Abstract
The occurrence of Listeria monocytogenes has been widely investigated in the poultry production chain from the processing plant to the final product. However, limited data are available on Listeria species, including Listeria monocytogenes, in the poultry farm environment. Therefore, fecal and soil samples from 37 pastured poultry flocks from 10 all-natural farms over 3 years were assessed to determine the prevalence and diversity of Listeria within these alternative poultry farm environments using standard cultural and molecular methods. Listeria species were isolated in 15% of poultry farm samples and included Listeria innocua (65.7%), L. monocytogenes (17.4%), and Listeria welshimeri (15.1%). Additional multiplex PCR serotyping showed group 1/2a-3a to be the most dominant L. monocytogenes serovar group. Based on these results, monoculture growth experiments were conducted on four Listeria soil isolates (three L. monocytogenes isolates representing the three recovered serovar groups and one L. innocua isolate) to determine if culture medium [tripticase soy broth (TSB) and University of Vermont modified Listeria enrichment broth (UVM)], inoculum concentration (102 or 105 CFU/ml), or incubation temperature (20, 30, and 42°C) differentially affected these Listeria species. Overall, very few significant growth differences were observed between the behavior of the three L. monocytogenes isolates (representing the three recovered serovar groups) under the growth conditions tested. Alternatively, at 30°C in UVM with the lower inoculum concentration, the L. innocua isolate had a significantly shorter lag phase than the L. monocytogenes isolates. In coculture growth studies under these same incubation conditions, the lag phase of L. innocua and L. monocytogenes was similar, but the final concentration of L. innocua was significantly higher than L. monocytogenes. However, cocultures in UVM for high inoculum concentration did not show preferential growth of L. innocua over L. monocytogenes. These results indicate that the use of UVM as an enrichment medium may preferentially allow L. innocua to outcompete L. monocytogenes at low concentrations, biasing the Listeria prevalence from these farm samples toward L. innocua and potentially underreporting the presence of L. monocytogenes in these environments.
Keywords: Listeria monocytogenes, Listeria innocua, pastured poultry, UVM enrichment medium, live production farms
Introduction
The genus Listeria is currently comprised of 17 species, including 11 Listeria species described since 2009 (1). However, the genus Listeria sensu stricto includes six species: Listeria innocua, Listeria ivanovii, Listeria grayii, Listeria monocytogenes, Listeria seeligeri, and Listeria welshimeri. These species are well documented and are known to be commonly found in different environments throughout the world (2–6). Among all the Listeria species, L. monocytogenes is recognized as one of the most important foodborne pathogens in many industrialized countries. This pathogen is responsible for listeriosis, a potentially fatal disease that may lead to abortion or serious cases of meningitis, encephalitis, and septicemia (7, 8). Although listeriosis infections are uncommon, mortality rates can reach 30% in at-risk population groups (9–11). In 2015 in the United States, L. monocytogenes was responsible for an estimated 116 cases of listeriosis, 111 hospitalizations, and 15 deaths (12).
Listeria species have been isolated from a wide variety of environments including nature (13) and urban areas (14), agricultural environments (15), food processing plants (16, 17), and retail food (18, 19). The occurrence of Listeria species is of special interest in the food production chain due to the significant threat that L. monocytogenes represents to public health (20, 21). Numerous studies have investigated the occurrence of L. monocytogenes in final products and in food-processing and retail environments, thought to be the main source of contamination for the final product (21, 22). However, limited information is available on Listeria prevalence in poultry farm production environments. In a farm-to-fork approach, it is necessary to assess the incidence of L. monocytogenes along the entire production chain and particularly at the primary production step (the farm environment), taking into account that it could be a potential source of this pathogen into food processing plants. Few studies have investigated and characterized Listeria species in the farm environment, with L. innocua being the predominant species found on grow-out farms, representing ≤78% of all isolated Listeria species (15, 23–26). Other species such as L. ivanovii, L. monocytogenes, L. welshimeri, and L. seeligeri have also been identified in environmental farm samples or chicken feces, but their detection remains infrequent (15, 23).
The initial isolation of L. monocytogenes may be difficult due to its low cell number within the larger indigenous microflora of environmental samples. Thus, the detection of Listeria species involves selective enrichment procedures. Numerous one-step and two-step enrichment broths have been described during the past 50 years (27), with the three most commonly used procedures being the (1) modified ISO 11290-1, (2) USDA-Food Safety Inspection Service (FSIS) Microbiology Laboratory Guide (MLG) method 8.10, and (3) U.S. Food and Drug Administration Bacteriological Analytical Method (FDA-BAM) method #10. Several studies have shown that the enrichment procedure can result in L. monocytogenes being overgrown by other non-pathogenic Listeria species in samples where multiple species are present (28–32). This has especially been demonstrated with L. innocua, whose presence may mask L. monocytogenes and lead to false negative results (33, 34). In addition, several studies have shown that among Listeria strains of food origins, L. innocua grows faster than L. monocytogenes in enrichment media cocultures or food matrices (28–32). These observations raised the question whether the higher prevalence of L. innocua observed in samples from the farm environment is due to a differential growth of Listeria species during the enrichment process or reflect their true distribution in the environment.
While commercial, conventional production represents the majority of the U.S. poultry market, alternative production systems (e.g., organic, all-natural) are becoming more prevalent and there is very limited information related to the prevalence of Listeria spp. within this type of farm environment (35). Therefore, the goal of this work was twofold: (1) determine the prevalence and distribution of Listeria spp. within poultry-related environmental samples (feces and soil) during live production on pastured poultry farms and (2) evaluate whether the distribution of recovered Listeria spp. could be explained by a differential growth in the enrichment broth used in this study, or accurately reflected the native species distribution in the environment.
Materials and Methods
Sample Collection
Ten farms within the southeastern United States were sampled over a period of 3 years (from 2014 to 2016), representing 37 pasture-raised broiler flocks. Farm descriptions are available in Table 1. Soil and feces samples were collected from the pasture where the flock was currently residing at the time of sampling. Samplings occurred three times during grow-out: (i) within a few days of being placed in the pasture, (ii) halfway through their time on pasture, and (iii) on the day the flock was processed. At each sampling time, the pasture area was divided into five separate sections, and five subsamples in each section were pooled into a single sample for each section (a total of five soil samples and five feces samples were collected on each sampling day). Soil samples were collected from the surface (0–7 cm) with sterile scoops, and feces samples were collected from fresh droppings on the soil surface. Gloves and scoops were changed between sample types and between sampling areas. Samples were transported back to the lab on ice and processed within 2 h of collection. A total of 1,110 samples (555 feces samples and 555 soil samples) were collected over the 3-year study period.
Table 1.
Characteristics of the 10 all-natural pastured poultry farms sampled over the 3-year period.
