Abstract
Head and neck squamous cell carcinoma (HNSCC) is the sixth most common cause of cancer mortality in the world. Some progress has been made in the therapy of HNSCC, however treatment remains unsatisfactory. Recent studies have shown that different types of long non-coding RNAs (lncRNAs) are dysregulated in HNSCC and correlate with tumor progression, lymph node metastasis, clinical stage and poor prognosis. lncRNAs are a class of functional RNA molecules that can not be translated into proteins but can modulate the activity of transcription factors or regulate changes in chromatin structure. The lncRNAs might have potential of biomarker in HNSCC diagnosis, prognosis, prediction and targeted treatment. In this review we describe the potential role of lncRNAs as new biomarkers and discuss their features including source of origin, extraction methods, stability, detection methods and data normalization and potential function as biomarkers in HNSCC.
Keywords: HNSCC, biomarkers, lncRNA, head and neck
Head and neck cancers
Head and neck squamous cell carcinoma (HNSCC) including tumors that occur in the oral cavity is the sixth most common cancer and one of the most common cause of cancer mortality worldwide. Tobacco smoking, alcohol consumption and human papilloma virus (HPV) or Epstein-Barr virus (EBV) infections are the main causes of these malignancies. HNSCC often develops within pre-neoplastic fields of genetically altered cells. HNSCC is divided into many types according to tumor localization: tongue squamous cell carcinoma (TSCC), oral squamous cell carcinoma (OSCC), laryngeal squamous cell carcinoma (LSCC) and nasopharyngeal carcinoma (NPC). For the last decade it has been mainly treated by surgical resection, radical radio(chemo)therapy or systemic treatment alone (e.g. cetuximab, cisplatin, 5-fluorouracyl or taxanes). The 5-year survival rate has persisted at approximately 43%. The most cases are not early diagnosed until cancer metastases to the regional lymph nodes of the neck what influences on the patients’ survival rate [1–4]. Recently pembrolizumab (anty-PD1) has been approved by the U.S. Food and Drug Administration in the second line treatment in patients with advanced HNSCC [5]. However, treatment results remain unsatisfactory despite these efforts. A high proportion of patients who do not respond to standard treatment could get a benefit from personalized therapy based on the molecular diagnostic or targeted therapies. Many studies have shown abnormal changes of many types of RNA (coding and non-coding) in HNSCC patients, which have pivotal role in the cancer biology. This suggests that coding and non-coding RNAs can serve as biomarkers for treatment response prediction or as diagnostic tools [1, 6–8]. However, the role of long non-coding RNA (lncRNA) are still not deeply understood.
lncRNA– biogenesis, function and role in cancer
Long non-coding RNAs are a class of functional RNA molecules that are not translated into proteins and consist of at least 200 nucleotides [9, 10]. However, this remains debatable, and some scientists postulate that lncRNAs have limited protein-coding ability – some transcripts can function in dual roles both as coding and non-coding RNA [11, 12]. It is thought that 93% of the human genome can be transcribed into RNAs [13]. Approximately 2% of these transcripts will be translated into proteins, and the remaining 98% – called non-coding RNAs (ncRNAs)-will rarely be transcribed [14].
The ncRNAs are classified into two groups. The first group is called constitutive RNA and it includes transfer RNAs (tRNAs), ribosome RNAs (rRNAs), small nuclear RNAs (snRNAs) and small nucleolar RNAs (snoRNAs). The second group, called regulatory RNAs, consists of small interfering RNAs (siRNAs), piRNAs, microRNAs (miRNAs), natural antisense transcripts (NATs), circular RNAs (circRNAs) and long non-coding RNAs (lncRNAs) [15–17]. All ncRNAs, except for tRNAs and rRNAs, are considered “transcriptional noise” [18]. However, lncRNAs are highly transcribed and believed to play roles in more complex biological functions, i.e. regulation of gene expression at the transcriptional level in nucleus (chromatin regulation, alternative splicing of pre-mRNA, DNA demethylation and nuclear organization) or posttranscriptional level in the cytoplasm [18]. It should be noted that large proportions of lncRNAs are closely connected with genes encoded near specific mRNAs and these ‘lncRNA-mRNA pairs’ influence of each other [19, 20]. Guigo’s group reported that lncRNAs exhibit unusual exonic structure and can be alternatively spliced [21]. Another classification of ncRNAs is based on their position relative to protein coding genes: intergenic, intragenic/intronic and anti-sense [9].
The location in the genome (overlapping with possibly important genes) and difficulties in finding an appropriate model make functional study of lncRNAs challenging. The choice of a suitable model is a problematic issue because of the lack of conservation at the nucleotide sequence level [16], tissue specific expression level [22], transcription initiation from regions rich in repeats [23] and mostly high isoform heterogeneity. The lncRNA isoforms can have different functions [24]. Moreover, recent studies have shown that lncRNAs may have cell-type specificity [25], and their function should be verified in different cell models.
Despite the fact that the vast majority of long non-coding RNAs remain functionally uncharacterized, some of them have been linked with a range biological processes including chromatin modification, regulation of transcription factors, mRNA processing and degradation as well as cell signaling [26]. They also have a vital role in cellular processing, and their deregulated expression has been associated with different types of cancers [27, 28]. Even though many cancer gene-profiling studies have revealed some cancer-associated lncRNAs, there are very few lncRNAs reported for HNSCC.
Features of lncRNA molecules and methods of analysis
Early diagnosis of cancer results in more effective therapy, and biomarkers to predict and monitor treatment response are urgently needed [29, 30]. The use of DNA or RNA as a potential biomarker is not innovative, but there are not many DNA or RNA-based markers translated to clinics. Detection of abnormal expression of lncRNAs from tissue, blood or urine samples seems to be easily performed using molecular biology methods nowadays [27, 31, 32].
