Abstract
Vertebrate embryonic patterning depends on signaling from Nodal, a TGFβ superfamily member. There are three Nodal orthologs in zebrafish; southpaw directs left-right asymmetries, while squint and cyclops function earlier to pattern mesendoderm. TGFβ member Vg1 is implicated in mesoderm formation but the role of the zebrafish ortholog, Growth differentiation factor 3 (Gdf3), has not been fully explored. We show that zygotic expression of gdf3 is dispensable for embryonic development, while maternally deposited gdf3 is required for mesendoderm formation and dorsal-ventral patterning. We further show that Gdf3 can affect left-right patterning at multiple stages, including proper development of regional cell morphology in Kupffer’s vesicle and the establishment of southpaw expression in the lateral plate mesoderm. Collectively, our data indicate that gdf3 is critical for robust Nodal signaling at multiple stages in zebrafish embryonic development.
Research organism: Zebrafish
Introduction
Transforming Growth Factor-β (TGFβ) superfamily proteins Nodal and Bone Morphogenetic Protein (Bmp) are secreted ligands that play essential roles in development, including maintaining pluripotency of embryonic stem cells and in the patterning of the early embryo (Morikawa et al., 2016). The well-conserved Nodal ligand binds to a heteromeric complex of receptors that phosphorylate Smad2/3, which enables its association with Smad4 (Hata and Chen, 2016). This Smad complex then interacts with transcription factors in the nucleus to activate the expression of target genes. In embryogenesis, Nodal signaling is required for the formation of the mesoderm and endoderm germ-layers (Iannaccone et al., 1992; Conlon et al., 1994; Jones et al., 1995; Feldman et al., 1998) and for proper establishment of left-right (L-R) asymmetry (Levin et al., 1995; Collignon et al., 1996; Lowe et al., 1996; Sampath et al., 1997; Long et al., 2003; Grimes and Burdine, 2017). In zebrafish, the Nodal orthologs Nodal-related1 (Ndr1; previously known as Squint) and Nodal-related 2 (Ndr2; previously known as Cyclops) primarily function in mesoderm and endoderm formation (Feldman et al., 1998) while a third ortholog, Southpaw (Spaw), is essential for proper L-R patterning (Long et al., 2003). The Bmp signaling pathway shares some common components with the Nodal pathway except Bmp binds to a different heteromeric complex of receptors that phosphorylates Smad1/5/8, enabling their association with Smad4 and ultimately activating Bmp-specific target genes (Hata and Chen, 2016). In the embryo, a ventral-to-dorsal decreasing Bmp activity gradient functions to specify ventral fates (De Robertis, 2009; Langdon and Mullins, 2011). Overall, the correct temporal and spatial signaling combinations from the Nodal (van Boxtel et al., 2015; Dubrulle et al., 2015; Sako et al., 2016) and Bmp (De Robertis, 2009; Schmid et al., 2000) pathways are essential along the dorsal-ventral axis for organization of the vertebrate embryo (Xu et al., 2014; Thisse and Thisse, 2015).
An additional TGFβ ligand, Vg1, has also been implicated in both mesoderm formation (Thomsen and Melton, 1993; Kessler and Melton, 1995; Dohrmann et al., 1996; Wall et al., 2000; Birsoy et al., 2006; Chen et al., 2006; Andersson et al., 2007) and L-R patterning (Hyatt et al., 1996; Hanafusa et al., 2000; Rankin et al., 2000; Wall et al., 2000; Peterson et al., 2013). The vg1 transcript was initially identified as being enriched in the vegetal cortex of Xenopus oocytes (Rebagliati et al., 1985) and was subsequently found to be able to induce mesoderm in animal cap assays (Thomsen and Melton, 1993; Kessler and Melton, 1995). Vg1 physically interacts with Nodal (Tanaka et al., 2007; Fuerer et al., 2014) and utilizes the same Nodal signaling pathway components (Cheng et al., 2003). The mammalian and Xenopus Vg1 orthologs may inhibit Bmp signaling in Xenopus and cultured cells (Birsoy et al., 2006; Levine and Brivanlou, 2006). Chimeric versions of the zebrafish Vg1 ortholog Gdf3 also induce mesoderm in animal cap assays (Dohrmann et al., 1996; Peterson et al., 2013), while morpholino oligonucleotide (MO)-mediated knockdown has demonstrated a requirement for Gdf3 in L-R patterning in zebrafish (Peterson et al., 2013).
gdf3 is both maternally and zygotically expressed in zebrafish (Helde and Grunwald, 1993; Dohrmann et al., 1996), but the potential role of the endogenous protein in early embryonic patterning events has not been examined. To address this issue and further investigate the roles of Gdf3 during embryogenesis, we generated zebrafish gdf3 mutants using CRISPR/Cas9 genome editing. Analysis of mutants revealed that maternally-supplied Gdf3 is essential for robust Nodal signaling in the early embryo, being required to form head and trunk mesoderm and endoderm (mesendoderm). Utilizing gdf3 mutants and MO knockdown, we confirm the role of Gdf3 in promoting spaw expression during L-R patterning and demonstrate a role for Gdf3 in the formation of the zebrafish L-R coordinator, Kupffer’s vesicle (KV). Taken together, our work supports a model of Gdf3 as an essential co-ligand for robust Nodal signaling throughout zebrafish development and, therefore, as an essential factor in vertebrate patterning.
Results
Maternal gdf3 is required for embryonic mesendoderm patterning
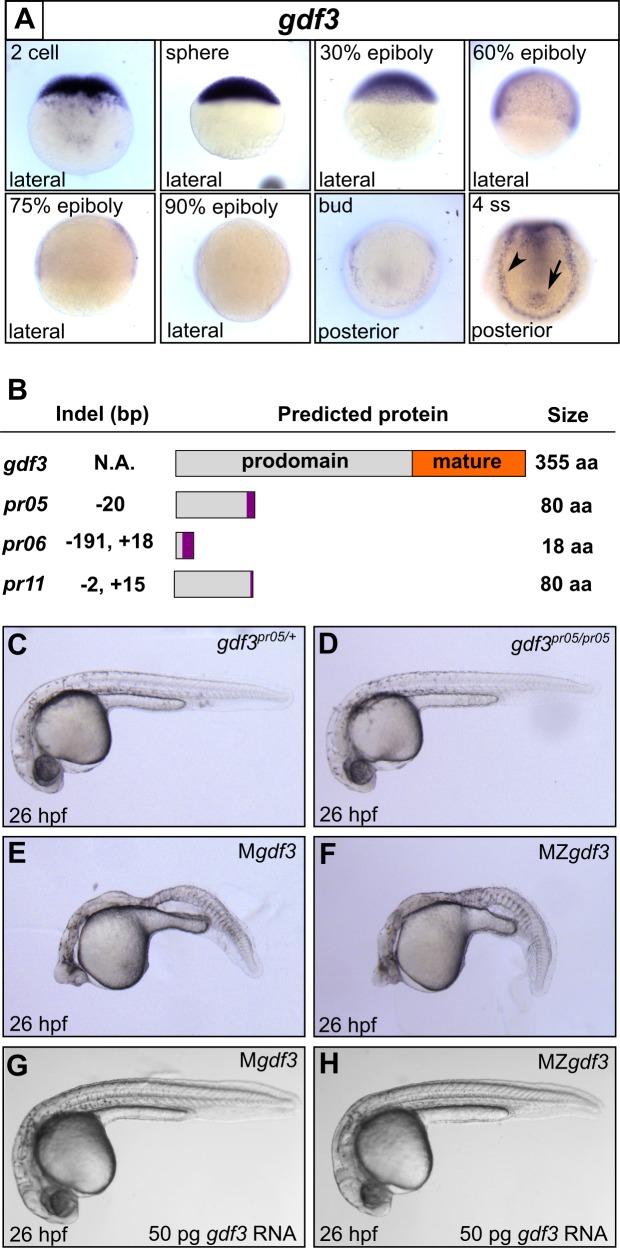
gdf3 is expressed maternally and ubiquitously during early stages of zebrafish development (Figure 1A) (Helde and Grunwald, 1993). The amount of gdf3 mRNA diminishes rapidly through the blastula and gastrula stages and disappears by approximately 90% epiboly (Helde and Grunwald, 1993). This is followed by the return of expression in a tissue-restricted manner starting at the late bud stage and progressing into somitogenesis, when expression is observed in the lateral plate mesoderm (LPM) and in cells around KV (Figure 1A) (Peterson et al., 2013). This temporal and positional variation suggests that gdf3 has multiple roles in zebrafish development.
Figure 1. Mutations in maternal gdf3 lead to defects in embryonic patterning.
(A) RNA in situ hybridization shows gdf3 mRNA is broadly expressed throughout early zebrafish development, followed by specific expression in Kupffer’s vesicle (arrow) and the lateral plate mesoderm (arrowhead). Views are indicated. (B) Schematic of gdf3 mutant alleles predicted to form truncated proteins due to early stop codons in the first exon. Purple regions indicate changes in amino acid sequence prior to the premature stop codon in each allele (see Material and Methods for more details). (C–F) Loss of maternally deposited Gdf3 causes patterning defects. Embryos heterozygous (C) or homozygous (D) for the pr05 allele appeared normal at 26 hpf. Heterozygous embryos from homozygous mothers (Mgdf) exhibited defects in mesoderm and endoderm formation (E). Maternal-zygotic gdf3 embryos (MZgdf3) had similar mesoderm and endoderm defects as Mgdf3 embryos (F). gdf3 mRNA injected at the one cell stage rescued defects in both Mgdf3 (G) and MZgdf3 (H) embryos. C-H are lateral views.
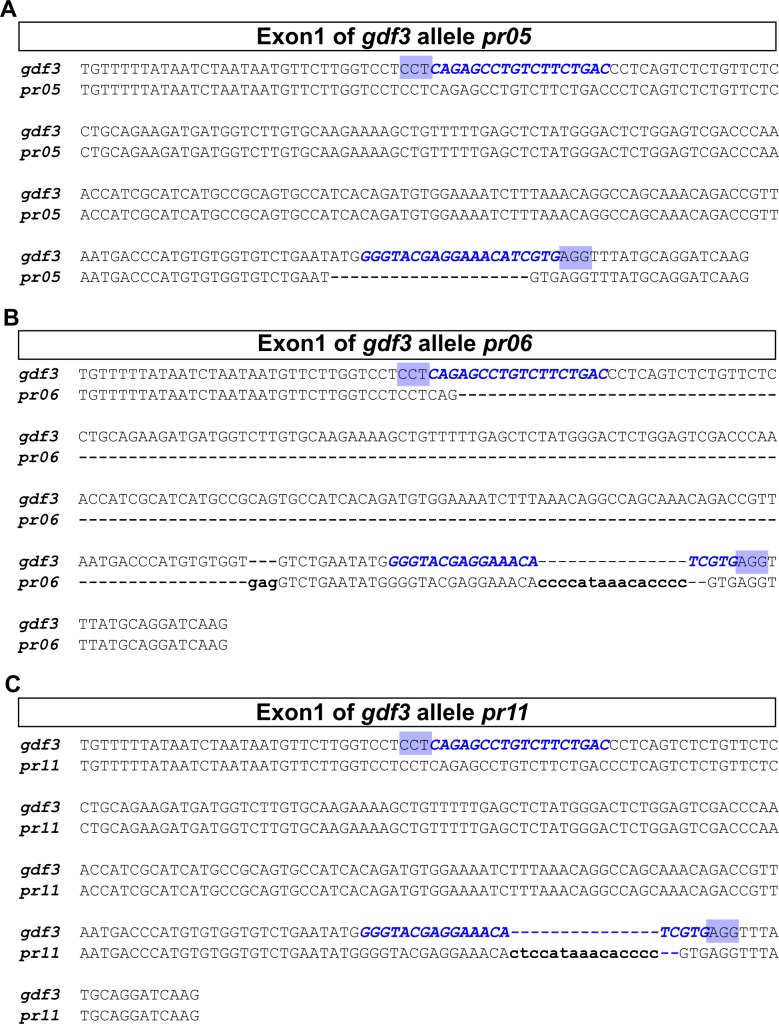
Figure 1—figure supplement 1. Schematic of gdf3 mutant allele sequences.
Figure 1—figure supplement 2. Multiple gdf3 mutant alleles have similar phenotypes.
