Abstract
Acute myeloid leukemia (AML) is the most common and severe form of acute leukemia diagnosed in adults. Owing to its heterogeneity, AML is divided into classes associated with different treatment outcomes and specific gene expression profiles. Based on previous studies on AML, in this study, we designed and generated an AML-array containing 900 oligonucleotide probes complementary to human genes implicated in hematopoietic cell differentiation and maturation, proliferation, apoptosis and leukemic transformation. The AML-array was used to hybridize 118 samples from 33 patients with AML of the M1 and M2 subtypes of the French-American-British (FAB) classification and 15 healthy volunteers (HV). Rigorous analysis of the microarray data revealed that 83 genes were differentially expressed between the patients with AML and the HV, including genes not yet discussed in the context of AML pathogenesis. The most overexpressed genes in AML were STMN1, KITLG, CDK6, MCM5, KRAS, CEBPA, MYC, ANGPT1, SRGN, RPLP0, ENO1 and SET, whereas the most underexpressed genes were IFITM1, LTB, FCN1, BIRC3, LYZ, ADD3, S100A9, FCER1G, PTRPE, CD74 and TMSB4X. The overexpression of the CPA3 gene was specific for AML with mutated NPM1 and FLT3. Although the microarray-based method was insufficient to differentiate between any other AML subgroups, quantitative PCR approaches enabled us to identify 3 genes (ANXA3, S100A9 and WT1) whose expression can be used to discriminate between the 2 studied AML FAB subtypes. The expression levels of the ANXA3 and S100A9 genes were increased, whereas those of WT1 were decreased in the AML-M2 compared to the AML-M1 group. We also examined the association between the STMN1, CAT and ABL1 genes, and the FLT3 and NPM1 mutation status. FLT3+/NPM1− AML was associated with the highest expression of STMN1, and ABL1 was upregulated in FLT3+ AML and CAT in FLT3− AML, irrespectively of the NPM1 mutation status. Moreover, our results indicated that CAT and WT1 gene expression levels correlated with the response to therapy. CAT expression was highest in patients who remained longer under complete remission, whereas WT1 expression increased with treatment resistance. On the whole, this study demonstrates that the AML-array can potentially serve as a first-line screening tool, and may be helpful for the diagnosis of AML, whereas the differentiation between AML subgroups can be more successfully performed with PCR-based analysis of a few marker genes.
Keywords: acute myeloid leukemia, French-American-British2, M1, M2, gene expression profiling, DNA microarrays, boutique arrays, RT-qPCR, real-time PCR, droplet digital PCR
Introduction
Acute myeloid leukemia (AML), the most common and severe form of acute leukemia in adults, is responsible for the highest mortality from leukemia in general (1). The origin of AML is multifactorial and has not yet been entirely elucidated. The disease begins in a bone marrow stem cell exposed to a complex interplay of hereditary and environmental factors. Disturbances in myeloid progenitor cell growth, differentiation and proliferation lead to the clonal expansion of bone marrow myeloblasts and their infiltration into the peripheral blood (1,2). Therefore, the number of immature, non-functional leukocytes is increased, and normal blood cell production is impaired.
AML is a heterogeneous type of cancer in which subsets of molecularly different types can be distinguished. According to the first classification of hematological disorders, the French-American-British (FAB) system (3), there are 8 types of AML (M0-M7), with specific morphological characteristics and differentiation stages. The more recent World Health Organization (WHO) classification (4) is based on a combination of clinical symptoms, cell morphology, immunophenotype and genetic abnormalities. In ~55% of patients with AML, clonal chromosome rearrangements are present (5); e.g., translocations t(8;21), t(15;17) and t(16;16), which result in the fusion genes, RUNX1/RUNX1T1, PML-RARα and CBFB-MYH11, respectively. In a large group of patients with AML (40–49%) with normal karyotypes (NK-AML or CN-AML, from cytogenetically normal AML) (5,6), recurrent small mutations have been identified. They usually occur in genes encoding signaling proteins, transcription factors and chromatin modifiers, which affect cell signaling or general gene expression. The most frequent are mutations in NPM1 (7), FLT3 (8), CEBPA (9), KIT (10), NRAS/KRAS (11), TET2 (12), DNMT3A (13) and IDH1/2 (14). Some of these mutations are clinically relevant as diagnostic or prognostic markers and potential therapeutic targets (15).
The development of high-throughput technologies, such as microarrays and next generation sequencing has contributed to progress in leukemia research (16–18). Since 1999, when the first applications of DNA microarrays in leukemia classification and outcome prediction were demonstrated (19,20), many publications based on gene expression profiling in hematological malignancies have appeared. Among these, several hundred have focused on AML [such as for example (21–23)]. Some have shown that certain genetic alterations correspond with specific gene expression signatures (24,25). Gene expression profiles have also been correlated with prognosis and treatment outcomes (26,27). However, in clinical practice, age, white blood cell (WBC) counts in the blood and karyotype abnormalities are still the key outcome determinants (6). Diagnostic tests based on single gene mutations [including one recently published by our group (28)], are being increasingly applied; however, the number of mutations and their detection methods are not standardized among laboratories. A reasonable compromise between a single gene test and a genome-wide tool, irrespective of the purpose (mutation detection or gene expression measurements), is a small dedicated microarray, also known as a boutique array (29,30). Based on our experience in boutique microarray design, production and data normalization (30–32), we decided to create a small microarray dedicated to gene expression profiling in AML (AML-array). The main aims of this study were to test the utility of this array, verify the selected results with 2 quantitative polymerase chain reaction (PCR) approaches, standard real-time PCR and droplet-digital PCR (ddPCR), and to examine gene expression in a new group of patients with AML. Into the analysis, we included Polish adult patients with de novo AML, prior to therapy, classified as M1 and M2, 2 FAB subtypes in which myeloid differentiation is arrested in the first stages of granulopoiesis. In the same group of patients, we have previously performed a comparative proteomics analysis of AML with and without maturation (33,34). The additional aim of the study was to compare presented here transcriptomic results with our earlier proteomic results and with other AML transcriptomic data. We hoped to find the novel factors, such as gene groups, gene expression patterns or gene associations with molecular or clinical characteristics, which can be correlated with AML pathogenesis. Such analyses are valuable as it was shown that AML pathogenesis can differ among individual patients (35) and our understanding of AML genomics is still incomplete.
Materials and methods
Samples
Peripheral blood (PB) and bone marrow (BM) samples were collected from 41 adult patients with AML-M1/M2 and from 20 adult healthy volunteers (HV). Each individual provided signed informed consent for treatment and for their participation in this study. Appropriate approval was also obtained from the Bioethical Commission of the Karol Marcinkowski Poznan University of Medical Sciences, Poznan, Poland. The patients were diagnosed and treated at the Department of Hematology and Bone Marrow Transplantation at the Poznan University Hospital of the Lord’s Transfiguration of the University of Medical Sciences in Poznan, Poland. Standard AML therapy using cytosine arabinoside plus daunorubicin (3′+7′) was administered to all patients to induce complete remission (CR), which was defined according to the European Leukemia Net guidelines (2). When available, the samples were collected at the 3 following time-points: when AML was diagnosed, prior to first therapy (T0), when CR was established [between day +21 and +28 after the start of induction therapy (T1)], and when the disease relapsed (T2). However, from the T1 samples, the number of cells was much lower than that of cells from the T0 and T2 samples, and often, there was not sufficient material to perform replicate experiments. Moreover, not all micro-array images met the required quality criteria and had to be filtered out. The material from the T1 time-point was the most heterogeneous, and preliminary microarray analysis revealed that the T1 samples did not cluster together and did not exhibit clear common characteristics. The T2 samples were generally similar to the T0 samples and we were not able to distinguish them by unsupervised hierarchical clustering. Therefore, we decided to limit our analyses to the samples collected at the T0 time-point, with the use of available information about the history of treatment. The advantage of this selection was that the T0 samples were the most homogenous (the fractions of blood and bone marrow-derived mononuclear cells isolated from patients with AML-M1 contained ~90% leukemic cells, whereas from those from the patients with AML-M2 contained ~70% myeloblasts). Table I presents the summarized information on the patient and HV samples.
Table I.
Sample summary.
| A, General information
| |||||||
|---|---|---|---|---|---|---|---|
| Total no. of samples | Age range (median) | Sex, F/M/unknown | WBC count [×109/l] range (median) | No. of microarray hybridizations (individual samples) | No. of samples in real-time PCR experiment | No. of samples in ddPCR experiment | |
| HV | 20 | 23–60 (36.4) | 10/07/03 | 5.1–25.3 (6.2) | 15 (15) | 14 | 16 |
| AML | 41 | 19–78 (48.8) | 17/24 | 0.5–345 (13.7) | 103 (33) | 21 | 37 |
| B, FAB classification and mutation status of AML samples
| ||||||
|---|---|---|---|---|---|---|
| FAB M1 | FAB M2 | RUNX1/RUNX1T1 [t(8;21)] | NPM1−/FLT3+ | NPM1+/FLT3− | NPM1+/FLT3+ | |
| Patients (micr.a) | 15 (11/31) | 26 (22/72) | 5 (5/15) | 3 (2/11) | 8 (6/23) | 2 (2/8) |
| C, Classification of AML samples in respect to treatment response
| |||||
|---|---|---|---|---|---|
| RES | CR | CR-short | CR-long | X | |
| Patients (micr.a) | 10 (9/26) | 20 (15/49) | 14 (12/40) | 6 (3/9) | 11 (9/28) |
The number of patients/samples included into the microarray experiment; WBC, white blood cell count; HV, healthy volunteers (controls); AML, acute myeloid leukemia; RES, resistance to therapy; CR, complete remission (CR-short, lasting up to 1 year; CR-long, -lasting >1 year); X, death during therapy; FLT3+−FLT3-ITD (internal tandem duplication in the FLT3 gene); RT-qPCR -reverse transcription-quantitative PCR; ddPCR, droplet digital PCR.
Cell separation
Mononuclear cells form the peripheral blood (termed PBMCs) and bone marrow (termed BMMCs) were separated through density gradient centrifugation (Gradisol L; Aqua-Medica, Bogucin, Poland) and washed 3 times with 1X phosphate-buffered saline (PBS; Ca and Mg-free; IBSS Biomed S.A., Warsaw, Poland). The cell pellet was suspended in lysis buffer from a mirVana miRNA Isolation kit (Ambion/Thermo Fisher Scientific, Waltham, MA, USA) and immediately frozen at −80°C.
RNA isolation
Total RNA was extracted from the PBMCs and BMMCs using a mirVana miRNA Isolation kit and the DNase-treated (TURBO DNA-free kit) (both from Ambion/Thermo Fisher Scientific). RNA integrity was evaluated with the use of a Bioanalyzer 2100 and Total RNA Nano assay (Agilent Technologies, Santa Clara, CA, USA). Only RNAs with an RNA integrity number (RIN) ≥7.5 were used for the gene expression analyses.
Microarray construction
Screening of the literature enabled us to identify the following groups of genes: i) Proven and postulated acute leukemia biomarkers; ii) general oncogenes; and iii) predicted to be specifically involved in leukemic transformation. In addition, we selected a set of control genes. Human housekeeping genes, not associated with oncogenesis, were used as the positive controls, whereas plant- and bacterial-specific genes served as the negative controls. We ordered 903 chemically synthesized, amino-modified microarray probes from 2 companies: Ocimum Biosolutions Ltd. (Hyderabad, India; 783 probes 50-nt-long) and Operon Biotechnologies GmbH (Cologne, Germany; 120 probes 70-nt-long). As an additional control, we used 4 short DNA probes (24-28-nt-long; Institute of Biochemistry and Biophysics, Warsaw, Poland), 8 probes complementary to external spike RNAs (AM1781; Ambion/Thermo Fisher Scientific) and one random oligo probe (Operon Biotechnologies GmbH). In total, the microarray contained 916 unique probes: 896 probes complementary to 838 human genes, 8 probes specific for 5 bacterial genes, 3 probes specific for 3 plant genes and 9 artificial control probes. The probes were diluted in Pronto Epoxide Spotting Solution to a 20 µM concentration and spotted in triplicates onto Epoxide-Coated Slides (both from Corning Inc., New York, NY, USA) with a SpotArray 24 instrument (Perkin-Elmer, Waltham, MA, USA). The design of our AML-array has been deposited in the ArrayExpress database (http://www.ebi.ac.uk/arrayexpress) under the accession no. A-MEXP-2220.
Labeling and microarray hybridization
A quantity of 10 µg of each total RNA sample was reverse transcribed using anchoredoligo(dT)20, aminoallyl-modified dNTPs and SuperScript III reverse transcriptase from a SuperScript Plus Indirect cDNA Labeling System (Invitrogen, Carlsbad, CA, USA). Amino-modified cDNAs from the patients with AML and HV were labeled with Alexa Fluor 647 and the reference cDNA from the HL60 cell line sample (obtained from Dr Marcin Schmidt from Poznan University of Life Sciences) with AlexaFluor 555. The labeled cDNA was purified (MinElute Reaction Cleanup kit; Qiagen, Hilden, Germany), dissolved in hybridization buffer (5X SSC, 0.1% SDS and 0.1 mg BSA/ml) and kept up to 30 min at 50°C prior to hybridization in Corning microarray hybridization chambers in a HybArray12 (Perkin-Elmer) or a water bath by using a step-down hybridization protocol (5 h/50°C, 5 h/45°C and 5 h/40°C). Three subsequent wash steps were applied: i) 2X SSC and 0.1% SDS at 40°C for 5 min; ii) 2X SSC at room temperature for 5 min; and iii) 0.2X SSC at room temperature for 5 min. The slides were dried through centrifugation with the Microarray High-Speed Centrifuge (Arrayit Corp., Sunnyvale, CA, USA; 5 sec, 2,000 × g, room temperature) and scanned with a ScanArrayExpress system (Perkin-Elmer) at 5-µm resolution.