| Farm A | Farm B | Farm C | Farm D | Farm E | |||
|---|---|---|---|---|---|---|---|
| Breed | Freedom ranger | Freedom ranger | Cornish cross | Freedom ranger | Freedom ranger | Freedom ranger | Cornish cross |
| Flock size | >500 | 50–75 | 50–75 | 50–75 | 50–75 | 50–75 | 50–75 |
| No. of flocks | 10 | 3 | 2 | 1 | 1 | 1 | 4 |
| Length of grow-out (weeks) | 10–11 | 13 | 13 | 12.5 | 11 | 11 | 9 |
| Multiuse farm? | Yes | Yes | Yes | No | No | Yes | Yes |
| Animal types | Layers, swine, beef cattle, and sheep | Layers, swine, horses, and goats | Layers, swine, horses, and goats | n/a | n/a | Layers, swine, beef cattle, and sheep | Layers, swine, beef cattle, and sheep |
| Farm I | Farm J | Farm K | Farm L | Farm M | |||
| Breed | Freedom ranger | Cornish cross | Freedom ranger | Cornish cross | Freedom ranger | Freedom ranger | Cornish cross |
| Flock size | 100–500 | 100–500 | 50–75 | 50–75 | 100–500 | >500 | 50–75 |
| No. of flocks | 5 | 3 | 1 | 1 | 2 | 2 | 1 |
| Length of grow-out (weeks) | 11–12 | 9 | 11 | 9 | 11 | 11–12 | 11 |
| Multiuse farm? | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| Animal types | Layers, swine, and goats | Layers, swine, and goats | Layers | Layers | Layers, beef cattle, and goats | Layers, swine, beef cattle, and sheep | Layers and swine |
Culture-Based Detection and Isolation of Listeria Species from Soil and Feces Samples
Enrichment and isolation of Listeria from these environmental samples were performed using a modified version of the USDA-FSIS MLG 8.10 method (36). Three grams of fresh soil or feces were added to 9 ml of buffered peptone water (Acumedia, Lansing, MI, USA) in a filtered stomacher bag and were vigorously shaken for 30 s. As a pre-enrichment step, the stomached homogenates remained in the filtered stomacher bag and were incubated overnight at 35°C. This pre-enrichment step was followed by two enrichments in University of Vermont Modified Listeria Enrichment Broth (UVM; Remel, Lenexa, KS, USA) and Fraser Broth (Oxoid CM0895, Basingstoke, UK), both requiring overnight incubation for 24 h at 30°C. One loopful of the Fraser’s enrichment culture was streaked on Listeria selective agar (LSA, Oxoid CM0856, Basingstoke, UK) for the isolation of Listeria colonies. These plates were incubated overnight at 30°C, and on each plate three Listeria-like colonies per positive samples were picked and kept for further identification tests. Stock cultures were prepared by growing Listeria strains in tripticase soy broth (TSB; Acumedia, Lansing, MI, USA) at 37°C. After washing in sterile water, the cell pellet was suspended in a brain heart infusion (BHI) broth (Acumedia, Lansing, MI, USA) with 25% of glycerol, aliquoted (300 µl in microtubes) and frozen at −80°C until further utilization.
Characterization of Listeria Species and L. monocytogenes Serovar Groups by Multiplex PCR
The species of presumptive Listeria colonies recovered on LSA were determined by multiplex PCR (37). In short, speciation occurred using two multiplex PCR reactions, based on the size of PCR amplicons. Pool 1 contained the primers for the identification of L. ivanovii, L. grayi, and L. innocua, and pool two contained primers for the identification of L. welshimeri, L. monocytogenes, and L. seeligeri. A 25 µl PCR reaction was composed of 1× EconoTaq PLUS 2× Master Mix (Lucigen Corporation, Middleton, WI, USA), 1 µM of each livN, Igr, and lin2 reverse and forward primers (for Pool 1) or 1 µM of each lwe, Lmo, and lse reverse and forward primers (for Pool 2) and quantity sufficient (qs) of water. For negative controls, sterile water was added instead of template DNA. The cycling program consisted of 1 cycle at 95°C for 9 min; 30 cycles at 94°C for 30 s, at 60°C for 30 s, and at 72°C for 1 min; and 1 cycle at 72°C for 7 min. The serovar group of isolates classified as L. monocytogenes was determined by multiplex PCR using five sets of primers (38). Briefly, one colony of L. monocytogenes isolates was thoroughly mixed in a 25 µl PCR reaction containing: 1× EconoTaq PLUS 2× Master Mix (Lucigen Corporation, Middleton, WI, USA), 1 µM of each Lmo0737, ORF2819, and ORF2110 reverse and forward primers, 1.5 µM of Lmo1118 reverse and forward primers, 0.2 µM of prs reverse and forward primers and qs water. For negative controls, sterile water was added instead of template DNA. PCR was performed with an initial denaturation step at 94°C for 3 min; 35 cycles of 94°C for 0.40 min, 53°C for 1.15 min and 72°C for 1.15 min; and 1 final cycle for 72°C for 7 min. PCR reactions were performed in an Eppendorf Mastercycler EP Gradient S (Eppendorf). After the completion of all cycles, 18 µl of PCR product was mixed with 3 µl of BlueJuice™ loading buffer (Invitrogen, Carlsbad, CA, USA) and separated on a 2% E-gel® with SYBR-safe™ (Invitrogen) along with 12 µl of E-Gel™ 1 kb Plus DNA Ladder (Invitrogen).
Bacterial Growth Experiments
Bacterial Strain Selection and Inoculum Preparation
Based on the two main Listeria species found from farm distribution data, three L. monocytogenes strains representing the different recovered serovar groups (1/2a-3a, 1/2b-3b-7, and 4b-4d-4c) and one L. innocua were selected for the growth experiments. Pre-cultures were prepared by inoculating 60 ml of TSB with 100 µl of the thawed stock culture and incubated for 24 h at 30°C while shaking (150 rpm). After 24 h, cell density was estimated spectrophotometrically by measuring the optical density (OD) at 600 nm (OD600nm) with the Thermo Scientific Spectronic 200™ (Fisher Scientific). Pre-cultures were initially diluted in TSB or UVM to a concentration of 106 CFU/ml, and then serially diluted in TSB or UVM to obtain final inoculum concentrations of 105 and 102 CFU/ml.
Monoculture Growth Experiments of Listeria Strains
A volume of 0.4 ml of each culture (105 and 102 CFU/ml) for the four Listeria strains was aliquoted into wells of a microplate (Honeycomb 2 cuvette plate; Labsystems, Inc., Franklin, MA, USA), with five repeats of each culture condition (strain × medium × concentration) per plate. Negative controls consisted on 0.4 ml of uninoculated TSB and UVM (five repeats) incubated along the cultures. For each culture, two independent plate repeats containing all treatment combinations were performed. The inoculated microplate was placed in a Bioscreen C microbiology reader (Thermo Electron Corp., West Palm Beach, FL, USA), which was operated by a computer with Growth Curves Software, v 2.28 (Transgalactic Ltd., Helsinki, Finland). The microbiology reader recorded the OD values of cultures at 20-min intervals after a plate shaking of 10 s at a medium speed (30 shakes/min). Three incubation temperatures were chosen, and the corresponding incubation times were adjusted to make sure that all growth curves reach the stationary phase by the end of the experiment. Plates were incubated at (i) 20°C, estimated soil temperature calculated upon the average of the atmospheric temperatures encountered during the sampling period (from March to August), for 48 h, (ii) 30°C, recommended temperature used for the enrichment procedure of Listeria spp. in UVM medium, for 24 h, and (iii) 42°C, expected temperature inside the chicken intestine, for 24 h.
Coculture Growth Experiments of L. monocytogenes Serovar Group 1/2a-3a and L. innocua
Since no significant growth differences were observed among the three L. monocytogenes isolates in the monoculture growth study, subsequent coculture growth studies with the L. innocua isolate were only performed with the L. monocytogenes 1/2a-3a isolate (the most prevalent serovar group found within the farm data). Three different coculture mixtures were used to observe coculture growth effects: (1) L. monocytogenes 1/2a-3a to L. innocua ratio of 102:102 CFU/ml in UVM, (2) L. monocytogenes 1/2a-3a to L. innocua ratio of 105:105 CFU/ml in UVM, and (3) L. monocytogenes 1/2a-3a to L. innocua ratio of 102:102 CFU/ml in TSB. As positive controls, monocultures of L. monocytogenes 1/2a-3a and L. innocua were tested under the same conditions (medium × concentration) as the coculture mixtures, while negative controls consisted of uninoculated TSB or UVM, respectively. Each coculture mixture was inoculated individually into a microplate along with positive and negative controls with a final inoculum volume of 0.4 ml for each condition. The OD was recorded at 30-min intervals after a brief plate shaking of 10 s at a medium speed (30 shakes/min) during the incubation at 30°C for 24 h. To quantify the growth of L. monocytogenes 1/2a-3a and L. innocua in coculture, 100 µl aliquots were sampled every hour from the microwell plate and serially 10-fold diluted. Appropriate dilutions were plated on Rapid’L.mono medium (Bio-Rad, Hercules, CA, USA). Blue and white colonies were enumerated as L. monocytogenes 1/2a-3a and L. innocua, respectively.