An ideal biomarker should be simple obtained from the diverse of sources and require simple measurement methodology [33]. However in the case of lncRNAs some problematic questions have arisen and they need to be clarified before implementation of these molecules in the clinical diagnosis.
First of all, it is thought that good biomarker should be easily available. lncRNAs are present in tissue, peripheral blood, serum, saliva, urine or some cell-derived exosomes [31, 32, 34, 35–37], but not all lncRNAs are present in every type of biological material. For example, Tang et al. observed the presence of HOTAIR, HULC, MALAT1, MEG-3, NEAT-1 and UCA1 in malignant and adjacent nonmalignant samples from OSCC patients, but in saliva only HOTAIR and MALAT1 were detectable [36].
High quantity and quality of biomarker molecules are also important. lncRNAs are supposed to be less stable and easier to degrade than miRNAs due to their length. However, Kraus et al. showed in their studies of postmortem brain tissues that some lncRNAs are more stable than miRNAs [38, 39]. Others also indicated that most lncRNAs are stable (half-life more than 16 h) – especially intragenic and cis-antisense lncRNAs compared with those derived from introns [10, 36]. The specific lncRNA half-life depends not only on its coding place in the genome and posttranscriptional modifications but also on subcellular localization and function [10]. Moreover, the presence of some lncRNAs in body fluids such as saliva [36], and resistance of plasma lncRNAs to RNase A digestion and overnight incubation at room temperature [37] confirm high stability of these transcripts. On the other hand, analysis of both long coding and non-coding RNA transcripts obtained from archived formalin-fixed paraffin-embedded (FFPE) blocks is difficult because of their low stability [40]. However, this problem can be solved by measuring the expression level of lncRNA by real-time PCR reaction with three different pairs of non-overlapping primers [41].
The next issue refers to the standardization of material sample and the RNA isolation method. There is a lack of specific methods to sample and store material and methodological differences occur. One can compare cancer tissue with adjacent non-cancer samples from the same patient or with samples from healthy donors without a history of cancer. In our opinion, adjacent non-cancer samples are not good reference because of the risk of tumor influence or inflammation. There is also a lack of RNA isolation methods dedicated to lncRNA analysis. lncRNA from tissue and cell lines is usually extracted using standard methods for total RNA isolation based on classical TRIzol or column-based methods [42]. The isolation method seems not affect lncRNA quantification results, but there is no available data supporting this statement. However, column-based approach seems to be better than TRIzol extraction especially in the case of RNA extraction from body fluid [43]. For circulating lncRNA, the sample choice, handling and processing as well as contamination of blood cells influence the sample preparation. Due to coagulation and hemolysis, blood cells release lncRNAs into the serum affecting the results [37, 43, 44]. However, the use of special blood collection tubes can minimize the level of background RNA and eliminate false results under quantification [45].
The detection of lncRNA, its quantification and determination of the transcriptional activity of the lncRNA gene (methylation) should be also considered. There are many methods to determine these: i) lncRNA immunoprecipitation; ii) lncRNA in-situ hybridization; iii) Au-NP assay (gold nanoparticle-based); iv) lncRNA northern blot analysis; v) methylation status using HRM (High Resolution Melting); vi) microarray or RNA sequencing; vii) and the most widely used qRT-PCR or new developed ddPCR [31, 32, 46–48]. The choice of proper analysis method depends on kind of study (screening or specific detection), type of material source and costs.
The most common methods in lncRNA studies are hybridization assays especially qRT-PCR. The available qRT-PCR lncRNA platforms allows simple and quick quantification of 90 lncRNAs based on CT analysis in one run, while one commercial lncRNA microarray platform can check the expression of more than 30,000 lncRNAs without sophisticated bioinformatics methods required for NGS (Next Generation Sequencing) data extraction [49, 50]. Moreover, microarray experiments seem to be more precise because of the well-validated technology, in contrast to lab-designed qRT-PCR primers, which can differ among laboratories [51]. In addition, the presence of lncRNA isoforms and their polymorphisms influences the function of specific lncRNA [52–54], but there is a lack of information about specific studies. This can make it difficult to compare results.
Microarray or NGS methods are expensive and data analyzing is demanding and probably they will be used only in biomarker research area. The simplicity of performing and low cost as well as availability of lncRNA quantification kits with well-defined workflow seem to make qRT-PCR as a gold-standard of lncRNA quantification.
The most popular qRT-PCR method used in lncRNA research is based on SYBR-Green dye and TaqMan probes [42]. qRT-PCR assay requires the right choice of cDNA synthesis method and the proper reference genes. There are no standardized methods for reverse transcription of lncRNA. Some lncRNAs have endogenous polyA tails but others do not possess these elements. Moreover, most lncRNA is present in low copy numbers, and this makes it difficult to quantitate with conventional methods. These lncRNA require the addition of polyA tails and annealing anchor dT adapters before cDNA synthesis. This approach allows enhance specificity and sensitivity of lncRNA quantification [55]. However, in most studies, the cDNA is prepared using kits containing mixtures of oligo(dT) and random hexamer primers.
Another very important issue is the preparation of RNA to cDNA synthesis, particularly circulating RNA. Qi et al. noted that quantification of circulating RNAs via the NanoDrop spectrophotometer is problematic. They recommended using the same volume of input rather than the same amounts of RNA. However, they showed no evidence supporting this statement [43].
