Abstract
An efficient and low-cost method of examining larval movement in Drosophila melanogaster is needed to study how mutations and/or alterations in the muscular, neural, and olfactory systems affect locomotor behavior. Here, we describe the implementation of wrMTrck, a freely available ImageJ plugin originally developed for examining multiple behavioral parameters in the nematode C. elegans. Our optimized method is rapid, reproducible and does not require automated microscope setups or the purchase of proprietary software. To demonstrate the utility of this method, we analyzed the velocity and crawling paths of two Drosophila mutants that affect muscle structure and/or function. Additionally, we show that this approach is useful for tracking the behavior of adult insects, including Tribolium castaneum and Drosophila melanogaster.
Keywords: larval locomotion, behavior assay, muscle, Drosophila, Tribolium
1. Introduction
Drosophila melanogaster has emerged as a critical model organism to understand how altered behavior is linked to human disease phenotypes and aging. Both the larval and adult stages are routinely used in assays to monitor locomotion, feeding, circadian, and learning behaviors (http://www.sdbonline.org/sites/fly/aimain/6behavior.htm). The flight ability of Drosophila adults makes accurately recording speed and trajectories difficult. However, larval movement is relatively low speed, making it feasible to track both velocity and crawling patterns. Larval behavior can be reliably monitored to examine the effects of drugs or altered expression of human disease genes (Pandey & Nichols, 2011, van der Voet et al., 2014). Additionally, analysis of the larval stage is necessary if mutants do not survive to adulthood. For example, the muscle mutants tiggrin (tig); thin (tn); and muscle LIM protein (mlp84B) are all pupal lethal (Bunch et al., 1998, LaBeau-DiMenna et al., 2012, Clark et al., 2007). Thus, phenotypic analysis is performed in the prior third larval instar (L3) stage.
There are numerous published assays for examining larval movement. A simple, yet effective method is manual tracking on a gridded surface, which counts the number of squares traversed by larvae in a specified time period (Wang et al., 2015). However, this method can be time-consuming and the tracks of individual larvae are not recorded. Computer-based tracking programs have been developed for Drosophila and other organisms that rely on visualizing pixel intensity differences between the specimen and background (Sinadinos et al., 2012, Jung et al., 2015, Gomez-Marin et al., 2012). Unfortunately, the semi-translucent nature of the Drosophila larval body results in low contrast between the organism and background on standard apple juice media. One approach to circumvent this problem is to feed larvae colored dyes to enhance contrast followed by movement analysis using the commercially available Matlab software (Khurana et al., 2010). Other groups have generated technological imaging-based approaches that can be used for the rapid, high throughput analysis of various larval locomotor behaviors. Risse et al., designed an imaging technique based upon frustrated total internal reflection (FTIR) to generate high resolution movies that permit forward genetic screens (Risse et al., 2013). A computerized, single-animal tracking system, called MaggotTracker, was also devised to measure and analyze over 20 different parameters of larval traits, including shape, size, and peristalsis movements (Aleman-Meza, et al., 2015). While these automatic phenotyping programs are useful for large scale projects in which analysis of multiple genotypes is needed, simpler tracking systems are required for basic locomotor analysis.
Our lab has modified a freeware version of the ImageJ Plugin wrMTrck (ImageJ: http://imagej.nih.gov/ij/) and wrMTrck: http://www.phage.dk/plugins/wrmtrck.html), originally designed to track the movement and speed of C. elegans crawling on agar plates or swimming in liquid (Husson et al., 2012). Herein, we have modified the optimal settings of the wrMTrck program for the detection and analysis of Drosophila L3 larvae. This system has several advantages over other larvae tracking setups. First, we use a simple camera setup that provides consistency from one experiment to the next while being accessible to any lab. Second, we image individuals on apple juice agar supplemented with a dye for contrast enhancement over background. Third, larval movement is recorded on agar molded in weigh dishes (used to measure laboratory chemicals), thus eliminating the high contrast edge of petri dishes that can be misconstrued as larvae in automated tracking systems. Finally, we are able to analyze multiple parameters of larval behavior, including relative speed, distance traveled, and track path movement. As proof of principle, we present data for two mutants that exhibit muscle and/or neurological phenotypes. We further show that this simple setup can be used to track the movement of other insects, including Tribolium castanuem.