Modeling the Microbial Growth Kinetics and Statistical Analysis
Growth curves were plotted based on OD values over time. Each bacterial growth curve was fitted to a modified Gompertz model using Matlab 2007b. The model equation is as follows:
where y is the OD value measured, t is the time (h), μmax is the maximum specific growth rate (h−1), A is the maximum OD value attained, and λ is the lag time (h). Within the m-file written in Matlab, the lsqcurvefit function (a nonlinear least-squares solver for data fitting) was utilized to fit the growth curves by first using the following Gompertz equation:
Then, the A, B, and C terms were used to determine the growth parameters of interest in the model equation as follows:
The raw data for each growth curve were graphed along with the resulting fit, and the R2 value (coefficient of determination) for each resulting fit was calculated. A four-way analysis of variance (ANOVA) was performed separately on each growth parameter (λ, μmax, and A), followed by Tukey’s post hoc test in R software v3.2.1. Factors included in the model were the Listeria strains, the culture medium, the inoculum concentration, and the incubation temperature. For the coculture experiments, μmax and stationary phase cell densities (equivalent to ODmax) were log10-transformed before ANOVA, and a Tukey’s post hoc test was used to group treatments. For all analyses, differences among groups were considered significant if p ≤ 0.05.
Results and Discussion
Prevalence and Distribution of Listeria Species in Soil and Feces Collected from Pastured Poultry Farms
A total of 1,110 samples (555 feces samples and 555 soil samples) were collected from 37 flocks on 10 pastured poultry farms over a 3-year period, and the distribution of Listeria species varied according to the sampling year (Figure 1A), broiler farm (Figure 1B), and sample type (Figure 1C). Overall, Listeria species were detected on all the farms and isolated in 15% of samples (83 from feces and 85 from soils), which is in the range of Listeria species prevalences reported in poultry-related environmental samples (from 1.4 to 53%) such as broiler litter, farm feed, farm drinking water, soil, and grass (25, 26, 39, 40) as well as in poultry feces (4.7–17%) (23, 25). In our study, three species were isolated including L. innocua (65.7%), L. monocytogenes (17.4%), and L. welshimeri (15.1%), and each of these species were recovered from at least half of the broiler farms (80, 50, and 90%, respectively; Figure 1B). Although different Listeria species distributions were observed between farms, at least two Listeria species were recovered from all but one farm (Farm M), and all three species were recovered from soil samples in all 3 years of the study (Figure 1C). Listeria innocua has been previously shown to be the predominant species isolated from the broiler farm environment (23–26, 40), while the detection of other non-pathogenic Listeria species, such as L. welshimeri, remains infrequent (23, 40), mostly because studies only focus on L. monocytogenes (41–43).
Figure 1.
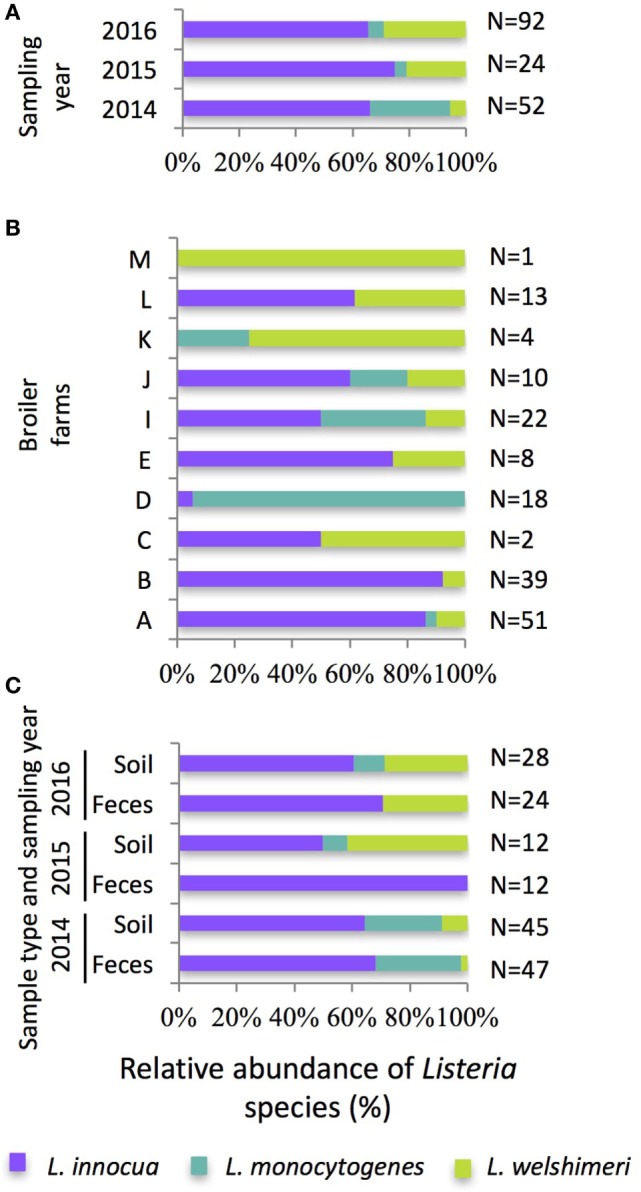
Relative abundance (% total Listeria species isolated) and distribution of the isolated Listeria innocua, Listeria monocytogenes, and Listeria welshimeri (A) according to the sampling year, (B) the broiler farm, and (C) the sample type over the 3-year sampling period. The number of Listeria species isolated per year/farm/sample type is indicated to the right of the bar.
Listeria monocytogenes was isolated from 5.8, 0.3, and 1.0% of all samples collected in 2014, 2015, and 2016, respectively, with 87% (26/30) recovered during 2014 (Figure 1A) and 57% (17/30) of all L. monocytogenes isolates coming from the only flock sampled on Farm D in 2014 (Figure 1B). Overall, three L. monocytogenes serovar groups were identified: 1/2a-3a (70%), 1/2b-3b-7 (20%), and 4b-4d-4e (10%). Interestingly, over the 3-year sampling period, only one L. monocytogenes-positive flock (Farm I, 2014) harbored more than one serotype, demonstrating the potential clonal nature of L. monocytogenes within a flock or on a farm (44). The overall prevalence of L. monocytogenes on these 10 farms was low compared with other grow-out farm environments where 0–46.2% of the environmental and feces samples were L. monocytogenes positive (45), but the distribution of the serotypes was consistent with other studies that have characterized L. monocytogenes serotypes in broiler flocks (26, 41, 43). The prevalence of L. monocytogenes contamination may be dependent on the type of production system. A significant difference between caged- and floor-reared hens was observed with a greater detection of L. monocytogenes in dust samples from floor-reared hens in L. monocytogenes-positive flocks (41). In alternative systems, broilers are raised in less controlled environments than conventional systems and are more likely to be in contact with L. monocytogenes known to be widely spread in soil and vegetation (35).
Poultry farms frequently have other animals (beef cattle, sheep, goats, or swine) and pets present on the production site (35). These animals can be reservoirs for and play a role in the proliferation and deposition of L. monocytogenes into the environment. In our study, all but two farms had other animals raised in close proximity to the broiler flocks during the sampling period, but we did not investigate the possible genotype matching between animal species. Generally, the presence of other animals on the farm increase the risk factor associated with pathogenic bacteria contamination of poultry flocks (42, 46). This has been shown with Campylobacter spp. where adjacent broiler flocks and cattle appear to be the most frequently identified animals with broiler-flock matching Campylobacter spp. isolates (47). Another study has reported an increased risk of L. monocytogenes contamination in laying hen flocks when pets were present on the production site (42).