The use of proper reference gene is still problematic in lncRNA expression measurement using qRT-PCR. There is a lack of standardized references, and most lncRNA studies are based on GAPDH, U6 (RNU6B) or ACTB [42]. The mismatched reference influences the results and makes it difficult to compare various studies. The problems with the normalization were observed in the case of miRNA expression studies. The snoRNAs used as normalization for miRNAs are not stably expressed and actually could serve as prognostic factors [51]. This situation suggests that proper normalization is an important step in data presentation and comparison. It should be verified if different types of tissue need specific normalization genes for examination of lncRNA. For example, different lncRNA references are suitable only for brain tissue studies. Some can be applied as universal references in profiling various gliomas and normal tissues [38, 39]. Thus, it should be verified if different cancers need specific lncRNA-related normalizing genes [43]. Fang et al. checked 16 different reference genes regarding cancer, normal and metastasis tissues (such as ACTB, TUBA3, KALPHA1, GAPDH or B2M); for example ACTB was selected as the best normalizer for MALAT1 [55]. One of the open questions is normalization of circulating lncRNA data. Dong et al. verified the utility of ACTB, GAPDH, HPRT, 18S RNA, CYC, and GUSB as the reference genes in the serum of healthy and cancer patients, and ACTB was selected as the best normalizing gene. Moreover, ACTB is stable after temperature changes in serum samples [56].
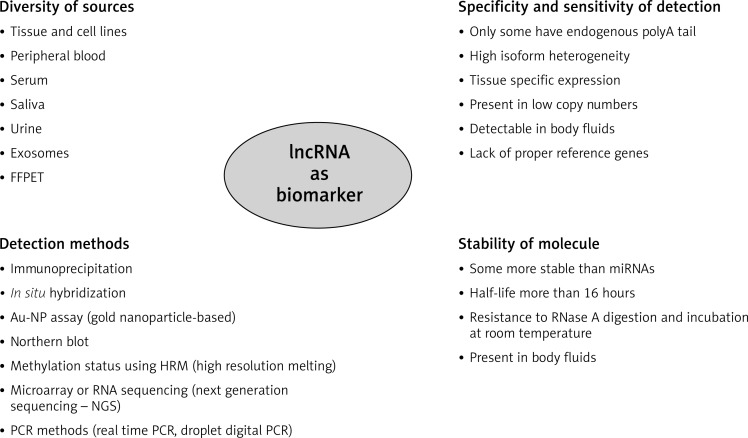
Despite the numerous problems, lncRNA under proper laboratory conditions seems to be a good candidate as biomarker and this statement is proven by many diagnostic studies [43]. The features of lncRNA molecules as biomarker were summarized in Fig. 1. However, standardization of procedures and definition of specific expression profiles bearing specific clinical information before using lncRNA as biomarkers are urgently needed and are the challenge for the future studies.
Fig. 1.
Characteristic of lncRNA molecules as potential biomarker
lncRNA biomarkers in HNSCC
Many studies have shown that deregulated expression of lncRNAs can be associated with diabetes [57], leukemia [58], solid cancers [59] or other diseases such as endometriosis [60]. The potential role of lncRNA as biomarkers in cancer such as gastric, colorectal, prostate, lung cancer or HNSCC has already been described [27, 42]. While HOTAIR is deregulated in many cancer types, a few lncRNAs are deregulated in a particular cancer type, i.e. prostate cancer antigen 3 (PCA3) is found only in prostate tissue [61]. The results are promising because PCA3 may be potentially used as a cancer-type specific biomarker.
To date, only nine independent studies show global analysis of lncRNA expression profile in HNSCC: three bioinformatic analysis of available data, four experimental microarray studies and two experimental next generation sequencing studies (Table 1).
Table 1.
Characteristic of global lncRNA profiling studies in HNSCC
| Study | Analysis | Results description |
|---|---|---|
| Zou et al. [62, 63] | Bioinformatic analysis of RNA-seq data sets (TCGA data from UCSC Cancer Genomics Hub) of 40-tumour-adjacent normal pairs and 363 additional unpaired tumors |
|
| Nohata et al. [64] | Bioinformatic analysis of RNA-seq data sets (TCGA data from The Atlas of Noncoding RNAs in Cancer – TANRIC) of 468 tumor samples Analysis of sequencing data of OPC-22 panel (cell lines) |
|
| Yang and Deng [65] | Microarray analysis (mRNA and lncRNA) of 6 pairs of NPC and chronic nasopharyngitis (CNP) samples and qRT-PCR validation |
|
| Zhang et al. [69] | Microarray analysis of randomly paired 3 metastatic and 4 primary NPC tumor samples and qRT-PCR validation |
|
| Zhou et al. [74] | Microarray analysis of 3 paired tumor and adjacent noncancerouse samples from hypopharyngeal squamose cell carcinoma (HSCC) patients and qRT-PCR validation | |
| Ren et al. [70] | Next generation sequencing and qRT-PCR validation |
|
| Zhang et al. [66] | Bioinformatic analysis of microarray data sets (GSE25099 from Gene Expression Omnibus database) of 57 OSCC samples and 22 normal sample |
|
| Li et al. [73] | Next generation sequencing and qRT-PCR of radio-resistant CNE-2-Rs and parental CNE-2 cell lines (nasopharengynal) and qRT-PCR validation |
|
| Zhang et al. [75] | Microarray analysis (lncRNA and mRNA) of 7 NPC tumor samples and adjacent non-tumor samples and qRT-PCR validation |
|
Profiling studies revealed that specific lncRNAs expression in cancer tissue is associated with HPV status, known mutations, cancer-related pathways and gene copy number changes [62–66]. We need to remember, that specific global expression analysis is based on the use of some bioinformatics tools [49, 50] and the results should be verified by different methodologies. Thus, most of these studies are not validated using different types of samples or in vitro models. Moreover, the inaccuracies of the results may be caused by differences in examined groups or samples (such as anatomical sites) reflecting genetic diversity. Biological role of only a few lncRNAs dysregulated in HNSCC is well known. The most studied lncRNAs both in vivo and in vitro in HNSCC are HOTAIR, HOTTIP, UCA1, LET, MEG3, MALAT1, H19 and NAG7. They are involved in many important cellular processes such as proliferation, migration and invasion, apoptosis or phenotype regulation. Their exact function in biology of HNSCC has been carefully described by us elsewhere [67].
