Abstract
Circular (circ)RNAs, naturally formed endogenous non-coding RNAs, have received extensive attention in recent years due to their special loop structures and specific function. circRNAs are formed with covalently closed continuous loops and are mainly generated by back-splicing processes or lariat introns from exons and/or introns. Usually, circRNAs are stable, abundant, and evolutionarily conserved in the cytoplasm. circRNAs often exhibit abnormal expression in different diseases, notably in human cancers, and the presence of abundant circRNAs in serum, saliva and exosomes renders them potential diagnostic or predictive biomarkers for diseases, including multiple types of cancer. Presently, certain circRNAs have been reported to function as microRNA sponges and RNA-binding protein sponges to regulate downstream gene transcription, which suggests a potential for circRNAs in cancer diagnosis, prognosis and clinical therapy. The present study assessed the latest advances in the study of circRNAs in cancer, summarized the functions of circRNAs in different types of cancer, highlighted the competing endogenous RNA function of circRNAs in the occurrence and development of human malignancies, and provided evidence for the future application of circRNAs in the diagnosis, prognosis and treatment of multiple types of cancer.
Keywords: circular RNA, competing endogenous RNA, microRNA, RNA-binding protein, cancer
1. Introduction
Cancer is a malignant disease associated with abnormal proliferation, aberrant differentiation and metastasis (1). A high proportion of patients with cancer present with varying degrees of metastasis at the point of diagnosis. Although efforts have been made to identify strategies to treat various types of cancer, numerous types continue to be significant contributors to mortality and high healthcare expenditures in populations (2,3). Therefore, identifying early diagnostic or prognostic biomarkers, and exploring effective treatment strategies to improve survival times in different types of cancer, is urgent.
Tumorigenesis is associated with the functional inhibition or loss of tumor suppressor genes and the overexpression of oncogenes (4–6). With the continuous improvement of high-throughput genome-wide and RNA-sequencing technologies, more non-coding (nc)RNAs associated with human cancer progression have been discovered (7,8). A threshold of 200 nucleotides divides regulatory ncRNAs into short ncRNAs (sncRNAs) and long ncRNAs (lncRNAs) (9,10). sncRNAs include microRNAs (miRNAs), endogenous short interfering RNAs (endo-siRNAs) and Piwi-interacting RNAs (piRNAs). lncRNAs are composed of >200 nucleotides, and include multiple well-studied ncRNAs, such as metastasis-associated lung adenocarcinoma transcript 1 (non-protein coding), hepatocellular carcinoma upregulated lncRNA and HOX transcript antisense RNA. Circular (circ)RNAs, a subset of ncRNAs that are generally composed of >200 nucleotides (with a minority <200 nucleotides), have also been grouped into lncRNAs (11).
circRNAs have become another focus in the field of ncRNA research (12). Previous studies have indicated that certain circRNAs are endogenous, stable, abundant and conserved in eukaryotic cells (13–16). Differing from the linear RNAs, which feature 5′ caps and 3′ tails as their termini, all known circRNAs possess covalently closed loop structures, may occur in any genomic region, including gene-bearing regions and intergenic regions, and range in length from a few hundred to thousands of nucleotides (11,15,17). Previously, certain circRNAs have been identified as potential biomarkers for aging in Drosophila (18), which suggested the potential of circRNAs as biomarkers for the diagnosis and treatment of cancer. With regard to the function of circRNAs, previous studies have indicated that circRNAs may serve as miRNA sponges and RNA-binding protein (RBP) sponges and could regulate miRNA-mediated signaling or RBP-associated transcription, and influence the function of certain types of cell (11,13,14,16,19).
The present study reviews circRNAs in cancer, summarizes the vital functions of circRNAs in different types of cancer, highlights the competing endogenous (ce)RNA function of circRNAs in regulating downstream gene expression, and provides evidence for the application of circRNAs in the future diagnosis and treatment of numerous types of cancer.
2. Landscape of circRNAs
In recent years, circRNAs have emerged as a novel class of functional molecules among lncRNAs (8,10,15,20). Differing from the linear RNAs with the typical termini consisting of 5′ caps and 3′ tails, all known circRNAs display covalently closed loop structures, without any 5′ to 3′ polarity or polyadenylation at the 3′ ends (21,22), leading to mass escape of circRNAs from general transcriptomic polyadenylated RNA profiling (23,24). However, RNA sequencing technology and deep bioinformatics analyses have previously revealed numerous novel circRNAs in different diseases and cell lines (11,12,15,18,25–27).
Origins of circRNAs
circRNAs may be found in serum, saliva and the exosomes of mammalian cells. Using a genome-wide approach, circRNAs were first detected in excised exons or introns (28) and were generally classified into three subtypes: Exonic circRNAs (ecircRNAs) (29), circular intronic RNAs (ciRNAs) (30), and exon-intron circRNAs (EIciRNAs) (31). Cytoplasmic ecircRNAs account for >90% of total circRNAs (11,13,21), whereas nuclear circRNAs are mainly ciRNAs and EIciRNAs (30,31), and exosomal circRNAs (exo-circRNAs), another type of circRNA, are distributed in the exosomes of human serum and in multiple human cancer cell lines (32).
To date, circRNAs have mainly been reported to form via back-splicing (22), which is characterized by the covalent binding of a 5′ splicing acceptor to a 3′ splicing donor (15). Jeck et al (13) proposed two models to assess the production of circRNAs; model one is referred to as ‘direct back-splicing’, while model two is termed ‘exon skipping’ or ‘lariat intermediate’. Generally, alternative circularization is generated from different numbers of exons (13,22,25,33), with or without an internal intron (14,22,31). Nevertheless, it remains unclear whether alternative circularization occurs co-transcriptionally or post-transcriptionally, and what factors affect circularization.
Characteristics of circRNAs
circRNAs exhibit specific characteristics distinct from linear RNAs. Firstly, circRNAs have particular covalently closed loop structures, rather than the typical 5′ caps and 3′ tails at the termini of linear RNAs (21,22), which render circRNAs more stable compared with their linear counterparts and more resistant to degradation by various endogenous RNA exonucleases (34,35). Additionally, certain circRNAs are widespread and more abundant compared with their linear counterparts (13,36). A previous study revealed that certain circRNAs were expressed in a cell type-specific manner; for example, hsa_circRNA_21 was only detected in cluster of differentiation (CD)19+ leukocytes and not in CD34+ leukocytes or neutrophils (11). The majority of circRNAs are evolutionarily conserved among different species (13,14,37), with the exception of intergenic or intronic circRNAs (30).
3. Association between circRNAs and multiple types of cancer
Since the function of one circRNA molecule was first proposed (11), the potential of circRNAs has invoked widespread interest (38). Numerous circRNAs are differentially expressed between cancerous and normal tissues, including in colorectal cancer (CRC), gastric cancer (GC), hepatocellular carcinoma (HCC), breast cancer, and bladder urothelial carcinoma.
