Abstract
Lung adenocarcinoma, characterized by its early and aggressive local invasion and high metastatic potential, is the most frequently observed histological type of non-small-cell lung cancer (NSCLC). Visceral pleural invasion (VPI) caused by peripheral lung adenocarcinomas is closely associated with the poor prognosis of patients with NSCLC. The association between VPI and some clinicopathological characteristics has been observed in the past few decades. However, the molecular mechanism of VPI in lung adenocarcinomas is unknown. In the present, the expression level of microRNA (miR-)135b and epidermal growth factor receptor (EGFR) mutations using the reverse transcription-quantitative polymerase chain reaction and DNA sequencing, respectively. In addition, the present study aimed at exploring the association between the miR-135b level, EGFR mutations and VPI in peripheral lung adenocarcinoma. The results of the present study demonstrated that miR-135b was significantly upregulated in lung adenocarcinoma compared with adjacent normal tissue and positively associated EGFR mutations in peripheral lung adenocarcinoma. Furthermore, it was identified that lung adenocarcinomas with EGFR mutations and miR-135b overexpression were more likely to invade visceral pleura. Taken together, these findings indicate that miR-135b overexpression is positively associated with mutations to EGFR, which may promote the development of peripheral lung adenocarcinomas by the formation of VPI. This indicates that the two factors may serve as prognostic markers and molecular targets for the treatment of peripheral lung adenocarcinomas.
Keywords: peripheral lung adenocarcinomas, epithelial growth factor receptor mutations, microRNA-135b, visceral pleural invasion, invasion
Introduction
Lung cancer, the most common cause of cancer-associated mortality in women and men worldwide (1,2), has a high invasive and metastatic potential, which leads to drug resistance and treatment failure (3). A total of 85% of all cases of lung cancers are of the non-small cell lung cancer (NSCLC) histotype, of which lung adenocarcinoma is the most frequent type (accounting for 40%) (4). Although advances in treatment have been made in recent years, patients with lung adenocarcinoma continue to have poor prognoses when the disease is detected at an advanced clinical stage (5). Owing to its growth site in the periphery of the lung, adenocarcinoma readily invades the pleural cavity (6). Visceral pleural invasion (VPI) has been regarded as a pathological descriptor and a poor independent prognostic factor for the past several decades in NSCLC (7–9). In addition to the current Tumor-Node-Metastasis (TNM) classification recommendations (10), in which T1 tumors with VPI are upgraded to T2a, T2a tumors with VPI should be classified as T2b (11). Evidence indicates that patients suffering with VPI had a poorer prognosis, even when adjusting for tumor size (12–14). In addition, VPI is also associated with extensive N2 involvement (9) and lymph node metastasis (9,15,16), which reflects the fact that malignant tumor cells can move to the cervical venous circulation through the mediastinal lymphatic vessels, leading to a greater incidence of mortality (8). VPI has long been recognized as an important clinical factor by thoracic surgeons, medical oncologists and radiation oncologists when treating patients with NSCLC. Although a number of studies have focused on VPI in NSCLC (7,8,14), few have assessed the role of VPI in lung adenocarcinomas, let alone explored the molecular mechanism.
The epidermal growth factor receptor (EGFR), one of the members of the ErbB receptor family, is activated by the binding of its extracellular domain to specific ligands, including EGF and transforming growth factor-α (TGFα), which causes trans-autophosphorylation of EGFR and activation of the intracellular tyrosine kinase activity ultimately leading to cell proliferation, survival, invasion, and metastasis (17,18). Approximately 10% of Western and 40% of Asian patients with lung cancer harbor somatic EGFR mutations (19), which have a notable role in the pathogenesis of lung cancer (20), particularly in lung adenocarcinomas. Tsai et al (21) found that lung adenocarcinomas harboring the EGFR-L858R mutation tended to invade the adjacent pleural cavity and be involved in the formation of malignant pleural effusion (MPE). Shi et al (22) analyzed the associations between EGFR mutations and clinicopathological factors in lung adenosquamous cell carcinoma cases and found that the incidence of positive pleural invasion was higher in patients positive for the EGFR-L858R mutation than that in those negative for the mutation. In addition, NSCLC patients with EGFR mutations are sensitive to EGFR tyrosine kinase inhibitors (TKIs) such as gefitinib (23). Of the different types of EGFR mutations, exon 19 deletions and missense mutations (L858R) in exon 21 are the two predominant types, and have been found to be the most powerful biologic predictors of EGFR TKI sensitivity (24). Although studies investigating clinical and histological characteristics of patients harboring EGFR mutations have been published in recent years (19–22), the prognostic value of EGFR mutations in patients with peripheral lung adenocarcinomas exhibiting VPI is uncertain.
MicroRNAs (miRNAs/miRs), a class of small non-coding, highly conserved single-stranded RNAs, can serve as oncogenes or tumor suppressors when the expression is dysregulated in malignancies (25). For example, miR-483-5p promotes the epithelial-mesenchymal transition (EMT), which is frequently accompanied by the invasive, metastatic transformation of lung adenocarcinoma cells (26). miR-543 inhibits the proliferation and metastasis of colorectal cancer (CRC) cells, which indicates that it may serve as a favorable diagnostic and biomarker for CRC metastasis (27). However, the potential to use molecules similar to these for cancer diagnosis has not yet been systematically explored. A previous study revealed that microRNA expression could be used to distinguish between tumors of different developmental origin, and between tumors and normal tissue, and reflect the state of cellular differentiation (28). miRNAs may serve as useful clinical biomarkers for the screening of high-risk populations and detection of solid tumors in the early stages of cancer progression, and may also be novel targets for cancer therapy (29,30). Valeri et al demonstrated that use of anti-miR-135b in CRC mouse models has substantial therapeutic potential, which indicated that miRNAs are a promising therapeutic target (30). Previous findings demonstrated that miR-135b was upregulated in many types of cancer, including colorectal cancer (31), hepatocellular carcinoma (32), head and neck squamous cell carcinoma (33) and NSCLC (3). Additionally, miR-135b participates in the development and progression of cancer by promoting the proliferation, invasion and metastasis of tumor cells, (34). Although Lin et al (3) demonstrated that miR-135b can promote lung cancer cell invasion in vitro and metastasis in vivo, the role of miR-135b in lung adenocarcinomas was not clear. In addition, the role of miR-135b in the formation of VPI has not yet been reported.
Despite the general appreciation of the role of VPI in lung cancer dissemination, little is known about the molecular characteristics and gene expression profile that may attract cancer cells to the pleural space (35). In order to study the association between the miR-135b level, EGFR mutations and VPI in peripheral lung adenocarcinoma, the present study examined the expression level of miR-135b and EGFR mutations using the reverse transcription-quantitative polymerase chain reaction (RT-qPCR) and DNA sequencing, respectively. The results of the present study demonstrated that miR-135b was significantly upregulated in lung adenocarcinoma compared with adjacent normal tissue and positively associated EGFR mutations in peripheral lung adenocarcinoma. Furthermore, it was identified that lung adenocarcinomas with EGFR mutations and miR-135b overexpression were more likely to invade visceral pleura. The present study highlights the significance of miR-135b and EGFR mutations with VPI; therefore, the present study provides potential drug targets in peripheral lung adenocarcinoma therapy.