Microarray data analysis
The microarray images were processed using GenePix Pro version 6.0 software (Molecular Devices, LLC, Sunnyvale, CA, USA). Spots that did not meet a set of criteria (SNR >5; Dia., 110–250 nm; F CV ≤100; SD ≥1000; satur. ≤1%) were filtered out. The raw data files (deposited in the ArrayExpress database under the accession no. E-MTAB-5434) were loaded into R Bioconductor version 3.2.0 (R Development Core Team 2009) and processed and analyzed with the limma package (36). Briefly, the foreground and background median signal values (excluding those with weights of 0) were background-corrected with the subtraction method and normalized with the global loess method. Probed replicates were averaged, and the replicate arrays from the same patient were merged. Differential gene expression was tested using a linear model suitable for experiments with common reference design. The final p-values were adjusted with FDR correction (37). Data clustering was conducted and visualized with the Heatplus R package.
Real-time PCR analysis
DNA-free RNA (3 µg per sample) was reverse transcribed using SuperScript III RT and oligo(dT) (Invitrogen). The reaction mixtures (20 µl vol) were incubated for 2 h at 50°C, and additional enzyme (1 µl) was added after the first hour. Following reverse transcription, the samples were incubated for 20 min at 70°C with 10 µl of 1 M NaOH. Subsequently, 10 µl of HCl 1 M were added for neutralization, and the cDNA was precipitated overnight at −20°C with 100 µl (2.5 vol) of 96% ethanol and 4 µl (1/10 vol) of 3 M sodium acetate, pH 5.2. The centrifuged pellet was washed twice with 70% ethanol and dissolved in with 60 µl of DEPC-H2O. A total of 1 µl of each cDNA, diluted 3-fold, served as a template for real-time PCR in reactions, including MESA Green qRT-PCR MasterMix Plus (Eurogentec, Seraing, Belgium) and primers specific for the STMN1, NPM1, S100A8 and S100A9 transcripts. From 4 reference genes [ACTB, glyceraldehyde 3-phosphate dehydrogenase (GAPDH), PGK1 and PPIA], the last 2 were selected as the most stable using geNorm (https://genorm.cmgg.be). The sequences of all the primers are presented in Table II. Each gene analysis was performed in duplicate. The standards were generated by PCR amplification of the HeLa cDNA template, obtained as a result of reverse transcription of control HeLa RNA with a SuperScript Plus Indirect cDNA Labeling system (Invitrogen). Real-time analysis was performed in a Rotor-Gene Q thermocycler (Qiagen), using the following program: initial denaturation (94°C, 2 min), 40 cycles of denaturation (94°C, 1 min), annealing (60°C, 1 min) and elongation (72°C, 30 sec), final elongation (72°C, 10 min) and a final hold (4°C). The melting curve was generated in the range of 60–95°C. The reaction volume was 20 µl. Each reaction was performed in triplicate. The real-time PCR products were verified by melting curve analysis. The product quantities were estimated using the two standard curves’ method. The values obtained for the triplicates were averaged. Subsequently, the results obtained for the 2 biological replicates (separate reverse transcription reactions and separate real-time analyses) were averaged and divided by the average of the replicates of the 2 reference genes.
Table II.
PCR primers.
| Gene | Gene ID | Description | NCBI nucleotide reference | Primer sequences (forward and reverse) | Tm | PCR product length name (bp) |
|---|---|---|---|---|---|---|
| ABL1 | 25 | ABL proto-oncogene 1 | NM_005157.5 | 5′-TCATATCAACCCGAGTGTCT-3′ 5′-AACTTGTTCCTCATTTGCTG-3′ |
56°C 54°C |
229 |
| ANXA3 | 306 | Annexin A3 | NM_005139.2 | 5′-CGCAATCAGGTGGAGTCGAG-3′ 5′-TCACTAGGGCCACCATGAGA-3′ |
61°C 60°C |
467 |
| ACTB | 60 | Actin beta | NM_001101.4 | 5′-TCCCTGGAGAAGAGCTACGA-3′ 5′-AGGAAGGAAGGCTGGAAGAG-3′ |
59°C 59°C |
98 |
| CAT | 847 | Catalase | NM_001752.3 | 5′-CTCCGGAACAACAGCCTTCT-3′ 5′-GATGAGCGGGTTACACGGAT -3′ |
60°C 60°C |
412 |
| GAPDH | 2597 | Glyceraldehyde-3-phosphate dehydrogenase | NM_002046.5 | 5′-CCGTCTAGAAAAACCTGCC-3′ 5′-AGCCAAATTCGTTGTCATACC-3′ |
56°C 57°C |
218 |
| NPM1 | 4869 | Nucleophosmin 1 | NM_001037738.2 | 5′-GCGCATTGAACAGTCCTGGG -3′ 5′-CCAGCCTGAAGAGGCATGGGT -3′ |
62°C 64°C |
172 |
| PGK1 | 5230 | Phosphoglycerate kinase 1 | NM_000291.3 | 5′-GGGAAAAGATGCTTCTGGGAA-3′ 5′-TTGGAAAGTGAAGCTCGGAAA-3′ |
58°C 58°C |
72 |
| PPIA | 5478 | Peptidylprolyl isomerase A | NM_021130.4 | 5′-CTGGACCCAACACAAATGGT-3′ 5′-GCCTTCTTTCACTTTGCCAAAC-3′ |
58°C 59°C |
98 |
| S100A8 | 6279 | S100 calcium binding protein A8 | NM_001319196.1 | 5′-TGAAGAAATTGCTAGAGAC-3′ 5′-CTTTATCACCAGAATGAGGA-3′ |
50°C 52°C |
131 |
| S100A9 | 6280 | S100 calcium binding protein A9 | NM_002965.3 | 5′-CCTGGACACAAATGCAGACAA-3′ 5′-CGTCACCCTCGTGCATCTT-3′ |
59°C 60°C |
101 |
| STMN1 | 3925 | Stathmin 1 | NM_203401.1 | 5′-GCCCTCGGTCAAAAGAATCTG-3′ 5′-TGCTTCAAGACCTCAGCTTCA-3′ |
59°C 60°C |
139 |
| WT1 | 7490 | Wilms tumor 1 | NM_000378.4 | 5′-ACAGGGTACGAGAGCGATAACCA-3′ 5′-CACACGTCGCACATCCTGAAT-3′ |
63°C 61°C |
105 |
The primer sequences of 3 reference genes (ACTB, PGK1 and PPIA) were from the study by Lossos et al (69). The STMN1 primers were from the study by Hussein et al (70). The WT1 primers were from the study by Kreuzer et al (71). The design of the remaining primers and melting temperature (Tm) calculation was supported by NCBI Primer Blast (www.ncbi.nlm.nih.gov/tools/primer-blast/).
Droplet digital PCR analysis
DNA-free RNA (2.5 µg per sample) was reverse transcribed and purified, as described above for real-time PCR, with the exception that the RT reaction lasted 1.5 h and only 1 µl of enzyme was used. The precipitated cDNA, dissolved in 50 µl of DEPC-H2O, served as a template for quantitative PCR with the use of a QX200 ddPCR system, QX200 EvaGreen ddPCR Supermix (Bio-Rad, Hercules, CA, USA), and primers specific for 4 candidate genes (ABL1, ANXA3, CAT and WT1) and one reference gene (PGK1). The reaction volume was 20 µl, with a primer concentration of 250 nM, and the volume of cDNA (1–3 µl) was optimized for each candidate gene and sample. The PCR conditions were as follows: initial denaturation (95°C, 5 min), 40 cycles of denaturation (95°C, 30 sec), annealing (60°C, 30 sec) and elongation (72°C, 45 sec), cooling (4°C, 5 min), final denaturation (90°C, 5 min) and final hold (12°C). The temperature ramping rate was 2°C/sec. Following PCR, the plates were directly analyzed with a QX200 Droplet Reader. The data were processed using Quanta Soft version 1.5.38.1118 software (Bio-Rad). The number of droplets for each candidate gene was divided by the number of droplets obtained for the reference gene analyzed in the same run (for ABL1, ANXA3 and WT1) or in the same multiplex reaction (CAT). Each reaction was performed in duplicate. The sequences of all the primers are presented in Table II.
Statistical analysis of RT-qPCR data
Statistical analyses and bar plots were made in R version 3.2.0 R and R Studio version 0.98.1102. Welch two sample t-tests (unpaired) were applied for pairwise comparisons and ANOVA was applied for comparisons across 2 or more groups. For two- or more-way ANOVA, Tukey honestly significant difference (HSD) tests based on multiple comparisons of means were applied to determine which pairwise comparisons were statistically significant. To test the correlations between the expression values of 2 genes or between a particular gene’s expression and clinical data (WBC, age), Pearson’s correlations were calculated. The threshold p-value was always set as 0.05.
Database screening
The following open-source software tools and databases were explored: NCBI databases (www.ncbi.nlm.nih.gov/), including PubMed, Gene and Gene Expression Omnibus (GEO) and GDSbrowser, Expression Atlas (www.ebi-ac-uk/gxa/) and DAVID (david.abcc.ncifcrf.gov/).
Results
Microarray analysis
To study gene expression in AML, we designed a small boutique AML-array, as deposited in the ArrayExpress database (A-MEXP-2220). The array contained >900 long oligonucleotide probes that are complementary to human genes implicated, inter alia, in hematopoietic cell differentiation and maturation, proliferation, apoptosis and leukemic transformation. Gene functional analysis with DAVID (38) revealed that our microarray was significantly enriched in genes linked with immune, infection and cancer classes of disease. Out of OMIM diseases, only AML was found to be statistically significant. The genes covered by the AML-array encode proteins from the following KEGG pathways: Cancer (111 genes), cytokine-cytokine receptor interaction (102 genes), Jak-STAT signaling (64 genes), chemokine signaling (59 genes), MAPK signaling (54 genes), focal adhesion (49 genes), hematopoietic cell lineage (42 genes) and regulation of actin cytoskeleton (41 genes).
The AML-array was used to estimate relative gene expression in PBMCs and/or BMMCs obtained from 33 de novo AML patients, prior to therapy, and 15 HV (Table I). The AML samples were classified as M1 or M2 FAB subtypes, which were characterized on the basis of the early blast cell differentiation arrest and either no myeloid cell maturation (M1) or partial maturation (M2). The microarray experiment was based on two-color hybridization, wherein each studied sample was matched with a common reference (HL60 cell line). Data from 118 hybridizations, as deposited in the ArrayExpress database under the accession no. E-MTAB-5434, were background-corrected and normalized, and technical replicates (of probes and arrays) were then merged and analyzed using the R Bioconductor limma package.
First, we assessed whether there were any differences between the transcriptomes of the BMMCs and PBMCs. Similar to the proteomics analysis results of our previous study (33), there were no statistically significant differences between the AML-PBMCs and AML-BMMCs (differential expression analysis with a limma linear model, adjusted p-value >0.11) (Table III). Therefore, in the following transcriptome analyses, we did not divide the samples into BMMC- and PBMC-derived groups.
Table III.
The summary of microarray data analysis results.