One example is homeodomain-interacting protein kinase (HIPK)3 circRNA (circ-HIPK3), which is derived from exon 2 of the HIPK3 gene, and demonstrates significant differential expression between cancerous and normal tissues (39).
Yang et al (40) revealed that the expression of forkhead box O3 (FOXO3), one of the members of the forkhead family of transcription factors, may be regulated by FOXO3 circRNA (circ-FOXO3) and FOXO3 pseudogene (FOXO3P). The aberrant expression of FOXO3 mRNA, circ-FOXO3 and FOXO3P may synergistically affect tumor growth and survival (40). Additionally, Du et al (41,42) demonstrated that circ-FOXO3 may retard cell cycle progression and induce cell apoptosis.
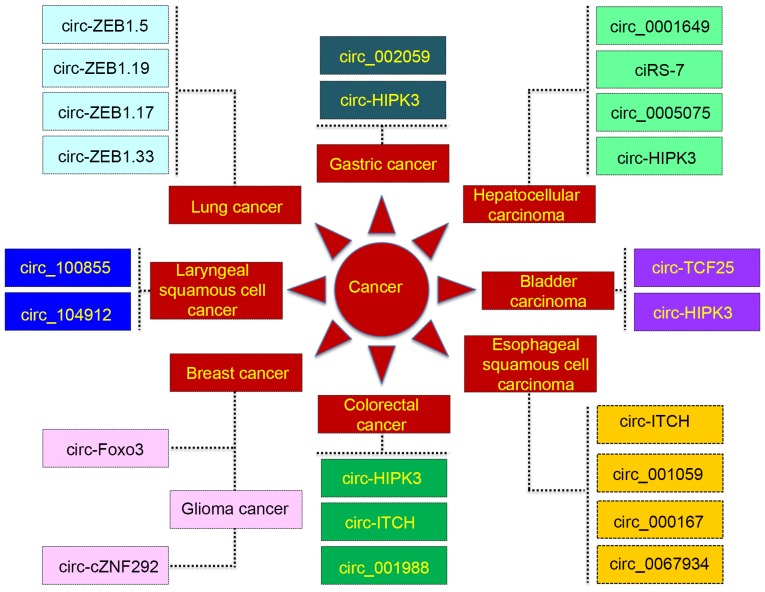
Li et al (32) reported that abundant exo-circRNAs were markedly enriched in the exosomes derived from multiple cancer cell lines, including lung, breast, colon, cervical and stomach cancer, and the detected exo-circRNAs derived from human serum further suggested their potential as novel diagnostic or prognostic markers for multiple types of cancer. Although the function of circRNAs in tumorigenesis is limited, studies explored in the present review have indicated that certain circRNAs may be associated with the development of various types of cancer (Fig. 1).
Figure 1.
Association between circular RNAs and multiple types of cancer. Circular RNAs associated with certain genes as well as circular RNAs that are yet to be formally named were included. ciRS-7 was named by the individual who first discovered it. circ, circular RNA; ZEB, zinc finger E-box binding homeobox; HIPK3, homeodomain interacting protein kinase 3; TCF25, transcription factor 25; ITCH, itchy E3 ubiquitin protein ligase; Foxo3, forkhead box O3; ZNF292, zinc finger protein 292.
circRNAs and CRC
In a previous study on CRC, 39 differentially expressed circRNAs were identified between CRC and matched normal colonic mucosa tissues (43). In addition, the ratio of circRNAs to linear RNA counterparts was decreased in CRC tissues compared with that in normal colon tissues, and even further decreased in CRC cell lines compared with that in normal colon tissues. Furthermore, there was a negative correlation between this ratio and the proliferation index, which provided the basis to explore their functions and mechanisms in CRC. In addition, the authors hypothesized that their findings were due to the dysfunction of back-splicing or dysregulation of oncogenic miRNAs, resulting in the degradation and downregulation of circRNAs in CRC (43).
Similarly, itchy E3 ubiquitin protein ligase (ITCH) circRNA (circ-ITCH) expression was revealed to be significantly downregulated in CRC tissues compared with that in peritumoral tissues, and circ-ITCH could upregulate the expression of its linear counterpart, ITCH, and thereafter affect the Wnt/β-catenin signaling pathway in CRC (44). Additionally, decreased expression of hsa_circ_001988 was identified in CRC, and the expression of hsa_circ_001988 was significantly correlated with differentiation and perineural invasion of CRC cells (45). Dou et al (46) also revealed that numerous circRNAs are downregulated in KRAS proto-oncogene mutant colon cancer cells and may be transferred to exosomes, and that circRNAs were more abundant in exosomes compared with cells.
circRNAs and HCC
Qin et al (47) revealed that hsa_circ_0001649 expression was significantly decreased in HCC and may represent a novel biomarker for HCC. Xu et al (48) demonstrated that, although the expression of CDR1 antisense RNA (CDR1-AS; also known as ciRS-7) was comparable between HCC and matched non-tumor tissues, the increased ciRS-7 expression in HCC tissues was significantly correlated with hepatic microvascular invasion and therefore associated with the deterioration of HCC. Furthermore, circRNA profiling revealed that hsa_circ_0005075, a circRNA biomarker, exhibited a significant difference in expression between HCC and normal tissues, and a pathological correlation analysis indicated that hsa_circ_0005075 was associated with the development of HCC (49).
circRNAs and esophageal squamous cell carcinoma (ESCC)
Li et al (50) demonstrated that the expression of circ-ITCH was downregulated in ESCC. Su et al (51) also performed expression profiling of circRNAs in radioresistant ESCC cells, and revealed that circRNA_001059 and circRNA_000167 were associated with the development of radiation resistance. In addition, Xia et al (52) demonstrated that, hsa_circ_0067934 expression was significantly upregulated in ESCC and could promote the proliferation of ESCC cells.
circRNAs and lung cancer
Expression patterns in CircNet revealed that expression of the circRNAs circ-zinc finger E-box binding homeobox (ZEB)1.5, circ-ZEB1.19, circ-ZEB1.17 and circ-ZEB1.33 were downregulated in lung cancer specimens compared with that in normal lung tissues (53). Another study demonstrated that the expression of circ-ITCH was significantly decreased in lung cancer tissue compared with normal lung tissue (54).
circRNAs and other types of cancer
By microarray analysis, Qu et al (55) identified that numerous circRNAs were dysregulated in pancreatic ductal adenocarcinoma (PDAC), and these results implied their vital functions in the progression of PDAC.
Hsa_circ_002059, a representative circRNA, was downregulated in GC and its differential expression was significantly correlated with distal metastasis and tumor-node-metastasis stage (56), suggesting it could potentially serve as a diagnostic biomarker for GC.