Materials and methods
Specimens
A total of 65 paraffin-embedded peripheral lung adenocarcinoma tissue and corresponding pericarcinomatous tissue that had been surgically excised between January 2012 and October 2014 from the Affiliated Zhoushan Hospital of Wenzhou Medical University (Zhejiang, China) were used in this study. No patient received radiotherapy, chemotherapy or any other antineoplastic therapy prior to surgery. The cases were divided into those of adenocarcinoma in situ/minimally invasive adenocarcinoma (AIS/MIA) and invasive adenocarcinoma (IAC); there were 40 cases (61.5%) of AIS/MIA and 25 cases (38.5%) of IAC. In total, 21 patients were male (32.3%) and 44 were female (67.7%). The median age was 56 years (range, 31–83 years) (Table I). At least three qualified and certified pathologists conducted strict identification of the tissues, the classification of each sample was undetermined until at least two-thirds of the recognized experts reached a consensus. Staging and pathological classification standard reference to the World Health Organization classification of lung cancer, 2011 (36). The present study was approved by the Ethics Committee of the Affiliated Zhoushan Hospital of Wenzhou Medical University.
Table I.
Patient characteristics and demographic data.
| Characteristics | Patients, n (%) |
|---|---|
| Sex | |
| Male | 21 (32.3) |
| Female | 44 (67.7) |
| Age, years | |
| ≥60 | 25 (38.5) |
| <60 | 40 (61.5) |
| Smoking status | |
| Yes | 14 (21.5) |
| No | 51 (78.5) |
| TNM clinical stage | |
| 0 | 26 (40.0) |
| IA | 22 (33.8) |
| IB-IV | 17 (21.2) |
| Tumor size, cm | |
| ≥3 | 60 (92.3) |
| >3 | 5 (7.7) |
| Site | |
| Left | 28 (43.1) |
| Right | 37 (56.9) |
| CEA, µg/ml | |
| ≥5 | 8 (12.3) |
| <5 | 57 (87.7) |
| VPI | |
| Yes | 17 (26.2) |
| No | 48 (73.8) |
| Histological types | |
| AIS/MIA | 40 (61.5) |
| IAC | 25 (38.5) |
TNM, Tumor-Node-Metastasis; CEA, carcinoembryonic antigen; VPI, visceral pleural invasion; AIS, adenocarcinoma in situ; MIA, minimally invasive adenocarcinoma; IAC, invasive adenocarcinoma.
Total RNA extraction
Total RNA, including miRNA, was extracted from 65 paraffin-embedded samples of tumor tissue and corresponding pericarcinomatous tissue using RecoverAll™ Total Nucleic Acid Isolation kit (Thermo Fisher Scientific, Inc., Waltham, MA, USA) and the concentration and purity of total RNA was measured using the Q-3000 micro-ultraviolet spectrophotometer (Quawell Technology, Inc., San Jose, CA, USA). Total RNA was extracted and stored at −80°C until use.
Taqman-based RT-qPCR
The isolated total RNA was reverse transcribed using TaqMan MicroRNA Reverse Transcription kit (Life Technologies; Thermo Fisher Scientific, Inc.) for mature microRNA-135b and U6 internal control by MyCycler Thermal Cycler (Bio-Rad Laboratories, Inc., Hercules, CA, USA). Reaction conditions were: 16°C for 30 min, 42°C for 30 min, and 85°C for 5 min. PCR was then performed on cDNA using TaqMan 2X Universal PCR Master Mix with no AmpErase Uracil-N-Glycosylase and TaqMan Human MiRNA Assay kits (cat nos. miR-135b, 002261; U6, 001973; Applied Biosystems; Thermo Fisher Scientific, Inc.) on an Applied Biosystems 7500 real-time PCR system (Thermocycling conditions were: 95°C for 10 min, followed by 40 cycles of 95°C for 15 sec and 60°C for 1 min). All qPCR reactions were performed in triplicate. The quantity of miR-135b in each peripheral lung adenocarcinoma tissue was calculated using the 2−ΔΔCq method (37) by SDS 2.0.1 software (Applied Biosystems; Thermo Fisher Scientific, Inc.), ΔΔCq=ΔCq (cancer group)-ΔCq(adjacent cancer group), where ΔCq=Cq(miR-135b)-Cq(U6).
Genomic DNA extraction
Genomic DNA was isolated from paraffin-embedded lung adenocarcinoma tissue using the DNA FFRPE Tissue kit (Qiagen, Inc., Valencia, CA, USA), according to the manufacturer's instructions. The concentration and purity of total DNA was measured using a Q-3000 micro-ultraviolet spectrophotometer (Quawell Technology, Inc.). DNA was authenticated by 1.2% agarose gel electrophoresis.
PCR and product purification
PCR was used to analyze exons 19 and 21 of the EGFR gene. The primers used were: exon 19 forward, 5′-CCCCAGCAATATCAGCCTTA-3′ and reverse, 5′-TGTGGGAGATGAGCAGGGTCT-3′; and exon 21 forward, 5′-GAATTCGGATGCAGAGCTTC-3′ and reverse, 5′-TCATTCACTGTCCCAGCAAG-3′. The reaction system was determined as follows: 10X Buffer 5 µl, 5X Q-Solution 10 µl, 100 mmol dNTP 0.1 µl (Applied Biosystems; Thermo Fisher Scientific, Inc.), 2.5 units Hot-Star Taq DNA polymerase (Qiagen, Inc.), 50 ng template DNA 5 µl, ddH2O up to 50 µl. PCR thermocycling conditions: 95°C for 15 min; 94°C for 15 sec, 60°C for 30 sec, 72°C for 1 min, 10 cycles; 94°C for 15 sec; 56°C for 30 sec, 72°C for 1 min, 25 cycles; 72°C for 10 min. PCR products was authenticated by 1.5% agarose gel electrophoresis and purified using the EasyPure PCR Purification kit (Beijing Transgen Biotech Co., Ltd., Beijing, China), according to the manufacturer's instructions.
Gene sequencing for EGFR mutation status
Sequencing reaction system: 1 µl BigDye, 0.5 ml BigDye seqBuffer, 1 M primer, 1 µl PCR products that have been purified, 1.5 µl ddH2O (96°C for 1 min; 96°C for 10 sec, 50°C for 5 sec, 60°C for 4 min, followed by 25 cycles) (Applied Biosystems; Thermo Fisher Scientific, Inc.). The PCR products of sequencing reaction were purified using the ethanol and EDTA precipitation method (19), and dissolved with Hi-Di Formamide, followed by 95°C for 4 min, ice water for 4 min, then separated by electrophoresis with ABI3100XL genetic analyzer. The original sequencing results were analyzed using ABI Sequencing Analysis V5.4 software (Applied Biosystems; Thermo Fisher Scientific, Inc.). The occurrence of EGFR mutations were confirmed by comparative analysis between the results on ABI SeqScap software version 5 (Applied Biosystems; Thermo Fisher Scientific, Inc.) and the reference EGFR gene sequence (19), in combination with manual checking.
Statistical analysis
Statistical analyses were performed using SPSS v.21.0 (IBM Corp., Armonk, New York) and GraphPad Prism 5 (GraphPad Software, Inc., La Jolla, CA, USA). Paired Student's t-test was used to analyze the difference in the expression of miR-135b between 65 pairs of peripheral lung adenocarcinomas and adjacent non-cancerous tissue. Mann-Whitney U-test was used to compare the expression level of miR-135b in different pathological classification of lung adenocarcinomas. Pearson's χ2 test and Fisher's exact test were used to evaluate the association between miR-135b expression and clinicopathological characteristics in peripheral lung adenocarcinomas, the association between EGFR mutation status and clinicopathological characteristics, the association between miR-135b overexpression and EGFR mutations, and the association between EGFR mutations with miR-135b overexpression and VPI. P<0.05 was considered to indicate a statistically significant difference.