To investigate these potential roles, we used CRISPR/Cas9 to generate stable zebrafish lines containing mutations in the first exon of the gdf3 locus (Figure 1B, Figure 1—figure supplement 1). Each of the three mutant alleles contained a premature stop codon in the first exon, prior to the mature ligand domain-encoding region, and are thus predicted to be null alleles. Homozygous gdf3 zygotic mutant embryos (Zgdf3) displayed no gross morphological phenotypes, were indistinguishable from heterozygous controls (Figure 1C and D), and grew to adulthood. By contrast, embryos lacking maternal gdf3 (Mgdf3) displayed overt developmental defects (Figure 1E) and did not survive beyond 3 days post fertilization (dpf). Mgdf3 and maternal-zygotic gdf3 (MZgdf3) embryos were indistinguishable from each other (Figure 1E and F), suggesting that the loss of Zgdf3 did not contribute further to the gross Mgdf3 phenotype. Moreover, these defects could be rescued with wild-type (WT) gdf3 mRNA (Figure 1G and H), a result that confirms the mutant phenotypes arose from disruption of the gdf3 locus.
The defects in Mgdf3 and in MZgdf3 embryos strongly resemble the phenotypes exhibited by double mutants for the zebrafish Nodal orthlogs ndr1 and ndr2 (Feldman et al., 1998) and MZ mutants for the essential Nodal co-receptor, teratocarcinoma-derived growth factor 1 (tdgf1; previously known as one eyed pinhead) (Gritsman et al., 1999). Similarities include the dorsal accumulation of cells following convergence and epiboly in Mgdf3 and MZgdf3 embryos at the tailbud stage, displacing the anterior most region of these embryos vegetally by 90 degrees compared to WT embryos (Figure 1—figure supplement 2E and F; compare to embryos in Gritsman et al., 1999). Additionally, Mgdf3 and MZgdf3 embryos at 26 hours post fertilization (hpf) are cyclopic and exhibit extensive defects in mesoderm and endoderm formation (Figure 1E and F and below), again reminiscent of ndr1;ndr2 double mutants and MZtdgf1 mutants (Feldman et al., 1998; Gritsman et al., 1999). Thus, maternally-supplied gdf3 is required for early embryogenesis, and loss of function cause defects that are consistent with strong reduction of early Nodal signaling.
Gdf3 is necessary for the expression of Nodal target genes responsible for mesendoderm formation
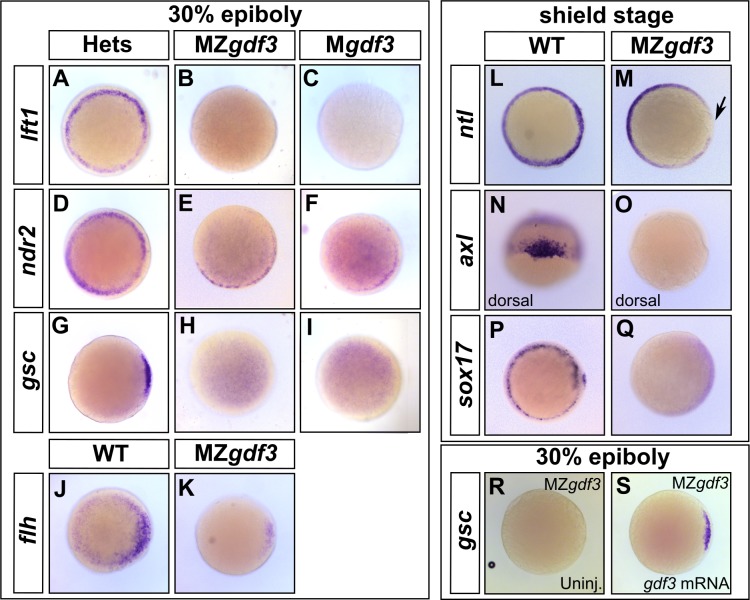
Since Mgdf3 and MZgdf3 mutants resemble embryos lacking Nodal signaling, we assessed Nodal target gene expression in these mutants. The Nodal target gene lefty1 (lft1) was absent in Mgdf3 and MZgdf3 embryos (Figure 2A–C). While expression of ndr2 initiated in Mgdf3 and MZgdf3 embryos, it was expressed at lower levels compared with controls (Figure 2D–F); since Nodal signals propagate ndr2 expression, this result and the absence of lft1 suggests that Nodal signaling is markedly reduced or lost in the absence of maternal gdf3. In agreement, Nodal targets required for mesoderm and endoderm formation (Feldman et al., 1998) were also lost or decreased in Mgdf3 and MZgdf3 mutants. goosecoid (gsc) (Stachel et al., 1993), a marker of progenitors of the prechordal plate, is absent in Mgdf3 and MZgdf3 embryos (Figure 2G–I). Expression of floating head (flh) (Talbot et al., 1995), a Nodal target that marks notochord progenitors (Gritsman et al., 2000), is decreased in Mgdf3 and MZgdf3 embryos (Figure 2J and K).
Figure 2. Gdf3 is necessary for the expression of Nodal target genes.
(A–K) RNA in situ hybridization of Nodal target gene expression at 30% epiboly. (A–C) lefty1 (lft1) is normally expressed in the margin (A), but is lost in maternal-zygotic gdf3 mutant embryos (MZgdf3, B) and heterozygous embryos derived from maternal homozygotes (Mgdf3, C). (D–F) cyclops (cyc) is normally expressed in the margin (D). Expression of cyc is reduced in MZgdf3 (E) and Mgdf3 (F) embryos. (G–I) goosecoid (gsc) is normally expressed in the dorsal region (G), but is absent in MZgdf3 (H) and Mgdf3 (I) embryos. (J–K) floating head (flh) is normally expressed in the dorsal region (J) but is diminished in MZgdf3 (K) embryos. (L–Q) RNA in situ hybridization of Nodal target gene expression at the shield stage. (L–M) no tail (ntla) is expressed along the margin in WT embryos (L), but is absent from the dorsal end of MZgdf3 embryos (M, arrow). (N, O) axial (axl) is expressed in the dorsal region of WT embryos (N) but absent in MZgdf3 embryos (O). (P, Q) sox17 is expressed in the margin of WT embryos (P) but absent in MZgdf3 embryos (Q). (R–S) Loss of gsc in MZgdf3 embryos (R) can be rescued by injection of 50 pg gdf3 mRNA. All views are animal, with dorsal to the right and ventral to the left,unless otherwise indicated.
Figure 2—figure supplement 1. Loss of Nodal target gene expression can be rescued with gdf3 mRNA injection.
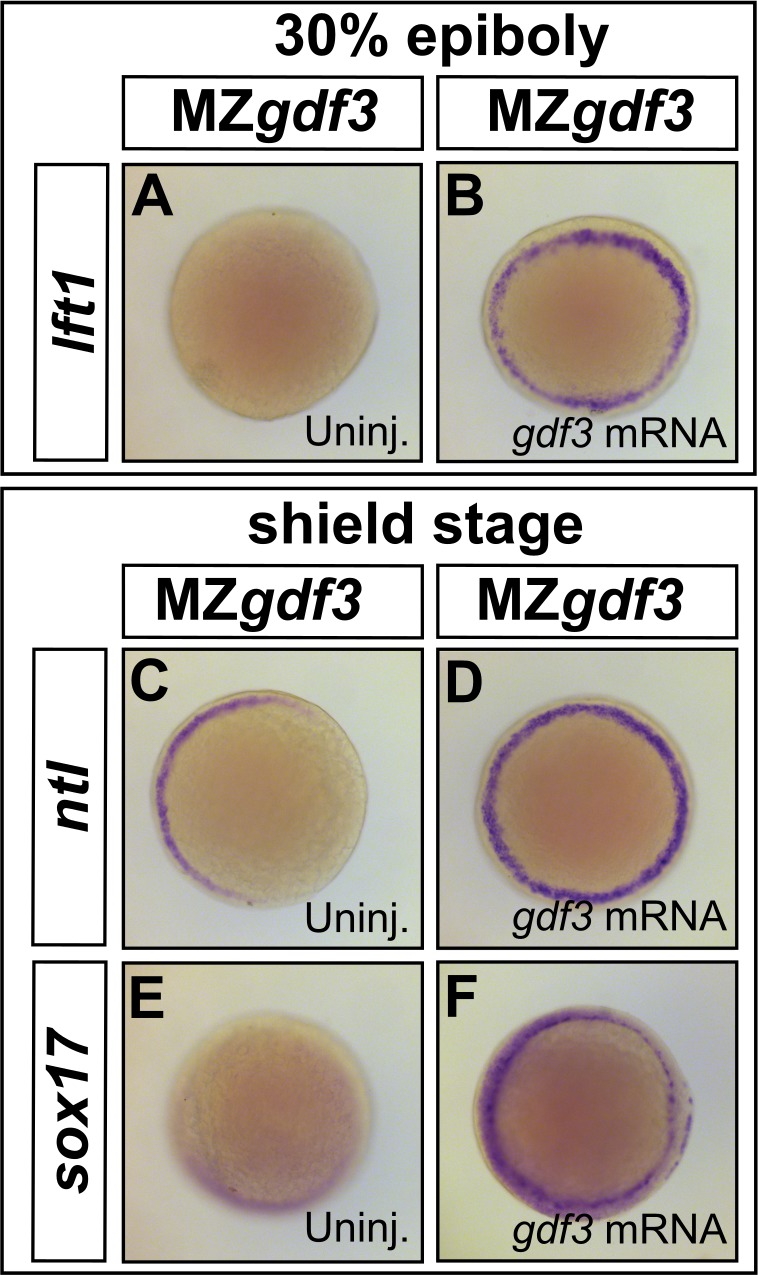
Similarly, expression of no tail (ntl) in the dorsal region (Schulte-Merker et al., 1992), a domain that requires the highest levels of Nodal signaling (Gritsman et al., 2000), is absent in Mgdf3 and MZgdf3 embryos (Figure 2L and M). axial (axl) (Strähle et al., 1993), a marker of progenitors of axial mesoderm and endoderm, and sox17 (Alexander and Stainier, 1999), a marker of presumptive endoderm, were both absent in Mgdf3 and MZgdf3 embryos (Figure 2N–Q). Taken together, these results suggest that Nodal signaling is strongly diminished or absent in Mgdf3 and MZgdf3 mutants and, as a result, mesoderm and endoderm are not induced. To confirm that these effects on Nodal target genes were a result of the loss of Gdf3, we injected gdf3 mRNA into MZgdf3 embryos at the one-cell stage. This led to the recovery of gsc (Figure 2R and S), lft1, ntl, and sox17 expression, (Figure 2—figure supplement 1A–F) confirming the role of gdf3 in regulating Nodal target genes.
