Abstract
Sub-optimal early life environments can influence life-long phenotypes, and epigenetic factors are thought to be key mediators of these effects. We show that early life protein restriction in mice induces a linear correlation between growth restriction and DNA methylation at a specific genetic variant of ribosomal DNA (rDNA), which is present long after exposure has ceased. Related effects are also found in models of maternal high-fat or obesogenic diets. Our work defines a unique example of gene-environment induced epigenetic dynamics, and establishes rDNA as the only known genomic target of nutritional insults found in mammalian and non-mammalian models.
There is now strong evidence that the in utero environment can influence adult phenotypes and disease outcomes in mammals, a process termed ‘developmental programming’ (1, 2). Consequently, there is great interest in identifying the molecular mechanisms that underlie developmental programming, and in this regard modulation of the epigenome has emerged as a plausible contributing factor (3, 4).
To explore epigenetic mechanisms involved in developmental programming effects, we used the well-established maternal protein restriction model (5). We set up G0 matings using C57BL/6J mice exclusively, and the following morning G0 females were assigned to either a PR (8% protein) or standard (20% protein) diet (Table S1) until their G1 offspring were weaned. Only male G1 offspring were studied in detail. From weaning onwards, both ‘PR’ and ‘Control’ G1 males were kept on standard diet until sacrifice at 16-20 weeks. A total of 146 animals were generated for analysis. At the phenotypic level, we found that G1-PR males were ~25% lighter than Controls at weaning (Fig 1A; P = 2 x 10-6, female pups were also lighter but not studied further, Fig S1), consistent with previous work (5). PRs also displayed reduced spontaneous locomotor activity (Fig S2) and reduced glucose-stimulated insulin secretion (Fig S3).
Figure 1. Maternal protein restriction (PR) induces a correlation between rDNA methylation and weaning weight.
A. Statistical tests were performed using litter means (large symbols (nL)), individuals are represented with small symbols (n). Control = black, PR = red. Litters of 5-10 pups included. Weaning weight of PR males (n = 83, nL = 17) were reduced compared to Control (n = 135, nL = 20) (left; t-test, P = 2 x10-6).
B. Methylation across the rDNA promoter and coding region was determined in sperm of using RRBS. PR sperm (red, n = 8) is relatively hypermethylated compared to Control (black, n = 8). Mean methylation % (line) and individual mice (points) are shown. The relative position of the transcriptional start site (TSS) and rRNA subunits are shown (top). The region of the rDNA 98% homologous to the differentially methylated region on chromosome 17 (mm10) is shaded blue.
C. The correlation coefficient (τ) between weaning weight and DNA methylation for each CpG site covered (> 100X) with RRBS is shown for the rDNA promoter and coding region in sperm. Control (black, n = 8), PR sperm (red, n = 8). Illustrations for the derivation of the correlation coefficient are provided when positive (green point), close to zero (purple point) and negative (orange point) on the left hand side. The relative position of the transcriptional start site (TSS) and rRNA subunits are shown (top). CpG-133 is circled in blue. Non coding regions include the promoter, ETS = external transcribed spacer, ITS = internal transcribed spacer.
We initially focussed on mature sperm in an effort to identify epigenetic perturbations that are found in the directly exposed individuals and are also potential candidates for intergenerational effects, although here we focus on the effects observed in the directly exposed G1 males. Several recent studies have provided evidence that DNA methylation profiles can be perturbed by developmental programming effects (1). We therefore performed genome-scale, single base resolution DNA methylation analysis in sperm from 8 PR and 8 Control G1s using reduced representation bisulfite sequencing (RRBS). After genome-wide correction, we identified a single 1916 bp differentially methylated region (DMR) hypermethylated in G1-PR males, that mapped to a gene annotated as Rn45s on chromosome 17 (mm10, Table S2) (NR_046233). Closer examination revealed that this DMR was 98% homologous to the 973-2883 bp region of the ribosomal DNA (rDNA) consensus (Fig 1B), which is excluded from genome assemblies because of its highly repetitivee, multi-copy structure. Re-mapping of RRBS data to the consensus sequence for mouse rDNA (BK000964) confirmed extensive hypermethylation in PR sperm across the entire rDNA promoter and coding region (~13.5 kb) (Fig 1B). Additional MeDIP-seq profiling (8 individual PR and 8 Control samples), and low coverage whole genome bisulfite sequencing (WGBS), did not yield any other genome-wide corrected hits (data not shown but available from GEO).
