Abstract
Fluorescence in situ hybridization (FISH) with DNA probes allows the visualization of gene copy number and localization of specific DNA targets with fluorescence microscopy. Cells in culture, metaphase chromosomes, and tissue sections are fixed and prepared on glass slides. Both the DNA in the cells and fluorescently labeled probe are denatured, and the labeled probe is allowed to hybridize to the cellular DNA. The slides are washed, counterstained, and viewed via fluorescence microscopy. We describe the basic method for preparing slides and probes for studies involving DNA copy number changes and structural chromosome rearrangements in formalin-fixed paraffin-embedded (FFPE) tissue sections and cell culture preparations.
Keywords: Fluorescence microscopy, DNA probes, Metaphase chromosomes, Formalin-fixed paraffin- embedded tissues
1 Introduction
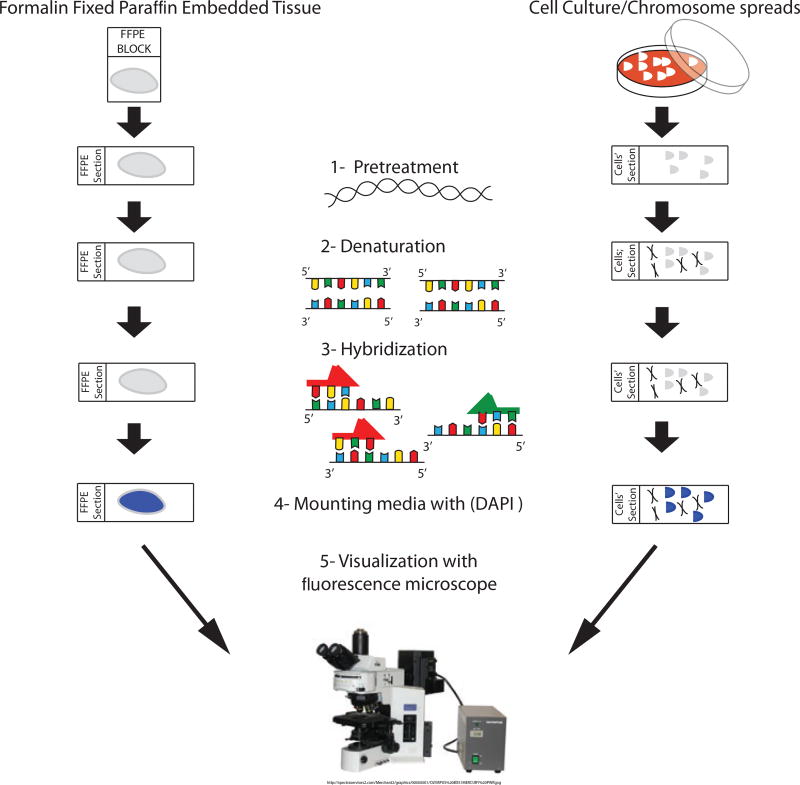
The application of fluorescently labeled probes used in fluorescence in situ hybridization (FISH) experiments has made a critical impact on the fields of genetics and cell biology. Experimental protocols that utilize DNA probes labeled with haptens or directly with fluorochromes have supported large-scale adoption in the clinical and research laboratory [1–3]. DNA probes for centromeres, telomeres, unique sequences, and whole chromosomes can be used to answer questions involving individual genes or entire chromosomes that influence diseases such as cancer and human developmental disorders [4, 5]. Complex clinical gene changes can be determined and monitored using blood samples, biopsy materials, and circulating cells as targets for the FISH probe hybridization [6–9]. Cells and metaphase chromosomes from cell culture do not require extensive pretreatment prior to FISH procedures. However, FFPE tissue sections require protein digestion and treatment with chaotropic agents to disrupt hydrogen bonding in order for the probe to access the target DNA, thus ensuring successful hybridization that can be visualized clearly [10, 11]. In this chapter, we provide detailed instruction on FISH methods for formalin-fixed paraffin-embedded (FFPE) tissue sections and cells and metaphase chromosomes from cell culture (Fig. 1).
Figure 1.
Step by step FISH experiment. The target DNA and fluorochrome-labeled DNA probe are denatured and allowed to hybridize. The slide is then washed, counterstained, and viewed with fluorescent microscopy
2 Materials
Instrument operation and reagent preparation and handling should be performed under standard operation procedures and safety regulations provided by the facility.
