Abstract
The cell and tissue engineering fields have profited immensely through the implementation of highly-structured biomaterials. The development and implementation of advanced biofabrication techniques has established new avenues for generating biomimetic scaffolds for a multitude of cell and tissue engineering applications. Among these, laser-based degradation of biomaterials has been implemented to achieve user-directed features and functionalities within biomimetic scaffolds. This review offers an overview of the physical mechanisms that govern laser-material interactions and specifically, laser-hydrogel interactions. The influence of both laser and material properties on efficient, high-resolution hydrogel degradation are discussed and the current application space in cell and tissue engineering is reviewed. This review aims to acquaint readers with the capability and uses of laser-based degradation of biomaterials, so that it may be easily and widely adopted.
Keywords: biofabrication, biomaterial, microfluidic, neuronal guidance, cell migration, biomimetic, microphysiological systems
1. Introduction
The rational design, development, and implementation of highly structured biomaterials for cell and tissue engineering applications has gained tremendous interest in the past few decades. Advances in biofabrication techniques have enabled chemical and physical manipulation of biomaterials across a broad size range, from subcellular to the tissue level, to spatially tailor biomaterial properties for desired applications.[1,2] The implementation of molecular design and materials science principles have enabled nano- to macro-scale engineering of cell-material interfaces, tissue constructs, and acellular platforms.[3,4] Engineered cell-matrix and cell-cell interactions, and hierarchical modulation of mechanical/structural properties of materials have facilitated recapitulation of tissue level features and function.[5–7] Key considerations in the fabrication of tissue-engineered platforms include vascularization for adequate transport of oxygen and metabolites, control over matrix architecture via regulation of porosity and mechanical properties, presentation of biological signaling motifs, and microenvironmental control of cell-cell and cell-matrix interactions.[8,9] A common goal of these engineered tissue constructs is to spatially influence the function of several cell types, often with the ability to generate constructs with heterogeneous properties and in many cases, with temporal control over these properties.[10,11]
A multitude of biofabrication techniques have been developed and implemented to generate biomimetic, tissue-engineered constructs including three-dimensional (3D) printing,[12,13] stereolithography,[1] multiphoton lithography,[1,14] direct-write assembly,[15] sacrificial micromolding,[16,17] emulsification,[18,19] and microfluidic chip-based fabrication[20,21] amongst others. These techniques provide direct control over a broad range of parameters including microarchitecture, matrix mechanical properties, bioactive ligand presentation, porosity, and overall scalability that ultimately influence cell viability, metabolic activity, and tissue-level function. While many advanced fabrication techniques exist, each approach has its own advantages and limitations that restrict their universal implementation for all applications.
Laser-based degradation of hydrogels, biomaterials, and other soft interfaces has started gaining attention as a technique for obtaining high-resolution control over user-defined architectures within 3D tissue constructs and acellular platforms (Fig. 1). Laser-based degradation and similar techniques (direct-writing, ablation, micromachining) have been implemented for topographical patterning of two-dimensional (2D) interfaces,[22,23] modification of material surface properties,[24,25] and for laser-based surgery.[26,27] Laser-based degradation has facilitated fabrication of biomaterials for many emerging applications including engineered cellular microenvironments,[28,29] regulating cell-matrix interactions,[30,31] structuring tissue scaffolds,[32,33] fabrication of hydrogel-embedded microfluidic networks,[34–36] and drug delivery.[37] A wide range of both synthetic and natural materials including glass,[23] poly(ethylene terephthalate) (PET),[38] poly(methyl methacrylate) (PMMA),[39] polyimide,[39] poly(dimethyl sulfoxide) (PDMS),[40] poly(ethylene glycol) (PEG),[34,35] collagen,[41] agar/agarose,[22,42,43] hyaluronic acid[44] and silk[29,45] are amenable to manipulation via laser-based degradation thereby establishing this technique as suitable for a broad range of applications from fundamental cell studies to fabrication of vascularized, tissue-engineered constructs (Fig. 1B–D).[46,47]
Figure 1. Overview of laser-based degradation in literature.
(A) There has been a steady increase in the number of publications involving laser-based degradation of hydrogels and biomaterials over the past two decades. (B) Trends in the use of different biomaterials manipulated via laser-based degradation. (C) Categorization of publications based on biomaterial type and (D) application. The graphs represent (A, B) the number of publications or (C, D) percentages, that contain the terms ‘laser degradation’, ‘laser structuring’, ‘laser micropatterning’, ‘laser ablation’, ‘hydrogels’, and ‘scaffolds’ from searches using NCBI Pubmed, Google Scholar, and the ISI Web of Knowledge.
In this two-part review, we focus on laser-based degradation of polymeric hydrogels commonly used in cell and tissue engineering applications. In the first part, we introduce and discuss laser features and the mechanisms that govern laser-based hydrogel degradation. To provide readers with a basic understanding of achievable resolution and degradation efficiency, we divide the most commonly used hydrogels into three broad categories based on the dominant degradation mechanism at play: (1) synthetic hydrogels with low two-photon absorption, (2) protein-based hydrogels with high two-photon absorption, and (3) hydrogels containing engineered photolabile groups. In the second part, we review recent applications of laser-based hydrogel degradation in cell and tissue engineering applications and discuss the technical limitations and potential applications of this fabrication technique. Overall, this review aims to acquaint readers with the capability, limitations, and potential uses of laser-based degradation of biomaterials, so that it may be easily and widely adopted for these applications.
PART 1: Fundamentals of Laser-Based Hydrogel Degradation
2. Physical Principles and Mechanisms of Laser-Based Hydrogel Degradation
To implement laser-based hydrogel degradation, it is useful to possess knowledge of the fundamental laser-material interactions that occur, as well as the various degradation mechanisms that exist, as both influence the process efficiency and achievable resolution. Unfortunately, thorough experimentation independently investigating the influences of each laser parameter and characteristics of soft polymers/hydrogels does not currently exist. To cope with this lack of experimental data, we discuss the theoretical aspects of laser-hydrogel interactions that have been determined in the context of laser-tissue interactions for laser-based surgery.[48–50] Since hydrated tissue is analogous to polymeric hydrogels in many ways, the degradation mechanisms that govern laser surgery can be helpful in understanding hydrogel degradation. We describe the characteristics of common lasers used for hydrogel degradation, the various modes of degradation that dominate based on laser properties and hydrogel composition, and the parameters that govern resolution and degradation efficiency. Equipped with this knowledge, readers will have the necessary background to successfully implement laser-based hydrogel degradation for desired applications.
2.1 Considerations for Laser Sources and Optical Systems
To implement laser-based hydrogel degradation for biofabrication, it is important to have a working knowledge of the laser operating parameters including the laser power, intensity, fluence, scan speed, repetition rate, and focal volume defined by the objective lens used for focusing. This information is necessary for choosing the appropriate laser source to efficiently degrade features with a desired resolution in different hydrogel compositions. Optimization and tuning of both the laser parameters and hydrogel composition allows users to efficiently degrade desired features, while also reducing the time and labor involved. Accordingly, some of the key parameters to keep in mind when comparing and selecting laser sources are discussed below.
Lasers can be classified into two general categories, continuous wave and pulsed, the main difference being the time scale over which energy is delivered to the focal volume. Continuous wave lasers output a constant optical power with minimal to no deviation from the average power whereas pulsed lasers deliver energy in discrete, high power pulses at a constant frequency or pulse rate with instantaneous power that deviates substantially from the average value. Continuous wave lasers are easily characterized by directly measuring the power output (P: Watts (W)) or the intensity of the laser beam (I: W.cm−2) (power output per unit cross-sectional area). For pulsed lasers, these power and intensity measurements are often temporal averages (PAVG and IAVG) (owing to discretization of energy delivery), which are useful in comparing lasers but do not provide information concerning the way energy is delivered. Hence, the pulse duration (t: s) and pulse frequency (f: Hz) must be considered as these properties influence the peak power and intensity achieved which subsequently influences the mechanisms dominating material degradation.
Pulse durations can vary from ultrashort pulses (femtoseconds, 10−15 s) to longer pulses (milliseconds, 10−3 s). Similarly, the pulse frequency can vary across a wide range from high frequency (MHz pulses, 106 pulses.s−1) to low frequency (kHz pulses, 103 pulses.s−1). The peak power and peak intensity achieved during each pulse can be orders of magnitude higher than the average power and average intensity measured and the peak values depend on both the pulse duration and frequency (Fig. 2). To account for this, the peak power (PPEAK: W), peak intensity (IPEAK: W.cm−2), and fluence (F: J.cm−2) are used to characterize pulsed lasers. Based on the assumption of a square-wave pulse, these metrics can be calculated from the measured average power using the following equations:
| (1) |
Figure 2. Theoretical power and intensity outputs for three different lasers over time as a function of pulse duration and pulse frequency.
(A) Comparison of the power and intensity over time for three different lasers (continuous wave, 1 ns pulsed, and 140 fs pulsed) operating at the same pulse frequency for the pulsed lasers (80 MHz), same average power (1 W), and same average intensity (1.34×108 W.cm−2) but with different pulse durations, 1 ns and 140 fs for the pulsed lasers. The fs pulsed laser outputs a higher peak power and intensity compared to the ns pulsed or continuous wave lasers to achieve the same average power and intensity. (B) Comparison of the power and intensity output over time for three different lasers (continuous wave, low frequency (8 MHz) fs pulsed, and high frequency (80 MHz) fs pulsed) operating at the same average power (1 W) and same average intensity (1.34×108 W.cm−2) with the same pulse duration for the pulsed lasers (140 fs) but at different frequencies. The laser operating at the lower frequency (8 MHz) outputs a higher peak power and intensity to achieve the same average output compared to the higher frequency pulse and continuous wave lasers. (IP: peak intensity, PP: peak power, IAVG: average intensity, PAVG: average power).
Where E is the pulse energy (energy delivered per pulse: J), PAVG is the measured average power, and f is the pulse frequency. The pulse energy, E, is also described by the following equation:
| (2) |
Where PPEAK is the peak power achieved during each pulse and t is the pulse duration. From the above relations, the peak power (PPEAK) can be determined as:
| (3) |
Thus, the peak power for each pulse can be calculated by measuring the average power and knowing the pulse frequency (f) and pulse duration (t).
Once peak power is obtained, the peak intensity (IPEAK) can be calculated using the area of the focused beam (AFB: cm2) from the following equations:
| (4) |
The area of the focused beam can be estimated from the Airy disk:[48]
| (5) |
| (6) |
where d is the diameter of the focused beam, λ is the wavelength of incident beam, and NA is the numerical aperture of the objective.
The laser fluence (F: J.cm−2), energy delivered per unit area, can be determined using the following equation:
| (7) |
The peak fluence (FPEAK: J.cm−2) per pulse can be derived from this relation:
| (8) |
An important consideration is the location in the beam path where the power measurement is acquired. The laser power can be measured at any location in the beam path, but substantial losses can occur between the laser source and the sample. For example, a commercially available laser-scanning confocal microscope equipped with a femtosecond pulsed laser that has an average power output of 3.8 W at the source is reduced to 0.612 W at the back aperture of the objective indicating an 84% loss in power in the optical path.[34] Though many studies report the rated power of the laser source, accounting for optical losses and determination of the power delivered to the sample is of critical importance. The best recommended practice is to measure the average power at the focal plane of the objective using a microscope slide sensor (if power losses through the objective are significant).
The resolution achievable during hydrogel degradation is influenced by a number of parameters including the numerical aperture (NA) and focusing angle which also influence the working distance. The objective NA is dependent on the refractive index of the medium between the lens and the sample (n) and the focusing half angle (θ) through the following relation:
| (9) |
Additionally, the focal volume excited assumes the shape of an ellipsoid with a short axis d (given by equation 6 above) and a long axis l (given by equation 10 below):[48]
| (10) |
Objectives with a higher NA have an increased focusing angle which typically results in a reduced working distance, which limits the distance from the hydrogel surface to which features can be degraded. An increase in the focusing angle results in a reduced focal volume, higher plasma energy density, increased absorption of energy, and therefore enhanced photodegradation and higher resolution.[51–53] There is a tradeoff between obtaining a maximum range of depth, desired resolution, and efficient degradation. These limitations can be partially mitigated by using a high refractive index medium, for example, a water-immersion objective (n = 1.33) instead of an air objective (n = 1.00). A 20X(NA 1.0) water-immersion objective allowed 200×15×15 μm rectangular channels to be degraded in PEG hydrogels at depths up to 1500 μm from the gel surface.[34] In another study, a 10X(NA 0.3) objective allowed degradation of 30 μm wide lines in silk hydrogels at a depth of 6 cm from the surface.[29] Careful selection of the objective determines the resolution achievable, depth of degradation possible, and degradation efficiency.
Other important aspects that mediate degradation efficiency are the repetition rate and scan speed of the pulsed laser. The repetition rate refers to the frequency of the laser pulses while the scan speed refers to the speed of relative displacement between the focal volume and the sample. Both parameters determine the amount of energy deposited within the focal volume and therefore influence the degree of degradation that occurs. A higher repetition rate or a slower scan speed at a given power and pulse duration induce a higher degree of photodisruption. The extent of degradation of PEG-based hydrogels can be controlled by varying the laser scan speed.[34] Varying the amount of energy delivered via the scan speed allowed for local control over the hydrogel pore size which was used to generate a hydrogel-embedded microfluidic channel with a linear gradient in pore size for biomolecule separation.[34]
While several metrics (average and peak power, intensity, and fluence) are used to characterize lasers, it is also important to identify the energy delivery profile for each system, which subsequently influences the degradation mechanisms at play and ultimately, the achievable resolution and degradation efficiency. As an example, we present theoretical data for three different laser sources (continuous wave, femtosecond pulsed, and nanosecond pulsed) operating at the same average power (1 W) and average intensity (1.34×108 W.cm−2) using an 800 nm wavelength source focused through a 20X(NA 1.0) water-immersion objective (Fig. 2A). The continuous wave laser outputs a constant power and intensity over the entire time interval. The nanosecond (pulse duration = 1 ns) and femtosecond laser (pulse duration = 140 fs) (both operating at 80 MHz frequency) provide significantly higher peak powers and peak intensities: 12.5 W and 1.67×109 W.cm−2 for the nanosecond pulsed laser and 89,286 W and 1.19×1013 W.cm−2 for the femtosecond laser. This demonstrates that laser sources operating at the same average power, average intensity, and pulse frequency can deliver energy in drastically different manners, which must be considered when implementing these systems for material degradation and when comparing results achieved with different laser sources and/or optics.
In a second example, the influence of pulse frequency at a constant pulse duration is demonstrated (Fig. 2B). Using the same average power, average intensity, wavelength, and objective characteristics as in Fig. 2A, the power output for three lasers was calculated (Fig. 2B). The continuous wave laser delivers a constant power and intensity as in Fig. 2A while the low frequency (8 MHz) femtosecond laser delivers a much higher peak power (892857 W) and peak intensity (1.19×1014 W.cm−2) compared to the higher frequency (80 MHz) femtosecond laser operating at the same pulse duration (140 fs) (89286 W and 1.19×1013 W.cm−2). Higher frequency pulses deliver energy at a faster rate but do not achieve the peak values attainable at lower frequencies. Since the peak power and intensity delivered to the focal volume dictate the dominant degradation mechanism and ultimately, the degradation efficiency, the influence of pulse frequency and pulse duration must be considered when choosing a laser source and when publishing degradation parameters.
As a final example, the output from two commercially available lasers used for hydrogel degradation is presented. In the first system, a femtosecond pulsed laser operating at 80 MHz with a 140 fs pulse duration at 790 nm wavelength and 21000 μm.s−1 scan speed focused through a 20X(NA 1.0) objective generates a peak intensity of 7.5×1012 W.cm−2 and a peak fluence of 3.77×106 μJ.cm−2.[34] In the second system, a nanosecond pulsed laser operating at 100 Hz with a 1 ns pulse duration at 355 nm wavelength and 95 μm.s−1 scan speed focused through a 10X(NA 0.25) objective achieves a peak intensity of 5×1010 W.cm−2 and a fluence of 1.13 μJ.cm−2.[35,54] The higher peak intensity and fluence achieved with the femtosecond pulsed laser operating at a higher pulse frequency enabled faster and hence more efficient hydrogel degradation allowing faster scan speeds.
While average powers and intensities of lasers are often reported in the literature, the influence of pulse duration and frequency on the peak power, intensity, fluence, and total energy delivered must be considered when choosing a laser source and comparing the results attained via different laser sources. Furthermore, when reporting the average power, intensity and fluence, consistency in the measurement location along the beam path should be maintained for obtaining comparable results. Since the energy delivered to the sample in the focal volume ultimately governs the degradation process, we propose reporting calculated or measured values at the focal plane of the objective.