2. Materials and methods
2.1. Animal husbandry
Flies used in this study were reared from egg to adulthood in standard 12h:12h light:dark cycles and maintained in 25 × 95 mm polystyrene vials containing approximately 10 ml of standard cornmeal, molasses, yeast, and agar medium at 25 °C unless otherwise indicated. The following fly stocks were obtained from the Bloomington (BL) Stock Center: y[1], w[1118] as the wild-type strain (BL-6598); w[*]; parkin [1]/TM3, Sb[1] Ser[1] (BL-34747); w[*]; how[24B]-GAL4 (BL-1767); parkin RNAi (BL-38333); w[1118]; wg[Gla-1]/CyO, twi-2xEGFP (BL-6662) and w[1118]; Dr[Mio]/TM3, twi-2xEGFP, Sb[1] (BL-6663). The tn RNAi lines #1 (v19291) and #2 (v19290) were obtained from the Vienna Drosophila Resource Center (VDRC). The tnΔA mutant was generated as described (LaBeau-DiMenna et al., 2012). The parkin1 and tnΔA alleles were balanced over the appropriate GFP-marked balancer chromosomes and mutant larvae were selected by the absence of GFP using a Nikon M165 fluorescent stereomicroscope. 100% of the progeny from the 24B-Gal4>th RNAi cross contained both the Gal4 element and UAS-tn RNAi. The UAS-parkin RNAi line was balanced over the TM6b, Tb balancer chromosome so larvae of the genotype 24B>parkin RNAi were identified as they lacked the Tb larval phenotype.
2.2 Larval staging and verification of RNAi knockdown
All crosses or analysis of mutant stocks used in this study were reared as described above. Specifically, crosses were set up in vials at 25 °C and maintained at standard densities (10–20 flies/vial). Parental adults were transferred to new vials approximately every two days after first instar (L1) larvae were observed crawling in the food. Thereafter, mutant stocks (tn or parkin mutants) were maintained at 25°C, while control, th or parkin RNAi crosses were incubated at 29°C to induce maximal GAL4 expression for the induction of RNAi knockdown. Individuals that reach the late, ‘wandering’ L3 larval stage climb out of the food towards the top of the vial to undergo pupation and can be verified by examining mouth hook morphology (Roberts, 1998). To ensure that only wandering L3 larvae were selected for locomotion analysis, development at this stage was carefully monitored. As shown in Table S1, the average time for WT or control larvae to wander out of the food and reach 0 h after puparium formation (APF) was 6–7 h. This wandering stage was longer in tn or parkin mutants and in tn or parkin RNAi knockdown, taking up to 11 h. Only individuals that were wandering out of the food during these intervals were selected for locomotor activity assays.
For quantitative real-time PCR (qPCR), three biological replicates (L3 larvae) of each genotype were evaluated. Total RNA was extracted and purified using the RNeasy Mini Kit (Qiagen). Single stranded cDNA was generated from 125 ng RNA using the SuperScript III First-Strand Synthesis System Kit (Invitrogen). For the qPCR reactions, each cDNA solution was diluted 1:25 and mixed with iQ SYBR Green Supermix (Bio-Rad). rp49 was used as the reference gene. Primers for the qPCR reactions were synthesized by Integrated DNA Technologies: tn: [F5′ – GAGCTGCATATCGAAATCACCG, R5′ = AGATAGGCTTTTTCCGAGCAAAC] parkin: [F5′ – CTTACGCACACGCTGAGTATT, R5′ = GCCACCAGTTCCTTTACGTTTTT] rp49: [F5′-GCCCAAGGGTATCGACAACA, R5′-GCGCTTGTTCGATCCGTAAC]. All qPCR assays were performed using the CFX96 Touch Real-Time PCR Detection System with CFX Manager Software (Bio-Rad). Results from three technical replicates were averaged to generate Ct values for each biological replicate. Then, the differential normalized fold change was performed using the ΔΔCt method.
2.3. Camera setup and acquisition
Appropriately staged wandering L3 individuals (Fig. 1A) were removed from the side of the vial using a metal probe or paintbrush and placed on an agar plate. Apple or grape juice-based agar recipes are typically utilized for the collection of embryos and staging of larvae. While grape juice plates provide a dark background for contrast enhancement, larval velocity was significantly reduced on grape juice agar compared to apple juice agar plates (Fig. S1). Thus, we chose to use apple juice agar medium dyed with bromophenol blue, which did not alter larval movement (Fig. S1), to enhance the contrast of individual larvae over apple juice agar alone. Standard weigh dishes (Fisherbrand polystyrene, 140 × 140 × 25 mm, natural) were used as molds to create apple juice + bromophenol blue agar plates (Fig. S1). These resulting agar plates were substituted for media poured into petri dishes, the latter being more costly. Moreover, the reflective nature of the petri dish plastic edges are difficult to remove during ImageJ processing and wrMTrck analysis, often being labeled as larvae.