As PR males showed both reduced weaning weight (Fig 1A) and hypermethylation of rDNA in sperm (Fig 1B), we assessed if these two variables are directly correlated (Fig 1C). PR males displayed a significantly greater negative correlation between weaning weight and DNA methylation compared to Control (Wilcoxon rank sum test; P < 2.2 x 10-16). This correlation was not confounded by weight or age at death (Fig S4). We therefore decided to perform a more detailed analysis of rDNA methylation dynamics in the rest of the G1 males in our study. In the C57BL/6J genome, rDNA is comprised of hundreds of copies in large arrays on chromosomes 12, 15, 18 and 19, but only a subset are actively transcribed (6). Constitutively silenced copies are methylated at a CpG site located 133 bp upstream of the 45S-rRNA transcriptional start site (Fig 1C), and this prevents binding of the transcription factor UBF and assembly of RNA polymerase I (7). We further investigated CpG-133 methylation states by high-throughput sequencing of bisulfite PCR amplicons generated on the Fluidigm microfluidics platform (henceforth termed bisPCR-seq; amplicons were ~300 bp in length and sequenced to >1000X coverage). BisPCR-seq analysis of the same samples profiled by RRBS revealed strong concordance between the two methods (Fig S5, τ = 0.77, P = 1 x 10-5).
As rDNA copies within a single genome are genetically polymorphic, and can be independently regulated (8), we designed the bisPCR-seq amplicon targeting CpG-133 to simultaneously assay previously documented genetic variation at position -104 (C or A, Fig 2A. Note: this variant does not overlap a CpG site) (9). Interestingly, CpG-133 methylation levels were substantially lower for the C-variant relative to the A-variant (Fig 2A) and there was no interaction between C-variant associated CpG-133 methylation and weaning weight in PR or Control sperm (Fig S6). On the other hand, CpG-133 methylation levels of A-variant rDNA (which we denote as CpG-133A) were negatively correlated with weaning weight (Fig 2B; τ = -0.43, P = 0.017). Figure 2B incorporates additional independent males (9 PR and 7 Controls from a range of different litters not represented in the RRBS data), reinforcing the negative correlation between weaning weight and total CpG-133 methylation observed in the discovery RRBS dataset. BisPCR-seq analysis of in vitro methylated samples confirmed that there was no amplification bias associated with either the A- or C-variants (Fig S7). We also confirmed sperm purity by analysis of several different parentally imprinted regions (Fig S8). BisPCR-seq analysis of liver showed strong correlation with sperm within individual Control (Fig S9; τ = 0.72, P = 0.00028) or PR animals (Fig S9; τ = 0.54, P = 0.0041). Like for sperm, liver CpG-133A methylation was negatively correlated with weaning weight in PRs (τ = -0.46, n = 24, P = 0.0016) but not in Controls (n = 26) (Fig 2C). Collectively, these data demonstrate that PR exposure induces not just rDNA hypermethylation, but also a linear relationship between a phenotypic outcome (weaning weight) and CpG-133A methylation in sperm and liver, long after exposure had ceased.