2.1 FFPE Tissue Sections
2.1.1 FFPE Tissue Section Cutting and Preparation
Microtome.
FFPE tissue blocks or FFPE tissue sections at 4 µm thickness on silanized or other adhesive-coated slides (see Note 1).
Microscope slides: silanized, frosted glass, excellent quality (see Note 2).
Small water bath: with thermostats set to 37 °C.
Incubator or lab oven with thermostat set to 65 °C.
2.1.2 Deparaffinizing FFPE Slides
Glass Coplin jars: 50 mL total volume.
Xylene: 40 mL, store in a vented cabinet at RT (see Note 3).
CitriSolv Hybrid Solvent and Clearing Agent (Thermo Fisher): 40 mL.
100% ethanol: 80 mL, can be stored for 1 month at RT if tightly capped with limited exposure to air.
2.1.3 Hematoxylin and Eosin (H&E) Stain
Hematoxylin nuclear stain.
Eosin Y solution.
95% ethanol: can be stored for 1 month at RT if tightly capped with limited exposure to air.
100% ethanol: can be stored for 1 month at RT if tightly capped with limited exposure to air.
Xylene: store in a vented cabinet at RT.
Mounting media: xylene based.
Glass cover slips: 22 × 50 mm, 25 × 25 mm, 22 × 22 mm, or 18 × 18 mm.
2.1.4 FFPE Slide Pretreatmen
Unstained, deparaffinized FFPE tissue sections (normal control and experimental slide).
Glass Coplin jars: 50 mL total volume.
Plastic Coplin jars: for steps that require temperatures over 50 °C.
Incubator or lab oven with thermostat set to 37 °C.
Water bath with thermostat set to 85 °C.
Water bath with thermostat set to 80 °C.
Slide warmer with surface thermometer or programmable slide warmer set to 45°C.
Pretreatment solution: Nonidet P-40, 20×SSC, and distilled water, pH 7. Store at room temperature (RT) 18–25 °C for up to 1 year (see Note 4).
Digestion solution: 25 mg/mL of proteinase in 50 mL of protease buffer.
Denaturation solution: 70% formamide/2×SSC, pH 7–8.
0.2 N hydrochloric acid (HCl).
20× saline sodium citrate (SSC). Store at room temperature. The solution must be pH 7. If necessary, use concentrated 12 N HCL and/or 1 N sodium hydroxide (NaOH) to adjust the pH.
Wash buffer 1: 0.4×SSC, 0.3% IGEPAL, pH 7 (SciGene).
Wash buffer 2: 2×SSC, 0.1% IGEPAL, pH 7 (SciGene).
4% formaldehyde in 1× phosphate-buffered saline (PBS).
Ethanol series: prepare 50 mL (v/v) solution of 70%, 80%, 95% ethanol in distilled water. Can be stored for 1 month at RT if tightly capped with limited exposure to air (see Note 5).
2.1.5 FISH Probe Preparation and Hybridization for FFPE
FISH probes (DNA probes) directly labeled with fluorochromes are commercially available in red, green, orange, gold, and aqua fluorophores. Probe sets can be customized by the investigator to target specific sequences, simultaneously (see Note 6).
Incubator or lab oven with thermostat set to 37 °C.
Small water baths with thermostats set to 80 °C and 85 °C.
Humidified chamber for incubation.
Denaturation solution: 70% formamide/2×SSC, pH 7–8.
Micropipettes and tips for P20–P200 mL pipettes.
Glass Coplin jars: 50 mL total volume.
Plastic Coplin jars: for steps that require temperatures over 50 °C.
Ethanol series: prepare 50 mL (v/v) solution of 70%, 80%, 95% ethanol in distilled water. Can be stored for 1 month at RT if tightly capped with limited exposure to air.
Glass cover slips: 22 × 22 mm.
Rubber cement for sealing cover slips.
Syringes without needles: 3 mL.
2.1.6 Post-Hybridization Washing for FFPE
Small water bath: with thermostats set to 73 °C (see Note 7).
Calibrated thermometer: measurable temperature range 65–85°C.
Wash buffer 1: 0.4×SSC, 0.3% IGEPAL, pH 7 (SciGene).
Wash buffer 2: 2×SSC, 0.1% IGEPAL, pH 7 (SciGene).
Mounting medium: Vectashield mounting medium for fluorescence with DAPI (4′,6-diamidino-2-phenylindole) (Vector Laboratories, Inc.).
