Abstract
Oligomeric assemblies of the amyloid β-protein, Aβ, are thought to be the proximate neurotoxic agents in Alzheimer’s disease (AD). Oligomer formation is a complex process that produces a polydisperse population of metastable structures. For this reason, formal structure–activity correlations, both in vitro and in vivo, have been difficult to accomplish. An analytical solution to this problem was provided by the application of a photochemical cross-linking method to the Aβ assembly system. This method, photo-induced cross-linking of unmodified proteins (PICUP), enabled the quantitative determination of the oligomer size distribution. We report here the integration of PICUP with SDS-PAGE and alkaline extraction procedures to create a method for the isolation of pure populations of oligomers of defined order. This method has been used successfully to provide material for formal structure–activity studies of Aβ oligomers.
Keywords: Amyloid β-protein, Oligomers, PICUP, Purification
1. Introduction
Alzheimer’s disease (AD) is a neurodegenerative disorder characterized by the formation of extracellular amyloid deposits in the brain parenchyma and vasculature and within neuronal cells (1). These deposits are composed of the amyloid β-protein, Aβ, and the microtubule-associated protein, tau, respectively (1, 2). An important current working hypothesis of AD causation posits that Aβ oligomers are the proximate pathologic agents (3–5). In vivo and in vitro studies have revealed a diversity of such assemblies (6), including dimers (7), Aβ*56 (8), Aβ-derived diffusible ligands (ADDLs) (9), paranuclei (10), protofibrils (11, 12), globulomers (13), and amylospheroids (14). To establish how each of these assemblies is involved in disease causation, structure–activity correlations must be established. However, achievement of this goal has been difficult due to the complexity of Aβ assembly, the metastability of Aβ oligomers, and the polydispersity of the oligomer population (6, 15).
Bitan et al. (16) applied the method of photo-induced cross-linking of unmodified proteins (PICUP) (17, 18) to “freeze” the oligomer equilibria and allow analytical studies of Aβ oligomerization. PICUP is a highly efficient, zero-length cross-linking method that can be applied to native (no pre facto protein modification is required) Aβ populations. Following cross-linking, monomer interchange among cross-linked oligomer species does not occur because the monomers are covalently bound to each other. This eliminates the metastability problem discussed above and allows quantitative determination of the polydispersity of the population. Bitan et al. observed that the shorter isoform of Aβ, Aβ40, and the longer isoform, Aβ42, each produced a distinct oligomer distribution when studied by SDS-PAGE (10). This distinction is correlated with the strong disease linkage of Aβ42.
The successful application of PICUP to the problem of quantitatively determining the Aβ oligomer size distribution suggested that PICUP might be incorporated into a protocol for the production of Aβ oligomers of defined order. Initial work in this area focused on coupling PICUP with size exclusion chromatography (SEC). SEC is a useful method for the separation of soluble proteins on the basis of Stokes radius. SEC has the advantage of being carried out in the solution phase, which results in sample fractions that can be used immediately in other experiments. Unfortunately, preliminary experiments showed that the molecular weight resolution of SEC was insufficient to produce pure populations of oligomers larger than dimers (unpublished results). For this reason, we attempted to combine PICUP with SDS-PAGE, which has very high molecular weight resolution. To do so, an efficient method of extraction of the oligomer populations from the gel matrix had to be developed. This goal was achieved through modification of an alkaline extraction protocol originally reported by Jin and Manabe (19). The ability to produce pure populations of oligomers of specific order enabled formal structure–activity studies of Aβ40 oligomers (20). We communicate here details of the method used in this study.
2. Materials
All solutions were prepared using water provided by a Milli-Q system (18 MΩ/cm, Millipore Corp., Bedford, MA). All reagents were of the highest purity available and were purchased from Sigma-Aldrich, unless otherwise noted.
2.1. PICUP
Ammonium persulfate (APS): 60 mM solution in water (13.7 mg/ml). Vortex until the APS has dissolved, place on ice, and then use immediately thereafter.
