Abstract
Bats host a large number of zoonotic viruses, including several viruses that are highly pathogenic to other mammals. The mechanisms underlying this rich viral diversity are unknown, but they may be linked to unique immunological features that allow bats to act as asymptomatic viral reservoirs. Vertebrates respond to viral infection by inducing interferons (IFNs), which trigger antiviral defenses through interferon-stimulated gene (ISG) expression. While the IFN system of several bats is characterized at the genomic level, less is known about bat IFN-mediated transcriptional responses. Here, we show that IFN signaling in bat cells from the black flying fox (Pteropus alecto) consists of both conserved and unique ISG expression profiles. In IFN-stimulated cells, bat ISGs comprise two unique temporal subclusters with similar early induction kinetics but distinct late-phase declines. By contrast, human ISGs lack this decline phase and remained elevated for longer periods. Notably, in unstimulated cells, bat ISGs were expressed more highly than their human counterparts. We also found that the antiviral effector RNASEL, which is not an ISG in humans, is highly IFN-inducible in black flying fox cells and contributes to cell intrinsic control of viral infection. These studies reveal distinctive innate immune features that may underlie a unique virus-host relationship in bats.
Keywords: bat, immunity, interferon, ISG, virus
Introduction
Bats are recognized as important viral reservoirs, and many highly pathogenic viruses including Nipah virus (NiV) (1), Hendra virus (HeV) (2), Marburg virus (3), SARS-like coronaviruses (4), and Ebolavirus (5) have been detected in various bat species. In experimental infection studies, certain bats can be productively infected with pathogenic viruses without obvious disease symptoms (6–9). A recent study identified bats as hosts for a greater proportion of zoonotic viruses than all other mammalian orders tested, with the highest viral richness found in flavi-, bunya- and rhabdoviruses (10). While the mechanisms underlying disease resistance are not known, it has been speculated that bats possess unique immune features that confer innate antiviral protection (9, 11). In vertebrates, one of the first lines of defense against viral pathogens is the interferon (IFN) response. Upon viral infection, pattern-recognition receptors (PRRs) sense viral components and initiate a signaling cascade that results in the secretion of IFNs. These IFNs bind their cognate receptors to activate the JAK-STAT signaling pathway, leading to the transcriptional induction of hundreds of interferon-stimulated genes (ISGs), many of which have antiviral activity (12).
The black flying fox (Pteropus alecto) is an asymptomatic natural reservoir for the highly lethal Hendra virus (2, 13, 14). Studies of the recently sequenced black flying fox genome revealed that genes for key components of antiviral immunity are conserved in bats, such as pathogen sensors (including toll-like receptors (15), RIG-I-like helicases, and NOD-like receptors), IFNs and their receptors, and ISGs (11, 16). Previous efforts to study bat-virus interactions have mainly focused on the host response to viral infection (17–20), but global transcriptional responses to type I IFN remain uncharacterized.
Since the black flying fox harbors pathogenic human viruses and has an annotated genome, we sought to characterize the IFN-induced transcriptional response in this species. Gene expression analyses revealed that bat cells induce a pool of common ISGs. However, they also induced a small number of apparent novel ISGs, including 2–5A-dependent endoribonuclease (RNASEL). Kinetic analyses revealed that bat ISGs can be categorized into distinct groups depending on their temporal gene expression patterns. Additionally, maintenance of ISG expression over time differed between bat and human cells, suggesting distinct mechanisms of gene regulation.
Methods & Materials
Cell lines
Black flying fox-derived PaBr (brain), PaLu (lung), and PaKi (kidney) immortalized cell lines (21) were grown at 37°C and 5% CO2 and passaged in DMEM/F-12 media (GIBCO #11320033) supplemented with 10% FBS. Human A549 lung adenocarcinoma and HEK293 cells were grown at 37°C and 5% CO2 and passaged in DMEM media (GIBCO #119995065) supplemented with 10% FBS and 1× non-essential amino acids (NEAA) (GIBCO #11140076).
Viruses
YFV-Venus (derived from YF17D-5C25Venus2AUbi) stocks were generated by electroporation of in vitro-transcribed RNA into STAT1−/− fibroblasts as previously described (12). VSV-GFP, Indiana serotype (provided by Jack Rose) was generated by passage in BHK-J cells. For both viruses, virus-containing supernatant was centrifuged to remove cellular debris and stored at −80°C until use.
Viral infection
Cells were seeded into 24-well plates at a density of 1×105 cells per well. Viral stocks were diluted into DMEM/F-12 media supplemented with 1% FBS to make infection media. Media was aspirated and replaced with 200 μL of infection media. Infections were performed at 37°C for 1h, then 800 μL DMEM/F-12 media supplemented with 10% FBS was added back to each well. After approximately one viral life cycle (5h for VSV, 25h for YFV), cells were harvested for flow cytometry.
Flow cytometry
Cells were detached using Accumax, fixed in 1% PFA for 10 min at room temperature, and pelleted by centrifugation at 800×g. Fixed cell pellets were resuspended in 200 μL FACS buffer (1X PBS supplemented with 3% FBS). Samples were run on a Stratedigm S1000 instrument using CellCapTure software and gated based on GFP fluorescence. Data analysis was done using FlowJo software (v9.7.6).
Interferon treatment and RNA isolation
Cells were seeded into 6-well plates at 3×105 cells per well. The following day, cells were treated with 2 mL of DMEM/F-12 media supplemented with 10% FBS and 50 units/mL of Universal Type I IFN Alpha (PBL Assay Science Cat. #11200). The cells were harvested by aspirating the media, washing twice with 2 mL of sterile 1X PBS, then lysing with 350 μL RLT buffer from the RNeasy Mini kit (Qiagen). The cell lysate was stored at −80°C until RNA isolation was completed using the RNeasy kit following the manufacturer’s protocol.
