Abstract
Culturomics investigates microbial diversity of the human microbiome by combining diversified culture conditions, matrix-assisted laser desorption/ionization time-of-flight mass spectrometry and 16S rRNA gene identification. The present study allowed identification of four putative new Clostridium sensu stricto species: ‘Clostridium amazonitimonense’ strain LF2T, ‘Clostridium massilidielmoense’ strain MT26T, ‘Clostridium nigeriense’ strain Marseille-P2414T and ‘Clostridium merdae’ strain Marseille-P2953T, which we describe using the concept of taxonogenomics. We describe the main characteristics of each bacterium and present their complete genome sequence and annotation.
Keywords: ‘Clostridium amazonitimonense’, ‘Clostridium massilidielmoense’, ‘Clostridium merdae’, ‘Clostridium nigeriense’, culturomics, emerging bacteria, human microbiota, taxonogenomics
Introduction
As showed by a remarkable number of studies, the human gut microbiota plays a tremendous role in human health and the homeostasis of the human body [1], [2], [3]. Conversely, an increasing number of links between the gut microbiota and several diseases has been uncovered; these diseases include nutritional disorders, metabolic diseases and pathologies of the intestine [1], [4], [5], [6]. Techniques used in exploratory studies have evolved alongside technology from culture-based techniques in early studies to today's culture-independent techniques [7]. A turning point in the study of the gut microbiota is in progress, with a return to culture-based techniques. In fact, cultured strains are needed for in vivo and in vitro experiments to apply the extensive knowledge acquired thanks to metagenomic studies. Culturomics, a method invented in our laboratory, is a part of this turning point [8]. It consists of the multiplication of culture conditions with a variation of media and physicochemical parameters such as atmosphere, temperature, pH and salinity in order to exhaustively explore a microbial ecosystem [9].
As part of a global exploratory study designed to improve the known and cultivated bacterial diversity of the human gut microbiota, several previously unknown species of Clostridium were isolated. Exploratory studies have shown that the healthy gut microbiota mostly consists of anaerobic bacteria, which represent 95% of the bacterial diversity [2]. Among these anaerobic bacteria, the genus Clostridium is one of the most represented in the human gut, although taxonomic chaos is associated with this genus and its higher ranks in taxonomy, such as the family Clostridiaceae and the Clostridiales order [7]. This genus was created in 1880 by Prazmowski with the type species Clostridium butyricum [10], [11]. In early taxonomy, which was based on phenotypic characteristics, all Gram-positive, anaerobic and spore-forming bacilli were attributed to the genus Clostridium. Consequently, with the advent of DNA sequencing and phylogenetic analysis, taxonomic chaos was uncovered in this genus, and several clusters were subsequently built, among which cluster I, Clostridium sensu stricto, was built around C. butyricum [11], [12]. Among the 211 Clostridium species with standing in nomenclature, only a third are Clostridium sensu stricto species [10], [12]. These mesophilic species are mostly strict anaerobes with Gram-positive staining and the ability to sporulate. Their metabolism is highly variable because they are able to metabolize various compounds such as carbohydrates, amino acids, alcohols, amino acids and purines. Acid and alcohol can be produced from carbohydrates or via protein metabolism [10], [11].
Here we propose the creation of four putative new Clostridium sensu stricto species: ‘Clostridium amazonitimonense’ strain LF2T (= CSUR P1445 = DSM 28600), ‘Clostridium massilidielmoense’ strain MT26T (= CSUR P2255 = CCUG 69893), ‘Clostridium nigeriense’ strain Marseille-P2414T (= CSUR P2414 = DSM 102218) and one new putative Clostridium cluster IV, ‘Clostridium merdae’ strain Marseille-P2953T (= CSUR P2953 = DSM 103669), which we describe using the concept of taxonogenomics [13]. This concept of new species description is based on the presentation of the phylogenetic, phenotypic and genomic features of the studied new species and their comparison to closely related species.
Materials and methods
Sample information
In order to explore the bacterial diversity of the human gut microbiota, stool samples were collected from several individuals: a healthy 8-month-old Senegalese boy, a healthy 44-month-old Senegalese girl, a healthy 37-year-old Amazonian woman from French Guyana and a healthy 29-year-old Senegalese woman. None of the donors was receiving antibiotic treatment at the time of sample collection. Oral consent was obtained from each adult. For the children, oral consent was provided by the parents. Samples were stored at −80°C upon arrival in the La Timone hospital in Marseille. This study received approval from the Institut Fédératif de Recherche 48 (Faculty of Medicine, Marseille, France) under agreement 09-022.
Strain identification
Each stool sample was cultured using the 18 culture conditions of standardized culturomics [8], [9]. For each condition, preincubation in the medium of an anaerobic blood culture bottle from BD BACTEC (Becton Dickinson [BD], San Diego, CA, USA) was performed during 30 days with seeding on 5% sheep's blood–enriched Colombia agar (bioMérieux, Marcy l’Étoile, France) every 3 days. Colonies were purified before identification using a Microflex spectrometer and a MTP 96 matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) target plate (Bruker Daltonics, Bremen, Germany) as described previously [14], [15]. Each obtained spectrum was matched against the database from Bruker and from the La Timone hospital using MALDI Biotyper 3.0 software by standard pattern matching (with default parameter settings). Matching scores under 1.7 indicated the absence of a match in the database. For these unidentified colonies, the 16S rRNA gene was sequenced using fD1 and rP2 primers as previously described [16]. Kim et al. [17] recommended the creation of a new species at the 16S similarity level threshold of 98.65% with the closest species with standing in nomenclature. The corresponding spectra are then incremented in the database and compared to the spectra of closely related species.
Growth conditions
For each species, optimal growth conditions were determined by testing various temperatures (25, 28, 37, 45 and 56°C), atmospheres (aerobic with or without 5% CO2, microaerophilic and anaerobic atmospheres using the GENbag anaer and GENbag microaer systems, respectively; bioMérieux), pH values (6, 6.5, 7 and 8.5) and NaCl concentrations (0.5%, 1%, 5%, 7.5% and 10%).
Morphologic, biochemical and antibiotic susceptibility tests
The morphologic characteristics of each species, namely Gram staining, motility and spore formation, were determined as previously described [8]. To determine the size, and to confirm the shape of the cells as well as spore formation, the cells were negatively stained and observed with electron microscopy as follows: cells were fixed with 2.5% glutaraldehyde in 0.1 M cacodylate buffer for at least 1 hour at 4°C. A drop of cell suspension was deposited for approximately 5 minutes on glow-discharged formvar carbon film on 400 mesh nickel grids (FCF400-Ni; Electron Microscopy Sciences (EMS), Hatfield, PA, USA). The grids were dried on blotting paper, and cells were negatively stained for 10 seconds with 1% ammonium molybdate solution in filtered water at room temperature. Electron micrographs were acquired with a Tecnai G20 Cryo (FEI Company, Limeil-Brevannes, France) transmission electron microscope operated at 200 keV. Cells were then measured by ImageJ software (Image Processing and Analysis in Java; US National Institutes of Health, Bethesda, MD, USA; http://imagej.nih.gov/) [18], [19].