| Comparison | The smallest adjusted p-value (top gene) | The no. of DEGs (over-/underexpressed) | Overexpressed genes | Underexpressed genes |
|---|---|---|---|---|
| AML-PBMC vs. AML-BMMC | 0.11 (FAS) | 0 | FAS, DUSP2 | PRG1, THY1, STAT4, CTSG, CCL7, PRG2, SRGN, SRP9 |
| AML vs. HV | 1.67×10−10 (STMN1) | 163 (85/78) | Genes from Table IV, and ABL1, BCR, CBL, CCL24, CCND1, CCND2, CCNDBP1, CCR10, CD34, CD38, CDKN1A, CFL1, CLU, CTSG, DNAJB4, DNMT3B, GAB1, GSTP1, HOXB5, HOXC8, HPH, HSPA4L, IFNGR2, INPP5D, IRS2, JUNB, METTL3, MME, MT1H, NUP98, PHB2, PPIF, PRG2, RAF1, RHOH, SLC25A1, SLC7A5, SLITRK6, SMAD3, TLE1, TRAF4, ZEB2 | Genes from Table IV, and AIF1, CBFB, CRIP2, CSF3, CYBB, DRAP1, E2F1, FAS, FOSB, GDI1, HIF1A, HLA-DPA1, HLA-DRB1, IGHM, IL12B, KDR, LGALS3, MLLT10, MTMR11, MYLPF, NFKB2, PAK1, PHB2, PROCR, RHAG, S100A8, SHC1, SLC29A1, SULT1E1, TNFRSF10A, XCL1, XCR1, ZFP36 |
| AML-M1 vs. AML-M2 | 0.35 (IL5B) | 0 | IL5B, CASP2, LCP1, FLT3, BIRC2 | PRG1, CST7, MAPK6, SRGN, CTSG |
| AML-M1 vs. AML-M2a | 0.11 (CASP2) | 0 | CASP2, IL5B, BIRC2, FLT3, TPM1 | PRG1, CST7, MAPK6, CTSG, SRGN |
| AML-M1 vs. HV | 2.84×10−08 (STMN1) | 53 (28/25) | Set 1, and ARHGEF12, ERCC2, HSP90AB1, HSPA8, HSPA9, IGFBP7, ANGTP1, INPP5D, ITGB4, KIT, KITLG, MN1, MYB, RHOH | Set 3, and BIRC3, BTG1, CCL5, FCER1G, HCK, IFNA1, LYZ, NFKBIA, PGK1, S100A9, ACTB, CST7, LBD1 |
| AML-M2 vs. HV | 2.09×10−09 (STMN1) | 146 (74/72) | Set 2, and ABCF1, ABL1, ANGTP1, ARF1, ATP6V0C, BCR, CBL, CCL24, CCND2, CCR10, CD34, CDKN1A, CFL1, CTSD, CTSG, DNAJB4, DNMT3B, ETV6, GAB1, GAL, GAPDH, HIGD1A, HPH, HSPA4L, HSPD1, IFNGR2, ITGB4, KIT, KITLG, MAP7, METTL3, MME, MN1, MPO, MYB, MYH9, NPM1, PHB2, PPIF, PRG1, PRG2, RAF1, SLC7A5, SMAD3, TUBB, ZEB2 | Set 4, and BTG1, FCER1G, HCK, HLA-DPB1, PGK1, PLBD1, CBFB, CD44, CD52, CD74, CRIP2, CX3CR1, CYP2E1, DRAP1, FAS, HIF1A, HLA-DRB1, IGHM, JUN, KDR, LCP1, LGALS3, MLLT10, MYLPF, NFKB2, PAK1, PROCR, PTPRE, PXN, RARA, RHAG, RHOB, SLC29A1, SPTBN1, STAT5B, STAU2, SULT1E1, TNFRSF10B, TNFSF10, XCR1, ZFP36 |
| RUNX1/RUNX1T1+ | 1.75×10−06 (STMN1) | 13 (7/6) | STMN1, CDK6, MYC, RPLP0, MPO, HSPA8, ATP6V0C | TMSB4X, LTB, IFITM1, PF4, FCN1, CD44 |
| AML vs. HV FLT3+ AML vs. HV |
3.32×10−07 (CDK6) | 17 (10/7) | CDK6, STMN1, RPLP0, ANGTP1, SET, MCM5, ENO1, RASSF5, DNTT, MYL9 | TRADD, TMSB4X, CAPN10, LTB, PDGFRB, XBP1, SPTBN1 |
| NPM1+ AML vs. HV | 7.35×10−07 (STMN1) | 97 (50/47) | Set 2, and ABCF1, AK2, ANGTP1, ARHGAP4, BCR, DAD1, GAB1, GAL, GAPDH, HSPA1A, HSPD1, IFNGR2, ITGB4, MAP7, MN1, NPM1, NUP98, PRG1, RHOH, SLC25A1, TRAF2, TUBB | Set 4, and HCK, HLA-DPB1, PGK1, PLBD1, CD52, CEBPB, CYBB, FOSB, GJB1, IL12B, IL9, PAK1, PRODH, PTPRE, SPTBN1, SWAP70 |
| CR AML vs. HV | 4.44×10−09 (STMN1) | 74 (31/43) | Set 1, and ADRA2C, HSP90AB1, HSPA8, HSPA9, IGFBP7, ANGTP1, ATP6V0C, GAL, GAPDH, HOXB5, HPH, ITGB4, KIT, MT1H, MYB, PRTN3, TUBB | Set 3, and ADD3, BTG1, CCL3, CCL5, CSF3R, FCER1G, HCK, HLA-DPB1, IFNA1, LYZ, NFKBIA, PLBD1, S100A9, XBP1, AGPAT1, AIF1, BIRC3, CBFB, CD74, CX3CR1, CYP2E1, HIF1A, PTPRE, RARA, RHOB, S100A8, SPTBN1, STAT5B, SULT1E1, TNFSF10 |
| RES AML vs. HV | 7.98×10−07 (STMN1) | 84 (41/43) | Set 2, and ABL1, ARF1, CBL, CD38, CDKN1A, DNAJB4, KIT, MYH9, PRG1, SMAD3, TBL1X, TLE1, TRAP1 | Set 3, and ADD3, BIRC3, CSF3, DUSP2, FCER1G, HLA-DPB1, IGF1, PGK1, PIM1, PLBD1, STK4, XBP1, CRIP2, CYP2E1, DRAP1, GDI1, HLA-DRB1, HOXA3, IGHM, IL8, KDR, MLLT10, NFKB2, PAK1, PXN, RARA, SLC29A1, STAU2, SWAP70, TNFSF10, XCR1 |
| X AML vs. HV | 5.57×10−07 (STMN1) | 79 (37/42) | Set 1, and ARHGEF12, ERCC2, HOXA10, HSPA8, IGFBP7, MYH11, PDE3B, PRKAR1B, ZNF22, ANGTP1, CD81, FUS, GAB1, HSPD1, IFNGR2, KIT, KITLG, MAP7, MN1, MYB, NPM1, RHOH, TUBB | Set 4, and BTG1, FCER1G, CD74, FOSB, IGHM, IL12B, PRAME, PXN, TIPARP, TNFRSF10A, ZFP36 |
| RUNX1/RUNX1T1+ AML vs. remaining AML | 0.99 (TERT) | 0 | TERT, MPO, OGFR, STK32B, CST7, IL5RA, SPI1, PLXNC1 | HPRT1, PML |
| RUNX1/RUNX1T1+ AML vs. remaining AMLa | 0.96 (HPRT1) | 0 | MPO, CXCL2, IL5RA, OGFR, STK32B, CST7, TERT, PLXNC1, SWAP70 | HPRT1 |
| FLT3+ AML vs. remaining AML | 0.48 (VIM) | 0 | VIM, DNTT, CDK6, CEACAM6 | IL4, G6PD, PSMA6, PMAIP1, TRADD, TUBB4 |
| FLT3+ AML vs. remaining AMLa | 0.088 (CDK6) | 0 | CDK6, VIM, CEACAM6, DNTT, CASP3 | PMAIP1, IL4, G6PD, MKI67, CDKN2A |
| NPM1+ AML vs. remaining AML | 0.99 (FOSB) | 0 | ANXA8, TRAF2, NAP1L1 | FOSB, HLA-DPA1, STOM, KLF4, MLLT1, PRODH, EBF1 |
| NPM1+ AML vs. remaining AMLa | 0.9 (FOSB) | 0 | TRAF2, ANXA8, RHOH, PGK2, HSPA1A | FOSB, KLF4, HLA-DPA1, EBF1, MLLT1 |
| NPM1+/FLT3+ AML vs. remaining AML | 0.26 (CPA3) | 0 | CPA3, CCL17, HOXA5, PLCG1, ANXA1, CLEC2B, TUBA1A, CCNG2, VIM | RTN2 |
| NPM1+/FLT3+ AML vs. remaining AMLa | 0.043 (CPA3) | 1 | CPA3, (CASP3, PKN2, CCL17, CLEC2B, PBX3, RB1) | PMAIP1, RTN2, LTA |
| NPM1+/FLT3− AML vs. remaining AML | 0.79 (TRAF2) | 0 | TRAF2, ANXA8, ITGB4, HSPA9, BCR, MYL9, RHOH | PRODH, GTF2E2, STOM |
| NPM1+/FLT3− AML vs. remaining AMLa | 0.28 (TRAF2) | 0 | TRAF2, RHOH, SLC25A1, ANXA8, NONO, ITGB4, ABCF1, BCR, MYL9 | PRODH |
| NPM1−/FLT3+ AML vs. remaining AML | 0.63 (PSMA6) | 0 | PTK7, SRP9, DNAJB5, CD247 | PSMA6, NUP88, CANX, NUBPL, BUD31, CASP2 |
| NPM1−/FLT3+ AML vs. remaining AMLa | 0.36 (CASP2) | 0 | PTK7, CDK6, CEACAM6, FGF13 | CASP2, APOC1, PSMA6, NUP88, NUBPL, CANX |
| NPM1+/FLT3− AML vs. NPM1−/FLT3+ AML | 0.77 (CANX) | 0 | CANX, PSMA6, NUP88, ITGB4, G6PD, TRAF2 | DNAJB5, CEACAM6, PTK7, CEBPB |
| NPM1+/FLT3− AML vs. NPM1−/FLT3+ AMLa | 0.42 (CDK6) | 0 | CASP2, CANX, APOC1, TRAF2, HSPA8, PSMA6 | CDK6, CEACAM6, PTK7, DNAJB5 |
| NPM1+/FLT3− AML vs. NPM1+/FLT3+ AML | 0.56 (RTN2) | 0 | RTN2, IL4, PKLR | CPA3, CCL17, PLCG1, GTF2E2, ITGA4, CCL1, RNH1 |
| NPM1+/FLT3− AML vs. NPM1+/FLT3+ AMLa | 0.17 (PMAIP1) | 0 | PMAIP1, RTN2, IL4 | CPA3, PRKCQ, CCL17, CCL1, PKN2, CASP3, PLCG1 |
| NPM1−/FLT3+ AML vs. NPM1+/FLT3+ AML | 0.83 (DNAJB5) | 0 | DNAJB5, FGF13, CD247 | APOC1, NUBPL, PKN2, NAP1L1 FOXP1, CCL17, BUD31 |
| NPM1−/FLT3+ AML vs. NPM1+/FLT3+ AMLa | 0.09 (PKN2) | 0 | DNAJB5, FGF13, ABR | PKN2, APOC1, CPA3, CASP2, NUBPL, JUN, CCL1 |
| CR AML vs. RES AML | 0.52 (TRAP1) | 0 | GAPDH, HOP, TERT, PXN, MT1H | TRAP1, NFKBIA, ABL1, AGPAT1, NUBPL |
| CR AML vs. RES | 0.46 (GAPDH) | 0 | GAPDH, MT1H, TERT, TUBA1 | NFKBIA, NUBPL, TRAP1, TBL1X, CFLAR, PRDX2 |
| CR AML vs. X AML | 0.72 (ARHGEF12) | 0 | PRAME, ROBO1, RBPMS, TERT | ARHGEF12, MEIS1, PTPN6, EPHB2, ANXA8, AVEN |
| CR AML vs. X AMLa | 0.71 (MEIS1) | 0 | PRAME, ROBO1, IGHM, IL6 | MEIS1, ARHGEF12, ERCC2, PTPN6, CFLAR, HIF1A |
| RES AML vs. X AML | 0.98 (AZU1) | 0 | AZU1, LYZ, SIRPA, STOM, GP1BB, TOP2B, HPH | MAFB, MLLT10, CACNA2D2 |
| RES AML vs. X AMLa | 0.95 (AZU1) | 0 | AZU1, HPH, TOP2B, SIRPA, CD7 | CD81, CACNA2D2, MLLT10, MAFB, BIRC2 |
| CR_long AML vs. CR_short AML | 0.98 (CAST) | 0 | CAST, BIRC5, MAF, FBXL10, PTPRC, SHQ1, MYH11, NPM1 | CRIP2, CREBBP |
| CR_long AML vs. CR_short AMLa | 0.98 (CRIP2) | 0 | CAST, BIRC5, MAF, HMGA2, MCM3, SHQ1, PTPRC | CRIP2, ID1, SQSTM1 |
The analysis repeated with AML samples only (without HV). All analyses were performed with the use of Bioconductor limma package and linear models. Differentially expressed genes (DEGs) were identified with a significance level of α 0.05. p-values were corrected for multiple testing. When no statistically significant genes were identified in a comparison, the table includes the top 10 genes ranked according to p-value (in parentheses). The sets of genes common for multiple comparisons - set 1: CDK6, CEBPA, EEF1G, ENO1, GSN, HOXB6, KRAS, MCM5, MYC, PKM2, RASSF5, RPLP0, SET, STMN1 (14 overexpressed genes common for 7 comparisons); set 2: set 1 and ADRA2C, ARHGEF12, ERCC2, HOXA10, HSP90AB1, HSPA8, HSPA9, IGFBP7, JUNB, MYH11, PDE3B, PRKAR1B, SRGN, ZNF22 (28 overexpressed genes common for 4 comparisons); set 3: B2M, CAPN10, CCL19, FCN1, HOXA9, IFITM1, LTB, MAP4K2, PDGFRB, PF4, TMSB4X, TRADD (12 underexpressed genes common for 7 comparisons); set 4: set 3 and ADD3, BIRC3, CCL3, CCL5, CSF3, CSF3R, DUSP2, E2F1, HLA-DPA1, IFNA1, IGF1, LYZ, NFKBIA, PIM1, RALGDS, S100A9, SHC1, STK4, XBP1 (31 underexpressed genes common for 4 comparisons).
Our next aim was to identify the differences between the AML and HV samples. Assuming a significance level α 0.05 as the threshold, we selected 163 differentially expressed genes (DEGs), of which 78 were underexpressed and 85 were overexpressed in AML (Table III). However, after sample clustering with these DEGs, 2 samples were still misclassified (one AML sample was included in the HV cluster, and one HV was in the AML cluster). The application of a more restrictive threshold (α 0.01) limited the list of DEGs to 83 (42 underexpressed and 41 overexpressed) (Table IV). In total, 36 genes from the list (43%, up- and downregulated in equal proportions) encoded proteins involved in signal transduction and transcriptional regulation, affecting processes, such as cell growth, proliferation, adhesion and apoptosis. Of note, 10 genes (12%, 4 up- and 6 downregulated) were associated with cytoskeletal organization and functioning, and 12 genes (14,5%, exclusively underexpressed in AML) were engaged in immune and inflammatory response. With the use of these 83 DEGs, we were able to perfectly distinguish the AML from the HV samples (Fig. 1A); the entire set of arrays was clearly divided into 2 clusters: 15 control HV samples constituted a separate cluster, whereas all 33 AML samples belonged to a second cluster.
Table IV.