RNA-Seq
Total RNA samples for each time point were prepared in three independent experiments as described above. The Agilent 2100 Bioanalyzer was used to determine RNA quality; all samples had a RIN Score >9. A Qubit fluorometer was used to determine RNA concentration. Libraries were prepared using the TruSeq Stranded mRNA LT Library Prep Kit (Illumina) following the manufacturer’s instructions, summarized as follows. Four μg of DNAse-treated total RNA was used as input. Poly-A RNA was purified and fragmented before strand specific cDNA synthesis. cDNA was then A-tailed and ligated with indexed adapters. Samples were then PCR amplified using the following parameters: 5 ul of PCR Primer Cocktail was added to 20 ul of adapter ligated library, and then 25 μl of PCR Master Mix was added to each sample. Samples were mixed by pipet, the plate sealed and cycled on the thermal cycler with a 100°C heated lid under the following conditions: initial denaturing at 98°C for 30 sec; 15 cycles of 98°C for 10 sec, 60°C for 30 sec, and 72°C for 30 sec; final extension at 72°C for 5 min, hold at 4°C. Samples were then purified with AmpureXP beads, and re-validated on the Agilent 2100 Bioanalyzer and Qubit. Normalized and pooled samples were run on the Illumina HiSeq 2500 using SBS v3 reagents.
Paired-end 100 bp read length FASTQ files were checked for quality using fastqc (http://www.bioinformatics.babraham.ac.uk/projects/fastqc) and fastq_screen (http://www.bioinformatics.babraham.ac.uk/projects/fastq_screen/) and were quality trimmed using fastq-mcf (https://github.com/ExpressionAnalysis/ea-utils/blob/wiki/FastqMcf.md). Trimmed fastq files were mapped to black flying fox assembly ASM32557v1 (https://ftp.ncbi.nih.gov/genomes/Pteropus_alecto/) using TopHat (22). Duplicates were marked using picard-tools (https://broadinstitute.github.io/picard/). Reference annotation based transcript assembly was done using cufflinks (23), and read counts were generated using featureCounts (24). Pairwise differential expression analysis was performed using edgeR (25), and longitudinal analysis was performed using time course (26) after data transformation by voom (27). Differentially-expressed unannotated genes were manually annotated using a nucleotide BLAST (28) search for homologous genes.
Data were deposited in the GEO database (https://www.ncbi.nlm.nih.gov/geo/) under accession number GSE102296.
Heatmaps
Heatmaps were constructed using Morpheus https://software.broadinstitute.org/morpheus/.
Nanostring analysis
A customized panel targeting both black flying fox and human genes (Schoggins_1_C4066, NanoString Technologies, Seattle, WA, USA) was used to measure the expression of 50 genes per species. Probes were designed to target as many known transcript variants as possible (see dataset S2 for accession numbers of targeted genes). Total RNA was isolated from IFN-treated samples as described above. 100 ng of total RNA in 5 μL was used as input in each reaction for NanoString assay. RNA samples were processed according to the NanoString nCounter XT Codeset Gene Expression Assay manufacturer’s protocol. Following hybridization, transcripts were quantitated using the nCounter Digital Analyzer.
RT-qPCR
Total RNA was prepared as described above. Reactions were prepared with the QuantiFast SYBR Green RT-PCR kit (Qiagen #204154) as recommended by the manufacturer, using 50 ng total RNA per reaction. Samples were run on the Applied Biosystems 7500 Fast Real-Time PCR System using 7500 Software v2.0.6. The program consisted of a reverse transcription step at 50°C for 10 min, a polymerase activation step at 95°C for 5 min, and cycling steps alternating between 95°C for 10 sec and 60°C for 30 sec (40 cycles). Melt curves were generated for all experiments by ramping up the temperature 1°C/minute from 60°C to 95°C. Only a single melt curve peak was observed for all primer sets. Primers used to amplify RNASEL (XM_006907762.2) are 5′-CCACCCTGGGGAAAATGTGA-3′ and 5′-GGAGGATCCTGCTTGCTTGT-3′. Primers used to amplify reference gene RPS11 (XM_006905029) are 5′-ATCCGCCGAGACTATCTCCA-3′ and 5′-GGACATCTCTGAAGCAGGGT-3′. Relative expression in IFN-treated samples versus untreated samples was calculated using the comparative CT method using RPS11 as the reference gene.
DNA constructs and plasmid propagation
lentiCRISPR v2 was a gift from Feng Zhang (Addgene plasmid # 52961). RNASEL targeting guides were generated by cloning annealed, complementary 20-bp oligos with Esp3I-compatible overhangs (5′-caccgAGACCCACACCCTCCAGCAG-3′ and 5′-aaacCTGCTGGAGGGTGTGGGTCTc-3′) targeting the black flying fox RNASEL gene into the lentiCRISPRv2 backbone as described in (29). CRISPR guide oligos were designed using CRISPRdirect (30). Proper assembly was confirmed using Sanger sequencing.
Lentiviral pseudoparticles
Lentiviral pseudoparticles were made as described in (31), with some modifications. Briefly, 2×106 HEK293T cells were seeded on a poly-lysine coated 10 cm plate. The following day, media was changed to 7.5 mL DMEM with 3% FBS and 1x NEAA. Cells were co-transfected with 5 μg lentiCRIPSRv2, 2.5 μg pCMV-VSVg and 3.5 μg pGag-pol. 4h post-transfection, media was changed to 7.5 mL fresh DMEM with 3% FBS and 1X NEAA. Supernatant was collected 48h post-transfection, filtered through 0.45 μm to remove debris, and stored at −80°C until use.
RNASEL KO bulk PaKi cell lines
3×105 PaKi cells were seeded on 6-well plates. The following day, media was changed to DMEM/F-12 supplemented with 3% FBS, 4 μg/mL polybrene and 20 mM HEPES. LentiCRISPRv2 lentiviral pseudoparticles were added and cells were spinoculated at 800×g for 45 min at 37°C. Media was changed to DMEM/F-12 with 10% FBS immediately following spinoculation. 48h after transduction, cells were pooled into a 10-cm dish and selected in DMEM/F-12 with 10% FBS and 5 μg/mL puromycin.