Cellular fatty acid methyl ester (FAME) analysis was performed by gas chromatography/mass spectrometry (GC/MS). Two samples were prepared with approximately 23 mg of bacterial biomass per tube collected from several culture plates. Fatty acid methyl esters were prepared as described by Sasser [20]. GC/MS analyses were carried out as described previously [21]. Briefly, fatty acid methyl esters were separated using an Elite 5-MS column and monitored by mass spectrometry (Clarus 500–SQ 8 S; PerkinElmer, Courtaboeuf, France). A spectral database search was performed using MS Search 2.0 operated with the Standard Reference Database 1A (National Institute of Standards and Technology, Gaithersburg, MD, USA) and the FAME mass spectral database (Wiley, Chichester, UK).
The metabolic capacities of the strains were determined using API ZYM, 20A and 50CH strips according to the manufacturer's instructions. After inoculation of the strips, they were incubated in anaerobic atmosphere for 4, 24 and 48 hours. Other enzymatic features were tested separately, namely oxidase and catalase, which were tested as previously described [8].
The disc diffusion method was used to test antibiotic susceptibility according to European Committee on Antimicrobial Susceptibility Testing recommendations [22].
Genomic DNA preparation and genome sequencing
Bacterial cell lysis was achieved using pretreatments such as lysozyme incubation at 37°C for 2 hours, followed by proteinase K action. DNA was extracted by the EZ1 biorobot (Qiagen, Germantown, MD, USA) with the EZ1 DNA tissues kit. The elution volume was 50 μL. Genomic DNA (gDNA) was quantified by a Qubit assay with the high sensitivity kit (Life Technologies, Carlsbad, CA, USA) to 123.7 ng/μL.
gDNA was sequenced with a MiSeq sequencer (Illumina, San Diego, CA, USA) using the mate-pair strategy. The gDNA was barcoded in order to be mixed with 11 other projects using the Nextera Mate Pair sample prep kit (Illumina). The mate-pair library was prepared with 1.5 μg of gDNA using the Nextera mate-pair Illumina guide. The gDNA sample was simultaneously fragmented and tagged with a mate-pair junction adapter. The fragmentation pattern was validated on an Agilent 2100 BioAnalyzer (Agilent Technologies, Santa Clara, CA, USA) with a DNA 7500 labchip. The DNA fragments ranged in size from 1.5 to 11 kb, with an optimal size at 4.031 kb. No size selection was performed, and 385.5 ng of tagmented fragments were circularized. The circularized DNA was mechanically sheared to small fragments with an optimal size of 1070 bp on the Covaris device S2 in T6 tubes (Covaris, Woburn, MA, USA). The library profile was visualized on a High Sensitivity Bioanalyzer LabChip (Agilent Technologies), and the final concentration library was measured at 2.40 nmol/L. The libraries were normalized at 2 nM and pooled. After a denaturation step and dilution at 15 pM, the pool of libraries was loaded onto the reagent cartridge and then onto the instrument along with the flow cell. The automated cluster generation and sequencing runs were performed in a single 2 × 301 bp run.
Genome annotation and analysis
Open reading frames (ORFs) were predicted using Prodigal [23] with default parameters, but the predicted ORFs were excluded if they spanned a sequencing gap region. The predicted bacterial protein sequences were searched against the GenBank [24] and Clusters of Orthologous Groups (COGs) databases using BLASTP (E value 1e-03, coverage 0.7, identity percentage of 30%). If no hit was found, it was searched against the NR database using BLASTP with an E value of 1e-03, a coverage of 0.7 and an identity percentage of 30%, and if the sequence length was smaller than 80 aa, we used an E value of 1e-05. The tRNAScanSE tool [25] was used to find tRNA genes, while ribosomal RNAs were found using RNAmmer [26]. Lipoprotein signal peptides and the number of transmembrane helices were predicted using Phobius [27]. Mobile genetic elements were predicted using PHAST [28] and RAST [29]. ORFans were identified if all the BLASTP performed did not give positive results (E value smaller than 1e-03 for ORFs with sequence size larger than 80 aa or E value smaller than 1e-05 for ORFs with sequence length smaller than 80 aa). Such parameter thresholds have already been used in previous studies to define ORFans. Artemis [30] and DNA Plotter [31] were used for data management and the visualization of genomic features, respectively. The Mauve 2.3.1 alignment tool was used for multiple genomic sequence alignment [32].
Comparator species for genomic comparison were identified in the 16S RNA tree using PhyloPattern software [33]. For each selected genome, the complete genome sequence, proteome genome sequence and ORFeome genome sequence were retrieved from the National Center for Biotechnology Information FTP site. An annotation of the entire proteome was performed to define the distribution of functional classes of predicted genes according to the clusters of orthologous groups of proteins (using the same method as for the genome annotation). Annotation and comparison processes were performed in the Multi-Agent software system DAGOBAH [34], which includes Figenix [35] libraries that provide pipeline analysis. To evaluate the genomic similarity between studied genomes, we determined two parameters: digital DNA-DNA hybridization (DDH), which exhibits a high correlation with DDH [36], [37]; and average genomic identity of orthologous gene sequences (AGIOS) [38], which was designed to be independent from DDH. The AGIOS score is the mean value of nucleotide similarity between all couples of orthologous proteins between the two studied genomes [38], [39].
Results
Strain identification and phylogenetic analyses
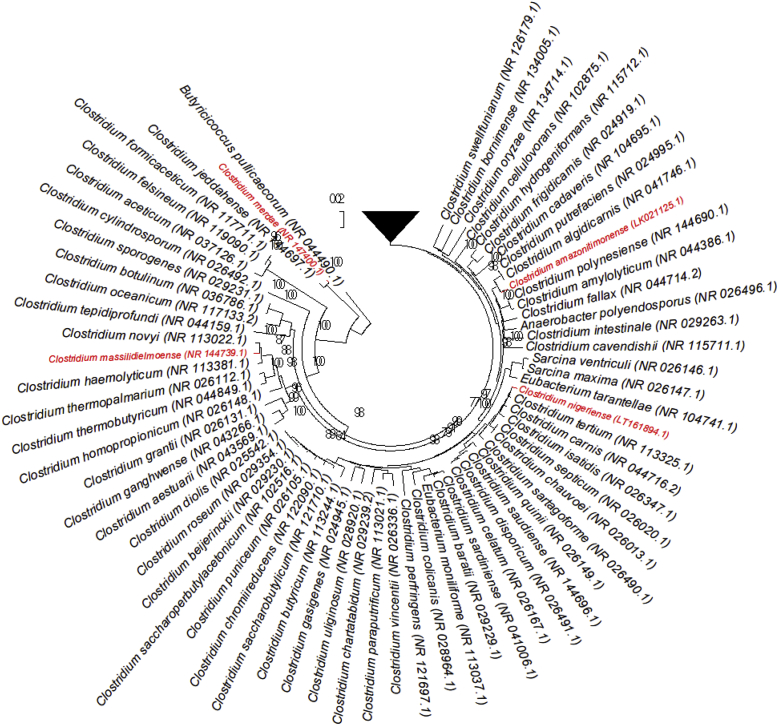
Isolation of strains Marseille-P2953 and LF2 occurred, respectively, after 7 days' and 30 days' preincubation in anaerobic blood culture bottles supplemented with rumen and sheep's blood, respectively. Strain Marseille-P2414 was first isolated after 15 days' preincubation in an aerobic blood culture bottle supplemented with rumen and sheep's blood. Finally, strain MT26 was first isolated after 30 days' preincubation in an anaerobic blood culture bottle supplemented with sheep's blood. Because the routine identification using MALDI-TOF MS failed, the 16S rRNA gene of each strain was sequenced. The 16S rRNA gene sequences of strains Marseille-P2953 (LT732642), LF2 (LK021125), Marseille-P2414 (LT161894) and MT26 (LN998063) revealed a 96.41%, 96.95%, 98.23% and 98.44% similarity level with, respectively, Clostridium sporosphaeroides strain DSM 1294, Clostridium putrefaciens strain DSM 1291, Clostridium tertium strain JCM 6289 and Clostridium novyi strain JCM 1406 (Fig. 1, Table 1). All these similarity levels were lower than the 98.65 cutoff defined to create a new species without performing DDH. We thus propose the creation of four new species within the genus Clostridium with the following names: ‘Clostridium merdae’ (strain Marseille-P2953), ‘Clostridium amazonitimonense’ (strain LF2), ‘Clostridium nigeriense’ (strain Marseille-P2414) and ‘Clostridium massilidielmoense’ (strain MT26). The MALDI-TOF MS spectra of the four strains in this study were incremented in our database and compared to those of their closest relatives (Fig. 2).