However, some lncRNAs have a strong prognostic ability for overall survival, disease-free survival, or recurrence-free survival in HNSCC. These lncRNAs described as potential biomarkers are presented in Table 2. Some of lncRNAs are proposed to be independent of gender, organ site, tumor stage or TP53 status [64, 66, 68]. lncRNAs can also be used as virus infection indicators [63], while others may serve as metastasis or disease progression markers [69].
Table 2.
lncRNAs described as potential biomarkers in HNSCC
| lncRNA | Description | Ref. |
|---|---|---|
| NEAT-1 |
|
[62, 36, 75, 76] |
| HOTAIR |
|
[36, 77–81] |
| HOTTIP |
|
[32] |
| UCA1 |
|
[36, 82] |
| AC026166.2-001 & RP11-169D4.1-001 |
|
[83] |
| GAS5 |
|
[62, 35, 51] |
| lnc-JPHl-7 |
|
[63] |
| LET |
|
[68] |
| lncRNA-ROR |
|
[72] |
| XIST |
|
[84] |
| MEG3 |
|
[36, 85] |
| MALAT1 |
|
[36, 55, 71, 86] |
| H19 |
|
[87] |
| lincRNA NAG7 (LINC00312) |
|
[12] |
| NKILA |
|
[88] |
Unfortunately, there is only one study indicating lncRNAs as biomarkers related to response to chemoradiotherapy. Fayda et al. showed that only plasma circulating GAS5 might be a useful predictive biomarker [35]. However, this study is based only on a small group of patients, and these results should be verified in a randomized trial before clinical use [43]. However, expression of lncRNA changes after exposure to chemotherapeutic drugs, and this could maintain drug resistance or sensitivity in cancer cell lines [70–72].
Only one study has indicated a role of lncRNA in radioresistant NPC cell lines. Li et al. used NGS technology and showed some previously known and some novel lncRNAs are dysregulated in radioresistant cell (Rs) line compared to the parental line. Three pairs of lncRNA-mRNA in CNE-2-Rs and 6-10B-Rs cell lines have been described. The discovered lncRNAs: n373932, n409627 and n386034, regulate SLITRK5, PRSS12 and RIMKLB mRNAs, respectively. Only CNE-2-Rs cells show a slight change in lncRNAs-mRNAs. In the case of 6-10B-RSs, there is strong down-regulation of n373932 lncRNA and up-regulation of STITRK5 mRNA. Moreover, the expression of n373932 and SLITRK5 is negatively correlated in NPC patients [73]. More in vivo and in vitro studies are needed to define exact role of specific lncRNA in chemo- and radioresponse and next validation of predictive lncRNA panel in clinical practice.
Conclusions and future perspectives
lncRNAs, including the ones described here, are aberrantly expressed in a number of different cancers and were characterized as molecules with a great impact. These recently discovered RNA molecules affect the hallmarks of carcinogenesis including proliferation, metastasis and apoptosis. Moreover, existing reports indicate the potential role of lncRNA as a new class of biomarkers. However, there are some challenges and problems to take and solve before use. First of all, there is only a few data regarding specific lncRNA in HNSCC. Despite the fact that several studies have shown global changes in lncRNA expression profile in HNSCC, the validated panel was not proposed. Moreover, most studies are based on a small study group, and it is difficult to compare them to TCGA-based analyzes. Some of lncRNAs described in this review play pivotal roles in HNSCC and might be biomarkers of treatment response. However, only a few studies have focused on lncRNA after irradiation or chemoexposure. The exact role of specific lncRNAs in regulation and response to radiation and chemotherapy used in HNSCC is unknown and requires future studies. The next issue is lack of well-validated methodology to use lncRNA in diagnostics such as different detection methods as well as use of non-specific or unsuitable normalization genes, which influence on the results.
We supposed, that future cancer diagnostic panels will likely consist of a wide variety of genes-both protein coding and non-coding. Such a platform will offer a detailed description of the nature of the tumor, and this will allow personalized treatment of HNSCC.
The examples presented here are just the tip of the iceberg. Knowledge about lncRNA is evolving. A lot of work was done to explain the role of this molecules in the cancer biology, but much more should be do to use this knowledge in clinical practice.
Footnotes
This work was supported by Greater Poland Cancer Centre – grant No.: 21/2015 (113) and grant No.: 13/2016 (128). English language revision and editing was made by American Manuscript Editors – americanmanuscripteditors.com.
The authors declare no conflict of interest.
