom S, Klimov A, Bresee JS, Jernigan DB, Cox N, Finelli L. 2013. Human infections with influenza A(H3N2) variant virus in the United States, 2011-2012. Clin Infect Dis 57(Suppl 1):S4–S11. doi: 10.1093/cid/cit272. [DOI] [PubMed] [Google Scholar]
- 4.Huang JW, Yang JM. 2011. Changed epitopes drive the antigenic drift for influenza A (H3N2) viruses. BMC Bioinformatics 12(Suppl 1):S31. doi: 10.1186/1471-2105-12-S1-S31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Xiao Y, Moghadas SM. 2013. Impact of viral drift on vaccination dynamics and patterns of seasonal influenza. BMC Infect Dis 13:589. doi: 10.1186/1471-2334-13-589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Anonymous. 2015. CDC presents updated estimates of flu vaccine effectiveness for the 2014-2015 season. Centers for Disease Control and Prevention, Atlanta, GA: http://www.cdc.gov/flu/news/updated-vaccine-effectiveness-2014-15.htm. [Google Scholar]
- 7.Belongia EA, Kieke BA, Donahue JG, Greenlee RT, Balish A, Foust A, Lindstrom S, Shay DK, Marshfield Influenza Study Group . 2009. Effectiveness of inactivated influenza vaccines varied substantially with antigenic match from the 2004-2005 season to the 2006-2007 season. J Infect Dis 199:159–167. doi: 10.1086/595861. [DOI] [PubMed] [Google Scholar]
- 8.Centers for Disease Control and Prevention. 2005. Seasonal influenza vaccine effectiveness. Centers for Disease Control and Prevention, Atlanta, GA. [Google Scholar]
- 9.Du X, Dong L, Lan Y, Peng Y, Wu A, Zhang Y, Huang W, Wang D, Wang M, Guo Y. 2012. Mapping of H3N2 influenza antigenic evolution in China reveals a strategy for vaccine strain recommendation. Nat Commun 3:709. doi: 10.1038/ncomms1710. [DOI] [PubMed] [Google Scholar]
- 10.Smith DJ, Lapedes AS, de Jong JC, Bestebroer TM, Rimmelzwaan GF, Osterhaus AD, Fouchier RA. 2004. Mapping the antigenic and genetic evolution of influenza virus. Science 305:371–376. doi: 10.1126/science.1097211. [DOI] [PubMed] [Google Scholar]
- 11.Lin YP, Xiong X, Wharton SA, Martin SR, Coombs PJ, Vachieri SG, Christodoulou E, Walker PA, Liu J, Skehel JJ, Gamblin SJ, Hay AJ, Daniels RS, McCauley JW. 2012. Evolution of the receptor binding properties of the influenza A(H3N2) hemagglutinin. Proc Natl Acad Sci USA 109:21474–21479. doi: 10.1073/pnas.1218841110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Crevar CJ, Ross TM. 2008. Elicitation of protective immune responses using a bivalent H5N1 VLP vaccine. Virol J 5:131. doi: 10.1186/1743-422X-5-131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Giles BM, Bissel SJ, Dealmeida DR, Wiley CA, Ross TM. 2012. Antibody breadth and protective efficacy are increased by vaccination with computationally optimized hemagglutinin but not with polyvalent hemagglutinin-based H5N1 virus-like particle vaccines. Clin Vaccine Immunol 19:128–139. doi: 10.1128/CVI.05533-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Giles BM, Crevar CJ, Carter DM, Bissel SJ, Schultz-Cherry S, Wiley CA, Ross TM. 2012. A computationally optimized hemagglutinin virus-like particle vaccine elicits broadly reactive antibodies that protect nonhuman primates from H5N1 infection. J Infect Dis 205:1562–1570. doi: 10.1093/infdis/jis232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Giles BM, Ross TM. 2011. A computationally optimized broadly reactive antigen (COBRA) based H5N1 VLP vaccine elicits broadly reactive antibodies in mice and ferrets. Vaccine 29:3043–3054. doi: 10.1016/j.vaccine.2011.01.100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Carter DM, Darby CA, Lefoley BC, Crevar CJ, Alefantis T, Oomen R, Anderson SF, Strugnell T, Cortes-Garcia G, Vogel TU, Parrington M, Kleanthous H, Ross TM. 2016. Design and characterization of a computationally optimized broadly reactive hemagglutinin vaccine for H1N1 influenza viruses. J Virol 90:4720–4734. doi: 10.1128/JVI.03152-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kiefer F, Arnold K, Kunzli M, Bordoli L, Schwede T. 2009. The SWISS-MODEL Repository and associated resources. Nucleic Acids Res 37:D387–D392. doi: 10.1093/nar/gkn750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Arnold K, Bordoli L, Kopp J, Schwede T. 2006. The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22:195–201. doi: 10.1093/bioinformatics/bti770. [DOI] [PubMed] [Google Scholar]