Fig. 1.
Phylogenetic tree highlighting position of ‘Clostridium amazonitimonense,’ ‘Clostridium merdae,’ ‘Clostridium massilidielmoense’ and ‘Clostridium nigeriense’ relative to other type strains within Clostridium genus. Strains and their corresponding GenBank accession numbers for 16S rRNA genes are indicated in brackets. Sequences were aligned using Clustal W (http://www.clustal.org/clustal2/), and phylogenetic inferences were obtained using maximum-likelihood method within MEGA6 software (http://www.megasoftware.net/mega.php). Numbers at nodes are percentages of bootstrap values obtained by repeating analysis 1000 times to generate majority consensus tree. Butyricicoccus pullicaecorum (NR_044490) was used as outgroup. Scale bar = 0.02% nucleotide sequence divergence.
Table 1.
Species information
| Characteristic | LF2 | MT26 | Marseille-P2953 | Marseille-P2414 |
|---|---|---|---|---|
| Patient information | Healthy 37-year-old Amazonian woman | Healthy 8-month-old Senegalese boy | Healthy 29-year-old Senegalese woman | Healthy 44-month-old girl from Niger |
| Isolation conditions | 30 days' anaerobic preincubation in blood culture bottle with rumen and sheep's blood | 30 days' anaerobic preincubation in blood culture bottle with sheep's blood | 7 days' anaerobic preincubation in blood culture bottle with rumen and sheep's blood | 15 days' aerobic preincubation in blood culture bottle with rumen and sheep's blood |
| Optimal growth temperature | 37°C | 37°C | 37°C | 37°C |
| Atmosphere | Strict anaerobe | Strict anaerobe | Strict anaerobe | Facultative anaerobe |
| Similarity percentage with closest species | 96.95% with Clostridium putrefaciens strain DSM 1291T (AF127024) | 98.44% with Clostridium novyi strain JCM 1406T (AB045606) | 96.41% with Clostridium sporosphaeroides strain DSM 1294T (X66002) | 98.23% with Clostridium tertium strain JCM 6289T (Y18174) |
Fig. 2.
Reference mass spectra from ‘Clostridium amazonitimonense,’ ‘Clostridium merdae,’ ‘Clostridium massilidielmoense’ and ‘Clostridium nigeriense.’ Spectra from 12 individual colonies were compared and each reference spectrum generated (A). Gel view comparing ‘Clostridium amazonitimonense,’ ‘Clostridium merdae,’ ‘Clostridium massilidielmoense’ and ‘Clostridium nigeriense’ to other species within genus Clostridium. Gel view displays raw spectra of loaded spectrum files arranged in pseudo–gellike look. x-axis records m/z value. Left y-axis displays running spectrum number originating from subsequent spectra loading. Peak intensity expressed by greyscale scheme code. Colour bar and right y-axis indicate relation between colour peaks, with peak intensity in arbitrary units. Displayed species are indicated at left (B).
Phenotypic description
Growth was observed between 28 and 45°C for all studied strains. While strain Marseille-P2414 grew in all tested atmospheres (namely aerobic, microaerophilic and anaerobic), growth was observed only in anaerobic atmosphere for strains Marseille-P2953, LF2 and MT26. All strains exhibited optimal growth at 37°C in anaerobic atmosphere with an incubation period of 24 hours for strain Marseille-P2414 and 48 hours for strains Marseille-P2953, LF2 and MT26. None of the strains was tolerant to NaCl, only growing at a concentration of 0.5%. Conversely, all strains grew at all tested pH concentrations (6, 6.5, 7, 8.5).
The phenotypic characteristics of the four strains in this study and their closest relatives are summarized in Table 2. All strains were Gram positive and were spore-forming, rod-shaped cells (Fig. 3). Strains LF2 and Marseille-P2414 were motile, whereas strains Marseille-P2953 and MT26 were nonmotile. Major fatty acids are presented in Table 3. We observed that strains Marseille-P2414, LF2 and MT26 present hexadecanoic acid (16:0) as the major fatty acid composing their cell envelope, confirming these three strains belong to the same group. For strain Marseille-P2953, its major fatty acid was 9-octadecenoic acid (18:1n9), which is minor or absent in the other strains. Although phylogeny clearly classified Marseille-P2953 as a Clostridium species, these data confirmed other phenotypical observations, suggesting that this strain could belong to another Clostridium cluster.
Table 2.
Differential characteristics of ‘Clostridium amazonitimonense,’ ‘Clostridium putrefaciens,’ ‘Clostridium merdae,’ ‘Clostridium sporosphaeroides,’ ‘Clostridium nigeriense,’ Clostridium tertium, ‘Clostridium massilidielmoense’ and Clostridium novyi[1], [2]
| Characteristic | ‘C. amazonitimonense’ | C. putrefaciens | ‘C. merdae’ | C. sporosphaeroides | ‘C. nigeriense’ | C. tertium | ‘C. massilidielmoense’ | C. novyi |
|---|---|---|---|---|---|---|---|---|
| Property | ||||||||
| Cell shape | Bacilli | Bacilli | Bacilli | Bacilli | Bacilli | Bacilli | Bacilli | Bacilli |
| Cell size (μm) | 2–2.5 | NA | 1.2–1.5 | 0.5–0.6 | 0.8–1.5 | 0.5–1.4 | 0.54 | 0.6–1.4 |
| Gram stain | + | + | + | + | + | + | + | + |
| Oxygen tolerance | − | − | − | − | + | + | − | − |
| Salt tolerance | − | − | − | − | − | − | − | − |
| Motility | + | − | + | − | + | + | − | + |
| Endospore formation | + | + | + | + | + | + | + | + |
| Major cellular fatty acid | 15:0 iso | NA | 16:00 | NA | 16:00 | NA | 16:00 | NA |
| Production of: | ||||||||
| Catalase | − | NA | − | NA | − | NA | − | NA |
| Oxidase | − | NA | − | NA | + | NA | − | NA |
| Alkaline phosphatase | + | NA | − | NA | + | NA | + | NA |
| Urease | − | NA | − | NA | − | NA | − | NA |
| β-Galactosidase | − | NA | − | NA | − | NA | − | NA |
| N-acetyl-glucosamine | + | NA | − | NA | + | NA | + | NA |
| Acid from: | ||||||||
| l-Arabinose | − | NA | − | NA | − | NA | − | NA |
| Ribose | + | NA | − | NA | + | NA | + | NA |
| Mannose | + | NA | − | NA | + | NA | + | NA |
| Mannitol | − | NA | − | NA | − | NA | − | NA |
| Sucrose | − | − | − | NA | + | NA | − | NA |
| d-Glucose | + | + | + | NA | + | NA | + | NA |
| d-Fructose | + | NA | − | NA | + | NA | + | NA |
| d-Maltose | − | − | + | NA | + | NA | + | NA |
| d-Lactose | − | − | − | NA | + | NA | − | NA |
| Habitat | Human gut | Hog muscle tissue | Human gut | Human gut | Human gut | Human gut | Human gut | Soil |
+, positive result; −, negative result; NA, data not available.