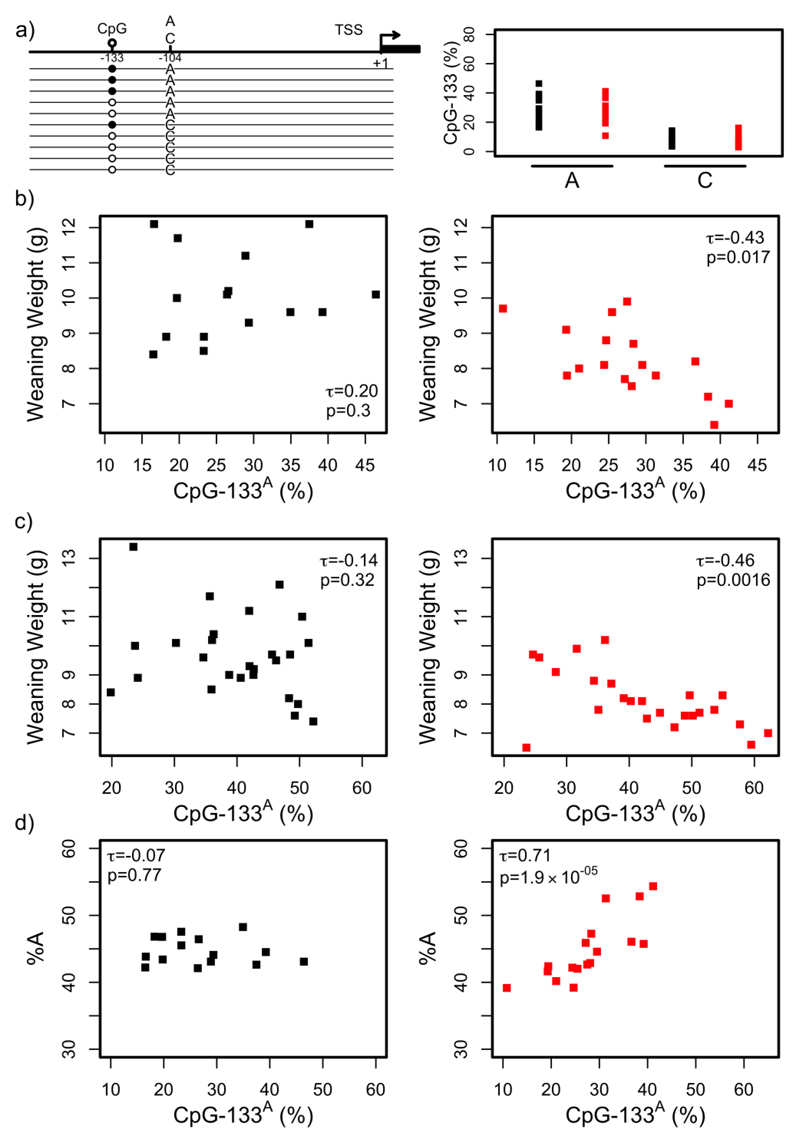
Figure 2. Diet induced dynamics are restricted to a specific genetic variant of rDNA and reflect the relative copy number of that variant.
A. BisPCR-seq amplicons were generated spanning both CpG-133 (methylation is indicated by black circle) and the genetic variant (A or C) at position -104, relative to the TSS (left panel). The percentage of CpG-133 methylation in sperm for each genetic variant is shown for Control sperm (black, n=15), and PR sperm (red, n=17).
B. Methylation of CpG-133 in A-variant rDNA copies (CpG-133A, x-axis) does not correlate with weaning weight (y-axis) in Control sperm (black, n=15; τ = 0.20, P = 0.30). Methylation of CpG-133 in A-variant rDNA copies (CpG-133A, x-axis) correlates negatively with weaning weight (y-axis) in PR sperm (red, n=17; τ = -0.43, P = 0.017).
C. Methylation of CpG-133 in A-variant rDNA copies (CpG-133A, x-axis) does not correlate with weaning weight (y-axis) in Control liver (black, n=26; τ = -0.14, P = 0.32). Methylation of CpG-133 in A-variant rDNA copies (CpG-133A, x-axis) correlates negatively with weaning weight (y-axis) in PR liver (red, n=24; τ = -0.46, P =0.0016).
D. Methylation of CpG-133 in A-variant rDNA copies (CpG-133A, x-axis) does not correlate with the percentage of total rDNA copies with an A-variant (%A, y-axis) in Control sperm (black, n=15; τ = -0.07, P = 0.77). Methylation of CpG-133 in A-variant rDNA copies (CpG-133A, x-axis) correlates positively with the percentage of total rDNA copies with an A-variant (%A, y-axis) in PR sperm (red, n=17; τ = 0.71, P = 1.9 x 10-5).