Genomic characterization of PaKi RNASEL KO bulk cell lines
Genomic DNA was isolated from wild-type or RNASEL KO bulk populations using the DNeasy Blood and Tissue Kit (Qiagen). The region flanking the CRISPR-targeted sequence was amplified via PCR using primers (5′-ATGGAGACCAAGAGCCACAACA-3′) and (5′-CGTCCTCGTCCTGGAAATTGA-3′). PCR products were gel-purified using the QIAquick Gel Extraction Kit (Qiagen) and subsequently used in TOPO cloning reactions using the TOPO TA Cloning Kit (Thermo Fisher). Several colonies were selected for each cell background and colony PCR was used to amplify the CRISPR-targeted region. Samples were then analyzed using Sanger sequencing.
rRNA degradation assay
3×105 PaKi cells were seeded on 6-well plates. The following day, universal IFN (100 U/mL) was added to the media. After 24h, cells were transfected with 100 ng/mL poly(I:C) in Optimem using Lipofectamine 3000. After 4h, RNA was harvested using the RNeasy Mini Kit (Qiagen) and RNA integrity was measured on a Total RNA Nano chip using an Agilent 2100 Bioanalyzer.
Results
Black flying fox-derived cell lines respond to exogenous type I interferon
Previous studies have shown that exogenous IFNα and IFNγ treatment of cells from the black flying fox can suppress Pteropine orthoreovirus Pulau virus (32) and Hendra virus (33), respectively. We treated immortalized black flying fox kidney-derived (PaKi) cells for 24h with universal IFNα, which is designed for activity across multiple species. We then infected the cells with two model GFP-expressing reporter viruses: a negative-sense RNA rhabdovirus, vesicular stomatitis virus (VSV), and a positive-sense RNA flavivirus, yellow fever virus (YFV). We observed a dose-dependent inhibition of both viruses with IFN treatment (Fig. 1A, B). VSV infection was maximally inhibited by only 50%, whereas YFV infection was suppressed completely at the highest IFN dose. In addition, we confirmed IFN-mediated dose-dependent and time-dependent induction of the canonical ISG MX1 (Fig. 1C, D) (34). These data confirm that PaKi cells are capable of mounting an antiviral response and highlight virus-specific sensitivities to IFN in this cellular background.
Fig 1. Bat-derived cells respond to Type I IFN.

(A) PaKi cells were treated with increasing doses of IFNα for 24h and then infected with VSV-GFP at an MOI of 1 for 5h. The percentage of infected cells was quantified via GFP fluorescence using flow cytometry and normalized to a mock-treated control. Data are represented as mean ± SD for three independent experiments. (B) Same as (A), using YFV-17D Venus for 25h. (C) PaKi cells were treated with increasing doses of IFNα and RNA was harvested at 8h. MX1 mRNA levels were quantified using qRT-PCR and normalized to reference gene RPS11. Data are represented as mean ± SD for three independent experiments. (D) PaKi cells were treated with IFNα (50U/mL) and RNA was harvested at several time points. MX1 mRNA levels were measured using qRT-PCR and normalized to reference gene RPS11. Data are represented as mean ± SD for three independent experiments.
IFN induces a classical ISG signature in PaKi cells
We next used total RNA sequencing (RNA-Seq) to profile the global transcriptional response of PaKi cells treated with IFNα over time (Fig. 2A). Transcriptome assembly analysis across all experimental conditions returned approximately 30,000 genes, of which 11,559 met a minimal read count threshold (mean log2CPM≥0). Differential gene expression analysis revealed that IFN induced 93 genes at 4h, 104 at 8h, 103 at 12h, and 103 at 24h (fold change (FC)≥1.5, FDR≤0.05) (Fig. 2B). There were no downregulated genes at 4h, 2 at 8h, 105 at 12h, and 279 at 24h. However, statistical significance of upregulated genes was more robust than the statistical significance of the downregulated genes. Overall, IFN treatment of PaKi cells produced a positive gene induction signature.
Fig 2. PaKi transcriptome response to IFN.

(A) Experimental schematic. (B) Volcano plots for all time points. Differentially-expressed genes are quantified on the top left. Grey bars indicate log2FC≥1.5 (vertical) or FDR≤0.05 (horizontal). Data is a result of three independent experiments (C) Heatmap including genes that meet the cutoffs mean log2CPM≥0, log2FC≥2, FDR≤0.05 for at least 2 time points. XLOC genes represent unannotated genes.
Next, heat maps were generated to assess individual gene induction, using a cutoff of FDR≤0.05 and FC≥4 for at least 2 time points (Fig. 2C). Many genes in this list have known roles in innate immunity, including well-known ISGs (IFIT1, MX1, OAS2, RSAD2/viperin, USP18), members of the JAK-STAT signaling cascade (STAT1, STAT2, IRF9), pattern recognition receptors (DDX58/RIG-I, IFIH1/MDA5, ZBP1) and transcription factors (ETV7, IRF7, SP110) (35). Notably, we detected induction of GVIN1 (interferon-induced very large GTPase), which is predicted to encode a protein in bats but is annotated as a pseudogene in humans. This list also included transcripts predicted to encode an endogenous retrovirus (ERVK25) and several transcripts predicted to be long non-coding RNAs.
Differentially-expressed genes were cross-referenced to the INTERFEROME v2.0 database (36) to determine if they had previously been reported as IFN-induced genes. At the 4h and 8h time points, more than 80% of the genes in our data set overlapped with INTERFEROME v2.0. Since the INTEFEROME database consists predominantly of human and mouse data sets, this result suggests that antiviral responses in bat cells include a conserved repertoire of IFN-inducible genes commonly found in other mammalian species.
Differential temporal regulation of black flying fox ISGs
We next used a clustering algorithm to group genes in the RNA-Seq data set based on induction kinetics, without imposing fold change or FDR cutoffs (26). This analysis revealed that genes were organized into eight subclusters (SC) based on changes in expression levels throughout the IFN time course (Fig. 3A, Supplemental Table 1). Genes in SC1 and SC2 increased in expression after 8h. Genes in SC3 and SC4 were induced by 4h and peaked at 4–8h, with genes in SC3 returning close to baseline by 12h and SC4 genes remaining elevated for at least 24h. These two SCs were highly-enriched for genes found in the INTERFEROME v2.0 database. Genes in SC5 increased slightly and peaked at 8h, returning to baseline levels by 12h. Levels of SC6 genes either increased or decreased by 4h, followed by decreased levels at 12–24h. SC7 gene expression levels decreased sharply by 8h, followed by a partial recovery by 12h and further decrease in expression by 24h. Finally, genes in SC8 exhibited a sustained decrease in expression levels starting at 8–12h. These data demonstrate that interferon induces distinct subsets of genes which are characterized by differing temporal expression patterns.
Fig 3. Temporal clustering analysis.