2.2 General Mechanisms of Laser-Based Degradation
The mechanisms through which radiant photonic exposure affects different materials, particularly, biomaterials and hydrogels, is an important consideration when implementing laser-based degradation. The dominant degradation mechanism depends on the operating parameters of the laser and the absorption properties and composition of the material. Ultimately, both the laser characteristics and material properties collectively influence the resolution, speed, and efficiency of the degradation process.
For laser-induced degradation to occur, free electrons must be generated in the focal volume. When a material is exposed to pulsed laser irradiation, electrons in the material absorb photons and become excited. Depending on the laser intensity and frequency, these electrons, known as ‘seed electrons’, can undergo two different ionization processes to generate more free electrons: multiphoton ionization (MPI) and tunneling ionization, as determined from the Keldysh parameter described below. Exposure to a high intensity laser imparts enough energy to allow electrons to transition from the valence band to the conduction band. When high intensity is coupled with a high frequency (~MHz range) (where time between pulses is very short), the electrons constantly get excited to a free state via MPI.[53] However, when excited at low frequency (~kHz range), the electrons can relax between pulses and MPI does not occur. In this case, the electric field of the laser suppresses the atomic potential barrier (Coulombic resistance), so that bound electrons tunnel through the barrier and reach a free state which is known as tunneling ionization. In both cases, a threshold intensity is required for ionization to occur.
The Keldysh parameter (γ) can be used to determine the dominant regime of photoionization (tunneling or multiphoton). Below a Keldysh value of 1.5, tunneling ionization dominates and for a value greater than 1.5, multiphoton ionization dominates.[53]
| (11) |
where ω is the pulse frequency, I is the laser intensity at the focal volume, m and e are the reduced mass and charge of the electron, c is the velocity of light, n is the refractive index of the material, ε0 is the permittivity of free space, and Eg is the band gap of the material (energy required to excite the electron from the valence to the conduction band).
Once seed electrons have been excited to the conduction band, they generate additional free electrons via ‘avalanche ionization’, ‘cascade ionization’, or ‘impact ionization’. A seed electron in the conduction band minimum absorbs incident photons (‘inverse Bremsstrahlung absorption’) and is excited to a higher energy in the conduction band. As the excited electron relaxes, it can collisionally ionize other electrons from the valence band, resulting in both electrons being present in the conduction band minimum. With continued absorption, additional electrons are collisionally excited, thereby leading to an avalanche increase in electron density in the conduction band and subsequent plasma formation in the focal volume (Fig. 3A).[48,49,53] During avalanche ionization, there may be energy losses through several mechanisms including electron cooling, recombination, diffusion out of the focal volume, and energy absorption via collision with heavier particles. Hence, the laser intensity must be high enough to overcome these losses for avalanche ionization to proceed.
Figure 3. Interplay of physical mechanisms influencing laser-based degradation.
(A) The sequential occurrence of photoionization, inverse Bremsstrahlung absorption, and impact ionization leads to plasma formation and recurring sequences in absorption and ionization events lead to avalanche growth of free electrons produced by laser excitation. Reproduced with permission.[48] Copyright 2005, Springer-Verlag. (B) Schematic illustration of (a) distribution of processes in the focal volume and (b) sequence of processes occurring over the course of a laser pulse. Zener ionization and Zener-seeded avalanche ionization cause increases in the electron density and plasma field intensity in the focal volume. The laser penetration depth (δ) becomes approximately equal to the wavelength (λ) at 1021 electrons/cm3. When plasma frequency (ωp) becomes equal to the laser frequency (ω), rapid ionization and heavy absorption take place and material over depth, δ, is vaporized. The pink region at the bottom indicates the duration of the laser pulse. EQ and EG refer to the electron quiver energy and the band gap respectively. Reproduced with permission.[55] Copyright 2004, The National Academy of Sciences of the USA. (C) (a) Overview of the range of laser-tissue interactions that occur at varying energy density, power density, and pulse duration and (b) inverse variation of threshold fluence with absorption coefficient over a range of wavelengths. (b) Solid black line indicates the absorption coefficient of water (cm−1), dashed line connecting solid filled circles indicates the absorption coefficient of tissue (cm−1), and dashed line connecting hollow circles indicates the threshold fluence for laser ablation (mJ.mm−2). Reproduced with permission.[50] Copyright 1991, Baillière Tindall.
In the case of nanosecond and picosecond pulses, the ionization process mainly proceeds via tunneling ionization and avalanche ionization leading to luminous plasma formation. At a low laser frequency, the time between pulses is long enough for the energy of the excited electrons in the plasma to be removed from the focal volume via thermal diffusion. Material damage occurs when the local temperature in the focal volume exceeds the melting temperature. Hence, the relative rates of generation and diffusion of thermal energy determines the damage threshold. For sub-picosecond pulses, seed electrons are excited at a faster rate, leading to avalanche ionization and high density plasma formation. Due to the short time between pulses, thermal diffusion does not occur. Instead, the plasma energy is deposited into the bulk material lattice in a shock-like manner, inducing photoablation. For an even shorter pulse duration (femtosecond range), multiphoton ionization dominates and produces sufficient electron excitation to cause direct photoablation of the material.[53] Due to the ultrashort duration between femtosecond pulses, there is no visible plasma formation and optical breakdown occurs via the sudden deposition of excited electron energy and formation of cavitation bubbles.[48,49] It has been argued that, at sub-picosecond pulse durations, laser-induced optical breakdown proceeds via Zener tunneling ionization and Zener-seeded saturation avalanche ionization rather than via multiphoton ionization (Fig. 3B).[55] However, this theory has been challenged by Vogel et al, on the basis that the time constraints required for avalanche ionization were not considered.[48,49]
Depending on the pulse duration, fluence, and intensity, a range of physical effects can be produced in hydrogels. For example, using a low fluence (<10−1 J.cm−2), a low intensity (<103 W.cm−2), and a long irradiation time (>10−6 s), laser-material interactions are usually limited to photochemical reactions since the energy threshold for degradation is not achieved. A reduction in pulse duration and corresponding increase in intensity gradually leads to direct vaporization. A further increase in intensity, and more importantly, reduction of pulse duration, results in deposition of extremely high energy in the focal volume thereby inducing strong controlled explosions of material and direct degradation (Fig. 3C).[50] This photoablation phenomenon can be suitably used for obtaining well-defined, high-resolution degraded features within biomaterials.
The threshold irradiance and threshold fluence required for photoablation depend strongly on the pulse duration. For longer pulse durations (>10−11 s, nanosecond range) threshold irradiance remains constant and threshold fluence decreases strongly. However, a further reduction of the pulse duration to the picosecond and femtosecond range causes a significant increase in threshold irradiance and a weaker reduction in threshold fluence.[48] Considering that the threshold energy for optical breakdown is proportional to the square root of the pulse duration, photoablation can be achieved with much lower energy using femtosecond pulses compared to picosecond or nanosecond pulsed lasers.[56] A detailed picture of the dependence of laser irradiance and fluence on pulse duration is provided in a study by Vogel et al.[51]
Another advantage of using femtosecond pulses over picosecond or nanosecond pulses for hydrogel degradation is that the shockwave generated by the lasing event and material degradation is confined to a smaller volume;[57] hence a high degree of control and precision over degraded features can be achieved. This was demonstrated by Seliktar et al where nanosecond or femtosecond laser pulses were used to ablate 3D channels within PEG-fibrinogen hydrogels.[31] Femtosecond pulses were characterized by a higher laser peak intensity (1012 W.cm−2 vs. 1010 W.cm−2) and the generation of smaller channels (lateral resolution of < 1 μm vs. 4 μm) compared to nanosecond pulses (Fig. 4A–B).
Figure 4. Effect of pulse duration on laser-material interactions.
(A) (a) Degradation using a femtosecond pulsed laser provides increased spatial resolution compared to a nanosecond pulsed as demonstrated by the lateral and axial dimensions of the degradation volume. (b) The visible laser-induced damage proportional to the light intensity (ΔI) as a function of the peak laser intensity for nanosecond and femtosecond pulsed lasers. (c) The measured (X symbols) and theoretical (curves) ablation threshold values versus pulse duration for degradation of PEG-fibrinogen hydrogels. (B) Comparison of visible damage within PEG-fibrinogen hydrogels caused by nanosecond and femtosecond pulsed lasers as a function of laser intensity. Scale bar = 100 μm. (A–B) Reproduced with permission.[31] Copyright 2009, The Biophysical Society. (C) Plasma, shock wave, and cavitation bubble formation in water produced by Nd:YAG laser pulses of different pulse duration and energy, imaged 44 ns after the optical breakdown. Scale bar = 100 μm. Reproduced with permission.[58] Copyright 1996, Acoustical Society of America.
The use of femtosecond vs picosecond and nanosecond pulses for laser ablation can also be comparatively evaluated based on the mechanism of energy dissipation. When energy is focused on a transparent medium (water, hydrogel matrices in water, etc.), the incident energy can either be transmitted, reflected, scattered, or absorbed. Only the absorbed portion of the energy is useful for degradation. More specifically, the absorbed energy can be classified as shock wave energy (due to generation of a mechanical shockwave), bubble energy (due to generation of a cavitation bubble), evaporation energy (due to photoablation), and other radiative losses (Fig. 4C). Amongst these, evaporation energy is of critical importance for obtaining true photoablation, while shock wave and bubble energy lead to disruptive breakdown and a loss in degradation efficiency and resolution.[51] With a reduction in pulse duration from nanosecond (~ 5 ns) to femtosecond (~ 100 fs), more incident laser energy is channeled towards evaporation energy thereby leading to improved and efficient photoablation.[51] For nanosecond and picosecond pulses, a significant fraction of the incident energy may pass the focal volume before it can be absorbed while in the case of femtosecond pulses, the pulse energy is more efficiently absorbed in the focal volume.[52]
The 2P absorption coefficient and 2P cross-section vary across the wide range of biomaterials used in tissue engineering and subsequently influence laser-biomaterial interactions and the dominate degradation mechanism (Fig. 5C, D). Prior knowledge of these biomaterial properties is beneficial for optimizing the efficiency of laser-based degradation to achieve desired features in a timely manner. The pulse duration also influences the 2P absorption coefficient through time evolution of the electron field concentration. In the case of nanosecond pulses, the electron concentration peaks early in the pulse due to avalanche ionization, leading to a higher absorption coefficient and lower transmission. With picosecond pulses, the peak electron concentration is achieved much later during the pulse leading to a decreased absorption coefficient. With femtosecond pulses, a high electron density is reached early in the pulse due to multiphoton ionization, leading to an increased absorption coefficient and resolution.[51] Laser pulse duration is critical in determining the mechanisms involved in hydrogel degradation as well as achieving micron-scale resolution.
Figure 5. Characterization of the two-photon excitation volume.
(A) Visualization of the excitation volume for single-photon (1P) and two-photon (2P) excitation of fluorescein using (a) a continuous wave laser at 488 nm and (b) a femtosecond pulsed laser at 960 nm focused through a NA 0.16 objective. (B) (a) Lateral and axial views of the point spread function using 1P and 2P excitation. (b) FWHM refers to the full-width half-maximum of the Gaussian fit and ω refers to the axial radius. (C) The 2P excitation volume calculated for a 1-GM and a 300-GM fluorophore excited using a 200 fs pulsed laser operating at 80 MHz focused through a 1.2 NA objective. Inset shows the point spread function for excitation of a 1 GM (left) and 300 GM (right) fluorophore at 20 mW. (A–C) Reproduced with permission.[59] Copyright 2003, Nature Publishing Group. (D) The distribution of the normalized intensity and electron density in a focal volume of a femtosecond pulsed laser during optical breakdown using a 1.3 NA objective and 800 nm light. Reproduced with permission.[48] Copyright 2005, Springer-Verlag.
Another important aspect of laser-based degradation is the use of one photon (1P) versus two-photon (2P) events. A majority of the mechanisms that mediate laser-based hydrogel degradation occur via 2P events within a focal volume that assumes a Gaussian profile (Fig. 5B) with lateral (wxy) and axial (wz) radii given by the following equations:[59]
| (12) |
| (13) |
| (14) |
where n is the refractive index of the medium. 2P events depend on the near-simultaneous (10−16 s) absorption of two photons, the probability of which is proportional to the square of the optical intensity and is extremely low under 1P excitation.[59,60] 2P excitation thus necessitates a high photon density and the spatiotemporal focusing of photons within a small volume. Hence, the excitation volume (Fig. 5A–B) (and subsequent degradation volume (Fig. 4A–B)) obtained in the 2P mode (0.1–1 μm3) is significantly smaller compared to the 1P mode (>100 μm3). Another advantage of 2P excitation is the choice of wavelengths used during the process. 2P mode requires two photons of longer, red-shifted wavelength for excitation compared to one photon at a shorter wavelength. The use of longer wavelengths enables reduced scattering and hence deeper photon penetration. This ability is particularly useful when utilizing thick samples to achieve a high photon density deep within the sample.[60–62] When considering the implementation of laser-based degradation in hydrogels, it is advantageous to use 2P absorption over 1P as it allows higher resolution manipulation over the region of interest and the generation of micron-scale features.
An understanding of the basic mechanisms that influence laser-material interactions is useful when applying laser-based degradation of hydrogels. Careful selection of the laser operating parameters and the characteristics of the target material dictate, to some extent, the dominant mechanism underlying degradation which can be optimized to achieve a desired degree of degradation in an efficient and timely manner at desired resolution.
2.3 Mechanisms Dominating Degradation Based on Hydrogel Composition
The mechanisms through which laser exposure affects different materials and in particular, hydrogels, is an important consideration when implementing laser-based degradation. Since a majority of studies investigating laser-material interactions have been conducted using hard materials, it would be interesting to elucidate laser irradiation effects on soft polymeric hydrogels. Since hydrogels inherently possess high water content, laser-induced optical breakdown of hydrogels can be modelled, to some extent, on the optical breakdown of water. Several studies have investigated the optical breakdown of water and simulated biofluids at nano-, pico- and femtosecond pulse durations and the effects of laser properties on shock wave emission, vapor generation, and cavitation.[52,56–58,63–67] However, since the polymeric component of hydrogels also has a wide range of physical, biochemical, structural, and optical properties, it is important to understand their role in laser-based degradation. Hydrogels commonly used for tissue engineering applications can be divided into three broad categories: 1) hydrogels with low 2P absorption, 2) hydrogels with high 2P absorption, and 3) hydrogels containing engineered photolabile groups. Hydrogels from all three categories have been used in laser-based degradation for a broad range of cell and tissue engineering applications.
2.3.1 Hydrogels with low 2P absorption
Hydrogels with low 2P absorption are usually derived from synthetic sources and include PEG, PMMA, and polyacrylamide amongst others. Laser-based degradation of these hydrogels relies primarily on the photodisruptive effect of the entrapped water. When irradiated at a high intensity, water in the hydrogel undergoes plasma formation through electron generation as described in the previous section. The liquid surrounding the plasma is compressed and flows radially outward leading to rapid thermo-elastic expansion and subsequent shock wave generation (for nanosecond pulsed excitation). As the compression wave propagates outward, more liquid is incorporated into the radial flow causing vapor bubble formation. The pressure of the cavitation vapor bubble causes it to expand even after the shock wave has subsided. However, with increasing bubble radius, the kinetic energy of the bubble is dissipated into the surrounding media, eventually leading to bubble collapse within 1 μs.[48,49] The dynamics of the cavitation bubble are highly influenced by the laser fluence, intensity, and pulse duration. A shorter pulse duration (picosecond range) leads to rapid bubble formation and significantly reduced photocavitation at lower energy compared to longer pulses (nanosecond range or larger). Photocavitation of the vapor bubble indirectly causes rupture of the chemical bonds in the polymer chains of the hydrogel leading to physical degradation of the hydrogel in the focal volume. When using a femtosecond pulse duration and reduced threshold fluence, it is also possible for the polymeric chains to undergo direct photoablation via plasma ionization. However, due to the high mass fraction of water, degradation primarily proceeds through bubble cavitation rather than ionization of polymeric chains (Fig. 6). The relative contribution of these parallel mechanisms during degradation of low 2P absorption hydrogels needs to be investigated in detail.
Figure 6. Schematic of the various modes of laser-hydrogel interactions.