Zhong et al (57) screened differential circRNA expression profiles in bladder carcinoma and revealed that the expression of family with sequence similarity 169 member A circRNA and tripartite motif-containing 24 circRNA were significantly downregulated in bladder carcinoma, while the expression levels of transcription factor (TCF)25 circRNA (circ-TCF25), zinc finger RNA-binding protein circRNA, protein tyrosine kinase 2 circRNA and BC048201 circRNA were significantly increased in carcinoma tissues. This suggested that circ-TCF25 could represent a novel biomarker for bladder cancer.
Ahmed et al (58) revealed that altered expression patterns of circRNAs were present in primary and metastatic sites of epithelial ovarian carcinoma. In addition, circRNA expression was inversely associated with linear RNA expression for numerous cancer-associated signaling pathways; the majority of these signaling pathways, such as the transforming growth factor β1 signaling pathway, were typically activated by linear RNA (mRNA) expression in metastases compared to primary site of origin, but repressed by circRNA expression (58).
Xuan et al (59) demonstrated that hsa_circ_100855 expression was significantly increased in laryngeal squamous cell cancer tissues compared with that in matched adjacent non-neoplastic tissues, and increased hsa_circ_100855 expression was revealed in patients with T3-4 stage disease, neck nodal metastasis, or advanced clinical stage. However, hsa_circ_104912 expression exhibited the reverse trend compared with hsa_circ_100855 (59).
In breast cancer, circRNA expression profiling results revealed that normal-adjacent tissues in patients with the estrogen receptor (ER)+ subtype were associated with an increased number of circRNAs compared with that in breast cancer tissues, and the number of circRNAs in normal-adjacent samples of ER+ subtype was inversely correlated with the risk-of-relapse proliferation score for proliferation-associated genes (60). Yang et al (61) revealed that zinc finger protein 292 circRNA could suppress human glioma tube formation via the Wnt/β-catenin signaling pathway and may potentially be applied as a therapeutic target and biomarker in glioma.
However, the association between certain circRNA and tumorigenesis is unclear and supported by sufficient evidence. Further studies will provide a more extensive awareness of circRNAs in different types of cancer, including their properties, functions and mechanisms.
4. Function of circRNAs as competing endogenous RNAs in cancer
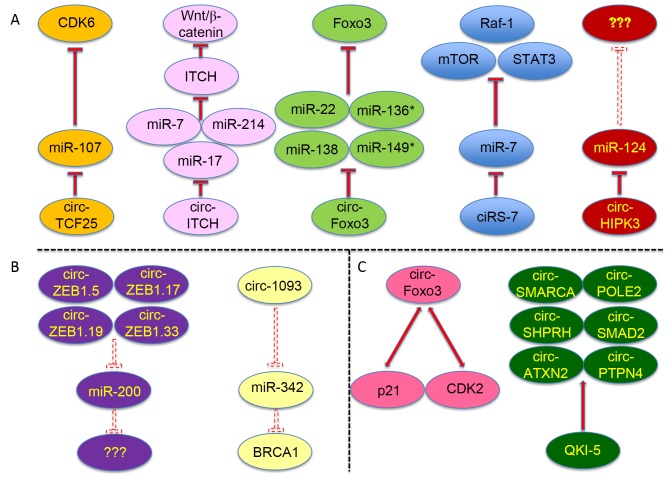
Currently, studies on the function and mechanism of circRNAs are few. Since circRNAs may be considered alternative splicing isoforms, they may regulate alternative splicing in certain diseases, including cancer (30,32). Certain circRNAs, including sirtuin (SIRT)7 circRNA and minichromosome maintenance complex component (MCM)5 circRNA, may enhance the expression of their parental mRNAs, while the parental linear mRNAs may be associated with tumorigenesis, functioning as a tumor suppressor (e.g. SIRT7) or promoter (e.g. MCM5) (30,32,62). circRNAs mainly function as ceRNAs in cancer by competing with other RNAs to bind miRNAs and RBPs (Fig. 2A-C).
Figure 2.
Circular RNAs function as competing endogenous RNAs in multiple types of cancer. The specific microRNAs and RNA-binding proteins sponged by certain circular RNAs are shown, as well as the protein targets of certain sponged miRNAs. (A) Specific and validated microRNAs sponged by certain circular RNAs, as well as the protein targets of certain sponged miRNAs. (B) Predicted microRNAs sponged by certain circular RNAs. (C) Validated and specific RNA-binding proteins sponged by certain circular RNAs. The dashed inhibition arrows represent predicted but not validated inhibitory effects, and the solid inhibition arrows represent that the inhibitory effects have been experimentally validated. ‘???’ denotes that the targeted genes of the microRNAs were not addressed in the corresponding reference. circ, circular RNA; miR, microRNA; CDK, cyclin-dependent kinase; TCF25, transcription factor 25; ITCH, itchy E3 ubiquitin protein ligase; Foxo3, forkhead box O3; Raf-1, Raf-1 proto-oncogene; mTOR, mechanistic target of rapamycin; STAT3, signal transducer and activator of transcription 3; HIPK3, homeodomain-interacting protein kinase 3; ZEB, zinc finger E-box binding homeobox; SMARCA, SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin, subfamily A; POLE2, DNA polymerase ε2, accessory subunit; SHPRH, SNF2 histone linker PHD RING helicase; SMAD2, SMAD family member 2; ATXN2, ataxin 2; PTPN4, protein tyrosine phosphatase, non-receptor type 4; QKI-5, Quaking 5.
circRNAs as miRNA sponges in cancer
miRNAs, a large class of sncRNAs, serve important functions in multiple types of cancer by repressing downstream cancer-associated mRNAs (63). Due to the complementarity between circRNAs and targeted genes, circRNAs may bind miRNAs and competitively suppress the function of targeted miRNAs, thereby permitting circRNAs to regulate the progression of cancer by sequestering specific miRNAs associated with proliferation, differentiation, metastasis and carcinogenesis.
Human ciRS-7/CDR1-AS is a typical example of a circRNA that functions as an miRNA sponge, and comprises as many as 74 miR-7-binding sites (11,19,62). miR-7 may serve as a tumor suppressor, and is downregulated in various types of cancer, including GC (64), HCC (65) and CRC (66). miR-7 is involved in numerous cancer-associated signaling pathways by downregulating the expression of crucial oncogenic factors, including Raf-1 proto-oncogene (67), mechanistic target of rapamycin (65) and signal transducer and activator of transcription 3 (68). Therefore, ciRS-7 could regulate cancer progression through miR-7-mediated downstream signaling pathways. However, other reports have revealed a tumor-promoting function of miR-7 in CRC (69), human papilloma virus-positive HeLa cells (70) and A549 cells (71). miR-7 is coupled to ciRS-7 through multiple miRNA response elements in ciRS-7, and ciRS-7 may either promote cancer progression or suppress carcinogenesis depending on the expression of miRNA targets (11,19,62).