(A) Temporal subclusters. The number of genes in each subcluster is indicated below the title. Blue lines represent mean log2(fold change) over time for a single gene. Black lines represent median of all genes in the subcluster. (B) RNA-seq validation using Nanostring. RNA was isolated from PaKi cells and gene expression over time was compared between RNA-seq data (left) and Nanostring data (right). Data presented is the mean of three independent experiments. HK = housekeeping genes.
Orthogonal validation of RNA-Seq data
To validate our RNA-seq results, we used Nanostring nCounter technology, which uses colorimetrically-barcoded DNA probes for direct detection of mRNA without the use of nucleic acid amplification. A customized nCounter codeset was designed containing 50 genes from several temporal SCs, with a focus on the ISG-rich SC3 and SC4 (Supplemental Figure 1, Supplemental Table 2). Temporal expression profiles using Nanostring were generally similar to the RNA-Seq data (Fig. 3B). SC1 and SC2 contained genes with increased expression levels at the later time points. SC3 and SC4 had genes with peak expression levels at 8h, with genes in SC4 exhibiting expression levels that remained elevated at 24h. Genes in SC8 decreased in expression over time, with the lowest expression levels observed at 24h.
Human vs Bat Temporal ISG Regulation
We next used Nanostring to compare temporal regulation of ISGs between bat and human cell lines. We first compared gene expression between PaKi and HEK293 as both cell lines are kidney-derived, but HEK293 cells responded poorly to type I IFN (Supplemental Figure 1). After screening for robust IFN responses in other human cell lines, we chose human A549 cells for comparative studies (Fig. 4A). A striking difference was observed between expression profiles of SC1 and SC2 when comparing PaKi and A549 cells. Of the 5 selected genes in SC1 and SC2, none were significantly upregulated in IFN-treated A549 cells. Similarly, the decreased expression levels observed for bat genes in SC8 were not observed in A549 cells.
Fig 4. ISG expression over time in bat and human cells.

(A) Nanostring gene expression for PaKi (left) and A549 cells (right). PaKi data is the same shown in Fig 3B (right), with additional time points. Data presented is the mean of three independent experiments. HK = housekeeping genes. (B) Expression over time for PaKi and A549 genes in SC3 and SC4. Each line represents the median log2(FC) for all genes in a SC. (C) Time at peak induction for PaKi and A549 genes in SC3 and SC4. (D) Percent genes induced to levels ≥50% of maximum induction at specified time points in SC3 and SC4. (E) Percent genes that recovered to levels ≤50% of maximum induction at specified time points in SC3 and SC4. (F) Baseline mRNA counts for PaKi and A549 genes in SC3 and SC4. Data are represented as mean + SD for three independent experiments. Darker symbols represent mean counts for each subcluster. * = p-value <0.05 using Student’s t-test. (G) Maximum mRNA counts for PaKi and A549 genes in SC3 and SC4. Data are represented as mean + SD for three independent experiments. Darker symbols represent mean counts for each subcluster. * = p-value <0.05 using Student’s t-test.
For further analysis, we chose to focus on previously-reported ISGs, particularly those found in SC3 and SC4. When comparing changes in gene expression in SC3, the median fold-change of all genes in SC3 followed similar trends between bat and human cell lines (Fig. 4B). In SC4, however, genes from A549 cells exhibited higher fold induction and remained elevated at later time points when compared to PaKi cells.
We next determined the time point at which we observed maximum gene induction (tmax). For both SC3 and SC4, approximately 80% of IFN-induced PaKi genes peaked at 8h (Fig. 4C). However, A549 genes had a bimodal pattern within both SCs, with most genes peaking at 4–8h and a smaller subset peaking at 12h or later.
To identify potential differences in induction kinetics, we calculated the percentage of genes that were induced to at least 50% of maximum expression for each time point. Greater than 80% of PaKi genes in both SCs were induced by 4h (Fig. 4D). A549 genes in SC3 behaved similarly, although a small subset of genes was induced by 2h. However, approximately 40% of A549 genes in SC4 were induced by 2h, indicating faster induction of this subset of genes. Genes in this list include HERC5, IFI6, IFIH1, MX1, NLRC5, OAS2, OASL, and PARP12. Notably, PaKi cells express higher baseline levels of ISGs such that by full induction at 8h, PaKi cells express similar or greater mRNA counts compared to A549 cells, despite faster induction in A549 cells (Fig. 4D, Fig. 4G, and Supplemental Fig. 1).
Next, we calculated the percentage of genes that had recovered to below 50% of maximum expression for each time point. In both SCs, recovery occurred earlier in PaKi cells, with most genes recovering by 16h (Fig. 4E). In contrast, the expression levels for most A549 genes remained elevated throughout the time course. Together, these data suggest that the regulatory mechanisms governing IFN-mediated gene induction and maintenance of gene expression differs in each cell type, particularly with genes in SC4.
It was recently reported that black flying fox tissues have constitutive and ubiquitous expression of IFNα, suggesting that cells from this species may be primed to inhibit viral infection due to constitutive expression of certain ISGs (32). Indeed, we observed overall higher ISG mRNA levels in unstimulated PaKi cells, particularly the SC4 genes (Fig. 4F). In addition, we found that over half of SC4 PaKi ISGs were induced to higher maximum counts than corresponding A549 ISGs (Fig. 4G).
Bat cells express multiple non-canonical ISGs, including an active RNASEL
Our gene expression profiling revealed several genes not previously reported to be ISGs. These included EMC2, FILIP1, IL17RC, OTOGL, SLC10A2, and SLC24A1 (Fig. 2A). It is unclear whether the induction of these genes is cell type or species specific. In addition, we observed IFN-mediated induction of RNASEL, which encodes a 2′–5′-oligoadenylate synthetase-dependent ribonuclease. This protein exerts its antiviral effect by degrading viral RNA in response to 2′–5′-linked oligoadenylates, which are generated by the IFN-inducible oligoadenylate synthase (OAS) family of enzymes upon stimulation by dsRNA in the cytosol (37). In human cells, RNASEL is a constitutively-expressed latent enzyme and is not IFN-inducible (Fig. 4A, Supplemental Fig. 1) (38). Interestingly, we observed a dose-dependent induction of RNASEL in IFN-treated PaKi cells (Fig. 5A). Of note, similar mRNA expression levels of RNASEL were observed in unstimulated human and bat cell lines (Fig. 4F and Supplementary Fig. 1). In addition, we observed IFN-mediated RNASEL induction in brain-derived (PaBr) and lung-derived (PaLu) black flying fox cell lines (Fig. 5B), suggesting IFN-mediated induction of RNASEL in the black flying fox is not cell type-specific.