The list of 83 differentially expressed genes between AML and HV samples.
| Gene name (ID) | Gene description | Gene function | Log2FC | Adjusted p-value |
|---|---|---|---|---|
|
A, Genes overexpressed in AML | ||||
| STMN1 (3925) | Stathmin 1 | Regulation of the microtubule filament system by destabilizing microtubules | 2.91 | 1.67×10−10 |
| KITLG (4254) | KIT ligand | The ligand of the tyrosine-kinase receptor, required in hematopoiesis | 2.38 | 0.0088481 |
| CDK6 (1021) | Cyclin-dependent kinase 6 | Cell cycle regulation, G1 phase progression and G1/S transition; altered expression in multiple human cancers | 1.51 | 1.88×10−9 |
| MCM5 (4174) | Minichromosome maintenance complex component 5 | DNA replication, cell cycle regulation; upregulated in the transition from the G0 to G1/S phase | 1.42 | 0.0001986 |
| KRAS (3845) | KRAS proto-oncogene, GTPase | Oncogene, member of the small GTPase superfamily; mutated in various malignancies | 1.42 | 0.0005584 |
| CEBPA (1050) | CCAAT/enhancer binding protein α | Transcription factor, modulator of the expression of genes involved in cell cycle regulation and body weight homeostasis; mutated in AML | 1.32 | 0.0006135 |
| MYC (4609) | MYC proto-oncogene, bHLH transcription factor | Regulation of cell cycle progression, apoptosis and cellular transformation; amplified in numerous human cancers, translocated in Burkitt’s lymphoma and multiple myeloma | 1.26 | 4.32×10−7 |
| ANGPT1 (284) | Angiopoietin 1 | Role in vascular development and angiogenesis, interactions between the endothelium and surrounding matrix, blood vessel maturation and stability | 1.25 | 0.0022923 |
| SRGN (5552) | Serglycin | Hematopoietic cell granule proteoglycan, associated with complex of granzymes and perforin, and granule-mediated apoptosis | 1.17 | 0.0064793 |
| RPLP0 (6175) | Ribosomal protein lateral stalk subunit P0 | Ribosomal protein, component of the 60S subunit, protein synthesis | 1.14 | 0.0000199 |
| ENO1 (2023) | Enolase 1 | Glycolytic enzyme, structural lens protein (tau-crystallin) | 1.14 | 0.0000584 |
| SET (6418) | SET nuclear proto-oncogene | Inhibitor of histone acetylases (HAT), nucleosome acetylation and transcription | 1.07 | 0.0000584 |
| MYB (4602) | MYB proto-oncogene, transcription factor | Transcription regulator, regulation of hematopoiesis, aberrently expressed, rearranged or translocated in leukemias and lymphomas | 0.88 | 0.0058191 |
| GAL (51083) | Galanin and GMAP prepropeptide | Nociception, feeding and energy homeostasis, osmotic regulation and water balance | 0.87 | 0.0015198 |
| HSPA8 (3312) | Heat shock protein family A (Hsp70) member 8 | Chaperone, correct folding of nascent polypeptides, disassembly of clathrin-coated vesicles in transport of cell membrane components (ATPase) | 0.72 | 0.0014811 |
| ADRA2C (152) | Adrenoceptor α 2C | G protein-coupled receptor, regulation of neurotransmitter release at lower levels of nerve activity | 0.70 | 0.0017407 |
| HSPD1 (3329) | Heat shock protein family D (Hsp60) member 1 | Chaperone, folding and assembly of newly imported proteins in the mitochondria, signalling molecule in the innate immune system | 0.61 | 0.0058191 |
| IGFBP7 (3490) | Insulin like growth factor (IGF) binding protein 7 | Regulation of IGF availability in body fluids and tissues and modulation of IGF binding to its receptors, stimulation of prostacyclin production and cell adhesion | 0.59 | 0.0014913 |
| RASSF5 (83593) | Ras association domain family member 5 | Tumor suppressor, inactivated in a variety of cancers, regulation of lymphocyte adhesion, cell growth suppression | 0.58 | 0.0016728 |
| EEF1G (1937) | Eukaryotic translation elongation factor 1 g | Asubunit of the elongation factor-1 complex, enzymatic delivery of aminoacyl tRNAs to the ribosome | 0.56 | 0.0002401 |
| GSN (2934) | Gelsolin | Calcium-regulated protein, assembly and disassembly of actin filaments | 0.55 | 0.0005526 |
| ZNF22 (7570) | Zinc finger protein 22 | DNA binding, transcription regulation | 0.51 | 0.0054429 |
| GAPDH (2597) | Glyceraldehyde-3-phosphate dehydrogenase | Carbohydrate metabolism, phosphorylation of glyceraldehyde-3-phosphate in the presence of inorganic phosphate and nicotinamide adenine dinucleotide (NAD) | 0.49 | 0.0015715 |
| KIT (3815) | KIT proto-oncogene receptor tyrosine kinase | Transmembrane receptor for MGF (mast cell growth factor, stem cell factor), mutated in gastrointestinal tumors, mast cell disease, AML and piebaldism | 0.46 | 0.0017591 |
| HOXA10 (3206) | Homeobox A10 | Transcription factor, regulation of gene expression, morphogenesis, differentiation, fertility, embryo viability, and hematopoietic lineage commitment | 0.43 | 0.0047731 |
| ERCC2 (2068) | ERCC excision repair 2, TFIIH core complex helicase subunit | DNA damage repair, transcription-coupled nucleotide excision repair, member of the basal transcription factor BTF2/TFIIH complex | 0.43 | 0.0005584 |
| ATP6V0C (527) | ATPase H+ transporting V0 subunit c | Component of vacuolar multisubunit enzyme ATPase (V-ATPase), acidification of eukaryotic intracellular organelles, necessary for protein sorting, zymogen activation, receptor-mediated endocytosis, synaptic vesicle proton gradient generation | 0.43 | 0.0080978 |
| HSPA9 (3313) | Heat shock protein family A (Hsp70) member 9 | Role in cell proliferation, stress response and maintenance of the mitochondria | 0.42 | 0.0026984 |
| ARHGEF12 (23365) | Rho guanine nucleotide exchange factor 12 | Numerous cellular processes initiated by extracellular stimuli through G protein-coupled receptors, fusion partner in AML | 0.42 | 0.0014811 |
| PDE3B (5140) | Phosphodiesterase 3B | Regulation of energy metabolism, energy homeostasis and energy intake | 0.42 | 0.0046975 |
| HOXB6 (3216) | Homeobox B6 | DNA binding, sequence-specific transcription factor, regulation of e.g. lung and skin development | 0.40 | 0.0000475 |
| HSP90AB1 (3326) | Heat shock protein 90α family class B member 1 | Signal transduction, protein folding and degradation, morphological evolution, role in gastric apoptosis and inflammation | 0.39 | 0.0006700 |
| ARF1 (375) | ADP ribosylation factor 1 | Phospholipase D activator, a role in vesicular trafficking, intra-Golgi transport | 0.38 | 0.0080978 |
| CTSD (1509) | Cathepsin D | Pepsin-like activity, role in protein turnover and proteolytic activation of hormones and growth factors | 0.37 | 0.0086744 |
| MAP7 (9053) | Microtubule associated protein 7 | Microtubule stabilization, essential for cell polarization, differentiation and spermatogenesis | 0.37 | 0.0058191 |
| ITGB4 (3691) | Integrin subunit β4 | Mediation in cell-matrix or cell-cell adhesion, gene expression and cell growth, receptor for the laminins, pivotal role in the biology of invasive carcinoma | 0.36 | 0.0058191 |
| MYH11 (4629) | Myosin heavy chain 11 | Major contractile protein, rearranged in AML of the M4Eo subtype | 0.35 | 0.0020442 |
| PRKAR1B (5575) | Protein kinase cAMP-dependent type I regulatory subunit β | Signaling pathway of the second messenger cAMP, regulation of ion transport, metabolism, and transcription | 0.32 | 0.0080978 |
| PKM (5315) | Pyruvate kinase M1/2 | Glycolysis, mediation of cellular metabolic effects induced by thyroid hormones, role in bacterial pathogenesis | 0.31 | 0.0006463 |
| TUBB (203068) | Tubulin β class I | Structural component of microtubules | 0.29 | 0.0081504 |
| MN1 (4330) | MN1 proto-oncogene, transcriptional regulator | Translocated in meningioma, myeloproliferation stimulation, leukemogenesis | 0.25 | 0.0070082 |
|
B, Genes underexpressed in AML | ||||
| IFITM1 (8519) | Interferon induced transmembrane protein 1 | Regulation of angiogenesis, tumorigenesis | −2.05 | 0.0000584 |
| LTB (4050) | Lymphotoxin β | Induction of the inflammatory response system, role in normal development of lymphoid tissue | −2.00 | 1.29×10−7 |
| FCN1 (2219) | Ficolin 1 | Plasma protein with elastin-binding activity, expressed in the peripheral blood leukocytes | −1.93 | 0.0000173 |
| BIRC3 (330) | BIculoviral IAP repeat containing 3 | Inhibition of apoptosis by binding to tumor TRAF1 and TRAF2 necrosis factor receptor-associated factors | −1.90 | 0.0014913 |
| LYZ (4069) | Lysozyme | Antimicrobial agent, cleavage of bacterial cell wall peptidoglycan | −1.85 | 0.0005526 |
| ADD3 (120) | Adducin 3 | Membrane skeletal protein, assembly of spectrin-actin network; expression restricted to brain and hematopoietic tissues | −1.60 | 0.0014913 |
| S100A9 (6280) | S100 calcium binding protein A9 | Regulation of cell cycle progression and differentiation, inhibition of casein kinase, antifungal and antibacterial activity | −1.59 | 0.0014811 |
| FCER1G (2207) | Fc fragment of IgE receptor Ig | Key molecule involved in allergic reactions | −1.39 | 0.0006135 |
| PTPRE (5791) | Protein tyrosine phosphatase, receptor type E | Signaling molecule, regulation of cell growth, differentiation, mitotic cycle and oncogenic transformation | −1.39 | 0.0024662 |
| CD74 (972) | CD74 molecule | Associates with class II major histocompatibility complex (MHC), chaperone regulating antigen presentation for immune response, mediation in survival pathways and cell proliferation | −1.23 | 0.0058191 |
| TMSB4X (7114) | Thymosin β4, X-linked | Actin sequestering protein, regulation of actin polymerization, cell proliferation, migration, and differentiation | −1.05 | 1.32×10−8 |
| PF4 (5196) | Platelet factor 4 | Member of the CXC chemokine family, involved in platelet aggregation, inhibitor of hematopoiesis, angiogenesis and T-cell function | −0.97 | 0.0002201 |
| B2M (567) | β-2-microglobulin | Serum protein associating with the MHC class I heavy chain on the surface of nucleated cells, antibacterial activity | −0.95 | 0.0006700 |
| HLA-DPB1 (3115) | Major histocompatibility complex, class II, DP β1 | Central role in the immune system by presenting peptides derived from extracellular proteins, expressed in B lymphocytes, dendritic cells and macrophages | −0.77 | 0.0025915 |
| CSF3R (1441) | Colony stimulating factor 3 receptor | Cytokine controlling production, differentiation, and function of granulocytes, involvement in cell surface adhesion and recognition processes | −0.71 | 0.0025443 |
| HOXA9 (3205) | Homeobox A9 | DNA-binding transcription factor, regulation of gene expression, morphogenesis, and differentiation, translocated in myeloid leukemias | −0.70 | 0.0006135 |
| CCL5 (6352) | C-C motif chemokine ligand 5 | Chemokine involved in immunoregulatory and inflammatory processes, chemoattractant for blood monocytes, memory T helper cells and eosinophils | −0.53 | 0.0033854 |
| CCL3 (6348) | C-C motif chemokine ligand 3 | Small inducible cytokine, macrophage inflammatory protein 1 α, role in inflammatory responses | −0.51 | 0.0034134 |
| RARA (5914) | Retinoic acid receptor α | Nuclear retinoic acid receptor, transcription regulation in a ligand-dependent manner, regulation of development, differentiation, apoptosis, granulopoeisis, and clock genes, translocated in acute promyelocytic leukemia | −0.49 | 0.0053657 |
| CCL19 (6363) | C-C motif Chemokine ligand 19 | Cytokine involved in immunoregulatory and inflammatory processes, antimicrobial, role in normal lymphocyte recirculation and homing | −0.46 | 0.0000612 |
| NFKBIA (4792) | NFKB inhibitor α | Interactions with REL dimers to inhibit NF-κB/REL complexes involved in inflammatory responses | −0.46 | 0.0054429 |
| TNFSF10 (8743) | TNF superfamily member 10 | Cytokine from the tumor necrosis factor (TNF) ligand family, preferential induction of apoptosis in transformed and tumor cells | −0.46 | 0.0033432 |
| CD52 (1043) | CD52 molecule | Lymphocyte surface antigene, involved in fertilisation and response to therapy in large granular lymphocytic leukemia, AML and MDS | −0.45 | 0.0095319 |
| PLBD1 (79887) | Phospholipase B domain containing 1 | Role in the defence against microorganisms, generation of lipid mediators of inflammation | −0.43 | 0.0058191 |
| CAPN10 (11132) | Calpain 10 | Calcium-dependent cysteine protease, associated with type 2 or non-insulin-dependent diabetes | −0.38 | 0.0005178 |
| STAT5B (6777) | Signal transducer and activator of transcription 5B | Transcription factor, signal transduction triggered by IL2, IL4, CSF1, and growth hormones, involved in apoptosis, adult mammary gland development, and sexual dimorphism of liver gene expression | −0.36 | 0.0026984 |
| PGK1 (5230) | Phosphoglycerate kinase 1 | Glycolytic enzyme, conversion of 1,3-diphosphoglycerate to 3-phosphoglycerate, involved in angiogenesis of tumor cells | −0.35 | 0.0064868 |
| RALGDS (5900) | Ral guanine nucleotide dissociation stimulator | Effector of Ras-related GTPase, signaling for a variety of cellular processes | −0.35 | 0.0070755 |
| SPTBN1 (6711) | Spectrin β, non-erythrocytic 1 | Actin crosslinking and molecular scaffold protein, cytoskeleton functioning, determination of cell shape, arrangement of transmembrane proteins, and organization of organelles | −0.35 | 0.0056936 |
| BTG1 (694) | BTG anti-proliferation factor 1 | Anti-proliferative activity, co-activator of cell differentiation, regulation of cell growth | −0.34 | 0.0079167 |
| TRADD (8717) | TNFRSF1A associated via death domain | Programmed cell death signaling and NF-κB activation, suppression of TRAF2-mediated apoptosis | −0.34 | 0.0005526 |
| CYP2E1 (1571) | Cytochrome P450 family 2 subfamily E member 1 | Involved in drug metabolism, synthesis of cholesterol, steroids and other lipids, induced by ethanol, the diabetic state, and starvation | −0.33 | 0.0052990 |
| MAP4K2 (5871) | Mitogen-activated protein kinase kinase kinase kinase 2 | Involved in B-cell differentiation, activated by TNF-α, interacting with TNF receptor-associated factor 2 (TRAF2) | −0.32 | 0.0001674 |
| IGF1 (3479) | Insulin like growth factor 1 | Involved in mediating growth and development | −0.32 | 0.0032632 |
| STK4 (6789) | Serine/threonine kinase 4 | Acting upstream of the stress-induced mitogen-activated protein kinase cascade, involved in apoptosis and chromatin condensation induction | −0.31 | 0.0022923 |
| HCK (3055) | HCK proto-oncogene, Src family tyrosine kinase | Hemopoietic protein, present myeloid and B-lymphoid cell lineages, role in activation of the respiratory burst, neutrophil migration and degranulation of neutrophils | −0.29 | 0.0059276 |
| PDGFRB (5159) | Platelet-derived growth factor receptor β | Cell surface tyrosine kinase receptor for growth factors, development of the cardiovascular system, rearrangement of the actin cytoskeleton | −0.28 | 0.0001623 |
| RHOB (388) | Ras homolog family member B | Tumor suppressor, regulation of cytoskeletal dynamics and vesicle trafficking, control of endothelial barrier function | −0.28 | 0.0048010 |
| XBP1 (7494) | X-box binding protein 1 | Transcription factor regulating MHC class II genes, increase of viral protein expression | −0.27 | 0.0014913 |
| IFNA1 (3439) | Interferon α1 | Antiviral activity, pathogen clearance and tissue inflammation | −0.26 | 0.0024163 |
| PIM1 (5292) | Pim-1 proto-oncogene, serine/threonine kinase | Signal transduction in blood cells, propagation of cell proliferation and survival, tumorigenesis, overexpressed in hematopoietic malignancies and prostate cancer | −0.24 | 0.0013396 |
| PXN (5829) | Paxillin | Cytoskeletal protein involved in actin-membrane attachment at sites of cell adhesion to the extracellular matrix (focal adhesion) | −0.24 | 0.0086744 |
The genes were identified with a significance level of α 0.01 and ranked according to the normalized expression fold change (Log2FC). p-values were corrected for multiple testing. Gene descriptions and functions come from the NCBI gene database.