Li et al (50) demonstrated that circ-ITCH may serve a tumor-suppressive function in ESCC and serve as a sponge for miR-7, miR-17 and miR-214, resulting in increased ITCH expression, thereby promoting ubiquitin-mediated degradation of disheveled segment polarity protein 2, and inhibiting the Wnt/β-catenin signaling pathway. This mechanism for circ-ITCH was also verified in lung cancer (54). Similarly, Yang et al (40) revealed that circ-FOXO3 may also function as a sponge of miRNAs to increase FOXO3 translation and thereby suppress tumor growth, cancer cell proliferation and survival. In addition, Zhong et al (57) demonstrated in bladder cancer that the overexpression of circ-TCF25 could decrease the expression of miR-103a-3p and miR-107, increase the expression of cyclin-dependent kinase (CDK)6 and promote the proliferation and migration of cancer cells in vitro and in vivo. Zheng et al (39) also revealed that circ-HIPK3 sponged 9 miRNAs and directly bound miR-124 and inhibited its activity.
Although multiple circRNAs have been demonstrated to regulate cancer progression in vitro or in vivo (53,17,72), certain predicted circRNAs remain to be validated. For example, Liu et al (53) proposed that circ-ZEB1.5, circ-ZEB1.19, circ-ZEB1.17 and circ-ZEB1.33 possess the potential to sponge miR-200. Zhang et al (17) revealed that circRNA_1093, with a short sequence of 108 bp, has 4 potential miR-342 binding sites, while miR-342 has been reported to regulate the expression of BRCA1 in breast cancer (72). These circRNAs require further study to confirm their function and molecular mechanisms. Certainly, the circRNAs that function as miRNA sponges provide a novel direction for diagnosing and treating multiple types of cancer and the circRNA-miRNA-gene regulatory networks in cancer require further study.
circRNAs as RBP sponges in cancer
circRNAs have also been associated with another important function: Acting as RBP sponges to regulate downstream gene transcription (12). Multiple studies have indicated that certain circRNAs may be associated with RBPs, such as argonaute (AGO) proteins (11,19), Quaking (QKI) (73), RNA polymerase II (30), mannose-binding lectin 2 (74) and eukaryotic translation initiation factor 4A3 (75). These circRNAs can serve as carriers to store, sort or deliver RBPs to particular subcellular locations, or regulate the function of RBPs by functioning as ceRNAs (11,76). Certain RBPs serve key regulatory functions in cell proliferation, differentiation, apoptosis and migration (77,78). Their dysregulated expression may affect cancer development (79,80). For example, AGO proteins may regulate tumorigenesis via miRNA-dependent or independent signaling pathways and AGO2 was shown to be associated with cancer progression by interacting with RISC-loading complex subunit TARBP2 and endoribonuclease Dicer (81).
As an RBP, QKI-5 has been characterized as a novel tumor suppressor in multiple types of cancer. For example, QKI-5 has been demonstrated to inhibit the proliferation of lung cancer cells (82), and the proliferation, cell cycle progression and invasion of prostate cancer cells (83). Furthermore, QKI-5 was shown to regulate the formation of circRNAs, and the upregulation of the majority of circRNAs during the epithelial-mesenchymal transition (EMT) suggests they may serve a crucial function in EMT (73). Since EMT serves an important function in initiating metastasis (84,85), further study is required to elucidate the function of QKI-mediated circRNAs in cancer. In addition, Du et al (41) also revealed that circ-FOXO3 could decelerate cell cycle progression by forming ternary complexes with p21 and CDK2 proteins.
These studies suggest that RBPs and circRNAs could interact reciprocally to affect cancer progression.
5. Prospects and challenges
Increasing numbers of circRNAs are being identified, and certain circRNAs have been revealed to be abundant in clinical blood samples (86), saliva (35) and exosomes (32). Considering their covalently closed loop structures, stability and cell/tissue type-specific manner (21,22,34,35,11,13,14,37), circRNAs are stable, sensitive, specific and are easily detectable. Detecting circRNAs early in samples of bodily fluids may render circRNAs promising diagnostic biomarkers for different types of cancer.
Currently, the concept of ceRNAs as regulators is novel (87,88) and controversial (89). However, the stability and other particular properties of circRNAs may confer ceRNA functions with additional advantages for effective binding with miRNA/RBP, exhibiting long-term effects. Given the important functions of circRNAs as miRNA and RBP sponges, the functional potential of circRNAs could be developed for cancer therapy. For example, an artificial circRNA (90,91) functioning as a ‘super-sponge’ or silencing the circRNA in cancer cells could regulate the expression of miRNAs and other RNAs or RBPs and increase the activities of suppressor genes or decrease the activities of oncogenes. Although synthetic miRNA inhibitors for treating human diseases have already been clinically trialed in phase 2a (92), small RNA-based drugs, including those based on miRNAs, circRNAs and piRNAs, are associated with different challenges due to the lack of targeted delivery strategies in cancer therapy (93). However, delivery systems mediated by nano materials or microbodies (including exosomes) are providing promising treatment directions (32).
Presently, multiple circRNAs are undergoing initial exploration and the molecular mechanisms underlying the functions of circRNAs in cancer progression remain to be fully understood. The strategies for assessing circRNAs remain limited and have developmental potential (13,15). High-throughput RNA sequencing technology combined with bioinformatics analysis may be applied to predict circRNAs, and multiple circRNA-associated databases have been developed for public use, including circRNAFinder (18), Circ2Traits (94), find_circ (11), CIRCexplorer (22), CIRI (95), starBase (96), CircNet (53) and circBase (97). Identifying more cancer-associated circRNAs and evaluating their underlying mechanisms may assist healthcare professionals in clinically applying circRNAs in cancer diagnosis and therapy.
Acknowledgements
The present study was supported by the International Cooperation Key Project of the National Natural Science Foundation of China (grant no. 81520108031), the National Natural Science Foundation of China (grant nos. 81303102, 81473478, 81473628, 81573749 and 81573805), the Key Project of Shanghai Municipal Science and Technology Commission (grant no. 16401970500), the Three Year Plan of Action for Innovation of Traditional Chinese Medicine in Shanghai (grant no. ZY3-CCCX-3-3012), the Program of Shanghai Academic Research Leader (grant no. 16XD1403600) and the Program for Shanghai Outstanding Medical Academic Leader.