Monoculture Growth Experiments of L. innocua and L. monocytogenes Isolated from Pastured Poultry Farm Soils
Growth Curve Modeling and Determination of Bacterial Growth Parameters
Our field results showed a higher prevalence of L. innocua compared with L. monocytogenes on pastured poultry grow-out farms, which has been supported by other studies reporting the incidence and characterization of Listeria species in the commercial poultry farm environment (15, 23, 24, 26). In terms of food safety interests, there is a question as to whether there is any physiological basis for the dominance of L. innocua over L. monocytogenes within the poultry farm environment, and whether this dominance related to preferential growth. To determine if this environmental dominance of L. innocua over L. monocytogenes may be linked to growth conditions (e.g., initial concentration, growth temperature, and growth medium), monoculture and coculture growth studies were performed. Three L. monocytogenes isolates (one strain of each serovar groups: 1/2a-3a, 1/2b-3b-7, and 4b-4d-4e) and one L. innocua isolate were selected to compare their growth capacity in liquid media. Bacterial growth was monitored by recording the OD of a culture in growth media (TSB and UVM) inoculated at different initial concentrations (102 and 105 CFU/ml) and incubated at 20°C (average environmental temperature), 30°C (UVM enrichment temperature according to USDA-FSIS MLG 8.10), or 42°C (broiler body temperature). Curve modeling was performed with the Gompertz function that fits the data with R2 values ranging from 0.674 to 0.998, indicating a good fit. From the modified Gompertz equation, three relevant parameters [lag time (λ), maximum specific growth rate (μmax), and maximum OD (ODmax)] were determined for each curve and subsequently used to statistically compare the bacterial growth of the Listeria strains under the different cultural conditions. Using a four-way ANOVA (Tables S1–S3 in Supplementary Material for λ, μmax, and ODmax, respectively), we investigated whether the culture medium, the inoculum concentration, and the incubation temperature could explain the global variation observed between the growth curves. In the same model, we more specifically examine the growth differences between the four Listeria isolates for a single culture condition. The parameters λ, μmax, and ODmax representing bacterial growth characteristics were used in the model.
Effect of Culture Medium, Inoculum Concentration, and Incubation Temperature on Lag Time (λ)
Unsurprisingly, λ was significantly shorter for the higher inoculum concentrations for all strains, enrichment media, and incubation temperatures (F = 801, p < 0.0001; Figure 2). This is in agreement with other studies that have evidenced the importance of inoculum concentration on the ability of a microbial population to initiate growth (48, 49). While Baranyi et al. showed that as the cell numbers in the inoculum decrease, λ increases (50, 51), other studies have reported an effect of the inoculum size only under stressful conditions (49, 52). Increasing incubation temperature significantly decreased λ (F = 174, p < 0.0001) in both TSB and UVM enrichment media for all Listeria strains, as has been showed in other growth media for both L. monocytogenes and L. innocua (28, 49, 52, 53). The temperature-dependent effect was significantly greater in the low initial concentration treatments compared with the higher initial inoculum treatments (F = 77, p < 0.0001). While lag time was significantly shorter in TSB compared with UVM, this was the weakest association of the major growth variables tested (F = 60, p < 0.05).
Figure 2.
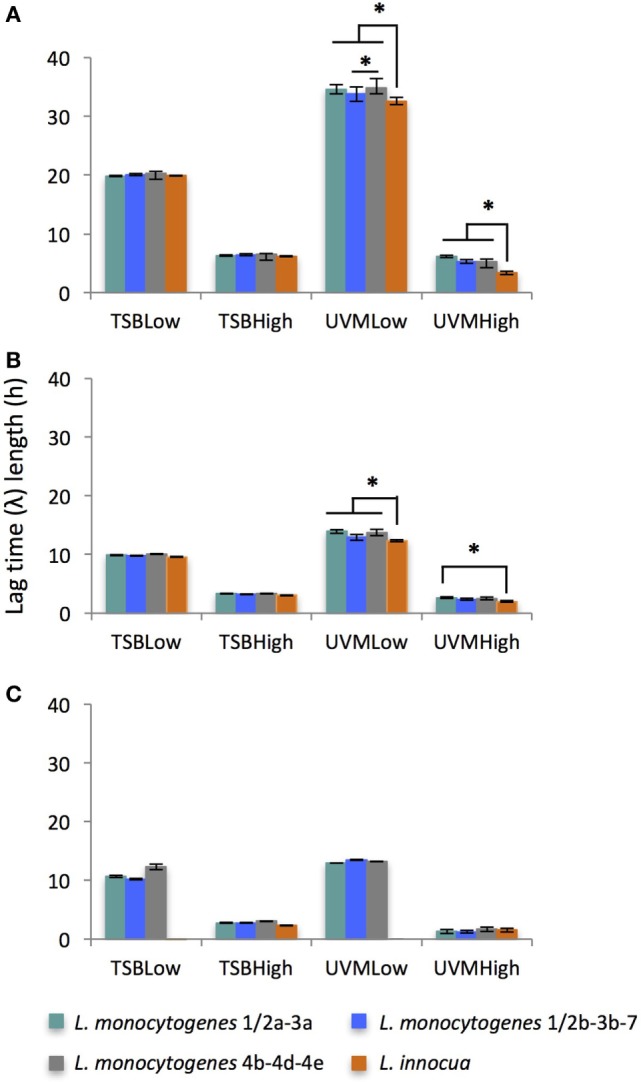
Average of lag time (λ) length of Listeria monocytogenes and Listeria innocua strains inoculated at low (102 CFU/ml) and high (105 CFU/ml) concentrations in TSB and UVM incubated at (A) 20°C, (B) 30°C, and (C) 42°C. *Significant differences between strains (p < 0.05). No growth observed for L. innocua in TSBLow and UVMLow at 42°C.
Effect of Culture Medium, Inoculum Concentration, and Incubation Temperature on Maximum Specific Growth Rate (μmax)
While many of the growth variables tested significantly effected μmax, by far the strongest association was to the enrichment medium (F = 2431, p < 0.0001), where the four Listeria strains grew faster in TSB than UVM (Figure 3). This result is in agreement with the general trend of Listeria strains from food origin showing a faster growth in general growth media (e.g., BHI, TSB + yeast extract) than Listeria enrichment media (UVM, Fraser, and Half-Fraser) (30, 33). We also observed that Listeria strains grew significantly faster at lower incubation temperatures, peaking at 30°C, and this effect was amplified in TSB medium (F = 81, p < 0.0001). This is in agreement with Duh and Schaffner (28), who showed that Listeria strains grew faster at temperatures below 41°C in general growth media (28). The growth variable with the weakest significant association to μmax was initial inoculum concentration, where its effect were only observed in the 42°C treatments (F = 25, p > 0.0001).
Figure 3.
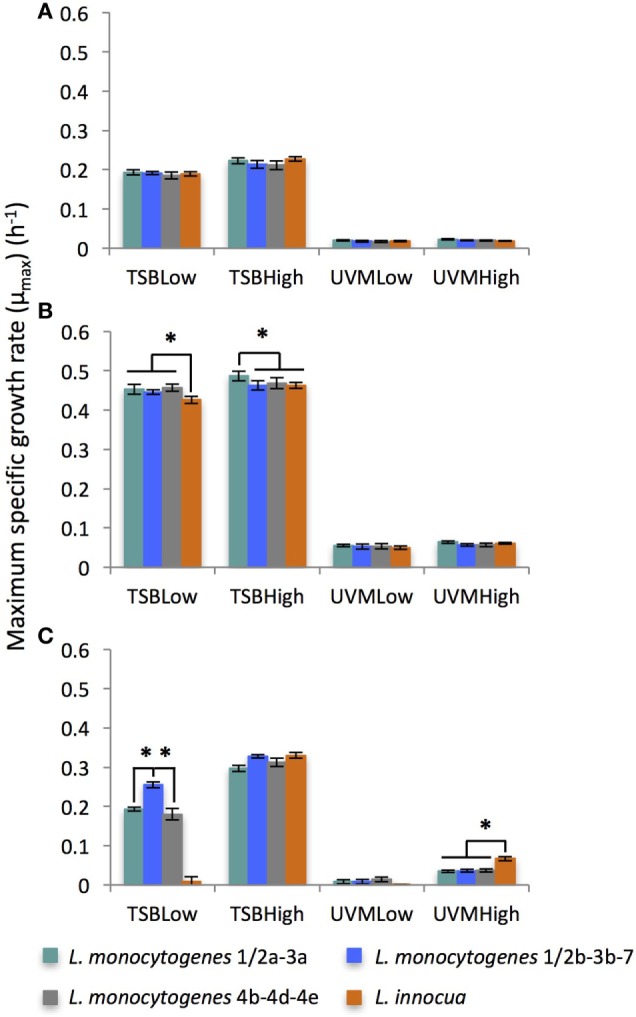
Average of maximum specific growth rate (μmax) of Listeria monocytogenes and Listeria innocua strains inoculated at low (102 CFU/ml) and high (105 CFU/ml) concentrations in TSB and UVM incubated at (A) 20°C, (B) 30°C, and (C) 42°C. *Significant differences between strains (p < 0.05). No growth observed for L. innocua in TSBLow and UVMLow at 42°C.