Fig 5. RNASEL is IFN-induced in P. alecto.
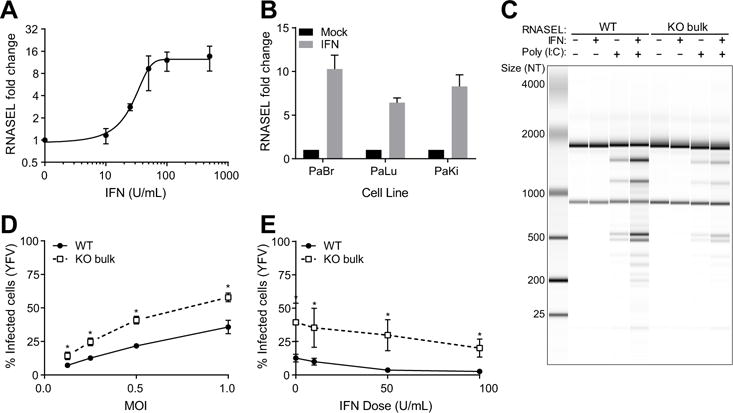
(A) PaKi cells were treated with increasing doses of IFNα and RNA was harvested at 8h. RNASEL mRNA levels were measured using qRT-PCR and normalized to untreated control using reference gene RPS11. Data are represented as mean ± SD for two independent experiments. (B) RNASEL induction in P. alecto brain, lung, and kidney cells. Cells were treated with IFNα (100U/mL) and RNA was harvested at 8h. RNASEL levels were quantified using qRT-PCR and normalized to untreated control using RPS11 as housekeeping control. Data are represented as mean ± SD for three independent experiments. (C) PaKi cells were treated with IFNα (100U/mL) or mock-treated for 24h, followed by transfection with poly(I:C) (100ng/mL) or mock-transfected for 4h and RNA integrity was monitored using a bionalyzer. 100 ng total RNA was run per lane. Data is representative of two independent experiments. (D) WT or RNASEL KO bulk PaKi cells were infected with YFV-17D Venus for 24h and viral infection was quantified via GFP fluorescence using flow cytometry. Data are represented as mean ± SD for three independent experiments. * = p-value < 0.05 using Student’s t-test. (E) WT or RNASEL KO bulk PaKi cells were treated with increasing doses of IFNα for 24h, then infected with YFV-17D Venus at an MOI of 0.5 for 24h. Viral infection was quantified using flow cytometry. Data are represented as mean ± SD for three independent experiments. * = p-value < 0.05 using Student’s t-test.
Next, we constructed RNASEL-deficient PaKi cells using CRISPR (39). Due to lack of a bat-specific RNASEL antibody, we were not able to monitor RNASEL protein levels. However, we detected modifications to the RNASEL gene in the PaKi knock-out bulk population (KO bulk) as compared to the parental wild-type (WT) population (Supplementary Table 3).
To test if RNASEL protein was functional, we activated RNASEL with poly (I:C) transfection after IFN or mock treatment and monitored RNA integrity (Fig. 5C) (40, 41). We observed rRNA degradation when cells were treated with poly(I:C), but not with IFN alone. Treating cells with IFN for 24h to induce RNASEL expression before poly (I:C) transfection resulted in increased RNA degradation and accumulation of smaller products, suggesting increased RNASEL activity. The RNA degradation observed in the WT cells was reduced in the bulk KO cells. Together, these data indicate that RNASEL is a functional ribonuclease that, unlike its human ortholog, is IFN-inducible.
To test if the presence of RNASEL is important in the context of viral infection, we infected both WT and bulk KO cells with YFV17D-Venus and quantified infectivity after one viral life cycle (Fig. 5D). RNASEL KO cells were more permissive to infection at all doses used, suggesting that RNASEL is important for suppression of initial viral infection. To test if RNASEL induction played a role in the protective effect of the IFN response, we treated PaKi WT and KO bulk cells with increasing doses of IFNα for 24h, then infected with YFV17D-Venus (Fig. 5E). Consistent with previous results, KO bulk cells were more permissive to infection than WT cells. In addition, KO bulk cells were resistant to the protective activity of IFN. IFNα (100U/mL) pre-treatment resulted in 80% reduction of infection in WT cells, but only 50% reduction in bulk KO cells. Together, these data suggest RNASEL plays a significant role in the inhibition of viral infection, particularly in the context of the IFN response.
Discussion
This study aimed to identify IFN-stimulated transcripts in a cell line from the black flying fox. Transcriptional analysis revealed over 100 genes induced in response to IFNα. Most of these genes have been previously described as ISGs, suggesting strong evolutionary conservation of the ISG pool, as would be predicted by previous genomic studies of immune genes in the black flying fox (11, 16).
We have provided a framework of black flying fox ISGs, organized by early, mid, and late responses to type I IFN. Temporal expression profiling delineated two separate ISG pools based on unique temporal induction profiles. While both SCs are characterized by similar peak mRNA levels and a subsequent decline by 12–24 hours, SC4 contained some genes that remained elevated. These genes may offer residual antiviral protection, even when IFN signaling has returned to basal levels. In addition, many genes in SC4 have both higher baseline and higher maximal induction levels in bat PaKi cells compared to human A549 cells, which could result in species-specific differences in susceptibility to viral infection. It remains unclear why only particular ISGs are differentially expressed in this context, but it would be interesting to see if there were unique features or functions of these ISGs that could explain their differing expression patterns. Compared to human A549 cells, bat PaKi cells have a more rapid decline in ISG levels, suggesting tightly-regulated expression kinetics. The reason for this strict transcriptional regulation remains unclear, but such a mechanism may exist to prevent excessive inflammation in a highly metabolically active host (11).