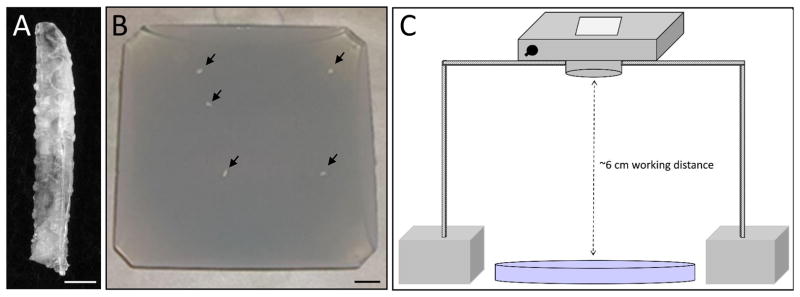
Fig. 1.
Larval locomotion recording setup. (A, B) Wandering Drosophila L3 larvae (A) were selected and placed upon apple juice agar plates dyed with bromophenol blue (B, arrows indicate larvae). (C) The camera was mounted upside down in an enclosed container to eliminate stray light. Scale bar, 0.05 mm (A), 1.0 mm (B).
Individuals were placed in the center of the apple juice agar plates and allowed to acclimate to their new surroundings for ~30 seconds before filming (Fig. 1B). A Kodak EasyShare C195 camera (14 megapixels; 640 × 480 video capture resolution) was placed upside down using a box (14 × 14 × 17 cm) as a stand (Fig. 1C) and raised to a height at which the entire apple juice plate was in the camera viewing area (~ 6 cm from the surface of the workbench). In addition, plates were placed on a black, uniform surface to reduce aberrant reflections during filming. Videos were captured for 60–360 seconds, depending on the type of analysis to be performed (Movies S3–5). Video files were uploaded onto a Dell Latitude E5540 Windows 7 (64-bit operating system) laptop computer.
2.4. Image J and Worm tracker analysis
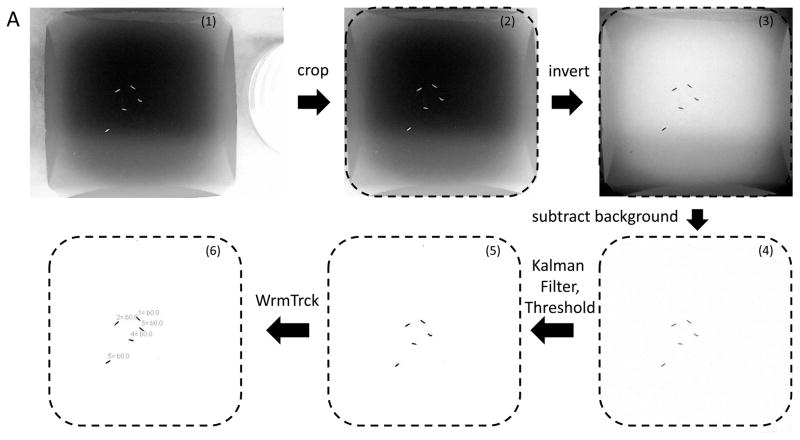
Fig. 2 depicts the flow and expected results at each step during ImageJ and wrMTrck analysis. Files (.avi) were opened in Image J and the ‘convert to grayscale’ option was selected in the ‘AVI Reader’ box (panel 1). Files that are not saved as uncompressed ‘.avi’ files by default can be converted using freely available conversion software (for example: http://www.anvsoft.com/any-video-converter-free.html). The rectangular selection tool was used to select an area of interest immediately adjacent to the edge of the agar (Image > Crop) (panel 2). The entire stack of images were then inverted so the larvae appeared dark on a light background (Edit > Invert) (panel 3). To analyze 60 sec worth of video data, substacks of 1788 frames, corresponding to 29.8 frames/sec, were created (Image > Stacks > Tools > Make Substack). Usually 10–100 frames from the beginning and end of the movie were omitted to avoid shaking of the camera when starting or stopping camera recording. For increased video lengths, the number of frames were adjusted accordingly.
Fig. 2.
ImageJ and WrMTrck analysis work flow. (A) After video collection, the resulting files were opened in ImageJ (panel 1), cropped to the area of interest (panel 2), and inverted so the larvae appear dark on a light background (panel 3). After applying the Kalman stack filter (panel 4), the image threshold was adjusted to increase the contrast of pixels corresponding to larvae (panel 5). Finally, application of the WrMTrck plugin assigned a number to each larva after data analysis.
wrMTrck measures speed as pixels/second. To convert to mm/sec, the length in mm of the agar plate was measured with a ruler from one edge to the other. In ImageJ, the same length was measured using the straight line tool: a line was drawn from one edge of the plate to the other and the length was measured in pixels (Analyze > Measure). The length in mm was divided by the length in pixels to give a conversion factor of mm/pixel. Multiplying the speed given in pixels/second by this conversion factor results in mm/second.