Effect of Culture Medium, Inoculum Concentration, and Incubation Temperature on ODmax
As was observed for μmax, the enrichment medium was the dominant growth variable affecting ODmax (F = 1481, p < 0.0001) with significantly higher maximum optical densities found in the treatments grown in TSB (Figure 4). This is consistent with other data reporting a higher final cell density in the non-selective culture medium BHI than in selective enrichment media UVM or Half-Fraser (34, 54). For all Listeria strains, the ODmax significantly increased with decreasing incubation temperatures, especially in the UVM treatments (F = 193, p < 0.0001). Unlike λ or μmax, initial inoculum concentrations did not have a significant effect of ODmax overall (F = 0.01, p > 0.05), although limited effects were observed at treatments incubated at 42°C.
Figure 4.
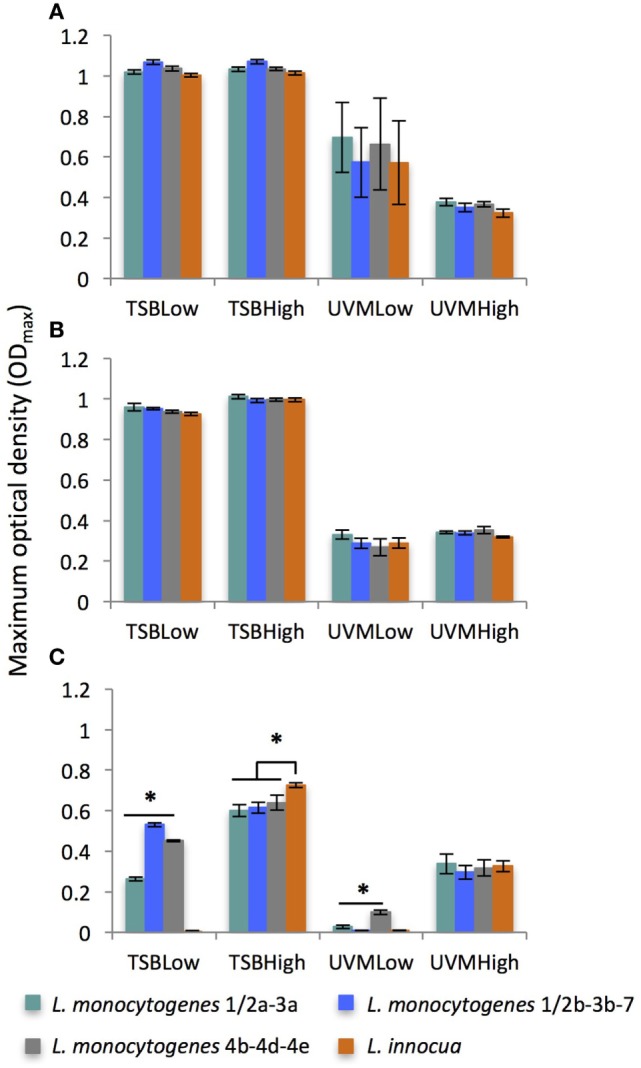
Average of maximum optical density (ODmax) of Listeria monocytogenes and Listeria innocua strains inoculated at low (102 CFU/ml) and high (105 CFU/ml) concentrations in TSB and UVM incubated at (A) 20°C, (B) 30°C, and (C) 42°C. *Significant differences between strains (p < 0.05). No growth observed for L. innocua in TSBLow and UVMLow at 42°C.
Comparing Monoculture Growth between L. monocytogenes and L. innocua Strains
While the general effects of the above variables on growth of Listeria spp. overall, the question of the differential effect between specific Listeria species still remained. While there were some exceptions, generally there were no significant differences between the three L. monocytogenes strains in terms of λ, μmax, or ODmax, and regardless of enrichment media, the L. innocua strain was unable to grow at broiler body temperature (42°C) when the initial inoculum concentrations was 102 CFU/ml (TSBLow and UVMLow). Significant differences in lag time between the three L. monocytogenes strains and the L. innocua strain varied based on the growth variables (Figure 2). At the average environmental and UVM enrichment temperatures (20 and 30°C, respectively), the lag time of L. innocua was significantly shorter than the L. monocytogenes strains in UVM, especially for low inoculum concentrations (p < 0.05; Figures 2A,B, respectively). However, at broiler body temperatures (42°C), L. innocua only grew in the high initial inoculum treatments (TSBHigh and UVMHigh), where there were no significant differences between the four Listeria strains (Figure 2C). No significant differences in λ between L. innocua and L. monocytogenes were found in any of the TSB treatments. Previous studies comparing the growth of L. monocytogenes and L. innocua strains mostly from food origins have reported shorter λ for L. innocua in Fraser (incubated at 30°C), and Half-Fraser (incubated at 37°C) enrichment media (31) and at lower incubation temperatures (≤8°C) (28).
While significant differences in μmax were observed between the four Listeria strains used in this study, there were no consistent trends based on initial inoculum concentration, incubation temperature or enrichment medium (Figure 3). Only in two treatment combinations were there species-specific significant differences in μmax, with L. monocytogenes growing faster than L. innocua in TSBLow at 30°C (Figure 3B) and L. innocua growing faster than L. monocytogenes at 42°C in the UVMHigh treatment (Figure 3C). In contrast to previously reported findings, there were no significant differences found in μmax between the L. monocytogenes and L. innocua isolates in UVM at 30°C, conditions used for the Listeria enrichment process (28, 31, 33). However, the studies comparing the generation time or the growth rate have shown a faster growth of L. innocua compared with L. monocytogenes at temperatures below 40°C only in certain culture media (28, 31, 33), which may explain the similar μmax between L. innocua and L. monocytogenes in our study. In addition, a high level of heterogeneity in growth behavior within L. innocua and L. monocytogenes strains can lead to equivalent μmax between the slowest L. innocua and the fastest L. monocytogenes (30, 55).
There were very few strain-specific differences in the maximum OD (ODmax) for any of the growth variables, with the significant differences found at 42°C (Figure 4C). Among those difference, only under one treatment condition (TSBHigh) were there significant differences between L. monocytogenes and L. innocua, so overall the maximum cell density in culture was unaffected by the Listeria species. No differences were observed between the ODmax of L. innocua and L. monocytogenes species in UVM at 30°C as reported in studies using Half-Fraser and Fraser (30, 31). However, these results are highly dependent on the experiment and opposite trends are also reported showing an higher final population density of L. innocua than L. monocytogenes in enrichment media (29, 54).
Differential Growth of L. monocytogenes 1/2a-3a and L. innocua in Coculture Growth Experiments
Using the cultural conditions for the initial enrichment step for the USDA-FSIS MLG 8.10 L. monocytogenes enrichment method (UVM, 30°C) we found that L. innocua exhibited a significantly shorter lag time than the L. monocytogenes strains in monocultures, especially for low initial inoculum concentrations (102 CFU/ml). To determine if L. innocua has any direct competitive growth advantages over L. monocytogenes in UVM, coculture experiments were performed using the L. innocua strain and the L. monocytogenes 1/2a-3a strain (the most prevalent serovar group from the farm surveys). When both strains were inoculated into the coculture at 105 CFU/ml (Figure 5A), there were no significant differences in λ, μmax, or stationary phase cell density (similar to ODmax), although L. monocytogenes densities did begin to exceed L. innocua cell densities after 24 h. When both strains started at the lower inoculum level (102 CFU/ml), while λ and μmax were similar, L. innocua reached a significantly higher stationary phase cell density compared with L. monocytogenes (F = 31, p < 0.01; Figure 5B). Conversely, when the cocultures inoculated at the lower initial concentrations were grown in TSB, L. monocytogenes demonstrated significantly higher stationary phase cell densities compared with L. innocua (F = 19, p < 0.05), with L. monocytogenes cell densities being ~3× greater than L. innocua (Figure 5C). When comparing the growth curve parameters among the three coculture experiments, only the stationary phase cell density was significantly effected at the species-level (Table 2).