References
- 1.Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, Parkin DM, Forman D, Bray F. Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359–E386. doi: 10.1002/ijc.29210. [DOI] [PubMed] [Google Scholar]
- 2.Dušek L, Mužík J, Malúšková D, Májek O, Pavlík T, Koptíková J, Gregor J, Brabec P, Abrahámová J. Epidemiology of screening-targeted cancers according to new data of the Czech National Cancer Registry. Klin Onkol. 2014;27(Suppl 2):S19–S39. doi: 10.14735/amko20142S19. [DOI] [PubMed] [Google Scholar]
- 3.Rosenberg AR, Kroon L, Chen L, Li CI, Jones B. Insurance status and risk of cancer mortality among adolescents and young adults. Cancer. 2015;121:1279–1286. doi: 10.1002/cncr.29187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.van der Weyden L, Arends MJ, Rust AG, Poulogiannis G, McIntyre RE, Adams DJ. Increased tumorigenesis associated with loss of the tumor suppressor gene Cadm1. Mol Cancer. 2012;11:29. doi: 10.1186/1476-4598-11-29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chan XH, Nama S, Gopal F, Rizk P, Ramasamy S, Sundaram G, Ow GS, Ivshina AV, Tanavde V, Haybaeck J, et al. Targeting glioma stem cells by functional inhibition of a prosurvival oncomiR-138 in malignant gliomas. Cell Rep. 2012;2:591–602. doi: 10.1016/j.celrep.2012.07.012. [DOI] [PubMed] [Google Scholar]
- 6.Shroff EH, Eberlin LS, Dang VM, Gouw AM, Gabay M, Adam SJ, Bellovin DI, Tran PT, Philbrick WM, Garcia-Ocana A, et al. MYC oncogene overexpression drives renal cell carcinoma in a mouse model through glutamine metabolism; Proc Natl Acad Sci USA; 2015; pp. 6539–6544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hashim A, Rizzo F, Marchese G, Ravo M, Tarallo R, Nassa G, Giurato G, Santamaria G, Cordella A, Cantarella C, Weisz A. RNA sequencing identifies specific PIWI-interacting small non-coding RNA expression patterns in breast cancer. Oncotarget. 2014;5:9901–9910. doi: 10.18632/oncotarget.2476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Sun QL, Zhao CP, Wang TY, Hao XB, Wang XY, Zhang X, Li YC. Expression profile analysis of long non-coding RNA associated with vincristine resistance in colon cancer cells by next-generation sequencing. Gene. 2015;572:79–86. doi: 10.1016/j.gene.2015.06.087. [DOI] [PubMed] [Google Scholar]
- 9.Brosnan CA, Voinnet O. The long and the short of noncoding RNAs. Curr Opin Cell Biol. 2009;21:416–425. doi: 10.1016/j.ceb.2009.04.001. [DOI] [PubMed] [Google Scholar]
- 10.Gibb EA, Brown CJ, Lam WL. The functional role of long non-coding RNA in human carcinomas. Mol Cancer. 2011;10:38. doi: 10.1186/1476-4598-10-38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer M, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495:333–338. doi: 10.1038/nature11928. [DOI] [PubMed] [Google Scholar]
- 12.Qu S, Yang X, Li X, Wang J, Gao Y, Shang R, Sun W, Dou K, Li H. Circular RNA: A new star of noncoding RNAs. Cancer Lett. 2015;365:141–148. doi: 10.1016/j.canlet.2015.06.003. [DOI] [PubMed] [Google Scholar]
- 13.Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, Marzluff WF, Sharpless NE. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19:141–157. doi: 10.1261/rna.035667.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Salzman J, Chen RE, Olsen MN, Wang PL, Brown PO. Cell-type specific features of circular RNA expression. PLoS Genet. 2013;9:e1003777. doi: 10.1371/annotation/f782282b-eefa-4c8d-985c-b1484e845855. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Guo JU, Agarwal V, Guo H, Bartel DP. Expanded identification and characterization of mammalian circular RNAs. Genome Biol. 2014;15:409. doi: 10.1186/s13059-014-0409-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Rybak-Wolf A, Stottmeister C, Glažar P, Jens M, Pino N, Giusti S, Hanan M, Behm M, Bartok O, Ashwal-Fluss R, et al. Circular RNAs in the mammalian brain are highly abundant, conserved, and dynamically expressed. Mol Cell. 2015;58:870–885. doi: 10.1016/j.molcel.2015.03.027. [DOI] [PubMed] [Google Scholar]
- 17.Zhang C, Wu H, Wang Y, Zhao Y, Fang X, Chen C, Chen H. Expression patterns of circular RNAs from primary kinase transcripts in the mammary glands of lactating rats. J Breast Cancer. 2015;18:235–241. doi: 10.4048/jbc.2015.18.3.235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Westholm JO, Miura P, Olson S, Shenker S, Joseph B, Sanfilippo P, Celniker SE, Graveley BR, Lai EC. Genome-wide analysis of drosophila circular RNAs reveals their structural and sequence properties and age-dependent neural accumulation. Cell Rep. 2014;9:1966–1980. doi: 10.1016/j.celrep.2014.10.062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, Kjems J. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495:384–388. doi: 10.1038/nature11993. [DOI] [PubMed] [Google Scholar]
- 20.Chen LL, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015;12:381–388. doi: 10.1080/15476286.2015.1020271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Jeck WR, Sharpless NE. Detecting and characterizing circular RNAs. Nat Biotechnol. 2014;32:453–461. doi: 10.1038/nbt.2890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Zhang XO, Wang HB, Zhang Y, Lu X, Chen LL, Yang L. Complementary sequence-mediated exon circularization. Cell. 2014;159:134–147. doi: 10.1016/j.cell.2014.09.001. [DOI] [PubMed] [Google Scholar]
- 23.Graveley BR. Molecular biology: Power sequencing. Nature. 2008;453:1197–1198. doi: 10.1038/4531197b. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wilhelm BT, Marguerat S, Watt S, Schubert F, Wood V, Goodhead I, Penkett CJ, Rogers J, Bähler J. Dynamic repertoire of a eukaryotic transcriptome surveyed at single-nucleotide resolution. Nature. 2008;453:1239–1243. doi: 10.1038/nature07002. [DOI] [PubMed] [Google Scholar]
- 25.Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One. 2012;7:e30733. doi: 10.1371/journal.pone.0030733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ivanov A, Memczak S, Wyler E, Torti F, Porath HT, Orejuela MR, Piechotta M, Levanon EY, Landthaler M, Dieterich C, Rajewsky N. Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell Rep. 2015;10:170–177. doi: 10.1016/j.celrep.2014.12.019. [DOI] [PubMed] [Google Scholar]