Table 1 lists the parameters used for variables that can affect the remaining analysis. The subtract background function was used to remove smooth, continuous backgrounds from all images (Process > Subtract background; rolling ball radius of 0.7; boxes corresponding to ‘light background’ and ‘sliding paraboloid’ were checked). Potential light flickering during video creation was removed using the Kalman Stack Filter option (Plugins > stacks > Kalman Stack; acquisition noise variance estimate = 0.05; bias to be placed on the prediction = 0.97) panel 4). The addition of the Kalman Stack Filter to eliminate excess light was critical for accurate quantification of larval velocity in individuals that demonstrated limited movement. For example, calculations of rate movement in heat-killed larvae were 0.22 mm/sec compared to 0.003 mm/sec with and without the Kalman filter, respectively (Fig. S2). WT larvae also exhibited a 23% increase in speed without application of the Kalman filter. wrMTrck requires a threshold image for analysis to further delineate larvae over background (Image>Adjust>Threshold; use default settings and deselect ‘dark background;’ click on Apply and OK in ‘Convert Stack to Binary’ box) (panel 5). Next we ran the wrMTrck plugin (Plugins>wrMTrck>wrMTrck) and modified the optimal settings for the tracking of L3 larvae as indicated in Table 1, specifically analyzing path length, path trace, and the raw data for movement parameters. The final wrMTrck output assigns a number to each larvae tracked (panel 6) that corresponds to the raw data displayed as a results table.
Table 1.
Optimized parameters to analyze larval or adult insect movement.1
| Parameters in ImageJ or WormTracker Analysis | Default | Drosophila L3 | Drosophila adult | Tribolium |
|---|---|---|---|---|
| Subtract background | ||||
| Rolling ball radius (pixels): | 10 | 0.7 | 5 | 10 |
| Kalman Stack Filter | ||||
| Acquisition noise variance estimate: | 0.05 | 0.05 | 0.05 | 0.05 |
| Bias to be placed on the prediction: | N/A | 0.97 | 0.5 | 0.80 |
| WrMTrck | ||||
| minSize2 – Minimum Object Area (pixels ∧2): | 100 | 10 | 5 | 10 |
| maxSize2 – Minimum Object Area (pixels ∧2): | 400 | 400 | 400 | 400 |
| maxVelocity – Maximum velocity (pixels/frame): | 10 | 10 | 10 | 10 |
| maxAreaChange – Maximum area change (%): | 30 | 200 | 100 | 100 |
| minTrackLength – Minimum track length (frames): | 50 | 500 | 50 | 50 |
| bendThreshold – Threshold for turn: | 2.0 | 2.0 | 2.0 | 2.0 |
| binSize – Size of bin for speed histogram (pixels/frame): | 0.0 | 0.0 | 0.0 | 0.0 |
| save Results File3 | - | - | - | |
| show Path Lengths | √ | √ | √ | |
| show Labels | √ | √ | √ | |
| show Positions3 | - | - | - | |
| show Paths | √ | √ | √ | |
| show Summary | √ | √ | √ | |
| round Coordinates3 | - | - | - | |
| smoothing | √ | √ | √ | |
| plotBendTrack3 | - | - | - | |
| rawData – (0=off, 5 = Bendcalc): | 0 | 0 | 0 | 0 |
| bendDetect – (0=off, 3 = AR + Histogram): | 2 | 2 | 2 | 2 |
| FPS – frames/sec (0= try to load from file): | 30 | 0 | 30 | 30 |
| backSub – background subtraction (0=off, 1=F1RB15): | 0 | 0 | 0 | 0 |
| threshMode – Thresholding method (only of backSub=0): | Otsu | Otsu | Otsu | Otsu |
| fontSize – Size of labeling font: | 16 | 16 | 16 | 16 |
The settings here are optimized for a 14 megapixel, 640 × 480 video capture resolution camera. Higher video resolution may require optimization of settings than depicted above.
Increasing video resolution would result in an increase in the minSize and maxSize.