Results
miR-135b is markedly upregulated in invasive lung adenocarcinoma compared to AIS/MIA
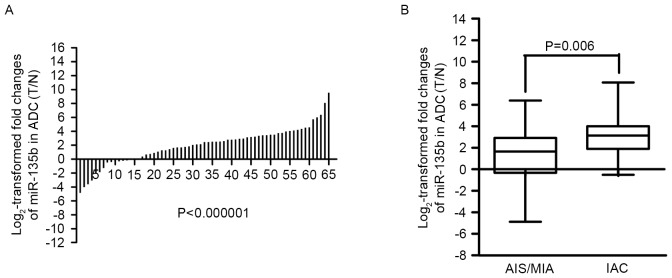
miR-135b expression was analyzed by TaqMan RT-qPCR, relative to the endogenous control U6 snRNA, in 65 paired lung adenocarcinomas and adjacent non-cancerous lung tissue specimens. Compared with the adjacent tissue, miR-135b expression was downregulated in 15 lung adenocarcinomas tissues (15/65, 23.1%) and upregulated in 50 (50/65, 76.9%). The results revealed that miR-135b is significantly upregulated in lung adenocarcinomas, compared with the adjacent non-cancerous tissue (P=7.1248×10–8) (Fig. 1A), indicating that miR-135b may promote the development of lung adenocarcinomas.
Figure 1.
Expression level of miR-135b in 65 paraffin-embedded samples. (A) miR-135b is significantly upregulated in lung adenocarcinomas compared with the corresponding para-carcinoma tissue (P<0.001). (B) miR-135b is markedly upregulated in invasive lung adenocarcinoma compared to AIS/MIA (P=0.006). miR-135b, microRNA-135b; AIS/MIA, adenocarcinoma in situ/minimally invasive adenocarcinoma; IAC, invasive adenocarcinoma; ADC, adenocarcinoma; T, lung adenocarcinoma tissue; N, adjacent normal tissue.
To evaluate the biological functions of miR-135b in peripheral lung adenocarcinomas, the expression of miR-135b was analyzed in AIS/MIA and IAC. The present study revealed that miR-135b was expressed at a higher level in IAC (P=0.006, Mann-Whitney U-test) (Fig. 1B) than AIS/MIA, which further indicated that miR-135b overexpression was associated with the invasiveness of lung adenocarcinomas.
Additionally, Pearson's χ2 test and Fisher's exact test were used to evaluate the association between miR-135b expression and clinicopathological characteristics in peripheral lung adenocarcinomas. There was no significant statistical difference between miR-135b expression and the following clinicopathological characteristics: gender, tumor size, smoking, tumor site, TNM clinical stage, carcinoembryonic antigen (CEA) levels and VPI. By contrast, higher levels of miR-135b were found in patients with peripheral lung adenocarcinoma of advanced age (≥60 years of age) (P=0.008) (Table II).
Table II.
Statistical analysis between miR-135b expression and clinicopathological characteristics.
| miR-135b Expression in ADC (T/N) | |||
|---|---|---|---|
| Variable | Overexpressing | Not overexpressing | P-valuea |
| Sex | 0.368 | ||
| Male | 17 | 4 | |
| Female | 31 | 13 | |
| Age, years | 0.008 | ||
| ≥60 | 23 | 2 | |
| <60 | 25 | 15 | |
| Tumor size, cm | 0.297 | ||
| ≥3 | 6 | 0 | |
| <3 | 42 | 17 | |
| Smoking | 0.425 | ||
| Yes | 12 | 2 | |
| No | 36 | 15 | |
| Site | 0.572 | ||
| Left | 22 | 6 | |
| Right | 26 | 11 | |
| TNM clinical stage | 0.058 | ||
| 0-IA | 32 | 16 | |
| IB-IV | 16 | 1 | |
| CEA, µg/ml | 0.171 | ||
| ≥5 | 8 | 0 | |
| <5 | 40 | 17 | |
| VPI | 0.058 | ||
| Yes | 16 | 1 | |
| No | 32 | 16 | |
Calculated using Pearson's χ2 test. miR-135b, microRNA-135b; TNM, Tumor-Node-Metastasis; CEA, carcinoembryonic antigen; VPI, visceral pleural invasion; ADC, adenocarcinoma; T, lung adenocarcinoma tissue; N, adjacent normal tissue.
EGFR mutation rate in invasive lung adenocarcinoma was higher than that in AIS/MIA
Of the 65 samples of peripheral lung adenocarcinoma in the present study, 23 were positive for EGFR mutations (35.4%), with 20% (13 cases) and 15.4% (10 cases) positive for mutations in exons 19 and 21, respectively. The in-frame deletions in exon 19 include 9 incidences of the ΔE746-A750 mutation, 1 incidence of the ΔL747-A750 mutation and 3 incidences of the ΔL747-S752 mutation, whereas the point mutations in exon 21 were 9 incidences of L858R and one incidence of L861Q.
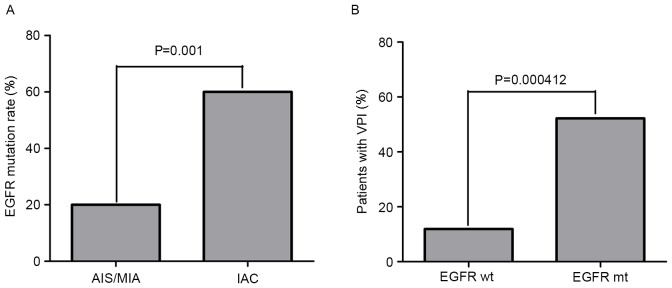
The EGFR mutation rate in IAC was 60%, which was significantly higher than that in AIS/MIA (20%) (P=0.001) (Fig. 2A), demonstrating that EGFR mutation was positively associated with the invasion of lung adenocarcinoma.
Figure 2.
EGFR mutation status in peripheral lung adenocarcinomas. (A) The EGFR mutation rate in invasive lung adenocarcinoma (60%) was higher than that in AIS/MIA (20%) (P=0.001). (B) Patients harboring EGFR mutations have a higher incidence of VPI (52.2%) than those with wild-type EGFR (11.9%) (P=0.000412). EGFR wt, patients with wild-type epithelial growth factor receptor; EGFR mt, patients harboring EGFR mutations; VPI, visceral pleural invasion; IAC, invasive adenocarcinoma.
Additionally, Pearson's χ2 test and Fisher's exact test were used to evaluate the association between EGFR mutation status and clinicopathological characteristics in peripheral lung adenocarcinomas. Higher EGFR mutation rates were observed in peripheral lung adenocarcinoma patients with characteristics including advanced age (P=0.035), large tumor (P=0.033), advanced TNM stage (P=0.001), high level of CEA (P=0.011) and visceral pleural invasion (P=0.001) (Table III). Of note is that patients harboring EGFR mutations had a higher incidence of VPI (52.2%) than those with wild-type EGFR (11.9%) (P=0.000412; Fig. 2B).
Table III.
Statistical analysis between EGFR mutation status and clinicopathological characteristics in peripheral lung adenocarcinomas.
| Mutation status of EGFR in ADC (T/N) | |||
|---|---|---|---|
| Variable | Mutated | Wild type | P-value |
| Sex | 0.154 | ||
| Male | 10 | 11 | |
| Female | 13 | 31 | |
| Age, years | 0.027 | ||
| ≥60 | 13 | 12 | |
| <60 | 10 | 30 | |
| Tumor size, cm | 0.033 | ||
| ≥3 | 5 | 1 | |
| <3 | 18 | 41 | |
| Smoking | 0.108 | ||
| Yes | 8 | 6 | |
| No | 15 | 36 | |
| Site | 0.105 | ||
| Left | 13 | 15 | |
| Right | 10 | 27 | |
| TNM stage | <0.001 | ||
| 0-IA | 11 | 37 | |
| IB-IV | 12 | 5 | |
| CEA, µg/ml | 0.011 | ||
| ≥5 | 6 | 1 | |
| <5 | 17 | 41 | |
| VPI | <0.001 | ||
| Yes | 12 | 5 | |
| No | 11 | 37 | |
Calculated using Pearson's χ2 test. miR-135b, microRNA-135b; ADC, adenocarcinoma; TNM, Tumor-Node-Metastasis; CEA, carcinoembryonic antigen; VPI, visceral pleural invasion.