- 27.Danan M, Schwartz S, Edelheit S, Sorek R. Transcriptome-wide discovery of circular RNAs in Archaea. Nucleic Acids Res. 2012;40:3131–3142. doi: 10.1093/nar/gkr1009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Yang L, Duff MO, Graveley BR, Carmichael GG, Chen LL. Genomewide characterization of non-polyadenylated RNAs. Genome Biol. 2011;12:R16. doi: 10.1186/gb-2011-12-2-r16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kelly S, Greenman C, Cook PR, Papantonis A. Exon skipping is correlated with exon circularization. J Mol Biol. 2015;427:2414–2417. doi: 10.1016/j.jmb.2015.02.018. [DOI] [PubMed] [Google Scholar]
- 30.Zhang Y, Zhang XO, Chen T, Xiang JF, Yin QF, Xing YH, Zhu S, Yang L, Chen LL. Circular intronic long noncoding RNAs. Mol Cell. 2013;51:792–806. doi: 10.1016/j.molcel.2013.08.017. [DOI] [PubMed] [Google Scholar]
- 31.Li Z, Huang C, Bao C, Chen L, Lin M, Wang X, Zhong G, Yu B, Hu W, Dai L, et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. 2015;22:256–264. doi: 10.1038/nsmb.2959. [DOI] [PubMed] [Google Scholar]
- 32.Li Y, Zheng Q, Bao C, Li S, Guo W, Zhao J, Chen D, Gu J, He X, Huang S. Circular RNA is enriched and stable in exosomes: A promising biomarker for cancer diagnosis. Cell Res. 2015;25:981–984. doi: 10.1038/cr.2015.82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Burd CE, Jeck WR, Liu Y, Sanoff HK, Wang Z, Sharpless NE. Expression of linear and novel circular forms of an INK4/ARF-associated non-coding RNA correlates with atherosclerosis risk. PLoS Genet. 2010;6:e1001233. doi: 10.1371/journal.pgen.1001233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Suzuki H, Tsukahara T. A view of pre-mRNA splicing from RNase R resistant RNAs. Int J Mol Sci. 2014;15:9331–9342. doi: 10.3390/ijms15069331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Bahn JH, Zhang Q, Li F, Chan TM, Lin X, Kim Y, Wong DT, Xiao X. The landscape of microRNA, Piwi-interacting RNA, and circular RNA in human saliva. Clin Chem. 2015;61:221–230. doi: 10.1373/clinchem.2014.230433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Lasda E, Parker R. Circular RNAs: Diversity of form and function. RNA. 2014;20:1829–1842. doi: 10.1261/rna.047126.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wang PL, Bao Y, Yee MC, Barrett SP, Hogan GJ, Olsen MN, Dinneny JR, Brown PO, Salzman J. Circular RNA is expressed across the eukaryotic tree of life. PLoS One. 2014;9:e90859. doi: 10.1371/journal.pone.0090859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Perkel JM. Assume nothing: The tale of circular RNA. Biotechniques. 2013;55:55–57. doi: 10.2144/000114061. [DOI] [PubMed] [Google Scholar]
- 39.Zheng Q, Bao C, Guo W, Li S, Chen J, Chen B, Luo Y, Lyu D, Li Y, Shi G, et al. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat Commun. 2016;7:11215. doi: 10.1038/ncomms11215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Yang W, Du WW, Li X, Yee AJ, Yang BB. Foxo3 activity promoted by non-coding effects of circular RNA and Foxo3 pseudogene in the inhibition of tumor growth and angiogenesis. Oncogene. 2016;35:3919–3931. doi: 10.1038/onc.2015.460. [DOI] [PubMed] [Google Scholar]
- 41.Du WW, Yang W, Liu E, Yang Z, Dhaliwal P, Yang BB. Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res. 2016;44:2846–2858. doi: 10.1093/nar/gkw027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Du WW, Fang L, Yang W, Wu N, Awan FM, Yang Z, Yang BB. Induction of tumor apoptosis through a circular RNA enhancing Foxo3 activity. Cell Death Differ. 2017;24:357–370. doi: 10.1038/cdd.2016.133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Bachmayr-Heyda A, Reiner AT, Auer K, Sukhbaatar N, Aust S, Bachleitner-Hofmann T, Mesteri I, Grunt TW, Zeillinger R, Pils D. Correlation of circular RNA abundance with proliferation-Exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci Rep. 2015;5:8057. doi: 10.1038/srep08057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Huang G, Zhu H, Shi Y, Wu W, Cai H, Chen X. cir-ITCH plays an inhibitory role in colorectal cancer by regulating the Wnt/β-catenin pathway. PLoS One. 2015;10:e0131225. doi: 10.1371/journal.pone.0131225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Wang X, Zhang Y, Huang L, Zhang J, Pan F, Li B, Yan Y, Jia B, Liu H, Li S, Zheng W. Decreased expression of hsa_circ_001988 in colorectal cancer and its clinical significances. Int J Clin Exp Pathol. 2015;8:16020–16025. [PMC free article] [PubMed] [Google Scholar]
- 46.Dou Y, Cha DJ, Franklin JL, Higginbotham JN, Jeppesen DK, Weaver AM, Prasad N, Levy S, Coffey RJ, Patton JG, Zhang B. Circular RNAs are down-regulated in KRAS mutant colon cancer cells and can be transferred to exosomes. Sci Rep. 2016;6:37982. doi: 10.1038/srep37982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Qin M, Liu G, Huo X, Tao X, Sun X, Ge Z, Yang J, Fan J, Liu L, Qin W. Hsa_circ_0001649: A circular RNA and potential novel biomarker for hepatocellular carcinoma. Cancer Biomark. 2016;16:161–169. doi: 10.3233/CBM-150552. [DOI] [PubMed] [Google Scholar]
- 48.Xu L, Zhang M, Zheng X, Yi P, Lan C, Xu M. The circular RNA ciRS-7 (Cdr1as) acts as a risk factor of hepatic microvascular invasion in hepatocellular carcinoma. J Cancer Res Clin Oncol. 2017;143:17–27. doi: 10.1007/s00432-016-2256-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Shang X, Li G, Liu H, Li T, Liu J, Zhao Q, Wang C. Comprehensive circular RNA profiling reveals that hsa_circ_0005075, a new circular RNA biomarker, is involved in hepatocellular crcinoma development. Medicine (Baltimore) 2016;95:e3811. doi: 10.1097/MD.0000000000003811. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Li F, Zhang L, Li W, Deng J, Zheng J, An M, Lu J, Zhou Y. Circular RNA ITCH has inhibitory effect on ESCC by supupressing the Wnt/β-catenin pathway. Oncotarget. 