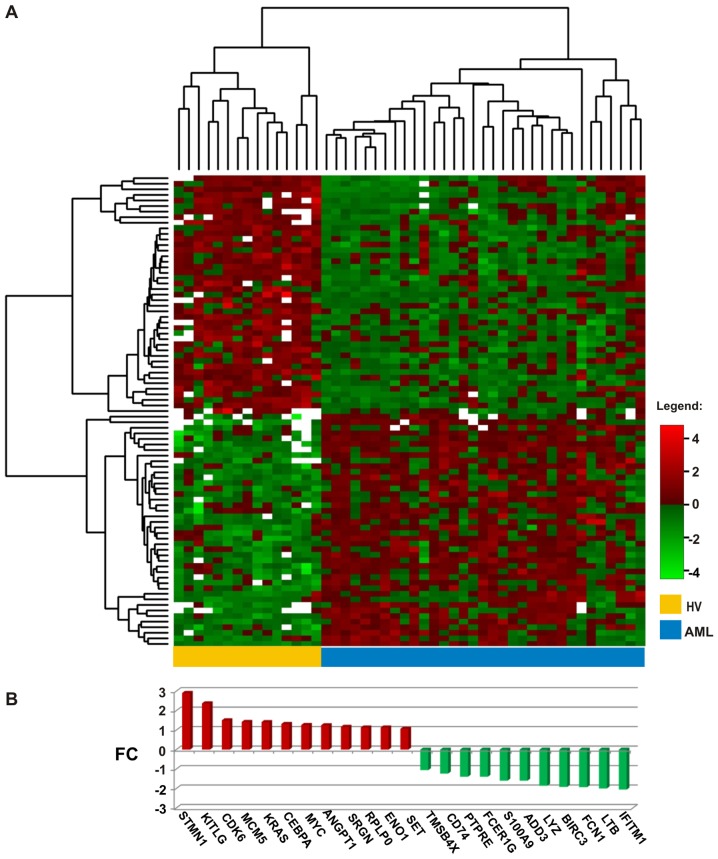
Figure 1.
(A) Heatmap presenting the normalized log2 expression values of 83 genes (horizontally) selected as being over- or underexpressed in acute myeloid leukemia (AML) when compared with healthy volunteers (HV) samples (vertically) with the adjusted p-value threshold of 0.01. Two dendrograms represent the results of clustering: Genes (left side of the heatmap) and samples (above the heatmap). White points mean missing values, referring to the probes with weight 0 that were excluded from the analysis because of poor quality. (B) Changes in the expression (log2FC values) of 23 genes selected as the most overexpressed (red) or underexpressed (green) in AML compared with the HV samples. The genes are ranked according to the log2FC value.
Considering another criterion, the level of expression, we selected from the list of 83 DEGs, 23 genes with the greatest change in expression (log2FC values >1.0 or <−1.0, equivalent to a >2-fold change in expression in both directions): 12 up- and 11 downregulated (Fig. 1B). The most significantly overexpressed gene in AML (log2FC close to 3, equal to an 8-fold enrichment) was STMN1, which encodes a protein involved in the regulation of the microtubule filament system, stathmin 1. STMN1 was also the top statistically significant gene in the comparison between the AML and HV samples (adjusted p-value 1.67×10−10) (Tables III and IV). Other upregulated genes encoded proteins associated with myeloid leukemia and cancer development: Oncogenes, cell cycle and apoptosis regulators [CCAAT/enhancer binding protein α (CEBPA), Kirsten rat sarcoma viral oncogene homolog (KRAS), v-myc myelocytomatosis viral oncogene homolog, avian (MYC), cyclin-dependent kinase 6 (CDK6), minichromosome maintenance complex component 5 (MCM5), SET nuclear proto-oncogene (SET), proto-oncogene c-Kit ligand (KITLG) and serglycin (SRGN)], proteins involved in angiogenesis [angiopoietin 1 (ANGPT1)], cell growth, glucose metabolism and the transcriptional regulator, enolase 1 (ENO1) and ribosomal protein RPLP0.
Among the downregulated genes, predominantly those encoding defense/immunity proteins, were the following: Cell surface receptors CD74 and Fc fragment of IgE (FCER1G), ficolin 1 (FCN1) typically expressed in the peripheral blood leukocytes, antimicrobial agent lysozyme (LYZ), tumor necrosis factor (TNF) family member lymphotoxin β (LTB), interferon induced transmembrane protein 1 (IFITM1) and calcium binding protein S100A9. Moreover, decreased gene expression in AML was observed for the cell cycle regulator protein tyrosine phosphatase (PTPRE), apoptosis suppressor baculoviral IAP repeat-containing 3 (BIRC3), regulator of actin polymerization, cell proliferation, migration and differentiation thymosin β4 (TMSB4X) and membrane skeletal protein adducin 3 (ADD3).
To determine whether AML FAB subtypes could be distinguished by using our boutique array, we compared the gene expression profiles obtained for 11 patients diagnosed as AML-M1 and for 22 patients classified as AML-M2. However, we did not detect any statistically significant differences (adjusted p-value >0.35) (Table III). A separate analysis of AML-M1 vs. HV and AML-M2 vs. HV revealed 54 and 149 DEGs, respectively (α=0.05). Among the 54 DEGs from the comparison between AML-M1 and HV, 50 were shared with the list of 149 DEGs obtained from the comparison between AML-M2 and HV, thus suggesting that the AML-M1 subtype is a subset of AML-M2 and the latter shows some unique features. The list of genes shared between the M1 and M2 FAB subtypes included all the genes shown in Fig. 1B apart from SRGN, CD74, PTPRE and ADD3, which were unique for the comparison between AML-M2 and HV. Other genes shared between M1 and M2 were ERCC2, ITGB4, HSPA8, EEF1G, HOXB6, IGFBP7, KIT, PKM2, RASSF5, MN1, ARHGEF12, HSP90AB1, MYB, HSPA9 and GSN (overexpressed), and CCL19, MAP4K2, PF4, CCL5, B2M, HCK, CAPN10, HOXA9, TRADD, PGK1, PLBD1, IFNA1, BTG1, NFKBIA and PDGFRB (underexpressed). To 4 genes unique for AML-M1 belonged 2 underexpressed (ACTB and CST7) and 2 overexpressed (INPP5D and RHOH) genes. The list of overexpressed genes unique for AML-M2 included inter alia CTSG, CTSD, GAL, MYH11, GAPDH, ADRA2C, ATP6V0C, MAP7, ARF1, METTL3, BCR, TUBB, ABL1, NPM1, CD34, JUNB and MPO, encoding myeloperoxidase, marker protein of AML with maturation.
As regards the effects of mutations, we analyzed the gene expression profiles from the following AML subgroups: RUNX1/RUNX1T1+ [5 patients with translocation t(8;21), typical of the AML-M2 FAB subtype], NPM1+ (8 patients with an NPM1 mutation) and FLT3+ (4 patients with FLT3-ITD). Owing to the co-existence of the FLT3-ITD and NPM1 mutation in some patients, we were also able to distinguish 3 additional subgroups: NPM1+/FLT3+ (2 patients), NPM1+/FLT3− (6 patients) and NPM1−/FLT3+ (2 patients). However, we were not able to identify any distinct clusters or detect significant differences in any of the comparisons (Table III).
The last result worth mentioning regarding the AML analysis using the AML-array is the classification of the patients according to their response to therapy. The association of gene expression with the prognosis and treatment outcome is a clinically important aspect of leukemia studies. Although all the studied samples were collected at the time of diagnosis, further monitoring of patients enabled to divide them into 3 general subgroups corresponding to their responses to therapy: Patients who reached complete remission (CR, 15 patients), patients resistant to therapy (RES, 9 patients) and patients who succumbed to the disease during therapy (X, 9 patients). The CR subgroup was further divided into 2 smaller subgroups: Patients with complete remission lasting up to 1 year (CR-short, 12 patients) and patients with complete remission lasting >1 year (CR-long, 3 patients). We did not detect any statistically significant differences in gene expression between any of the above subgroups (Table III).
In all analyses performed with the AML-array, significant differences were detected only when an AML subgroup was compared with HV. Notably, the obtained lists of DEGs, despite being of variable length, consisted a high number of shared genes, and STMN1 was always at the top of the ranking (Table III). To eliminate the HV impact, we repeated all the analyses, focusing solely on the AML samples. The result was the same: No statistically significant differences were detected between AML subgroups apart from one: The overexpression of the CPA3 gene, encoding a carboxypeptidase, in NPM1+/FLT3+ AML vs. remaining AML samples (p=0.043) (Table III). Unsupervised hierarchical clustering of the data also indicated that the patients with AML were generally a homogenous group, despite individual variations (Fig. 2).
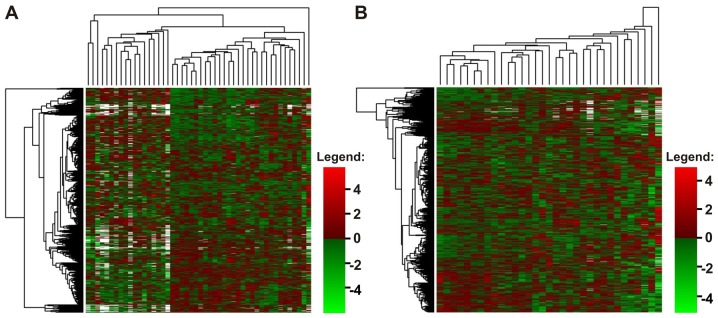
Figure 2.
Heatmaps presenting the normalized log2 expression values of all acute myeloid leukemia (AML)-array genes (horizontally) in all studied samples, (A) AML and healthy volunteers (HV), and (B) only in AML samples, excluding HV. Dendrograms represent the results of clustering: Genes (left side of the heatmap) and samples (above the heatmap). White points indicate missing values, referring to the probes with weight 0 that were excluded from the analysis because of poor quality. (A) Reveals the existence of outlier samples (first sample from the right and three samples from the left) and two major clusters (in the middle), with all HV samples collected at the left side of a heatmap. (B) Shows general homogeneity of AML samples. No clear sample clusters cannot be distinguished.
Real-time PCR analysis
To verify the results of the microarray analysis, we selected 4 genes, STMN1, NPM1, S100A9 and S100A8, and quantified their expression with the use of real-time PCR and a Rotor-Gene Q instrument. In this case, our sample set contained 14 HV and 21 AML samples. The STMN1 gene was selected as the most upregulated gene in AML compared with HV and NPM1 was selected as upregulated only in some AML subgroups (AML-M2, AML-NPM1+ and AML X), whereas S100A9 was selected as one of the most downregulated genes in AML and different AML subgroups compared with HV (Table III). S100A8 was underexpressed with statistical significance only in AML and in the AML-CR subgroup (Table III); however, we were interested whether 2 genes belonging to the same gene family present similar expression patterns.
First, owing to the variable expression of many so-called ‘housekeeping genes’, we tested 4 commonly used reference genes: ACTB, GAPDH, PGK1 and PPIA. The expression stability of each gene across the samples was estimated with geNorm. PGK1 and PPIA were indicated as the most stable, whereas GAPDH was the most variable, showing a trend toward upregulation in the AML samples (data not shown). This finding was consistent with the results of microarray analysis, in which GAPDH was in the list of DEGs between the AML and HV samples. Finally, 2 reference genes, PGK1 and PPIA, were used as normalization factors. Gene expression was analyzed in the context of AML subtypes, mutation status and response to therapy (Figs. 3 and 4).
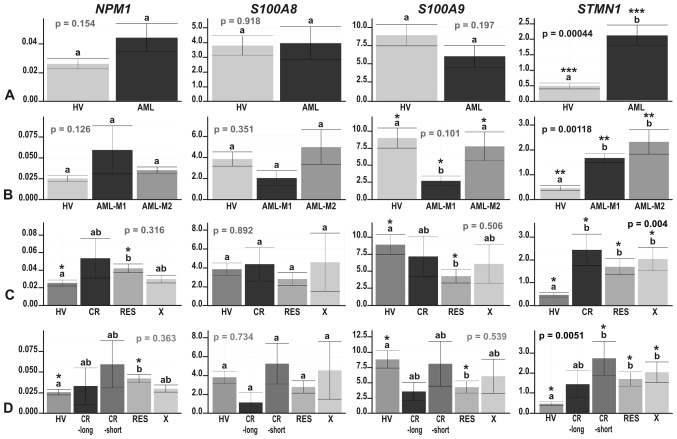
Figure 3.
The results of real-time PCR-based expression analysis of 4 genes (NPM1, S100A8, S100A9 and STMN1) in (A) acute myeloid leukemia (AML) compared with healthy volunteers (HV) samples; (B) in the context of the AML French-American-British (FAB) subtypes, M1 and M2; and in the context of the response to therapy, with the AML divided into (C) 3 (CR, RES, X) subgroups or (D) 4 subgroups (CR_long, CR_short, RES, X). The y-axes show relative expression values of a studied gene compared to the reference genes. Each plot contains the ANOVA p-value obtained for a whole test. Pair-wise comparisons with statistically significant differences, verified with a t-test and HSD Tukey test, are indicated by letters and asterisk symbols above each bar. Bars showing the same letter indicate no significant difference (p>0.05). The numbers of asterisks denote the level of statistical significance: *p≤0.05, **p≤0.01 and ***p≤0.001.