- 19.Biasini M, Bienert S, Waterhouse A, Arnold K, Studer G, Schmidt T, Kiefer F, Gallo Cassarino T, Bertoni M, Bordoli L, Schwede T. 2014. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res 42:W252–W258. doi: 10.1093/nar/gku340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.DeLano WL. 2002. The PyMOL molecular graphics system: MacPyMOL v1.7.6.2. DeLano Scientific, San Carlos, CA. [Google Scholar]
- 21.Lewis NS, Anderson TK, Kitikoon P, Skepner E, Burke DF, Vincent AL. 2014. Substitutions near the hemagglutinin receptor-binding site determine the antigenic evolution of influenza A H3N2 viruses in U.S. swine. J Virol 88:4752–4763. doi: 10.1128/JVI.03805-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Stray SJ, Pittman LB. 2012. Subtype- and antigenic site-specific differences in biophysical influences on evolution of influenza virus hemagglutinin. Virol J 9:91. doi: 10.1186/1743-422X-9-91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Arevalo MT, Wong TM, Ross T. 2 June 2016. Expression and purification of virus-like particles for vaccination. J Vis Exp 112:e54041. doi: 10.3791/54041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Tan GS, Lee PS, Hoffman RM, Mazel-Sanchez B, Krammer F, Leon PE, Ward AB, Wilson IA, Palese P. 2014. Characterization of a broadly neutralizing monoclonal antibody that targets the fusion domain of group 2 influenza A virus hemagglutinin. J Virol 88:13580–13592. doi: 10.1128/JVI.02289-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Jang YH, Seong BL. 2014. Options and obstacles for designing a universal influenza vaccine. Viruses 6:3159–3180. doi: 10.3390/v6083159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zhang H, Wang L, Compans RW, Wang BZ. 2014. Universal influenza vaccines, a dream to be realized soon. Viruses 6:1974–1991. doi: 10.3390/v6051974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Carter DM, Bloom CE, Nascimento EJ, Marques ET, Craigo JK, Cherry JL, Lipman DJ, Ross TM. 2013. Sequential seasonal H1N1 influenza virus infections protect ferrets against novel 2009 H1N1 influenza virus. J Virol 87:1400–1410. doi: 10.1128/JVI.02257-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kirchenbaum GA, Ross TM. 2014. Eliciting broadly protective antibody responses against influenza. Curr Opin Immunol 28:71–76. doi: 10.1016/j.coi.2014.02.005. [DOI] [PubMed] [Google Scholar]
- 29.Tsang TK, Cauchemez S, Perera RA, Freeman G, Fang VJ, Ip DK, Leung GM, Malik Peiris JS, Cowling BJ. 2014. Association between antibody titers and protection against influenza virus infection within households. J Infect Dis 210:684–692. doi: 10.1093/infdis/jiu186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Corti D, Suguitan AL Jr, Pinna D, Silacci C, Fernandez-Rodriguez BM, Vanzetta F, Santos C, Luke CJ, Torres-Velez FJ, Temperton NJ, Weiss RA, Sallusto F, Subbarao K, Lanzavecchia A. 2010. Heterosubtypic neutralizing antibodies are produced by individuals immunized with a seasonal influenza vaccine. J Clin Investig 120:1663–1673. doi: 10.1172/JCI41902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Hensley SE, Das SR, Bailey AL, Schmidt LM, Hickman HD, Jayaraman A, Viswanathan K, Raman R, Sasisekharan R, Bennink JR, Yewdell JW. 2009. Hemagglutinin receptor binding avidity drives influenza A virus antigenic drift. Science 326:734–736. doi: 10.1126/science.1178258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Hai