Expression level of miR-135b in EGFR mutation group is higher than that in EGFR wild-type group
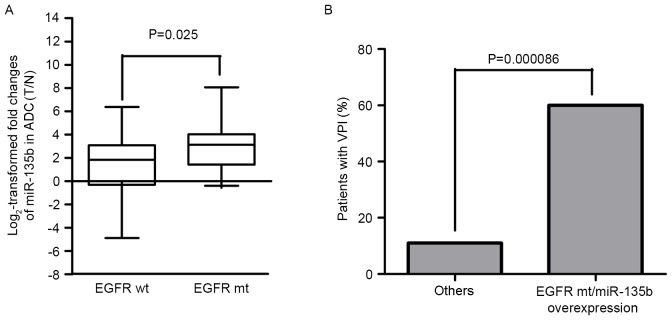
According to the aforementioned results, it appears that high expression levels of miR-135b and EGFR mutations were positively associated with the invasion of peripheral lung adenocarcinoma. To investigate whether miR-135b was associated with mutations to EGFR, the 65 cases were divided into the EGFR mutation and EGFR wild-type groups. The expression levels of miR-135b in the EGFR mutation group were significantly higher than that in EGFR wild-type group by χ2 test (P=0.020; Table IV) and Mann-Whitney U-test (P=0.025; Fig. 3A), which revealed that miR-135b expression was positively associated with mutations to EGFR. Thus, we can also consider that the peripheral lung adenocarcinomas patients harboring EGFR mutations usually have higher expression levels of miR-135b.
Table IV.
Patients harboring EGFR mutations have higher expression levels of miR-135b than those with wild-type EGFR.
| miR-135b Expression in ADC (T/N) | |||
|---|---|---|---|
| Variable | Overexpressing | Underexpressing | P-valuea |
| Mutant EGFR | 21 | 2 | 0.020 |
| Wild-type EGFR | 27 | 15 | |
Calculated using Pearson's χ2 test. miR-135b, microRNA-135b; ADC, adenocarcinoma; EGFR, epithelial growth factor receptor.
Figure 3.
Association between EGFR mutations, miR-135b and VPI. (A) Patients harboring EGFR mutations have exhibited higher expression levels of miR-135b than ones with wild-type EGFR. (P=0.025). (B) Patients harboring EGFR mutations with microRNA-135b overexpression are more likely to invade visceral pleura (P=0.000086). A total of 60% of patients harboring EGFR mutation with miR-135b overexpression had pleural invasion vs. 11% of patients that did not. EGFR wt, patients with wild-type epithelial growth factor receptor; EGFR mt, patients harboring EGFR mutations; ADC, adenocarcinoma; mir-135b, microRNA-135b; VPI, visceral pleural invasion.
Peripheral lung adenocarcinomas harboring EGFR mutations with miRNA-135b overexpression are more likely to invade visceral pleura
As a poor prognostic indicator in lung cancer, VPI can be caused by invasion of lung cancer. As aforementioned, the expression level of miR-135b was positively associated with mutations to EGFR; the two had a synergistic role in the invasion of peripheral lung adenocarcinoma. Thus, the association between miR-135b expression, EGFR mutation status and VPI was analyzed. Analysis using Pearson's χ2 and Fisher's exact tests revealed that peripheral lung adenocarcinomas harboring EGFR mutations with miR-135b overexpression (60%) were more likely to invade the visceral pleura (Fig. 3B) compared with other groups (11%) (P=0.000086; Table V). Table VI shows the association between clinicopathological characteristics and VPI. Statistical analysis revealed the following clinicopathological factors as significant: Sex, age, tumor size, smoking, TNM stage, serum CEA level, EGFR mutation status, which demonstrated that VPI is a poor prognostic predictor for patients with lung adenocarcinomas In addition, patients harboring EGFR mutations with miR-135b overexpression have higher incidence in the elderly (P=0.000119), lager tumor size (P=0.019), advanced TNM classification (P=0.000086) and VPI (P=0.000086) (Table VII).
Table V.
Peripheral lung adenocarcinomas harboring EGFR mutations with miR-135b overexpression are more likely to invade visceral pleura.
| Variable | Total | VPI | No VPI | P-valuea |
|---|---|---|---|---|
| miR-135b overexpression, mutated EGFR | 21 | 12 | 9 | <0.001 |
| No miR-135b overexpression, mutated EGFR | 2 | 0 | 2 | 0.970 |
| miR-135b overexpression, wild-type EGFR | 27 | 4 | 23 | 0.080 |
| No miR-135b overexpression, wild-type EGFR | 15 | 1 | 14 | 0.105 |
Calculated using Pearson's χ2 test. miR-135b, microRNA-135b; miR-135b, microRNA-135b; VPI, visceral pleural invasion.
Table VI.
The association between clinicopathological characteristics and VPI.
| Variable | VPI | No VPI | P-value |
|---|---|---|---|
| Sex | 0.034 | ||
| Male | 9 | 12 | |
| Female | 8 | 36 | |
| Age, years | 0.002 | ||
| ≥60 | 12 | 13 | |
| <60 | 5 | 35 | |
| Tumor size, cm | <0.001 | ||
| ≥3cm | 6 | 0 | |
| <3cm | 11 | 48 | |
| Smoking | 0.008 | ||
| Yes | 8 | 6 | |
| No | 9 | 42 | |
| Site | 0.127 | ||
| Left | 10 | 18 | |
| Right | 7 | 30 | |
| TNM stage | <0.001 | ||
| 0-IA | 0 | 48 | |
| IB-IV | 17 | 0 | |
| CEA (µg/ml) | <0.001 | ||
| ≥5 | 7 | 0 | |
| <5 | 10 | 38 | |
| EGFR mutation status | <0.001 | ||
| Mutated | 12 | 11 | |
| Wild type | 5 | 37 | |
| miR-135b | 0.058 | ||
| Overexpressed | 16 | 32 | |
| Not overexpressed | 1 | 16 |
Calculated using Pearson's χ2 test. TNM, Tumor-Node-Metastasis; CEA, carcinoembryonic antigen; VPI, visceral pleural invasion; miR-135b, microRNA-135b.
Table VII.
Association between clinicopathological characteristics and patients harboring EGFR mutations with miR-135b overexpression.
| miR-135b expression status | |||
|---|---|---|---|
| Variable | Overexpressed | Not overexpressed | P-value |
| Sex | |||
| Male | 10 | 11 | 0.068 |
| Female | 11 | 33 | |
| Age, years | |||
| ≥60 | 12 | 13 | 0.032 |
| <60 | 9 | 31 | |
| Tumor size, cm | |||
| ≥3 | 5 | 1 | 0.019 |
| <3 | 16 | 43 | |
| Smoking status | |||
| Yes | 8 | 6 | 0.055 |
| No | 13 | 38 | |
| Site | |||
| Left | 12 | 16 | 0.114 |
| Right | 9 | 28 | |
| TNM stage | |||
| 0-IA | 9 | 39 | <0.001 |
| IB-IV | 12 | 5 | |
| CEA, µg/ml | |||
| ≥5 | 7 | 5 | 0.055 |
| <5 | 16 | 42 | |
| VPI | |||
| Yes | 12 | 5 | <0.001 |
| No | 9 | 39 | |
Calculated using Pearson's χ2 test. EGFR, epithelial growth factor receptor; miR-135b, microRNA-135b; TNM, Tumor-Node-Metastasis; CEA, carcinoembryonic antigen; VPI, visceral pleural invasion.