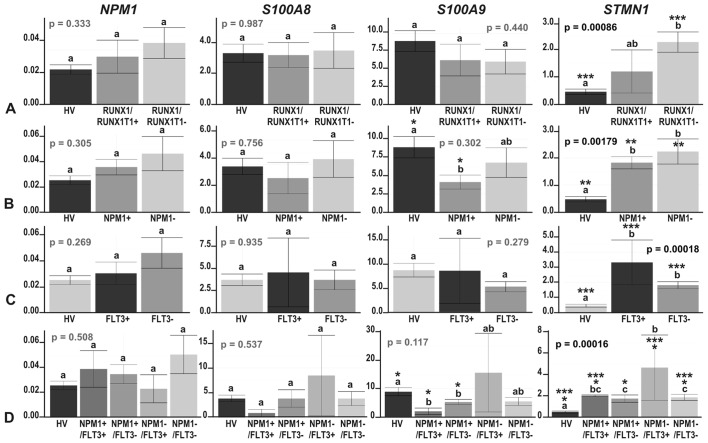
Figure 4.
The results of real-time PCR-based expression analysis of 4 genes (NPM1, S100A8, S100A9 and STMN1) in acute myeloid leukemia (AML) compared with healthy volunteers (HV) samples, with the AML divided according to the mutation status of (A) RUNX1/RUNX1T1; (B) NPM1; (C) FLT3; and (D) NPM1 and FLT3 together. The y-axes show relative expression values of a studied gene compared to the reference genes. Each plot contains the ANOVA p-value obtained for a whole test. Pair-wise comparisons with statistically significant differences, verified with a t-test and HSD Tukey test, are indicated by letters and asterisk symbols above each bar. Bars showing the same letter indicate no significant difference (p>0.05). The numbers of asterisks denote the level of statistical significance: *p≤0.05, **p≤0.01 and ***p≤0.001.
The most consistent results between the microarray and real-time PCR experiments were obtained for the STMN1 gene, which was significantly induced in AML compared with HV (p=0.00044), although the fold change was somewhat lower (~4.5-fold compared with an almost 8-fold change in the microarray data analysis) (Fig. 3A). Similar to the microarray data analysis results, STMN1 upregulation was visible in each AML subgroup compared with HV (t-test p=0.038–0.000019) (Figs. 3 and 4). There were no significant differences in STMN1 expression between particular AML subgroups with the following two exceptions: NPM1−/FLT3− vs. NPM1−/FLT3+ (p=0.015) and NPM1−/FLT3+ vs. NPM1+/FLT3− (p=0.029). The highest STMN1 expression was observed in the NPM1−/FLT3+ subgroup (Fig. 4D).
As regards NPM1 gene expression, we observed a trend toward an upregulation in AML (Fig. 3A); however, the increase in NPM1 expression was statistically significant in only one comparison (RES vs. HV; t-test p=0.021) (Fig. 3C and D).
Real-time PCR-based analysis of S100A8 and S100A9 genes confirmed their general suppression trend in AML or at least some AML subgroups compared to HV. The expression levels of these 2 relative genes, coding for calcium-binding proteins, highly correlated (Pearson’s correlation, 1; p-value <2.2e−16); however, a statistically significant decrease was observed only for S100A9: In the AML-M1 subtype compared with HV (t-test p=0.0012), as well as with AML-M2 (t-test p=0.038) (Fig. 3B), in NPM1-mutated AML vs. HV (t-test p=0.013) (Fig. 4B), irrespective of the FLT3 mutation status (t-test p=0.048 for HV vs. NPM1+/FLT3− and t-test p=0.01 for HV vs. NPM1+/FLT3+) (Fig. 4D), and in the RES subgroup vs. HV (t-test p=0.018) (Fig. 3C and D).
Comparisons of the results from the transcriptomic and proteomic studies
As the same group of AML samples was also previously analyzed in a proteomic analysis in our laboratory (33,34), in this study, we attempted to compare the results of the AML transcriptome and proteome analyses. However, owing to the limitations of both approaches, the comparison could be made for only a small subset of genes/proteins. The AML-array consisted of probes that were complementary for >900 genes, representing ~4% of all human genes. However, the 2-DE-MS analysis of AML proteomes enabled the detection of <200 proteins (33), representing much <1% of all the human proteins. In detail, from the 184 unique proteins detected in the proteomic study, only 52 (28%) had corresponding gene-specific probes on the AML-array. From the 184 identified proteins, 40 were determined to be differentially accumulated in the studied samples, of which 20 were represented on the AML-array: 13 by gene-specific probes and 7 by the probes complementary to the related genes (from the same gene family). Half of the results of the proteomics and transcriptomic analyses were concordant or partially concordant, but the other half was discordant (Table V).
Table V.
Comparisons of the results from the transcriptomic and proteomic studies.
| Protein identified in the proteomic study (33) | Corresponding gene | Microarray probe | Protein accumulation | Gene expression | Concordant |
|---|---|---|---|---|---|
| Actin γ1 | ACTG1 | Target-specific | In HV present only in BM, in AML higher in CR-short | In HV higher in PB, in AML higher in CR-short | Partially |
| α-actinin | ACTN4 | For ACTN1 | Higher in AML | Slightly lower in AML | No |
| Fructose-bisphosphate aldolase A | ALDOA | For ALDOC | In HV present only in BM | In HV higher in PB | No |
| Annexin I | ANXA1 | Target-specific | In HV higher in BM, higher in AML vs. HV and in CR vs. RES AML | In HV higher in BM, no difference between AML and HV, lower in CR vs. RES AML | No |
| Annexin III | ANXA3 | Target-specific | In HV higher in PB, in AML present only in M2 | In HV higher in BM, higher in AML vs. HV, higher in AML M1 vs. M2 | No |
| Rho GDP-dissociation inhibitor 2 | ARHGDIB | For ARHGAP4, ARHGEF1, ARHGEF12 | In HV higher in BM, higher in AML vs. HV | In HV higher in PB, ARHGEF12 and ARHGAP4 - higher in AML, ARHGEF1 - slightly lower in AML | Partially |
| Catalase | CAT | Target-specific | In HV higher in PB, higher in AML and in AML M2 vs. M1 | In HV higher in PB, slightly higher in AML vs. HV, higher in AML M2 vs. M1 | Yes |
| Cofilin-1 | CFL1 | Target-specific | In HV higher in PB | In HV slightly higher in PB | Yes |
| α-enolase (ENO1 protein) | ENO1 | Target-specific | In HV higher in BM | In HV higher in PB, significantly higher in AML vs. HV | No |
| Glutathione transferase ω | GSTO (1,2) | For GSTP1 | Present only in AML CR | Significantly higher in AML, slightly lower in CR | No |
| Tumor rejection antigen (Gp96) | HSP90B1 | Target-specific | In HV higher in PB, lower in AML vs. HV | In HV higher in PB, higher in AML vs. HV | Partially |
| L-Plastin (lymphocyte cytosolic protein 1) | LCP1 | Target-specific | Present only in AML M2 | Significantly lower in AML vs. HV, higher in AML M1 | No |
| Pyruvate kinase | PKM | Target-specific | Much higher in AML vs. HV | Significantly higher in AML vs. HV | Yes |
| Purine nucleoside phosphorylase | PNP | Target-specific | In HV higher in BM, lower in AML | In HV higher in PB, higher in AML | No |
| Acetyl-cypa:cyclosporine complex [peptidylprolyl isomerase A (cyclophilin A)] | PPIA | For PPIF | In HV higher in PB | In HV no difference between PB and BM, higher in AML vs. HV | No |
| Peroxiredoxin-2 | PRDX2 | Target-specific | In HV higher in BM, higher in AML vs. HV PB, lower in AML vs. HV BM | In HV higher in BM, slightly lower in AML vs. HV | Partially |
| Histone-binding protein RBBP4 (retinoblastoma binding protein) | RBBP4 | Target-specific | Higher in AML, in HV merely detectable | Slightly lower in AML vs. HV | No |
| Tropomyosin α | TPM1 | Target-specific | In HV present only in PB, much lower in AML vs. HV | In HV higher in BM, lower in AML vs. HV | Partially |
| Tubulin β | TUBB | Target-specific | In HV higher in BM, lower in AML vs. HV | In HV higher in BM, higher in AML vs. HV | Partially |
| 14-3-3 protein ζ/δ | YWHAZ | For YWHAQ | In HV higher in PB, lower in AML vs. HV | In HV higher in PB, slightly lower in AML vs. HV | Yes |
Droplet digital PCR analysis
To verify some of the discrepancies between the transcriptomic and proteomic results, we applied another quantitative PCR method, ddPCR, which is one of the most precise, sensitive and accurate PCR types. We expanded our research group to 20 HV samples and 41 AML samples. For the analysis, performed with a QX200 Droplet Digital PCR system (Bio-Rad), we selected 4 genes. Two of these, ANXA3, encoding Annexin 3, and CAT, encoding catalase, were determined as being differentially accumulated in the proteomics analyses. None of these genes were identified with the AML-array as being differentially expressed, and the expression trend was concordant for CAT, but was discordant for ANXA3. As a type of positive control, we used the ABL1 oncogene, which was slightly, but significantly, increased in AML-M2 and AML-RES compared with HV in the microarray experiment. Additionally, we analyzed Wilms tumor 1 gene (WT1), which has recently been described as overexpressed and prognostically relevant in most acute leukemias (39,40), but was not included in our AML-array probe set. As a reference gene, we selected PGK1, on the basis of our previous real-time PCR analysis. As regards the CAT gene, we were able to multiplex reactions with a reference gene, due to the differences in PCR product sizes (Table II). For the ABL1, ANXA3 and WT1 genes, a reference gene was analyzed in a separate reaction, but always within the same run. The data from duplicate experiments were separately processed in Quanta Soft version 1.5.38.1118 software (Bio-Rad). The relative expression value of each gene was calculated as the ratio of droplet numbers for the studied gene vs. the reference gene. The results of the statistical analysis, performed in R Bioconductor, are presented in Figs. 5 and 6.
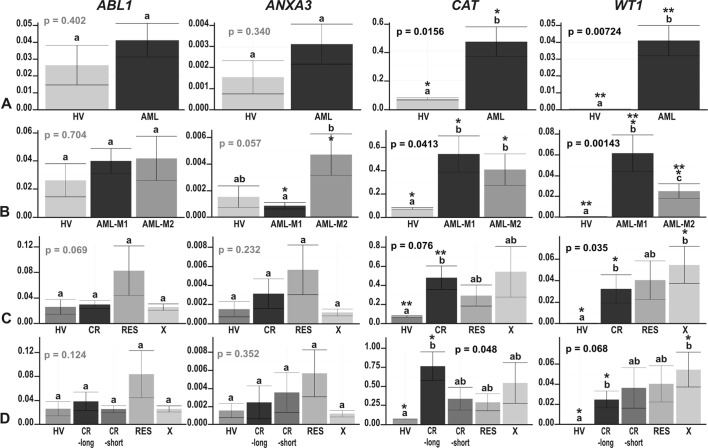
Figure 5.
The results of ddPCR-based expression analysis of 4 genes (ABL1, ANXA3, CAT and WT1) in (A) acute myeloid leukemia (AML) compared with healthy volunteers (HV) samples; (B) in the context of the AML French-American-British (FAB) subtypes, M1 and M2; and in the context of the response to therapy, with the AML divided into (C) 3 (CR, RES, X) subgroups or (D) 4 subgroups (CR_long, CR_short, RES, X). The y-axes show relative expression values of a studied gene compared to a reference gene. Each plot contains the ANOVA p-value obtained for a whole test. Pair-wise comparisons with statistically significant differences, verified with a t-test and HSD Tukey test, are indicated by letters and asterisk symbols above each bar. Bars showing the same letter indicate no significant difference (p>0.05). The numbers of asterisks denote the level of statistical significance: *p≤0.05 and **p≤0.01.
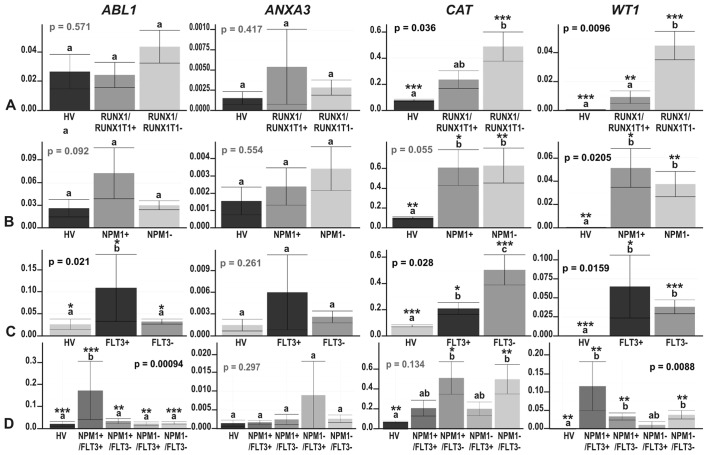
Figure 6.
The results of ddPCR-based expression analysis of 4 genes (ABL1, ANXA3, CAT and WT1) in acute myeloid leukemia (AML) compared with healthy volunteers (HV) samples, with the AML divided according to the mutation status of (A) RUNX1/RUNX1T1; (B) NPM1; (C) FLT3; and (D) NPM1 and FLT3 together. The y-axes show relative expression values of a studied gene compared to a reference gene. Each plot contains the ANOVA p-value obtained for a whole test. Pair-wise comparisons with statistically significant differences, verified with a t-test and HSD Tukey test, are indicated by letters and asterisk symbols above each bar. Bars showing the same letter indicate no significant difference (p>0.05). The numbers of asterisks denote the level of statistical significance: *p≤0.05, **p≤0.01 and ***p≤0.001.