Fig. 3.
Electron micrographs of ‘Clostridium nigeriense’ (A), ‘Clostridium amazonitimonense’ (B), ‘Clostridium massilidielmoense’ (C) and ‘Clostridium merdae’ (D).
Table 3.
Cellular fatty acid composition (%)
| Fatty acid | Name | ‘Clostridium nigeriense’ | ‘Clostridium merdae’ | ‘Clostridium massilidielmoense’ | ‘Clostridium amazonitimonense’ |
|---|---|---|---|---|---|
| 16:00 | Hexadecanoic acid | 38.9 ± 0.8 | 38.5 ± 2.1 | 55.0 ± 2.5 | 15.3 ± 2.3 |
| 14:00 | Tetradecanoic acid | ND | ND | 15.7 ± 1.7 | 12.0 ± 0.6 |
| 18:00 | Octadecanoic acid | 6.7 ± 1.1 | 3.2 ± 0.6 | 3.6 ± 0.9 | TR |
| 13:00 | Tridecanoic acid | ND | ND | 3.0 ± 0.6 | ND |
| 12:00 | Dodecanoic acid | 1.8 ± 0.1 | TR | 1.5 ± 1.0 | TR |
| 04:00 | Butanoic acid | ND | ND | ND | 1.7 ± 1.0 |
| 15:00 | Pentadecanoic acid | TR | ND | 2.9 ± 0.6 | ND |
| 18:1n9 | 9-Octadecenoic acid | 20.1 ± 0.4 | 35.4 ± 0.4 | 14.4 ± 1.0 | 2.3 ± 0.6 |
| 13:0 iso | 11-methyl-Dodecanoic acid | ND | ND | ND | 22.7 ± 1.8 |
| 5:0 iso | 3-methyl-Butanoic acid | ND | ND | ND | 11.2 ± 2.1 |
| 18:1n7 | 11-Octadecenoic acid | 7.9 ± 0.2 | ND | ND | TR |
| 14:01 | Tetradecanoic acid | 6.6 ± 0.3 | 5.3 ± 1.4 | ND | ND |
| 18:2n6 | 9.12-Octadecadienoic acid | 5.5 ± 0.4 | 15.2 ± 0.6 | 3.9 ± 1.3 | TR |
| 16:1n7 | 9-Hexadecenoic acid | 5.3 ± 0.2 | TR | ND | ND |
| 16:1n9 | 7-Hexadecenoic acid | 3.9 ± 0.2 | ND | ND | ND |
| 13:0 anteiso | 10-methyl-Dodecanoic acid | ND | ND | ND | 2.3 ± 0.3 |
| 15:0 anteiso | 12-methyl-Tetradecanoic acid | TR | TR | ND | 4.8 ± 0.5 |
| 15:0 iso | 13-methyl-Tetradecanoic acid | TR | TR | ND | 22.0 ± 1.5 |
ND, not detected; TR, traces.
Genome description and comparison
The genomic properties of all strains are presented in Table 4 and Fig. 4. Genome sizes of the strains ranged from 2.75 Mb for strain Marseille-P2953 to 3.8 Mb for strain Marseille-P2414, while the G+C content ranged from 28.2% for strain Marseille-P2414 to 45.4% for strain Marseille-P2953. The proportion of protein-coding genes was similar in all four genomes, and the distribution into COGs categories was also similar (Table 5). However, we observed that the genome of ‘Clostridium merdae’ strain Marseille-P2953 presented a G+C% level significantly higher than the genomes of the other presented strains (Table 6). The G+C content of strain Marseille-P2953 sets it apart from the three other studied strains, as previously observed by its phenotypical features. Thus, the very large family of Clostridiaceae contains species with great phenotypical and genomic diversity, suggesting that this family, which was created before modern sequencing methods were in use, needs to be greatly reclassified. These observations demonstrate that phenotypical observations are no longer sufficient to classify all the newly discovered species. Whole genome sequencing must be used to characterize new bacterial species.
Table 4.
Nucleotide content and gene count levels of genomes
| Attribute |
‘Clostridium amazonitimonense’ |
‘Clostridium merdae’ |
‘Clostridium nigeriense’ |
‘Clostridium massilidielmoense’ |
||||
|---|---|---|---|---|---|---|---|---|
| No. of reads |
5 235 372 |
1 841 718 |
1 832 750 |
3 097 182 |
||||
| Scaffolds/contigs | 4/15 |
4/4 |
4/19 |
9/9 |
||||
| Value | % of totala | Value | % of totala | Value | % of totala | Value | % of totala | |
| Size (bp) | 3 761 417 | 100 | 2 748 860 | 100 | 3 799 474 | 100 | 2 759 335 | 100 |
| G+C content (%) | 1 145 890 | 30.55 | 1 246 258 | 45.4 | 1 064 556 | 28.2 | 782 307 | 28.3 |
| Coding region (bp) | 3 129 293 | 83.2 | 2 373 508 | 86.3 | 3 322 705 | 87.4 | 2 453 243 | 88.9 |
| Total genes | 3553 | 100 | 2699 | 100 | 3659 | 100 | 2772 | 100 |
| RNA genes | 147 | 4.1 | 59 | 2.2 | 187 | 5.1 | 91 | 3.3 |
| Protein-coding genes | 3406 | 95.9 | 264 | 97.8 | 3472 | 94.9 | 2681 | 96.7 |
| Genes with function prediction | 2477 | 72.7 | 1966 | 74.4 | 245 | 70.5 | 1989 | 74.2 |
| Genes assigned to COGs | 2241 | 65.8 | 167 | 63.2 | 2149 | 61.9 | 1639 | 61.1 |
| Genes with peptide signals | 270 | 7.9 | 271 | 10.2 | 307 | 8.8 | 228 | 2.8 |
| ORFans genes | 189 | 5.5 | 117 | 4.4 | 119 | 3.4 | 40 | 1.5 |
| Genes with transmembrane helices | 849 | 24.9 | 569 | 21.5 | 935 | 26.9 | 639 | 23.8 |
| Genes associated with PKS or NRPS | 3 | 0.08 | 0 | 0 | 5 | 0.1 | 3 | 0.1 |
| No. of antibiotic resistance genes | 1 | 0.03 | 0 | 0 | 0 | 0 | 0 | 0 |
| No. of genes associated with Pfam-A domains | NA | NA | 2386 | 88 | 3182 | 86 | 246 | 88 |
COGs, Clusters of Orthologous Groups database; NRPS, nonribosomal peptide synthase; ORF, open reading frame; PKS, polyketide synthase.