Bmp and residual Nodal signaling promote tail mesoderm formation in Mgdf3 and MZgdf3 embryos
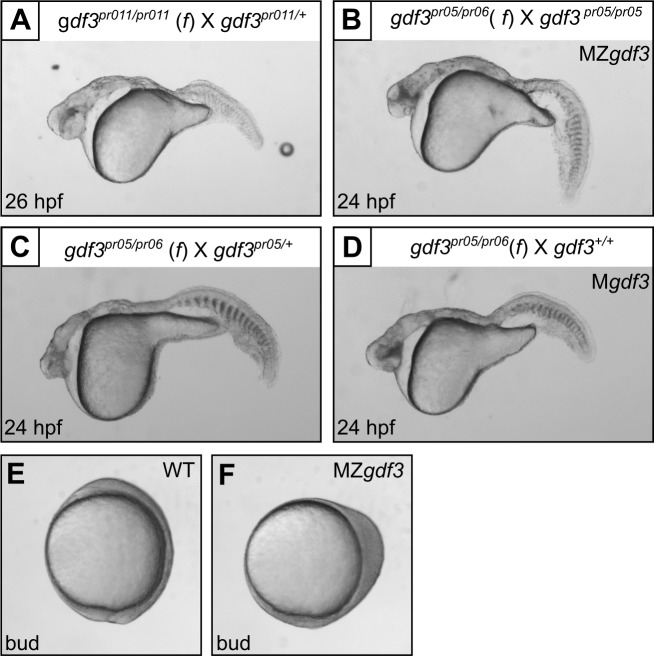
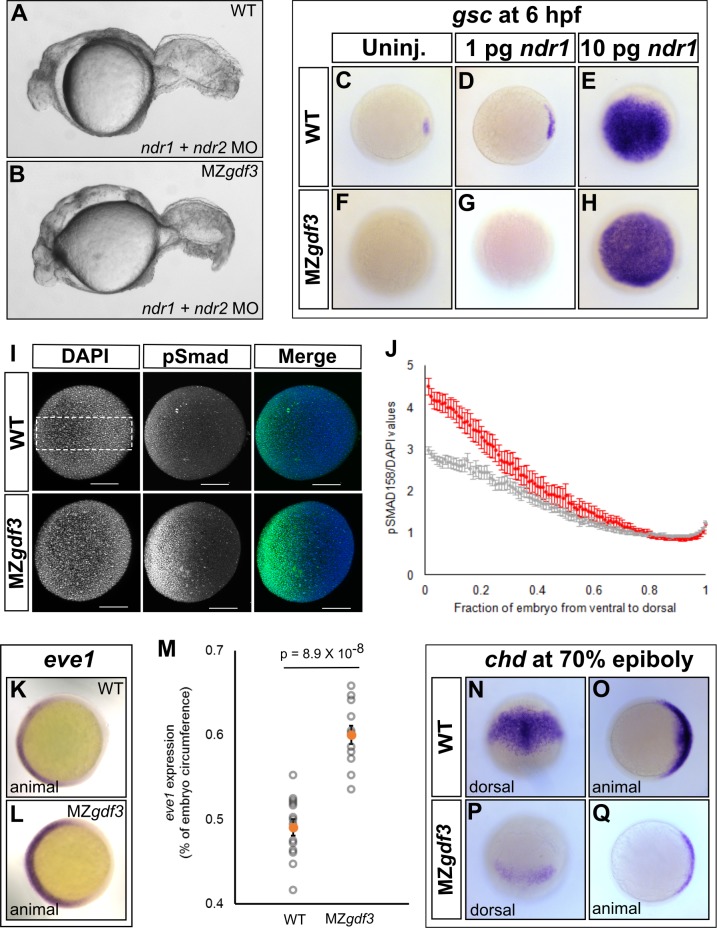
While Mgdf3 and MZgdf3 mutants strongly resemble ndr1:ndr2 double mutants, Mgdf3 and MZgdf3 embryos consistently exhibit longer tails. This phenotype resembles that of MZfoxh1p1r/pr1 mutants in which a subset of Nodal signaling is eliminated by loss of the Foxh1 transcription factor, but signaling through other Nodal-activated transcription factors is retained (Slagle et al., 2011). To determine if low levels of Nodal signaling remain in the absence of maternal Gdf3, we injected Mgdf3 and MZgdf3 embryos with ndr1:ndr2 MOs. Injection of the ndr1:ndr2 MOs into MZgdf3 mutants led to a tail defect that closely resembled the tail of ndr1:ndr2 double morphants (Figure 3A and B) and ndr1:ndr2 double mutants (Feldman et al., 1998). We then overexpressed Nodal using exogenous ndr1 mRNA to examine whether higher amounts of Nodal could activate target gene expression in MZgdf3 mutants. Although 1 pg of ndr1 led to a modest expansion of gsc expression in WT embryos (Figure 3C and D), it did not activate gsc expression in MZgdf3 embryos (Figure 3F and G). By contrast, 10 pg of injected ndr1 mRNA led to broad gsc expression in both WT and MZgdf3 mutant embryos (Figure 3E and H). These data, combined with our marker gene expression analysis, suggests that there is residual low-level Nodal activity present in the Mgdf3 and MZgdf3 mutant embryos that is inadequate for the formation of head and trunk endoderm.
Figure 3. Loss of Gdf3 leads to expansion of Bmp signaling.
(A) Knockdown of nodal related 1 (ndr1) and nodal related 2 (ndr2) in wild-type (WT) embryos causes complete loss of head and trunk mesendoderm and defects in tail patterning. (B) ndr1:ndr2 knockdown in MZgdf3 embryos causes loss of residual tail patterning, producing embryos that more closely resemble those completely lacking Nodal signaling (A). (C-E) Overexpression of 1 pg or 10 pg ndr1 led to increased expression of the Nodal target gene goosecoid (gsc) in WT embryos. (F-H) Overexpression of 10 pg ndr1, but not 1 pg, produced widespread expression of gsc in MZgdf3 mutants. (I) Immunostaining of phosphorylated Smad1/5/8 (green) counterstained with DAPI (blue). Ventral pSmad levels were increased in MZgdf3 mutants. (J) Plotting the normalized pSmad1/5/8 intensity over position on the ventral-dorsal axis confirms that MZgdf3 embryos (n = 14) have increased BMP signaling in the ventral region compared to WT (n = 13). (K-M) RNA in situ hybridization of eve1 showed expansion of expression in the ventral region of MZgdf3 embryos (K) compared to WT (L). (M) Quantitative comparison of WT (n = 14) and MZgdf3 (n = 12) embryos confirmed expansion of the ventral eve1 expression domain in MZgdf3 embryos. (N-Q) RNA in situ hybridization of chordin (chd) revealed its expression is reduced in MZgdf3 embryos. All views are animal unless otherwise indicated. In M, p-value obtained by Student’s t-test (two-sided, homoscedastic). Error bars in J and M, standard error of the mean. Matlab code and data are available as Figure 3—source code 1, 2, and 3 and Figure 3—source data 1 and 2.
Low levels of Nodal signaling act in concert with Bmp signaling to correctly form the posterior structures in the zebrafish embryo (Fauny et al., 2009). It has been reported that Gdf3 actively inhibits Bmp signaling in Xenopus and in cultured stem cells (Levine and Brivanlou, 2006). Additionally, the depletion of Gdf3 in Xenopus results in reduced expression of the Bmp inhibitor chordin (Birsoy et al., 2006). We therefore examined Bmp signaling in Mgdf3 and MZgdf3 embryos by assessing levels of phosphorylated Smad1/5/8 (p-Smad1/5/8), a modification induced by Bmp signaling. At 70% epiboly, a ventral-to-dorsal increase in p-Smad1/5/8 was observed in MZgdf3 mutants compared to controls, with the ventral end exhibiting the largest increase in phosphorylation (Figure 3I and J). Consequently, we observed an expansion in expression of the Bmp target and ventral mesoderm marker eve1 (Joly et al., 1993) towards the dorsal side of the embryo (Figure 3K–M). These data suggest that Gdf3 plays a role in modulating Bmp signaling and restricting the expansion of the ventral mesoderm. We also note that dorsal expression of the Bmp inhibitor chordin (chd) (Figure 3N–Q) is decreased in Mgdf3 and MZgdf3 embryos; this would also allow for the increased Bmp signaling observed in Mgdf3 and MZgdf3 embryos.
Gdf3 is required for proper KV architecture and the expression of LPM spaw
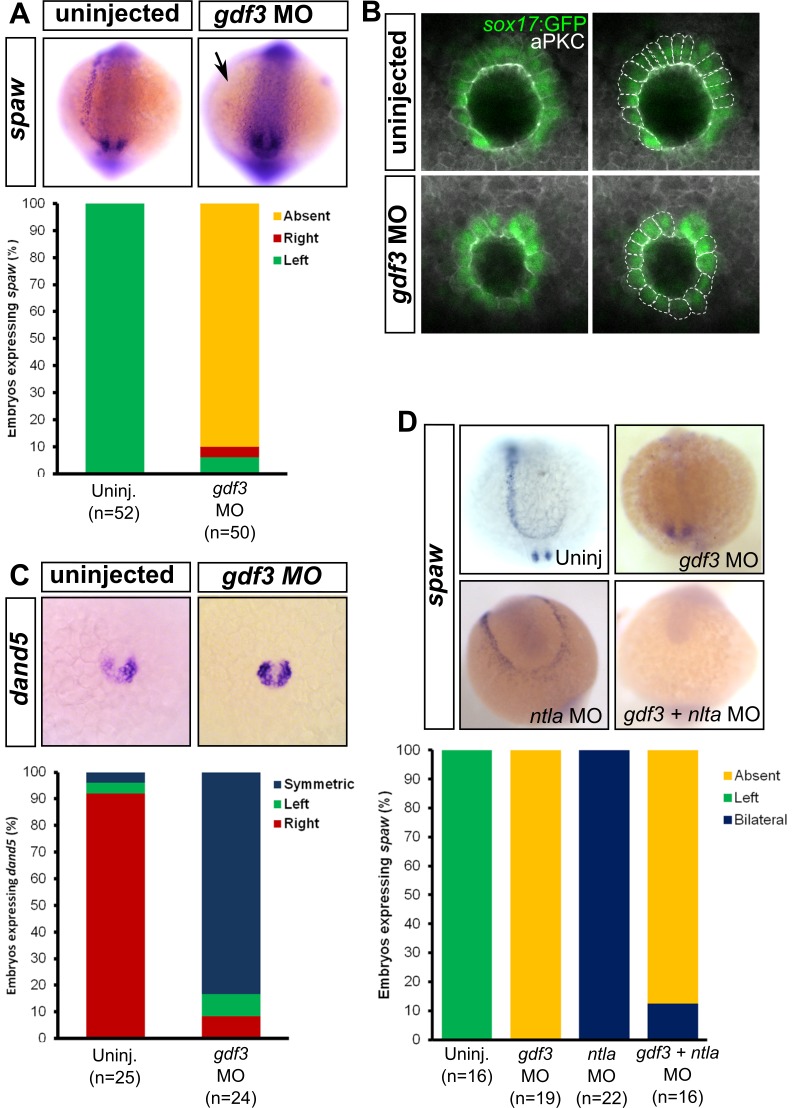
Nodal signaling has at least two roles in L-R patterning at different stages of zebrafish development: first at an early stage by directing correct patterning of the dorsal cells to properly form KV (Aamar and Dawid, 2010; Compagnon et al., 2014), and second, by inducing expression of the nodal ortholog spaw in the LPM (Long et al., 2003; Peterson et al., 2013). Gdf3 is also required for Nodal signaling in the LPM, since MO knockdown of Gdf3 reduces the expression of the nodal ortholog spaw in this tissue (Figure 4A) (Peterson et al., 2013). Therefore, we expected L-R patterning defects in our mutants. However, Zgdf3 mutants did not exhibit defects in the laterality of heart jogging (data not shown), an asymmetric morphogenetic event that acts as a measure of L-R patterning (Baker et al., 2008; Smith et al., 2008; de Campos-Baptista et al., 2008). Gdf3 is a stable protein since it remains detectable in the zebrafish embryo as late as 2.5 dpf, well beyond stages of reported mRNA expression (Helde and Grunwald, 1993; Dohrmann et al., 1996). In the absence of zygotic Gdf3, maternally supplied Gdf3 might compensate in L-R patterning events. Unfortunately, Mgdf3 and MZgdf3 mutants cannot be assessed for L-R patterning defects as they do not produce LPM or mesendoderm-derived organs. Therefore, we used a reported translation-blocking MO (Ye et al., 2010) to reduce the amount of maternal and zygotic Gdf3 to levels that allow for mesendoderm formation, and yet allow us to further analyze the role of Gdf3 in L-R patterning.
Figure 4. Attenuated production of Gdf3 leads to left-right patterning defects.
(A) spaw expression is altered in gdf3 morphants, with most lacking expression in the lateral plate mesoderm (arrow), as shown by RNA in situ hybridization. (B) Immunostaining of aPKC (white) and GFP driven by the sox17 regulatory region (green) in Kupffer’s Vesicle (KV) at 10 ss. The anterior-posterior asymmetry in cell morphology is lost in gdf3 morphants. Cells are outlined by white dashed lines. (C) RNA in situ hybridization reveals dand5 expression is symmetric around KV in gdf3 morphants at 10 ss. (D) Loss of ntla leads to robust spaw expression in both the left and right LPM and a loss of KV and associated gene expression. spaw expression in ntla + gdf3 double morphants is completely lost, supporting the idea that Gdf3 is required for robust Nodal expression given Nodal is unable to activate in the absence of inhibitors. Raw data provided in Figure 4—source data 1–3.
KV development is aberrant in MO-injected embryos. Normally, the cells in the anterior-dorsal region of KV are columnar and more tightly packed as compared to the ventral region (Figure 4B, see uninjected). This architecture is proposed to generate left-ward asymmetric flow that is faster across the anterior-dorsal portion of KV (Prabhat et al., 2007; Okabe et al., 2008; Supatto et al., 2008; Wang et al., 2012; Grimes and Burdine, 2017). In Gdf3 knockdowns, this architecture is lost and all the cells exhibited a cuboidal morphology (Figure 4B, see gdf3 MO). These data demonstrate that Gdf3 knockdown can adversely affect the formation of KV, which would prevent the generation of normal asymmetric fluid flow.