Figure 5.
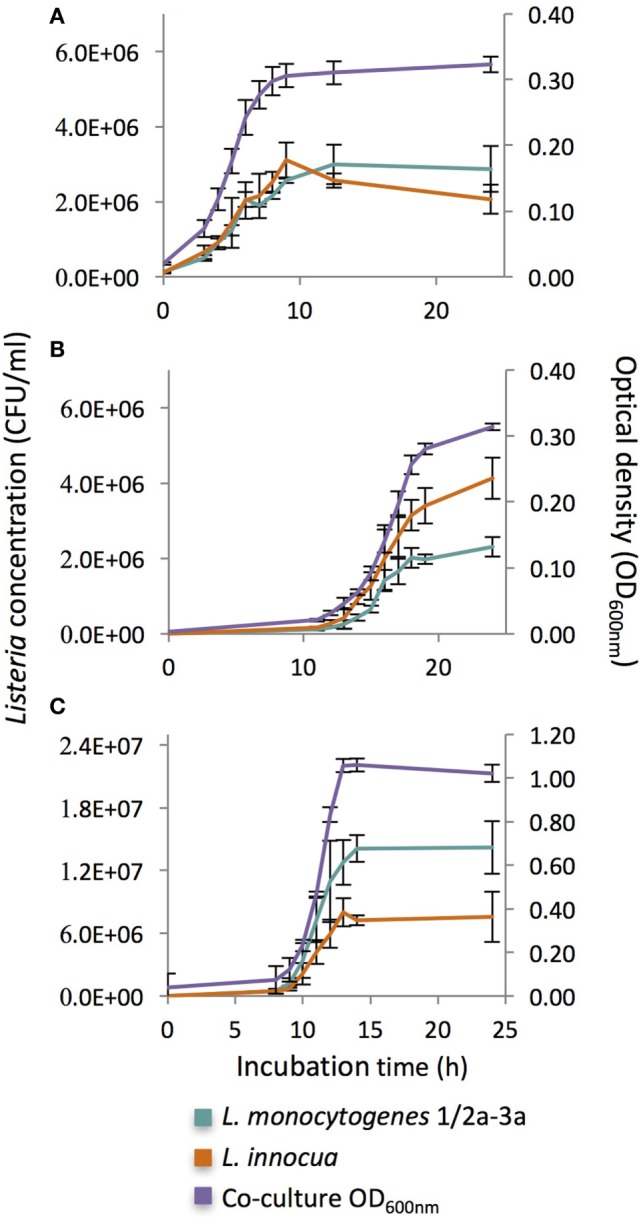
Growth curves of cocultures of Listeria monocytogenes serovar groups 1/2a-3a and Listeria innocua at 30°C inoculated at (A) high initial concentrations (105 CFU/ml) in UVM, (B) low initial concentrations in UVM (102 CFU/ml), and (C) low initial concentrations (102 CFU/ml) in TSB.
Table 2.
Growth parameters lag time (λ), maximum specific growth rate (μmax) and stationary phase cell density for triplicate cocultures studies of Listeria monocytogenes and Listeria innocua grown at 30°C and inoculated at two inoculum ratios (Low:Low and High:High) in two different growth media (UVM and TSB).
| Growth medium | Inoculum ratio (L. monocytogenes: L. innocua)a | Listeria species | λ (h)b | μmax (h−1)b | Stationary phase cell density (log10 CFU/ml)b |
|---|---|---|---|---|---|
| UVM | High:High | monocytogenes | 1.95 ± 0.17A | 5.65 ± 0.04A | 6.47 ± 0.02A |
| Innocua | 2.56 ± 0.43A | 5.82 ± 0.06A | 6.41 ± 0.02A | ||
| UVM | Low:Low | monocytogenes | 12.74 ± 0.45B | 5.68 ± 0.11A | 6.38 ± 0.05AB |
| innocua | 13.20 ± 0.45B | 5.79 ± 0.07A | 6.64 ± 0.07B | ||
| TSB | Low:Low | monocytogenes | 9.27 ± 0.45C | 6.64 ± 0.11B | 7.17 ± 0.06C |
| innocua | 9.24 ± 0.76C | 6.44 ± 0.12B | 6.89 ± 0.09B |
aHigh = 105 CFU/ml; Low = 102 CFU/ml.
bSuperscript letters “A–C” indicate significant differences (p < 0.05) in a single column.
Unlike the monoculture results using the UVM protocol from the USDA-FSIS MLG 8.10 method, no significant lag time difference was observed between L. innocua and L. monocytogenes at low initial inoculum concentrations; however, L. innocua still maintained a competitive growth advantage under these cultural conditions represented by significantly higher stationary phase cell densities (Table 2; Figure 5B). When grown under the same conditions in TSB (Figure 5C), L. monocytogenes grew at significantly higher levels than L. innocua, indicating that there is a enrichment media-specific effect on Listeria growth within these cocultures. When looking at the differences in stationary phase cell densities across all cocultures, there was no significant difference for L. innocua between UVM and TSB in the Low:Low cocultures, but there was over a 1 log reduction in stationary phase cell density for L. monocytogenes grown in TSB (7.17 ± 0.06 log10 CFU/ml) compared with UVM (6.38 ± 0.05 log10 CFU/ml) under those growth conditions (Table 2). While it appears that L. innocua has a competitive growth advantage in UVM with low initial inoculum concentrations, it is possible that this advantage comes more from a disadvantage of L. monocytogenes growing under these conditions, rather than a specific advantage that L. innocua possesses, and previous studies have shown that enrichment/culture media can differentially effect L. innocua and L. monocytogenes (28, 31, 33).
However, considering the UVM enrichment is the first of two enrichments in the USDA-FSIS MLG 8.10 protocol, having significantly higher densities of L. innocua compared with L. monocytogenes would artificially increase the likelihood of isolating L. innocua from samples with equivalent levels of L. innocua and L. monocytogenes. Considering the UVM enrichment is used within the USDA-FSIS MLG 8.10 method, and USDA-FSIS is responsible for the testing of foodborne pathogens from broiler production farms and processing plants, this enrichment bias could potentially or partially explain the prevalence of L. innocua as the dominant Listeria spp. found on poultry farms (15, 23–26).
Conclusion
In our study, we found that L. innocua is more prevalent than the foodborne pathogen L. monocytogenes in soil and feces samples collected from pastured poultry farms, which is consistent with conventional poultry farms. Mono- and coculture growth experiments showed that under cultural conditions used in the first enrichment step of the USDA-FSIS MLG 8.10 L. monocytogenes method (UVM, 30°C), L. innocua had a significantly shorter lag phase (monoculture) and a significantly higher stationary phase cell density (coculture) compared with L. monocytogenes; these growth advantages occurred at low initial inoculum concentrations simulating the low levels of Listeria species encountered in the environment. Based on these results, it is possible that UVM enrichment medium either preferentially supports L. innocua growth over L. monocytogenes, or preferentially restricts L. monocytogenes growth, and that this enrichment step may be biasing the recovery of L. innocua over L. monocytogenes from live production samples. Considering the public health importance of accurately identifying the source of L. monocytogenes outbreaks, future work will need to understand the cultural and molecular mechanisms of this preferential L. innocua growth in UVM, and alternative enrichment methods for L. monocytogenes may need to be considered.
Author Contributions
AL and MR helped to develop experiments, analyze data, and prepare manuscript. ML helped in data analysis and manuscript preparation.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The reviewer KG declared a shared affiliation, with no collaboration, with the authors to the handling editor.
Acknowledgments
The authors would like to acknowledge Laura Lee Rutherford and Cheryl Pearson Gresham for their assistance in sample acquisition and processing as well as Tori McIntosh for the molecular analyses of Listeria isolates. They would also like to thank Dr. Arthur Hinton for access his equipment and Kimberly Ingram for training on the microplate growth reader equipment and software.
Footnotes
Funding. This work was funded under USDA ARS CRIS # 6040-32000-011-00-D entitled “Reduction of Invasive Salmonella enterica in Poultry through Genomics, Phenomics, and Field Investigations of Small Multi-Species Farm Environments.”