2015;6:6001–6013. doi: 10.18632/oncotarget.3469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Su H, Lin F, Deng X, Shen L, Fang Y, Fei Z, Zhao L, Zhang X, Pan H, Xie D, et al. Profiling and bioinformatics analyses reveal differential circular RNA expression in radioresistant esophageal cancer cells. J Transl Med. 2016;14:225. doi: 10.1186/s12967-016-0977-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Xia W, Qiu M, Chen R, Wang S, Leng X, Wang J, Xu Y, Hu J, Dong G, Xu PL, Yin R. Circular RNA has_circ_0067934 is upregulated in esophageal squamous cell carcinoma and promoted proliferation. Sci Rep. 2016;6:35576. doi: 10.1038/srep35576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Liu YC, Li JR, Sun CH, Andrews E, Chao RF, Lin FM, Weng SL, Hsu SD, Huang CC, Cheng C, et al. CircNet: A database of circular RNAs derived from transcriptome sequencing data. Nucleic Acids Res. 2016;44:D209–D215. doi: 10.1093/nar/gkv940. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Wan L, Zhang L, Fan K, Cheng ZX, Sun QC, Wang JJ. Circular RNA-ITCH suppresses lung cancer proliferation via inhibiting the Wnt/β-catenin pathway. Biomed Res Int. 2016;2016:1579490. doi: 10.1155/2016/1579490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Qu S, Song W, Yang X, Wang J, Zhang R, Zhang Z, Zhang H, Li H. Microarray expression profile of circular RNAs in human pancreatic ductal adenocarcinoma. Genom Data. 2015;5:385–387. doi: 10.1016/j.gdata.2015.07.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Li P, Chen S, Chen H, Mo X, Li T, Shao Y, Xiao B, Guo J. Using circular RNA as a novel type of biomarker in the screening of gastric cancer. Clin Chim Acta. 2015;444:132–136. doi: 10.1016/j.cca.2015.02.018. [DOI] [PubMed] [Google Scholar]
- 57.Zhong Z, Lv M, Chen J. Screening differential circular RNA expression profiles reveals the regulatory role of circTCF25-miR-103a-3p/miR-107-CDK6 pathway in bladder carcinoma. Sci Rep. 2016;6:30919. doi: 10.1038/srep30919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Ahmed I, Karedath T, Andrews SS, Al-Azwani IK, Mohamoud YA, Querleu D, Rafii A, Malek JA. Altered expression pattern of circular RNAs in primary and metastatic sites of epithelial ovarian carcinoma. Oncotarget. 2016;7:36366–36381. doi: 10.18632/oncotarget.8917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Xuan L, Qu L, Zhou H, Wang P, Yu H, Wu T, Wang X, Li Q, Tian L, Liu M, Sun Y. Circular RNA: A novel biomarker for progressive laryngeal cancer. Am J Transl Res. 2016;8:932–939. [PMC free article] [PubMed] [Google Scholar]
- 60.Nair AA, Niu N, Tang X, Thompson KJ, Wang L, Kocher JP, Subramanian S, Kalari KR. Circular RNAs and their associations with breast cancer subtypes. Oncotarget. 2016;7:80967–80979. doi: 10.18632/oncotarget.13134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Yang P, Qiu Z, Jiang Y, Dong L, Yang W, Gu C, Li G, Zhu Y. Silencing of cZNF292 circular RNA suppresses human glioma tube formation via the Wnt/β-catenin signaling pathway. Oncotarget. 2016;7:63449–63455. doi: 10.18632/oncotarget.11523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Chen I, Chen CY, Chuang TJ. Biogenesis, identification, and function of exonic circular RNAs. Wiley Interdiscip Rev RNA. 2015;6:563–579. doi: 10.1002/wrna.1294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Romero-Cordoba SL, Salido-Guadarrama I, Rodriguez-Dorantes M, Hidalgo-Miranda A. miRNA biogenesis: Biological impact in the development of cancer. Cancer Biol Ther. 2014;15:1444–1455. doi: 10.4161/15384047.2014.955442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Kong D, Piao YS, Yamashita S, Oshima H, Oguma K, Fushida S, Fujimura T, Minamoto T, Seno H, Yamada Y, et al. Inflammation-induced repression of tumor suppressor miR-7 in gastric tumor cells. Oncogene. 2012;31:3949–3960. doi: 10.1038/onc.2011.558. [DOI] [PubMed] [Google Scholar]
- 65.Fang Y, Xue JL, Shen Q, Chen J, Tian L. MicroRNA-7 inhibits tumor growth and metastasis by targeting the phosphoinositide 3-kinase/Akt pathway in hepatocellular carcinoma. Hepatology. 2012;55:1852–1862. doi: 10.1002/hep.25576. [DOI] [PubMed] [Google Scholar]
- 66.Zhang N, Li X, Wu C, Dong Y, Cai M, Mok M, Wang H, Chen J, Ng SS, Chen M, et al. microRNA-7 is a novel inhibitor of YY1 contributing to colorectal tumorigenesis. Oncogene. 2013;32:5078–5088. doi: 10.1038/onc.2012.526. [DOI] [PubMed] [Google Scholar]
- 67.Webster RJ, Giles KM, Price KJ, Zhang PM, Mattick JS, Leedman PJ. Regulation of epidermal growth factor receptor signaling in human cancer cells by microRNA-7. J Biol Chem. 2009;284:5731–5741. doi: 10.1074/jbc.M804280200. [DOI] [PubMed] [Google Scholar]
- 68.Zhang H, Cai K, Wang J, Wang X, Cheng K, Shi F, Jiang L, Zhang Y, Dou J. MiR-7, inhibited indirectly by lincRNA HOTAIR, directly inhibits SETDB1 and reverses the EMT of breast cancer stem cells by downregulating the STAT3 pathway. Stem Cells. 2014;32:2858–2868. doi: 10.1002/stem.1795. [DOI] [PubMed] [Google Scholar]
- 69.Nakagawa Y, Akao Y, Taniguchi K, Kamatani A, Tahara T, Kamano T, Nakano N, Komura N, Ikuno H, Ohmori T, et al. Relationship between expression of onco-related miRNAs and the endoscopic appearance of colorectal tumors. Int J Mol Sci. 2015;16:1526–1543. doi: 10.3390/ijms16011526. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Honegger A, Schilling D, Bastian S, Sponagel J, Kuryshev V, Sültmann H, Scheffner M, Hoppe-Seyler K, Hoppe-Seyler F. Dependence of intracellular and exosomal microRNAs on Viral E6/E7 oncogene expression in HPV-positive tumor cells. PLoS Pathog. 2015;11:e1004712. doi: 10.1371/journal.ppat.1004712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Meza-Sosa KF, Pérez-García EI, Camacho-Concha N, López-Gutiérrez O, Pedraza-Alva G, Pérez-Martínez L. MiR-7 promotes epithelial cell transformation by targeting the tumor suppressor KLF4. PLoS One. 2014;9:e103987. doi: 10.1371/journal.pone.0103987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Crippa E, Lusa L, De Cecco L, Marchesi E, Calin GA, Radice P, Manoukian S, Peissel B, Daidone MG, Gariboldi M, Pierotti MA. miR-342 regulates BRCA1 expression through modulation of ID4 in breast cancer. PLoS One. 2014;9:e87039. doi: 10.1371/journal.pone.0087039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Conn SJ, Pillman KA, Toubia J, Conn VM, Salmanidis M, Phillips CA, Roslan S, Schreiber AW, Gregory PA, Goodall GJ. The RNA binding protein quaking regulates formation of circRNAs. Cell. 2015;160:1125–1134. doi: 10.1016/j.cell.2015.02.014. [DOI] [PubMed] [Google Scholar]