Laser-induced degradation of hydrogels can follow various modes depending on the type and composition of polymeric material. Scheme 1: the water within the polymeric network is excited by the laser and undergoes electron generation, plasma formation, and vapor bubble generation. The vapor bubble expands radially along with shock wave propagation and heat dissipation and subsequently causes physical breakage of the network due to thermoelastic stress. Scheme 2: the polymeric network is excited by the incident laser light and undergoes plasma formation which subsequently leads to physical breakage of polymer network. Scheme 3: hydrogels with a high two-photon cross-section or those containing photolabile groups undergo direct chemical scission which induces rapid dissociation of the polymer network.
2.3.2 Hydrogels with high 2P absorption
Hydrogels with high 2P absorption undergo photoablation primarily via MPI and include natural protein-based hydrogels (collagen, silk, agar/agarose) or semi-synthetic materials with high protein content (PEG-fibrinogen) where the presence of amino acid residues imparts a high degree of molar absorptivity.[68] Laser-based degradation proceeds via two parallel mechanisms for this category of hydrogels. When exposed to nanosecond pulses, the incident laser energy is strongly absorbed by the protein fraction leading to rapid heating and chemical denaturation of the protein matrix. The denaturation process involves breakage of hydrogen bonds and disassembly of the primary, secondary, and tertiary protein structure. Due to strong absorption, heat generated in the focal volume is dissipated into the surrounding matrix leading to local protein damage. Simultaneously, the entrapped water undergoes plasma formation and photocavitation via vapor bubble formation (Fig. 6). The combination of protein breakage and photocavitation leads to material degradation. However, at femtosecond pulse durations, the high frequency of the pulses does not allow heat dissipation to the regions surrounding the focal volume, thereby leading to accumulation of heat and subsequent degradation confined to extremely small volumes and high-resolution degradation (on the order of nm).[31]
2.3.3 Hydrogels containing engineered photolabile groups
The third category of hydrogels contain engineered photolabile groups that undergo photochemical scission (Fig. 6).[28,30,36,69–72] A major advantage of these hydrogels is that they are amenable to degradation via both single- and two-photon exposure and require much lower energy compared to the other two hydrogel categories. In one study, a bis(azide) di-functionalized polypeptide linker containing the photodegradable nitrobenzyl ether (NBE) moiety was covalently coupled to PEG monomeric chains enabling photocleavage using single photon (λ = 365 nm) or two-photon (λ = 740 nm) exposure.[73] The ability to specifically cleave the polymeric chains of the hydrogel matrix at low energy provides a high level of spatiotemporal control over the degradation process and enables the generation of well-defined, user-controlled 3D features within bulk matrices.
An important consideration for these three classes of hydrogels is the efficiency with which they can be degraded. It is often desirable to obtain complete hydrogel degradation at the fastest laser scan speed available while still maintaining high-resolution features. Hydrogels with photolabile groups exhibit the fastest rate of degradation owing to direct chemical scission of the polymer backbone upon exposure to light and most likely the highest resolution. This class of hydrogels can be subjected to higher scan speeds while still achieving complete degradation with high resolution. Protein-rich hydrogels also demonstrate relatively fast rates of degradation owing to the strong absorption of the protein backbone. However, hydrogels with low 2P absorption require more energy. Since degradation in this class of hydrogels depends on indirect vapor bubble generation and cavitation, a slower scan speed is required in order to deposit a higher amount of energy per unit volume per unit time compared to the previous two classes of hydrogels. For example, photolabile PEG-tetraDIFO3 hydrogels containing UV-sensitive nitrobenzyl ether moieties underwent 2P-induced degradation at a scan rate of ~1 m.s−1.[28] In comparison, 2P-induced degradation of PEG-based hydrogels underwent complete degradation at a scan speed of 0.02 m.s−1.[34] In addition to hydrogel composition, the scan speed necessary to achieve complete degradation also depends on the pulse duration of the laser. In one study, PEG-based hydrogels were degraded using a nanosecond-pulsed laser at a scan speed of ~0.0001 m.s−1 compared to 0.02 m.s−1 achieved using a femtosecond pulsed laser for a similar hydrogel composition.[35] The same nanosecond laser system was used to degrade features in PEG-fibrinogen hydrogels that contain a higher 2P cross-section at a similar speed of 0.0001 m.s−1.[31]
Hydrogel composition and the laser operating parameters govern the physical mechanism(s) of degradation that occur which in turn dictates degradation efficiency, the fastest scan speed attainable, and resolution of degraded features. Prior to implementation, the roles of laser operating parameters and mechanisms of degradation must be carefully considered to achieve efficient and high-resolution degradation. As a general guideline, the following table (Table 1) briefly lists the published operating parameters for different biomaterials. A more detailed table is provided as Supplementary Table 1 and provides published resolutions achieved thus far.
Table 1. Laser parameters for laser-based degradation of various hydrogels and biomaterials based on composition.
A brief list of studies employing laser-based degradation for natural, synthetic, and semi-synthetic materials is provided (λ: wavelength of laser light, Ti:S: Titanium-sapphire, Nd:YAG: Neodymium-yttrium aluminum garnet, PVP: polyvinyl pyrrolidone, PEG: poly(ethylene glycol)).
| Material | Degradation Parameters Source / λ / Pulse Duration / Pulse Frequency / Speed / Power |
References |
|---|---|---|
| Collagen | Ti:S / 775–830 nm / 75–140 fs / 0.001–82 MHz / 0–100 μm·s−1 / 0.4–1130 W | [41,74,75] |
| Collagen – Gold Nanorods | Ti:S / 800 nm / 100 fs / 80 MHz / 750–2000 μm·s−1 / 0.1–0.29 W | [76] |
| Collagen-Elastin Blend | Ti:S / 800 nm / 30 fs / 0.001 MHz / --- μm·s−1 / ~0.8 W | [77] |
| Gelatin | Ti:S / 800 nm / 30 fs / 0.001 MHz / --- μm·s−1 / --- W | [78] |
| Agar | Nd:YAG / 1053–1064 nm / 1 ps / 133 MHz / 0–20 μm·s−1 / 0.1 W | [42,79] |
| Agarose | ArF / 193 nm / --- fs / 0.00001 MHz / --- μm·s−1 / 0.7 W | [22] |
| Agarose-Alginate Blend | Thulium / 1920 nm / --- fs / --- MHz / 500–5000 μm·s−1 / --- W | [80] |
| Silk Fibroin | Ti:S / 810–1030 nm / 100–230 fs / 0.1–80 MHz / 25–100 μm·s−1 / --- W | [29,45,81] |
| PVP | Ti:S / 800 nm / 30 fs / 0.001 MHz / --- μm·s−1 / --- W | [78] |
| PVP-Chitosan Blend | Ti:S / 800 nm / 30 fs / 0.001 MHz / --- μm·s−1 / --- W | [78] |
| PEG | Ti:S / 790 nm / 140 fs / 80 MHz / 21000 μm·s−1 / 0.612 W N2 / 355 nm / 1 ns / 0.0001 MHz / 95 μm·s−1 / --- W |
[34,82] [35] |
| Photolabile PEG |
Ti:S / 405–740 nm / --- fs / --- MHz / 890000 μm·s−1 / 0.008–1.8 W | [28,30,69–71,73,83] |
| PEG-Fibrinogen | Ti:S / 700–920 nm / 100 fs / --- MHz / 100 μm·s−1 / --- W N2 / 355 nm / 1 ns / 0.0001 MHz / 97–100 μm·s−1 / --- W |
[31,84] [31,85] |
| PEG-Albumin | N2 / 355 nm / 1 ns / 0.0001 MHz / 100 μm·s−1 / --- W | [85] |
| PEG-Gelatin | N2 / 355 nm / 1 ns / 0.0001 MHz / 100 μm·s−1 / --- W | [85] |
PART 2: Applications of Laser-Based Hydrogel Degradation in Cell and Tissue Engineering
3. Applications of Laser-Based Hydrogel Degradation in Cell and Tissue Engineering
Laser-based micro- and nano-scale manipulation of hydrogels has received increasing attention with the advent of improved laser systems and associated infrastructure. Common laser-based techniques used in biofabrication include stereolithography,[54,86] 3D micro- and nano-patterning of biochemical moieties,[14,87–91] 2D texturing of biomaterial interfaces to modify surface properties,[25,38] and microstructuring/micromachining bulk materials to generate user-defined features.[92–94] In addition to these established techniques, laser-based degradation has been progressively applied in various aspects of tissue engineering including fabrication of microchannels for vascular and neural studies, probing cell-material interactions, and for characterization of scaffolds and matrices as discussed in the following sections.
3.1 Laser-Based Degradation of Hydrogels to Control 3D Cellular Architecture
Spatiotemporal monitoring of cell migration, self-assembly, and morphogenesis in both synthetic and natural hydrogels has received significant attention owing to its importance in numerous physiological phenomena including organ and tissue development, cardiovascular diseases, and cancer progression.[95–98] Two broad approaches have been adopted to investigate these processes: 1) self-assembly of cells (including multiple cell types) from a “zero-state” initial encapsulation condition[99,100] and 2) directed organization of cells into desired configurations using a pre-defined architecture.[34,101] Self-assembly relies on the ability of cells to actively remodel the surrounding matrix, often through proteolytic degradation,[102] and facilitates the investigation of several processes including normal vasculogenic and tumor-associated angiogenic progression,[103,104] spheroid formation of cancer cells,[19,105] collective cell migration,[106,107] and tissue morphogenesis.[98,108] Though self-assembly provides significant mechanistic insights into developmental processes, it suffers from limitations including minimal to no control over the final architecture, dependence on cells with high proteinase activity, and poor-user control over micron-scale ECM features.
To overcome some of these limitations, directed spatial organization of cells or multicellular constructs using pre-defined biochemical or mechanical structures to guide and directly control 3D cellular architecture has been adopted.[14,34] This approach often utilizes well-defined, micron- to centimeter scale channels or conduits to guide cellular organization and offers a high level of reproducibility between constructs. While simple geometric patterns are often utilized, the implementation of image-guided approaches has allowed fabrication of biomimetic architectures that closely recapitulate in vivo organization.[14,34] These highly reproducible biomimetic architectures are of interest in generating in vitro platforms that mimic physiological and pathological tissue architectures. In this context, laser-based hydrogel degradation facilitates the formation of user-defined and in vivo-derived tissue architectures and offers a unique opportunity to guide 3D cellular organization over multiple length scales.
3.1.1 Neuronal Guidance
There has been a long-standing interest in the ability to spatially guide neuronal outgrowth in tissue engineered constructs for regenerative medicine applications and for the development of in vitro microphysiological models.[109,110] Currently, no effective treatments for repair of damaged nerves in spinal cord and brain injuries exist; hence, methods to effectively culture neural cells in implantable biomimetic matrices while guiding their architecture and connectivity are of great interest for neuronal regeneration.[111] Additionally, development of advanced neuronal cell culture platforms could significantly aid the study of neurite extension and morphogenesis, whereas methods to improve proliferation, migration, differentiation, survival, and controlled 3D architecture of neuronal connections in vitro could lend themselves towards the formation of microphysiological systems as disease models.
Cell culture methods to direct and guide axonal outgrowth have employed 2.5D tracks on bioadhesive surfaces,[22,112] large micromolded, planar channels in hydrogels,[113] or patterned 3D tracks of cell-adhesive GRGDS peptides in hydrogels.[114] Complementary approaches utilizing laser-based degradation to generate 3D conduits embedded in hydrogels for guidance of neuronal extension outgrowth have also been developed. These approaches allow for high resolution control over axon bundle diameter and 3D architecture and could potentially be extended to spatially control where axonal connections and neuromuscular junctions form.
Laser-based degradation of guidance channels in agarose-based hydrogels for neuro-morphogenesis enabled the formation of mature neuron bundles and neural circuits.[22] 1 cm long, 60–400 μm wide guidance channels were degraded in agarose using an ArF excimer laser (193 nm wavelength, 1 μJ energy per pulse, 10 Hz repetition rate, 30 μm scan size), functionalized with extracellular matrix proteins, and implemented to study proliferation, alignment, and differentiation of B35 rat neuroblasts into neural bundles. The thickness of the neural bundles was directly proportional to the width of the guidance channel which, in turn, was dependent on the laser operating parameters during degradation. Hence, optimization of laser parameters could be used for generating neural bundles of desired length and thickness embedded in hydrogels.[22] Similarly, guided neural network formation was investigated using agarose-based, on-chip platforms where individual microchambers were connected via microchannels using laser-based, photothermal-induced degradation. In this case, agarose degradation was achieved through heating an adjoining chromium layer using a Nd:YAG laser (1064 nm wavelength, 20–48 mW laser power, 1–1000 μm·s−1 scan speed) thereby inducing the formation of microchannels with widths of 5–45 μm and heights of 5–20 μm connecting adjacent microchambers.[43,115] Neural cells or rat adult hippocampal cells cultured in microchambers formed neural networks in the microchannels between chambers.[42] Well-developed neuronal connections were established irrespective of channel width, length, or microchamber shape, demonstrating the applicability of the on-chip platform coupled with laser-based fabrication to directly control the architecture and dynamics of neuronal connections (Fig. 7A). However, photothermal melting of agarose could potentially cause cellular disruption and heat shock protein generation and hence care needs to be taken during stage movement and the laser properties must be tightly controlled to mitigate these risks. This phenomenon was observed where S5Y5 neuroblastoma cells were encapsulated in 3D agarose-alginate hydrogels and subjected to laser-based microchannel fabrication.[80] Cells residing in the direct ablation path died while cells in the adjoining regions did not suffer significant damage and were able to fully recover metabolic activity.
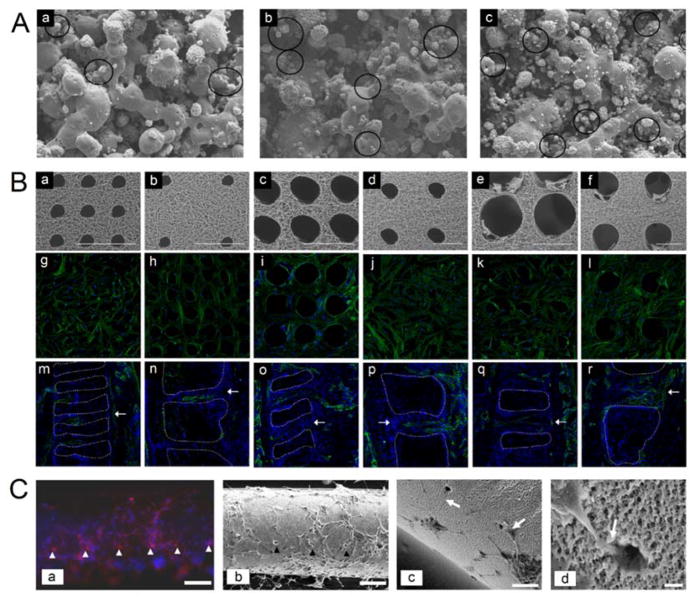
Figure 7. Neuronal guidance using laser-degraded networks.
(A) Degradation of microchannels between agar microchambers enables the formation of functional connections between rat hippocampal cells. (a) Microchannels (indicated by white arrows) are fabricated 1 day after cells are introduced, (b) cellular connections are formed over 5 days, (c) photothermal etching to create new channels in the presence of cells, and (d) neurites connecting cells in the new channels after 10 days. Scale bar = 100 μm. Reproduced with permission.[42] Copyright 2004, Elsevier. (B) Directing axonal growth in photolabile PEG hydrogels under various degradation conditions. (a) Embryonic stem cell-derived motor neuron embryoid body encapsulated within photolabile PEG hydrogels on day 0 and (b) 48 hours later demonstrate axonal growth into fully degraded channels. The power density was varied from 15 to 110 mW.μm−3 to control the extent of degradation in the channels. Channels marked with * theoretically undergo incomplete degradation and hence axonal growth is hampered. Scale bar = 100 μm. (c) Fork-shaped patterns were degraded in the path of extending neuronal projections to provide directional choice in growth. (d) A majority of axons entered the 0° or ±45° forks, indicating persistence in outgrowth under the absence of external stimuli. Scale bar = 10 μm. Reproduced with permission.[71] Copyright 2014, ACS. (C) Degraded channels in PEG-fibrinogen hydrogels in different configurations ((a–b) along the word “nerve” and (c–d) a cylindrical shaped channel) allow dorsal root ganglion cells to form neuronal projections over 3 days. Scale bar = 100 μm. Reproduced with permission.[31] Copyright 2009, Biophysical Society. (D) Comparative potential of PEGylated proteins (PEG-fibrinogen, PEG-albumin, PEG-gelatin) and PEG as a control to support dorsal root ganglion axonal outgrowth in laser-degraded microchannels. Cells were labeled for βIII-tubulin (red) and a DAPI counterstain (blue). Scale bar = 100 μm. Reproduced with permission.[85] Copyright 2015, Elsevier.