Our positive control gene, ABL1, revealed a trend towards an upregulation in AML, statistically significant in FLT3- and NPM1-mutated AML vs. HV (HSD Tukey test p=0.02 for HV vs. FLT3+; HSD Tukey test p=0.0003 for HV vs. NPM1+/FLT3+) (Fig. 6C and D). ABL1 gene expression was also significantly upregulated in the FLT3+ samples when compared with the FLT3− samples (HSD Tukey test p=0.02) (Fig. 6C) and significantly different in the following comparisons: NPM1−/FLT3− vs. NPM1+/FLT3+ (HSD Tukey test p=0.0003), NPM1+/FLT3+ vs. NPM1+/FLT3− (HSD Tukey test p=0.002) and NPM1+/FLT3+ vs. NPM1−/FLT3+ (HSD Tukey test p=0.0097). The ANXA3 gene was generally expressed at a very low level, with a trend towards overexpression in the AML vs. HV; however, the greatest increase in ANXA3 expression was observed in AML-M2 (Fig. 5B). The only statistically significant difference in the expression of this gene was observed between AML-M1 and AML-M2 (t-test p=0.023) (Fig. 5B), what definitively confirmed the proteomics results. Partially concordant results were obtained for the CAT gene. In agreement with the proteomics results, we observed a significant increase in CAT expression in AML and some AML subgroups compared with HV (Figs. 5 and 6, t-test p=0.0096 for HV vs. AML-M1; t-test p=0.022 for HV vs. AML-M2; t-test p=0.00085 for HV vs. RUNX1/RUNX1T1−; p=0.006 for HV vs. NPM1−; p=0.02 for HV vs. NPM1+; p=0.00085 for HV vs. FLT3−; t-test p=0.04 for HV vs. FLT3+; t-test p=0.008 for HV vs. NPM1−/FLT3−; t-test p=0.03 for HV vs. NPM1+/FLT3−). We also detected a significant increase in CAT expression in the FLT3− compared to the FLT3+ samples (t-test p=0.023). However, the differences in the CAT gene expression levels between the M1 and M2 AML subtypes were subtle and not significant (t-test p=0.52), in contrast to the proteomics findings (Fig. 5B). As regards the response to therapy, the only statistically significant difference, and the most elevated CAT expression was observed in the CR and CR-long AML subgroups (t-test p=0.0044 for HV vs. CR and t-test p=0.013 for HV vs. CR-long) (Fig. 5C and D). As regards the WT1 gene, the ddPCR experiment revealed an evident and statistically significant upregulation of this gene in all AML subgroups, whereas in the HV samples, its expression was scarcely detectable (t-test p=0.046–0.000066) (Figs. 5 and 6). An interesting observation was the increasing trend in WT1 expression in the therapy-related AML subgroups: The higher the WT1 level, the less responsive was AML to therapy (Fig. 5C and D). High levels of WT1 were also detected in the AML-M1 (Fig. 5B) and AML samples without the RUNX1/RUNX1T1 mutation (Fig. 6A), but harboring NPM1, FLT3 or both of these mutations (Fig. 6B–D). The differences across the particular AML subgroups were not significant, apart from 2 comparisons: AML-M1 vs. AML-M2, where WT1 was upregulated in AML-M1 (HSD Tukey’s test p= 0.039) (Fig. 5B) and RUNX1/RUNX1T1+ vs. RUNX1/RUNX1T1−, where WT1 was upregulated in AML without translocation t(8;21) (t-test p=0.0023) (Fig. 6A).
Correlations among the molecular and clinical features of AML
In order to identify links between gene expression measured by RT-qPCR, the mutation status and the clinical characteristics of the patients with AML (WBC, age and sex), we performed a series of statistical tests (t-test, one- and two-way ANOVA, Tukey multiple comparison of means and Pearson’s correlation). The strongest correlations were observed between the WBC counts and the AML FAB subtypes (one-way ANOVA, p=0.0101, WBC counts were higher in AML-M1), WBC counts and NPM1 mutation status (one-way ANOVA, p=0.0229), WBC counts and FLT3 mutation status (one-way ANOVA, p=0.0308), and WBC counts and both FLT3 and NPM1 mutation status (two-way ANOVA, p=0.0160), and the highest WBC count was observed in mutated AML samples.
As regards the effects of age, we observed a moderately positive correlation between patient age and NPM1 gene expression (Pearson’s correlation, 0.348; p=0.1222), a moderately negative correlation between age and the expression of the 2 genes, S100A9 (Pearson’s correlation, −0.380; p=0.0899) and ANXA3 (Pearson’s correlation, −0.33; p=0.046) and a correlation between age and the mutation status of NPM1 (one-way ANOVA, p=0.0411) or NPM1 and FLT3 together (two-way ANOVA, p=0.0446). There were no correlations between the RT-qPCR-determined expression of any other gene and age (Pearson’s correlations equal to −0.23, 0.019, 0.15, 0.07 and 0.2 for S100A8, STMN1, ABL1, CAT and WT1, respectively). We also did not find any correlations between the gene expression levels and sex (one-way ANOVA, p-values equal to 0.45, 0.659, 0.286, 0.555, 0.633, 0.302, 0.628 and 0.263 for NPM1, S100A8, S100A9, STMN1, ABL1, ANXA3, CAT and WT1, respectively) or WBC count (Pearson’s correlations equal to −0.02, −0.25, −0.27, −0.02, 0.07, −0.19, −0.18 and 0.28 for NPM1, S100A8, S100A9, STMN1, ABL1, ANXA3, CAT and WT1, respectively).
Of note, there was no correlation between NPM1 gene expression and the NPM1 mutation status (one-way ANOVA; p=0.634). The presence of the translocation t(8;21) and fusion gene RUNX1/RUNX1T1 did not correlate with any molecular or clinical characteristic.
Gene expression database exploration
To compare our results with other AML gene expression profiling results, we explored the Gene Expression Omnibus (GEO) database (www.ncbi.nlm.nih.gov/sites/GDSbrowser) to identify the most relevant datasets. We selected 7 datasets, including 6 derived from Affymetrix Gene Chips, the most commonly used microarray platform, and one from Agilent microarrays. Among these datasets, only 3 (GDS3057, GDS1059 and GDS2251) contained healthy control samples. These 3 datasets were screened with 238 names of genes, which we had identified as differentially expressed between AML (or particular AML subgroups) and HV by using the AML-array, RT-qPCR and proteomics. Table VI shows the results of comparative analysis. The percentage of genes with an expression trend concordant in our data and any GEO dataset, was in the range 47.8–51.9% (mean, 49.2%) or in the range 51.8–78.3% (mean, 65.3%), when including partially concordant genes, for which expression trends were concordant for some (but not all) conditions/cell lines/microarray probes. Interestingly, the level of coincidence between GEO datasets was much lower: The percentage of concordant genes remained in the range 35.3–43.4% (mean, 39.3%) or in the range 47.4–64.2% (mean, 56.6%), when including partially concordant genes. In reference to discordant genes, an opposite effect could be observed, although the difference was smaller: The percentage of discordant genes in comparative analysis between our data and any GEO data set was in the range 15.5–31.3% (mean, 22.8%) while within GEO datasets in the range 10.0–27.1% (mean, 19.3%). Analyzing 238 gene expression trends, we were able to select 45 strong candidate genes with expression trends concordant in our dataset and all 3 GEO datasets. In total, 26 genes (ABCF1, ABL1, AK2, ANGPT1, BCR, CD34, CDK6, CEBPA, HOXA10, HSP90AB1, HSPA8, HSPA9, IGFBP7, MAP7, MCM5, MT1H, MYB, NPM1, RBBP4, RPLP0, SET, SLC25A1, SMAD3, STMN1, TRAF4 and WT1) were consequently increased in AML when compared to the normal control, whereas 19 were decreased in AML (BTG1, CEBPB, CSF3R, CX3CR1, CYBB, FCER1G, FCN1, HCK, IGHM, LCP1, LGALS3, LYZ, PF4, S100A8, S100A9, STAT5B, STK4, TMSB4X and TRADD). The above-mentioned list contained 14 (10 increased and 4 decreased) from the 23 genes presented in Fig. 1B, identified as the genes with the highest expression fold change in AML-array-based analysis, and one gene (RBBP4) identified in our data only by proteomic approach. Not all of these genes were up-to-date described as AML biomarkers. It is also worth mentioning that 2 members of the CCAAT/enhancer binding protein family of transcription factors, CEBPA and CEBPB, exhibited opposite expression trends in all studied datasets.
Table VI.
The results of comparative analysis of our data and 3 GEO datasets.
| Dataset | Our data | GDS3057 | GDS1059 | GDS2251 |
|---|---|---|---|---|
| Our data | 47.8 (51.8) | 51.9 (59.8) | 47.8 (78.3) | |
| GDS3057 | 15.5 | 43.3 (47.4) | 39.3 (58.1) | |
| GDS1059 | 31.3 | 20.9 | 35.3 (64.2) | |
| GDS2251 | 21.7 | 10.0 | 27.1 |
Only 238 genes identified as differentially expressed in our data (using AML-array, RT-qPCR and proteomics) were considered. The numbers refer to the percentage of concordant genes (the same trend in expression; regular font; the numbers in parentheses include also partially concordant genes) or discordant genes (the opposite trend; italics). Each percentage value was calculated based on the number of genes represented in both compared data sets (214–229).
As regards FAB subtypes, we analyzed the expression trends of 6 genes, including 5 genes encoding proteins determined as differentially accumulated between AML-M1 and AML-M2 by proteomic approach (ANXA3, CAT, LCP1, PGD and PRDX6) (33) and one gene identified with ddPCR (WT1). In comparison to the GDS1064 dataset, referring to AML in different stages of maturation, we observed concordant results for only 2 genes (33%), WT1 and ANXA3, the only 1 gene out of 5 selected by proteomic approach.
Discussion
Taking advantage of genomic tools, tremendous progress in the molecular studies of AML has been achieved (16,41). Although many studies have already been published in this area, to the best of our knowledge, our research is the first transcriptomics analysis of the Polish adult AML population. Moreover, for the same group of patients, a proteomics analysis has also been performed (33). The number of samples was not high; however, the advantage was the preselection of the M1 and M2 FAB subtypes and a significant enrichment in CN-AML samples. Previous microarray-based AML analyses have often included more subtypes, but fewer patients. The present study is also one of the few reports of custom-made array applications in AML studies.
The majority of AML research has been conducted with the use of Affymetrix Gene Chips (19,22,42–44), and some studies have been conducted with large cDNA microarrays (21,45,46). Both platforms utilize arrays constructed from probes corresponding to at least several thousand genes. The advantages of boutique arrays are their lower price, higher flexibility and smaller laboratory infrastructure requirements. Comparative analysis of our results with the results of 3 other gene expression studies based on Affymetrix Gene Chips, proved the utility and reliability of the AML-array. The level of concordance was even higher for our results and any GEO dataset than between GEO datasets. However, we still observed discordant results for ~20% of genes, irrespectively of the datasets compared. There are 3 main reasons for this discrepancy: Different patients, different laboratories, and varying numbers of samples, all of which contribute to technical and biological bias.
The application of the boutique array presented herein enabled us to identify a set of genes differentiating AML and HV. However, we were not able to detect statistically significant differences between AML subgroups. The lack of success can be explained by a few factors: The method was not effective enough, the studied samples were not an accurate representation of the population, or the sample size was not sufficient. The limitation of the method is a pre-selected set of genes, which could be not adequate to distinguish between particular AML subgroups. Home-made microarray production could also contribute to more technical bias comparing to commercial gene chips. Based on our experience with microarray design, production, hybridization and data normalization, we performed many technical replicates, carefully inspected microarray images and filtered out all slides which did not respect quality criteria. As regards patients with AML, we did not apply additional selection criteria; we included all AML patients classified as M1 or M2 FAB subtypes at the time of our project. Unsupervised clustering revealed that the AML subpopulation was generally homogenous. In addition, some of the studied AML subgroups were represented by only a few samples, and thus statistical significance was more difficult to achieve. Probably, a substantial increase in the sample size could overcome this problem. Taking into account the above-mentioned limitations, we consider that microarray-based analysis should be always treated as a first-line screening tool. When precise gene expression measurements are needed, RT-qPCR, in particular its more sophisticated version, ddPCR, is more reliable.
Among the genes overexpressed in AML in the presented AML-array-based study, there were already-established AML markers, e.g., KIT, MYH11, MN1, MPO, SET and HOXA10 (44,47,48), genes associated with AML or potential biomarkers, such as STMN1, CDK6 and ANGPT1, or genes not yet discussed in the context of AML, such as RPLP0 or ENO1, reported in neoplasms other than leukemia (49,50). As these genes often play fundamental roles in cell metabolism and function, their upregulation may be explained by the needs of the highly proliferating cancer cells. The STMN1 gene, which encodes stathmin 1, formerly known as oncoprotein 18 (Op18), was the most significantly overexpressed gene that we observed in AML. The upregulation of this gene, confirmed by real-time PCR, is consistent with the results of other gene expression studies. STMN1, highly expressed in acute leukemias, lymphomas and intensively proliferating cells (51,52) and decreased after the induction of cell differentiation (53), is a good AML biomarker candidate and potential therapeutic target. Other possible candidates are CDK6, which encodes one of the cyclin dependent kinase and cell cycle regulators and prevents differentiation and apoptosis (54), and ANGPT1, which encodes a proangiogenic protein abundant in AML (55) and early myeloblast cell lines (56).
Similarly, the suppression of numerous genes in the AML samples detected with the AML-microarray was consistent with data in the literature. The examples are PF4, which is among the 10 hematopoietic-specific genes found to be significantly downregulated in CD34+ cells obtained from patients with therapy-related AML (57), or TMSB4X, which encodes thymosin β4 and is a regulator of actin polymerization, cell proliferation, migration and differentiation. As TMSB4X expression was higher in the lymphoid than the granulocytic lineage, and its transcript levels increased during granulocyte maturation, it has been proposed to be a marker of differentiation in hematopoietic cells (58). For some genes, such as FCN1, which encodes ficolin 1 and was the second most downregulated gene in AML in our microarray experiment, we did not find any reports associating its gene function with AML. The role of this gene may therefore be the subject of further studies.