Further exploration of the bisPCR-seq data revealed unexpected inter-individual variation in the relative copy number of rDNA associated with the A-variant at position -104, even in an inbred genetic background (C57BL/6J). This underlying variation in relative copy number (which we denote as ‘%A’ i.e. the percentage of A-variant reads relative to all reads in the bisPCR-seq amplicon targeting this site) was positively correlated between sperm and liver of both Control (Fig S10; τ = 0.77, P = 7 x 10-5) and PR animals (Fig S10; τ = 0.73, P = 3.7 x 10-5). The accuracy of the bisPCR-seq derived estimates of %A were confirmed by whole genome re-sequencing of 6 different animals (Fig S11; τ = 1, P = 0.0028). Furthermore, we found that in PR G1 males, CpG-133A methylation correlated positively with %A in both the sperm (Fig 2D; τ = 0.71, P = 1.9 x 10-5) and liver (Fig S12; τ = 0.31, P = 0.034), but there was no such interaction in the sperm (Fig 2D) or liver (Fig S12) of Controls. Therefore, early life PR induces an interdependence between underlying variation in the relative abundance of a specific genetic variant of rDNA and methylation state of a linked and functionally relevant CpG site.
rDNA copies that lack methylation at CpG-133 have the potential to be transcriptionally active (7). As most methylation is localised to A-variant rDNA in our model, both the level of methylation at CpG-133A and the relative abundance of this variant (i.e. %A) will contribute towards transcriptional competency. This interaction can be represented as the percentage of total rDNA copies that are both A-variant and unmethylated at CpG-133 (which we denote as ‘%AUN’. Note: %AUN is different to simply considering the percentage of CpG-133A that is unmethylated). As expected, %AUN correlates between the sperm and liver of Controls and PR mice (Fig S13). To confirm the functional significance of %AUN, we analysed a regulatory non-coding RNA (promoter associated RNA; pRNA) that spans the rDNA promoter (Fig 3A). pRNA is transcribed from early replicating and unmethylated rDNA copies (10). It functions in trans to recruit nucleolar chromatin remodelling complex and DNA methyltransferase to silenced rDNA copies (11). Using RT-qPCR, we generated a pRNA-derived amplicon spanning the genetic polymorphism at position -104, and determined the percentage of A-variant reads after high throughput sequencing (pRNA(%A)). The pRNA(%A) reads in liver were consistently and positively correlated with %AUN (Fig 3B), but not %A, (Fig S14). Therefore, %AUN is indicative of transcriptional competency at rDNA.
Figure 3. Functional consequences of altered rDNA dynamics.
A. pRNA is transcribed from early replicating rDNA copies (assumed to be unmethylated at CpG-133). If this assumption is true, the percentage of pRNA reads that encode an A at position -104 (pRNA(%A); indicated in blue; right) is a reflection of not only the relative abundance of A-variant rDNA copies (%A; indicated by A in black; left), but also the methylation of the A-variant (CpG-133A, methylation is indicated by black circles).
B. The percentage of pRNA reads that encode an A at position -104 (pRNA(%A); y-axis) positively correlates with the percentage of unmethylated A-variant reads as a total of all rDNA copies (%AUN) in both Control (black) and PR (red) liver (total, n=23, τ = 0.61, P = 1.4 x10-5).
C. The percentage of unmethylated A-variant reads as a total of all rDNA copies (%AUN, x-axis) is not correlated with the abundance of unprocessed 45S-rRNA (y-axis) in liver of Control (black; n=14, τ = 0.03, P = 0.91), but is positively correlated in liver of PR (red; n=12, τ = 0.52, P = 0.021).
We then examined the consequences of transcriptional competency on 45S-rRNA expression. The 45S-rRNA is co-transcriptionally cleaved at position +650 within the 5’ external transcribed spacer, and the first 650 nt is then rapidly degraded (12). We assessed the abundance of the nascent, uncleaved 45S-rRNA precursor via RT-qPCR targeting the first 650 nt. In the liver of Control animals, 45S-rRNA abundance did not correlate with CpG-133A methylation, %A, or %AUN (Fig 3C, Fig S15). In PR males, 45S-rRNA levels did not correlate with CpG-133A methylation or %A, but correlated positively with %AUN (Fig 3C; τ = 0.52, P = 0.021, Fig S15). Therefore, PR exposure induces a correlation between transcriptional competency and 45S-rRNA levels.