R, Krammer F, Tan GS, Pica N, Eggink D, Maamary J, Margine I, Albrecht RA, Palese P. 2012. Influenza viruses expressing chimeric hemagglutinins: globular head and stalk domains derived from different subtypes. J Virol 86:5774–5781. doi: 10.1128/JVI.00137-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Krammer F, Hai R, Yondola M, Tan GS, Leyva-Grado VH, Ryder AB, Miller MS, Rose JK, Palese P, Garcia-Sastre A, Albrecht RA. 2014. Assessment of influenza virus hemagglutinin stalk-based immunity in ferrets. J Virol 88:3432–3442. doi: 10.1128/JVI.03004-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Margine I, Krammer F, Hai R, Heaton NS, Tan GS, Andrews SA, Runstadler JA, Wilson PC, Albrecht RA, Garcia-Sastre A, Palese P. 2013. Hemagglutinin stalk-based universal vaccine constructs protect against group 2 influenza A viruses. J Virol 87:10435–10446. doi: 10.1128/JVI.01715-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Ross TM, Xu Y, Bright RA, Robinson HL. 2000. C3d enhancement of antibodies to hemagglutinin accelerates protection against influenza virus challenge. Nat Immunol 1:127–131. doi: 10.1038/77802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Green TD, Montefiori DC, Ross TM. 2003. Enhancement of antibodies to the human immunodeficiency virus type 1 envelope by using the molecular adjuvant C3d. J Virol 77:2046–2055. doi: 10.1128/JVI.77.3.2046-2055.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Blom N, Sicheritz-Ponten T, Gupta R, Gammeltoft S, Brunak S. 2004. Prediction of post-translational glycosylation and phosphorylation of proteins from the amino acid sequence. Proteomics 4:1633–1649. doi: 10.1002/pmic.200300771. [DOI] [PubMed] [Google Scholar]
- 38.Huang ZZ, Yu L, Huang P, Liang LJ, Guo Q. 2017. Charged amino acid variability related to N-glycosylation and epitopes in A/H3N2 influenza: hemagglutinin and neuraminidase. PLoS One 12:e0178231. doi: 10.1371/journal.pone.0178231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.WHO Global Influenza Surveillance Network. 2011. Manual for the laboratory diagnosis and virological surveillance of influenza. World Health Organization, Geneva, Switzerland. [Google Scholar]
- 40.Katz JM, Hancock K, Xu X. 2011. Serologic assays for influenza surveillance, diagnosis and vaccine evaluation. Expert Rev Anti Infect Ther 9:669–683. doi: 10.1586/eri.11.51. [DOI] [PubMed] [Google Scholar]
- 41.Oh DY, Barr IG, Mosse JA, Laurie KL. 2008. MDCK-SIAT1 cells show improved isolation rates for recent human influenza viruses compared to conventional MDCK cells. J Clin Microbiol 46:2189–2194. doi: 10.1128/JCM.00398-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.European Medicines Agency. 2014. Guideline on influenza vaccines: non-clinical and clinical module (draft). European Medicines Agency, London, United Kingdom. [Google Scholar]
- 43.Matrosovich M, Matrosovich T, Garten W, Klenk HD. 2006. New low-viscosity overlay medium for viral plaque assays. Virol J 3:63. doi: 10.1186/1743-422X-3-63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Sullivan K, Kloess J, Qian C, Bell D, Hay A, Lin YP, Gu Y. 2012. High throughput virus plaque quantitation using a flatbed scanner. J Virol Methods 179:81–89. doi: 10.1016/j.jviromet.2011.10.003. [DOI] [PubMed] [Google Scholar]
- 45.Walls HH, Harmon MW, Slagle JJ, Stocksdale C, Kendal AP. 1986. Characterization and evaluation of monoclonal antibodies developed for typing influenza A and influenza B viruses. J Clin Microbiol 23:240–245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Dereeper A, Guignon V, Blanc G, Audic S, Buffet S, Chevenet F, Dufayard JF, Guindon S, Lefort V, Lescot M, Claverie JM, Gascuel O. 2008. Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res 36:W465–W469. doi: 10.1093/nar/gkn180. [DOI] [PMC free article] [PubMed] [Google Scholar]