Optional parameters
2.5. Locomotor parameters analyzed
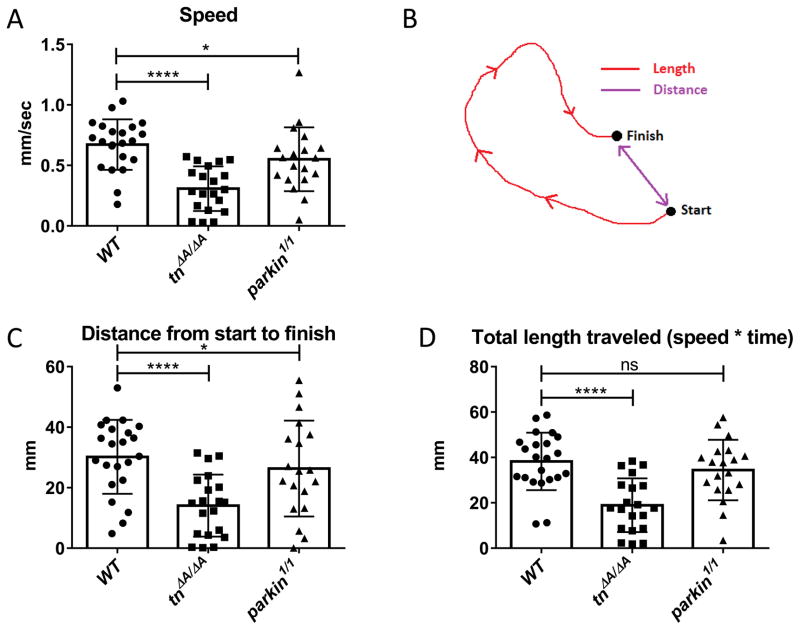
To demonstrate the utility and reproducibility of our experimental setup, we tested mutants known or predicted to exhibit locomotor defects. Larvae that are mutant for Drosophila parkin, a gene mutated in patients that have sporadic Parkinson’s disease (PD), are already known to exhibit locomotion defects (Vincent et al., 2012). Trim32 is an E3 ubiquitin ligase and mutations that cluster in the NHL domains result in human limb-girdle muscular dystrophy type 2H (LGMD2H) (Shieh et al., 2011). Although not yet tested, mutations in the Drosophila ortholog of Trim32 called thin (tn), would be expected to exhibit locomotor defects due to the degenerative muscle phenotype observed in tn L3 mutants (LaBeau-DiMenna et al., 2012). Indeed, the rate of larval crawling was decreased in both tn and parkin mutants compared to WT larvae (Fig. 3A). We also analyzed the distance from start to finish (Fig. 3B, C) and the total length traveled by each larva (Fig. 3B, D). Each of these parameters is decreased in tn mutant larvae.
Fig. 3.
Locomotion is reduced in tn and parkin mutants. (A) Individuals homozygous for mutations in tn and parkin travel slower than their WT counterparts. (B) Diagram of path parameters, including the length (red) and distance (purple) from start to finish measured in WrMTrck. (C, D) Quantitation of the distance (C) and total length traveled (D). Both are reduced in tn, but not parkin mutants, compared to WT larvae. Data was analyzed using one-way ANOVA with Dunnett’s post hoc test (mean = +/− SEM) (* p < 0.05; **** p < 0.0001; ns = not significant).
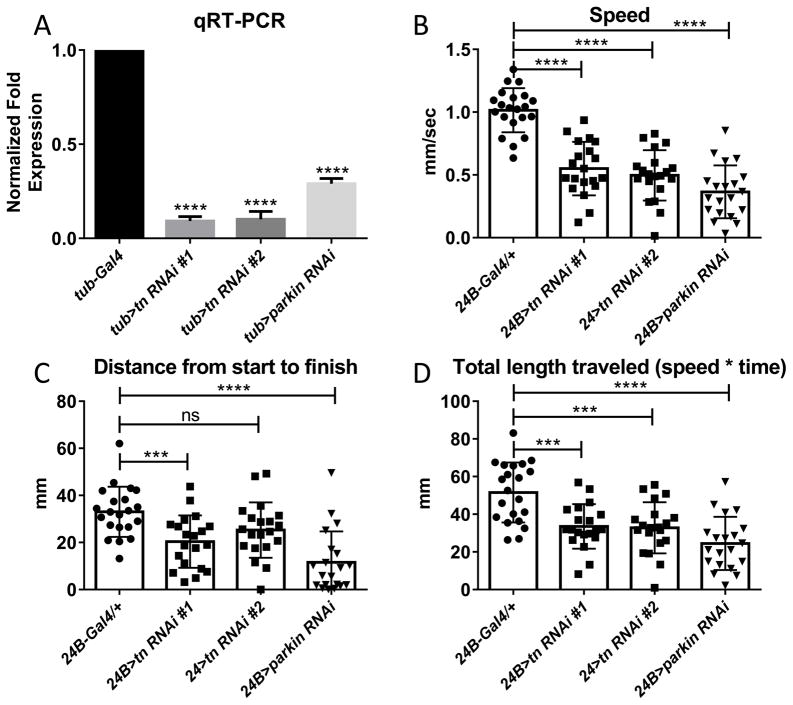
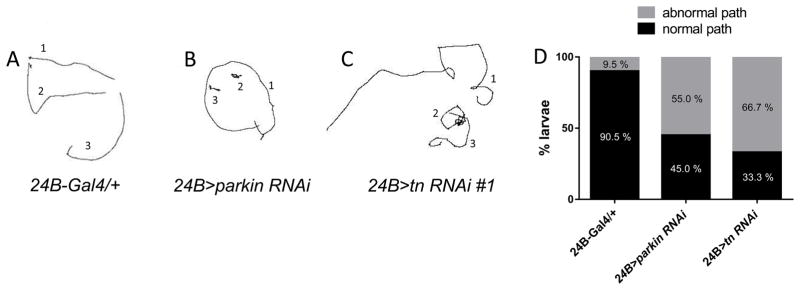
To verify the locomotor defects independently, we utilized muscle-specific RNAi knockdown of tn and parkin. Consistent with defects observed in tn and parkin mutants, RNAi mutant larvae also had slower crawling speeds (Fig. 4A). These mutants also ended up closer to their starting points (Fig. 4B) and traveled less overall distance (Fig. 4C) than their control counterparts. These data suggest the mutant travel paths may be altered. To test this possibility, we enabled the ‘show paths’ box during