We also identified several previously unrecognized, or non-canonical ISGs, including RNASEL. Notably, while our manuscript was under review, another transcriptome study of IFN-treated black flying fox cells also uncovered RNASEL as an ISG (42). In addition, another transcriptomic study done in Jamaican fruit bats (Artibeus jamaicensis) reported that RNASEL levels are increased in the spleen following experimental infection with Tacaribe virus (TCRV), indicating RNASEL induction can be observed in certain bat species in vivo (43). The OAS/RNASEL pathway, by which OAS proteins use viral dsRNA to create short oligonucleotides that act as second messengers to activate the constitutively-expressed latent enzyme RNASEL, is used to clear viral genetic material from the cell. Due to cleavage of both viral and cellular RNA, RNASEL activation can also lead to apoptosis of infected cells (44). In addition, the short RNA fragments created by RNASEL can potentiate the IFN response by activating the cytosolic RNA sensor RIG-I (45). The induction of RNASEL in response to IFN in bats may provide an additional layer of antiviral protection. Indeed, knockout of RNASEL increased viral susceptibility of black flying fox-derived cells. Although induction itself does not result in significant nuclease activity, stimulation with poly (I:C) is sufficient to cause degradation of total RNA in the cell. Unlike in humans, where only the upstream OAS proteins are IFN-induced, bat cells and tissues co-induce both components of the OAS/RNASEL pathway, likely creating a more rapid and potent effect that would inhibit viral replication before extensive viral spread could occur. Induction of RNASEL could also be a way of attenuating the effect of viruses that affect RNASEL activity either by direct inhibition, as seen with the L* protein of murine Theiler’s virus (46), or via increased expression of an RNASEL inhibitor as seen with HIV(47) and EMCV(48).
There is some evidence of modest (~2-fold) RNASEL induction in certain mouse cell lines (49, 50). One mouse cell line with low endogenous levels of RNASEL, however, has been shown to increase RNASEL levels around 10-fold in response to high doses of IFN (51). Rodents are also important viral reservoirs (10), but additional studies are needed to determine if IFN-mediated RNASEL induction plays a role in the host-virus interactions in mice.
We acknowledge several limitations of this study. First, temporal kinetics analysis was performed between one bat kidney cell line and one human lung cell line. It is possible that some differences observed may be due to intrinsic differences between cell lines and not due to differences in species. Studies in multiple cell backgrounds from both species may help determine whether kinetic differences between bat and human ISGs can be generalized to the whole species. Second, in using Universal IFN in place of black flying fox IFN, we made the assumption that the IFN response would be comparable between both types of IFN. Although we observed a transcriptional profile that is expected for IFN-treated cells, we cannot rule out possible differences in downstream signaling kinetics or magnitude of the response between universal and bat-derived IFN. Nonetheless, a recent transcriptomic study done in PaKi cells using black flying fox IFN showed comparable ISG profiles to those observed in our study (42).
Overall, this work lays the foundation for future investigation into the potential unique features of the bat IFN response. While this study focused on ISG induction, we know little about whether bat ISG-encoded effectors possess unique antiviral properties. Uncovering mechanisms of bat ISGs will provide insight into the innate immune responses of an important viral reservoir and may inform research and development of antiviral therapies.
Supplementary Material
Acknowledgments
We would like to thank Jeanine Baisch and Cynthia Silverman from the Genomics Core at Baylor Research Institute for assistance with Nanostring samples, and the McDermott Center Sequencing Core and Bioinformatics Core for sequencing and analysis.
Footnotes
Financial Support
This work was supported in part by NIH grant AI117922 (J.W.S.), the UT Southwestern Endowed Scholars program (J.W.S.), the UT Southwestern High Impact/High Risk Grant Program (J.W.S.), the William F. and Grace H. Kirkpatrick Award (P.C.D.) and the NRF-CRP grant NRF2012NRF-CRP001–056 (L-F.W.). C.X. was partially supported by NIH grant UL1TR001105.
References
- 1.Chua KB, Koh CL, Hooi PS, Wee KF, Khong JH, Chua BH, Chan YP, Lim ME, Lam SK. Isolation of Nipah virus from Malaysian Island flying-foxes. Microbes Infect. 2002;4:145–151. doi: 10.1016/s1286-4579(01)01522-2. [DOI] [PubMed] [Google Scholar]
- 2.Halpin K, Young PL, Field HE, Mackenzie JS. Isolation of Hendra virus from pteropid bats: a natural reservoir of Hendra virus. J Gen Virol. 2000;81:1927–1932. doi: 10.1099/0022-1317-81-8-1927. [DOI] [PubMed] [Google Scholar]
- 3.Towner JS, Pourrut X, Albarino CG, Nkogue CN, Bird BH, Grard G, Ksiazek TG, Gonzalez JP, Nichol ST, Leroy EM. Marburg virus infection detected in a common African bat. PLoS One. 2007;2:e764. doi: 10.1371/journal.pone.0000764. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Li W, Shi Z, Yu M, Ren W, Smith C, Epstein JH, Wang H, Crameri G, Hu Z, Zhang H, Zhang J, McEachern J, Field H, Daszak P, Eaton BT, Zhang S, Wang LF. Bats are natural reservoirs of SARS-like coronaviruses. Science. 2005;310:676–679. doi: 10.1126/science.1118391. [DOI] [PubMed] [Google Scholar]
- 5.Leroy EM, Kumulungui B, Pourrut X, Rouquet P, Hassanin A, Yaba P, Delicat A, Paweska JT, Gonzalez JP, Swanepoel R. Fruit bats as reservoirs of Ebola virus. Nature. 2005;438:575–576. doi: 10.1038/438575a. [DOI] [PubMed] [Google Scholar]
- 6.Williamson MM, Hooper PT, Selleck PW, Westbury HA, Slocombe RF. Experimental hendra virus infectionin pregnant guinea-pigs and fruit Bats (Pteropus poliocephalus) J Comp Pathol. 2000;122:201–207. doi: 10.1053/jcpa.1999.0364. [DOI] [PubMed] [Google Scholar]