References
- 1.Zhi IX, Lamperska K, Golusinski P, Schork NJ, Luczewski L, Kolenda T, et al. Gene expression analysis of head and neck squamous cell carcinoma survival and recurrence. Oncotarget. 2015;6:547–55. doi: 10.18632/oncotarget.2772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Tsang CM, Tsao SW. The role of Epstein-Barr virus infection in the pathogenesis of nasopharyngeal carcinoma. Virol Sin. 2015;30:107–21. doi: 10.1007/s12250-015-3592-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Marur S, Forastiere AA. Head and Neck Squamous Cell Carcinoma: Update on Epidemiology, Diagnosis, and Treatment. Mayo Clin Proc. 2016;91:386–96. doi: 10.1016/j.mayocp.2015.12.017. [DOI] [PubMed] [Google Scholar]
- 4.van Oijen MG, Slootweg PJ. Oral field cancerization: carcinogen – induced independent events or micrometastatic deposits? Cancer Epidemiol Biomarkes Prev. 2000;9:249–56. [PubMed] [Google Scholar]
- 5.Fuereder T. Immunotherapy for head and neck squamous cell carcinoma. Memo. 2016;9:66–9. doi: 10.1007/s12254-016-0270-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Victoria Martinez B, Dhahbi JM, et al. Circulating small non coding RNA signature in head and neck squamous cell carcinoma. Oncotarget. 2015;6:19246–63. doi: 10.18632/oncotarget.4266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lamperska K, Kozlowski P, Kolenda T, et al. Unpredictable changes of selected miRNA in expression profile of HNSCC. Cancer Biomark. 2016;16:55–64. doi: 10.3233/CBM-150540. [DOI] [PubMed] [Google Scholar]
- 8.Kolenda T, Przybyła W, Teresiak A, Mackiewicz A, Lamperska K. The mystery of let-7d – a small RNA with great power. Contemp Oncol (Pozn) 2014;18:293–301. doi: 10.5114/wo.2014.44467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009;10:155–9. doi: 10.1038/nrg2521. [DOI] [PubMed] [Google Scholar]
- 10.Clark MB, Johnston RL, Inostroza-Ponta M, Fox AH, Fortini E, Moscato P, Dinger ME, Mattick JS. Genome – wide analysis of long noncoding RNA stability. Genome Res. 2012;22:885–98. doi: 10.1101/gr.131037.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Dinger ME, Pang KC, Mercer TR, Mattick JS. Differentiating protein coding and noncoding RNA: challenges and ambiguities. PLoS Comput Biol. 2008;4:e1000176. doi: 10.1371/journal.pcbi.1000176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Zhang W, Huang C, Gong Z, et al. Expression of LINC00312, a long intergenic non-coding RNA, is negatively correlated with tumor size but positively correlated with lymph node metastasis in nasopharyngeal carcinoma. J Mol Histol. 2013;44:545–54. doi: 10.1007/s10735-013-9503-x. [DOI] [PubMed] [Google Scholar]
- 13.Birney E, Stamatoyannopoulos JA, Dutta A, Guigó R, Gingeras TR, et al. ENCODE Project Consortium Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature. 2007;447:799–816. doi: 10.1038/nature05874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Carninci P, Kasukawa T, Katayama S, et al. The transcriptional landscape of the mammalian genome. Science. 2005;309:1559–63. doi: 10.1126/science.1112014. [DOI] [PubMed] [Google Scholar]
- 15.Zhang Y, Yang L, Chen LL. Life without a tail: new formats of long noncoding RNAs. Int J Biochem Cell Biol. 2014;54:338–49. doi: 10.1016/j.biocel.2013.10.009. [DOI] [PubMed] [Google Scholar]
- 16.Furuno M, Pang KC Ninomiya N, et al. Clusters of internally primed transcripts reveal novel long noncoding RNAs. PLoS Genet. 2006;2:e37. doi: 10.1371/journal.pgen.0020037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Memczak S, Jens M, Elefsinioti A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495:333–8. doi: 10.1038/nature11928. [DOI] [PubMed] [Google Scholar]
- 18.Kun Y, Arfat Y, Li D, Zhao F, et al. Structure Prediction: New Insights into Decrypting Long Noncoding RNAs. Int J Mol Sci. 2016;17:132.. doi: 10.3390/ijms17010132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kong XP, Yao J, Luo W, Feng FK, Ma JT, Ren YP, Wang DL, Bu RF. The expression and functional role of a FOXC1 related mRNA-lncRNA pair in oral squamous cell carcinoma. Mol Cell Biochem. 2014;394:177–86. doi: 10.1007/s11010-014-2093-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Sigova AA, Mullen AC, Molinie B, et al. Divergent transcription of long noncoding RNA/mRNA gene pairs in embryonic stem cells. Proc Natl Acad Sci U S A. 2013;110:2876–81. doi: 10.1073/pnas.1221904110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Derrien T, Johnson R, Bussotti G, et al. The GENCODE v7 catalog of human long noncoding RNAs: Analysis of their gene structure, evolution, and expression. Genome Res. 2012;22:1775–89. doi: 10.1101/gr.132159.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Cabili MN, Trapnell C, Goff L, Koziol M, Koziol M, Tazon-Vega B, Regev A, Rinn JL. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 2011;25:1915–27. doi: 10.1101/gad.17446611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kelley D, Rinn J. Transposable elements reveal a stem cell-specific class of long noncoding RNAs. Genome Biol. 2012;13:R107. doi: 10.1186/gb-2012-13-11-r107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hoffmann MJ, Dehn J, Droop J, et al. Truncated Isoforms of lncRNA ANRIL Are Overexpressed in Bladder Cancer, But Do Not Contribute to Repression of INK4 Tumor Suppressors. Non-Coding RNA. 2015;1:266–84. doi: 10.3390/ncrna1030266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Huarte M, Guttman M, Feldser D, et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell. 2010;142:409–19. doi: 10.1016/j.cell.2010.06.040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Chang HY, Schmitt AM. Long Noncoding RNAs in Cancer Pathways. Cancer Cell. 2016;29:452–63. doi: 10.1016/j.ccell.2016.03.