Peptide-coupled, photolabile PEG-based hydrogels have also been modified with laser-induced degradation to guide neuro-morphogenesis and neural circuit formation.[71] Embryonic stem cell-derived motor neurons (ESMNs) encapsulated in YIGSR-functionalized, photolabile (containing nitrobenzyl ether moieties) PEG-based hydrogels were degraded using a power density ranging from 15–110 mW.μm−3 to create channels with widths varying from 2–50 μm. Axons extended into channels as small as 2×2 μm but the invasion speed was significantly slower in 2 μm channels compared to invasion into 5–50 μm channels due to physical impedance of the axons. When presented with multiple channels at different branching angles, motor axons demonstrated directional persistence toward 45° channels. Interestingly, guided axonal growth from ESMN embryoid bodies toward myotube bundles formed functional synapses over millimeter-scale distances, thereby demonstrating the ability to direct and control the organization of neural circuits (Fig. 7B).[71]
Semi-synthetic hydrogel blends including PEGylated protein hydrogels have also been subjected to laser-based degradation for neuronal guidance. Using PEG-conjugated ECM protein hydrogels[31,84,85,116] and cell culture medium supplemented with nerve growth factor[117], 20–70 μm diameter channels in PEG-fibrinogen hydrogels supported outgrowth of neurites and glial cells from dorsal root ganglia (DRG)[85] (Fig. 7C). The comparative potential of PEGylated protein hydrogels with different compositions (PEG-fibrinogen, PEG-gelatin, and PEG-albumin) to support laser-guided channel fabrication for neurite growth was investigated. PEG-gelatin displayed significant ultraviolet (UV) light attenuation which induced an intensity loss with increasing depth into the hydrogel. This attenuation reduced the total energy delivered resulting in decreased channel diameters with increased depth from the gel surface.[85] In contrast, PEG-fibrinogen (PEG-Fb), PEG-albumin, and PEGDA were optically transparent and ablated channel diameters remained consistent with increased depth. The laser parameters (355 nm, 1 nm pulse duration, 90 μJ per pulse, 100 Hz repetition rate, maximum intensity 5×1010 W.cm−2) were modified (60–100% of maximum intensity) to control the channel diameter (10–70 μm) adjacent to dorsal root ganglion (DRG) cell clusters; with increasing laser energy producing larger channels. DRG cell migration and outgrowth occurred quicker in channels larger than 30 μm, while 10 μm diameter channels induced outgrowth at a slower rate. Cells also displayed higher persistence outgrowth in PEG-Fb and PEG-albumin hydrogels compared to PEG-gelatin (Fig. 7D).[85] These results further demonstrate that the appropriate choice of material properties and laser operating parameters require careful consideration to achieve desired neuronal growth and development.
PEG-Fb hydrogels were further investigated for their ability to support guided neurite outgrowth toward functional nerve regeneration[116] and brain-on-a-chip applications.[84] The influence of femtosecond and nanosecond pulsed laser sources on microchannel characteristics in PEG-Fb hydrogels with varying fibrinogen and PEG concentrations were investigated.[31] Hydrogel degradation using a nanosecond pulsed laser (1 ns pulses, 100 Hz frequency, 355 nm, 97 μm.s−1 scan speed, and intensity varying from 0.05×1010 to 5.0×1010 W.cm−2) or a femtosecond pulsed laser (100 fs pulses, 880 nm, 100 μm.s−1 scan speed, and intensity varying from 0.6×1012 to 7.8×1012 W.cm−2) was implemented to investigate the role of laser properties on degradation resolution. Laser-based degradation using the femtosecond pulsed laser produced higher resolution structures compared to the nanosecond pulsed laser owing to a higher peak intensity and a smaller spot size. Accordingly, modulation of laser intensity resulted in variations in neurite outgrowth and invasion. When laser intensity was reduced below 1×1010 W.cm−2 (for the nanosecond laser) or 5×1012 W.cm−2 (for the femtosecond laser), neurite outgrowth was severely restricted owing to the small diameters of the fabricated channels. However, channels with a larger cross-section (>20μm) supported neural outgrowth and maturation over 23 days in culture, as evidenced by the migration of βIII-tubulin-positive neurites and supportive, s100-positive glial cells, in both simple and complex geometries, including non-planar networks.[31]
An important consideration in these studies is the presence of other cell types in addition to neural cells and their ability to actively remodel the surrounding ECM. Glial or non-neuronal cells present in the DRG cell cluster could potentially degrade the PEG-Fb matrix adjoining the fabricated channels and display transverse growth instead of being unilaterally restricted along the channel.[31] Tuning the PEG-Fb hydrogel composition by controlling the relative concentration of fibrinogen or PEG moieties was necessary to confine cellular outgrowth to the channels.[31] Overall, the use of PEG-conjugated ECM protein hydrogels in combination with laser-based degradation allows for the simultaneous investigation of biochemical and physical cues and their role in the progression of neuronal development.
Controlled laser-based degradation of micron-scale features in natural, synthetic, and natural-synthetic hybrid hydrogels can be implemented to guide the formation of cell-cell interactions and complex neural structures to improve neural functionality and maturation of fabricated in vitro neural tissues. This ability to guide neural organization enhances researchers’ ability to study distinct mechanisms of neuronal development and to accelerate neural regeneration of damaged tissue post-implantation of fabricated scaffolds.
3.1.2 Fabrication of Vascular Networks
Another important application of laser-based hydrogel degradation is the fabrication of vasculature and guidance of endothelial network formation. The generation of hydrogel embedded vascular networks is essential for long-term culture of large-volume tissue constructs and maintenance of high cell viability via adequate transport of nutrients and cellular metabolites.[118–120] Additionally, fabrication of well-defined, repeatable vascular architectures lends itself to the development of fluidized disease models. In this respect, laser-based degradation of hydrogels has facilitated the fabrication of in vitro biomimetic microfluidic networks for subsequent tissue vascularization.[34–36,76,82]
In one study, a nanosecond pulsed laser (1 ns pulses, 100 Hz frequency, 355 nm, 95 μm.s−1 stage speed) was used to generate 2D planar and 3D microfluidic channels in a number of hydrogel matrices including peptide-coupled PEG, collagen I, gelatin, fibrin, agarose, and alginate.[35] This approach was used to recreate the architecture of a planar capillary bed, form 3D “evolvable” networks within cell-laden hydrogels, and to generate lumenized, mature vasculature composed of human umbilical vein endothelial cells (HUVECs) with tight cell-cell junctions. An advantage of this method is that it facilitates degradation in situ, in the presence of encapsulated cells, without inducing damage to cells within ~20 μm of the laser path and also allowed spatiotemporal control over fluidic channel formation and flow and transport patterns (Fig. 8A). The ability to generate new microfluidic channels or new connections between existing networks in cell-containing constructs provides unprecedented spatiotemporal control over the delivery and presentation of soluble factors to encapsulated cells and the potential to probe the spatiotemporal influence of soluble factors on tissue formation and maturation.
Figure 8. Fabrication of vascular networks in hydrogels via laser-based degradation.
(A) (a) Schematic implementing laser-based hydrogel degradation to generate microchannels in cell-laden hydrogels and formation of additional channels on-demand. (b) Digital mask of a 2D capillary bed and (c–d) resulting microfluidic network generated via image-guided degradation. Scale bar = 100 μm. (e) Confocal 3D reconstruction of a lumenized channel formed by HUVECs in collagen type I, (f) transverse and (g–i) frontal plane of the channel. Cells stained for F-actin (green), CD-31 (purple), VE-cadherin (red) and DAPI (blue). Reproduced with permission.[35] Copyright 2016, Wiley. (B) (a) Schematic of image-guided, laser-based degradation in micromolded PEGDA hydrogels. (b) Hydrogel embedded, 3D, cerebral cortex-derived microfluidic networks demonstrate correlation with in vivo vasculature. (c) 3D confocal reconstruction of lumenized microchannels within PEGDA hydrogels formed by bEnd.3 mouse brain endothelial cells labelled with ZO-1 (tight junctions, green) and DAPI (nuclei, blue). Reproduced with permission.[34] Copyright 2016, Wiley. (C) (a) Schematic of a nanorod-embedded collagen matrix undergoing photothermal degradation to form vascular channels. (b–c) bEnd.3 cell migration and tube formation in the laser degraded channels over 14 days. (d) A 3D confocal reconstruction of the lumenized channels with hollow cores. Scale bar = 50 μm. Reproduced from reference.[76]
An image-guided, laser-based degradation technique has also been developed that allows for accurate recapitulation of the 3D architecture of microvascular networks in hydrogels. [34,82] In this approach any 3D configuration can be utilized from those generated using computer-aided design (CAD) to 3D image stacks of in vivo vasculature. A 3D image stack of mouse cerebral cortex microvasculature was utilized as a digital template to fabricate a 3D biomimetic microfluidic network.[34] A series of digital, virtual masks that contained the network spatial information was used to guide the laser position during 3D degradation of PEG-based hydrogels using a femtosecond pulsed laser (790 nm, 140 fs pulse duration, 37.7 nJ.μm−2 laser fluence).[34] This approach allowed fabrication of microfluidic networks whose architecture closely matched the in vivo architecture used to define the network in terms of total network length, number of branch points, and average vessel diameter across many networks. Post functionalization with the cell-adhesive peptide, RGDS, facilitated adhesion and spreading of bEnd.3 mouse brain endothelial cells which subsequently formed lumenized vascular networks with tight cell-cell junctions (Fig. 8B).
Another approach for vascular channel formation is via femtosecond photothermal degradation (100 fs, 800 nm wavelength, 80 MHz, 50–100 mW laser power, 0.25–2.0 mm.s−1 writing speed) of cell-laden collagen hydrogels containing gold nanorods.[76] Instead of directly degrading the collagen, the gold nanorods absorb the photon energy to convert light to heat thereby causing local thermal denaturation of the collagen matrix. Optimization of laser power and scan speed (100 mW, 2.0 mm.s−1) were necessary to avoid extensive cell death while simultaneously maintaining channel fidelity. Hollow, tube-like channels were degraded within collagen hydrogels to guide the migration and formation of a lumenized endothelium by bEnd.3 mouse endothelial cells (Fig. 8C).
Much research has focused on the generation of hydrogel-embedded cardiovasculature to support viability and function of engineered microtissues for disease modeling and regenerative medicine applications. However, the role of the lymphatic system, which is responsible for regulating metabolic transport and vessel stabilization,[121] has been relatively understudied. To simulate local transport between the cardiovascular and lymphatic system, image-guided, laser-based degradation has been used to generate two microfluidic networks that occupy the same volume but do not directly connect to generate a simple model of cardiovascular-lymphatic transport.[34] This multi-network device allowed for quantification of transport between networks and provides a platform to begin investigating in vivo-like transport in engineered tissues.
Laser-based hydrogel degradation facilitates fabrication of simple, and more complex, in vivo-derived vascular networks which can subsequently be used for generation of large-volume tissues and fluidized microphysiological systems. The ability of endothelial cells to form user-defined, 3D networks in hydrogels provides some advantages over other platforms that rely on self-assembly through vasculogenic or angiogenic processes. Hence, employing laser-based degradation for in vitro vascular studies could provide a useful platform for studying vascular phenomena including interactions with immune cells, circulating tumor cells, and other blood components.
3.1.3 Guided Cell Migration
In addition to neural and vascular applications, laser-based hydrogel degradation has been applied to investigate cell migration through hydrogel-embedded channels. This approach provides a high degree of control over channel dimensions and the ability to regulate specific events and/or mechanisms of cell migration.
hMSCs and fibroblasts encapsulated in silk fibroin hydrogels containing laser-micromachined features were observed to align to the features thereby demonstrating the role of cell-matrix contact guidance in regulating 3D cell morphology (Fig. 9A).[29] Implantation of these constructs into mice resulted in cellular infiltration along the 3D channels, thereby providing proof-of-principle of the applicability of this technique in regenerative medicine. 3D channels also enable the investigation of chemokine-induced cell migration (chemotaxis).[35] hMSCs encapsulated in PEG-based hydrogels with laser-degraded linear channels were subjected to platelet derived growth factor-BB (PDGF-BB) perfusion, which resulted in a higher cellular migration distance and directional persistence compared to perfusion without PDGF-BB (Fig. 9B). Temporal control over chemokine-induced gradients was also demonstrated via in situ fabrication of new microchannels within desired locations of cell-encapsulated hydrogels. This approach allows for temporal control over soluble factor gradients and could significantly aid the investigation of soluble factor influences on cell- and tissue-level function.
Figure 9. Laser-based degradation for cell migration studies.
(A) (a) Schematic of the laser-based degradation of cell-laden silk hydrogels. (Inset) Bright-field image of the degraded region. Scale bar = 250 μm. (b–d) Confocal images of the cell-laden gel 76 μm below, 62 μm above, and in the plane of the degraded features, respectively. Dashed lines indicate the degraded region. Scale bar = 250 μm. (e–f) Cells irradiated by the beam above (e) and below (f) the focal plane. Green indicates live cells and red indicates dead cells. Reproduced with permission.[29] Copyright 2015, The National Academy of Sciences of the USA (B) (a) Representative bright-field image of pre-starved hMSCs overlapped with respective cell tracks migrating towards a microfluidic channel (indicated by white dashed line) perfused with a chemoattractant, platelet derived growth factor-BB, compared to (d) control (perfused with media only). (b,c) Measured cell migration distances and directionality of individual cells in response to chemoattractant perfusion as compared to the (e,f) control group. Reproduced with permission.[35] Copyright 2016, Wiley. (C) Migration of a fibrosarcoma cell (white rectangle) through a channel in photolabile PEG and its corresponding position trace. Scale bar = 50 μm. Reproduced with permission.[30] Copyright 2009, AAAS. (D) (a) Laser-degraded microchannels functionalized with RGD direct migration of 3T3 fibroblasts in photolabile PEG hydrogels. (b) 3D cell outgrowth can be directed via RGD-functionalized channels, inset shows top-down projection. (c) Y-shaped microchannels with one arm functionalized with RGD shows preferential cellular outgrowth compared to the non-functionalized arm. Dashed polygons represent RGD functionalized regions. Hydrogel is shown in red, F-actin in green and cell nuclei in blue. Scale bars = 100 μm. Reproduced with permission.[28] Copyright 2011, Macmillan.
In addition to modeling physiological conditions, modeling of pathological processes such as the metastatic cascade has received significant attention in recent years with a number of hydrogel formulations being used for 3D culture of cancer cells.[96,122,123] Laser-based degradation has been implemented to generate well-defined migration tracks in 3D collagen matrices to elucidate the role of MMP-dependent and -independent cancer cell invasion.[41] Laser-based degradation (400 mW power, 830 nm wavelength) was implemented to generate microtracks of varying width (3–30 μm) and the invasion of mouse mammary tumor (MMT) cells along the ablated tracks was quantified. Interestingly, MMT cells, owing to high degree of proteolysis, were able to invade and degrade the surrounding collagen matrix beyond the generated tracks. However, small molecule inhibition of MMP activity resulted in MMP-independent invasion of cells, which resulted in confinement of cell migration within the tracks. Additionally, invasion path thickness increased with the width of the ablated tracks, thereby demonstrating the utility of laser-guided degradation in investigating cancer cell invasion mechanisms. This approach was used to study the guided migration of human fibrosarcoma cells and 3T3 fibroblasts along laser-degraded channels in photolabile PEG-based hydrogels (Fig. 9C).[28,30] 3T3 fibroblasts within a fibrin clot were encapsulated in photolabile hydrogels and microchannels were degraded and selectively functionalized with cell-adhesive RGDS motifs. Cells were observed to migrate specifically to RGDS-functionalized channels, thereby demonstrating user-specific spatial control over 3D cell migration (Fig. 9D).
Laser-based degradation of hydrogels can be suitably applied to create user-defined and controlled microstructures within 3D biomaterial matrices, which can subsequently be used to investigate cellular behavior pertaining to physiological or pathological conditions. Guided or directed cell growth and alignment could be extended towards reconstruction of native tissue microarchitectures for regenerative medicine applications. Additionally, fabrication of these features within in vitro platforms could also be used to study disease mechanisms involving cell invasion and migration.