Discussion
The latest classification of lung adenocarcinoma was raised by the International Association for the Study of Lung Cancer/American Thoracic Society/European Respiratory Society in 2011, which suggested three adenocarcinoma subtypes: AIS, MIA, and IAC (38). It is hypothesized that the development of lung adenocarcinomas may be a stepwise process triggered by the sequential accumulation of genetic and epigenetic changes from atypical adenomatous hyperplasia (AAH) to AIS, and ultimately to IAC (36). A previous study indicated that the overall survival (OS) rate of AIS was 100% and patients with MIA had ~100% OS rate if the disease was completely resected (39); however, lepidic-predominant ADC, acinar-predominant ADC, papillary predominant ADC, micropapillary predominant ADC, solid with mucin production-predominant ADC and invasive mucinous ADC exhibited 93, 67, 74, 62, 58 and 76% OS rate if the disease was completely resected, respectively (40). Owing to the highly similar patient outcomes, AIS and MIA are combined into a single category (36). On the other hand, Tsuta et al (40) suggested that AIS and MIA should be reclassified as tumor in situ. Samples were therefore divided into two groups: AIS/MIA and IAC. Peripheral lung adenocarcinoma, which arises in the peripheral lung parenchyma, represents the dominant histological subtype of all lung cancers (41) and frequently leads to aggressive local invasion and metastatic phenotypes (42,43). In this retrospective study, 65 peripheral lung adenocarcinoma samples were collected and tested for EGFR mutation status and miR-135b expression level. Mutations to EGFR and miR-135b overexpression were positively associated with the invasion of lung adenocarcinomas, which may supplement the molecular mechanism of invasiveness.
Lung cancer arises through the progressive accumulation of mutations in oncogenes and tumor-suppressor genes (42). Mutation of EGFR, the most common targetable driver mutation in advanced lung adenocarcinoma with the mutation rate varying between 43.5 and 76.6% in Asian cohorts and between 9.6 and 29.8% in non-Asian cohorts, has attracted increasing attention (44). Numerous studies have detailed the close association between lung cancer invasion and EGFR mutation status (45–47). Kadota et al (46) found that lepidic predominant adenocarcinoma tumors were statistically more likely to have activating EGFR mutations compared with other subtypes, with no EGFR mutations observed in AIS specimens and low mutation levels in MIA specimens. Soh et al (47) also found the number of EGFR mutations was increased in 39.5 and 50% of non-invasive and invasive lesions, respectively. Yoshida et al found that EGFR mutations in AAH and bronchioloalveolar carcinoma (BAC) were less frequent than those detected in invasive adenocarcinoma (48), which indicated that EGFR mutations might have a role in the multi-stage pathogenesis of lung adenocarcinoma. The present study observed the same phenomenon: Patients with invasive lesions had higher EGFR mutation rates than those without such lesions. However, EGFR mutations were not significantly associated with sex or smoking status in the present study, which is inconsistent with previous findings that EGFR mutations were more prevalent in women and non-smokers (19). Another study revealed that EGFR mutations facilitated the migration of cancer cells into the pleural cavity (49) and had largely focused on advanced-stage lung adenocarcinoma (50), while the present study found that EGFR mutations were more prevalent in the elderly (≥60 years of age), and those with larger tumors (≥3 cm), aggravated tumor TNM classification (IB-IV), higher CEA levels (≥5 µg/ml) and VPI, which demonstrated the positive association between EGFR mutations and lung adenocarcinoma progression. The number of patients analyzed in the present study was small, and EGFR mutation rates may fluctuate owing to variations in geographic patient location and methods of examination. Further studies are therefore required to confirm the aforementioned results.
miR-135b has long been regarded as an oncogenic miRNA in different cancer types (3,31–33). In the present study, the expression level of miR-135b was significantly higher in lung adenocarcinoma samples than that in corresponding non-cancerous tissue. Just as previous research has shown that miR-135b can promote lung cancer cell invasion in vitro (3), the present study found miR-135b was markedly upregulated in invasive lung adenocarcinoma compared to AIS/MIA, which may further demonstrate that miR-135b can promote the development of lung cancer by enhancing the invasiveness of tumor cells in vitro and in vivo. Thus, miR-135b may be a novel biomarker for lung cancer progression (30). In addition, miR-135b overexpression was pervasive in the elderly according to our study and patients with VPI tended to have higher levels of miR-135b (P=0.058).
Different oncogenic pathways can converge to affect the same miRNA, for example, miR-135b overexpression is associated with gene mutations in many pathways. Valeri et al (30) revealed that miR-135b overexpression is associated with mutations in specific colorectal cancer (CRC) pathways [mutations to adenomatous polyposis coli, phosphatidylinositol 3-kinase (PI3K), and SRC proto-oncogene, non-receptor tyrosine kinase]. PI3K/protein kinase B, rat sarcoma viral oncogene homolog (RAS)/mitogen-activated protein kinase and signal transducer and activator of transcription 3 (STAT3) are three major downstream pathways activated by EGFR phosphorylation, which eventually results in tumor cell proliferation, migration and metastasis, and evasion from apoptosis. A previous study also revealed that EGFR signaling is required for tumor maintenance in human lung adenocarcinoma-expressing EGFR mutants (51). EGFR mutations are hypothesized to promote the expression of miR-135b through the activation of STAT3; thus, miR-135b may be a biomarker and driver of resistance to TKI therapy (52–55). The present study found that miR-135b overexpression is significantly associated with EGFR mutation status, generally lung adenocarcinoma patients harboring EGFR mutations have higher miR-135b expression levels. As miRNAs often act as downstream effectors of protein kinases or driver genes mutated in cancer (56), miR-135b may mediate its carcinogenic role by participating in these signaling pathways. Targeting miR-135b may therefore represent a strategy to increase the specificity and overcome drug resistance to EGFR-mutation-targeted therapies.
In the present study, VPI was observed in 26.2% (17/65) of the surgically resected NSCLC specimens, which nearly matched the 26.8% reported by Kimihiro et al (55). VPI has a higher incidence in patients with large tumors, high CEA levels, advanced TNM classification, being elderly (≥60 years of age), EGFR mutations and men in the present study, which conformed with previous studies that found that VPI was observed significantly more frequently in tumors with factors indicative of tumor aggressiveness, including poorer histologic grade (57), larger tumor size (8) and high serum CEA level (57). In general, VPI in lung adenocarcinoma patients indicates an invasive and aggressive biology. Although many studies have reported the existence of an association between clinicopathological characteristics and VPI in NSCLC (34,58,59), little is known regarding the genetic changes or signaling pathways involved in the development of VPI in lung adenocarcinoma. Numerous studies have shown that the incidence of EGFR mutations in patients with MPE is significantly higher than in those without (49,60); however, few studies have indicated the association between VPI and EGFR mutations. The KRAS gene, a member of the RAS family, is a downstream factor of the EGFR signaling pathway (47), and Raparia et al (61) found that peripheral lung adenocarcinomas harboring KRAS mutations are more likely to invade the visceral pleura, suggesting that there may be an association between EGFR signaling pathway and visceral pleura involvement. In the present study, VPI was positively associated with mutations to EGFR. Furthermore, miR-135b is significantly associated with EGFR mutation status and may be a downstream effector in EGFR signaling pathway; therefore, we hypothesized that EGFR mutations and miR-135b overexpression can promote the invasion of lung adenocarcinoma through EGFR signaling pathway. Finally, lung adenocarcinoma patients harboring EGFR mutations were more likely to exhibit invasion of the visceral pleura when miR-135b was overexpressed, which further indicated the potential signaling pathway of EGFR mutations/miR-135b may promote the formation of VPI and miR-135b may be a good biomarker of the tumor progression. Additionally, targeting miR-135b may minimize drug resistance to EGFR mutations and increase therapeutic potential.