As rDNA expression is sensitive to general nutrient availability (13), the types of effects we describe could be a conserved feature of other developmental programming models of nutritional insults. To explore this further, we identified a recent study by Cannon et al (2104) in which the authors fed C57BL/6J females a low- or high-fat (LF and HF respectively) diet from 3 weeks prior to pregnancy up till when the male G1 offspring of each group were weaned onto either LF or HF diet (14). The authors did not identify any diet-induced effects on DNA methylation from RRBS data generated from the liver of offspring at 9 weeks of age. We mapped their raw sequencing reads to rDNA and found that in utero exposure to HF induces CpG-133A hypermethylation in the G1s, regardless of post-weaning diet (Fig 4A, P = 0.0098 for post-weaning LF diet and P = 0.0079 for post-weaning HF diet). Unfortunately, there were insufficient mice to examine correlations between %A and methylation or weaning weight. Encouraged by our re-analysis of Cannon et al’s data, we performed bisPCR-seq analysis of G1 male C57BL/6J mice we previously generated as part of another study of maternal obesogenic diet (this contains elevated fat and sugar content and hence is thought to better mimic obesity promoting diets in Western societies). C57BL/6J G0 females were fed either control or obesogenic diet 6 weeks prior to mating. The G1 offspring, after weaning, were placed on either control or obesogenic diets until sacrifice at 6 months, resulting in 4 different groups of G1 mice (Figure 4B). G1s exposed in utero to obesogenic diet and then kept on control diet post-weaning showed hypermethylation at CpG-133A (P = 0.017) relative to G1 animals exposed to control diet only. Collectively, these results demonstrate that a variety of suboptimal early life dietary exposures elicit DNA methylation responses at specific genetic variants of rDNA in the C57BL/6J strain.
Figure 4. High fat or Obesogenic diet perturbs rDNA and a model for PR induced rDNA interactions.
A. RRBS raw sequencing reads (obtained from Cannon et al., 2014) were mapped to the rDNA consensus. Dams were fed either a low (ML) or high fat (MH) diet prior to conception and up until the pups were weaned. At weaning pups were placed onto either a low (AL) or high fat (AH) diet, to give rise to four dietary categories as indicated (x-axis). Data is from liver of 9 week old male offspring. A-variant CpG-133 methylation (CpG-133A, y-axis) was significantly higher in the liver of 9 week old male offspring fed MHAL (t-test, P = 0.0098) and MHAH (t-test, P = 0.0079), compared to MLAL. n=10 for each dietary group.
B. Dams were fed either a Control (C) or Obesogenic (O) diet 6 weeks prior to conception and up until the pups were weaned. At weaning pups were placed onto either a Control (C) or Obesogenic (O) diet, to give rise to four dietary categories as indicated (x-axis). Data is from liver of 6 month old male offspring. A-variant CpG-133 methylation (CpG-133A, y-axis) was significantly higher in the liver of 6 month old male offspring exposed in early development to an obesogenic diet, but weaned onto control diet (OC, t-test, P = 0.017), compared to CC. (CC: n=7, CO: n=8, OC: n=8, OO: n=7).
C. There is natural variation in the relative copy number of rDNA with an A-variant at position -104 (%A, indicated left, grey). Methylation is established independently of copy number shortly after fertilisation and this is maintained throughout development in Control fed animals. However, in animals fed a PR diet, animals with higher %A silence more A-variants through methylation at CpG-133 and this correlates with reduced weaning weight. Once established, this methylation pattern is maintained into adulthood, in the absence of further exposure.
Here we have described a unique example of a mammalian ‘epiallele’ – a non-transposable endogenous genomic region whose epigenetic state is influenced by in utero gene-environment interactions, concomitant with transcriptional and phenotypic outcomes. A schematic model is presented in Fig 4C. Interestingly, differential methylation at Rn45S was also recently identified in an outbred mouse model of in utero caloric restriction (CR) (4), although the authors did not characterise the region further. Therefore, our work in combination with previous demonstrations in flies and yeast (15, 16), identifies rDNA as the only known genomic target of nutritional insults that is conserved amongst non-mammalian and mammalian models.