Proper KV formation and flow is important for generating asymmetric expression of the Nodal inhibitor dand5 in KV (Hashimoto et al., 2004; Hojo et al., 2007; Schweickert et al., 2010; Lopes et al., 2010). In contrast to WT embryos which exhibit a right-biased expression of dand5, Gdf3 knockdowns expressed dand5 symmetrically (Figure 4C). To determine if the symmetrical expression of dand5 leads to the loss of spaw expression in the left LPM of Gdf3 knockdowns, we prevented both the formation of KV and the expression of dand5 by also knocking down ntla with a MO. Loss of ntla leads to loss of KV, dand5 expression, and notochord (Amack et al., 2007; Hashimoto et al., 2004; Odenthal et al., 1996). In ntla mutants and morphants, spaw expression occurs at high levels in both the left and right LPM, presumably due to the loss of dand5 at KV and the Nodal inhibitor lefty1 in the midline (Long et al., 2003; Lenhart et al., 2011; Burdine and Grimes, 2016) (Figure 4D). We found that coinjection of ntla MO did not restore spaw expression in the LPM of gdf3 morphants (Figure 4D). This suggests that Gdf3 is required for robust Spaw signaling, and without Gdf3, Spaw is unable to induce its own expression, even in the absence of midline-derived Spaw inhibitors.
Rescue of mesendoderm defects in MZgdf3 embryos reveals a role for zygotic Gdf3 in L-R patterning
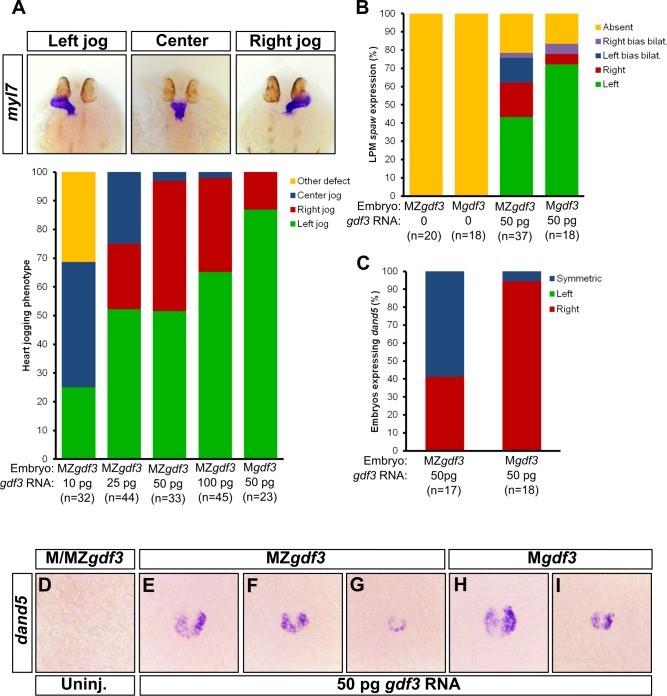
To confirm our Gdf3 knockdown results, and to further examine the potential role of zygotic gdf3 in L-R patterning, we overexpressed gdf3 mRNA to rescue the early mesendoderm defects in Mgdf3 and MZgdf3 embryos, and then analyzed L-R asymmetry in the embryos. A similar strategy was utilized to examine L-R patterning in MZtdgf1 embryos (Yan et al., 1999). This approach additionally allows us to examine potential differences in L-R patterning between Mgdf3 and MZgdf3 mutants, i.e. in the presence or absence of zygotic gdf3, respectively. Injection of 50 pg of gdf3 mRNA in Mgdf3 embryos, which retain zygotic expression of gdf3, rescued mesendoderm formation and resulted in ~90% of embryos with normal L-R patterning based on the direction of heart jogging (Figure 5A). Injection of 50 pg of gdf3 mRNA in MZgdf3 embryos also rescued mesendoderm defects but these embryos exhibited defects in heart jogging laterality, with ~50% of hearts jogging incorrectly to the right or remaining midline (Figure 5A). These data suggest that zygotic gdf3 facilitates left-right patterning in the absence of sufficient maternal gdf3 contribution. However, since Zgdf3 mutants exhibit no L-R defects, zygotic Gdf3 is not required for L-R patterning in the presence of maternal Gdf3. In further support of the hypothesis that Gdf3 produced early in development is stable and sufficient to function in L-R patterning, injecting increasing amounts of gdf3 mRNA increased the number of MZgdf3 embryos with correct leftward jogged hearts (Figure 5A).
Figure 5. Zygotic Gdf3 can restore proper left-right patterning to embryos rescued from early mesendoderm defects.
(A) Heart jogging defects improved with increasing amounts of injected gdf3 mRNA in MZgdf3 embryos. Zygotic gdf3 (present in Mgdf3 embryos) further rescued heart jogging defects in embryos injected with 50 pg of gdf3 mRNA. (B) M/MZgdf3 embryos lack spaw expression at 12 ss. Embryos injected with 50 pg of gdf3 mRNA rescued spaw expression in the LPM. Proper spaw expression in the left LPM was further restored in the presence of zygotic gdf3. (C) dand5 expression was recovered in MZgdf3 embryos injected with gdf3 mRNA, with most embryos containing symmetric expression of dand5 in KV (F,G). (C) Most Mgdf3 embryos injected with gdf3 mRNA contained proper right-biased asymmetric expression of dand5 (H,I). (D) dand5 expression was absent in all uninjected M/MZgdf3 embryos. Raw data provided in Figure 5—source data 1–3.
We next analyzed molecular markers of L-R asymmetry in rescued mutants. While the LPM does not form in Mgdf3 and MZgdf3 embryos, presence of LPM and expression of spaw in the LPM is rescued in mutant embryos injected with gdf3 mRNA (Figure 5B). In injected MZgdf3 embryos, LPM spaw laterality was irregular with ~40% of embryos displaying correct left-sided expression at the 12 somite stage (ss). In injected Mgdf3 embryos, where zygotic gdf3 is still present, the number of embryos exhibiting correct left-sided expression of spaw increased to ~70%. This further supports the conclusion that zygotic gdf3 contributes to correct L-R patterning in the absence of maternal Gdf3.
Since embryos treated with gdf3 MO exhibited defects in KV cell morphology and functionality, we analyzed KV functionality in mesendoderm-rescued Mgdf3 and MZgdf3 embryos. While correct asymmetric dand5 expression in KV was restored in rescued Mgdf3 embryos (Figure 5C and H–I), rescued MZgdf3 embryos expressed dand5 symmetrically (Figure 5C and F–G), indicating loss of functional flow in KV, likely owing to aberrant KV morphology. These data suggest that zygotic Gdf3 is required for proper formation and function of KV in the absence of maternal Gdf3.
Discussion
Signaling by multiple members of the TGFβ superfamily of ligands is important for patterning of early vertebrate embryos. Here, we show that the gene encoding the Vg1 ortholog in zebrafish, gdf3, is a maternal effect gene for early embryonic patterning. We provide evidence that the role of maternally supplied Gdf3 is to promote robust levels of Nodal signaling. Others have reported that Nodal and Vg1 orthologs can bind to one another (Tanaka et al., 2007; Fuerer et al., 2014). Furthermore, Vg1 requires the same pathway components as Nodal ligands for signaling, including the essential co-receptor Tdgf1 (Cheng et al., 2003). Thus, we hypothesize that Gdf3 dimerizes with Nodal to facilitate Nodal signaling during development. Our evidence suggests that lower levels of Nodal signaling still occur in the absence of maternal Gdf3. For instance, Mgdf3 and MZgdf3 embryos produce more posterior tail structures than embryos completely lacking Nodal signaling. Although Nodal can signal independently of Gdf3, target genes required for head and trunk mesendoderm are not activated without Gdf3 unless Nodal is overexpressed beyond physiological levels. Moreover, we find correct spatial expression of mesendoderm marker genes in MZgdf3 embryos injected with gdf3 mRNA, which suggests that Gdf3 plays a permissive role in mesendoderm formation while spatial restriction of ndr1 and ndr2 expression is critical for patterning. Gdf3 has also been proposed to act as a direct inhibitor of Bmp signaling (Birsoy et al., 2006; Levine and Brivanlou, 2006), and we observe an increase in Bmp signaling in Mgdf3 and MZgdf3 embryos by assaying for pSMAD1/5/8. However, we see reduction of Bmp antagonists in these embryos, such as chordin, and thus additional studies are warranted to determine if Gdf3 is playing a direct or indirect role in modulating Bmp signaling in the early embryo.
We demonstrate that knockdown of Gdf3 can affect L-R patterning at two crucial steps: in formation of the L-R coordinator KV and in promoting robust expression of Nodal/Spaw in the LPM. For KV to form properly, correct expression of sox17 is required (Aamar and Dawid, 2010), which itself requires robust Nodal signaling during germ layer formation (Sako et al., 2016). Furthermore, regional cell shape differences in KV depend on the presence of the neighboring notochord (Compagnon et al., 2014) which is absent in Nodal-signaling deficient embryos (Feldman et al., 1998). In our Gdf3 knockdown experiments, KV fails to acquire the regional cell shape differences that promote correct fluid flow, which is required for proper asymmetric expression of dand5 (Figure 6C and D) (Schweickert et al., 2010; Lopes et al., 2010). In addition to this phenotype, knockdown of Gdf3 also prevents spaw expression from occurring in the LPM even in the absence of Nodal inhibitors Dand5 and Lefty1 (Figure 6E). These knockdown experiments suggest that Gdf3 plays a role both in KV organization and downstream, in the activation of LPM spaw. A previous report suggested that Gdf3 knockodwn did not affect KV, and that the loss of Spaw expression in the LPM of these embryos could be rescued by injection of dand5 or lefty1 morpholinos (Peterson et al., 2013). However, given that our data suggest maternal Gdf3 is responsible for proper L-R patterning, we suggest the differences in our results reflect the amount by which maternal Gdf3 is reduced with the two different AUG morpholinos that were used.
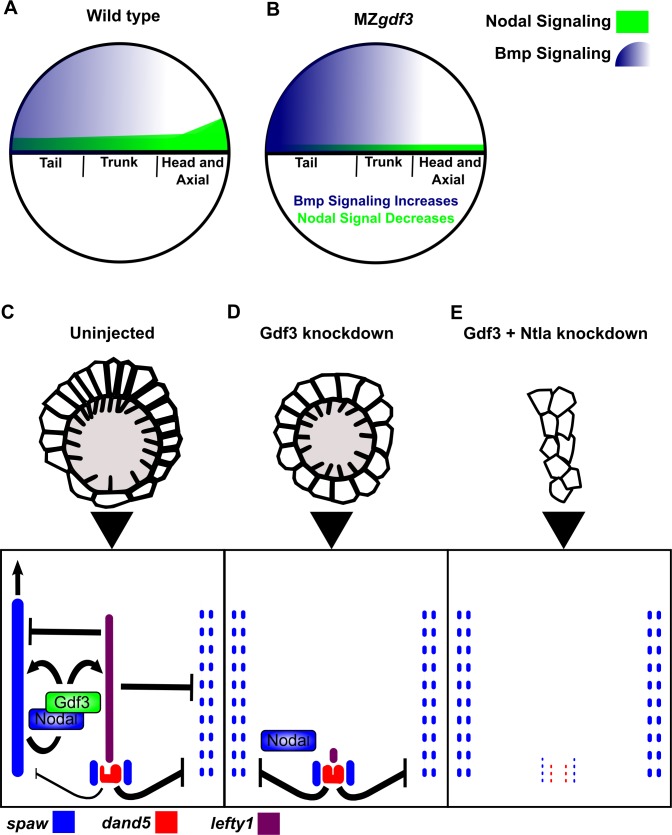
Figure 6. Model of Gdf3 function in early zebrafish development.