Glass cover slips: 22 × 22 mm.
2.2 FISH on Cell and Chromosome Spreads from Cell Culture
2.2.1 Cell and Slide Preparation
Cell culture: T75 flask.
Cell culture medium.
Phase contrast microscope.
Pasteur pipette.
Colcemid (see Note 8).
CMF-PBS: Dulbecco’s PBS without calcium or magnesium.
Sterile 15 mL conical tube.
0.05 trypsin-EDTA.
Hypotonic solution: 0.075 M potassium chloride (KCl) or one part cell culture medium plus two parts sterile water.
Fixative: 3:1 methyl alcohol and glacial acetic acid, 15 mL (see Note 9).
Glass microscope slides cleaned in 100% ethanol alcohol.
Ice-cold mixture of 40% methanol and 60% distilled water.
2.2.2 Slide Pretreatment from Cell Culture
Glass Coplin jars: four jars.
Small water bath with thermostats set to 37 °C.
Pretreatment solution: Nonidet P-40, 20×SSC, and distilled water, pH 7. Store at room temperature (RT) 18–25 °C for up to 1 year.
Ethanol series: prepare 50 mL (v/v) solution of 70%, 80%, 95% ethanol in distilled water. Can be stored for 1 month at RT if tightly capped with limited exposure to air.
2.2.3 FISH Probe Preparation and Hybridization from Cell Culture
Cell smears: slides prepared from cell culture specimens.
FISH probes (DNA probes) directly labeled with fluorochromes are commercially available in red, green, orange, gold, and aqua fluorophores. Probe sets can be customized by the investigator to target specific sequences, simultaneously (see Note 6).
Incubator or lab oven with thermostat set to 37 °C.
Small water baths with thermostats set to 70 °C and 80 °C.
Humidified chamber for incubation.
Denaturation solution: 70% formamide/2×SSC, pH 7–8.
Micropipettes and tips for P20–P200 mL pipettes.
Glass Coplin jars: 50 mL total volume.
Plastic Coplin jars: for steps that require temperatures over 50 °C.
Ethanol series: prepare 50 mL (v/v) solution of 70%, 80%, 95% ethanol in distilled water. Can be stored for 1 month at RT if tightly capped with limited exposure to air.
Glass cover slips: 22 × 22 mm.
2.2.4 Post-Hybridization Wash from Cell Culture
Small water bath: with thermostats set to 73 °C (see Notes 7 and 16).
Calibrated thermometer: measurable temperature range 65–85°C.
Wash buffer 1: 0.4×SSC, 0.3% IGEPAL, pH 7 (SciGene).
1× phosphate-buffered detergents (PBD): 4×SSC, 0.05% NP-40.
Mounting medium: Vectashield mounting medium for fluorescence with DAPI (4′,6-diamidino-2-phenylindole) (Vector Laboratories, Inc.).
Glass cover slips: 22 × 22 mm.
2.3 Microscopy and Scoring
Fluorescence microscope, equipped with a 100-W mercury arc or argon lamp or equivalent.
DAPI (blue), red, and green double or triple band-pass filters.
Immersion oil (see Note 10).
63× and 100× objectives.
2.3.1 Digital Imaging Microscopy
Fluorescence microscope, equipped with a 100-W mercury arc or argon lamp and DAPI, red, green, and double or triple band- pass filters.
Digital camera and imaging software.
3 Methods
3.1 FFPE Section
All steps are performed at room temperature (RT) unless otherwise specified.
3.1.1 FFPE Tissue Section Cutting and Preparation
Fixation is a critical step to preserve nucleic acids in paraffin-embedded tissue. Therefore, tissue should be fixed with 10% neutral buffered formalin or paraformaldehyde, and precipitating fixatives such as alcohol should be avoided. At least five serial sections cut at 4 µm thickness onto silanized or other adhesive-coated slides should be obtained from each case. One section will be used for H&E staining for examination under a light microscope, to characterize the site or region of interest, such as normal tissue vs. the tumor tissue, and to localize the connective tissue and blood vessels. This H&E slide will be used for section orientation to ensure that the correct area in an adjacent slide has been selected for FISH scoring. The additional slides can be used as a backup if the FISH staining does not work properly and needs to be repeated.