- 7.Lau SK, Li KS, Huang Y, Shek CT, Tse H, Wang M, Choi GK, Xu H, Lam CS, Guo R, Chan KH, Zheng BJ, Woo PC, Yuen KY. Ecoepidemiology and complete genome comparison of different strains of severe acute respiratory syndrome-related Rhinolophus bat coronavirus in China reveal bats as a reservoir for acute, self-limiting infection that allows recombination events. J Virol. 2010;84:2808–2819. doi: 10.1128/JVI.02219-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Calisher CH, Childs JE, Field HE, Holmes KV, Schountz T. Bats: important reservoir hosts of emerging viruses. Clin Microbiol Rev. 2006;19:531–545. doi: 10.1128/CMR.00017-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Brook CE, Dobson AP. Bats as ‘special’ reservoirs for emerging zoonotic pathogens. Trends Microbiol. 2015;23:172–180. doi: 10.1016/j.tim.2014.12.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Olival KJ, Hosseini PR, Zambrana-Torrelio C, Ross N, Bogich TL, Daszak P. Host and viral traits predict zoonotic spillover from mammals. Nature. 2017 doi: 10.1038/nature22975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Zhang G, Cowled C, Shi Z, Huang Z, Bishop-Lilly KA, Fang X, Wynne JW, Xiong Z, Baker ML, Zhao W, Tachedjian M, Zhu Y, Zhou P, Jiang X, Ng J, Yang L, Wu L, Xiao J, Feng Y, Chen Y, Sun X, Zhang Y, Marsh GA, Crameri G, Broder CC, Frey KG, Wang LF, Wang J. Comparative analysis of bat genomes provides insight into the evolution of flight and immunity. Science. 2013;339:456–460. doi: 10.1126/science.1230835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Schoggins JW, Wilson SJ, Panis M, Murphy MY, Jones CT, Bieniasz P, Rice CM. A diverse range of gene products are effectors of the type I interferon antiviral response. Nature. 2011;472:481–485. doi: 10.1038/nature09907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Field H, Young P, Yob JM, Mills J, Hall L, Mackenzie J. The natural history of Hendra and Nipah viruses. Microbes Infect. 2001;3:307–314. doi: 10.1016/s1286-4579(01)01384-3. [DOI] [PubMed] [Google Scholar]
- 14.Wang LF, Mackenzie JS, Broder CC. Henipaviruses. In: Knipe DM, Howley PM, editors. Fields Virology. 6th. Lippincott Williams & Wilkins; Philadelphia: 2013. pp. 268–313. [Google Scholar]
- 15.Cowled C, Baker M, Tachedjian M, Zhou P, Bulach D, Wang LF. Molecular characterisation of Toll-like receptors in the black flying fox Pteropus alecto. Dev Comp Immunol. 2011;35:7–18. doi: 10.1016/j.dci.2010.07.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Papenfuss AT, Baker ML, Feng ZP, Tachedjian M, Crameri G, Cowled C, Ng J, Janardhana V, Field HE, Wang LF. The immune gene repertoire of an important viral reservoir, the Australian black flying fox. BMC Genomics. 2012;13:261. doi: 10.1186/1471-2164-13-261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Holzer M, Krahling V, Amman F, Barth E, Bernhart SH, Carmelo VA, Collatz M, Doose G, Eggenhofer F, Ewald J, Fallmann J, Feldhahn LM, Fricke M, Gebauer J, Gruber AJ, Hufsky F, Indrischek H, Kanton S, Linde J, Mostajo N, Ochsenreiter R, Riege K, Rivarola-Duarte L, Sahyoun AH, Saunders SJ, Seemann SE, Tanzer A, Vogel B, Wehner S, Wolfinger MT, Backofen R, Gorodkin J, Grosse I, Hofacker I, Hoffmann S, Kaleta C, Stadler PF, Becker S, Marz M. Differential transcriptional responses to Ebola and Marburg virus infection in bat and human cells. Sci Rep. 2016;6:34589. doi: 10.1038/srep34589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Wu L, Zhou P, Ge X, Wang LF, Baker ML, Shi Z. Deep RNA sequencing reveals complex transcriptional landscape of a bat adenovirus. J Virol. 2013;87:503–511. doi: 10.1128/JVI.02332-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Wynne JW, Shiell BJ, Marsh GA, Boyd V, Harper JA, Heesom K, Monaghan P, Zhou P, Payne J, Klein R, Todd S, Mok L, Green D, Bingham J, Tachedjian M, Baker ML, Matthews D, Wang LF. Proteomics informed by transcriptomics reveals Hendra virus sensitizes bat cells to TRAIL-mediated apoptosis. Genome Biol. 2014;15:532. doi: 10.1186/s13059-014-0532-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Glennon NB, Jabado O, Lo MK, Shaw ML. Transcriptome Profiling of the Virus-Induced Innate Immune Response in Pteropus vampyrus and Its Attenuation by Nipah Virus Interferon Antagonist Functions. J Virol. 2015;89:7550–7566. doi: 10.1128/JVI.00302-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Crameri G, Todd S, Grimley S, McEachern JA, Marsh GA, Smith C, Tachedjian M, De Jong C, Virtue ER, Yu M, Bulach D, Liu JP, Michalski WP, Middleton D, Field HE, Wang LF. Establishment, immortalisation and characterisation of pteropid bat cell lines. PLoS One. 2009;4:e8266. doi: 10.1371/journal.pone.0008266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013;14:R36. doi: 10.1186/gb-2013-14-4-r36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Roberts A, Pimentel H, Trapnell C, Pachter L. Identification of novel transcripts in annotated genomes using RNA-Seq. Bioinformatics. 2011;27:2325–2329. doi: 10.1093/bioinformatics/btr355. [DOI] [PubMed] [Google Scholar]
- 24.Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014;30:923–930. doi: 10.1093/bioinformatics/btt656. [DOI] [PubMed] [Google Scholar]
- 25.Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–140. doi: 10.1093/bioinformatics/btp616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Tai YCAS, Terrence P. A multivariate empirical Bayes statistic for replicated microarray time course data. The Annals of Statistics. 2006;34:2387–2412. [Google Scholar]