(A) In wild-type (WT) embryos, Bmp signaling (blue) is established in a ventral-to-dorsal gradient during blastula and gastrula stages, while Nodal signaling (green) is established along the margin, with strongest expression on the dorsal side. (B) In MZgdf3 embryos, Nodal signaling is weak leading to reduced expression of Nodal target genes including the Bmp inhibitor Chordin. Bmp signaling is increased and leads to an extension of ventral mesoderm markers. However, as Nodal signaling is depleted or lost, the majority of mesoderm and endodermal cell fates fail to be maintained. (C) Proper development of Kupffer’s vesicle includes the generation of asymmetries in cell shape along the anterior-posterior axis of the structure. Anterior cells, closest to the notochord, take on columnar shapes and become more tightly packed, leading to a greater number of cilia in this region. This architecture is required for asymmetric fluid flow in KV and proper dand5 expression. (D) Attenuation of maternal Gdf3 leads to improper cell morphology during KV development, which in turn leads to irregular dand5 expression. Dysregulation of dand5 expression on the left may then inhibit spaw in the left LPM. (E) Removal of Nodal antagonists Lefty1 and Dand5 via ntla morpholino-mediate knockdown does not restore spaw expression in the LPM. This suggests that Gdf3 plays a more direct role in regulating spaw expression, independent of its effects on KV development. Dotted lines (C, D, E) represent lost gene expression.
Analysis of L-R patterning events in mesendoderm-rescued Mgdf3 and MZgdf3 complemented our Gdf3 knockdown experiments, and confirmed roles for Gdf3 in L-R patterning. Importantly, we demonstrated that the presence of zygotic gdf3 improved the rescue of L-R patterning defects in mesendoderm-rescued mutants. The lack of L-R patterning phenotypes in Zgdf3 mutants was unexpected, especially since gdf3 is zygotically expressed in very strict domains that also express crucial L-R patterning genes such as the KV region and the LPM. It could be that while maternal Gdf3 is sufficient for proper L-R patterning, zygotic expression provides robustness to the system. In the conditions used to raise zebrafish in the laboratory, such as controlled temperature, abundant food, and lack of predators, maternally supplied gdf3 is sufficient for this process. However, in situations where lower levels of maternal product are supplied, zygotic Gdf3 might then be required alongside maternal Gdf3 for robust L-R patterning.
Our results demonstrate that Gdf3 acts permissively to promote robust Nodal signaling, and yet is spatially restricted to KV and LPM during L-R patterning. Given that other L-R patterning components are expressed in these distinct domains at this time, gdf3 expression could be restricted to these same domains by similar patterning mechanisms, even if spatial restriction of its expression is not required. Taken together, we propose that Gdf3 and Spaw dimerize to promote robust Nodal signaling in the LPM. In the future, it will be interesting to see if dimerization of these ligands allows Spaw to more efficiently travel to, and propagate through the LPM.
Collectively, our data suggest that Gdf3 functions in multiple developmental processes. During the blastula and gastrula stages, Gdf3 affects Nodal activity required for organization of the vertebrate embryo as well as the formation of KV. During somitogenesis it is required for Nodal signaling in the LPM. This agrees with work performed in mouse, where lines with targeted mutations in the Vg1 mouse orthologs Gdf1 and Gdf3 exhibited defects in mesoderm formation and L-R patterning (Rankin et al., 2000; Andersson et al., 2006; 2007). Our work also suggests that Gdf3 functions as a co-ligand for Nodal signaling. Heterodimerization of other zebrafish TGFβ ligands in BMP signaling (Little and Mullins, 2009), lead us to propose a similar function for Gdf3 and Nodal ligands in Nodal signaling. Indeed, there is strong evidence that Vg1 orthologs can heterodimerize with Nodal to allow for robust or more efficient Nodal signaling (Tanaka et al., 2007; Fuerer et al., 2014). Overall, we conclude that Gdf3 facilitates Nodal ligands in generating a robust level of signaling activity required for both mesendoderm formation and proper L-R patterning.
Materials and methods
Key resources table.
| Reagent type (species) or resource |
Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| gene (Danio rerio) |
growth differentiation
factor 3 (gdf3) |
NA | ZFIN:ZDB-GENE-980526–389 | |
| gene (Danio rerio) | nodal-related 1 (ndr1) | NA | ZFIN:ZDB-GENE-990415–256 | |
| gene (Danio rerio) | nodal-related 2 (ndr2) | NA | ZFIN:ZDB-GENE-990415–181 | |
| gene (Danio rerio) | southpaw (spaw) | NA | ZFIN:ZDB-GENE-030219–1 | |
| gene (Danio rerio) |
DAN domain family,
member 5 (dand5) |
NA | ZFIN:ZDB-GENE-040421–2 | |
| genetic reagent (D. rerio) |
gdf3 allele pr05 | this paper | CRISPR/Cas9 generated deletion, predicted null | |
| genetic reagent (D. rerio) |
gdf3 allele pr06 | this paper | CRISPR/Cas9 generated indel, predicted null | |
| genetic reagent (ID. rerio) |
gdf3 allele pr11 | this paper | CRISPR/Cas9 generated indel, predicted null | |
| antibody | Anti-Digoxigenin-AP | Roche, 11093274910 | 1:3500 | |
| antibody | Anti-phosphorylated SMAD1/5/8 |
Cell Signaling Technologies, 9511S |
1:100 | |
| antibody | anti-PKCzeta (aPKC) | Santa Cruz Biotechnology, SC-216 |
1:400 | |
| antibody | Goat anti-Rabbit Alexa Fluor 647 |
Invitrogen, A21246 | 1:400 | |
| antibody | Donkey anti-Rabbit Alexa Fluor 647 secondary antibody |
Invitrogen, A31573 | 1:400 | |
| recombinant DNA reagent |
pCS2-nCas9n | Addgene plasmid # 47929 |
||
| recombinant DNA reagent |
pT7-gRNA | Addgene plasmid # 46759 |
||
| recombinant DNA reagent |
pCS2(+)-gdf3-WT | this paper | ||
| recombinant DNA reagent |
pCS2-3XFlag-sqt | PMID:27101364 | ||
| recombinant DNA reagent |
gdf3 in situ
probe plasmid |
PMID: 8405668 | ||
| recombinant DNA reagent |
ndr2 in situ
probe plasmid |
PMID: 9707578 | ||
| recombinant DNA reagent |
flh in situ
probe plasmid |
PMID: 7477317 | ||
| recombinant DNA reagent |
ntla in situ
probe plasmid |
PMID: 1295726 | ||
| recombinant DNA reagent |
gsc in situ
probe plasmid |
PMID: 8104775 | ||
| recombinant DNA reagent |
lft1 in situ
probe plasmid |
PMID: 10375514 | ||
| recombinant DNA reagent |
axial in situ
probe plasmid |
PMID: 7687227 | ||
| recombinant DNA reagent |
sox17 in situ
probe plasmid |
PMID: 10531029 | ||
| recombinant DNA reagent |
spaw in situ
probe plasmid |
PMID: 12702646 | ||
| recombinant DNA reagent |
dand5 in situ
probe plasmid |
PMID: 15084459 | ||
| sequence-based reagent |
gdf3 morpholino | Genetools | 5’-GCTCTGAGGAGGACCAAGAACATTA-3’ | |
| sequence-based reagent |
ntla morpholino | Genetools | 5’- GACTTGAGGCAGGCATATTTCCGAT-3’ | |
| sequence-based reagent |
ndr1 morpholino | Genetools | 5’-ATGTCAAATCAAGGTAATAATCCAC-3’ | |
| sequence-based reagent |
ndr2 morpholino | Genetools | 5’-GCGACTCCGAGCGTGTGCATGATG-3’ | |
| peptide, recombinant protein |
T4 PNK | NEB, M0201S | ||
| peptide, recombinant protein |
T4 DNA ligase | NEB, M0202S | ||
| peptide, recombinant protein |
BsmBI | NEB, R0580S | ||
| peptide, recombinant protein |
BglII | NEB, R0144S | ||
| peptide, recombinant protein |
SalI | NEB, R0138S | ||
| peptide, recombinant protein |
BamHI | NEB, R0136S | ||
| peptide, recombinant protein |
XbaI | NEB, R0145S | ||
| commercial assay or kit |
Qiaquick PCR Purification Kit |
Qiagen, 28104 | ||
| commercial assay or kit |
QIAquick Gel Extraction Kit |
Qiagen, 28704 | ||
| commercial assay or kit |
mMessage mMachine SP6 |
ThermoFisher, AM1340 |
||
| commercial assay or kit |
MEGAshortscript T7 Transcription kit |
ThermoFisher, AM1354 |
||
| commercial assay or kit |
QIAprep Spin Miniprep kit |
Qiagen, 27106 | ||
| commercial assay or kit |
Phusion High- Fidelity PCR Kit |
EB, E0553L | ||
| software, algorithm | Imaris | Image analysis and quantification | ||
| software, algorithm | ImageJ | Image analysis and quantification | ||
| software, algorithm | MATLAB | Image analysis and quantification |
Fish care
Zebrafish husbandry was performed using protocols in accordance with Princeton University Institutional Animal Care and Use Committee. Embryos were maintained at 28°C in Egg Water (300 mg/L Instant Ocean Sea Salts and 0.02 mg/L Methylene Blue).
Generation of gdf3 mutants
Insertions and/or deletions were produced in the first exon of zebrafish gdf3 using the CRISPR-Cas9 genome editing system. Target sites GGGTACGAGGAAACATCGTG (Chr17:4086653–4086672; Ensembl 88:10; genome assembly GRCz10) and GGGTCAGAAGACAGGCTCTG (17:4086856–4086875 Ensembl 88:10; genome assembly GRCz10) were selected using CHOPCHOP (Montague et al., 2014) (Figure 1—figure supplement 1). The sgRNAs were made with the guidance of published protocols (Cong et al., 2013; Jao et al., 2013). Oligos for each target site were annealed (5’-TAGGGTACGAGGAAACATCGTG-3’ annealed to 5’-AAACCACGATGTTTCCTCGTAC-3’)(5’-TAGGGTCAGAAGACAGGCTCTG-3’ annealed to 5’-AAACCAGAGCCTGTCTTCTGAC-3’) and phosphorylated at the 5’-end by incubating a 10 µl reaction mixture (1X T4 ligase buffer (NEB, B0202S), 10 µM each of the two oligos from a target-site, and 5 units of T4 PNK (NEB, M0201S)) with the following heating profile: 30 min at 37°C, 5 min at 95°C, a 2 °C/s decrease in temperature until 85°C, and cooled at 0.1 °C/s until 25°C. The annealed oligos were cloned into pT7-gRNA (Jao et al., 2013) by incubating a 10 µl reaction mixture (40 ng/µl of pT7-gRNA, a 1:20 dilution of the annealed oligos for an individual target site, 1X NEB buffer 3.1 (NEB, B7203S), 1X T4 ligase buffer (NEB, B0202S), 5 units of BsmBI (NEB, R0580S), 5 units of BglII (NEB, R0144S), 5 units of SalI (NEB, R0138S), and 200 units of T4 DNA ligase (NEB, M0202S)) with the following heating profile: 3 cycles of 20 min at 37°C, 15 min at 16°C, 10 min at 37°C, and 15 min at 55°C. Plasmids with the correct insertion were confirmed by Sanger sequencing. Plasmids were transformed into E.coli. DH5α, grown, and isolated using a QIAprep Spin Miniprep kit (Qiagen, 27106). The sgRNAs were then synthesized from BamHI (NEB, R0136S) digested plasmids using a MEGAshortscript T7 Transcription kit (ThermoFisher, AM1354). Cas9 mRNA was synthesized from NotI (NEB, R0189S) digested pCS2-nCas9n (Jao et al., 2013) using an mMessage mMachine SP6 Transcription Kit (ThermoFisher, AM1340). Both pT7-gRNA (Addgene plasmid # 46759) and pCS2-nCas9n (Addgene plasmid # 47929) were gifts from Wenbiao Chen.