Acknowledgements
The present study was supported by grants from the Scientific Research Foundation of Ministry of Public Health of China/Major Science and Technology Program of Medicine and Health of Zhejiang Province of China (no. WKJ2014-2-021), the National Natural Science Foundation of China (no. 81502106), the Natural Science Foundation of Zhejiang Province (no. LY15H160047) the Medical Bureau of Zhejiang Province (nos. 2015ZDA032 and 2016341029), the Science and Technology Bureau of Zhoushan (no. 2015C31029) and the Science Technology Department of Zhejiang Province (no. 2016C37008).
References
- 1.Siegel R, Naishadham D, Jemal A. Cancer statistics, 2013. CA Cancer J Clin. 2013;63:11–30. doi: 10.3322/caac.21166. [DOI] [PubMed] [Google Scholar]
- 2.Meoni G, Cecere FL, Lucherini E, Di Costanzo F. Medical treatment of advanced non-small cell lung cancer in elderly patients: A review of the role of chemotherapy and targeted agents. J Geriatr Oncol. 2013;4:282–290. doi: 10.1016/j.jgo.2013.04.005. [DOI] [PubMed] [Google Scholar]
- 3.Lin CW, Chang YL, Chang YC, Lin JC, Chen CC, Pan SH, Wu CT, Chen HY, Yang SC, Hong TM, Yang PC. MicroRNA-135b promotes lung cancer metastasis by regulating multiple targets in the hippo pathway and LZTS1. Nat Commun. 2013;4:1877. doi: 10.1038/ncomms2876. [DOI] [PubMed] [Google Scholar]
- 4.Travis WD, Brambilla E, Riely GJ. New pathologic classification of lung cancer: Relevance for clinical practice and clinical trials. J Clin Oncol. 2013;31:992–1001. doi: 10.1200/JCO.2012.46.9270. [DOI] [PubMed] [Google Scholar]
- 5.Siegelin MD, Borczuk AC. Epidermal growth factor receptor mutations in lung adenocarcinoma. Lab Invest. 2014;94:129–137. doi: 10.1038/labinvest.2013.147. [DOI] [PubMed] [Google Scholar]
- 6.Lee YC, Light RW. Management of malignant pleural effusions. Respirology. 2004;9:148–156. doi: 10.1111/j.1440-1843.2004.00566.x. [DOI] [PubMed] [Google Scholar]
- 7.Rami-Porta R, Bolejack V, Crowley J, Ball D, Kim J, Lyons G, Rice T, Suzuki K, Thomas CF, Jr, Travis WD, et al. The IASLC lung cancer staging project: Proposals for the revisions of the t descriptors in the forthcoming eighth edition of the tnm classification for lung cancer. J Thorac Oncol. 2015;10:990–1003. doi: 10.1097/JTO.0000000000000559. [DOI] [PubMed] [Google Scholar]
- 8.Ou SH, Zell JA, Ziogas A, Anton-Culver H. Prognostic significance of the non-size-based AJCC T2 descriptors: Visceral pleura invasion, hilar atelectasis, or obstructive pneumonitis in stage IB non-small cell lung cancer is dependent on tumor size. Chest. 2008;133:662–669. doi: 10.1378/chest.07-1306. [DOI] [PubMed] [Google Scholar]
- 9.Manac'h D, Riquet M, Medioni J, Le Pimpec-Barthes F, Dujon A, Danel C. Visceral pleura invasion by non-small cell lung cancer: An underrated bad prognostic factor. Ann Thorac Surg. 2001;71:1088–1093. doi: 10.1016/S0003-4975(00)02649-7. [DOI] [PubMed] [Google Scholar]
- 10.Rami-Porta R, Bolejack V, Crowley J, Ball D, Kim J, Lyons G, Rice T, Suzuki K, Thomas CF, Jr, Travis WD, et al. The IASLC lung cancer staging project: Proposals for the revisions of the T descriptors in the forthcoming eighth edition of the tnm classification for lung cancer. J Thorac Oncol. 2015;10:990–1003. doi: 10.1097/JTO.0000000000000559. [DOI] [PubMed] [Google Scholar]
- 11.Kawase A, Yoshida J, Miyaoka E, Asamura H, Fujii Y, Nakanishi Y, Eguchi K, Mori M, Sawabata N, Okumura M, et al. Visceral pleural invasion classification in non-small-cell lung cancer in the 7th edition of the tumor, node, metastasis classification for lung cancer: Validation analysis based on a large-scale nationwide database. J Thorac Oncol. 2013;8:606–611. doi: 10.1097/JTO.0b013e31828632b8. [DOI] [PubMed] [Google Scholar]
- 12.Yoshida J, Nagai K, Asamura H, Goya T, Koshiishi Y, Sohara Y, Eguchi K, Mori M, Nakanishi Y, Tsuchiya R, et al. Visceral pleura invasion impact on non-small cell lung cancer patient survival: Its implications for the forthcoming TNM staging based on a large-scale nation-wide database. J Thorac Oncol. 2009;4:959–963. doi: 10.1097/JTO.0b013e3181a85d5e. [DOI] [PubMed] [Google Scholar]
- 13.Kudo Y, Saji H, Shimada Y, Nomura M, Matsubayashi J, Nagao T, Kakihana M, Usuda J, Kajiwara N, Ohira T, Ikeda N. Impact of visceral pleural invasion on the survival of patients with non-small cell lung cancer. Lung cancer. 2012;78:153–160. doi: 10.1016/j.lungcan.2012.08.004. [DOI] [PubMed] [Google Scholar]
- 14.Fibla JJ, Cassivi SD, Brunelli A, Decker PA, Allen MS, Darling GE, Landreneau RJ, Putnam JB. Re-evaluation of the prognostic value of visceral pleura invasion in Stage IB non-small cell lung cancer using the prospective multicenter ACOSOG Z0030 trial data set. Lung cancer. 2012;78:259–262. doi: 10.1016/j.lungcan.2012.09.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Takizawa T, Terashima M, Koike T, Watanabe T, Kurita Y, Yokoyama A, Honma K. Lymph node metastasis in small peripheral adenocarcinoma of the lung. J Thorac Cardiovasc Surg. 1998;116:276–280. doi: 10.1016/S0022-5223(98)70127-8. [DOI] [PubMed] [Google Scholar]
- 16.Kang JH, Kim KD, Chung KY. Prognostic value of visceral pleura invasion in non-small cell lung cancer. Eur J Cardiothorac Surg. 2003;23:865–869. doi: 10.1016/S1010-7940(03)00119-2. [DOI] [PubMed] [Google Scholar]
- 17.Hirsch FR, Scagliotti GV, Langer CJ, Varella-Garcia M, Franklin WA. Epidermal growth factor family of receptors in preneoplasia and lung cancer: Perspectives for targeted therapies. Lung cancer. 2003;1(41 Suppl):S29–S42. doi: 10.1016/S0169-5002(03)00137-5. [DOI] [PubMed] [Google Scholar]
- 18.Kazandjian D, Blumenthal GM, Yuan W, He K, Keegan P, Pazdur R. FDA approval of gefitinib for the treatment of patients with metastatic EGFR mutation-positive non-small cell lung cancer. Clin Cancer Res. 2016;22:1307–1312. doi: 10.1158/1078-0432.CCR-15-2266. [DOI] [PubMed] [Google Scholar]
- 19.Dacic S, Shuai Y, Yousem S, Ohori P, Nikiforova M. Clinicopathological predictors of EGFR/KRAS mutational status in primary lung adenocarcinomas. Mod Pathol. 2010;23:159–168. doi: 10.1038/modpathol.2009.154. [DOI] [PubMed] [Google Scholar]