Very recently, Shea et al., (2015) reported a study in which they exposed male C57BL/6J mice to one of three different diets (PR, HF and CR) post-weaning (17). They identified substantial inter-individual genetic and methylomic variability at rDNA, but no consistent diet induced effects. Although part of the reason for the discrepant conclusions could be that they don’t discriminate between the ‘A’ or ‘C’ genetic variants, we propose that the more likely explanation is differences in developmental timing of the dietary insults. All models in our study have a component of in utero exposure. In contrast, all of Shea et al.’s models involve exposure post-weaning only. Indeed, previous human epidemiological and animal studies suggest that fetal life is a critical time window when suboptimal nutritional exposures can have a long-term effect on the physiology and metabolism of the offspring (18).
Epigenetic silencing of rDNA in response to nutrient levels has been demonstrated in human cells (13), and rDNA copies display an order of magnitude variability in human populations (19). Our preliminary analysis of human rDNA sequences from the 1000 genomes project suggests surprisingly extensive genetic variation in humans (data not shown). This, taken together with the gene-environment induced epiallelic effects at rDNA we report here, leads us to speculate that systematic characterisation of genetic-epigenetic interactions at rDNA could provide novel insights into the molecular basis of some human complex phenotypes and diseases.
Supplementary Material
Acknowledgements
This work was supported by the following grants and fellowships; Biotechnology and Biological Sciences Research Council, UK (BB/M012494/1) to VKR, and (BB/G00711/X/1) to VKR and CG. Research Council UK Academic Fellowship to MLH. RL is supported by EU-FP7 BLUEPRINT. British Heart Foundation (FS/12/64/30001) and MRC (MC_UU_12012/4) to SEO. We thank King’s College London FWB Genomics Centre and Barts and The London Genome Centre for performing high throughput sequencing.
Footnotes
Author contributions
MLH, CG, PWC, VKR, AAMC and EL performed all experiments. RL and GC conducted the bioinformatic analyses. VKR and SEO provided reagents. MLH, RL, VKR conceived the study and prepared the manuscript. All authors discussed the results and interpretation and approved the final manuscript.
Data deposition statement
Whole Genome sequencing data and pRNA sequencing was submitted to BioProject under ID (PRJNA293403). RRBS, RNA-Seq and Bis-PCR were all submitted to the NCBI Gene Expression Omnibus under accession (GSE72610). All data will be linked under the BioProject ID.
Competing interest declaration
We declare no conflict of interest
References
- 1.Fernandez-Twinn DS, Constancia M, Ozanne SE. Intergenerational epigenetic inheritance in models of developmental programming of adult disease. Seminars in cell & developmental biology. 2015 doi: 10.1016/j.semcdb.2015.06.006. published online EpubJun 30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Hanson MA, Gluckman PD. Early developmental conditioning of later health and disease: physiology or pathophysiology? Physiological reviews. 2014;94:1027–1076. doi: 10.1152/physrev.00029.2013. published online EpubOct. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Tobi EW, Goeman JJ, Monajemi R, Gu H, Putter H, Zhang Y, Slieker RC, Stok AP, Thijssen PE, Muller F, van Zwet EW, et al. DNA methylation signatures link prenatal famine exposure to growth and metabolism. Nature communications. 2014;5:5592. doi: 10.1038/ncomms6592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Radford EJ, Ito M, Shi H, Corish JA, Yamazawa K, Isganaitis E, Seisenberger S, Hore TA, Reik W, Erkek S, Peters AH, et al. In utero effects. In utero undernourishment perturbs the adult sperm methylome and intergenerational metabolism. Science. 2014;345 doi: 10.1126/science.1255903. 1255903; published online EpubAug 15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Vucetic Z, Totoki K, Schoch H, Whitaker KW, Hill-Smith T, Lucki I, Reyes TM. Early life protein restriction alters dopamine circuitry. Neuroscience. 2010;168:359–370. doi: 10.1016/j.neuroscience.2010.04.010. published online EpubJun 30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Dev VG, Tantravahi R, Miller DA, Miller OJ. Nucleolus organizers in Mus musculus subspecies and in the RAG mouse cell line. Genetics. 1977;86:389–398. published online EpubJun. [PMC free article] [PubMed] [Google Scholar]