Embryos were injected into the single cell at the one cell stage with a ~1.8 nl RNA mixture consisting of 240 pg of Cas9 mRNA and 10 pg of each sgRNA. The embryos were raised and incrossed to check for mutant phenotypes. Candidates containing potential germline transmission of a gdf3 mutant allele were isolated by outcrossing to WT zebrafish. Homozygous mutants for a gdf3 mutant allele were generated by incrossing heterozygous fish for that allele. Compound heterozygous fish with two different mutant alleles were generated by crossing heterozygous fish containing different mutant alleles. Mutations in the gdf3 locus were confirmed using PCR genotyping. Tail fin-clips were added to 50 μl of Lysis Buffer (10 mM Tris-HCl at pH 8.0, 50 mM KCl, 0.3% Tween 20, 0.3% NP40, and 1 mM EDTA) and incubated at 98°C for 10 min. The samples received 11 μl of 10 mg/ml ProK and were incubated overnight at 55°C. The samples were then incubated at 98°C for 10 min, cooled on ice, and diluted 1:18 with nuclease free water. PCR was performed using 5 μl of the diluted DNA extract. Phusion High-Fidelity PCR Kit (NEB, E0553L) was used with forward primer 5’-CAGATCCCAAAATCTTCATGC-3’ and reverse primer 5’-AAGTGTGGTCCGTAAGATCCTC-3’ to amplify the region containing potential mutations (Heating profile: 98°C for 30 s, followed by 30 cycles of 98°C for 10 s, 66°C for 30 s, and 72°C for 30 s, followed by 72°C for 5 min). DNA was size separated on a 3% Metaphor gel (Lonza, 50181) containing ethidium bromide and visualized using a UV light source. The bands were excised, gel purified using a Qiaquick Gel Extraction Kit (Qiagen, 28704), and sequenced.
Three mutations in gdf3 were obtained (Figure 1—figure supplement 1). Allele pr05 contains a 20 base pair (bp) deletion beginning at base 218 of the coding region, which causes a frame shift in eight amino acids (aa) and leads to a premature stop codon. This allele is predicted to produce an 80 aa truncated version of Gdf3. Allele pr06 contains a 189 bp deletion beginning at base 21 of the coding region immediately followed by a 3 bp insertion, and a second lesion that consists of a 15 bp insertion at base 236 immediately followed by a 2 bp deletion. The first lesion causes a frameshift that changes the sequence of 11 aa prior to a premature stop codon which is predicted to produce an 18 aa truncated version of Gdf3. Allele pr11 contains a 15 bp insertion after 235 bases in the coding region immediately followed by a 2 bp deletion. This indel causes a framshift that changes two aa prior to a premature stop codon which is predicted to produce an 80 aa truncated version of Gdf3.
Since, Mgdf3 and MZgdf3 embryos exhibited the same phenotype, they were used interchangeably. Additionally, all three gdf3 mutant alleles caused the same M/MZgdf3 phenotypes and did not complement each other (Figure 1—figure supplement 2A–D, data not shown), which confirmed that the defects in embryonic patterning we observed were due to a dysfunctional gdf3 locus. Since all the mutants exhibited the same phenotype, gdf3 mutants containing the pr05 allele were used for most this work.
Images of gross morphological phenotypes were acquired using a Leica DFC365 FX camera attached to a Leica M205 FA stereomicroscope.
Morpholino and RNA rescue injections
Morpholino injections
The previously published gdf3 AUG morpholino (Ye et al., 2010) (5’-GCTCTGAGGAGGACCAAGAACATTA-3’; Genetools), was used to knockdown translation of Gdf3. Each injection contained 5 ng of gdf3 AUG morpholino and 5 ng of p53 AUG morpholino (used to limit off-target effects [Ye et al., 2010]). Approximately 2 ng of the previously published ntla AUG morpholino (Nasevicius and Ekker, 2000) (5’- GACTTGAGGCAGGCATATTTCCGAT-3’; Genetools), was used to knockdown translation of Ntla. A mixture containing 10 ng of cyc (Karlen and Rebagliati, 2001) and 10 ng of sqt (Feldman and Stemple, 2001) AUG morpholinos (5’-GCGACTCCGAGCGTGTGCATGATG-3’, 5’-ATGTCAAATCAAGGTAATAATCCAC-3’; Genetools) was used to knockdown translation of Ndr1 and Ndr2. For all knockdowns, a morpholino mixture of ~1.8 nl was injected into the yolk of one cell stage embryos. All morpholino mixtures contained Danieau’s Buffer and 0.5 mg/ml phenol red.
RNA rescue injections and overexpression
The pCS2(+)-gdf3-WT construct was generated by inserting the complementary sequence of WT gdf3 mRNA into a pCS2(+) plasmid construct. The gdf3 insert was amplified from cDNA using primers with a 5’-end BamHI site (5’-AAGGATCCTGTTTTTATAATCTAATAATGTTCTTGG-3’) and a 5’-end XbaI site (5’-AATCTAGAATAGTAAAAGTTTTTATTATTACATTACAATG-3’). The amplicon was gel purified using a QIAquick Gel Extraction Kit (Qiagen, 28704) and both the gdf3 insert and the pCS2(+) destination plasmid were digested with BamHI (NEB, R0136S) and XbaI (NEB, R0145S). The digested DNA was purified using a Qiaquick PCR Purification Kit (Qiagen, 28104) and ligated using T4 DNA ligase (NEB, M0202S). The resulting plasmid was sequenced to confirm the insertion of gdf3 into the multiple cloning site of pCS2(+).The plasmid was digested with XbaI, and capped gdf3 mRNA was synthesized using an mMessage mMachine SP6 Transcription Kit. A 1.8 nl volume gdf3 mRNA in 0.1 M KCl was injected into the cell of a one cell stage Mgdf3 or MZgdf3 embryo. RNA in situ hybridization was used to analyze gsc, lft1, ntl, and sox17 expression. Nodal ovexpression was performed using nodal mRNA generated from a pCS2(+) plasmid, given to us as a gift from the Sampath Lab, containing the sequence for ndr1 linked to the sequence for 3XFlag (Wang et al., 2016). Capped nodal mRNA was synthesized and injected similar to the gdf3 mRNA rescues.
RNA in situ hybridization
Embryos were fixed at the 10 ss stage in 4% PFA overnight at 4°C. These embryos were washed with in PBST (1X PBS containing 0.1% Tween-20), dechorionated, transitioned to 100% methanol, and stored at −20°C for at least 1 day. The transition to methanol was done by performing five-minute washes in 75% 1X PBST:25% methanol, 50% 1X PBST:50% methanol, 25% 1X PBST:75% methanol, and 100% methanol. The embryos were then transitioned into 1X PBST by performing 5 min washes in 25% 1X PBST:75% methanol, 50% 1X PBST:50% methanol, and 75% 1X PBST:25% methanol. Embryos were then washed four times in 1X PBST with 5 min per wash. Somite stage embryos were incubated for 1 min in 1X PBST containing 0.01 mg/ml Proteinase K (Sigma-Aldrich, P2308) followed by a 20 min incubation in 1X PBST containing 4% paraformaldehyde. These embryos were then washed five more times in 1X PBST with 5 min per wash. Blastula and gastrula stage embryos did not undergo this Proteinase K treatment, extra fixation with 4% paraformaldehyde, or the extra five washes with 1X PBST. Embryos were incubated in HYB (50% formamide, 5X SSC, 500 µg/ml torula yeast RNA, 50 µg/ml heparin 0.1% Tween 20, and 9 mM Citric Acid (pH 6.0)) for 2 hr at 68°C. Embryos were then incubated overnight in HYB containing an insitu hybridization probe at 68°C. The next day, embryos were washed at 68°C in HYB, 75% HYB: 25% 2X SSC, 50% HYB: 50% 2X SSC, 25% HYB: 75% 2X SSC, and 2X SSC for 10 min each wash. Embryos were then washed twice in 0.2X SSC for 30 min each wash. The remaining washes were performed at room temperature. Embryos were washed in 75% 0.2X SSC: 25% 1X PBST, 50% 0.2X SSC: 50% 1X PBST, 25% 0.2X SSC: 75% 1X PBST, and 1X PBST for 5 min each wash. Next, embryos were incubated on a rocker for 2 hr in 1X PBST containing 2 mg/ml BSA and 2% normal sheep serum (NSS). Embryos were then incubated overnight on a rocker in 1X PBST containing 2 mg/ml BSA, 2% NSS, and 1:3500 of Anti-Digoxigenin-AP (Roche, 11093274910). The next day, the embryos were washed quickly in 1X PBST followed by six additional 15 min 1X PBST washes on a rocker. Embryos were then washed three times in NTMT (0.1 M Tris-Cl ph 9.5, 0.1 M NaCl, 0.05 M MgCl2, 0.1% Tween 20) and stained with 5 µl of NBT (Roche, 11383213001) and 3.75 µl BCIP (Roche, 11383221001) per 1 ml of NTMT. Staining was stopped by washing the embryos three times with NTMT, a 5 min wash with 1X PBST, and a 4°C overnight incubation in 1X PBST containing 4% paraformaldehyde. The embryos were then transitioned to methanol using the same four-step PBST:methanol washes listed above. Embryos were stored in −20°C or cleared in 2:1 Benzyl Benzoate:Benzyl Alcohol prior to imaging. Canada Balsam containing 10% methyl salicylate was used to mount cleared embryos on a slide. RNA in situ hybridization staining was visualized using a Leica DMRA2 microscope and images were acquired using a Leica DFC450 C camera. The following probes were used for the in situ hybridizations: gdf3 (Helde and Grunwald, 1993), nodal-related 2 (Rebagliati et al., 1998), lefty1 (Bisgrove et al., 1999), goosecoid (Stachel et al., 1993), floating head (Talbot et al., 1995), no tail (Schulte-Merker et al., 1992), axial (Strähle et al., 1993), sox17 (Alexander and Stainier, 1999), southpaw (Long et al., 2003), dand5 (Hashimoto et al., 2004), myl7 (Yelon et al., 1999).
Immunofluorescence
Immunofluorescence for Bmp signaling
Embryos were fixed at 75% epiboly stage in 4% PFA overnight at 4°C. These embryos were washed in PBST (1X PBS containing 0.1% Tween-20), transitioned to 100% methanol, and stored at −20°C for at least 1 day. The transition to methanol was done by performing five-minute washes in 75% PBST:25% methanol, 50% PBST:50% methanol, 25% PBST:75% methanol, and 100% methanol. The embryos were then transitioned into PBST by performing 5 min washes in 25% PBST:75% methanol, 50% PBST:50% methanol, 75% PBST:25% methanol, and 100% PBST. After three more five-minute washes in PBST, the embryos were washed once more in PBDT (PBST containing 1% DMSO) after which they were incubated in blocking solution of PBDT containing 10% normal goat serum (NGS) for 2 hr. Then, 1:100 anti-phosphorylatedSMAD1/5/8 antibody (Cell Signaling Technologies, 9511S) was added and incubated overnight at 4°C. The embryos were washed in PBDT containing 1% NGS and 0.1M NaCl for 1 min followed by six more 30 min washes at room temperature, after which the embryos underwent an overnight incubation in PBDT containing 10% NGS and 1:400 Donkey anti-Rabbit Alexa Fluor 647 secondary antibody (Invitrogen, A31573) at 4°C. After DAPI was added and the sample was rocked at room temperature for 20 min, the embryos were then washed in PBDT containing 1% NGS and 0.1M NaCl for one minute followed by five 30 min washes. The final PBDT wash was for 30 min.
Imaging
The embryos were positioned with the animal side toward the coverslip of a 30 mm dish (MatTek, P35G-1.5–14 C) in 1.5% low-melt agarose. All the WT and mutant embryos were imaged in one session on the Nikon A1 inverted confocal microscope using the same parameters.
Immunofluorescence for KV morphology
Immunofluorescence to examine KV morphology was performed with small changes to the protocol listed above for Bmp signaling. The embryos were blocked in PBDT containing 10% NSS (blocking solution). The first antibody incubation was performed in blocking solution containing 1:200 anti-PKCzeta (aPKC) (Santa Cruz Biotechnology, SC-216) primary antibody. The second antibody incubation was performed in PBDT containing 10% NSS and 1:400 Goat anti-Rabbit Alexa Fluor 647 (Invitrogen, A21246) secondary antibody. The subsequent washes were performed in PBDT containing 1% NSS and 0.1M NaCl. Analysis of KV morphology was performed in a Tg(sox17:GFP)s870 line (Sakaguchi et al., 2006).
Imaging
The embryos were positioned with the posterior end toward the coverslip (KV facing down) of a 30 mm dish (MatTek, P35G-1.5–14 C) in 1.5% low-melt agarose. All the uninjected and morpholino injected embryos were imaged in one session on the Nikon A1 using the same parameters.
Quantification
Quantification of phosphorylated SMAD1/5/8
The resulting z-stacks were opened in Imaris and positioned so that the D-V axis of the embryo was parallel to the left-right axis of the screen using the Free Rotate command. Snapshots in grayscale were taken of the DAPI channel and the Cy5 channel individually. For each embryo, a rectangle 200 pixels high and 650 pixels wide was taken so as to completely cover the middle of the embryo and the Plot Profile command was used to extract intensity of the DAPI and Cy5 channels along the D-V axis. A MATLAB script was used to align the intensity profiles and report the Cy5/DAPI ratio along the embryo from the ventral side (0) to the dorsal side (1). See provided source files for MATLAB script and data sourse files: Figure 3—source code 1, 2, 3.