Supplementary Material
The Supplementary Material for this article can be found online at http://www.frontiersin.org/articles/10.3389/fvets.2017.00227/full#supplementary-material.
References
- 1.Orsi RH, Wiedmann M. Characteristics and distribution of Listeria spp., including Listeria species newly described since 2009. Appl Microbiol Biotechnol (2016) 100:5273–87. 10.1007/s00253-016-7552-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Hofer E, Ribeiro R, Feitosa DP. Species and serovars of the genus Listeria isolated from different sources in Brazil from 1971 to 1997. Mem Inst Oswaldo Cruz (2000) 95:615–20. 10.1590/S0074-02762000000500005 [DOI] [PubMed] [Google Scholar]
- 3.Chapin TK, Nightingale KK, Worobo RW, Wiedmann M, Strawn LK. Geographical and meteorological factors associated with isolation of Listeria species in New York State produce production and natural environments. J Food Prot (2014) 77:1919–28. 10.4315/0362-028X.JFP-14-132 [DOI] [PubMed] [Google Scholar]
- 4.Linke K, Rückerl I, Brugger K, Karpiskova R, Walland J, Muri-Klinger S, et al. Reservoirs of Listeria species in three environmental ecosystems. Appl Environ Microbiol (2014) 80:5583–92. 10.1128/AEM.01018-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Mcauley CM, Mcmillan K, Moore SC, Fegan N, Fox EM. Prevalence and characterization of foodborne pathogens from Australian dairy farm environments. J Dairy Sci (2014) 97:7402–12. 10.3168/jds.2014-8735 [DOI] [PubMed] [Google Scholar]
- 6.Stea EC, Purdue LM, Jamieson RC, Yost CK, Hansen LT. Comparison of the prevalences and diversities of Listeria species and Listeria monocytogenes in an urban and a rural agricultural watershed. Appl Environ Microbiol (2015) 81:3812–22. 10.1128/AEM.00416-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Farber J, Peterkin P. Listeria monocytogenes, a food-borne pathogen. Microbiol Rev (1991) 55:476–511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Low J, Donachie W. A review of Listeria monocytogenes and listeriosis. Vet J (1997) 153:9–29. 10.1016/S1090-0233(97)80005-6 [DOI] [PubMed] [Google Scholar]
- 9.Siegman-Igra Y, Levin R, Weinberger M, Golan Y, Schwartz D, Samra Z, et al. Listeria monocytogenes infection in Israel and review of cases worldwide. Emerg Infect Dis (2002) 8:305–10. 10.3201/eid0803.010195 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wing EJ, Gregory SH. Listeria monocytogenes: clinical and experimental update. J Infect Dis (2002) 185:S18–24. 10.1086/338465 [DOI] [PubMed] [Google Scholar]
- 11.Painter J, Slutsker L. Listeriosis in humans. In: Ryser ET, Marth EH, editors. Listeria, Listeriosis and Food Safety. Boca Raton, FL: CRC Press; (2007). p. 75–95. [Google Scholar]
- 12.CDC. Foodborne Diseases Active Surveillance Network (FoodNet): FoodNet 2015 Surveillance Report (Final Data). Atlanta, GA: U.S. Department of Health and Human Services, Centers for Disease Control and Prevention; (2017). [Google Scholar]
- 13.Sauders BD, Wiedmann M. Ecology of Listeria species and L. monocytogenes in the natural environment. In: Ryser ET, Marth EH, editors. Listeria, Listeriosis and Food Safety – Third Edition. Boca Raton, FL: CRC Press; (2007). p. 21–53. [Google Scholar]
- 14.Sauders BD, Overdevest J, Fortes E, Windham K, Schukken Y, Lembo A, et al. Diversity of Listeria species in urban and natural environments. Appl Environ Microbiol (2012) 78(12):4420–33. 10.1128/AEM.00282-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Dahshan H, Merwad A, Mohamed TS. Listeria species in broiler poultry farms: potential public health hazards. J Microbiol Biotechnol (2016) 26:1551–6. 10.4014/jmb.1603.03075 [DOI] [PubMed] [Google Scholar]
- 16.Martín B, Perich A, Gómez D, Yangüela J, Rodríguez A, Garriga M, et al. Diversity and distribution of Listeria monocytogenes in meat processing plants. Food Microbiol (2014) 44:119–27. 10.1016/j.fm.2014.05.014 [DOI] [PubMed] [Google Scholar]
- 17.Sasaki Y, Haruna M, Murakami M, Hayashida M, Takahashi N, Urushiyama T, et al. Contamination of poultry products with Listeria monocytogenes at poultry processing plants. J Vet Med Sci (2014) 76:129–32. 10.1292/jvms.13-0267 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ahmed OM, Pangloli P, Hwang C-A, Zivanovic S, Wu T, D’Souza D, et al. The occurrence of Listeria monocytogenes in retail ready-to-eat meat and poultry products related to the levels of acetate and lactate in the products. Food Control (2015) 52:43–8. 10.1016/j.foodcont.2014.12.015 [DOI] [Google Scholar]
- 19.Kramarenko T, Roasto M, Keto-Timonen R, Mäesaar M, Meremäe K, Kuningas M, et al. Listeria monocytogenes in ready-to-eat vacuum and modified atmosphere packaged meat and fish products of Estonian origin at retail level. Food Control (2016) 67:48–52. 10.1016/j.foodcont.2016.02.034 [DOI] [Google Scholar]
- 20.Tompkin R. Control of Listeria monocytogenes in the food-processing environment. J Food Prot (2002) 65:709–25. 10.4315/0362-028X-65.4.709 [DOI] [PubMed] [Google Scholar]
- 21.Ferreira V, Wiedmann M, Teixeira P, Stasiewicz M. Listeria monocytogenes persistence in food-associated environments: epidemiology, strain characteristics, and implications for public health. J Food Prot (2014) 77:150–70. 10.4315/0362-028X.JFP-13-150 [DOI] [PubMed] [Google Scholar]
- 22.Lianou A, Sofos JN. A review of the incidence and transmission of Listeria monocytogenes in ready-to-eat products in retail and food service environments. J Food Prot (2007) 70:2172–98. 10.4315/0362-028X-70.9.2172 [DOI] [PubMed] [Google Scholar]
- 23.Iida T, Kanzaki M, Maruyama T, Inoue S, Kaneuchi C. Prevalence of Listeria monocytogenes in intestinal contents of healthy animals in Japan. J Vet Med Sci (1991) 53:873–5. 10.1292/jvms.53.873 [DOI] [PubMed] [Google Scholar]
- 24.Fenlon D, Wilson J, Donachie W. The incidence and level of Listeria monocytogenes contamination of food sources at primary production and initial processing. J Appl Bacteriol (1996) 81:641–50. 10.1111/j.1365-2672.1996.tb01966.x [DOI] [PubMed] [Google Scholar]
- 25.Petersen L, Madsen M. Listeria spp. in broiler flocks: recovery rates and species distribution investigated by conventional culture and the EiaFoss method. Int J Food Microbiol (2000) 58:113–6. 10.1016/S0168-1605(00)00258-0 [DOI] [PubMed] [Google Scholar]
- 26.Milillo S, Stout J, Hanning I, Clement A, Fortes E, Den Bakker H, et al. Listeria monocytogenes and hemolytic Listeria innocua in poultry. Poult Sci (2012) 91:2158–63. 10.3382/ps.2012-02292 [DOI] [PubMed] [Google Scholar]
- 27.Donnelly CW, Nyachuba DG. Conventional methods to detect and isolate Listeria monocytogenes. In: Ryser ET, Marth EH, editors. Listeria, Listeriosis and Food Safety – Third Edition. Boca Raton, FL: CRC Press; (2007). 215 p. [Google Scholar]
- 28.Duh Y-H, Schaffner DW. Modeling the effect of temperature on the growth rate and lag time of Listeria innocua and Listeria monocytogenes. J Food Prot (1993) 56:205–10. 10.4315/0362-028X-56.3.205 [DOI] [PubMed] [Google Scholar]