- 20.Ikeda K, Nomori H, Ohba Y, Shibata H, Mori T, Honda Y, Iyama K, Kobayashi T. Epidermal growth factor receptor mutations in multicentric lung adenocarcinomas and atypical adenomatous hyperplasias. J Thorac Oncol. 2008;3:467–471. doi: 10.1097/JTO.0b013e31816b4b14. [DOI] [PubMed] [Google Scholar]
- 21.Tsai MF, Chang TH, Wu SG, Yang HY, Hsu YC, Yang PC, Shih JY. EGFR-L858R mutant enhances lung adenocarcinoma cell invasive ability and promotes malignant pleural effusion formation through activation of the CXCL12-CXCR4 pathway. Sci Rep. 2015;5:13574. doi: 10.1038/srep13574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Shi X, Wu H, Lu J, Duan H, Liu X, Liang Z. Screening for major driver oncogene alterations in adenosquamous lung carcinoma using PCR coupled with next-generation and Sanger sequencing methods. Sci Rep. 2016;6:22297. doi: 10.1038/srep22297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Mok TS, Wu YL, Thongprasert S, Yang CH, Chu DT, Saijo N, Sunpaweravong P, Han B, Margono B, Ichinose Y, et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N Engl J Med. 2009;361:947–957. doi: 10.1056/NEJMoa0810699. [DOI] [PubMed] [Google Scholar]
- 24.Yang Y, Shi C, Sun H, Yin W, Zhou X, Zhang L, Jiang G. Elderly male smokers with right lung tumors are viable candidates for KRAS mutation screening. Sci Rep. 2016;6:18566. doi: 10.1038/srep18566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Gasparini P, Cascione L, Landi L, Carasi S, Lovat F, Tibaldi C, Alì G, D'Incecco A, Minuti G, Chella A, et al. microRNA classifiers are powerful diagnostic/prognostic tools in ALK-, EGFR-, and KRAS-driven lung cancers. Proc Natl Acad Sci USA. 2015;112:14924–14929. doi: 10.1073/pnas.1520329112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Song Q, Xu Y, Yang C, Chen Z, Jia C, Chen J, Zhang Y, Lai P, Fan X, Zhou X, et al. miR-483-5p promotes invasion and metastasis of lung adenocarcinoma by targeting RhoGDI1 and ALCAM. Cancer Res. 2014;74:3031–3042. doi: 10.1158/0008-5472.CAN-13-2193. [DOI] [PubMed] [Google Scholar]
- 27.Fan C, Lin Y, Mao Y, Huang Z, Liu AY, Ma H, Yu D, Maitikabili A, Xiao H, Zhang C, et al. MicroRNA-543 suppresses colorectal cancer growth and metastasis by targeting KRAS, MTA1 and HMGA2. Oncotarget. 2016;7:21825–21839. doi: 10.18632/oncotarget.7989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A, Ebert BL, Mak RH, Ferrando AA, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834–838. doi: 10.1038/nature03702. [DOI] [PubMed] [Google Scholar]
- 29.Yang WB, Chen PH, Hsu T I, Fu TF, Su WC, Liaw H, Chang WC, Hung JJ. Sp1-mediated microRNA-182 expression regulates lung cancer progression. Oncotarget. 2014;5:740–753. doi: 10.18632/oncotarget.1608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Valeri N, Braconi C, Gasparini P, Murgia C, Lampis A, Paulus-Hock V, Hart JR, Ueno L, Grivennikov SI, Lovat F, et al. MicroRNA-135b promotes cancer progression by acting as a downstream effector of oncogenic pathways in colon cancer. Cancer cell. 2014;25:469–483. doi: 10.1016/j.ccr.2014.03.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Nagel R, le Sage C, Diosdado B, van der Waal M, Vrielink Oude JA, Bolijn A, Meijer GA, Agami R. Regulation of the adenomatous polyposis coli gene by the miR-135 family in colorectal cancer. Cancer Res. 2008;68:5795–5802. doi: 10.1158/0008-5472.CAN-08-0951. [DOI] [PubMed] [Google Scholar]
- 32.Li Y, Xu D, Bao C, Zhang Y, Chen D, Zhao F, Ding J, Liang L, Wang Q, Liu L, et al. MicroRNA-135b, a HSF1 target, promotes tumor invasion and metastasis by regulating RECK and EVI5 in hepatocellular carcinoma. Oncotarget. 2015;6:2421–2433. doi: 10.18632/oncotarget.2965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Zhang L, Sun ZJ, Bian Y, Kulkarni AB. MicroRNA-135b acts as a tumor promoter by targeting the hypoxia-inducible factor pathway in genetically defined mouse model of head and neck squamous cell carcinoma. Cancer Lett. 2013;331:230–238. doi: 10.1016/j.canlet.2013.01.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Wu W, Wang Z, Yang P, Yang J, Liang J, Chen Y, Wang H, Wei G, Ye S, Zhou Y. MicroRNA-135b regulates metastasis suppressor 1 expression and promotes migration and invasion in colorectal cancer. Mol Cell Biochem. 2014;388:249–259. doi: 10.1007/s11010-013-1916-z. [DOI] [PubMed] [Google Scholar]
- 35.Agalioti T, Giannou AD, Stathopoulos GT. Pleural involvement in lung cancer. J Thorac Dis. 2015;7:1021–1030. doi: 10.3978/j.issn.2072-1439.2015.04.23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Travis WD, Brambilla E, Noguchi M, Nicholson AG, Geisinger KR, Yatabe Y, Beer DG, Powell CA, Riely GJ, Van Schil PE, et al. International association for the study of lung cancer/american thoracic society/european respiratory society international multidisciplinary classification of lung adenocarcinoma. J Thorac Oncol. 2011;6:244–285. doi: 10.1097/JTO.0b013e318206a221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25:402–408. doi: 10.1006/meth.2001.1262. [DOI] [PubMed] [Google Scholar]
- 38.Kadota K, Villena-Vargas J, Yoshizawa A, Motoi N, Sima CS, Riely GJ, Rusch VW, Adusumilli PS, Travis WD. Prognostic significance of adenocarcinoma in situ, minimally invasive adenocarcinoma, and nonmucinous lepidic predominant invasive adenocarcinoma of the lung in patients with stage I disease. Am J Surg Pathol. 2014;38:448–460. doi: 10.1097/PAS.0000000000000134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yoshizawa A, Motoi N, Riely GJ, Sima CS, Gerald WL, Kris MG, Park BJ, Rusch VW, Travis WD. Impact of proposed IASLC/ATS/ERS classification of lung adenocarcinoma: Prognostic subgroups and implications for further revision of staging based on analysis of 514 stage I cases. Mod Pathol. 2011;24:653–664. doi: 10.1038/modpathol.2010.232. [DOI] [PubMed] [Google Scholar]
- 40.Tsuta K, Kawago M, Inoue E, Yoshida A, Takahashi F, Sakurai H, Watanabe S, Takeuchi M, Furuta K, Asamura H, Tsuda H. The utility of the proposed IASLC/ATS/ERS lung adenocarcinoma subtypes for disease prognosis and correlation of driver gene alterations. Lung cancer. 2013;81:371–376. doi: 10.1016/j.lungcan.2013.06.012. [DOI] [PubMed] [Google Scholar]