Tris (2,2′-bipyridyl)dichlororuthenium (II) hexahydrate (Ru (Bpy)): 3 mM solution in water (2.24 mg/ml). Vortex the solution until the solid is dissolved and then place on ice, wrapped in aluminum foil, to protect from light. Use immediately.
Sodium hydroxide (NaOH): 60 mM (2.4 g/l) in water, pH 11.
Sodium phosphate, dibasic buffer (Na2HPO4): 20 mM (2.84 g/l) in water, pH 7.4. Prepare and store at room temperature.
β-Mercaptoethanol: 5% (v/v) solution in 2× sample buffer (Cat. Num. LC1676, Invitrogen, Carlsbad, CA).
200-W incandescent lamp: model 170-D (Dolan-Jenner, Lawrence, MA) (see Note 1, Fig. 1).
Sonicator (model 1510R-DTH; Branson Ultrasonics).
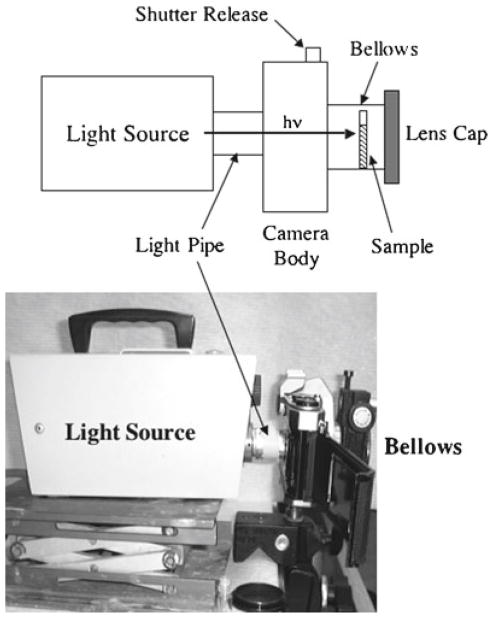
Fig. 1.
The irradiation system. A light source is linked to the shutter side (back ) of a 35-mm SLR camera body through a cylindrical tube (light pipe ). The sample is placed within a bellows attached to the lens opening, after which a lens cap closes the open end of the bellows. Irradiation then occurs by actuation of the shutter release, with the time adjusted using the shutter speed control of the camera.
2.2. SDS-PAGE and Gel Staining
10–20% Tricine gels, 1-mm thick, 10 wells (Cat. Num. EC6625BOX, Invitrogen, Carlsbad, CA) (Note 2). Store at 4°C.
Tricine SDS Running Buffer (10×) (Cat. Num. LC1675, Invitrogen, Carlsbad, CA). Dilute 50 ml of 10× buffer in 450 ml water. Mix thoroughly. Store at room temperature.
Mark12 unstained standard: protein standard (Cat. Num. LC5677, Invitrogen, Carlsbad, CA). 2.5–200 kDa standard. Store at 4°C.
Coomassie Blue: SimplyBlue™ (Cat. Num. LC6060, Invitrogen, Carlsbad, CA).
Orbital shaker, model ZD-9556 (Madell Technology Corporation, Ontario, CA).
2.3. Alkaline Extraction
1.5-ml Disposable pellet pestle (Cat. Num. K749521-1590, Fisher Scientific, Rockford, IL).
Ammonium hydroxide (NH4OH): 0.1 M (0.35% w/v) in water.
Rotator (Mini LabRoller, Labnet International, Inc., Woodbridge, NJ) (8.4″×4″×5″, 20–24 rpm).
2.4. Purification
SDS Removal: SDSOut (Pierce, Rockford, IL).
Dialysis membrane: Spectra/Por Biotech CE Dialysis Membranes; 2,000 MWCO (molecular weight cutoff); 7.5-mm diameter (Spectrum Laboratories, Rancho Dominguez, CA).
Urea: 10 M (60% w/v) in water.
Silver staining kit: SilverXpress(Cat. Num. LC6100, Invitrogen, Carlsbad, CA).