- 29.Petran RL, Swanson KM. Simultaneous growth of Listeria monocytogenes and Listeria innocua. J Food Prot (1993) 56:616–8. 10.4315/0362-028X-56.7.616 [DOI] [PubMed] [Google Scholar]
- 30.Cornu M, Kalmokoff M, Flandrois J-P. Modelling the competitive growth of Listeria monocytogenes and Listeria innocua in enrichment broths. Int J Food Microbiol (2002) 73:261–74. 10.1016/S0168-1605(01)00658-4 [DOI] [PubMed] [Google Scholar]
- 31.Besse NG, Barre L, Buhariwalla C, Vignaud ML, Khamissi E, Decourseulles E, et al. The overgrowth of Listeria monocytogenes by other Listeria spp. in food samples undergoing enrichment cultivation has a nutritional basis. Int J Food Microbiol (2010) 136:345–51. 10.1016/j.ijfoodmicro.2009.10.025 [DOI] [PubMed] [Google Scholar]
- 32.Carvalheira A, Eusébio C, Silva J, Gibbs P, Teixeira P. Influence of Listeria innocua on the growth of Listeria monocytogenes. Food Control (2010) 21:1492–6. 10.1016/j.foodcont.2010.04.021 [DOI] [Google Scholar]
- 33.Curiale MS, Lewus C. Detection of Listeria monocytogenes in samples containing Listeria innocua. J Food Prot (1994) 57:1048–51. 10.4315/0362-028X-57.12.1048 [DOI] [PubMed] [Google Scholar]
- 34.Oravcova K, Trnčíková T, Kuchta T, Kaclikova E. Limitation in the detection of Listeria monocytogenes in food in the presence of competing Listeria innocua. J Appl Microbiol (2008) 104:429–37. 10.1111/j.1365-2672.2007.03554.x [DOI] [PubMed] [Google Scholar]
- 35.Rothrock MJ, Hiett KL, Guard JY, Jackson CR. Antibiotic resistance patterns of major zoonotic pathogens from all-natural, antibiotic-free, pasture-raised broiler flocks in the southeastern United States. J Environ Qual (2016) 45:593–603. 10.2134/jeq2015.07.0366 [DOI] [PubMed] [Google Scholar]
- 36.USDA-FSIS. Isolation and Identification of Listeria monocytogenes from Red Meat, Poultry, Ready-to-Eat Siluriformes (Fish) and Egg Products, and Environmental Samples. (2017). Aviable from: https://www.fsis.usda.gov/wps/wcm/connect/1710bee8-76b9-4e6c-92fc-fdc290dbfa92/MLG-8.pdf?MOD=AJPERES
- 37.Huang B, Eglezos S, Heron BA, Smith H, Graham T, Bates J, et al. Comparison of multiplex PCR with conventional biochemical methods for the identification of Listeria spp. isolates from food and clinical samples in Queensland, Australia. J Food Prot (2007) 70:1874–80. 10.4315/0362-028X-70.8.1874 [DOI] [PubMed] [Google Scholar]
- 38.Doumith M, Buchrieser C, Glaser P, Jacquet C, Martin P. Differentiation of the major Listeria monocytogenes serovars by multiplex PCR. J Clin Microbiol (2004) 42:3819–22. 10.1128/JCM.42.8.3819-3822.2004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Jones D, Anderson K, Guard J. Prevalence of coliforms, Salmonella, Listeria, and Campylobacter associated with eggs and the environment of conventional cage and free-range egg production. Poult Sci (2012) 91:1195–202. 10.3382/ps.2011-01795 [DOI] [PubMed] [Google Scholar]
- 40.Dhama K, Verma AK, Rajagunalan S, Kumar A, Tiwari R, Chakraborty S, et al. Listeria monocytogenes infection in poultry and its public health importance with special reference to food borne zoonoses. Pak J Biol Sci (2013) 16:301–8. 10.3923/pjbs.2013.301.308 [DOI] [PubMed] [Google Scholar]
- 41.Chemaly M, Toquin M-T, Le Nôtre Y, Fravalo P. Prevalence of Listeria monocytogenes in poultry production in France. J Food Prot (2008) 71:1996–2000. 10.4315/0362-028X-71.10.1996 [DOI] [PubMed] [Google Scholar]
- 42.Aury K, Le Bouquin S, Toquin M-T, Huneau-Salaün A, Le Nôtre Y, Allain V, et al. Risk factors for Listeria monocytogenes contamination in French laying hens and broiler flocks. Prev Vet Med (2011) 98:271–8. 10.1016/j.prevetmed.2010.11.017 [DOI] [PubMed] [Google Scholar]
- 43.Aury-Hainry K, Le Bouquin S, Labbé A, Petetin I, Chemaly M. Listeria monocytogenes contamination in French breeding and fattening turkey flocks. J Food Prot (2011) 74:1096–103. 10.4315/0362-028X.JFP-10-540 [DOI] [PubMed] [Google Scholar]
- 44.Haase JK, Didelot X, Lecuit M, Korkeala H, Achtman M. The ubiquitous nature of Listeria monocytogenes clones: a large-scale Multilocus Sequence Typing study. Environ Microbiol (2014) 16:405–16. 10.1111/1462-2920.12342 [DOI] [PubMed] [Google Scholar]
- 45.Rothrock MJ, Davis ML, Locatelli A, Bodie A, Mcintosh TG, Donaldson JR, et al. Listeria occurrence in poultry flocks: detection and potential implications. Front Vet Sci (2017) 4:125. 10.3389/fvets.2017.00125 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Torralbo A, Borge C, Allepuz A, Garcia-Bocanegra I, Sheppard SK, Perea A, et al. Prevalence and risk factors of Campylobacter infection in broiler flocks from southern Spain. Prev Vet Med (2014) 114:106–13. 10.1016/j.prevetmed.2014.01.019 [DOI] [PubMed] [Google Scholar]
- 47.Agunos A, Waddell L, Leger D, Taboada E. A systematic review characterizing on-farm sources of Campylobacter spp. for broiler chickens. PLoS One (2014) 9:e104905. 10.1371/journal.pone.0104905 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Pascual C, Robinson T, Ocio M, Aboaba O, Mackey B. The effect of inoculum size and sublethal injury on the ability of Listeria monocytogenes to initiate growth under suboptimal conditions. Lett Appl Microbiol (2001) 33:357–61. 10.1046/j.1472-765X.2001.01012.x [DOI] [PubMed] [Google Scholar]
- 49.Robinson TP, Aboaba OO, Kaloti A, Ocio MJ, Baranyi J, Mackey BM. The effect of inoculum size on the lag phase of Listeria monocytogenes. Int J Food Microbiol (2001) 70:163–73. 10.1016/S0168-1605(01)00541-4 [DOI] [PubMed] [Google Scholar]
- 50.Baranyi J. Comparison of stochastic and deterministic concepts of bacterial lag. J Theor Biol (1998) 192:403–8. 10.1006/jtbi.1998.0673 [DOI] [PubMed] [Google Scholar]
- 51.Baranyi J, Pin C. Estimating bacterial growth parameters by means of detection times. Appl Environ Microbiol (1999) 65:732–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Koutsoumanis KP, Sofos JN. Effect of inoculum size on the combined temperature, pH and aw limits for growth of Listeria monocytogenes. Int J Food Microbiol (2005) 104:83–91. 10.1016/j.ijfoodmicro.2005.01.010 [DOI] [PubMed] [Google Scholar]
- 53.Francois K, Devlieghere F, Smet K, Standaert A, Geeraerd A, Van Impe J, et al. Modelling the individual cell lag phase: effect of temperature and pH on the individual cell lag distribution of Listeria monocytogenes. Int J Food Microbiol (2005) 100:41–53. 10.1016/j.ijfoodmicro.2004.10.032 [DOI] [PubMed] [Google Scholar]
- 54.Bruhn JB, Vogel BF, Gram L. Bias in the Listeria monocytogenes enrichment procedure: lineage 2 strains outcompete lineage 1 strains in University of Vermont selective enrichments. Appl Environ Microbiol (2005) 71:961–7. 10.1128/AEM.71.2.961-967.2005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.MacDonald F, Sutherland AD. Important differences between the generation times of Listeria monocytogenes and List. innocua in two Listeria enrichment broths. J Dairy Res (1994) 61:433–6. 10.1017/S0022029900030879 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