wrMTrck analysis, which provided a trace of each larval path. Control L3 individuals (24B-Gal4/+) traversed relatively straight paths (Fig. 5A). However, knockdown of either parkin (Fig. 5B) or tn (Fig. 5C) in muscle tissue generally resulted in less overall distance traveled and jagged or sharp turns. Quantitation of the total number of normal vs. circular paths is depicted in Fig. 5D, where more than half of parkin or tn RNAi larvae had paths altered from their WT counterparts.
Fig. 4.
Muscle-specific RNAi knockdown of tn and parkin result in measurable locomotion defects. (A) qRT-PCR results demonstrate that tn and parkin transcripts are reduced upon ubiquitous induction of RNAi by the tubulin promoter (tub-Gal4). (B–D) A reduction in tn and parkin RNA levels by a muscle Gal4 driver (24B-Gal4) results in decreased larval locomotion speeds (B), a shorter distance traversed from start to end points (C), and a loss of total distance traveled (D) compared to 24B-Gal4/+ controls. Data was analyzed using one-way ANOVA with Kruskal-Wallis test (A) or Dunnett’s post hoc test (B–D) (mean = +/− SEM) (*** p < 0.0005; **** p < 0.0001; ns = not significant).
Fig. 5.
Traces of larval path routes. (A–C) L3 larvae of the indicated genotypes were monitored for 120 sec and path lengths analyzed using ImageJ and WrMTrck. Compared to WT larvae which traverse relatively straight paths (A), both muscle-specific parkin RNAi (B) and tn RNAi (C) larvae make frequent turns and paths appear uncoordinated. (D) Quantitation of normal vs. abnormal paths traveled by the larvae of the indicated genotypes.
2.6. Use of Worm Tracker for other insect analysis
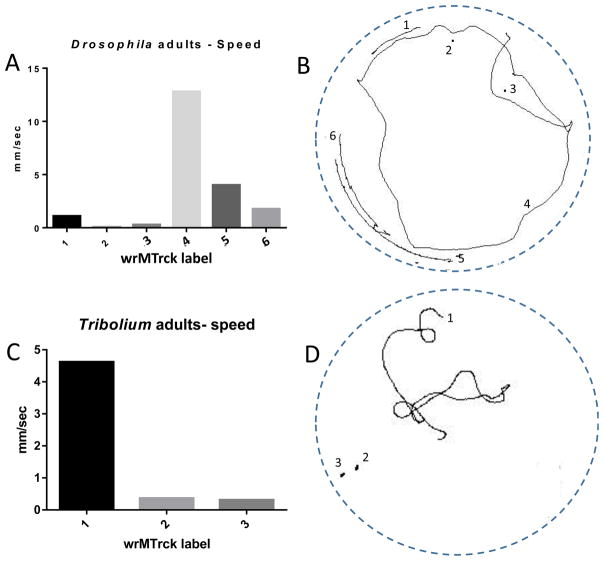
This experimental setup can also be used for the analysis of locomotor behaviors in adult Drosophila or other insects. The walking behavior of Drosophila adults or Tribolium castaneum adults were measured in petri dishes with lids to avoid adult insects from taking flight using the parameters in Table 1. Since adult insects move faster than larvae, only 10–30 sec of video needed to be analyzed. Overall, speeds (Fig. 6A) and paths traveled (Fig. 6B) varied widely from one individual to the next. This same trend was evident upon analyzing Tribolium adults.