Use a microtome to cut five formalin-fixed, paraffin wax-embedded tissue sections into 4 µm thick sections. Cut sections from normal tissue blocks and the experimental tissue block. A normal tissue block should be used as a negative control.
Float the sections in warmed distilled water. Then, scoop a silanized glass microscope slide under the floating tissue section, and lift the slide with the attached tissue section up and out of the water.
Leave sections on the slide to air-dry.
Bake the slides at 65 °C for 8–16 h.
Slides can be stored at −20 °C; however, long-term storage at −20 °C can affect the pretreatment conditions.
3.1.2 Deparaffinizing FFPE Slides
The following steps must be performed under a fume hood. Solutions can be placed in separate Coplin jars or in a HistoTek apparatus designed for sequential slide treatments:
Prepare four Coplin jars: each jar should contain one of the following chemicals, 40 mL xylene, 40 mL CitriSolv, 40 mL 100% ethanol jar #1, 40 mL 100% ethanol jar #2.
Place up to four slides in a Coplin jar containing xylene. Incubate the slides for 10 min.
Remove the slides from xylene and place into the CitriSolv. Incubate the slides for 10 min.
Remove the slides from the CitriSolv and place in 100% ethanol solution #1 for 5 min.
Remove the slides from ethanol solution #1 and place them in 100% ethanol #2 for 5 min.
Remove the slides from the ethanol and allow air-drying. Processing slides after this step for FISH will enhance the quality of the staining.
3.1.3 Hematoxylin and Eosin (H&E) Staining
These steps will be performed to stain one of the FFPE sections for light microscope examination:
Rinse slide in deionized water for 2 min.
Place slide in hematoxylin for 5 min.
Remove slide from hematoxylin and rinse under running tap water until no more blue stain runs off the slide.
Immerse slide in distilled water for 30–60 s.
Rinse slide with running tap water for 30 s.
“Blue” slide with running tap water for 1 min.
Rinse slide in 95% ethanol solution.
Immerse slide in eosin Y for 2 min.
Dehydrate slide in 100% ethanol solution for 1 min intervals, three times.
Immerse slide in three sequential xylene baths for 1 min intervals.
Allow the slide to air-dry.
Place several drops of xylene-based mounting media on the tissue section.
Apply a glass cover slip to the slide.
3.1.4 FFPE Slide Pretreatment
One unstained FFPE tissue slide each from the negative control and the experimental specimen will be pretreated prior to adding the DNA probes and hybridization. Pretreatment solutions are commercially available (SciGene) or can be freshly prepared on the day of use. Before starting the pretreatment, make sure to do the first four steps:
Pre-warm pretreatment solution (50 mL) in a plastic Coplin jar at 80 °C in water bath.
Verify pH of the protease buffer. Adjust to pH 2.0 using 12 N HCL and/or 1.0 N NaOH.
Pre-warm the protease buffer (part of the digestion solution) to 37 °C. Ensure that protease buffer is at 37 °C before adding 25 mg of protease to the buffer.
Pre-warm denaturation buffer to 85 °C.
Immerse the FFPE slides that will be used for FISH into the 0.2 N HCl for 20 min.
Remove slides from the 0.2 N HCl and place in purified water for 3 min.
Immerse slides in wash buffer 1 for 3 min.
Place slides in the pretreatment solution at 80 °C for 60 min (see Note 11).
Immerse slides in purified water for 1 min.
Immerse slides in wash buffer 1 for 5 min and in wash buffer 2 for 5 min; total 10 min.
Remove excess buffer by blotting the back of slides with paper towel.
Immerse slides in digestion (protease) solution at 37 °C for 20 min (see Note 12).
Immerse slides in wash buffer 1 for 5 min and in wash buffer 2 for 5 min for a total of 10 min.
Dry slides for 2–5 min on 45 °C slide warmer.
Place slides in 4% formaldehyde in 1× PBS for 10 min. Or drop formaldehyde directly onto the slide and cover with a plastic cover slip.
Immerse slides in wash buffer 1 for 5 min and in wash buffer 2 for 5 min. Total 10 min.
Dry slides for 2–5 min on 45 °C slide warmer.
Place slides in denaturation buffer at 80 °C for 5 min (see Note 13).
Dehydrate slides in ascending dilution series of ethanol solutions, 70%, 80%, and 95% for 1 min each.
Gently dry the back of the slides, and place on 45 °C slide warmer until ethanol evaporates.