3.1.4 Manipulation of Cell Behavior and Cell-Material Interactions
Control over cell-cell and cell-matrix interactions in tissue-engineered constructs are critical towards close recapitulation of native microenvironmental conditions and for achieving desired cellular level, and ultimately tissue level, function. The introduction/modification of micron-scale features in hydrogels via laser-based degradation facilitates the longitudinal study of these interactions. Alteration of key parameters including ligand presentation, crosslinking density, porosity, and others can influence cellular morphology, migration, and differentiation thereby providing controlled in vitro investigation of cell-matrix interactions [30,83] To this end, laser-based hydrogel degradation can be implemented to introduce or modify micron or sub-micron scale features in bioengineered matrices to manipulate or to specifically investigate 2D or 3D cell behavior in response to matrix modifications.
Laser-based erosion of hydrogels facilitates investigation of cellular interactions on 2D hydrogel surfaces where adhesiveness and detachment of human mesenchymal stem cells (hMSCs) was characterized.[83] A photolabile macromer, PEGdiPDA (PDA: photodegradable acrylate), containing an ortho-nitrobenzylether (o-NBE) moiety, with a 2P cross-section of 0.01–0.03 GM at 740 nm, was used to form photolabile hydrogels. Post cell spreading on the surfaces, specific regions of (~1–100 μm in size) were subjected to laser erosion. When regions corresponding to cell attachment sites or filopodial extensions were selectively eroded, cells were observed to dynamically detach and retract leading to rounded cell morphologies. The cellular retraction kinetics was modeled based on cell spread area, which revealed that cellular detachment is dependent on substrate elasticity and that hMSC detachment from softer substrates was ~6 fold slower than detachment from stiffer substrates (Fig. 10A). An advantage of using 2P laser erosion for studying cell detachment dynamics is that this method allows investigation of the full signaling cascade induced by disruption of adhesive junctions as compared to other methods which selectively target specific aspects of the cascade. Additionally, this method could be further used to study adhesion force dynamics between multiple cell types.[124] For example, in a co-culture or tri-culture 2D model, the maturation of individual cell-cell adhesions and associated signaling transduction cascades could be investigated by femtosecond laser ablation of the desired adhesive junctions followed by assessment of the energy required to disrupt these junctions via contact force measurements while simultaneously imaging changes in signal transduction using fluorescent protein reporters.
Figure 10. Laser-based manipulation of cellular behavior and cell-material interactions.
(A) Laser-based degradation of photolabile PEG hydrogel substrates beneath cultured cells’ adhesion sites was used to dynamically induce cell retraction. (a) Cell adhesion was disrupted by degradation under cell adhesions at the anterior (yellow circle), posterior (purple oval) or individual filopodial sites (red triangle). (b) Time course of erosion of the photolabile hydrogel substrate (outlined in gray) and subsequent retraction of GFP-actin transfected mesenchymal stem cells (MSCs). Scale bar = 20 μm. Reproduced with permission.[83] Copyright 2010, RSC. (B) (a) hMSCs encapsulated in photolabile PEG hydrogels appear rounded (top panel) but undergo spreading (indicated by white arrows) after hydrogel degradation using UV irradiation (480 s, 365 nm at 10 mW.cm−2) (bottom panel). Scale bar = 50 μm. (b) Photolytic removal of tethered RGDS adhesive peptides regulates αvβ3 integrin expression (top panel) and chondrogenic differentiation (bottom panel) of hMSCs. Scale bar = 100 μm. Reproduced with permission.[30] Copyright 2009, AAAS. (C) Field emission microscopy images of NIH3T3 fibroblasts cultured over 3 days on laser-degraded collagen/elastin layers. (a) Cells on unmodified surfaces migrate without any preferred direction. (b–d) Cells on laser-modified surfaces attach and align preferentially along the grooves (indicated by white dashed lines). White dashed circles indicate individual laser shots and white arrows indicate attached cells. Scale bar = 100 μm (a–c) and 10 μm (d). Reproduced with permission.[77] Copyright 2016, Elsevier.
Hydrogel erosion was also applied to hMSCs encapsulated in photolabile, bioactive PEGdiPDA hydrogels that were monitored for their ability to spread and undergo chondrogenic differentiation in response to photolytic degradation.[30] hMSCs initially exhibited a rounded morphology, but upon local photolytic degradation, formed filopodial extensions and displayed enhanced spreading (Fig. 10B). This phenomenon occurred primarily due to a reduced crosslinking density and subsequently increased molecular porosity due to degradation as was demonstrated with hMSC spreading in gradient degraded hydrogels.[69] The cell-adhesive peptide sequence, RGDS, was conjugated to NBE moieties to create hydrogels with dynamically tunable cellular adhesivity.[30] In comparison to hMSCs cultured in constant RGDS hydrogels, hMSCs cultured in photocleavable RGDS hydrogels demonstrated higher GAG (glycosaminoglycans) production, reduced αvβ3 integrin expression (due to loss of cell adhesion), reduced expression of CD105 (hMSC marker), and increased expression of COLII (chondrocyte marker), demonstrating enhanced chondrogenic differentiation through dynamic photomanipulation of ECM composition without altering mechanical properties (Fig. 10B).[30] These studies demonstrate that controlled material degradation and biochemical modification of hydrogel matrices can locally regulate the ECM composition which in turn can be used to obtain desired cellular morphologies or function and spatiotemporal control over tissue characteristics. More importantly, the decoupling of hydrogel mechanical properties from ligand concentration can be achieved via 2P photolysis, and their individual roles on cellular phenotype and 3D behavior investigated.
Another important application in this area of research is the control and manipulation of cell adhesion on laser-treated biomaterial interfaces. Laser structuring of biomaterial surfaces creates micron-scale or sub-micron scale features that influence surface wettability, roughness, and hydrophobicity.[25,125,126] These surface characteristics modulate how cells sense the biomaterial surface through focal adhesions and lamellipodial/filopodial extensions. A number of biomaterial surfaces including poly(dimethyl siloxane) (PDMS), photocurable ORMOCER (organically modified ceramic), and biodegradable poly(lactide-co-glycolide) (PLG) were subjected to laser microstructuring under a range of laser fluences (0.34–1.69 J.cm−2) to obtain surfaces with varying feature densities, roughness, and hydrophobicities.[127] NIH-3T3 fibroblasts and PC12 neuronal cells demonstrated cell type-dependent spreading and morphology which was significantly different from those on smooth, polished surfaces.[127] NIH-3T3 fibroblasts were more responsive to surface roughness compared to PC12 cells, forming a higher density of lamellipodia and focal adhesion sites on surfaces with lower roughness and appearing more round and less adhesive on surfaces with higher roughness. Considering that surface roughness and hydrophilicity are interdependent, it is necessary to attain an optimum level of roughness (and hence, hydrophilicity) through careful manipulation of laser fluence that would eventually facilitate a high degree of cell attachment. However, introduction of very sharp surface features can adversely affect cell viability in two ways: 1) reduction of cell-substrate contact and focal adhesion site formation and 2) in extreme cases, puncturing or rupturing of cell membranes.[128]
Though a majority of cell-adhesion studies have been conducted using stiff biomaterial surfaces, the principle of laser-based structuring can also be extended to softer hydrogel interfaces or ECM protein-coated substrates to interrogate cell adhesion and interactions.[129,130] Laser-based erosion or structuring has also been extended towards formation of linear or patterned features on hydrogel surfaces to guide cellular alignment, morphogenesis, and differentiation (Fig. 10C).[77] The ability to selectively modulate these features in 3D scaffolds allows for the study of dynamic cellular behavior in response to user-defined matrix microarchitectural features and potentially opens new avenues of research in cell-material interactions.
3.2 Fabrication of Scaffolds and Biomaterial Interfaces
The design and fabrication of bio-inspired scaffolds from natural, synthetic, and synthetic-natural hybrids has facilitated their use in tissue engineering applications requiring 3D cell culture, cellular infiltration, and host integration. Important criteria to consider for successful implementation of these scaffolds include cell viability and proliferation, host-biomaterial interactions, scaffold microarchitecture, and structural integrity amongst others. In this regard, laser-based degradation techniques have been suitably exploited for the fabrication and development of bioengineered scaffolds for skeletal tissue engineering, neuronal guidance, on-chip applications, and cell infiltration studies.[24,32,33,131,132] Laser-based fabrication of tissue scaffolds has been implemented two ways: 1) selective laser sintering (SLS) of microparticles to create porous, continuous scaffolds via partial melting and 2) laser-based degradation of scaffolds to create micron-scale porous features on scaffold surfaces.
In SLS, a blend of polymer microparticles is subjected to a laser beam that is selectively scanned over the powder surface at a controlled power and scan rate. Depending on the operating parameters, the laser-polymer interaction can result in nano-scale features on the polymer surface or selective melting and subsequent fusing of the polymer microparticles to form a continuous, solid mass. The microarchitectural characteristics (pore size and density, interconnectivity) of these solid scaffolds can be tuned by modulating the laser operating parameters or choosing the appropriate blend of polymeric particles that result in the desired scaffold biofunctionality. Biocomposite blends of poly(vinyl alcohol) (PVA) and hydroxyapatite (HA) were subjected to SLS treatment to generate scaffolds with defined microstructures.[33] The laser power and scan speed were varied between 4–20 W and 1270–5080 mm.s−1 respectively. Too high laser power can cause ignition of samples; hence an ideal power of 15 W was selected for SLS treatment. Tuning these parameters resulted in scaffolds that exhibited homogenous dispersion of HA microparticles in the PVA matrix (PVA acts as the polymeric blender in the biocomposite mixture) with well-defined pore interconnectivity, that promoted osteoclast proliferation for treating craniofacial and joint defects (Fig. 11A). In a similar study, polyetheretherketone (PEEK)-hydroxyapatite (HA) biocomposite blends were implemented for scaffold fabrication via SLS using laser powers ranging from 9–28 W and corresponding sintering temperatures of 110°C–140°C.[24] Optimization of these parameters and HA polymer concentration resulted in successful incorporation of HA into the polymer matrix and a high degree of pore interconnectivity. Notably, the “necking” phenomenon (fusing of adjacent polymer microparticles into a continuous mass) plays a major role in the overall polymer matrix formation and the degree of necking can be suitably modulated via the laser operating parameters to control the scaffold microarchitecture and structural integrity.
Figure 11. Laser-based modification of scaffolds and biomaterial interfaces.
(A) Selective laser sintering of hydroxyapatite(HA)/polyvinyl alcohol polymer blends fabricated using 15 W laser power and a scan speed of 1270 mm.s−1 with (a) 10 wt% HA, (b) 20 wt% HA and (c) 30 wt% HA, 100× magnification. Circles indicate HA particles. Reproduced with permission.[33] Copyright 2004, KAP. (B) Fabrication of pores via femtosecond laser ablation of electrospun poly(L-lactide) nanofibrous scaffolds for directing cell adhesion, proliferation, and infiltration. (a–f) SEM images of laser-ablated nanofibrous scaffolds of varying hole size and density (a) 50 μm diameter/50 μm spacing; (b) 50 μm diameter/200 μm spacing; (c) 100 μm diameter/50 μm spacing; (d) 100 μm diameter/200 μm spacing; (e) 200 μm diameter/50 μm spacing; and (e) 200 μm diameter/200 μm spacing. Scale bar = 200 μm. (g–l) hMSCs on corresponding ablated scaffolds labeled for actin (green) and DAPI (blue) for nuclei. (m–r) Endothelial cell infiltration into nanofibrous scaffolds in vivo after 2 weeks. White arrows indicate the location of ablated holes and white dashed lines indicate scaffold edges. Endothelial cells are labeled for CD31 (green) and all cells were counterstained with DAPI (blue) for nuclei. Scale bar = 100 μm. Reproduced with permission.[32] Copyright 2012, Elsevier. (C) Excimer laser modification of polyethersulfone hollow fibers to generate microchannels for directing the differentiation of primary adult rat neural progenitor cells (arrowheads indicate channels and white arrows indicate axonal growth into channels). (a) β-III-tubulin (red) labeled neuronal processes with Hoechst nuclei counterstain (blue), Scale bar = 100 μm. (b) SEM image, Scale bar = 100 μm; (c) SEM image, Scale bar = 50 μm; and (d) SEM image, scale bar = 5 μm. Reproduced with permission.[131] Copyright 2008, Elsevier.
Another laser-based technique for scaffold fabrication implements laser ablation to create micron-scale pores on biomaterial surfaces for cell infiltration and subsequent morphogenesis. Electrospun poly(L-lactide) (PLLA) nanofibrous scaffolds were subjected to ablation via a femtosecond pulsed laser to create structured holes for cell infiltration studies in vivo.[32] Modulation of the laser energy (10–80 μJ) and number of pulses (1–10 pulses, 35 μJ pulse energy) resulted in holes with diameters of 20–60 μm and corresponding depths of 20–120 μm. These structured scaffolds were used to study hMSC adhesion and proliferation with larger diameter holes promoting more aligned cellular morphologies. Implantation of these structured scaffolds in mice demonstrated M2 macrophage and endothelial cell infiltration through the ablated holes over two weeks, thereby establishing the role of the micron-scale pores in cellular ingrowth and scaffold remodeling (Fig. 11B).[32] This approach was also used to structure the surface of polyethersulfone (PES) hollow fibers to facilitate PC12 neuronal cell compartmentalized growth and adult rat neural progenitor cell (NPC) differentiation. Using a laser fluence of 1.2 J.cm−2 and pulse repetition of 35–50 pulses, 5 μm diameter circular channels were created on the fiber walls. The laser ablated holes facilitated axonal ingrowth of PC12 and NPC cells toward the inside fiber walls while restricting the cell bodies to the outside fiber walls thereby promoting cellular compartmentalization and neural maturation (Fig. 11C).[131]
These studies demonstrate that laser-based manipulation of biomaterial scaffolds can be utilized to generate scaffolds with controlled micro-scale features and mechanical properties. In addition to the applications discussed above, laser-structured scaffolds could potentially be used to study angiogenic ingrowth and microvascular lumen formation, fibroblast or immune cell infiltration, cancer cell invasion, and other cellular phenomena of interest.
3.3 Other Applications
Laser-based degradation and photocavitation have also been applied toward drug delivery,[37] investigation and regulation of flow in engineered constructs,[34] and miniaturized mechanical operations [133–135] amongst others.
Transport and diffusion processes within in vitro platforms are an important consideration for maintenance of cell viability and function. Microfluidic channels within these constructs are essential for adequate metabolite exchange to prevent tissue hypoxia and necrosis.[136,137] Though indirect studies of nutrient diffusion can be evaluated using cell viability, proliferation, or hypoxia assays, constructs with laser-degraded conduits that facilitate improved mass transport are preferred for creation of large-volume tissue constructs.[138,139] In this regard, laser-based degradation of hydrogels can be used to create ‘smart’ microfluidic systems for material transport and diffusion-based studies. Fluidic flow within fabricated microchannels can be tracked in real-time by imaging fluorescent particles and/or small molecules as they flow through the microchannels and diffuse into the surrounding bulk hydrogel constructs (Fig. 12A–C).[34–36,82] Transport of fluorescent moieties from one microchannel to another adjacent microchannel can also be investigated in these systems (Fig. 12A).[34] This flow phenomenon is important for our understanding of interstitial fluid flow (IFF). IFF, a critical physiological phenomenon present in healthy tissues, is driven by increased fluid pressure in the cardiovascular network and concurrent lower pressure in the lymphatic vasculature, and is a key regulator of biomolecular transport in vivo. Recapitulation of IFF through tissue engineered constructs is important towards studying the role of metabolite exchange on various cellular processes.[140,141]
Figure 12. Macromolecular flow and transport in microfluidic constructs.
(A) Flow and diffusion of dextran (red) and bovine serum albumin (BSA) (green) in two independent, yet intertwining channels as modeled through a CAD simulation and observed experimentally within laser-degraded channels in PEGDA hydrogels. Time-lapse confocal microscopy of diffusion phenomena between channels and quantification of fluorescence intensities of molecules through and between the channels over time. Reproduced with permission.[34] Copyright 2016, Wiley. (B) (a) Time-lapse of dextran flow and (b) a 10 μm polystyrene bead flow (indicated by white arrows) through a laser-degraded rectangular channel embedded in a PEGDA hydrogel. Scale bars = 50 μm. (c) 2 μm polystyrene microspheres flowing through microchannels in a PEGDA hydrogel under static and flow conditions. Scale bars = 20 μm. (d) Time-lapse of fluorescent dextran diffusing from a planar microfluidic network into the surrounding PEGDA. Scale bar = 50 μm. (White arrows indicate direction of flow in (c) and (d)). Reproduced with permission.[82] Copyright 2017, JoVE. (C) (a) Schematic of FITC-dextran diffusion through a single-source model and (b) a source-sink model. Arrows indicate direction of flow through laser-generated microchannels in PEG hydrogels (shown by white dashed lines). Corresponding time-lapse fluorescence images of diffusion and analysis of diffusion profiles for both models. (c–d) Brightfield images of superimposed and aligned microfluidic networks with flowing polystyrene microbeads through the upper and lower channels. (e–f) Evolving microfluidics via in situ formation of new microchannels to redirect flow of FITC-dextran from one channel to another. Scale bars = 100 μm. Reproduced with permission.[35] Copyright 2016, Wiley.