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Yarmishyn AA, Kurochkin IV. Long noncoding RNAs: a potential novel class of cancer biomarkers. Front Genet. 2015;6:145. doi: 10.3389/fgene.2015.00145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Jiang C, Li X, Zhao H, Liu H. Long non-coding RNAs: potential new biomarkers for predicting tumor invasion and metastasis. Mol Cancer. 2016;15:62. doi: 10.1186/s12943-016-0545-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Ahlawat P, Rawat S, Kakria A, Pal M, Chauhan D, Tandon S, Jain S. Tumour volumes: Predictors of early treatment response in locally advanced head and neck cancers treated with definitive chemoradiation. Rep Pract Oncol Radiother. 2016;21:419–26. doi: 10.1016/j.rpor.2016.04.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.González Ferreira JA, Jaén Olasolo J, Azinovic I, Jeremic B. Effect of radiotherapy delay in overall treatment time on local control and survival in head and neck cancer: Review of the literature. Rep Pract Oncol Radiother. 2015;20:328–39. doi: 10.1016/j.rpor.2015.05.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Eissa S, Matboli M, Essawy NO, Shehta M, Kotb YM. Rapid detection of urinary long non-coding RNA urothelial carcinoma associated one using a PCR-free nanoparticle-based assay. Biomarkers. 2015;20:212–7. doi: 10.3109/1354750X.2015.1062918. [DOI] [PubMed] [Google Scholar]
- 32.Zhang H, Zhao L, Wang YX, Xi M, Liu SL, Luo LL. Long non-coding RNA HOTTIP is correlated with progression and prognosis in tongue squamous cell carcinoma. Tumour Biol. 2015;36:8805–9. doi: 10.1007/s13277-015-3645-2. [DOI] [PubMed] [Google Scholar]
- 33.Kolenda T, Teresiak A, Kapałczyńska M, Przybyła W, Zajączkowska M, Bliźniak R, Masternak MM, Golusinski P, Golusinski W. Let-7d and miR-18a as biomarkers of head and neck cancers. Lett Oncol Sci. 2015;12:37–47. [Google Scholar]
- 34.Gezer U, Özgür E, Cetinkaya M, Isin M, Dalay N. Long non-coding RNAs with low expression levels in cells are enriched in secreted exosomes. Cell Biol Int. 2014;38:1076–9. doi: 10.1002/cbin.10301. [DOI] [PubMed] [Google Scholar]
- 35.Fayda M, Isin M, Tambas M, et al. Do circulating long non-coding RNAs (lncRNAs) (LincRNA-p21, GAS 5, HOTAIR) predict the treatment response in patients with head and neck cancer treated with chemoradiotherapy? Tumour Biol. 2016;37:3969–78. doi: 10.1007/s13277-015-4189-1. [DOI] [PubMed] [Google Scholar]
- 36.Tang H, Wu Z, Zhang J, Su B. Salivary lncRNA as a potential marker for oral squamous cell carcinoma diagnosis. Mol Med Rep. 2013;7:761–6. doi: 10.3892/mmr.2012.1254. [DOI] [PubMed] [Google Scholar]
- 37.Zhou X, Yin C, Dang Y, Ye F, Zhang G. Identification of the long non-coding RNA H19 in plasma as a novel biomarker for diagnosis of gastric cancer. Sci Rep. 2015;5:11516. doi: 10.1038/srep11516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kraus TF, Greiner A, Guibourt V, Lisec K, Kretzschmar HA. Identification of Stably Expressed lncRNAs as Valid Endogenous Controls for Profiling of Human Glioma. J Cancer. 2015;6:111–9. doi: 10.7150/jca.10867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kraus TF, Greiner A, Guibourt V, Kretzschmar HA. Long non-coding RNA normalisers in human brain tissue. J Neural Transm (Vienna) 2015;122:1045–54. doi: 10.1007/s00702-014-1352-6. [DOI] [PubMed] [Google Scholar]
- 40.Kokkat TJ, Patel MS, McGarvey D, LiVolsi VA, Baloch ZW. Archived formalin-fixed paraffin-embedded (FFPE) blocks: A valuable underexploited resource for extraction of DNA, RNA, and protein. Biopreserv Biobank. 2013;11:101–6. doi: 10.1089/bio.2012.0052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kong H, Zhu M, Cui F, Wang S, Gao X, Lu S, Wu Y, Zhu H. Quantitative assessment of short amplicons in FFPE-derived long-chain RNA. Sci Rep. 2014;4:7246. doi: 10.1038/srep07246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Shi T, Gao G, Cao Y. Long Noncoding RNAs as Novel Biomarkers Have a Promising Future in Cancer Diagnostics. Dis Markers. 2016;2016:9085195. doi: 10.1155/2016/9085195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Qi P, Zhou XY, Du X. Circulating long non-coding RNAs in cancer: current status and future perspectives. Mol Cancer. 2016;15:39. doi: 10.1186/s12943-016-0524-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Pritchard CC, Kroh E, Wood B, Arroyo JD, Dougherty KJ, Miyaji MM, Tait JF, Tewari M. Blood cell origin of circulating microRNAs: a cautionary note for cancer biomarker studies. Cancer Prev Res (Phila) 2012;5:492–7. doi: 10.1158/1940-6207.CAPR-11-0370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Qin J, Williams TL, Fernando MR. A novel blood collection device stabilizes cell-free RNA in blood during sample shipping and storage. BMC Res Notes. 2013;6:380. doi: 10.1186/1756-0500-6-380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Feng Y, Hu X, Zhang Y, Zhang D, Li C, Zhang L. Methods for the Study of Long Noncoding RNA in Cancer Cell Signaling Methods. Methods Mol Biol. 2014;1165:115–43. doi: 10.1007/978-1-4939-0856-1_10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Wojdacz TK, Dobrovic A, Algar EM. Rapid detection of methylation change at H19 in human imprinting disorders using methylation-sensitive high-resolution melting. Hum Mutat. 2008;29:1255–60. doi: 10.1002/humu.20779. [DOI] [PubMed] [Google Scholar]