- 74.Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O, Hanan M, Evantal N, Memczak S, Rajewsky N, Kadener S. circRNA biogenesis competes with pre-mRNA splicing. Mol Cell. 2014;56:55–66. doi: 10.1016/j.molcel.2014.08.019. [DOI] [PubMed] [Google Scholar]
- 75.Dudekula DB, Panda AC, Grammatikakis I, De S, Abdelmohsen K, Gorospe M. CircInteractome: A web tool for exploring circular RNAs and their interacting proteins and microRNAs. RNA Biol. 2016;13:34–42. doi: 10.1080/15476286.2015.1128065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Hentze MW, Preiss T. Circular RNAs: Splicing's enigma variations. EMBO J. 2013;32:923–925. doi: 10.1038/emboj.2013.53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Glisovic T, Bachorik JL, Yong J, Dreyfuss G. RNA-binding proteins and post-transcriptional gene regulation. FEBS Lett. 2008;582:1977–1986. doi: 10.1016/j.febslet.2008.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Abdelmohsen K, Kuwano Y, Kim HH, Gorospe M. Posttranscriptional gene regulation by RNA-binding proteins during oxidative stress: Implications for cellular senescence. Biol Chem. 2008;389:243–255. doi: 10.1515/BC.2008.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Kim MY, Hur J, Jeong S. Emerging roles of RNA and RNA-binding protein network in cancer cells. BMB Rep. 2009;42:125–130. doi: 10.5483/BMBRep.2009.42.3.125. [DOI] [PubMed] [Google Scholar]
- 80.Calabretta S, Richard S. Emerging roles of disordered sequences in RNA-binding proteins. Trends Biochem Sci. 2015;40:662–672. doi: 10.1016/j.tibs.2015.08.012. [DOI] [PubMed] [Google Scholar]
- 81.Ye Z, Jin H, Qian Q. Argonaute 2: A novel rising star in cancer research. J Cancer. 2015;6:877–882. doi: 10.7150/jca.11735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Zong FY, Fu X, Wei WJ, Luo YG, Heiner M, Cao LJ, Fang Z, Fang R, Lu D, Ji H, Hui J. The RNA-binding protein QKI suppresses cancer-associated aberrant splicing. PLoS Genet. 2014;10:e1004289. doi: 10.1371/journal.pgen.1004289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Zhao Y, Zhang G, Wei M, Lu X, Fu H, Feng F, Wang S, Lu W, Wu N, Lu Z, Yuan J. The tumor suppressing effects of QKI-5 in prostate cancer: A novel diagnostic and prognostic protein. Cancer Biol Ther. 2014;15:108–118. doi: 10.4161/cbt.26722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Rokavec M, Öner MG, Li H, Jackstadt R, Jiang L, Lodygin D, Kaller M, Horst D, Ziegler PK, Schwitalla S, et al. IL-6R/STAT3/miR-34a feedback loop promotes EMT-mediated colorectal cancer invasion and metastasis. J Clin Invest. 2014;124:1853–1867. doi: 10.1172/JCI73531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Gujral TS, Chan M, Peshkin L, Sorger PK, Kirschner MW, MacBeath G. A noncanonical Frizzled2 pathway regulates epithelial-mesenchymal transition and metastasis. Cell. 2014;159:844–856. doi: 10.1016/j.cell.2014.10.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Memczak S, Papavasileiou P, Peters O, Rajewsky N. Identification and characterization of circular RNAs as a new class of putative biomarkers in human blood. PLoS One. 2015;10:e0141214. doi: 10.1371/journal.pone.0141214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Ebert MS, Sharp PA. Emerging roles for natural microRNA sponges. Curr Biol. 2010;20:R858–R861. doi: 10.1016/j.cub.2010.08.052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Cheng DL, Xiang YY, Ji LJ, Lu XJ. Competing endogenous RNA interplay in cancer: Mechanism, methodology, and perspectives. Tumour Biol. 2015;36:479–488. doi: 10.1007/s13277-015-3093-z. [DOI] [PubMed] [Google Scholar]
- 89.Denzler R, Agarwal V, Stefano J, Bartel DP, Stoffel M. Assessing the ceRNA hypothesis with quantitative measurements of miRNA and target abundance. Mol Cell. 2014;5:766–776. doi: 10.1016/j.molcel.2014.03.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Errichelli L, Modigliani S Dini, Laneve P, Colantoni A, Legnini I, Capauto D, Rosa A, De Santis R, Scarfò R, Peruzzi G, et al. FUS affects circular RNA expression in murine embryonic stem cell-derived motor neurons. Nat Commun. 2017;8:14741. doi: 10.1038/ncomms14741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Tatomer DC, Liang D, Wilusz JE. Inducible expression of eukaryotic circular RNAs from plasmids. Methods Mol Biol. 2017;1648:143–154. doi: 10.1007/978-1-4939-7204-3_11. [DOI] [PubMed] [Google Scholar]
- 92.Janssen HL, Reesink HW, Lawitz EJ, Zeuzem S, Rodriguez-Torres M, Patel K, van der Meer AJ, Patick AK, Chen A, Zhou Y, et al. Treatment of HCV infection by targeting microRNA. N Engl J Med. 2013;368:1685–1694. doi: 10.1056/NEJMoa1209026. [DOI] [PubMed] [Google Scholar]
- 93.Gong Z, Yang J, Li J, Yang L, Le Y, Wang S, Lin HK. Novel insights into the role of microRNA in lung cancer resistance to treatment and targeted therapy. Curr Cancer Drug Targets. 2014;14:241–258. doi: 10.2174/1568009614666140305104845. [DOI] [PubMed] [Google Scholar]
- 94.Ghosal S, Das S, Sen R, Basak P, Chakrabarti J. Circ2Traits: A comprehensive database for circular RNA potentially associated with disease and traits. Front Genet. 2013;4:283. doi: 10.3389/fgene.2013.00283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Gao Y, Wang J, Zhao F. CIRI: An efficient and unbiased algorithm for de novo circular RNA identification. Genome Biol. 2015;16:4. doi: 10.1186/s13059-014-0571-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Li JH, Liu S, Zhou H, Qu LH, Yang JH. starBase v2.0: Decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014;42(Database Issue):D92–D97. doi: 10.1093/nar/gkt1248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Glažar P, Papavasileiou P, Rajewsky N. circBase: A database for circular RNAs. RNA. 2014;20:1666–1670. doi: 10.1261/rna.043687.113. [DOI] [PMC free article] [PubMed] [Google Scholar]