3. Methods
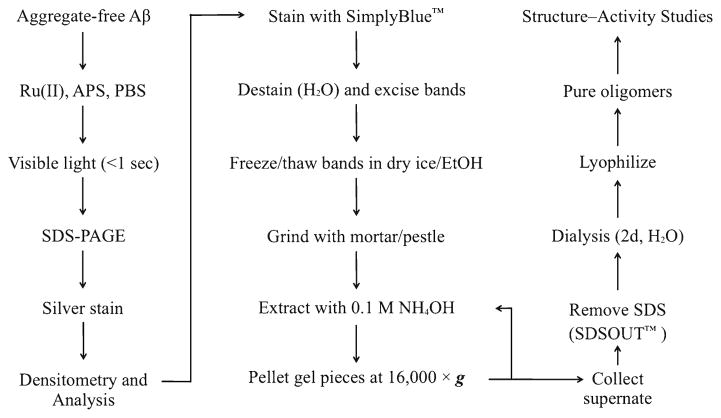
The method described involves the production of multiple samples of cross-linked peptide, followed by the pooling of the samples, their fractionation by SDS-PAGE, and their extraction from the resultant gel (Fig. 2).
Fig. 2.
Outline of the method.
3.1. PICUP
Dissolve Aβ40 (see Note 3) to a concentration of 90 μM in a solution of 6 mM NaOH/9 mM Na2HPO4. First, dissolve 195 μg of the peptide in 50 μl of 60 mM NaOH gently agitating the tube to aid in dissolution. Add 225 μl water, continuing to gently agitate, until the solution is clear. Finally, add 225 μl of 20 mM Na2HPO4, pH 7.4 (see Notes 4 and 5).
Sonicate the Aβ40 solution for 1 min.
Centrifuge the solution at 16,000 × g for 10 min, then aliquot the supernate into 0.2-ml clear, thin-walled PCR tubes (18 μl/tube).
Add 1 μl of APS and 1 μl Ru(Bpy) into one reaction tube. Place the tube into the camera bellows in the irradiation system (Fig. 2), cap the end, and then irradiate the tube for 1 s. Immediately quench the reaction with 20 μl of β-mercaptoethanol/sample buffer.
Multiple reactions may be done to produce larger amounts of cross-linked peptide (see Note 6). These reactions may be pooled and stored at −20°C if subsequent experiments are not to be done immediately (see Note 7).
3.2. SDS-PAGE
Prepare the gel for electrophoresis by creating a two-lane gel, one lane being an outermost lane and the other being the remaining lanes in the gel from which you have removed the intervening polyacrylamide teeth between the sample wells. This can be done using a small scalpel. Wash the two sample wells with running buffer and assemble the gel system.
Heat the cross-linked oligomers at 100°C for 10 min. This can be done in a boiling-water bath or a heating block. Centrifuge the pooled oligomers briefly in a microcentrifuge (16,000 × g).
Load molecular weight markers (10 μl) in the small well and the supernate of the cross-linked oligomers in the large well (up to 400 μl).
Electrophorese at 100 V, constant voltage, until the sample has stacked at the interface between the stacking and separating gels. Electrophorese the samples at 120 V, constant voltage, until the dye front has reached the bottom of the gel.
After electrophoresis, open the gel cartridge by prying it open with a spatula, knife, or other thin, flat implement. Carefully detach the gel from the bottom plate of the cartridge into a staining tray filled with water.
Add enough water to cover the gel by a few centimeters and gently agitate on an orbital shaker for 5 min. Discard the water and replace it with fresh water. Repeat wash three times.
Pour out the last water wash and add sufficient SimplyBlue™ stain solution to cover the gel. Place the gel on the orbital shaker for 1 h. After staining, pour out the staining solution and replace it with water. Place the gel on the orbital shaker for 1 h to destain the gel and to reveal the protein bands.
3.3. Alkaline Extraction
Place the stained gel on a glass plate (scrupulously cleaned with soap and water, then water, and finally with methanol or ethanol) and excise the oligomer bands with a scalpel or razor blade (see Note 8).
Dice each band into small (1 mm) cubes and place the cubes into a 1.5-ml microcentrifuge tube (see Note 9). Wash the gel cubes with 1 ml of water three times. Briefly centrifuge (16,000×g) and remove the supernatant water each time.