- 48.Dodd DW, Gagnon KT, Corey DR. Digital quantitation of potential therapeutic target RNAs. Nucleic Acid Ther. 2013;23:188–94. doi: 10.1089/nat.2013.0427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Oleksiewicz U, Tomczak K, Woropaj J, Markowska M, Stępniak P, Shah PK. Computational characterisation of cancer molecular profiles derived using next generation sequencing. Contemp Oncol (Pozn) 2015;19(1A):A78–A91. doi: 10.5114/wo.2014.47137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Tomczak K, Czerwińska P, Wiznerowicz M. The Cancer Genome Atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol (Pozn) 2015;19(1A):A68–77. doi: 10.5114/wo.2014.47136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Gee HE, Buffa FM, Camps C. The small-nucleolar RNAs commonly used for microRNA normalisation correlate with tumour pathology and prognosis. Br J Cancer. 2011;104:1168–77. doi: 10.1038/sj.bjc.6606076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Desai S, Ding M, Wang B, Lu Z, Zhao Q, Shaw K, et al. Tissue-specific isoform switch and DNA hypomethylation of the pyruvate kinase PKM gene in human cancers. Oncotarget. 2014;5:8202–10. doi: 10.18632/oncotarget.1159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Pang KC, Frith MC, Mattick JS. Rapid evolution of noncoding RNAs: lack of conservation does not mean lack of function. Trends Genet. 2006;22:1–5. doi: 10.1016/j.tig.2005.10.003. [DOI] [PubMed] [Google Scholar]
- 54.Hüttenhofer A, Schattner P, Polacek N. Non-coding RNAs: hope or hype? Trends Genet. 2005;21:289–97. doi: 10.1016/j.tig.2005.03.007. [DOI] [PubMed] [Google Scholar]
- 55.Fang Z, Zhang S, Wang Y, et al. Long non-coding RNA MALAT-1 modulates metastatic potential of tongue squamous cell carcinomas partially through the regulation of small proline rich proteins. BMC Cancer. 2016;16:706. doi: 10.1186/s12885-016-2735-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Dong L, Qi P, Xu MD, et al. Circulating CUDR, LSINCT-5 and PTENP1 long noncoding RNAs in sera distinguish patients with gastric cancer from healthy controls. Int J Cancer. 2015;137:1128–35. doi: 10.1002/ijc.29484. [DOI] [PubMed] [Google Scholar]
- 57.Reddy MA, Chen Z, Park JT, et al. Regulation of inflammatory phenotype in macrophages by a diabetes-induced long noncoding RNA. Diabetes. 2014;63:4249–61. doi: 10.2337/db14-0298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Trimarchi T, Bilal E, Ntziachristos P, Fabbri G, Dalla-Favera R, Tsirigos A, Aifantis I. Genome-wide mapping and characterization of Notch-regulated long noncoding RNAs in acute leukemia. Cell. 2014;158:593–606. doi: 10.1016/j.cell.2014.05.049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Huarte M. The emerging role of lncRNAs in cancer. Nature Med. 2015;21:1253–61. doi: 10.1038/nm.3981. [DOI] [PubMed] [Google Scholar]
- 60.Wang WT, Sun YM, Huang W, He B, Zhao YN, Chen YQ. Genome-wide long non-coding RNA analysis identified circulating LncRNAs as novel non-invasive diagnostic biomarkers for gynecological disease. Sci Rep. 2016;6:23343. doi: 10.1038/srep23343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Bussemakers MJ, van Bokhoven A, Verhaegh GW, et al. DD3: a new prostate-specific gene, highly overexpressed in prostate cancer. Cancer Res. 1999;59:5975–9. [PubMed] [Google Scholar]
- 62.Zou AE, Ku J, Honda TK, et al. Transcriptome sequencing uncovers novel long noncoding and small nucleolar RNAs dysregulated in head and neck squamous cell carcinoma. RNA. 2015;21:1122–34. doi: 10.1261/rna.049262.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Zou AE, Zheng H, Saad MA, Rahimy M, Ku J, Kuo SZ, et al. The non-coding landscape of head and neck squamous cell carcinoma. Oncotarget. 2016;7:51211–22. doi: 10.18632/oncotarget.9979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Nohata N, Abba MC, Gutkind JS. Unraveling the oral cancer lncRNAome: Identification of novel lncRNAs associated with malignant progression and HPV infection. Oral Oncol. 2016;59:58–66. doi: 10.1016/j.oraloncology.2016.05.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Yang QQ, Deng YF. Genome-wide analysis of long non-coding RNA in primary nasopharyngeal carcinoma by microarray. Histopathology. 2015;66:1022–30. doi: 10.1111/his.12616. [DOI] [PubMed] [Google Scholar]
- 66.Zhang S, Tian L, Ma P, Sun Q, Zhang K, Liu H, et al. Potential role of differentially expressed lncRNAs in the pathogenesis of oral squamous cell carcinoma. Arch Oral Biol. 2015;60:1581–7. doi: 10.1016/j.archoralbio.2015.08.003. [DOI] [PubMed] [Google Scholar]
- 67.Kolenda T, Guglas K, Ryś M, Bogaczyńska M, Teresiak A, Bliźniak R, et al. Biological role of lncRNA in head and neck cancers. Rep Pract Oncol Radiother. 2017;22:378–388. doi: 10.1016/j.rpor.2017.07.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Sun Q, Liu H, Li L, Zhang S, Liu K, Liu Y, Yang C. Long noncoding RNA-LET, which is repressed by EZH2, inhibits cell proliferation and induces apoptosis of nasopharyngeal carcinoma cell. Med Oncol. 2015;32:226.. doi: 10.1007/s12032-015-0673-0. [DOI] [PubMed] [Google Scholar]