3.1.5 FISH Probe Preparation and Hybridization for FFPE Tissue
Probes are directly labeled with fluorochromes and are available commercially in red, green, orange, gold, orange, and aqua. Each probe will have specific preparation directions described by the manufacturer, which should be followed.
Before starting, make sure to perform the following steps. Prepare a humidified chamber for hybridization and pre-warm at 37 °C, 30 min before hybridization (see Note 14). Pre-warm denaturation solution (70% formamide/2×SSC, pH 7–8) in Coplin jar at 85 °C (this is for the tissue) for 30 min:
Dilute probe with hybridization solution as described by manufacturers.
Denature probe in 80 °C water bath for 10 min, and then spin in a mini-centrifuge for 2–5 s to collect the contents in the bottom of the type. Place on ice until ready to hybridize.
Immerse slides in denaturation solution at 85 °C for 5 min.
Dehydrate slides in ascending dilution series of ethanol solution, 70%, 80%, and 95% for 1 min each.
Gently dry the back of the slide and place on slide warmer until ethanol evaporates.
Apply 10 µL of probe mixture to the slide.
Apply a clean 22 × 22 mm glass cover slip to the slide, and seal cover slip edges with rubber cement delivered in plastic disposable syringe without a needle (see Note 15).
Place slide in pre-warmed humid chamber, and allow to hybridize overnight in 37 °C oven or incubator for 16 h.
3.1.6 Post-Hybridization Washing for FFPE Tissue
Before starting, pre-warm wash buffer 1 (0.4×SSC/0.3% IGEPAL, pH 7) at 73 °C, and be sure to measure the temperature of the wash buffer with a calibrated thermometer within the Coplin jar itself (see Note 16). Wash buffer 2 (2×SSC/0.1% IGEPAL, pH 7) should be at room temperature:
Carefully, remove cover slips and place slides in pre-warmed wash buffer 1 at 73 °C, and let it stand exactly for 2 min. This step is critical, and attention to the temperature and time will insure results that are able to be scored/quantified.
Remove slides and place in wash buffer 2 at room temperature for 1 min.
Rinse once in distilled water at RT to remove excess salts and detergents.
Allow slides to dry at RT in the dark.
Apply 18 µL of DAPI counterstain.
Place a 22 × 22 mm glass cover slip over the section.
Store slide in a dark slide box at −20 °C. The slide can be stored up to a year; however, it is recommended to view the slides within 2 days of hybridization.
3.2 FISH on Cells and Chromosome Spreads from Cell Culture
Specimens that have been stained with nuclear stains and dyes such as Giemsa should not be used. In the unusual event there is only one stained sample to use for analysis, carefully destain the slide in 3:1 methanol/glacial acetic acid for 20 min at room temperature.
3.2.1 Cells and Slide Preparation
Grow cells in a T75 flask until they are at 80% confluency(see Note 17).
Aspirate medium, and incubate for 4 h in cell culture media containing colcemid at a final concentration of 0.06 mg/mL (see Note 8).
Aspirate medium and save in a sterile tube, and rinse the monolayer with CMF-PBS, and decant. Then add 3 mL of 0.05% trypsin-EDTA.
View the cells with a tissue culture microscope until detached, and then add the cell suspension to the tube containing medium saved from step 3, and centrifuge the suspension at 200 × g for 3 min.
Aspirate the supernatant, and suspend the cell pellet in 5 mL of hypotonic solution consisting of 0.075 M KCl or one part of cell culture media and two parts of distilled water, and leave for 7–10 min at RT. Centrifuge at 200 × g for 3 min.
Remove the hypotonic solution, and add 5 mL of fixative (3:1 methyl alcohol and glacial acetic acid, freshly prepared, 5 mL for each sample) to the centrifuge tube without disturbing the cell pellet, and let it stand for 20 min.
Suspend the cells with a Pasteur pipette and centrifuge at 200 × g for 3 min.
Aspirate the supernatant and resuspend cells in a fresh fixative. Centrifuge at 200 × g for 3 min.
Aspirate the supernatant and resuspend cells in 0.5 mL of fresh fixative.
Dip alcohol-cleaned slides in an ice-cold mixture of 40% methanol and 60% distilled water. Remove the slides and drain off the excess of fluid, but do not shake it off.
Hold the slide at a 45° angle toward the floor. Place a drop of the final cell suspension on the upper portion of the slide, and let the cell suspension spread downward. If the cell suspension is too thin, add more drops as needed on each slide.