Quantification of eve1 domain extension
For the WT and mutant images, a combination of red, green, and blue pixel values was chosen to threshold the set of images in MATLAB, so as to detect the purple in situ signal in an unbiased manner. Given an input image, the MATLAB script also produced a modified image with the in situ signal in white and the rest of image darkened, which was used for the next step. In ImageJ, an oval was manually traced out around the embryo and the extent of the circumference of embryo with in situ signal was manually identified using the Oval_Profile plugin with Number of Points as 360 and Analysis mode as Radial Sums. From the plot, the extent of degrees that the in situ stain occupied was measured and divided by 360 to get extent of the expression domain. See supplement for MATLAB script (Figure 3—source code 1, 2 and 3).
Acknowledgements
We are grateful for the exemplary collegiality of Alex Schier and Joe Yost in coordinating publications. We thank Victoria L Patterson and Daniel T Grimes for the constructive comments, helpful advice, and assistance in editing the manuscript. We thank Philip Johnson for zebrafish care, and Dr. Gary Laevsky and the Molecular Biology Confocal Microscopy Facility, which is a Nikon Center of Excellence, for microscopy support. GAJ acknowledges support from National Science Foundation Graduate Research Fellowship Grant DGE 1148900. This work was supported by funding awarded to RDB from the National Institute of Child Health and Development, grant R01HD048584. The content of this manuscript is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Funding Statement
The content of this manuscript is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health or National Science Foundation.
Contributor Information
Rebecca D Burdine, Email: rburdine@princeton.edu.
Deborah Yelon, University of California, San Diego, United States.
Funding Information
This paper was supported by the following grants:
Eunice Kennedy Shriver National Institute of Child Health and Human Development R01HD048584 to Rebecca D Burdine.
National Science Foundation Graduate Research Fellowship DGE 1148900 to Granton A Jindal.
Additional information
Competing interests
No competing interests declared.
Author contributions
Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Writing—original draft, Writing—review and editing.
Formal analysis, Validation, Investigation, Visualization, Methodology, Writing—review and editing.
Conceptualization, Supervision, Funding acquisition, Writing—original draft, Project administration, Writing—review and editing.
Ethics
Animal experimentation: Zebrafish husbandry was performed in strict accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals. All protocols are approved (Burdine #1915-16) by Princeton's Institutional Animal Care and Use Committee (IACUC).
Additional files
References
- Aamar E, Dawid IB. Sox17 and chordin are required for formation of Kupffer's vesicle and left-right asymmetry determination in zebrafish. Developmental Dynamics. 2010;239:2980–2988. doi: 10.1002/dvdy.22431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alexander J, Stainier DY. A molecular pathway leading to endoderm formation in zebrafish. Current Biology. 1999;9:1147–1157. doi: 10.1016/S0960-9822(00)80016-0. [DOI] [PubMed] [Google Scholar]
- Amack JD, Wang X, Yost HJ. Two T-box genes play independent and cooperative roles to regulate morphogenesis of ciliated Kupffer's vesicle in zebrafish. Developmental Biology. 2007;310:196–210. doi: 10.1016/j.ydbio.2007.05.039. [DOI] [PubMed] [Google Scholar]
- Andersson O, Bertolino P, Ibáñez CF. Distinct and cooperative roles of mammalian Vg1 homologs GDF1 and GDF3 during early embryonic development. Developmental Biology. 2007;311:500–511. doi: 10.1016/j.ydbio.2007.08.060. [DOI] [PubMed] [Google Scholar]
- Andersson O, Reissmann E, Jörnvall H, Ibáñez CF. Synergistic interaction between Gdf1 and Nodal during anterior axis development. Developmental Biology. 2006;293:370–381. doi: 10.1016/j.ydbio.2006.02.002. [DOI] [PubMed] [Google Scholar]
- Baker K, Holtzman NG, Burdine RD. Direct and indirect roles for Nodal signaling in two axis conversions during asymmetric morphogenesis of the zebrafish heart. PNAS. 2008;105:13924–13929. doi: 10.1073/pnas.0802159105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Birsoy B, Kofron M, Schaible K, Wylie C, Heasman J. Vg 1 is an essential signaling molecule in Xenopus development. Development. 2006;133:15–20. doi: 10.1242/dev.02144. [DOI] [PubMed] [Google Scholar]
- Bisgrove BW, Essner JJ, Yost HJ. Regulation of midline development by antagonism of lefty and nodal signaling. Development. 1999;126:3253–3262. doi: 10.1242/dev.126.14.3253. [DOI] [PubMed] [Google Scholar]
- Burdine RD, Grimes DT. Antagonistic interactions in the zebrafish midline prior to the emergence of asymmetric gene expression are important for left-right patterning. Philosophical Transactions of the Royal Society B: Biological Sciences. 2016;371:20150402. doi: 10.1098/rstb.2015.0402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen C, Ware SM, Sato A, Houston-Hawkins DE, Habas R, Matzuk MM, Shen MM, Brown CW. The Vg1-related protein Gdf3 acts in a Nodal signaling pathway in the pre-gastrulation mouse embryo. Development. 2006;133:319–329. doi: 10.1242/dev.02210. [DOI] [PubMed] [Google Scholar]
- Cheng SK, Olale F, Bennett JT, Brivanlou AH, Schier AF. EGF-CFC proteins are essential coreceptors for the TGF-beta signals Vg1 and GDF1. Genes & Development. 2003;17:31–36. doi: 10.1101/gad.1041203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Collignon J, Varlet I, Robertson EJ. Relationship between asymmetric nodal expression and the direction of embryonic turning. Nature. 1996;381:155–158. doi: 10.1038/381155a0. [DOI] [PubMed] [Google Scholar]
- Compagnon J, Barone V, Rajshekar S, Kottmeier R, Pranjic-Ferscha K, Behrndt M, Heisenberg CP. The notochord breaks bilateral symmetry by controlling cell shapes in the zebrafish laterality organ. Developmental Cell. 2014;31:774–783. doi: 10.1016/j.devcel.2014.11.003. [DOI] [PubMed] [Google Scholar]
- Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339:819–823. doi: 10.1126/science.1231143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Conlon FL, Lyons KM, Takaesu N, Barth KS, Kispert A, Herrmann B, Robertson EJ. A primary requirement for nodal in the formation and maintenance of the primitive streak in the mouse. Development. 1994;120:1919–1928. doi: 10.1242/dev.120.7.1919. [DOI] [PubMed] [Google Scholar]
- de Campos-Baptista MI, Holtzman NG, Yelon D, Schier AF. Nodal signaling promotes the speed and directional movement of cardiomyocytes in zebrafish. Developmental Dynamics. 2008;237:3624–3633. doi: 10.1002/dvdy.21777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Robertis EM. Spemann's organizer and the self-regulation of embryonic fields. Mechanisms of Development. 2009;126:925–941. doi: 10.1016/j.mod.2009.08.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dohrmann CE, Kessler DS, Melton DA. Induction of axial mesoderm by zDVR-1, the zebrafish orthologue of Xenopus Vg1. Developmental Biology. 1996;175:108–117. doi: 10.1006/dbio.1996.0099. [DOI] [PubMed] [Google Scholar]
- Dubrulle J, Jordan BM, Akhmetova L, Farrell JA, Kim S-H, Solnica-Krezel L, Schier AF. Response to Nodal morphogen gradient is determined by the kinetics of target gene induction. eLife. 2015;4:e05042. doi: 10.7554/eLife.05042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fauny JD, Thisse B, Thisse C. The entire zebrafish blastula-gastrula margin acts as an organizer dependent on the ratio of Nodal to BMP activity. Development. 2009;136:3811–3819. doi: 10.1242/dev.039693. [DOI] [PubMed] [Google Scholar]
- Feldman B, Gates MA, Egan ES, Dougan ST, Rennebeck G, Sirotkin HI, Schier AF, Talbot WS. Zebrafish organizer development and germ-layer formation require nodal-related signals. Nature. 1998;395:181–185. doi: 10.1038/26013. [DOI] [PubMed] [Google Scholar]
- Feldman B, Stemple DL. Morpholino phenocopies of sqt, oep, and ntl mutations. Genesis. 2001;30:175–177. doi: 10.1002/gene.1058. [DOI] [PubMed] [Google Scholar]
- Fuerer C, Nostro MC, Constam DB. Nodal·Gdf1 heterodimers with bound prodomains enable serum-independent nodal signaling and endoderm differentiation. Journal of Biological Chemistry. 2014;289:17854–17871. doi: 10.1074/jbc.M114.550301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grimes DT, Burdine RD. Left-Right Patterning: Breaking Symmetry to Asymmetric Morphogenesis. Trends in Genetics. 2017;33:616–628. doi: 10.1016/j.tig.2017.06.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gritsman K, Talbot WS, Schier AF. Nodal signaling patterns the organizer. Development. 2000;127:921–932. doi: 10.1242/dev.127.5.921. [DOI] [PubMed] [Google Scholar]
- Gritsman K, Zhang J, Cheng S, Heckscher E, Talbot WS, Schier AF. The EGF-CFC protein one-eyed pinhead is essential for nodal signaling. Cell. 1999;97:121–132. doi: 10.1016/S0092-8674(00)80720-5. [DOI] [PubMed] [Google Scholar]
- Hanafusa H, Masuyama N, Kusakabe M, Shibuya H, Nishida E. The TGF-beta family member derrière is involved in regulation of the establishment of left-right asymmetry. EMBO Reports. 2000;1:32–39. doi: 10.1093/embo-reports/kvd008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hashimoto H, Rebagliati M, Ahmad N, Muraoka O, Kurokawa T, Hibi M, Suzuki T. The Cerberus/Dan-family protein Charon is a negative regulator of Nodal signaling during left-right patterning in zebrafish. Development. 2004;131:1741–1753. doi: 10.1242/dev.01070. [DOI] [PubMed] [Google Scholar]
- Hata A, Chen YG. TGF-β signaling from receptors to smads. Cold Spring Harbor Perspectives in Biology. 2016;8:a022061. doi: 10.1101/cshperspect.a022061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Helde KA, Grunwald DJ. The DVR-1 (Vg1) transcript of zebrafish is maternally supplied and distributed throughout the embryo. Developmental Biology. 1993;159:418–426. doi: 10.1006/dbio.1993.1252. [DOI] [PubMed] [Google Scholar]
- Hojo M, Takashima S, Kobayashi D, Sumeragi A, Shimada A, Tsukahara T, Yokoi H, Narita T, Jindo T, Kage T, Kitagawa T, Kimura T, Sekimizu K, Miyake A, Setiamarga D, Murakami R, Tsuda S, Ooki S, Kakihara K, Naruse K, Takeda H. Right-elevated expression of charon is regulated by fluid flow in medaka Kupffer's vesicle. Development, Growth & Differentiation. 2007;49:395–405. doi: 10.1111/j.1440-169X.2007.00937.x. [DOI] [PubMed] [Google Scholar]
- Hyatt BA, Lohr JL, Yost HJ. Initiation of vertebrate left-right axis formation by maternal Vg1. Nature. 1996;384:62–65. doi: 10.1038/384062a0. [DOI] [PubMed] [Google Scholar]
- Iannaccone PM, Zhou X, Khokha M, Boucher D, Kuehn MR. Insertional mutation of a gene involved in growth regulation of the early mouse embryo. Developmental Dynamics. 1992;194:198–208. doi: 10.1002/aja.1001940305. [DOI] [PubMed] [Google Scholar]