The total is based on either the size of the genome in base pairs or the total number of protein-coding genes in the annotated genome.
Fig. 4.
Graphical circular map of genome of ‘Clostridium nigeriense’ (A), ‘Clostridium amazonitimonense’ (B), ‘Clostridium massilidielmoense’ (C) and ‘Clostridium merdae’ (D). From outside to center: Contigs (red/grey), COGs category of genes on forward strand (three circles), genes on forward strand (blue circle), genes on reverse strand (red circle), COGs category on reverse strand (three circles), G+C content. COGs, Clusters of Orthologous Groups database.
Table 5.
Number of genes associated with 25 general COGs functional categories
| Code | Description |
‘Clostridium merdae’ |
‘Clostridium nigeriense’ |
‘Clostridium massilidielmoense’ |
‘Clostridium amazonitimonense’ |
||||
|---|---|---|---|---|---|---|---|---|---|
| Value | % of total | Value | % of total | Value | % of total | Value | % of totala | ||
| [J] | Translation | 189 | 7.16 | 217 | 6.25 | 192 | 7.16 | 181 | 5.31 |
| [A] | RNA processing and modification | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| [K] | Transcription | 171 | 6.48 | 191 | 5.50 | 125 | 4.66 | 238 | 6.99 |
| [L] | Replication, recombination and repair | 77 | 2.91 | 85 | 2.45 | 87 | 3.24 | 135 | 3.96 |
| [B] | Chromatin structure and dynamics | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0.03 |
| [D] | Cell cycle control, mitosis and meiosis | 31 | 1.17 | 40 | 1.15 | 39 | 1.45 | 24 | 0.70 |
| [Y] | Nuclear structure | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| [V] | Defense mechanisms | 53 | 2.01 | 75 | 2.16 | 58 | 2.16 | 105 | 3.08 |
| [T] | Signal transduction mechanisms | 97 | 3.67 | 117 | 3.37 | 94 | 3.50 | 135 | 3.96 |
| [M] | Cell wall/membrane biogenesis | 75 | 2.84 | 129 | 3.71 | 106 | 3.95 | 104 | 3.05 |
| [N] | Cell motility | 43 | 1.63 | 53 | 1.52 | 47 | 1.75 | 61 | 1.79 |
| [Z] | Cytoskeleton | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| [W] | Extracellular structures | 7 | 0.26 | 6 | 0.17 | 7 | 0.26 | 0 | 0 |
| [U] | Intracellular trafficking and secretion | 20 | 0.76 | 26 | 0.75 | 25 | 0.93 | 34 | 1 |
| [O] | Posttanslational modification, protein turnover, chaperones | 61 | 2.31 | 79 | 2.27 | 75 | 2.8 | 74 | 2.17 |
| [X] | Mobilome: prophages, transposons | 15 | 0.57 | 8 | 0.23 | 36 | 1.34 | NA | NA |
| [C] | Energy production and conversion | 106 | 4.01 | 139 | 4.00 | 93 | 3.47 | 137 | 4.02 |
| [G] | Carbohydrate transport and metabolism | 142 | 5.38 | 271 | 7.80 | 125 | 4.66 | 165 | 4.84 |
| [E] | Amino acid transport and metabolism | 212 | 8.03 | 195 | 5.61 | 150 | 5.59 | 219 | 6.43 |
| [F] | Nucleotide transport and metabolism | 58 | 2.19 | 85 | 2.45 | 68 | 2.53 | 72 | 2.11 |
| [H] | Coenzyme transport and metabolism | 90 | 3.41 | 104 | 2.99 | 111 | 4.14 | 88 | 2.58 |
| [I] | Lipid transport and metabolism | 69 | 2.61 | 84 | 2.42 | 64 | 2.39 | 62 | 1.82 |
| [P] | Inorganic ion transport and metabolism | 87 | 3.29 | 133 | 3.83 | 87 | 3.24 | 153 | 4.49 |
| [Q] | Secondary metabolite biosynthesis, transport and catabolism | 28 | 1.06 | 30 | 0.86 | 13 | 0.48 | 35 | 1.02 |
| [R] | General function prediction only | 158 | 5.98 | 202 | 5.82 | 151 | 5.63 | 320 | 9.39 |
| [S] | Function unknown | 86 | 3.26 | 133 | 3.83 | 91 | 3.39 | 190 | 5.58 |
| — | Not in COGs | 970 | 36.74 | 1.323 | 38.10 | 1.042 | 38.87 | 1.165 | 34.20 |
COGs, Clusters of Orthologous Groups database; NA, not applicable.
The total is based on either the size of the genome in base pairs or the total number of protein-coding genes in the annotated genome.
Table 6.
Genome comparison of closely related species
| Organism | INSDC | Size (Mb) | G+C (%) | Total genes |
|---|---|---|---|---|
| Clostridium beijerinckiia | CP000721 | 6.48 | 30 | 5.425 |
| Clostridium clariflavum | CP003065 | 4.90 | 35.72 | 3.892 |
| Clostridium cellulovorans | CP002160 | 56 | 311 | 4.524 |
| Clostridium perfringensa | CP000246 | 35 | 28.38 | 2.876 |
| ‘Clostridium massilidielmoense’ | FXLO00000000 | 2.76 | 28.35 | 2.681 |
| Clostridium acetireducens | LZFO00000000 | 2.42 | 26.75 | 2.238 |
| Clostridium acetobutylicum | AE001437 | 43 | 30.93 | 3.848 |
| Clostridium argentinense | AYSO00000000 | 4.74 | 28.6 | 4.279 |
| Clostridium carboxidivorans | CP011803 | 5.75 | 29.93 | 4.973 |
| Clostridium drakei | JIBU00000000 | 5.58 | 29 | 5.029 |
| Clostridium haemolyticum | JENX00000000 | 2.61 | 27.43 | 2.201 |
| Clostridium homopropionicum | LHUR00000000 | 3.65 | 31.09 | 3.5 |
| Clostridium novyi | JENM00000000 | 2.46 | 28.58 | 1.833 |
| Clostridium tetani | JRGJ00000000 | 2.79 | 28.57 | 2.516 |
| ‘Clostridium nigeriense’ | FLKB00000000 | 3.79 | 28.7 | 3.472 |
| Clostridium baratii | JZTY00000000 | 37 | 27.95 | 2.839 |
| Clostridium diolis | AQQG00000000 | 5.84 | 29.74 | 5.196 |
| Clostridium paraputrificum | AUJC00000000 | 3.56 | 29.64 | 3.491 |
| Clostridium sartagoforme | ASRV00000000 | 3.98 | 27.91 | 3.714 |
| ‘Clostridium merdae’ | FXLN00000000 | 2.75 | 45.34 | 2.64 |
| Clostridium leptum | ABCB00000000 | 2.82 | 505 | 2.482 |
| Clostridium thermocellum | CP000568 | 3.84 | 38.99 | 3.173 |
| Eubacterium siraeum | ARNC00000000 | 2.62 | 44.85 | 2.308 |
| Flavonifractor plautii | AGCK00000000 | 3.82 | 61.07 | 4.278 |
| Intestinimonas butyricproducens | CP011307 | 3.37 | 59.06 | 3.381 |
| Pseudoflavonifractor capillosus | AAXG00000000 | 44 | 591 | 4.829 |
| Ruminococcus albus | CP002403 | 4.33 | 43.62 | 3.991 |
| ‘Clostridium amazonitimonense’ | CCNN00000000 | 3.76 | 30.56 | 3.046 |
INSDC, International Nucleotide Sequence Database Collaboration.