Allow slides to air-dry.
3.2.2 Slide Pretreatment from Cell Culture
Place the prepared slide in the Coplin jar with pre-warmed pretreatment solution at 37 °C for 30 min (see Note 18).
Dehydrate the slide in ascending dilution series of ethanol solution, 70%, 80%, and 95% for 2 min each.
Allow the slide to air-dry.
3.2.3 FISH Probe Preparation and Hybridization from Cell Culture
Before starting, prepare a humid chamber for hybridization, and pre-warm at 37 °C, 30 min before hybridization. Pre-warm denaturation solution (70% formamide/2×SSC, pH 7–8) in the Coplin jar at 70 °C for 30 min. 70°C is the pre-warm temperature for cells on a slide, rather than 80°C for tissue sections:
Dilute probe as directed by manufacturer with hybridization solution.
Denature probe by heating it at 80 °C for 10 min, and then spin it briefly (2–5 s) in a mini-centrifuge to collect the contents in the bottom of the tube. Place the probe on ice until ready to hybridize.
Immerse slide in pre-warmed denaturation solution at 70 °C for 2 min (see Note 19).
Dehydrate slide in ascending dilution series of ethanol solution, 70%, 80%, and 95% for 2 min each in ice (see Note 20).
Gently dry the back of the slide and allow to air-dry.
Apply 10 µL of probe mixture to the slide (according to manufacturer instructions), and cover it with a glass cover slip.
Place slide in pre-warmed humidified chamber and allow to hybridize 6 h or overnight in 37 °C oven or incubator.
3.2.4 Post-Hybridization Wash from Cell Culture
Before staring, preincubate wash buffer 1 (0.4×SSC/0.3% IGEPAL, pH 7) at 72 °C. Measure the temperature of the wash buffer with a calibrated thermometer within the Coplin jar itself:
Carefully, remove cover slip, and place slide in pre-warm wash buffer 1 at 72 °C, and let it stand exactly for 5 min (see Note 21).
Remove slide and place in 1× PBD (phosphate buffer detergents) at RT for 2 min.
Rinse once in distilled water at RT to remove excess salts and detergents. Do not allow slide surface to dry.
Apply 10 µL of DAPI counterstain (see Note 22).
Place a 22 × 22 mm glass cover slip over the cells, and press gently to remove any excess fluid.
Store slides in a dark slide box at −20 °C. Slides can be stored up to 2 years or more and retain signal. It is recommended that slides are viewed within 1 week of hybridization.
3.3 Microscopy and Scoring
Locate region of interest by scanning with a low magnification objective with a DAPI filter.
Apply immersion oil to the slide and view with 63× or 100× objective (see Note 10).
Score control probe for each cell and record. In a normal diploid cell, there should be two signals. If there are not two signals, choose another cell for analysis.
Score test probe and record.
Repeat until the required numbers of cells have been analyzed. This varies according to the purpose of the study and can range from 10 to 200 cells.
If scoring for gene amplification, divide the control by the test specimen signals. A score of 1.8 indicates amplification.
3.3.1 Digital Imaging Microscopy
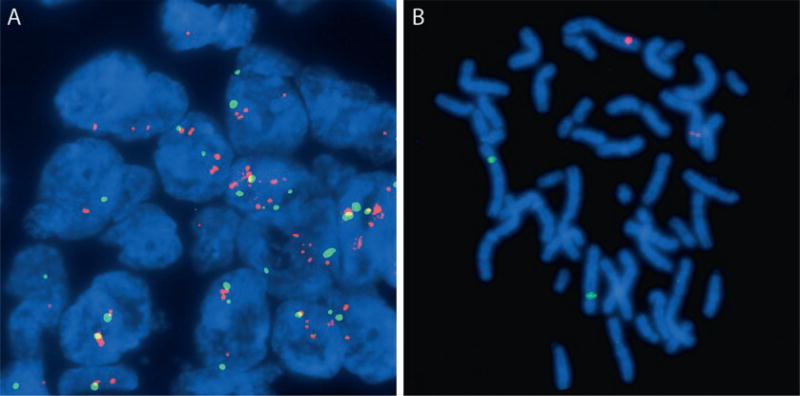
Document two representative cells using photography. Many fluorescent microscopes are equipped with cameras and imaging software that assist with this step (Fig. 2).