- Jao LE, Wente SR, Chen W. Efficient multiplex biallelic zebrafish genome editing using a CRISPR nuclease system. PNAS. 2013;110:13904–13909. doi: 10.1073/pnas.1308335110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Joly JS, Joly C, Schulte-Merker S, Boulekbache H, Condamine H. The ventral and posterior expression of the zebrafish homeobox gene eve1 is perturbed in dorsalized and mutant embryos. Development. 1993;119:1261–1275. doi: 10.1242/dev.119.4.1261. [DOI] [PubMed] [Google Scholar]
- Jones CM, Kuehn MR, Hogan BL, Smith JC, Wright CV. Nodal-related signals induce axial mesoderm and dorsalize mesoderm during gastrulation. Development. 1995;121:3651–3662. doi: 10.1242/dev.121.11.3651. [DOI] [PubMed] [Google Scholar]
- Karlen S, Rebagliati M. A morpholino phenocopy of the cyclops mutation. Genesis. 2001;30:126–128. doi: 10.1002/gene.1046. [DOI] [PubMed] [Google Scholar]
- Kessler DS, Melton DA. Induction of dorsal mesoderm by soluble, mature Vg1 protein. Development. 1995;121:2155–2164. doi: 10.1242/dev.121.7.2155. [DOI] [PubMed] [Google Scholar]
- Prabhat. Kreiling JA, Williams G, Creton R. Analysis of Kupffer's vesicle in zebrafish embryos using a cave automated virtual environment. Developmental Dynamics. 2007;236:1963–1969. doi: 10.1002/dvdy.21191. [DOI] [PubMed] [Google Scholar]
- Langdon YG, Mullins MC. Maternal and zygotic control of zebrafish dorsoventral axial patterning. Annual Review of Genetics. 2011;45:357–377. doi: 10.1146/annurev-genet-110410-132517. [DOI] [PubMed] [Google Scholar]
- Lenhart KF, Lin SY, Titus TA, Postlethwait JH, Burdine RD. Two additional midline barriers function with midline lefty1 expression to maintain asymmetric Nodal signaling during left-right axis specification in zebrafish. Development. 2011;138:4405–4410. doi: 10.1242/dev.071092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levin M, Johnson RL, Stern CD, Kuehn M, Tabin C. A molecular pathway determining left-right asymmetry in chick embryogenesis. Cell. 1995;82:803–814. doi: 10.1016/0092-8674(95)90477-8. [DOI] [PubMed] [Google Scholar]
- Levine AJ, Brivanlou AH. GDF3, a BMP inhibitor, regulates cell fate in stem cells and early embryos. Development. 2006;133:209–216. doi: 10.1242/dev.02192. [DOI] [PubMed] [Google Scholar]
- Little SC, Mullins MC. Bone morphogenetic protein heterodimers assemble heteromeric type I receptor complexes to pattern the dorsoventral axis. Nature Cell Biology. 2009;11:637–643. doi: 10.1038/ncb1870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Long S, Ahmad N, Rebagliati M. The zebrafish nodal-related gene southpaw is required for visceral and diencephalic left-right asymmetry. Development. 2003;130:2303–2316. doi: 10.1242/dev.00436. [DOI] [PubMed] [Google Scholar]
- Lopes SS, Lourenço R, Pacheco L, Moreno N, Kreiling J, Saúde L. Notch signalling regulates left-right asymmetry through ciliary length control. Development. 2010;137:3625–3632. doi: 10.1242/dev.054452. [DOI] [PubMed] [Google Scholar]
- Lowe LA, Supp DM, Sampath K, Yokoyama T, Wright CV, Potter SS, Overbeek P, Kuehn MR. Conserved left-right asymmetry of nodal expression and alterations in murine situs inversus. Nature. 1996;381:158–161. doi: 10.1038/381158a0. [DOI] [PubMed] [Google Scholar]
- Montague TG, Cruz JM, Gagnon JA, Church GM, Valen E. CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Research. 2014;42:W401–W407. doi: 10.1093/nar/gku410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morikawa M, Derynck R, Miyazono K. TGF-β and the TGF-β Family: Context-Dependent Roles in Cell and Tissue Physiology. Cold Spring Harbor Perspectives in Biology. 2016;8:a021873. doi: 10.1101/cshperspect.a021873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nasevicius A, Ekker SC. Effective targeted gene 'knockdown' in zebrafish. Nature Genetics. 2000;26:216–220. doi: 10.1038/79951. [DOI] [PubMed] [Google Scholar]
- Odenthal J, Haffter P, Vogelsang E, Brand M, van Eeden FJ, Furutani-Seiki M, Granato M, Hammerschmidt M, Heisenberg CP, Jiang YJ, Kane DA, Kelsh RN, Mullins MC, Warga RM, Allende ML, Weinberg ES, Nüsslein-Volhard C. Mutations affecting the formation of the notochord in the zebrafish, Danio rerio. Development. 1996;123:103–115. doi: 10.1242/dev.123.1.103. [DOI] [PubMed] [Google Scholar]
- Okabe N, Xu B, Burdine RD. Fluid dynamics in zebrafish Kupffer's vesicle. Developmental Dynamics. 2008;237:3602–3612. doi: 10.1002/dvdy.21730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peterson AG, Wang X, Yost HJ. Dvr1 transfers left-right asymmetric signals from Kupffer's vesicle to lateral plate mesoderm in zebrafish. Developmental Biology. 2013;382:198–208. doi: 10.1016/j.ydbio.2013.06.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rankin CT, Bunton T, Lawler AM, Lee SJ. Regulation of left-right patterning in mice by growth/differentiation factor-1. Nature Genetics. 2000;24:262–265. doi: 10.1038/73472. [DOI] [PubMed] [Google Scholar]
- Rebagliati MR, Toyama R, Haffter P, Dawid IB. cyclops encodes a nodal-related factor involved in midline signaling. PNAS. 1998;95:9932–9937. doi: 10.1073/pnas.95.17.9932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rebagliati MR, Weeks DL, Harvey RP, Melton DA. Identification and cloning of localized maternal RNAs from Xenopus eggs. Cell. 1985;42:769–777. doi: 10.1016/0092-8674(85)90273-9. [DOI] [PubMed] [Google Scholar]
- Sakaguchi T, Kikuchi Y, Kuroiwa A, Takeda H, Stainier DY. The yolk syncytial layer regulates myocardial migration by influencing extracellular matrix assembly in zebrafish. Development. 2006;133:4063–4072. doi: 10.1242/dev.02581. [DOI] [PubMed] [Google Scholar]
- Sako K, Pradhan SJ, Barone V, Inglés-Prieto Á, Müller P, Ruprecht V, Čapek D, Galande S, Janovjak H, Heisenberg CP. Optogenetic control of Nodal signaling reveals a temporal pattern of Nodal signaling regulating cell fate specification during gastrulation. Cell Reports. 2016;16:866–877. doi: 10.1016/j.celrep.2016.06.036. [DOI] [PubMed] [Google Scholar]
- Sampath K, Cheng AM, Frisch A, Wright CV. Functional differences among Xenopus nodal-related genes in left-right axis determination. Development. 1997;124:3293–3302. doi: 10.1242/dev.124.17.3293. [DOI] [PubMed] [Google Scholar]
- Schmid B, Fürthauer M, Connors SA, Trout J, Thisse B, Thisse C, Mullins MC. Equivalent genetic roles for bmp7/snailhouse and bmp2b/swirl in dorsoventral pattern formation. Development. 2000;127:957–967. doi: 10.1242/dev.127.5.957. [DOI] [PubMed] [Google Scholar]
- Schulte-Merker S, Ho RK, Herrmann BG, Nüsslein-Volhard C. The protein product of the zebrafish homologue of the mouse T gene is expressed in nuclei of the germ ring and the notochord of the early embryo. Development. 1992;116:1021–1032. doi: 10.1242/dev.116.4.1021. [DOI] [PubMed] [Google Scholar]
- Schweickert A, Vick P, Getwan M, Weber T, Schneider I, Eberhardt M, Beyer T, Pachur A, Blum M. The nodal inhibitor Coco is a critical target of leftward flow in Xenopus. Current Biology. 2010;20:738–743. doi: 10.1016/j.cub.2010.02.061. [DOI] [PubMed] [Google Scholar]
- Slagle CE, Aoki T, Burdine RD. Nodal-dependent mesendoderm specification requires the combinatorial activities of FoxH1 and Eomesodermin. PLoS Genetics. 2011;7:e1002072. doi: 10.1371/journal.pgen.1002072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith KA, Chocron S, von der Hardt S, de Pater E, Soufan A, Bussmann J, Schulte-Merker S, Hammerschmidt M, Bakkers J. Rotation and asymmetric development of the zebrafish heart requires directed migration of cardiac progenitor cells. Developmental Cell. 2008;14:287–297. doi: 10.1016/j.devcel.2007.11.015. [DOI] [PubMed] [Google Scholar]
- Stachel SE, Grunwald DJ, Myers PZ. Lithium perturbation and goosecoid expression identify a dorsal specification pathway in the pregastrula zebrafish. Development. 1993;117:1261–1274. doi: 10.1242/dev.117.4.1261. [DOI] [PubMed] [Google Scholar]
- Strähle U, Blader P, Henrique D, Ingham PW. Axial, a zebrafish gene expressed along the developing body axis, shows altered expression in cyclops mutant embryos. Genes & Development. 1993;7:1436–1446. doi: 10.1101/gad.7.7b.1436. [DOI] [PubMed] [Google Scholar]
- Supatto W, Fraser SE, Vermot J. An all-optical approach for probing microscopic flows in living embryos. Biophysical Journal. 2008;95:L29–L31. doi: 10.1529/biophysj.108.137786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Talbot WS, Trevarrow B, Halpern ME, Melby AE, Farr G, Postlethwait JH, Jowett T, Kimmel CB, Kimelman D. A homeobox gene essential for zebrafish notochord development. Nature. 1995;378:150–157. doi: 10.1038/378150a0. [DOI] [PubMed] [Google Scholar]
- Tanaka C, Sakuma R, Nakamura T, Hamada H, Saijoh Y. Long-range action of Nodal requires interaction with GDF1. Genes & Development. 2007;21:3272–3282. doi: 10.1101/gad.1623907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thisse B, Thisse C. Formation of the vertebrate embryo: Moving beyond the Spemann organizer. Seminars in Cell & Developmental Biology. 2015;42:94–102. doi: 10.1016/j.semcdb.2015.05.007. [DOI] [PubMed] [Google Scholar]
- Thomsen GH, Melton DA. Processed Vg1 protein is an axial mesoderm inducer in Xenopus. Cell. 1993;74:433–441. doi: 10.1016/0092-8674(93)80045-G. [DOI] [PubMed] [Google Scholar]
- van Boxtel AL, Chesebro JE, Heliot C, Ramel MC, Stone RK, Hill CS. A temporal window for signal activation dictates the dimensions of a Nodal signaling domain. Developmental Cell. 2015;35:175–185. doi: 10.1016/j.devcel.2015.09.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wall NA, Craig EJ, Labosky PA, Kessler DS. Mesendoderm induction and reversal of left-right pattern by mouse Gdf1, a Vg1-related gene. Developmental Biology. 2000;227:495–509. doi: 10.1006/dbio.2000.9926. [DOI] [PubMed] [Google Scholar]
- Wang G, Manning ML, Amack JD. Regional cell shape changes control form and function of Kupffer's vesicle in the zebrafish embryo. Developmental Biology. 2012;370:52–62. doi: 10.1016/j.ydbio.2012.07.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y, Wang X, Wohland T, Sampath K. Extracellular interactions and ligand degradation shape the nodal morphogen gradient. eLife. 2016;5:e13879. doi: 10.7554/eLife.13879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu PF, Houssin N, Ferri-Lagneau KF, Thisse B, Thisse C. Construction of a vertebrate embryo from two opposing morphogen gradients. Science. 2014;344:87–89. doi: 10.1126/science.1248252. [DOI] [PubMed] [Google Scholar]
- Yan YT, Gritsman K, Ding J, Burdine RD, Corrales JD, Price SM, Talbot WS, Schier AF, Shen MM. Conserved requirement for EGF-CFC genes in vertebrate left-right axis formation. Genes & Development. 1999;13:2527–2537. doi: 10.1101/gad.13.19.2527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ye M, Berry-Wynne KM, Asai-Coakwell M, Sundaresan P, Footz T, French CR, Abitbol M, Fleisch VC, Corbett N, Allison WT, Drummond G, Walter MA, Underhill TM, Waskiewicz AJ, Lehmann OJ. Mutation of the bone morphogenetic protein GDF3 causes ocular and skeletal anomalies. Human Molecular Genetics. 2010;19:287–298. doi: 10.1093/hmg/ddp496. [DOI] [PubMed] [Google Scholar]
- Yelon D, Horne SA, Stainier DY. Restricted expression of cardiac myosin genes reveals regulated aspects of heart tube assembly in zebrafish. Developmental Biology. 1999;214:23–37. doi: 10.1006/dbio.1999.9406. [DOI] [PubMed] [Google Scholar]