The total is based on either the size of the genome in base pairs or the total number of protein-coding genes in the annotated genome.
The DDH comparison presented in Table 7 demonstrates that the four studied new species are clearly distinct new species. Indeed, the DDH level between our four new species and the other Clostridium species comprises a range delimiting a distinct bacterial species.
Table 7.
Pairwise comparison of four studied species with nine other species using GGDC
| 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 19.90 ± 2.2 | 18.10 ± 2.2 | 19.00 ± 2.2 | 19.40 ± 2.2 | 18.40 ± 2.1 | 22.60 ± 2.2 | 21.00 ± 2.2 | 20.30 ± 2.2 | 24.40 ± 2.3 | 39.90 ± 2.5 | 23.70 ± 2.3 | 23.20 ± 2.3 | 24.80 ± 2.3 |
| 2 | 100.00 ± 0.0 | 19.40 ± 2.2 | 24.80 ± 2.3 | 39.80 ± 2.5 | 19.70 ± 2.2 | 21.30 ± 2.3 | 19.50 ± 2.2 | 17.40 ± 2.1 | 26.70 ± 2.3 | 26.40 ± 2.4 | 26.00 ± 2.3 | 19.60 ± 2.2 | 25.70 ± 2.3 |
| 3 | 100.00 ± 0.0 | 19.00 ± 2.2 | 18.10 ± 2.1 | 18.50 ± 2.1 | 21.30 ± 2.2 | 20.70 ± 2.2 | 20.00 ± 2.2 | 24.30 ± 2.3 | 26.70 ± 2.4 | 28.70 ± 2.3 | 20.10 ± 2.2 | 26.60 ± 2.4 | |
| 4 | 100.00 ± 0.0 | 24.70 ± 2.3 | 19.50 ± 2.2 | 21.30 ± 2.2 | 19.10 ± 2.2 | 18.30 ± 2.1 | 28.40 ± 2.4 | 26.10 ± 2.3 | 28.30 ± 2.3 | 20.00 ± 2.2 | 23.70 ± 2.3 | ||
| 5 | 100.00 ± 0.0 | 18.30 ± 2.1 | 21.30 ± 2.3 | 19.40 ± 2.1 | 18.40 ± 2.2 | 23.30 ± 2.3 | 25.70 ± 2.3 | 26.70 ± 2.3 | 20.50 ± 2.2 | 26.80 ± 2.3 | |||
| 6 | 100.00 ± 0.0 | 22.10 ± 2.2 | 19.40 ± 2.1 | 18.00 ± 2.1 | 22.90 ± 2.3 | 25.80 ± 2.3 | 28.40 ± 2.4 | 21.20 ± 2.2 | 24.10 ± 2.3 | ||||
| 7 | 100.00 ± 0.0 | 22.20 ± 2.3 | 23.60 ± 2.3 | 23.00 ± 2.3 | 24.70 ± 2.3 | 26.00 ± 2.3 | 24.90 ± 2.3 | 24.30 ± 2.4 | |||||
| 8 | 100.00 ± 0.0 | 20.20 ± 2.2 | 25.30 ± 2.3 | 25.00 ± 2.3 | 28.20 ± 2.4 | 25.10 ± 2.3 | 25.10 ± 2.4 | ||||||
| 9 | 100.00 ± 0.0 | 25.10 ± 2.3 | 24.70 ± 2.3 | 28.10 ± 2.4 | 23.40 ± 2.3 | 25.40 ± 2.3 | |||||||
| 10 | 100.00 ± 0.0 | 22.50 ± 2.2 | 29.10 ± 2.3 | 26.80 ± 2.4 | 29.20 ± 2.4 | ||||||||
| 11 | 100.00 ± 0.0 | 25.10 ± 2.3 | 23.60 ± 2.3 | 29.70 ± 2.4 | |||||||||
| 12 | 100.00 ± 0.0 | 27.30 ± 2.4 | 25.70 ± 2.3 | ||||||||||
| 13 | 100.00 ± 0.0 | 23.80 ± 2.3 | |||||||||||
| 14 | 100.00 ± 0.0 |
DDH, DNA-DNA hybridization; GGDC, genome-to-genome distance; HSP, high-scoring pair.
Data derived from formula 2 (DDH estimates based on identities/HSP length). Confidence intervals indicate inherent uncertainty in estimating DDH values from intergenomic distances based on models derived from empirical test data sets (which are always limited in size). These results are in accordance with 16S rRNA (Fig. 1) and phylogenetic analyses as well as GGDC results.
1, Clostridium cellulovorans; 2, Clostridium massilidielmoense; 3, Clostridium acetireducens; 4, Clostridium haemolyticum; 5, Clostridium novyi; 6, Clostridium tetani; 7, Clostridium nigeriense; 8, Clostridium diolis; 9, Clostridium sartagoforme; 10, Clostridium merdae; 11, Clostridium leptum; 12, Clostridium thermocellum; 13, Clostridium amazonitimonense; 14, Ruminococcus albus.
Strain Marseille-P2414 shared 1342, 1434, 1553, 1564, 1628 and 1664 orthologous proteins with Clostridium perfringens, Clostridium baratii, Clostridium diolis, Clostridium beijerinckii, Clostridium paraputrificum and Clostridium sartagoforme, respectively (Supplementary Table S1a). Strain MT26 shared 1111, 1114, 1151, 1165, 1253, 1254, 1261, 1294 and 1372 orthologous proteins with Clostridium acetireducens, Clostridium acetobutylicum, Clostridium argentinense, Clostridium tetani, Clostridium carboxidivorans, Clostridium drakei, Clostridium homopropionicum, Clostridium novyi and Clostridium haemolyticum, respectively (Supplementary Table S1b). Strain LF2 shared 774, 1005, 1017 and 1138 orthologous proteins with Clostridium clariflavum, Clostridium perfringens, Clostridium cellulovorans and Clostridium beijerinckii, respectively (Supplementary Table S1c).
Finally, strain Marseille-P2953 shared 735 orthologous proteins with Ruminococcus albus and Clostridium thermocellum as well as 740, 810, 833, 838, 867, 889 and 907 orthologous proteins with Eubacterium siraeum, Clostridium leptum, Pseudoflavonifractor capillosus, Intestinimonas butyriciproducens, Flavonifractor plautii, Oscillibacter ruminantium and Oscillibacter valericigenes, respectively (Supplementary Table S1d). Strain Marseille-P2953 shared fewer orthologous proteins than the other described genomes, as this strain belongs to Clostridium cluster IV and has been compared to more closely related species with available genomes, which mostly belong to other genera, unlike the other described strains. AGIOS value ranges were similar among species with standing in nomenclature with compared species and with the described genomes, thus confirming that the described strains are new species.
Conclusion
Considering the 16S rRNA gene sequence similarity level with the closest species, all under the 98.65% cutoff to define a new species without performing DDH; considering the phenotypic characteristics distinguishing these strains from other known Clostridium species; and confirming that they belong to the genus Clostridium and have the genomic characteristics of these strains, we propose the creation of four new Clostridium species: ‘Clostridium nigeriense,’ ‘Clostridium massilidielmoense,’ ‘Clostridium amazonitimonense’ and ‘Clostridium merdae.’