Store photographic records on a separate hard drive.
Figure 2.
Fluorescent in situ hybridization images. (a) Breast cancer tissue hybridized with a control chromosome 8 centromeric probe labeled in green and a MYC probe labeled in red showing low-level amplification. DAPI counterstain in blue. (b) Normal prostate cell metaphase chromosomes labeled with red and green unique sequence DNA probes. DAPI counterstain in blue
4 Notes
Tissue sections (recuts) can be ordered from the tissue bank or pathologist with a hematoxylin and eosin (H&E) stained slide for determining the region of interest. The histomorphology can be reviewed using the H&E stained slide.
The slides used for FISH should be of the highest quality. The glass slides should be pre-cleaned ones and labeled as such from the manufacturer. The glass slides used for the tissue sections need to have an adhesive surface so that the adherent sections do not slide off through the many washes.
Xylene is effective at removing paraffin from the blocks, but is toxic and must be used in a fume hood. If this remains a concern, less toxic clearing solutions, such as Histoclear, may be used.
1 M sodium thiocyanate (NaSCN) may also be used as a pretreatment reagent (SciGene).
The graded ethanol series and denaturation solutions can be stored at 4 °C for one or 2 days and used again if they have been carefully covered with a jar top or parafilm. If alcohols are exposed to ambient air, they will become diluted.
Investigators should first determine which fluorescence filters are fitted on the microscope that is planned for use so that probes or probe set choices for your experiments are consistent. It is convenient to have a double band-pass or a triple band-pass filter if studies involve multicolor scoring of many cells.
The post-hybridization wash step at 73°C is the most important step in FISH protocols. A calibrated thermometer should be used to measure the temperature of the solution for this step.
Colcemid (N-desacetyl-N-methylocolchicine) is a microtubule destabilizing agent, preventing spindle formation during mitosis, causing cell cycle arrest in metaphase [12].
When 3:1 ethanol/glacial acetic acid fixative is to be used for treating specimens, it must be made up fresh each day and stored in a glass bottle with a cap.
Fluorescence detection requires the use of glass cover slips and specialized low autofluorescence immersion oil to avoid autofluorescence that will interfere with image quality.
The pretreatment time may vary based on tissue type.
Monitoring protein digestion of tissues can be achieved by staining the section with propidium iodide which can be viewed with the fluorescent microscope and will not interfere with the experiment. Tissues that show cellular borders will provide optimal hybridization and analysis. Time may vary based on the tissue type, and the concentration and incubation times will need to be determined for in the lab. Propidium iodide intercalates with DNA and can cause mutations, so personal protective gear and disposal guidelines should be followed.
The denaturation step can be performed in a Coplin jar with heated solution if a programmable slide warmer is not available. It is usually more convenient to co-denature on the slide warmer when working with tissues, because the elevated temperatures (85°C) required for denaturing the DNA will eventually cause the glass Coplin jar to crack.
The hybridization oven or block should be humidified and preheated in preparation for the hybridization. Background hybridization may occur as well as difficulty in removing the hybridization cover slip.
A 22 × 22 mm glass cover slip is optimal for hybridization. When applying the hybridization cover slip with rubber cement, a 3 mL syringe is useful to quickly and liberally cover and seal the edges.
The post-wash step is the most critical of the FISH experiment. The hybridized probe can be washed away if the temperature is too high or the incubation is too long. When performing a study with a new probe set, it is recommended to use a lower stringency wash of 1×SSC. It is simple to rewash the slide if the background staining is too great. The cover slips are not permanently attached, so rewashing is possible.
Cell culture preparations should be inspected using phase microscopy before being used in a hybridization assay to assure that the density of cells is not too dilute or concentrated. An adjustment of the cell density can be achieved by centrifuging and resuspending cells in either more or less medium, as needed. If metaphase spreads have been prepared for analysis, assure that the chromosomes are well separated on the slide.
The pretreatment solution artificially ages the chromosomes, making them less sensitive to over-denaturation. This step is important for obtaining optimal signal.
Do not agitate. Time and temperature are very important for the maintenance of chromosome morphology. Over- denatured slides will not hybridize properly.
Cold ethanol washes quickly stop denaturation while dehydrating the slides.
The post-hybridization washing step is critical, and attention to the temperature and time will insure scorable results. Also, reducing washing time may increase the signaling background.