Additionally, laser-based degradation can be performed at varying scan speeds and intensities to degrade hydrogels to differing extents and thereby investigate the influence of matrix porosity and void volume on macromolecular transport and size-based molecular separation (Fig. 13A).[34] Changes in crosslinking density have also been altered to facilitate control of species transport (Fig. 13B).[69] These post-gelation modifications of porosity are important tools in engineering of sized-based separation of particulates and biomolecules.
Figure 13. Controlled laser-based hydrogel degradation to spatially regulate hydrogel porosity.
(A) (a–b) Control of hydrogel porosity through variation of the laser scan speed at constant fluence as evidenced through quantification of fluorescence intensities of Eosin Y, PEG-RGDS, and diffusion of 10 kDa and 2000 kDa dextran through the degraded channels. (c) Size-based separation of fluorescent species by local control of hydrogel porosity across a linear distance under a flow rate of 10 μl.min−1 and quantitative measurement of fluorescence intensities of separated species. Scale bars = 50 μm. Reproduced with permission.[34] Copyright 2016, Wiley. (B) (a) Schematic of the different modes of degradation of photolabile hydrogels via flood irradiation and their corresponding attenuated intensity profile through the depth of the hydrogel. (b) Induction of degradation gradients in photolabile hydrogels via gradient irradiation or via attenuation of flood irradiation through the hydrogel thickness over time (2.5, 5, or 8 min) and corresponding changes in normalized crosslinking densities. Scale bars = 100 μm. Reproduced with permission.[69] Copyright 2010, Wiley.
The implementation of laser-based ablation and associated thermo-mechanical events can also be extended to micro-manipulation and microfluidic operations. Two broad approaches have been employed to direct these processes: 1) generation of micron-scale features on the surface or within 3D material interfaces to enable guided manipulation of liquid behavior and 2) micro-mechanical perturbation of fluidic systems using laser-induced photocavitation and shock wave generation. As an example of the first approach, groove-like features were patterned on the surface of microfluidic channels of a PDMS slab via femtosecond laser ablation which facilitated mixing of two fluids via induction of 3D twisting co-axial flow.[142] Another group explored the potential of creating 3D high aspect ratio microchannels in glass to induce liquid mixing under laminar flow.[92,143,144] Liquid mixing via enhancement of diffusion phenomena, though made possible by laser-based microfabrication techniques, are still limited towards creation of high aspect ratio channels requiring intricate design features and complex fluid flow patterns which necessitates the generation of local turbulence and mechanical disruptions to induce rapid mixing.
In the second approach, laser-based photocavitation and subsequent bubble collapse have been implemented for turbulent fluid mixing and induction of local fluid pumping.[134,145] A focused laser beam is used to induce local thermal heating of one fluid phase leading to vapor generation and bubble formation. The mechanical shock associated with subsequent bubble collapse is exploited for directing turbulence, fluid shear stress, and mixing phenomena in very small but concentrated regions.[134] Optimization of laser power, focal position, and microfluidic design can induce variations in bubble size and dynamics, mechanical shock effects and subsequent mixing efficiency[145], which could eventually be used to control chemical reactions in small-volume systems.[133] These laser-based fluid manipulation techniques have also been applied to other processes including droplet generation via microfluidic channel fabrication in polystyrene[146] or via laser-induced photocavitation in PDMS microdevices,[135] microfluidic fluorescence activated cell sorting,[145] and characterization of protein adsorption and immunosorption kinetics within laser-ablated PET microchannels.[147]
One potential application of laser-based degradation of materials lies in selective release of entrapped biomolecules and drug delivery. Laser-based degradation for drug delivery provides controlled release of encapsulated biomolecules within polymeric scaffolds and has been applied in in vitro systems[148,149] and in cancer and ophthalmologic treatments.[37,150] Though theoretically implementable, these techniques suffer from some limitations which make it practically difficult for clinical translation. These limitations include: 1) the depth of the implanted, drug-loaded scaffold within the tissue that can still be reliably subjected to 2P degradation, 2) poor optical clarity and a high degree of scattering when passing through tissue, 3) non-specific release of entrapped drug even in the absence of laser excitation, 4) associated side-effects on surrounding healthy tissue due to repeated application of a 2P laser, and 5) optimal setup of bulky 2P laser systems with current intra-operative techniques.
Laser-based degradation, ablation, photolysis, and photocavitation have been implemented over a broad application space via rational design and implementation of laser-material interactions, optimization of laser operating parameters, and with a high degree of user-defined, spatiotemporal control. Biofabrication of microfluidic platforms, on-chip devices, and micron-scale functional units can be realized using these techniques. Micro-mechanical operations employing laser-induced mechanical and thermal excitation phenomena can facilitate fluid operation processes on small volumes and thereby miniaturize medium- or large-scale applications requiring preliminary testing at high-throughput scales.
4. Future Perspectives and Challenges
Tissue-engineered constructs employ 3D culture of hydrogel-embedded cell populations, thereby facilitating enhanced recapitulation of native microphysiology and improved in vitro functionality. Laser-based degradation, though useful in creating controlled 3D features within hydrogel matrices, could potentially present adverse side effects on encapsulated cells. Hence, optimization of laser operating parameters is necessary to maintain desired tissue function and cell viability when using laser-based degradation to structure cell-laden hydrogels.
Early studies employing laser-based degradation of cell-encapsulated hydrogels created channels adjacent to encapsulated dorsal root ganglia to encourage neuronal outgrowth in PEGylated protein hydrogels,[31,84,85,116] or from tumor spheroids to monitor collective cancer cell migration in collagen hydrogels.[41] Later attempts to create void space or microfluidic networks employed laser-based degradation to remove hydrogel (and any encapsulated cells in the focal volume) from specific regions, leaving the bulk of the material (and encapsulated cells) intact.[29] Degradation of hMSC-laden, silk-fibroin hydrogels using a Ti:S laser (wavelength: 810 nm, pulse duration: 100 fs, frequency: 80 MHz, pulse energy: 2–5 nJ) reported that cell death only occurred in the direct laser path and that cells above, below, and adjacent to the degraded volume maintained normal viability.[29] Similarly, hMSCs encapsulated in PEG hydrogels that were directly adjacent to the degraded volume also survived when using a 355 nm nanosecond pulsed laser at 0.0001 MHz.[35]
Laser-based degradation has also been used for cell surgery applications, including cytoskeletal disruption,[151] mitochondrial removal,[151,152] cell membrane perforation for DNA transfection,[153] and intranuclear scission of chromosomes.[154] The primary concern in cell surgery applications is the minimization of any generated thermal load; this can be achieved by decreasing the repetition rate to 1 MHz or less (to allow sufficient time for heat to dissipate between successive pulses), and by ensuring that the pulse energy does not exceed 2 to 3 nJ.[151] Furthermore, the criticality of the scanning mode for a particular application needs to be considered, as scanning irradiation can cause more cellular damage than locally confined irradiation.[48]
For applications involving laser-based degradation of cell-laden, non-photolabile hydrogels, the potential exposure of cells to laser-generated plasma, cavitation bubbles, shock waves, and temperature increases should be considered. As the degradation process is driven by laser-induced optical breakdown, the threshold for material degradation and cell damage are both of interest. Laser parameters including wavelength, power, scan speed, pulse frequency, pulse duration, and focusing conditions including objective magnification and NA, have a synergistic influence on the total energy delivered, and subsequently on the degradation threshold. The threshold for optical breakdown being proportional to the square root of the pulse duration,[56] lower energy is required to ablate hydrogels using femtosecond pulses rather than pico- or nanosecond pulses. Careful tuning of these parameters helps minimize cellular damage and allows cells to recover over time despite undergoing some degree of laser-induced damage.
The study of laser-based degradation and the impact of cavitation bubbles and resultant shock waves have been investigated for ophthalmic surgery to minimize collateral damage to nearby tissues.[57] Early development of ophthalmic surgery used pulsed laser sources with nanosecond pulse durations,[57] which resulted in a large diameter shock zone and cavitation bubbles on the millimeter scale, thus diminishing the surgical resolution of the technique.[58] Using picosecond (and even better, femtosecond) pulse trains, reduces the diameter of both the shock zone and cavitation bubble to the micron scale.[56,155,156] The numerical aperture of the objective also has a large impact on the degradative process. Use of a high NA objective results in a smaller focal volume, and thus, due to localized energy deposition, demands a lower amount of input energy for degradation to occur and also higher resolution features.[151]
More recently, emphasis has been placed on reducing the total energy delivered to the material, while still allowing the critical degradation threshold to be met. To illustrate, a Ti:S laser operating at 800 nm with 100 fs pulses produced pulses with energies of 1, 10, and 30 mJ; and the resultant shock wave extended 11, 17, and 20 μm, respectively from the focal volume.[57] In contrast, the pulse energy for the Ti:S laser source used in a more recent study was 37.5 nJ/pulse, or a 1×106 order decrease in the amount of energy delivered per pulse, which should minimize any resultant shock wave.[34] Degradation at this pulse energy did not extend outside the intended region.[34] Due to a wide range of potential optical systems and experimental setups available for laser-based hydrogel degradation, it is recommended that cell integrity be quantified in each scenario using a cell viability stain,[29,35] an AlamarBlue assay for metabolic activity[70] or propidium iodide to dynamically monitor nuclear and cell membrane integrity.[157,158]
To avoid any unwanted cellular exposure to cavitation bubbles or shock waves altogether, photolabile gels are preferred which undergo rapid chemical scission under very low energy compared to other degradation mechanisms. The most widely used photodegradable hydrogels employ a nitrobenzyl ether group in polymer backbone[30,69,70,83,159] that can absorb and degrade under UV flood light (with or without a photomask) and focused 1P or 2P laser sources.[70] In the presence of cells, these photolabile gels have been degraded to dynamically control cellular migration[28] and/or axonal extension from embedded clusters of cells,[71] cellular migration into or within channels,[30] cell adhesion to hydrogel surfaces,[83,160] cell spreading within 3D hydrogels,[30,69] and chondrogenic cellular differentiation.[30] These hydrogels can also be exposed to a flood source of light to recover cell populations from particular regions within a gel for further study and analysis.[28,161] A PEG-based hydrogel formulation has been developed to allow photopolymerization via efficient light-based activation of a photoinitiator above 400 nm and photodegradation via scission of a commercially-available photolabile peptide moiety when exposed to 365 nm light, thereby allowing efficient photopolymerization and photodegradation in the same hydrogel.[162]
Some additional aspects of laser-based degradation that need to be considered for practical implementation include feature resolution, scalability, reproducibility, time needed for efficient degradation, and depth of light penetration; all of which are dependent on the laser and target material characteristics. In terms of resolutions achievable, laser-based degradation techniques are comparable with other biofabrication techniques including 3D bioprinting and stereolithography. In the case of 3D bioprinting, feature resolutions vary from ~50 μm (inkjet printers) to < 5 μm (laser-based printers) based on a wide variety of currently available bioinks.[12,163,164] Current improvements in stereolithographic techniques have enabled resolutions of < 5 μm.[1,165] Laser-based degradation techniques have also facilitated feature generation with resolutions of 3–10 μm for PEGDA gels,[34,35] 1–5 μm for photolabile gels and collagen gels,[69,70,75,83] and even < 1 μm for silk and PEG-fibrinogen gels.[29,31] However, the resolution attained depend upon the laser properties and optical system (objective NA and refractive index of the medium) used, in addition to the material characteristics. The matrix characteristics including stiffness, crosslinking density, and swelling ratio may also influence the resolution and efficiency of the degradation process. Synthetic hydrogels with a low two-photon cross-section which degrade via water-dominated mechanisms may be sensitive to these factors. An increase in the relative polymer content or crosslinking density results in higher matrix stiffness and lower swelling. A reduced water content and subsequent increase in polymer density may require increased energy for degradation although these material influences have not been thoroughly investigated.
Image-guided degradation offers unique opportunities in tissue-engineering that have remained underutilized. The ability to implement user-developed CAD architectures or high-resolution 3D images of tissue have provided the ability to model lymphatic-cardiovascular transport and biomimetic microvascular networks respectively.[34] Combining image-guided degradation with microfluidic devices that provide multiple inlets and outlets could be implemented to recapitulate the multicellular architecture of complex tissues. Using non-enzymatically degradable hydrogels could allow for multiple cell types to be cultured in desired 3D spatial organizations, with each cell type confined to its own architecture, via microfluidic delivery or migration processes. The same approach could be applied using MMP-sensitive hydrogel formulations for directed self-assembly processes where cells are originally deposited in a desired 3D spatial organization at an initial time point using guidance conduits and then allowed to self-assemble into their own, but guided, final architecture. In either case, multiple rounds of image-guided degradation, followed by photopolymerization of cells suspended in a prepolymer solution flown into the degraded architectures, would allow fabrication of complex tissue constructs containing multiple cell types in physiological architectures. Additionally, this approach could allow for each cell type to be encapsulated in its own microenvironment as dictated by the composition of the prepolymer solution it was suspended and crosslinked in. These approaches to creating high-resolution, heterogeneous tissue constructs containing multiple cell types in predefined, physiological architectures could significantly advance the field of tissue engineering.
However, laser-based degradation is limited in two aspects: 1) scalability and 2) depth of light penetration (or working distance). In its present form, laser-based degradation is restricted to small volumes and surface areas depending on the laser scan speed and exposure time which limits its use in generating large-volume tissue constructs. Hence, it would be prudent to complement laser-based degradation with other biofabrication techniques such as 3D printing or stereolithography to obtain user-defined features over multiple length scales in a large volume construct in a timely and efficient manner. Additionally, laser-based degradation requires the target material to be optically transparent (particularly to the wavelength of laser light source of interest) to obtain deeper light penetration with minimal attenuation and scattering.
The scalability of laser based degradation is also associated with the time requirements of the degradation process. To provide a perspective of the time scale, consider the following three cases: 1) a 5% w/v 3.4 kDa PEGDA hydrogel (water-dominant) degraded at 790 nm using a laser power of 700 mW,[82] 2) a 2.5 mg/ml collagen I hydrogel (protein-rich) degraded at 900 nm using a laser power of 370 mW, and 3) a 10% w/v PEGdiPDA hydrogel (photolabile) degraded at 740 nm using a laser power of 100 mW.[70] The respective laser scan speeds for complete degradation in these three scenarios are 1) 20000 μm.s−1, 2) 40000 μm.s−1, and 3) 114000 μm.s−1, assuming a uniform lateral resolution of 0.18 μm.pixel−1 (unpublished data). To degrade a rectangular channel of dimensions 1000 μm × 100 μm × 100 μm, the time required to degrade the PEGDA, collagen, and PEGdiPDA hydrogels is estimated to be ~8, 4, and 1.5 minutes respectively, not including the time the laser to move between end locations during scanning, which is typically negligible if set up properly. Similarly, assuming a scale-up of the degradation process to fabricate a complex 3D cerebrovascular capillary bed, the time required for a PEGDA, collagen, and PEGdiPDA gel is approximately 1.4, 0.7, and 0.25 hours respectively.[82] As discussed earlier, the choice of laser, material, and the optical setup influence the dominant degradation mechanism and play significant roles in determining the degradation efficiency. Photolabile hydrogels tend to degrade the fastest with the highest resolution due to direct chemical scission of the polymer chains while protein-rich hydrogels degrade intermediately due to a high 2P absorption coefficient. PEGDA hydrogels, owing to their low 2P absorption coefficient, degrade indirectly via water-mediated mechanisms and hence display the slowest degradation rate and lowest resolution. Overall, for the implementation of laser-based degradation for cell and tissue engineering applications, the operating parameters must be optimized to obtain the fastest and most efficient biomaterial degradation while still maintaining high cell viability and function in the bulk construct.