Description of the new species ‘Clostridium nigeriense’
‘Clostridium nigeriense’ (ni.ge.rien'se, L. gen. adj., from nigeriense, ‘of Niger,’ referring to where the stool sample was collected) was isolated from the stool sample of a healthy girl from Niger and was deposited in the DSMZ collection under accession number DSM 102218 and in the CSUR collection under accession number CSUR P2414.
‘Clostridium nigeriense’ is a Gram-positive bacillus that is motile, spore forming and facultative anaerobic. The test of catalase activity was negative but was positive for oxidase. ‘Clostridium nigeriense’ forms translucent colonies with a mean diameter of 3 mm. Its major cellular fatty acid is hexadecanoic acid (16:0).
Its genome is 3 799 474 bp long with 28.27% G+C content. The 16S rRNA gene and genome sequences are available in the European Molecular Biology Laboratory–European Bioinformatics Institute (EMBL-EBI) database under accession numbers LT161894 and FLKB00000000, respectively.
Strain Marseille-P2414T is the type strain of the new species ‘Clostridium nigeriense.’
Description of the new species ‘Clostridium amazonitimonense’
‘Clostridium amazonitimonense’ (am.a.zo.ni.ti.mo.nen'se, N.L. neut. adj., from amazoni (a.ma.zo.ni, N.L. neut. adj.), referring to the Amazonian forest, the location where the stool sample was collected, and timonense (ti.mo.nen'se, N.L. neut. adj.), referring to the La Timone hospital, where the strain was isolated) was isolated from the stool sample of a healthy middle-aged woman from Amazonia and is deposited in the DSMZ collection under accession number DSM 28600 and in the CSUR collection under accession number CSUR P1445.
Strain LF2 is a Gram-positive bacillus that is motile, spore forming and a strict anaerobe. It is negative for catalase and oxidase activity. ‘Clostridium amazonitimonense’ forms white colonies with a mean diameter of 5 mm.
The genome is 3 761 417 bp long with 30.56% G+C content. The 16S rRNA gene and genome sequences are available in the EMBL-EBI database under accession numbers LK021125 and CCNN00000000, respectively.
The type strain LF2T is the type strain of the new species ‘Clostridium amazonitimonense.’
Description of the new species ‘Clostridium massilidielmoense’
‘Clostridium massilidielmoense’ (mas.si.li.di.el.mo.en'se, L. neut. adj., a combination of massili (ma.ssi.li, N.L. neut. adj.), from Massilia, the Roman name of Marseille, where strain MT26 was isolated; and dielmoense (di.el.mo.en'se, N.L. neut. adj.), from Dielmo, the Senegalese village where the boy from whom the strain was cultivated lived) was isolated from the stool sample of a healthy 8-month-old Senegalese boy and is deposited in the CCUG collection under accession number CCUG 69893 and in the CSUR collection under accession number CSUR P2255.
Strain MT26 is a Gram-positive bacillus that is nonmotile, spore forming and a strict anaerobe. It shows positive catalase activity but no oxidase activity. ‘Clostridium massilidielmoense’ forms a translucent biofilm with cells of an average diameter of 0.54 μm.
The genome is 2 759 335 bp long with 28.35% G+C content. The 16S rRNA gene and genome sequences are available in the EMBL-EBI database under accession numbers LN998063 and FXLO00000000, respectively.
The type strain MT26T is the type strain of the new species ‘Clostridium massilidielmoense.’
Description of the new species ‘Clostridium merdae’
‘Clostridium merdae’ (mer'dae, L. gen. n., from merdae, ‘of faeces,’ referring to the fact that the strain was isolated from a stool sample) was isolated from the stool sample of a healthy 29-year-old Senegalese woman and is deposited in the DSMZ collection under accession number DSM 103669 and in the CSUR collection under accession number CSUR P2953.
Strain Marseille-P2953 is a Gram-positive bacillus that is motile, spore forming and a strict anaerobe. It does not show catalase or oxidase activity. ‘Clostridium merdae’ forms small white colonies with a mean diameter of 1 mm.
The genome is 2 748 860 bp long with 45.34% G+C content. The 16S rRNA gene and genome sequences are available in the EMBL-EBI database under accession numbers LT732642 and FXLN00000000, respectively.
The type strain Marseille-P2953 is the type strain of the new species ‘Clostridium merdae.’
Conflict of interest
None declared.
Footnotes
Supplementary data related to this article can be found at https://doi.org/10.1016/j.nmni.2017.11.003.
Appendix A. Supplementary data
The following is the supplementary data related to this article:
References
- 1.Salazar N., Arboleya S., Valdés L., Stanton C., Ross P., Ruiz L. The human intestinal microbiome at extreme ages of life. Dietary intervention as a way to counteract alterations. Front Genet. 2014;5:406. doi: 10.3389/fgene.2014.00406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Lagier J.C., Million M., Hugon P., Armougom F., Raoult D. Human gut microbiota: repertoire and variations. Front Cell Infect Microbiol. 2012;2:136. doi: 10.3389/fcimb.2012.00136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Sankar S.A., Lagier J.C., Pontarotti P., Raoult D., Fournier P.E. The human gut microbiome; a taxonomic conundrum. Syst Appl Microbiol. 2015;38:276–286. doi: 10.1016/j.syapm.2015.03.004. [DOI] [PubMed] [Google Scholar]
- 4.Tidjani Alou M., Lagier J.C., Raoult D. Diet influence on the gut microbiota and dysbiosis related to nutritional disorders. Hum Microbiome J. 2016;1:3–11. [Google Scholar]
- 5.Million M., Diallo A., Raoult D. Gut microbiota and malnutrition. Microb Pathog. 2017;106:127–138. doi: 10.1016/j.micpath.2016.02.003. [DOI] [PubMed] [Google Scholar]
- 6.Angelakis E., Lagier J.C. Samples and techniques highlighting the links between obesity and microbiota. Microb Pathog. 2017;106:119–126. doi: 10.1016/j.micpath.2016.01.024. [DOI] [PubMed] [Google Scholar]
- 7.Rajilić-Stojanović M., de Vos W.M. The first 1000 cultured species of the human gastrointestinal microbiota. FEMS Microbiol Rev. 2014;38:996–1047. doi: 10.1111/1574-6976.12075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lagier J.C., Hugon P., Khelaifia S., Fournier P.E., La Scola B., Raoult D. The rebirth of culture in microbiology through the example of culturomics to study human gut microbiota. Clin Microbiol Rev. 2015;28:237–264. doi: 10.1128/CMR.00014-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Lagier J.C., Armougom F., Million M., Hugon P., Pagnier I., Robert C. Microbial culturomics: paradigm shift in the human gut microbiome study. Clin Microbiol Infect. 2012;18:1185–1193. doi: 10.1111/1469-0691.12023. [DOI] [PubMed] [Google Scholar]
- 10.Parte A.C. LPSN—list of prokaryotic names with standing in nomenclature. Nucleic Acids Res. 2014;42:D613–D616. doi: 10.1093/nar/gkt1111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.De Vos P., Garrity G.M., Jones D., Krieg N.R., Ludwig W., Rainey F.A., editors. Systematic bacteriology. Springer; New York: 2009. [Google Scholar]