Pre-heat a water bath to 70°C and prepare a small vessel (of geometry appropriate for immersing 1.5-ml microcentrifuge tubes) containing dry ice and ethanol. Subject the gel cubes to three cycles of rapid freezing and thawing by alternately placing the tubes in the water and dry ice/ethanol baths. The cubes will become brittle during this process, after which they are crushed into a homogeneous state using a 1.5-ml pellet pestle.
Extract the crushed gel pieces in 1 ml of 0.1 M NH4OH for 10 min at room temperature while rotating at 24 rpm. Centrifuge at 16,000×g for 5 min and collect the supernate. Repeat the extraction, rotation, and centrifugation twice more.
3.4. Purification
Pool the final supernates of the extracted gel pieces and treat with SDS-Out™ to remove SDS. Transfer the protein sample to one of the microcentrifuge tubes provided with the kit. Add the SDS-Out Precipitation Reagent to the protein sample (1:20, v/v) and vortex to mix. Incubate on ice for 20 min, then centrifuge at 16,000 × g for 10 min. Transfer up to 500 μl of the supernate to one of the spin cup columns provided with the kit and centrifuge at 16,000 × g for 1 min.
Prepare dialysis membrane by cutting into short lengths (suitable for up to 1 ml of sample) and soaking in enough water to fully immerse the membrane for 30 min at room temperature (this removes sodium azide and other chemical contaminants). Without letting the membrane dry out, fold one end of the tubing back onto itself and clamp the tubing closed across the double-thickness region using a dialysis tube clamp. Introduce up to 1 ml of the SDS-Out™-treated mixture into each piece of dialysis tubing and then clamp this end as specified above. When this end of the membrane is folded, ensure that as little air as possible is left inside the bag.
Dialyze the 1 ml samples in dialysis membranes against 5 L of 10 M urea at 4°C for 12 h. Remove the urea by dialysis twice with 5 L of water for 12 h at 4°C each time.
After dialysis, open one end of the bag, collect the solution, and lyophilize. Store the lyophilizate at −20°C (see Note 10).
Lyophilizates can be reconstituted in a variety of solvents. We routinely evaluate protein amount and purity by reconstitution of the lyophilizate in 200 μl of 50 mM sodium bicarbonate, after which 5 μl of this solution is mixed with 5 μl of 2× sample buffer/5% 2-mercaptoethanol, boiled for 10 min, and then analyzed by SDS-PAGE and silver staining (Fig. 3).
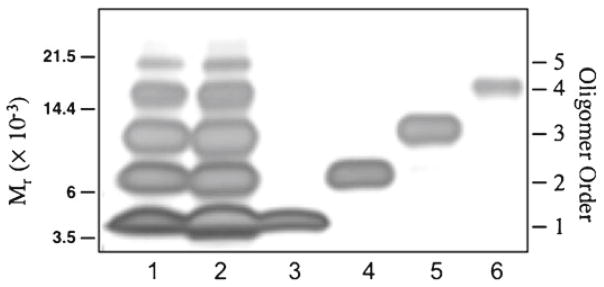
Fig. 3.
Evaluation of the method by SDS-PAGE and silver staining. Lanes: 1, cross-linked Aβ 40 prior to purification; 2, cross-linked Aβ 40, for which the entire lane from the first Coomassie-stained gel was subjected to the purification procedure; 3, final product from the monomer band of the first gel; 4, final product from the dimer band; 5, final product from the trimer band; and 6, final product from the tetramer band.
Acknowledgments
We gratefully acknowledge the support of NIH grants AG027818, NS038328, the Jim Easton Consortium for Alzheimer’s Drug Discovery and Biomarkers at UCLA, and the California Department of Public Health, Alzheimer’s Disease Program, grant #07-65806.
Footnotes
The camera body/bellows system is a convenient means of precisely irradiating samples for chosen amounts of time. Any system that can accomplish the same thing can be used. The incandescent lamp provides visible light to photooxidize the RuII in the Ru(Bpy) complex. The critical considerations here are the wavelength distribution of the light source and the photon flux. Adjustments to these parameters generally are not possible, but the modification of irradiation time is a simple and effective method for optimizing cross-linking efficiency and minimizing radical damage to the protein to be cross-linked (see (16) for a complete discussion of these points).