Fig. 6.
Tracking and analysis of adult Drosophila and Tribolium. (A, C) wrMTrck can be used to monitor the velocity and paths traveled in Drosophila (A, B) and Tribolium (C, D) adults.
3. Results and Discussion
Using our video setup and analysis parameters, we are able to monitor the velocity, total distance traveled, and relative path taken by Drosophila L3 larvae. While many tracking systems require a microscopy setup, the present setup requires only a camera and the experiment can be performed on any horizontal surface at room temperature. The inexpensive equipment required for experimental setup and freely available ImageJ and wrMTrck software allows access to a variety of users. Further, our system also allows for 3–6 larvae to be followed at one time, thus providing an efficient, time-saving, single-tracking system. Single larvae can also be examined in our system to avoid potential social interactions in different genetic backgrounds that may affect locomotor behavior (Durisko et al., 2014).
wrMTrck was unofficially released to the worm community in 2007 and multiple other platforms that vary in cost and efficiency have been designed since this time (http://www.wormbook.org/chapters/www_tracking/tracking.html). Nematodes crawl in sinusoidal waves, while Drosophila larvae migrate using peristalsis. These latter movements can be efficiently tracked using the wrMTrck software with minor adjustments to parameters used to recognize and track pixels corresponding to crawling larvae (Table 1). In addition, we have optimized tracking parameters to monitor the crawling behavior of adult Drosophila and Tribolium. This could be effective for tracking these insects under different conditions, such as the time required to navigate to a nutrient or odorant source, as movement behavior of individuals is a critical factor in the population ecology of Tribolium (Young et al., 2013).
Our lab is primarily interested in understanding how mutations in the locomotor system, whether muscular or neuronal in origin, contribute to underlying mobility defects. Analysis of multiple mutants using the manual counting method allowed us to measure only locomotion speed (Wang et al., 2015). Altering parameters in wrMTrck has made it possible to monitor larval crawling length in a set amount of time and to analyze crawling path patterns. It has been long recognized that mutations in neuronal genes can lead to altered path defects, often reminiscent of neurodegenerative disease phenotypes (Jakubowski et al., 2012). In addition to decreased velocity, loss of parkin alters larval locomotion paths. Control larvae exhibit directional wandering in relatively straight paths, while parkin mutants perform frequent turns with no apparent destination (Fig. 4). A previous report suggests that only the neuronal contribution of parkin affects larval locomotion speed (Vincent et al., 2012), while our data provide evidence that a muscle-specific decrease in Parkin alters locomotion speed and path attributes. These data support the neuronal and muscle phenotypes observed in parkin mutants (Guo, 2012). Unexpectedly, tn mutants also show gross defects in path wandering. Some larvae make small turns not observed in WT larval paths, while others exhibit small, uncoordinated traces reminiscent of parkin mutants. It would be interesting to examine the paths of additional mutants that affect muscle structure and/or function to see if altered path traces are a common feature.
Supplementary Material
Acknowledgments
Thank you to Cheryl Clark, Nicole Green, and Zong-heng Wang for reading the manuscript and providing insightful comments. Also, a special thanks to the Integrated Genomics Facility in the Department of Plant Pathology at Kansas State University for assistance with the qPCR results.
Funding
Research reported in this publication was supported by the National Institute of Arthritis and Musculoskeletal and Skin Diseases of the National Institutes of Health under Awards Number R01AR060788 to ERG and R21AR066264 to SB. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. This work was supported by NIH grants R01AR060788 to ERG and R21AR066264 to SB. JK is funded through an AHA predoctoral fellowship (14PRE18850079).