- 7.Santoro R, Grummt I. Molecular mechanisms mediating methylation-dependent silencing of ribosomal gene transcription. Molecular cell. 2001;8:719–725. doi: 10.1016/s1097-2765(01)00317-3. published online EpubSep. [DOI] [PubMed] [Google Scholar]
- 8.Tseng H, Chou W, Wang J, Zhang X, Zhang S, Schultz RM. Mouse ribosomal RNA genes contain multiple differentially regulated variants. PloS one. 2008;3 doi: 10.1371/journal.pone.0001843. e1843. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Shiao YH, Leighty RM, Wang C, Ge X, Crawford EB, Spurrier JM, McCann SD, Fields JR, Fornwald L, Riffle L, Driver C, et al. Ontogeny-driven rDNA rearrangement, methylation, and transcription, and paternal influence. PloS one. 2011;6 doi: 10.1371/journal.pone.0022266. e22266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Mayer C, Schmitz KM, Li J, Grummt I, Santoro R. Intergenic transcripts regulate the epigenetic state of rRNA genes. Molecular cell. 2006;22:351–36. doi: 10.1016/j.molcel.2006.03.028. published online EpubMay 5. [DOI] [PubMed] [Google Scholar]
- 11.Santoro R, Schmitz KM, Sandoval J, Grummt I. Intergenic transcripts originating from a subclass of ribosomal DNA repeats silence ribosomal RNA genes in trans. EMBO reports. 2010;11:52–58. doi: 10.1038/embor.2009.254. published online EpubJan. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kass S, Craig N, Sollner-Webb B. Primary processing of mammalian rRNA involves two adjacent cleavages and is not species specific. Molecular and cellular biology. 1987;7:2891–2898. doi: 10.1128/mcb.7.8.2891. published online EpubAug. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Murayama A, Ohmori K, Fujimura A, Minami H, Yasuzawa-Tanaka K, Kuroda T, Oie S, Daitoku H, Okuwaki M, Nagata K, Fukamizu A, et al. Epigenetic control of rDNA loci in response to intracellular energy status. Cell. 2008;133:627–639. doi: 10.1016/j.cell.2008.03.030. published online EpubMay 16. [DOI] [PubMed] [Google Scholar]
- 14.Cannon MV, Buchner DA, Hester J, Miller H, Sehayek E, Nadeau JH, Serre D. Maternal nutrition induces pervasive gene expression changes but no detectable DNA methylation differences in the liver of adult offspring. PloS one. 2014;9 doi: 10.1371/journal.pone.0090335. e90335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Jack CV, Cruz C, Hull RM, Keller MA, Ralser M, Houseley J. Regulation of ribosomal DNA amplification by the TOR pathway. Proceedings of the National Academy of Sciences of the United States of America. 2015;112:9674–9679. doi: 10.1073/pnas.1505015112. published online EpubAug 4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Aldrich JC, Maggert KA. Transgenerational inheritance of diet-induced genome rearrangements in Drosophila. PLoS genetics. 2015;11 doi: 10.1371/journal.pgen.1005148. e1005148; published online EpubApr. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Shea JM, Serra RW, Carone BR, Shulha HP, Kucukural A, Ziller MJ, Vallaster MP, Gu H, Tapper AR, Gardner PD, Meissner A, et al. Genetic and Epigenetic Variation, but Not Diet, Shape the Sperm Methylome. Developmental cell. 2015;35:750–758. doi: 10.1016/j.devcel.2015.11.024. published online EpubDec 21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.El Hajj N, Schneider E, Lehnen H, Haaf T. Epigenetics and life-long consequences of an adverse nutritional and diabetic intrauterine environment. Reproduction. 2014;148:R111–120. doi: 10.1530/REP-14-0334. published online EpubDec. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Gibbons JG, Branco AT, Godinho SA, Yu S, Lemos B. Concerted copy number variation balances ribosomal DNA dosage in human and mouse genomes. Proceedings of the National Academy of Sciences of the United States of America. 2015;112:2485–2490. doi: 10.1073/pnas.1416878112. published online EpubFeb 24. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.