- 27.Law CW, Chen Y, Shi W, Smyth GK. voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 2014;15:R29. doi: 10.1186/gb-2014-15-2-r29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Sanjana NE, Shalem O, Zhang F. Improved vectors and genome-wide libraries for CRISPR screening. Nat Methods. 2014;11:783–784. doi: 10.1038/nmeth.3047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Naito Y, Hino K, Bono H, Ui-Tei K. CRISPRdirect: software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics. 2015;31:1120–1123. doi: 10.1093/bioinformatics/btu743. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelsen TS, Heckl D, Ebert BL, Root DE, Doench JG, Zhang F. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science. 2014;343:84–87. doi: 10.1126/science.1247005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zhou P, Tachedjian M, Wynne JW, Boyd V, Cui J, Smith I, Cowled C, Ng JH, Mok L, Michalski WP, Mendenhall IH, Tachedjian G, Wang LF, Baker ML. Contraction of the type I IFN locus and unusual constitutive expression of IFN-alpha in bats. Proc Natl Acad Sci U S A. 2016;113:2696–2701. doi: 10.1073/pnas.1518240113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Janardhana V, Tachedjian M, Crameri G, Cowled C, Wang LF, Baker ML. Cloning, expression and antiviral activity of IFNgamma from the Australian fruit bat, Pteropus alecto. Dev Comp Immunol. 2012;36:610–618. doi: 10.1016/j.dci.2011.11.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Zhou P, Cowled C, Wang LF, Baker ML. Bat Mx1 and Oas1, but not Pkr are highly induced by bat interferon and viral infection. Dev Comp Immunol. 2013;40:240–247. doi: 10.1016/j.dci.2013.03.006. [DOI] [PubMed] [Google Scholar]
- 35.Mi H, Poudel S, Muruganujan A, Casagrande JT, Thomas PD. PANTHER version 10: expanded protein families and functions, and analysis tools. Nucleic Acids Res. 2016;44:D336–342. doi: 10.1093/nar/gkv1194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Rusinova I, Forster S, Yu S, Kannan A, Masse M, Cumming H, Chapman R, Hertzog PJ. Interferome v2.0: an updated database of annotated interferon-regulated genes. Nucleic Acids Res. 2013;41:D1040–1046. doi: 10.1093/nar/gks1215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Silverman RH. Viral encounters with 2′,5′-oligoadenylate synthetase and RNase L during the interferon antiviral response. J Virol. 2007;81:12720–12729. doi: 10.1128/JVI.01471-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Zhou A, Molinaro RJ, Malathi K, Silverman RH. Mapping of the human RNASEL promoter and expression in cancer and normal cells. J Interferon Cytokine Res. 2005;25:595–603. doi: 10.1089/jir.2005.25.595. [DOI] [PubMed] [Google Scholar]
- 39.Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM. RNA-guided human genome engineering via Cas9. Science. 2013;339:823–826. doi: 10.1126/science.1232033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Wreschner DH, James TC, Silverman RH, Kerr IM. Ribosomal RNA cleavage, nuclease activation and 2-5A(ppp(A2′p)nA) in interferon-treated cells. Nucleic Acids Res. 1981;9:1571–1581. doi: 10.1093/nar/9.7.1571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Li Y, Banerjee S, Wang Y, Goldstein SA, Dong B, Gaughan C, Silverman RH, Weiss SR. Activation of RNase L is dependent on OAS3 expression during infection with diverse human viruses. Proc Natl Acad Sci U S A. 2016;113:2241–2246. doi: 10.1073/pnas.1519657113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Zhang Q, Zeng LP, Zhou P, Irving AT, Li S, Shi ZL, Wang LF. IFNAR2-dependent gene expression profile induced by IFN-alpha in Pteropus alecto bat cells and impact of IFNAR2 knockout on virus infection. PLoS One. 2017;12:e0182866. doi: 10.1371/journal.pone.0182866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Gerrard DL, Hawkinson A, Sherman T, Modahl CM, Hume G, Campbell CL, Schountz T, Frietze S. Transcriptomic Signatures of Tacaribe Virus-Infected Jamaican Fruit Bats. mSphere. 2017;2 doi: 10.1128/mSphere.00245-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Li G, Xiang Y, Sabapathy K, Silverman RH. An apoptotic signaling pathway in the interferon antiviral response mediated by RNase L and c-Jun NH2-terminal kinase. J Biol Chem. 2004;279:1123–1131. doi: 10.1074/jbc.M305893200. [DOI] [PubMed] [Google Scholar]
- 45.Malathi K, Dong B, Gale M, Jr, Silverman RH. Small self-RNA generated by RNase L amplifies antiviral innate immunity. Nature. 2007;448:816–819. doi: 10.1038/nature06042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Sorgeloos F, Jha BK, Silverman RH, Michiels T. Evasion of antiviral innate immunity by Theiler’s virus L* protein through direct inhibition of RNase L. PLoS Pathog. 2013;9:e1003474. doi: 10.1371/journal.ppat.1003474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Martinand C, Montavon C, Salehzada T, Silhol M, Lebleu B, Bisbal C. RNase L inhibitor is induced during human immunodeficiency virus type 1 infection and down regulates the 2-5A/RNase L pathway in human T cells. J Virol. 1999;73:290–296. doi: 10.1128/jvi.73.1.290-296.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Martinand C, Salehzada T, Silhol M, Lebleu B, Bisbal C. RNase L inhibitor (RLI) antisense constructions block partially the down regulation of the 2-5A/RNase L pathway in encephalomyocarditis-virus-(EMCV)-infected cells. Eur J Biochem. 1998;254:248–255. doi: 10.1046/j.1432-1327.1998.2540248.x. [DOI] [PubMed] [Google Scholar]
- 49.Essers MA, Offner S, Blanco-Bose WE, Waibler Z, Kalinke U, Duchosal MA, Trumpp A. IFNalpha activates dormant haematopoietic stem cells in vivo. Nature. 2009;458:904–908. doi: 10.1038/nature07815. [DOI] [PubMed] [Google Scholar]
- 50.Mostafavi S, Yoshida H, Moodley D, LeBoite H, Rothamel K, Raj T, Ye CJ, Chevrier N, Zhang SY, Feng T, Lee M, Casanova JL, Clark JD, Hegen M, Telliez JB, Hacohen N, De Jager PL, Regev A, Mathis D, Benoist C, C. Immunological Genome Project Parsing the Interferon Transcriptional Network and Its Disease Associations. Cell. 2016;164:564–578. doi: 10.1016/j.cell.2015.12.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Jacobsen H, Czarniecki CW, Krause D, Friedman RM, Silverman RH. Interferon-induced synthesis of 2-5A-dependent RNase in mouse JLS-V9R cells. Virology. 1983;125:496–501. doi: 10.1016/0042-6822(83)90222-2. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