The DAPI is already diluted 1:1 with antifade reagent. The DAPI and propidium iodide intercalate with DNA and can cause mutations, so personal protective gear and disposal guidelines should be followed.
References
- 1.Kallioniemi A, Visakorpi T, Karhu R, Pinkel D, Kallioniemi OP. Gene copy number analysis by fluorescence in situ hybridization and comparative genomic hybridization. Methods. 1996;9(1):113–121. doi: 10.1006/meth.1996.0015. [DOI] [PubMed] [Google Scholar]
- 2.Pinkel D, Gray JW, Trask B, van den Engh G, Fuscoe J, van Dekken H. Cytogenetic analysis by in situ hybridization with fluorescently labeled nucleic acid probes. Cold Spring Harb Symp Quant Biol. 1986;51(Pt 1):151–157. doi: 10.1101/sqb.1986.051.01.018. [DOI] [PubMed] [Google Scholar]
- 3.Waye JS, Willard HF. Chromosome-specific alpha satellite DNA: nucleotide sequence analysis of the 2.0 kilobasepair repeat from the human X chromosome. Nucleic Acids Res. 1985;13(8):2731–2743. doi: 10.1093/nar/13.8.2731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Cajulis RS, Frias-Hidvegi D, Yu GH, Eggena S. Detection of numerical chromosomal abnormalities by fluorescence in situ hybridization of interphase cell nuclei with chromosome-specific probes on archival cytologic samples. Diagn Cytopathol. 1996;14(2):178–181. doi: 10.1002/(SICI)1097-0339(199603)14:2<178::AID--DC14>3.0.CO;2-J. [DOI] [PubMed] [Google Scholar]
- 5.Cremer T, Lichter P, Borden J, Ward DC, Manuelidis L. Detection of chromosome aberrations in metaphase and interphase tumor cells by in situ hybridization using chromosome-specific library probes. Hum Genet. 1988;80(3):235–246. doi: 10.1007/BF01790091. [DOI] [PubMed] [Google Scholar]
- 6.Mayer JA, Pham T, Wong KL, Scoggin J, Sales EV, Clarin T, Pircher TJ, Mikolajczyk SD, Cotter PD, Bischoff FZ. FISH-based determination of HER2 status in circulating tumor cells isolated with the microfluidic CEE platform. Cancer Genet. 2011;204(11):589–595. doi: 10.1016/j.cancergen.2011.10.011. [DOI] [PubMed] [Google Scholar]
- 7.Pinkel D, Albertson DG. Array comparative genomic hybridization and its applications in cancer. Nat Genet. 2005;37(Suppl):S11–S17. doi: 10.1038/ng1569. [DOI] [PubMed] [Google Scholar]
- 8.Ventura RA, Martin-Subero JI, Jones M, McParland J, Gesk S, Mason DY, Siebert R. FISH analysis for the detection of lymphoma-associated chromosomal abnormalities in routine paraffin-embedded tissue. J Mol Diagn. 2006;8(2):141–151. doi: 10.2353/jmoldx.2006.050083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wang N. Methodologies in cancer cytogenetics and molecular cytogenetics. Am J Med Genet. 2002;115(3):118–124. doi: 10.1002/ajmg.10687. [DOI] [PubMed] [Google Scholar]
- 10.Blancato JK, Williams MS, Dickson RB. Fluorescence in situ hybridization assessment of c-myc gene amplification in breast tumor tissues. Methods Mol Med. 2006;120:297–307. doi: 10.1385/1-59259-969-9:297. [DOI] [PubMed] [Google Scholar]
- 11.Chin SF, Daigo Y, Huang HE, Iyer NG, Callagy G, Kranjac T, Gonzalez M, Sangan T, Earl H, Caldas C. A simple and reliable pretreatment protocol facilitates fluorescent in situ hybridisation on tissue microarrays of paraffin wax embedded tumour samples. Mol Pathol. 2003;56(5):275–279. doi: 10.1136/mp.56.5.275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Saraiva NZ, Perecin F, Meo SC, Ferreira CR, Tetzner TA, Garcia JM. Demecolcine effects on microtubule kinetics and on chemically assisted enucleation of bovine oocytes. Cloning Stem Cells. 2009;11(1):141–152. doi: 10.1089/clo.2008.0044. http://luckylion.de/vasco/tutorials/indesignthesis/mol_toolbox.php. [DOI] [PubMed] [Google Scholar]