10–20% Gradient gels also can be prepared manually. We have chosen to purchase pre-cast gels for convenience.
The Aβ used in our experiments is synthesized “in house,” as described (20). The method of synthesis and the source of Aβ is not critical, as long as the peptide is chemically pure. Our peptides are characterized by HPLC, amino acid analysis, and mass spectrometry. Peptide purity is >90%, as determined by HPLC. The protein content of our peptide lyophilizates generally is >85%.
The calculation used here should provide enough material to run two gels of cross-linked material, but it can be scaled up or down to fit the needs of the user. The volume of NaOH used should be 10% of the final solution volume. Add water to 55% of the final volume. Add the final 45% solution volume as 20 mM Na2HPO4, pH 7.4.
Any remaining unsolubilized peptide should be solubilized following addition of the water. After addition of the phosphate buffer, the pH should change to 7.6. Any unsolubilized peptide should be removed at this point by centrifugation for 15 min at 16,000 × g at room temperature.
It is important to do one reaction at a time. Do not add the APS and Ru(Bpy) reagents to all the tubes and then irradiate them one at a time. Only add reagents to a tube once the irradiation and quenching of the prior tube has been completed.
The −20°C freezer should not have an auto-defrost function because this function produces freeze–thaw cycles that can affect peptide structure.
Place a plain white sheet of paper under the glass plate to enhance contrast and make the oligomer bands easier to distinguish.
Diced gel pieces can be stored at −20°C for several weeks prior to use.
Oligomers should be stable indefinitely in an anhydrous state at −20°C under N2 or Ar gas.
References
- 1.Selkoe DJ. The molecular pathology of Alzheimer’s disease. Neuron. 1991;6:487–498. doi: 10.1016/0896-6273(91)90052-2. [DOI] [PubMed] [Google Scholar]
- 2.Selkoe DJ. Cell biology of the amyloid β-protein precursor and the mechanism of Alzheimer’s disease. Annu Rev Cell Biol. 1994;10:373–403. doi: 10.1146/annurev.cb.10.110194.002105. [DOI] [PubMed] [Google Scholar]
- 3.Haass C, Selkoe DJ. Soluble protein oligomers in neurodegeneration: lessons from the Alzheimer’s amyloid β-peptide. Nat Rev Mol Cell Biol. 2007;8:101–112. doi: 10.1038/nrm2101. [DOI] [PubMed] [Google Scholar]
- 4.Kirkitadze MD, Bitan G, Teplow DB. Paradigm shifts in Alzheimer’s disease and other neurodegenerative disorders: the emerging role of oligomeric assemblies. J Neurosci Res. 2002;69:567–577. doi: 10.1002/jnr.10328. [DOI] [PubMed] [Google Scholar]
- 5.Walsh DM, Selkoe DJ. Aβ oligomers - a decade of discovery. J Neurochem. 2007;101:1172–1184. doi: 10.1111/j.1471-4159.2006.04426.x. [DOI] [PubMed] [Google Scholar]
- 6.Roychaudhuri R, Yang M, Hoshi MM, Teplow DB. Amyloid β-protein assembly and Alzheimer disease. J Biol Chem. 2009;284:4749–4753. doi: 10.1074/jbc.R800036200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Shankar GM, Li S, Mehta TH, Garcia-Munoz A, Shepardson NE, Smith I, Brett FM, Farrell MA, Rowan MJ, Lemere CA, Regan CM, Walsh DM, Sabatini BL, Selkoe DJ. Amyloid-β protein dimers isolated directly from Alzheimer’s brains impair synaptic plasticity and memory. Nat Med. 2008;14:837–842. doi: 10.1038/nm1782. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lesne S, Koh MT, Kotilinek L, Kayed R, Glabe CG, Yang A, Gallagher M, Ashe KH. A specific amyloid-β protein assembly in the brain impairs memory. Nature. 2006;440:352–357. doi: 10.1038/nature04533. [DOI] [PubMed] [Google Scholar]