References
- Bunch TA, Graner MW, Fessler LI, Fessler JH, Schneider KD, Kerschen A, Choy LP, Burgess BW, Brower DL. The PS2 integrin ligand tiggrin is required for proper muscle function in Drosophila. Development. 1998;125:1679–89. doi: 10.1242/dev.125.9.1679. [DOI] [PubMed] [Google Scholar]
- Clark KA, Bland JM, Beckerle MC. The Drosophila muscle LIM protein, Mlp84B, cooperates with D-titin to maintain muscle structural integrity. J Cell Sci. 2007;120:2066–77. doi: 10.1242/jcs.000695. [DOI] [PubMed] [Google Scholar]
- Durisko Z, Kemp R, Mubasher R, Dukas R. Dynamics of social behavior in fruit fly larvae. PLoS One. 2014;9:e95495. doi: 10.1371/journal.pone.0095495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gomez-Marin A, Partoune N, Stephens GJ, Louis M, Brembs B. Automated tracking of animal posture and movement during exploration and sensory orientation behaviors. PLoS One. 2012;7:e41642. doi: 10.1371/journal.pone.0041642. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo M. Drosophila as a model to study mitochondrial dysfunction in Parkinson’s disease. Cold Spring Harb Perspect Med. 2012:2. doi: 10.1101/cshperspect.a009944. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Husson SJ, Costa WS, Schmitt C, Gottschalk A. Keeping track of worm trackers. WormBook. 2012:1–17. doi: 10.1895/wormbook.1.156.1. [DOI] [PMC free article] [PubMed]
- Jakubowski BR, Longoria RA, Shubeita GT. A high throughput and sensitive method correlates neuronal disorder genotypes to Drosophila larvae crawling phenotypes. Fly (Austin) 2012;6:303–8. doi: 10.4161/fly.21582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jung SK, Qu X, Aleman-Meza B, Wang T, Riepe C, Liu Z, Li Q, Zhong W. Multi-endpoint, high-throughput study of nanomaterial toxicity in Caenorhabditis elegans. Environ Sci Technol. 2015;49:2477–85. doi: 10.1021/es5056462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khurana S, Li WK, Atkinson NS. Image enhancement for tracking the translucent larvae of Drosophila melanogaster. PLoS One. 2010;5:e15259. doi: 10.1371/journal.pone.0015259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Labeau-Dimenna EM, Clark KA, Bauman KD, Parker DS, Cripps RM, Geisbrecht ER. Thin, a Trim32 ortholog, is essential for myofibril stability and is required for the integrity of the costamere in Drosophila. Proc Natl Acad Sci U S A. 2012;109:17983–8. doi: 10.1073/pnas.1208408109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pandey UB, Nichols CD. Human disease models in Drosophila melanogaster and the role of the fly in therapeutic drug discovery. Pharmacol Rev. 2011;63:411–36. doi: 10.1124/pr.110.003293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Risse B, Thomas S, Otto N, Löpmeier T, Valkov D, Jiang X, Klämbt C. FIM, a novel FTIR-based imaging method for high throughput locomotion analysis. PLoS One. 2013;8:e53963. doi: 10.1371/journal.pone.0053963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roberts D. Drosophila: A Practical Approach. Oxford University Press; 1998. [Google Scholar]
- Shieh PB, Kudryashova E, Spencer MJ. Limb-girdle muscular dystrophy 2H and the role of TRIM32. Handb Clin Neurol. 2011;101:125–33. doi: 10.1016/B978-0-08-045031-5.00009-8. [DOI] [PubMed] [Google Scholar]
- Sinadinos C, Cowan CM, Wyttenbach A, Mudher A. Increased throughput assays of locomotor dysfunction in Drosophila larvae. J Neurosci Methods. 2012;203:325–34. doi: 10.1016/j.jneumeth.2011.08.037. [DOI] [PubMed] [Google Scholar]
- Van Der Voet M, Nijhof B, Oortveld MA, Schenck A. Drosophila models of early onset cognitive disorders and their clinical applications. Neurosci Biobehav Rev. 2014;46(Pt 2):326–42. doi: 10.1016/j.neubiorev.2014.01.013. [DOI] [PubMed] [Google Scholar]
- Vincent A, Briggs L, Chatwin GF, Emery E, Tomlins R, Oswald M, Middleton CA, Evans GJ, Sweeney ST, Elliott CJ. parkin-induced defects in neurophysiology and locomotion are generated by metabolic dysfunction and not oxidative stress. Hum Mol Genet. 2012;21:1760–9. doi: 10.1093/hmg/ddr609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang ZH, Rabouille C, Geisbrecht ER. Loss of a Clueless-dGRASP complex results in ER stress and blocks Integrin exit from the perinuclear endoplasmic reticulum in Drosophila larval muscle. Biol Open. 2015;4:636–48. doi: 10.1242/bio.201511551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young H, Reid T, Randall L, Lachowsky L, Foster D, Pengelly C, Latty T, Reid M. Influences of Movement Behavior on Animal Distributions at Edges of Homogeneous Patches. International Journal of Zoology 2013 [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.