- 12.Lawson P.A., Citron D.M., Tyrrell K.L., Finegold S.M. Reclassification of Clostridium difficile as Clostridioides difficile (Hall and O'Toole 1935) Prévot 1938. Anaerobe. 2016;40:95–99. doi: 10.1016/j.anaerobe.2016.06.008. [DOI] [PubMed] [Google Scholar]
- 13.Fournier P.E., Lagier J.C., Dubourg G., Raoult D. From culturomics to taxonomogenomics: a need to change the taxonomy of prokaryotes in clinical microbiology. Anaerobe. 2015;36:73–78. doi: 10.1016/j.anaerobe.2015.10.011. [DOI] [PubMed] [Google Scholar]
- 14.Seng P., Drancourt M., Gouriet F., La Scola B., Fournier P.E., Rolain J.M. Ongoing revolution in bacteriology: routine identification of bacteria by matrix-assisted laser desorption ionization time-of-flight mass spectrometry. Clin Infect Dis. 2009;49:543–551. doi: 10.1086/600885. [DOI] [PubMed] [Google Scholar]
- 15.Seng P., Abat C., Rolain J.M., Colson P., Lagier J.C., Gouriet F. Identification of rare pathogenic bacteria in a clinical microbiology laboratory: impact of matrix-assisted laser desorption ionization–time of flight mass spectrometry. J Clin Microbiol. 2013;51:2182–2194. doi: 10.1128/JCM.00492-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Drancourt M., Bollet C., Carlioz A., Martelin R., Gayral J.P., Raoult D. 16S ribosomal DNA sequence analysis of a large collection of environmental and clinical unidentifiable bacterial isolates. J Clin Microbiol. 2000;38:3623–3630. doi: 10.1128/jcm.38.10.3623-3630.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kim M., Oh H.S., Park S.C., Chun J. Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol. 2014;64:346–351. doi: 10.1099/ijs.0.059774-0. [DOI] [PubMed] [Google Scholar]
- 18.Schindelin J., Rueden C.T., Hiner M.C., Eliceiri K.W. The ImageJ ecosystem: an open platform for biomedical image analysis. Mol Reprod Dev. 2015;82:518–529. doi: 10.1002/mrd.22489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Schneider C.A., Rasband W.S., Eliceiri K.W. NIH Image to ImageJ: 25 years of image analysis. Nat Methods. 2012;9:671–675. doi: 10.1038/nmeth.2089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Sasser M. Microbial ID; Newark, NY: 2006. Bacterial identification by gas chromatographic analysis of fatty acids methyl esters (GC-FAME) [Google Scholar]
- 21.Dione N., Sankar S.A., Lagier J.C., Khelaifia S., Michele C., Armstrong N. Genome sequence and description of Anaerosalibacter massiliensis sp. nov. New Microbe New Infect. 2016;10:66–76. doi: 10.1016/j.nmni.2016.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Matuschek E., Brown D.F.J., Kahlmeter G. Development of the EUCAST disk diffusion antimicrobial susceptibility testing method and its implementation in routine microbiology laboratories. Clin Microbiol Infect. 2014;20:O255–O266. doi: 10.1111/1469-0691.12373. [DOI] [PubMed] [Google Scholar]
- 23.Hyatt D., Chen G.L., Locascio P.F., Land M.L., Larimer F.W., Hauser L.J. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 2010;11:119. doi: 10.1186/1471-2105-11-119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Benson D.A., Karsch-Mizrachi I., Clark K., Lipman D.J., Ostell J., Sayers E.W. GenBank. Nucleic Acids Res. 2012;40:D48–D53. doi: 10.1093/nar/gkr1202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Lowe T.M., Eddy S.R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997;25:955–964. doi: 10.1093/nar/25.5.955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Lagesen K., Hallin P., Rødland E.A., Staerfeldt H.H., Rognes T., Ussery D.W. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res. 2007;35:3100–3108. doi: 10.1093/nar/gkm160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Käll L., Krogh A., Sonnhammer E.L.L. A combined transmembrane topology and signal peptide prediction method. J Mol Biol. 2004;338:1027–1036. doi: 10.1016/j.jmb.2004.03.016. [DOI] [PubMed] [Google Scholar]
- 28.Zhou Y., Liang Y., Lynch K.H., Dennis J.J., Wishart D.S. PHAST: a fast phage search tool. Nucleic Acids Res. 2011;39:W347–W352. doi: 10.1093/nar/gkr485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Aziz R.K., Bartels D., Best A.A., DeJongh M., Disz T., Edwards R.A. The RAST Server: rapid annotations using subsystems technology. BMC Genomics. 2008;9:75. doi: 10.1186/1471-2164-9-75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Rutherford K., Parkhill J., Crook J., Horsnell T., Rice P., Rajandream M.A. Artemis: sequence visualization and annotation. Bioinforma. 2000;16:944–945. doi: 10.1093/bioinformatics/16.10.944. [DOI] [PubMed] [Google Scholar]
- 31.Carver T., Thomson N., Bleasby A., Berriman M., Parkhill J. DNAPlotter: circular and linear interactive genome visualization. Bioinforma. 2009;25:119–120. doi: 10.1093/bioinformatics/btn578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Darling A.C.E., Mau B., Blattner F.R., Perna N.T. Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genome Res. 2004;14:1394–1403. doi: 10.1101/gr.2289704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Gouret P., Thompson J.D., Pontarotti P. PhyloPattern: regular expressions to identify complex patterns in phylogenetic trees. BMC Bioinform. 2009;10:298. doi: 10.1186/1471-2105-10-298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Gouret P., Paganini J., Dainat J., Louati D., Darbo E., Pontarotti P. Integration of evolutionary biology concepts for functional annotation and automation of complex research in evolution: the multi-agent software system DAGOBAH. In: Pontarotti P., editor. Evolutionary biology—concepts, biodiversity, macroevolution and genome evolution. Springer; Berlin: 2011. pp. 71–87. [Google Scholar]
- 35.Gouret P., Vitiello V., Balandraud N., Gilles A., Pontarotti P., Danchin E.G. FIGENIX: intelligent automation of genomic annotation: expertise integration in a new software platform. BMC Bioinform. 2005;6:198. doi: 10.1186/1471-2105-6-198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Auch A.F., von Jan M., Klenk H.-P., Göker M. Digital DNA-DNA hybridization for microbial species delineation by means of genome-to-genome sequence comparison. Stand Genomic Sci. 2010;2:117–134. doi: 10.4056/sigs.531120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Meier-Kolthoff J.P., Auch A.F., Klenk H.-P., Göker M. Genome sequence–based species delimitation with confidence intervals and improved distance functions. BMC Bioinform. 2013;14:60. doi: 10.1186/1471-2105-14-60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Ramasamy D., Mishra A.K., Lagier J.C., Padhmanabhan R., Rossi M., Sentausa E. A polyphasic strategy incorporating genomic data for the taxonomic description of novel bacterial species. Int J Syst Evol Microbiol. 2014;64:384–391. doi: 10.1099/ijs.0.057091-0. [DOI] [PubMed] [Google Scholar]
- 39.Sturges W.S., Drake E.T. A complete description of Clostridium putrefaciens (McBryde) J Bacteriol. 1927;14:175–179. doi: 10.1128/jb.14.3.175-179.1927. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.