- 9.Lambert MP, Barlow AK, Chromy BA, Edwards C, Freed R, Liosatos M, Morgan TE, Rozovsky I, Trommer B, Viola KL, Wals P, Zhang C, Finch CE, Krafft GA, Klein WL. Diffusible, nonfibrillar ligands derived from Aβ1–42 are potent central nervous system neurotoxins. Proc Natl Acad Sci USA. 1998;95:6448–6453. doi: 10.1073/pnas.95.11.6448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Bitan G, Kirkitadze MD, Lomakin A, Vollers SS, Benedek GB, Teplow DB. Amyloid β-protein (Aβ) assembly: Aβ40 and Aβ42 oligomerize through distinct pathways. Proc Natl Acad Sci USA. 2003;100:330–335. doi: 10.1073/pnas.222681699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Harper JD, Wong SS, Lieber CM, Lansbury PT. Observation of metastable Aβ amyloid protofibrils by atomic force microscopy. Chem Biol. 1997;4:119–125. doi: 10.1016/s1074-5521(97)90255-6. [DOI] [PubMed] [Google Scholar]
- 12.Walsh DM, Lomakin A, Benedek GB, Condron MM, Teplow DB. Amyloid β-protein fibrillogenesis. Detection of a protofibrillar intermediate. J Biol Chem. 1997;272:22364–22372. doi: 10.1074/jbc.272.35.22364. [DOI] [PubMed] [Google Scholar]
- 13.Barghorn S, Nimmrich V, Striebinger A, Krantz C, Keller P, Janson B, Bahr M, Schmidt M, Bitner RS, Harlan J, Barlow E, Ebert U, Hillen H. Globular amyloid β-peptide1–42 oligomer - a homogenous and stable neuropathological protein in Alzheimer’s disease. J Neurochem. 2005;95:834–847. doi: 10.1111/j.1471-4159.2005.03407.x. [DOI] [PubMed] [Google Scholar]
- 14.Hoshi M, Sato M, Matsumoto S, Noguchi A, Yasutake K, Yoshida N, Sato K. Spherical aggregates of β-amyloid (amylospheroid) show high neurotoxicity and activate tau protein kinase I/glycogen synthase kinase-3β. Proc Natl Acad Sci USA. 2003;100:6370–6375. doi: 10.1073/pnas.1237107100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Teplow DB. Preparation of amyloid β-protein for structural and functional studies. Methods Enzymol. 2006;413:20–33. doi: 10.1016/S0076-6879(06)13002-5. [DOI] [PubMed] [Google Scholar]
- 16.Bitan G, Lomakin A, Teplow DB. Amyloid β-protein oligomerization: prenucleation interactions revealed by photo-induced cross-linking of unmodified proteins. J Biol Chem. 2001;276:35176–35184. doi: 10.1074/jbc.M102223200. [DOI] [PubMed] [Google Scholar]
- 17.Fancy DA. Elucidation of protein-protein interactions using chemical cross-linking or label transfer techniques. Curr Opin Chem Biol. 2000;4:28–33. doi: 10.1016/s1367-5931(99)00047-2. [DOI] [PubMed] [Google Scholar]
- 18.Fancy DA, Kodadek T. Chemistry for the analysis of protein-protein interactions: Rapid and efficient cross-linking triggered by long wavelength light. Proc Natl Acad Sci USA. 1999;96:6020–6024. doi: 10.1073/pnas.96.11.6020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Jin Y, Manabe T. High-efficiency protein extraction from polyacrylamide gels for molecular mass measurement by matrix-assisted laser desorption/ionization-time of flight-mass spectrometry. Electrophoresis. 2005;26:1019–1028. doi: 10.1002/elps.200410187. [DOI] [PubMed] [Google Scholar]
- 20.Ono K, Condron MM, Teplow DB. Structure-neurotoxicity relationships of amyloid β-protein oligomers. Proc Natl Acad Sci USA. 2009;106:14745–14750. doi: 10.1073/pnas.0905127106. [DOI] [PMC free article] [PubMed] [Google Scholar]