- 41.Cagle PT, Allen TC, Dacic S, Beasley MB, Borczuk AC, Chirieac LR, Laucirica R, Ro JY, Kerr KM. Revolution in lung cancer: New challenges for the surgical pathologist. Arch Pathol Lab Med. 2011;135:110–116. doi: 10.5858/2010-0567-RA.1. [DOI] [PubMed] [Google Scholar]
- 42.Eto T, Suzuki H, Honda A, Nagashima Y. The changes of the stromal elastotic framework in the growth of peripheral lung adenocarcinomas. Cancer. 1996;77:646–656. doi: 10.1002/(SICI)1097-0142(19960215)77:4<646::AID-CNCR10>3.0.CO;2-0. [DOI] [PubMed] [Google Scholar]
- 43.Nakanishi H, Matsumoto S, Iwakawa R, Kohno T, Suzuki K, Tsuta K, Matsuno Y, Noguchi M, Shimizu E, Yokota J. Whole genome comparison of allelic imbalance between noninvasive and invasive small-sized lung adenocarcinomas. Cancer Res. 2009;69:1615–1623. doi: 10.1158/0008-5472.CAN-08-3218. [DOI] [PubMed] [Google Scholar]
- 44.Clay TD, Russell PA, Do H, Sundararajan V, Conron M, Wright GM, Dobrovic A, Moore MM, McLachlan SA. Associations between the IASLC/ATS/ERS lung adenocarcinoma classification and EGFR and KRAS mutations. Pathology. 2016;48:17–24. doi: 10.1016/j.pathol.2015.11.002. [DOI] [PubMed] [Google Scholar]
- 45.Yatabe Y, Takahashi T, Mitsudomi T. Epidermal growth factor receptor gene amplification is acquired in association with tumor progression of EGFR-mutated lung cancer. Cancer Res. 2008;68:2106–2111. doi: 10.1158/0008-5472.CAN-07-5211. [DOI] [PubMed] [Google Scholar]
- 46.Kadota K, Yeh YC, D'Angelo SP, Moreira AL, Kuk D, Sima CS, Riely GJ, Arcila ME, Kris MG, Rusch VW, et al. Associations between mutations and histologic patterns of mucin in lung adenocarcinoma: Invasive mucinous pattern and extracellular mucin are associated with KRAS mutation. Am J Surg Pathol. 2014;38:1118–1127. doi: 10.1097/PAS.0000000000000246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Soh J, Toyooka S, Ichihara S, Asano H, Kobayashi N, Suehisa H, Otani H, Yamamoto H, Ichimura K, Kiura K, et al. Sequential molecular changes during multistage pathogenesis of small peripheral adenocarcinomas of the lung. J Thorac Oncol. 2008;3:340–347. doi: 10.1097/JTO.0b013e318168d20a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Yoshida Y, Shibata T, Kokubu A, Tsuta K, Matsuno Y, Kanai Y, Asamura H, Tsuchiya R, Hirohashi S. Mutations of the epidermal growth factor receptor gene in atypical adenomatous hyperplasia and bronchioloalveolar carcinoma of the lung. Lung cancer. 2005;50:1–8. doi: 10.1016/j.lungcan.2005.04.012. [DOI] [PubMed] [Google Scholar]
- 49.Han HS, Eom DW, Kim JH, Kim KH, Shin HM, An JY, Lee KM, Choe KH, Lee KH, Kim ST, et al. EGFR mutation status in primary lung adenocarcinomas and corresponding metastatic lesions: Discordance in pleural metastases. Clin Lung Cancer. 2011;12:380–386. doi: 10.1016/j.cllc.2011.02.006. [DOI] [PubMed] [Google Scholar]
- 50.Mok T, Wu YL, Zhang L. A small step towards personalized medicine for non-small cell lung cancer. Discov Med. 2009;8:227–231. [PubMed] [Google Scholar]
- 51.Mitsudomi T, Yatabe Y. Mutations of the epidermal growth factor receptor gene and related genes as determinants of epidermal growth factor receptor tyrosine kinase inhibitors sensitivity in lung cancer. Cancer Sci. 2007;98:1817–1824. doi: 10.1111/j.1349-7006.2007.00607.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Alvarez JV, Greulich H, Sellers WR, Meyerson M, Frank DA. Signal transducer and activator of transcription 3 is required for the oncogenic effects of non-small-cell lung cancer-associated mutations of the epidermal growth factor receptor. Cancer Res. 2006;66:3162–3168. doi: 10.1158/0008-5472.CAN-05-3757. [DOI] [PubMed] [Google Scholar]
- 53.Matsuyama H, Suzuki HI, Nishimori H, Noguchi M, Yao T, Komatsu N, Mano H, Sugimoto K, Miyazono K. miR-135b mediates NPM-ALK-driven oncogenicity and renders IL-17-producing immunophenotype to anaplastic large cell lymphoma. Blood. 2011;118:6881–6892. doi: 10.1182/blood-2011-05-354654. [DOI] [PubMed] [Google Scholar]
- 54.Takata S, Takigawa N, Segawa Y, Kubo T, Ohashi K, Kozuki T, Teramoto N, Yamashita M, Toyooka S, Tanimoto M, Kiura K. STAT3 expression in activating EGFR-driven adenocarcinoma of the lung. Lung Cancer. 2012;75:24–29. doi: 10.1016/j.lungcan.2011.05.015. [DOI] [PubMed] [Google Scholar]
- 55.Khatri R, Subramanian S. MicroRNA-135b and its circuitry networks as potential therapeutic targets in colon cancer. Front Oncol. 2013;3:268. doi: 10.3389/fonc.2013.00268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Croce CM. Causes and consequences of microRNA dysregulation in cancer. Nat Rev Genet. 2009;10:704–714. doi: 10.1038/nrg2634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Shimizu K, Yoshida J, Nagai K, Nishimura M, Ishii G, Morishita Y, Nishiwaki Y. Visceral pleural invasion is an invasive and aggressive indicator of non-small cell lung cancer. J Thorac Cardiovasc Surg. 2005;130:160–165. doi: 10.1016/j.jtcvs.2004.11.021. [DOI] [PubMed] [Google Scholar]
- 58.Riquet M, Badoual C, Le Pimpec Barthes F, Lhote FM, Souilamas R, Hubsch JP, Danel C. Visceral pleura invasion and pleural lavage tumor cytology by lung cancer: A prospective appraisal. Ann Thorac Surg. 2003;75:353–355. doi: 10.1016/S0003-4975(02)04403-X. [DOI] [PubMed] [Google Scholar]
- 59.Butnor KJ, Cooper K. Visceral pleural invasion in lung cancer: Recognizing histologic parameters that impact staging and prognosis. Adv Anat Pathol. 2005;12:1–6. doi: 10.1097/01.pap.0000151266.26814.02. [DOI] [PubMed] [Google Scholar]
- 60.Zou J, Bella AE, Chen Z, Han X, Su C, Lei Y, Luo H. Frequency of EGFR mutations in lung adenocarcinoma with malignant pleural effusion: Implication of cancer biological behaviour regulated by EGFR mutation. J Int Med Res. 2014;42:1110–1117. doi: 10.1177/0300060514539273. [DOI] [PubMed] [Google Scholar]
- 61.Raparia K, Villa C, Raj R, Cagle PT. Peripheral lung adenocarcinomas with KRAS mutations are more likely to invade visceral pleura. Arch Pathol Lab Med. 2015;139:189–193. doi: 10.5858/arpa.2013-0759-OA. [DOI] [PubMed] [Google Scholar]