- 69.Zhang W, Wang L, Zheng F, et al. Long Noncoding RNA Expression Signatures of Metastatic Nasopharyngeal Carcinoma and Their Prognostic Value. BioMed Res Intl. 2015;2015:618924. doi: 10.1155/2015/618924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Ren S, Li G, Liu C, et al. Next generation deep sequencing identified a novel lncRNA n375709 associated with paclitaxel resistance in nasopharyngeal carcinoma. Oncol Rep. 2016;36:1861–7. doi: 10.3892/or.2016.4981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Chen H, Xin Y, Zhou L, Huang JM, Tao L, Cheng L, Tian J. Cisplatin and paclitaxel target significant long noncoding RNAs in laryngeal squamous cell carcinoma. Med Oncol. 2014;31:246.. doi: 10.1007/s12032-014-0246-7. [DOI] [PubMed] [Google Scholar]
- 72.Li L, Gu M, You B, Shi S, Shan Y, Bao L, You Y. Long non-coding RNA ROR promotes proliferation, migration and chemoresistance of nasopharyngeal carcinoma. Cancer Sci. 2016;107:1215–22. doi: 10.1111/cas.12989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Li G, Liu Y, Liu C, et al. Genome – wide analyses of long noncoding RNA expression profiles correlated with radioresistance in nasopharyngeal carcinoma via nest – generation deep sequencing. BMC Cancer. 2016;16:719. doi: 10.1186/s12885-016-2755-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Zhou J, Li M, Yu W, et al. AB209630, a long non-coding RNA decreased expression in hypopharyngeal squamous cell carcinoma, influences proliferation, invasion, metastasis and survival. Oncotarget. 2016;7(12):14628–38. doi: 10.18632/oncotarget.7403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Zhang B, Wang D, Wu J, et al. Expression profiling and functional prediction of long noncoding RNAs in nasopharyngeal nonkeratinizing carcinoma. Discov Med. 2016;21:239–50. [PubMed] [Google Scholar]
- 76.Wang P, Wu T, Zhou H, et al. Long noncoding RNA NEAT1 promotes laryngeal squamous cell cancer through regulating miR-107/CDK6 pathway. J Exp Clin Cancer Res. 2016;35:22.. doi: 10.1186/s13046-016-0297-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Gonzalez-Ramirez I, Soto-Reyes E, Sanchez-Perez Y, Herrera LA, Garcia-Cuellar C. Histones and long non-coding RNAs: The new insight of epigenetic deregulation involved in oral cancer. Oral Oncol. 2014;50:691–5. doi: 10.1016/j.oraloncology.2014.04.006. [DOI] [PubMed] [Google Scholar]
- 78.Wu Y, Zhang L, Wang Y, et al. Long non-coding RNA HOTAIR promotes tumor cell invasion and metastasis by recruiting EZH2 and repressing E-cadherin in oral squamous cell carcinoma. Int J Oncol. 2015;46:2586–94. doi: 10.3892/ijo.2015.2976. [DOI] [PubMed] [Google Scholar]
- 79.Wang J, Zhou Y, Lu J, Sun Y, Xiao H, Liu M, Tian L. Combined detection of serum exosomal miR-21 and HOTAIR as diagnostic and prognostic biomarkers for laryngeal squamous cell carcinoma. Med Oncol. 2014;31:148.. doi: 10.1007/s12032-014-0148-8. [DOI] [PubMed] [Google Scholar]
- 80.Nie Y, Liu X, Qu S, Song E, Zou H, Gong C. Long non-coding RNA HOTAIR is an independent prognostic marker for nasopharyngeal carcinoma progression and survival. Cancer Sci. 2013;104:458–64. doi: 10.1111/cas.12092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Fu WM, Lu YF, Hu BG, et al. Long noncoding RNA Hot air mediated angiogenesis in nasopharyngeal carcinoma by direct and indirect signalling pathways. Oncotarget. 2016;7:4712–23. doi: 10.18632/oncotarget.6731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Fang Z, Wu L, Wang L, Yang Y, Meng Y, Yang H. Increased expression of the long non-coding RNA UCA1 in tongue squamous cell carcinomas: a possible correlation with cancer metastasis. Oral Surg Oral Med Oral Pathol Oral Radiol. 2014;117:89–95. doi: 10.1016/j.oooo.2013.09.007. [DOI] [PubMed] [Google Scholar]
- 83.Shen Z, Li Q, Deng H, Lu D, Song H, Guo J. Long non-coding rna profiling in laryngeal squamous cell carcinoma and its clinical significance: potential biomarkers for LSCC. PLoS One. 2014;9:e108237. doi: 10.1371/journal.pone.0108237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Song P, Ye LF, Zhang C, Peng T, Zhou XH. Long non-coding RNA XIST exerts oncogenic functions in human nasopharyngeal carcinoma by targeting miR-34a-5p. Gene. 2016;592:8–14. doi: 10.1016/j.gene.2016.07.055. [DOI] [PubMed] [Google Scholar]
- 85.Jia LF, Wei SB, Gan YH, Gou YK, Gong K, Mitchelson K, Cheng J, Yu GY. Expression, regulation and roles of miR-26a and MEG3 in tongue squamous cell carcinoma. Int J Cancer. 2014;135:2282–93. doi: 10.1002/ijc.28667. [DOI] [PubMed] [Google Scholar]
- 86.Liang J, Liang L, Ouyang K, Li Z, Yi X. MALAT1 induces tongue cancer cells’ EMT and inhibits apoptosis through Wnt/β-catenin signaling pathway. J Oral Pathol Med. 2016;46:98–105. doi: 10.1111/jop.12466. [DOI] [PubMed] [Google Scholar]
- 87.Wu T, Qu L, He G, et al. Regulation of laryngeal squamous cell cancer progression by the lncRNA H19/miR-148a-3p/DNMT1 axis. Oncotarget. 2016;7:11553–66. doi: 10.18632/oncotarget.7270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Huang W, Cui X, Chen J, Feng Y, Song E, Li J, Liu Y. Long non-coding RNA NKILA inhibits migration and invasion of tongue squamous cell carcinoma cells via suppressing epithelial – mesenchymal transition. Oncotarget. 2016;7:62520–32. doi: 10.18632/oncotarget.11528. [DOI] [PMC free article] [PubMed] [Google Scholar]