5. Conclusions
Laser-based degradation of hydrogels and biomaterials provides new capabilities which are of interest in many fields of increasing attention including vascular modeling, neural regeneration, 3D microfluidics, cell-material interactions, and microphysiological models. Implementation of these techniques will require consideration and careful optimization of laser and material parameters. Tuning of these parameters should help achieve the desired degree of degradation at the fastest and most efficient manner while minimizing unwanted side effects. Though the application of laser-based degradation within hydrogels is in its nascent stage, the continued improvement in laser and optical systems, combination with other larger-scale fabrication techniques, and wide adoption for biofabrication will ultimately provide significant advancements in the field of cell and tissue engineering and regenerative medicine.
Supplementary Material
Acknowledgments
The authors thank Dr. Matt Doty and Dr. Bruce Chase for insightful conversations concerning photodegradation mechanisms and Dr. Mark Pierce for providing insights concerning general optics and laser characterization. J.L.S. is supported by a NSF IGERT Fellowship (1144726). The work of S.P., K.A.K. and J.H.S is supported by grants from the National Institutes of Health (R21CA214299), the W.M. Keck Foundation (15A00396), the Delaware Bioscience Center for Advanced Technology (15A01570), and the University of Delaware Research Foundation (17A00429).
References
- 1.Bajaj P, Schweller RM, Khademhosseini A, West JL, Bashir R. Annu Rev Biomed Eng. 2014;16:247. doi: 10.1146/annurev-bioeng-071813-105155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Guan X, Avci-Adali M, Alarçin E, Cheng H, Kashaf SS, Li Y, Chawla A, Jang HL, Khademhosseini A. Biotechnol J. 2017:1600394. doi: 10.1002/biot.201600394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Burdick JA, Murphy WL. Nat Commun. 2012;3:1269. doi: 10.1038/ncomms2271. [DOI] [PubMed] [Google Scholar]
- 4.Zhu J. Biomaterials. 2010;31:4639. doi: 10.1016/j.biomaterials.2010.02.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Tibbitt MW, Anseth KS. Biotechnol Bioeng. 2009;103:655. doi: 10.1002/bit.22361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Drury JL, Mooney DJ. Biomaterials. 2003;24:4337. doi: 10.1016/s0142-9612(03)00340-5. [DOI] [PubMed] [Google Scholar]
- 7.Slater JH, Banda OA, Heintz KA, Nie HT. In: Carbon Nanomater Biomed Appl. Zhang M, Naik RR, Dai L, editors. Springer International Publishing; 2016. pp. 543–569. [Google Scholar]
- 8.Rouwkema J, Khademhosseini A. Trends Biotechnol. 2016;34:733. doi: 10.1016/j.tibtech.2016.03.002. [DOI] [PubMed] [Google Scholar]
- 9.Annabi N, Nichol JW, Zhong X, Ji C, Koshy S, Khademhosseini A, Dehghani F. Tissue Eng Part B Rev. 2010;16:371. doi: 10.1089/ten.teb.2009.0639. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wang L, Li Y, Huang G, Zhang X, Pingguan-Murphy B, Gao B, Lu TJ, Xu F. Crit Rev Biotechnol. 2016;36:553. doi: 10.3109/07388551.2014.993588. [DOI] [PubMed] [Google Scholar]
- 11.Sant S, Hancock MJ, Donnelly JP, Iyer D, Khademhosseini A. Can J Chem Eng. 2010;88:899. doi: 10.1002/cjce.20411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Murphy SV, Atala A. Nat Biotechnol. 2014;32:773. doi: 10.1038/nbt.2958. [DOI] [PubMed] [Google Scholar]
- 13.Zhu W, Ma X, Gou M, Mei D, Zhang K, Chen S. Curr Opin Biotechnol. 2016;40:103. doi: 10.1016/j.copbio.2016.03.014. [DOI] [PubMed] [Google Scholar]
- 14.Culver JC, Hoffmann JC, Poché RA, Slater JH, West JL, Dickinson ME. Adv Mater. 2012;24:2344. doi: 10.1002/adma.201200395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Barry RA, Shepherd RF, Hanson JN, Nuzzo RG, Wiltzius P, Lewis JA. Adv Mater. 2009;21:2407. [Google Scholar]
- 16.Cerchiari A, Garbe JC, Todhunter ME, Jee NY, Pinney JR, LaBarge MA, Desai TA, Gartner ZJ. Tissue Eng Part C Methods. 2015;21:541. doi: 10.1089/ten.tec.2014.0450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Golden AP, Tien J. Lab Chip. 2007;7:720. doi: 10.1039/b618409j. [DOI] [PubMed] [Google Scholar]
- 18.Franco CL, Price J, West JL. Acta Biomater. 2011;7:3267. doi: 10.1016/j.actbio.2011.06.011. [DOI] [PubMed] [Google Scholar]
- 19.Pradhan S, Clary JM, Seliktar D, Lipke EA. Biomaterials. 2017;115:141. doi: 10.1016/j.biomaterials.2016.10.052. [DOI] [PubMed] [Google Scholar]
- 20.van Duinen V, Trietsch SJ, Joore J, Vulto P, Hankemeier T. Curr Opin Biotechnol. 2015;35:118. doi: 10.1016/j.copbio.2015.05.002. [DOI] [PubMed] [Google Scholar]
- 21.Bhatia SN, Ingber DE. Nat Biotechnol. 2014;32:760. doi: 10.1038/nbt.2989. [DOI] [PubMed] [Google Scholar]
- 22.Doraiswamy A, Patz T, Narayan RJ, Dinescu M, Modi R, Auyeung RCY, Chrisey DB. Appl Surf Sci. 2006;252:4748. [Google Scholar]
- 23.Schaffer CB, Brodeur A, García JF, Mazur E. Opt Lett. 2001;26:93. doi: 10.1364/ol.26.000093. [DOI] [PubMed] [Google Scholar]
- 24.Tan K, Chua C, Leong K, Cheah C, Cheang P, Abu Bakar M, Cha S. Biomaterials. 2003;24:3115. doi: 10.1016/s0142-9612(03)00131-5. [DOI] [PubMed] [Google Scholar]
- 25.Chen F, Zhang D, Yang Q, Yong J, Du G, Si J, Yun F, Hou X. ACS Appl Mater Interfaces. 2013;5:6777. doi: 10.1021/am401677z. [DOI] [PubMed] [Google Scholar]
- 26.Yalcin HC, Shekhar A, Nishimura N, Rane AA, Schaffer CB, Butcher JT. AJP Heart Circ Physiol. 2010;299:H1728. doi: 10.1152/ajpheart.00495.2010. [DOI] [PubMed] [Google Scholar]
- 27.Nishimura N, Schaffer CB, Friedman B, Tsai PS, Lyden PD, Kleinfeld D. Nat Methods. 2006;3:99. doi: 10.1038/nmeth844. [DOI] [PubMed] [Google Scholar]
- 28.DeForest CA, Anseth KS. Nat Chem. 2011;3:925. doi: 10.1038/nchem.1174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Applegate MB, Coburn J, Partlow BP, Moreau JE, Mondia JP, Marelli B, Kaplan DL, Omenetto FG. Proc Natl Acad Sci. 2015;112:12052. doi: 10.1073/pnas.1509405112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kloxin AM, Kasko AM, Salinas CN, Anseth KS. Science. 2009;324:59. doi: 10.1126/science.1169494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sarig-Nadir O, Livnat N, Zajdman R, Shoham S, Seliktar D. Biophys J. 2009;96:4743. doi: 10.1016/j.bpj.2009.03.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lee BLP, Jeon H, Wang A, Yan Z, Yu J, Grigoropoulos C, Li S. Acta Biomater. 2012;8:2648. doi: 10.1016/j.actbio.2012.04.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Chua CK, Leong KF, Tan KH, Wiria FE, Cheah CM. J Mater Sci Mater Med. 2004;15:1113. doi: 10.1023/B:JMSM.0000046393.81449.a5. [DOI] [PubMed] [Google Scholar]
- 34.Heintz KA, Bregenzer ME, Mantle JL, Lee KH, West JL, Slater JH. Adv Healthc Mater. 2016 doi: 10.1002/adhm.201600351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Brandenberg N, Lutolf MP. Adv Mater. 2016 doi: 10.1002/adma.201601099. [DOI] [PubMed] [Google Scholar]
- 36.Arakawa CK, Badeau BA, Zheng Y, DeForest CA. Adv Mater. :1703156. doi: 10.1002/adma.201703156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Lambertz A, Klink CD, Röth A, Schmitz D, Pich A, Feher K, Bremus-Köbberling E, Neumann UP, Junge K. J Surg Res. 2014;192:312. doi: 10.1016/j.jss.2014.07.036. [DOI] [PubMed] [Google Scholar]
- 38.Rossier JS, Bercier P, Schwarz A, Loridant S, Girault HH. Langmuir. 1999;15:5173. [Google Scholar]
- 39.Urech L, Lippert T. In: Photochem Photophysics Polym Mater. Allen NS, editor. John Wiley And Sons; 2010. [Google Scholar]
- 40.Yong J, Chen F, Yang Q, Du G, Bian H, Zhang D, Si J, Yun F, Hou X. ACS Appl Mater Interfaces. 2013;5:9382. doi: 10.1021/am402923t. [DOI] [PubMed] [Google Scholar]
- 41.Ilina O, Bakker GJ, Vasaturo A, Hoffman RM, Friedl P. Phys Biol. 2011;8:029501. doi: 10.1088/1478-3975/8/1/015010. [DOI] [PubMed] [Google Scholar]
- 42.Sugio Y, Kojima K, Moriguchi H, Takahashi K, Kaneko T, Yasuda K. Sens Actuators B Chem. 2004;99:156. [Google Scholar]
- 43.Moriguchi H, Takahashi K, Sugio Y, Wakamoto Y, Inoue I, Jimbo Y, Yasuda K. Electr Eng Jpn. 2004;146:37. [Google Scholar]
- 44.Markus L, Dmitri O, Jons H, Peter G, Katja H, Marica M, Jurgen S, Robert L, Aleksandr O. Front Bioeng Biotechnol. 2016;4 doi: 10.3389/conf.FBIOE.2016.01.02498. [DOI] [Google Scholar]
- 45.Maximova K, Wang X, Balčytis A, Fan L, Li J, Juodkazis S. Biomicrofluidics. 2016;10:054101. doi: 10.1063/1.4962294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Ovsianikov A, Mironov V, Stampfl J, Liska R. Expert Rev Med Devices. 2012;9:613. doi: 10.1586/erd.12.48. [DOI] [PubMed] [Google Scholar]
- 47.Zhang W, Chung PH, Zhang A, Chen S. In: Laser Technol Biomim. Schmidt V, Belegratis MR, editors. Springer Berlin Heidelberg; Berlin, Heidelberg: 2013. pp. 237–257. [Google Scholar]
- 48.Vogel A, Noack J, Hüttman G, Paltauf G. Appl Phys B. 2005;81:1015. [Google Scholar]
- 49.Vogel A, Venugopalan V. Chem Rev. 2003;103:577. doi: 10.1021/cr010379n. [DOI] [PubMed] [Google Scholar]
- 50.Müller G, Dörschel K, Kar H. Lasers Med Sci. 1991;6:241. [Google Scholar]
- 51.Vogel A, Noack J, Nahen K, Theisen D, Busch S, Parlitz U, Hammer DX, Noojin GD, Rockwell BA, Birngruber R. Appl Phys B Lasers Opt. 1999;68:271. [Google Scholar]
- 52.Hammer DX, Jansen ED, Frenz M, Noojin GD, Thomas RJ, Noack J, Vogel A, Rockwell BA, Welch AJ. Appl Opt. 1997;36:5630. doi: 10.1364/ao.36.005630. [DOI] [PubMed] [Google Scholar]
- 53.Schaffer CB. Meas Sci Technol. 2001;12 [Google Scholar]
- 54.Mironi-Harpaz I, Hazanov L, Engel G, Yelin D, Seliktar D. Adv Mater. 2015;27:1933. doi: 10.1002/adma.201404185. [DOI] [PubMed] [Google Scholar]
- 55.Joglekar AP, Liu H-h, Meyhofer E, Mourou G, Hunt AJ. Proc Natl Acad Sci. 2004;101:5856. doi: 10.1073/pnas.0307470101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Juhasz T, Kastis GA, Suárez C, Bor Z, Bron WE. Lasers Surg Med. 1996;19:23. doi: 10.1002/(SICI)1096-9101(1996)19:1<23::AID-LSM4>3.0.CO;2-S. [DOI] [PubMed] [Google Scholar]
- 57.Glezer EN, Schaffer CB, Nishimura N, Mazur E. Opt Lett. 1997;22:1817. doi: 10.1364/ol.22.001817. [DOI] [PubMed] [Google Scholar]
- 58.Vogel A, Busch S, Parlitz U. J Acoust Soc Am. 1996;100:148. [Google Scholar]
- 59.Zipfel WR, Williams RM, Webb WW. Nat Biotechnol. 2003;21:1369. doi: 10.1038/nbt899. [DOI] [PubMed] [Google Scholar]
- 60.So PT, Dong CY, Masters BR, Berland KM. Annu Rev Biomed Eng. 2000;2:399. doi: 10.1146/annurev.bioeng.2.1.399. [DOI] [PubMed] [Google Scholar]
- 61.Svoboda K, Yasuda R. Neuron. 2006;50:823. doi: 10.1016/j.neuron.2006.05.019. [DOI] [PubMed] [Google Scholar]
- 62.Mulligan SJ, Macvicar BA. Two-Photon Fluorescence Microscopy: Basic Principles, Advantages and Risks. n.d. [Google Scholar]
- 63.Garen W, Hegedűs F, Kai Y, Koch S, Meyerer B, Neu W, Teubner U. Shock Waves. 2016;26:385. [Google Scholar]
- 64.Hansen A, Géneaux R, Günther A, Krüger A, Ripken T. Biomed Opt Express. 2013;4:852. doi: 10.1364/BOE.4.000852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Zysset B, Fujimoto JG, Deutsch TF. Appl Phys B. 1989;48:139. [Google Scholar]
- 66.Eickmans JH, Hsieh WF, Chang RK. Opt Lett. 1987;12:22. doi: 10.1364/ol.12.000022. [DOI] [PubMed] [Google Scholar]
- 67.Noack J, Vogel A. Appl Opt. 1998;37:4092. doi: 10.1364/ao.37.004092. [DOI] [PubMed] [Google Scholar]
- 68.Gill SC, von Hippel PH. Anal Biochem. 1989;182:319. doi: 10.1016/0003-2697(89)90602-7. [DOI] [PubMed] [Google Scholar]
- 69.Kloxin AM, Tibbitt MW, Kasko AM, Fairbairn JA, Anseth KS. Adv Mater. 2010;22:61. doi: 10.1002/adma.200900917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Kloxin AM, Tibbitt MW, Anseth KS. Nat Protoc. 2010;5:1867. doi: 10.1038/nprot.2010.139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.McKinnon DD, Brown TE, Kyburz KA, Kiyotake E, Anseth KS. Biomacromolecules. 2014;15:2808. doi: 10.1021/bm500731b. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Tibbitt MW, Kloxin AM, Sawicki LA, Anseth KS. Macromolecules. 2013;46:2785. doi: 10.1021/ma302522x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Wong DY, Griffin DR, Reed J, Kasko AM. Macromolecules. 2010;43:2824. [Google Scholar]
- 74.Liu Y, Sun S, Singha S, Cho MR, Gordon RJ. Biomaterials. 2005;26:4597. doi: 10.1016/j.biomaterials.2004.11.033. [DOI] [PubMed] [Google Scholar]
- 75.Smith NI, Fujita K, Nakamura O, Kawata S. Appl Phys Lett. 2001;78:999. [Google Scholar]
- 76.Hribar KC, Meggs K, Liu J, Zhu W, Qu X, Chen S. Sci Rep. 2015;5:17203. doi: 10.1038/srep17203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Daskalova A, Nathala CSR, Kavatzikidou P, Ranella A, Szoszkiewicz R, Husinsky W, Fotakis C. Appl Surf Sci. 2016;382:178. [Google Scholar]
- 78.Oujja M, Pérez S, Fadeeva E, Koch J, Chichkov BN, Castillejo M. Appl Phys Lett. 2009;95:263703. [Google Scholar]
- 79.Qian Z, Mordovanakis A, Schoenly JE, Covarrubias A, Feng Y, Lilge L, Marjoribanks RS. Biomed Opt Express. 2014;5:208. doi: 10.1364/BOE.5.000208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.De Maria C, Grassi L, Vozzi F, Ahluwalia A, Vozzi G. Rapid Prototyp J. 2014;20:490. [Google Scholar]
- 81.Applegate MB, Alonzo C, Georgakoudi I, Kaplan DL, Omenetto FG. Appl Phys Lett. 2016;108:241903. [Google Scholar]
- 82.Heintz KA, Mayerich D, Slater JH. J Vis Exp. 2017;119 doi: 10.3791/55101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Tibbitt MW, Kloxin AM, Dyamenahalli KU, Anseth KS. Soft Matter. 2010;6:5100. doi: 10.1039/C0SM00174K. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Livnat N, Sarig-Nadir O, Seliktar D, Shoham S. 2009 4th Int. IEEEEMBS Conf. Neural Eng; IEEE; 2009. pp. 116–119. [Google Scholar]
- 85.Berkovitch Y, Yelin D, Seliktar D. Eur Polym J. 2015;72:473. [Google Scholar]
- 86.Seck TM, Melchels FPW, Feijen J, Grijpma DW. J Controlled Release. 2010;148:34. doi: 10.1016/j.jconrel.2010.07.111. [DOI] [PubMed] [Google Scholar]
- 87.Slater JH, Frey W. J Biomed Mater Res A. 2008;87A:176. doi: 10.1002/jbm.a.31725. [DOI] [PubMed] [Google Scholar]
- 88.Shukla A, Slater JH, Culver JC, Dickinson ME, West JL. ACS Appl Mater Interfaces. 2016;8:21883. doi: 10.1021/acsami.5b08978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Slater JH, Culver JC, Long BL, Hu CW, Hu J, Birk TF, Qutub AA, Dickinson ME, West JL. ACS Nano. 2015;9:6128. doi: 10.1021/acsnano.5b01366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Slater JH, Boyce PJ, Jancaitis MP, Gaubert HE, Chang AL, Markey MK, Frey W. ACS Appl Mater Interfaces. 2015;7:4390. doi: 10.1021/am508906f. [DOI] [PubMed] [Google Scholar]
- 91.Slater JH, West JL. Methods Cell Biol. 2014;119:193. doi: 10.1016/B978-0-12-416742-1.00011-1. [DOI] [PubMed] [Google Scholar]
- 92.Masuda M, Sugioka K, Cheng Y, Aoki N, Kawachi M, Shihoyama K, Toyoda K, Helvajian H, Midorikawa K. Appl Phys Mater Sci Process. 2003;76:857. [Google Scholar]
- 93.Gattass RR, Mazur E. Nat Photonics. 2008;2:219. [Google Scholar]
- 94.Gittard SD, Narayan RJ. Expert Rev Med Devices. 2010;7:343. doi: 10.1586/erd.10.14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Vu LT, Jain G, Veres BD, Rajagopalan P. Tissue Eng Part B Rev. 2015;21:67. doi: 10.1089/ten.teb.2013.0782. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Pradhan S, Hassani I, Seeto WJ, Lipke EA. J Biomed Mater Res A. 2017;105:236. doi: 10.1002/jbm.a.35899. [DOI] [PubMed] [Google Scholar]
- 97.Pradhan S, Hassani I, Clary JM, Lipke EA. Tissue Eng Part B Rev. 2016;22:470. doi: 10.1089/ten.TEB.2015.0567. [DOI] [PubMed] [Google Scholar]
- 98.Zhu X, Gojgini S, Chen TH, Fei P, Dong S, Ho CM, Segura T. J Biol Eng. 2017;11:12. doi: 10.1186/s13036-017-0055-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Cuchiara MP, Gould DJ, McHale MK, Dickinson ME, West JL. Adv Funct Mater. 2012;22:4511. doi: 10.1002/adfm.201200976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Chen TH, Zhu X, Pan L, Zeng X, Garfinkel A, Tintut Y, Demer LL, Zhao X, Ho CM. Biomaterials. 2012;33:9019. doi: 10.1016/j.biomaterials.2012.08.067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Kachouie NN, Du Y, Bae H, Khabiry M, Ahari AF, Zamanian B, Fukuda J, Khademhosseini A. Organogenesis. 2010;6:234. doi: 10.4161/org.6.4.12650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Lu P, Weaver VM, Werb Z. J Cell Biol. 2012;196:395. doi: 10.1083/jcb.201102147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Bray LJ, Binner M, Holzheu A, Friedrichs J, Freudenberg U, Hutmacher DW, Werner C. Biomaterials. 2015;53:609. doi: 10.1016/j.biomaterials.2015.02.124. [DOI] [PubMed] [Google Scholar]
- 104.Song HHG, Park KM, Gerecht S. Adv Drug Deliv Rev. 2014;79–80:19. doi: 10.1016/j.addr.2014.06.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Gencoglu MF, Barney LE, Hall CL, Brooks EA, Schwartz AD, Corbett DC, Stevens KR, Peyton SR. ACS Biomater Sci Eng. 2017 doi: 10.1021/acsbiomaterials.7b00069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Raeber GP, Lutolf MP, Hubbell JA. Biophys J. 2005;89:1374. doi: 10.1529/biophysj.104.050682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Paul CD, Hung WC, Wirtz D, Konstantopoulos K. Annu Rev Biomed Eng. 2016;18:159. doi: 10.1146/annurev-bioeng-071114-040654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Chung IM, Enemchukwu NO, Khaja SD, Murthy N, Mantalaris A, García AJ. Biomaterials. 2008;29:2637. doi: 10.1016/j.biomaterials.2008.03.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Georgiou M, Bunting SCJ, Davies HA, Loughlin AJ, Golding JP, Phillips JB. Biomaterials. 2013;34:7335. doi: 10.1016/j.biomaterials.2013.06.025. [DOI] [PubMed] [Google Scholar]
- 110.Winter CC, Katiyar KS, Hernandez NS, Song YJ, Struzyna LA, Harris JP, Cullen DK. Acta Biomater. 2016;38:44. doi: 10.1016/j.actbio.2016.04.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Turunen S, Haaparanta AM, Äänismaa R, Kellomäki M. J Tissue Eng Regen Med. 2013;7:253. doi: 10.1002/term.520. [DOI] [PubMed] [Google Scholar]
- 112.Hosokawa C, Kudoh SN, Kiyohara A, Hosokawa Y, Okano K, Masuhara H, Taguchi T. Appl Phys A. 2008;93:57. [Google Scholar]
- 113.Shepard JA, Stevans AC, Holland S, Wang CE, Shikanov A, Shea LD. Biotechnol Bioeng. 2012;109:830. doi: 10.1002/bit.24355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Musoke-Zawedde P, Shoichet MS. Biomed Mater. 2006;1:162. doi: 10.1088/1748-6041/1/3/011. [DOI] [PubMed] [Google Scholar]
- 115.Moriguchi H, Wakamoto Y, Sugio Y, Takahashi K, Inoue I, Yasuda K. Lab Chip. 2002;2:125. doi: 10.1039/b202569h. [DOI] [PubMed] [Google Scholar]
- 116.Livnat N, Sarig-Nadir O, Zajdman R, Seliktar D, Shoham S. 2007 3rd Int. IEEEEMBS Conf. Neural Eng; IEEE; 2007. pp. 101–103. [Google Scholar]
- 117.Sarig-Nadir O, Seliktar D. Tissue Eng Part A. 2008;14:401. doi: 10.1089/tea.2007.0029. [DOI] [PubMed] [Google Scholar]
- 118.Miller JS, Stevens KR, Yang MT, Baker BM, Nguyen DHT, Cohen DM, Toro E, Chen AA, Galie PA, Yu X, Chaturvedi R, Bhatia SN, Chen CS. Nat Mater. 2012;11:768. doi: 10.1038/nmat3357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Leong MF, Toh JKC, Du C, Narayanan K, Lu HF, Lim TC, Wan ACA, Ying JY. Nat Commun. 2013;4:2353. doi: 10.1038/ncomms3353. [DOI] [PubMed] [Google Scholar]
- 120.Blinder YJ, Freiman A, Raindel N, Mooney DJ, Levenberg S. Sci Rep. 2015;5:17840. doi: 10.1038/srep17840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Wong KHK, Truslow JG, Khankhel AH, Chan KLS, Tien J. J Biomed Mater Res A. 2013;101A:2181. doi: 10.1002/jbm.a.34524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Kim J, Tanner K. Front Oncol. 2015;5 doi: 10.3389/fonc.2015.00170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Jeon JS, Bersini S, Gilardi M, Dubini G, Charest JL, Moretti M, Kamm RD. Proc Natl Acad Sci. 2015;112:214. doi: 10.1073/pnas.1417115112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Iino T, Hagiyama M, Furuno T, Ito A, Hosokawa Y. Biophys J. 2016;111:2255. doi: 10.1016/j.bpj.2016.09.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Hao L, Lawrence J, Phua YF, Chian KS, Lim GC, Zheng HY. J Biomed Mater Res B Appl Biomater. 2005;73B:148. doi: 10.1002/jbm.b.30194. [DOI] [PubMed] [Google Scholar]
- 126.Martínez-Calderon M, Manso-Silván M, Rodríguez A, Gómez-Aranzadi M, García-Ruiz JP, Olaizola SM, Martín-Palma RJ. Sci Rep. 2016;6 doi: 10.1038/srep36296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Koufaki N, Ranella A, Aifantis KE, Barberoglou M, Psycharakis S, Fotakis C, Stratakis E. Biofabrication. 2011;3:045004. doi: 10.1088/1758-5082/3/4/045004. [DOI] [PubMed] [Google Scholar]
- 128.Ranella A, Barberoglou M, Bakogianni S, Fotakis C, Stratakis E. Acta Biomater. 2010;6:2711. doi: 10.1016/j.actbio.2010.01.016. [DOI] [PubMed] [Google Scholar]
- 129.Paital SR, Cao Z, He W, Dahotre NB. Biofabrication. 2010;2:025001. doi: 10.1088/1758-5082/2/2/025001. [DOI] [PubMed] [Google Scholar]
- 130.Fadeeva E, Deiwick A, Chichkov B, Schlie-Wolter S. Interface Focus. 2013;4:20130048. doi: 10.1098/rsfs.2013.0048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Brayfield CA, Marra KG, Leonard JP, Tracy Cui X, Gerlach JC. Acta Biomater. 2008;4:244. doi: 10.1016/j.actbio.2007.10.004. [DOI] [PubMed] [Google Scholar]
- 132.Uzel SGM, Platt RJ, Subramanian V, Pearl TM, Rowlands CJ, Chan V, Boyer LA, So PTC, Kamm RD. Sci Adv. 2016;2:e1501429. doi: 10.1126/sciadv.1501429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Hellman AN, Rau KR, Yoon HH, Bae S, Palmer JF, Phillips KS, Allbritton NL, Venugopalan V. Anal Chem. 2007;79:4484. doi: 10.1021/ac070081i. [DOI] [PubMed] [Google Scholar]
- 134.Dijkink R, Ohl CD. Lab Chip. 2008;8:1676. doi: 10.1039/b806912c. [DOI] [PubMed] [Google Scholar]
- 135.Park SY, Wu TH, Chen Y, Teitell MA, Chiou PY. Lab Chip. 2011;11:1010. doi: 10.1039/c0lc00555j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Bettinger CJ, Borenstein JT. Soft Matter. 2010;6:4999. [Google Scholar]
- 137.Annabi N, Tamayol A, Uquillas JA, Akbari M, Bertassoni LE, Cha C, Camci-Unal G, Dokmeci MR, Peppas NA, Khademhosseini A. Adv Mater. 2014;26:85. doi: 10.1002/adma.201303233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Choi NW, Cabodi M, Held B, Gleghorn JP, Bonassar LJ, Stroock AD. Nat Mater. 2007;6:908. doi: 10.1038/nmat2022. [DOI] [PubMed] [Google Scholar]
- 139.Cuchiara MP, Allen ACB, Chen TM, Miller JS, West JL. Biomaterials. 2010;31:5491. doi: 10.1016/j.biomaterials.2010.03.031. [DOI] [PubMed] [Google Scholar]
- 140.Pisano M, Triacca V, Barbee KA, Swartz MA. Integr Biol. 2015;7:525. doi: 10.1039/c5ib00085h. [DOI] [PubMed] [Google Scholar]
- 141.Wiig H, Swartz MA. Physiol Rev. 2012;92:1005. doi: 10.1152/physrev.00037.2011. [DOI] [PubMed] [Google Scholar]
- 142.Wolfe DB, Ashcom JB, Hwang JC, Schaffer CB, Mazur E, Whitesides GM. Adv Mater. 2003;15:62. [Google Scholar]
- 143.Liao Y, Song J, Li E, Luo Y, Shen Y, Chen D, Cheng Y, Xu Z, Sugioka K, Midorikawa K. Lab Chip. 2012;12:746. doi: 10.1039/c2lc21015k. [DOI] [PubMed] [Google Scholar]
- 144.Liu C, Liao Y, He F, Shen Y, Chen D, Cheng Y, Xu Z, Sugioka K, Midorikawa K. Opt Express. 2012;20:4291. doi: 10.1364/OE.20.004291. [DOI] [PubMed] [Google Scholar]
- 145.Wu TH, Chen Y, Park SY, Hong J, Teslaa T, Zhong JF, Di Carlo D, Teitell MA, Chiou PY. Lab Chip. 2012;12:1378. doi: 10.1039/c2lc21084c. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Li H, Fan Y, Kodzius R, Foulds IG. Microsyst Technol. 2012;18:373. [Google Scholar]
- 147.Rossier JS, Gokulrangan G, Girault HH, Svojanovsky S, Wilson GS. Langmuir. 2000;16:8489. [Google Scholar]
- 148.Griffin DR, Schlosser JL, Lam SF, Nguyen TH, Maynard HD, Kasko AM. Biomacromolecules. 2013;14:1199. doi: 10.1021/bm400169d. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Griffin DR, Kasko AM. ACS Macro Lett. 2012;1:1330. doi: 10.1021/mz300366s. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Kim HC, Härtner S, Behe M, Behr TM, Hampp NA. J Biomed Opt. 2006;11:034024. doi: 10.1117/1.2209564. [DOI] [PubMed] [Google Scholar]
- 151.Shen N, Datta D, Schaffer CB, LeDuc P, Ingber DE, Mazur E. Mech Chem Biosyst MCB. 2005;2:17. [PubMed] [Google Scholar]
- 152.Watanabe W, Arakawa N, Matsunaga S, Higashi T, Fukui K, Isobe K, Itoh K. Opt Express. 2004;12:4203. doi: 10.1364/opex.12.004203. [DOI] [PubMed] [Google Scholar]
- 153.Tirlapur UK, König K. Nature. 2002;418:290. doi: 10.1038/418290a. [DOI] [PubMed] [Google Scholar]
- 154.Koenig K, Riemann I, Fischer P, Halbhuber K. Cell Mol Biol. 1999;45:195. [PubMed] [Google Scholar]
- 155.Noack J, Hammer DX, Noojin GD, Rockwell BA, Vogel A. J Appl Phys. 1998;83:7488. [Google Scholar]
- 156.Vogel A, Linz N, Freidank S, Paltauf G. Phys Rev Lett. 2008;100 doi: 10.1103/PhysRevLett.100.038102. [DOI] [PubMed] [Google Scholar]
- 157.Yuan F, Yang C, Zhong P. Proc Natl Acad Sci. 2015;112:E7039. doi: 10.1073/pnas.1518679112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Takizawa N, Okano K, Uwada T, Hosokawa Y, Masuhara H. In: Jacques SL, Roach WP, Thomas RJ, editors. 2008. p. 685411. [Google Scholar]
- 159.Peng K, Tomatsu I, van den Broek B, Cui C, Korobko AV, van Noort J, Meijer AH, Spaink HP, Kros A. Soft Matter. 2011;7:4881. [Google Scholar]
- 160.Pasparakis G, Manouras T, Selimis A, Vamvakaki M, Argitis P. Angew Chem. 2011;123:4228. doi: 10.1002/anie.201007310. [DOI] [PubMed] [Google Scholar]
- 161.Griffin DR, Kasko AM. J Am Chem Soc. 2012;134:13103. doi: 10.1021/ja305280w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162.Ki CS, Shih H, Lin CC. Polymer. 2013;54:2115. doi: 10.1016/j.polymer.2013.02.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Li J, Chen M, Fan X, Zhou H. J Transl Med. 2016;14 doi: 10.1186/s12967-016-1028-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Wang Z, Abdulla R, Parker B, Samanipour R, Ghosh S, Kim K. Biofabrication. 2015;7:045009. doi: 10.1088/1758-5090/7/4/045009. [DOI] [PubMed] [Google Scholar]
- 165.Raman R, Bhaduri B, Mir M, Shkumatov A, Lee MK, Popescu G, Kong H, Bashir R. Adv Healthc Mater. 2016;5:610. doi: 10.1002/adhm.201500721. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.