From a diagnostic point of view, it is important to correlate gene expression with the classes and types of a disease. For example, the distinction between M1 and M2, the two most frequent and closely related AML FAB subtypes, is challenging. Microarray analyses of AML usually result in the clear separation of AML M3 (APL) and AML with the most frequent cytogenetic aberrations (42,21,22). The M4 subtype has additionally been distinguished by (43) and (21), whereas the M0-M2 subtypes are usually indistinguishable (43). By analyzing the AML-M1 and M2 proteomes, we succeeded in determining five proteins with higher accumulation levels in AML M2 (33). For three of those proteins (catalase, Annexin 3 and L-plastin) we had gene-specific probes on the AML-array, but none of the genes were differentially expressed between the M1 and M2 FAB subtypes. However, with the use of ddPCR, one of the most sensitive types of quantitative PCR, we demonstrated that ANXA3 expression, despite being generally low and not detectable in all samples, was definitely higher in M2 compared with M1 as well as HV. These results prove the usefulness of Annexin 3 as a potential biomarker at both the protein and transcript levels. In the case of 4 other proteins, inconsistent results were found both in our data and GEO data, what implicates utility of these markers only at the protein level. This inconsistency may be explained by the following: Gene expression is not exclusively regulated by transcription; instead, the abundance of proteins is predominantly controlled at the translational level (59). The lifetimes of the proteins and corresponding transcripts vary due to different synthesis and degradation rates, moreover, from one transcript more than one protein isoform can be synthesized through alternative splicing and posttranslational modifications. Vogel and Marcotte (60) claimed that only ~40% of the protein concentration variation is determined by mRNA abundance. Weak correlation or a lack of correlation between gene expression in the same cells or tissues has also been demonstrated by (61,62).
The presented RT-qPCR-based analyses enabled the identification of 2 other genes with statistically significant differential expression between the M1 and M2 FAB subtypes: S100A9 and WT1, which were decreased and increased in AML-M1, respectively. WT1 overexpression in AML-M1 compared with -M2 is consistent with the observation that the expression of Wilms’ tumor gene is limited to immature leukemias and inversely correlates with cell differentiation (63). As regards the S100A genes, which encode calcium binding proteins implicated in cell cycle regulation, cell differentiation, growth, inflammation and progression of cancer, the data in the literature are contradictory. The upregulation of S100A8 and S100A9 has been reported in many solid tumor types (64,65) and in the late myeloblast and monoblast leukemic cell lines (56); their downregulation has been reported in childhood AML (E-GEOD-2191). Our data indicated a high variance of S100A gene expression within the AML samples. Similar conclusions can be drawn from the results obtained by (21), in which NK-AML samples clustered into 2 major subgroups; one associated with very high expression of the S100A9 gene and the second associated with very low expression. In agreement with our results, the majority of the AML-M1 samples had low expression levels of S100A9.
Whole-genome analyses of AML have enabled the determination of gene expression signatures specific for AML subgroups harboring the most frequent mutations (24,21). With the use of AML-array, we were able to identify only one gene that exhibited a statistically significant difference in the expression level between AML samples, which was CPA3, encoding mast cell carboxypeptidase A3. This gene, whose expression was 16-fold higher in the FLT3+/NPM1+ samples compared with the remaining AML, has also been identified by (66) as the fourth most upregulated gene in the comparison between FLT3-ITD and FLT3-wt in older AML patients. Our RT-qPCR analyses of the AML samples also revealed significantly increased levels of 4 other genes: STMN1 in FLT3+/NPM1− samples, ABL1 in FLT3+ and FLT3+/NPM1+ AML samples, CAT in FLT3− AML samples and the WT1 gene in FLT3+/NPM1+ AML. The WT1 expression pattern is consistent with the results of the study by Whitman et al (66), who reported WT1 as one of the most upregulated genes in AML with FLT3-ITD compared with wild-type AML. In contrast to the upregulation of CAT in FLT3− AML described herein, other authors have linked the elevated expression of this gene to the presence of FLT3-ITD (22,66). The association of the STMN1 and ABL1 genes with the FLT3 and NPM1 mutations has not yet been discussed.
While analyzing therapy-related AML subgroups, we observed differential expression in only two genes by using ddPCR: CAT and WT1. CAT exhibited the highest expression in the CR-long group, whereas WT1 was gradually increased with the resistance to AML treatment. The results obtained for catalase remain consistent with the described catalase downregulation in the doxorubicin-resistant AML subline (67). High expression of the WT1 gene, the postulated tumor suppressor, has been described in lymphomas and leukemias (68). The correlation of WT1 expression and AML prognosis has also been reported (39,40).
In conclusion, the present study demonstrates that a small AML-dedicated array, designed and generated in our laboratory, despite some limitations, is suitable for preliminary gene expression studies of AML. Although many of the genes identified with the use of AML-array, are confirmed in the literature, we show that some genes encoding basic metabolism proteins (e.g., RPLP0, ENO1, GAPDH, PDE3B, PKM and FCN1), may also contribute to leukemogenesis, though were not yet discussed in this context. The raw data were deposited in publically available repository and can be downloaded and re-analyzed by any researcher. Despite the lack of success in distinguishing between particular AML subgroups, AML-array can be potentially applied at the stage of first diagnosis of AML based on gene expression profile. It can also be modified to elaborate an improved array version with better rate of diagnostic and prognostic success. However, for more accurate transcript measurement, quantitative PCR approaches, in particular ddPCR, are more appropriate. Application of this strategy enabled us to obtain results valuable form the diagnostic and prognostic point of view. We identified three genes (S100A9, ANXA3 and WT1), whose expression levels can be used for discrimination between two AML FAB subtypes, M1 and M2, which are usually difficult to distinguish. We showed for the first time relationship of the STMN1 and ABL1 genes with the FLT3 and NPM1 mutation status, and correlation of high CAT expression with positive response to AML treatment.
Acknowledgments
This study was financially supported by the Polish Ministry of Science and Higher Education (grant no. PBZ_MniI-2/1/2005). This study was also supported by the Polish Ministry of Science and Higher Education under the SPUB (no. 5536/E-63/SPUB/2015) and KNOW programs. We are grateful to Dr Marcin Schmidt from the Poznan University of Life Sciences for providing the HL60 cells. We would also like to thank Dr Joanna Zyprych-Walczak, Dr Alicja Szabelska-Beręsewicz and Professor Idzi Siatkowski from Poznan University of Life Sciences for their assistance with microarray data normalization.
Abbreviations
- AML
acute myeloid leukemia
- CN-AML
cytogenetically normal AML
- ddPCR
droplet digital PCR
- FAB
French-American-British (classification of leukemia)
- PCR
polymerase chain reaction
Footnotes
Competing interests
The authors declare that they have no competing interests.
References
- 1.Estey E, Döhner H. Acute myeloid leukaemia. Lancet. 2006;368:1894–1907. doi: 10.1016/S0140-6736(06)69780-8. [DOI] [PubMed] [Google Scholar]
- 2.Döhner H, Estey EH, Amadori S, Appelbaum FR, Büchner T, Burnett AK, Dombret H, Fenaux P, Grimwade D, Larson RA, et al. European LeukemiaNet Diagnosis and management of acute myeloid leukemia in adults: Recommendations from an international expert panel, on behalf of the European LeukemiaNet. Blood. 2010;115:453–474. doi: 10.1182/blood-2009-07-235358. [DOI] [PubMed] [Google Scholar]
- 3.Bennett JM, Catovsky D, Daniel MT, Flandrin G, Galton DA, Gralnick HR, Sultan C. Proposed revised criteria for the classification of acute myeloid leukemia. A report of the French-American-British Cooperative Group. Ann Intern Med. 1985;103:620–625. doi: 10.7326/0003-4819-103-4-620. [DOI] [PubMed] [Google Scholar]
- 4.Vardiman JW, Thiele J, Arber DA, Brunning RD, Borowitz MJ, Porwit A, Harris NL, Le Beau MM, Hellström-Lindberg E, Tefferi A, et al. The 2008 revision of the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia: Rationale and important changes. Blood. 2009;114:937–951. doi: 10.1182/blood-2009-03-209262. [DOI] [PubMed] [Google Scholar]
- 5.Döhner H. Hematology Am Soc Hematol Educ Program. Washington, DC: 2007. Implication of the molecular characterization of acute myeloid leukemia; pp. 412–419. [DOI] [PubMed] [Google Scholar]
- 6.Grimwade D, Hills RK. Hematology Am Soc Hematol Educ Program. Washington, DC: 2009. Independent prognostic factors for AML outcome; pp. 385–395. [DOI] [PubMed] [Google Scholar]
- 7.Falini B. Acute myeloid leukemia with mutated nucleophosmin (NPM1): Molecular, pathological, and clinical features. Cancer Treat Res. 2010;145:149–168. doi: 10.1007/978-0-387-69259-3_9. [DOI] [PubMed] [Google Scholar]
- 8.Nakao M, Yokota S, Iwai T, Kaneko H, Horiike S, Kashima K, Sonoda Y, Fujimoto T, Misawa S. Internal tandem duplication of the flt3 gene found in acute myeloid leukemia. Leukemia. 1996;10:1911–1918. [PubMed] [Google Scholar]
- 9.Lin LI, Chen CY, Lin DT, Tsay W, Tang JL, Yeh YC, Shen HL, Su FH, Yao M, Huang SY, et al. Characterization of CEBPA mutations in acute myeloid leukemia: Most patients with CEBPA mutations have biallelic mutations and show a distinct immunophenotype of the leukemic cells. Clin Cancer Res. 2005;11:1372–1379. doi: 10.1158/1078-0432.CCR-04-1816. [DOI] [PubMed] [Google Scholar]
- 10.Care RS, Valk PJ, Goodeve AC, Abu-Duhier FM, Geertsma-Kleinekoort WM, Wilson GA, Gari MA, Peake IR, Löwenberg B, Reilly JT. Incidence and prognosis of c-KIT and FLT3 mutations in core binding factor (CBF) acute myeloid leukaemias. Br J Haematol. 2003;121:775–777. doi: 10.1046/j.1365-2141.2003.04362.x. [DOI] [PubMed] [Google Scholar]
- 11.Tyner JW, Erickson H, Deininger MWN, Willis SG, Eide CA, Levine RL, Heinrich MC, Gattermann N, Gilliland DG, Druker BJ, et al. High-throughput sequencing screen reveals novel, transforming RAS mutations in myeloid leukemia patients. Blood. 2009;113:1749–1755. doi: 10.1182/blood-2008-04-152157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Delhommeau F, Dupont S, Della Valle V, James C, Trannoy S, Massé A, Kosmider O, Le Couedic JP, Robert F, Alberdi A, et al. Mutation in TET2 in myeloid cancers. N Engl J Med. 2009;360:2289–2301. doi: 10.1056/NEJMoa0810069. [DOI] [PubMed] [Google Scholar]
- 13.Ley TJ, Ding L, Walter MJ, McLellan MD, Lamprecht T, Larson DE, Kandoth C, Payton JE, Baty J, Welch J, et al. DNMT3A mutations in acute myeloid leukemia. N Engl J Med. 2010;363:2424–2433. doi: 10.1056/NEJMoa1005143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Aref S, Kamel Areida S, Abdel Aaal MF, Adam OM, El-Ghonemy MS, El-Baiomy MA, Zeid TA. Prevalence and clinical effect of IDH1 and IDH2 mutations among cytogenetically normal acute myeloid leukemia patients. Clin Lymphoma Myeloma Leuk. 2015;15:550–555. doi: 10.1016/j.clml.2015.05.009. [DOI] [PubMed] [Google Scholar]
- 15.Mrózek K, Marcucci G, Paschka P, Whitman SP, Bloomfield CD. Clinical relevance of mutations and gene-expression changes in adult acute myeloid leukemia with normal cytogenetics: Are we ready for a prognostically prioritized molecular classification? Blood. 2007;109:431–448. doi: 10.1182/blood-2006-06-001149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Bacher U, Kohlmann A, Haferlach T. Gene expression profiling for diagnosis and therapy in acute leukaemia and other haematologic malignancies. Cancer Treat Rev. 2010;36:637–646. doi: 10.1016/j.ctrv.2010.05.002. [DOI] [PubMed] [Google Scholar]
- 17.Riva L, Luzi L, Pelicci PG. Genomics of acute myeloid leukemia: The next generation. Front Oncol. 2012;2:40. doi: 10.3389/fonc.2012.00040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kohlmann A, Grossmann V, Nadarajah N, Haferlach T. Next-generation sequencing - feasibility and practicality in haematology. Br J Haematol. 2013;160:736–753. doi: 10.1111/bjh.12194. [DOI] [PubMed] [Google Scholar]
- 19.Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP, Coller H, Loh ML, Downing JR, Caligiuri MA, et al. Molecular classification of cancer: Class discovery and class prediction by gene expression monitoring. Science. 1999;286:531–537. doi: 10.1126/science.286.5439.531. [DOI] [PubMed] [Google Scholar]
- 20.Alizadeh A, Eisen M, Davis RE, Ma C, Sabet H, Tran T, Powell JI, Yang L, Marti GE, Moore DT, et al. The lymphochip: A specialized cDNA microarray for the genomic-scale analysis of gene expression in normal and malignant lymphocytes. Cold Spring Harb Symp Quant Biol. 1999;64:71–78. doi: 10.1101/sqb.1999.64.71. [DOI] [PubMed] [Google Scholar]
- 21.Bullinger L, Döhner K, Bair E, Fröhling S, Schlenk RF, Tibshirani R, Döhner H, Pollack JR. Use of gene-expression profiling to identify prognostic subclasses in adult acute myeloid leukemia. N Engl J Med. 2004;350:1605–1616. doi: 10.1056/NEJMoa031046. [DOI] [PubMed] [Google Scholar]
- 22.Valk PJ, Verhaak RG, Beijen MA, Erpelinck CAJ, Barjesteh van Waalwijk van Doorn-Khosrovani S, Boer JM, Beverloo HB, Moorhouse MJ, van der Spek PJ, Löwenberg B, et al. Prognostically useful gene-expression profiles in acute myeloid leukemia. N Engl J Med. 2004;350:1617–1628. doi: 10.1056/NEJMoa040465. [DOI] [PubMed] [Google Scholar]
- 23.Bacher U, Kohlmann A, Haferlach C, Haferlach T. Gene expression profiling in acute myeloid leukaemia (AML) Best Pract Res Clin Haematol. 2009;22:169–180. doi: 10.1016/j.beha.2009.04.003. [DOI] [PubMed] [Google Scholar]
- 24.Alcalay M, Tiacci E, Bergomas R, Bigerna B, Venturini E, Minardi SP, Meani N, Diverio D, Bernard L, Tizzoni L, et al. Acute myeloid leukemia bearing cytoplasmic nucleophosmin (NPMc+ AML) shows a distinct gene expression profile characterized by up-regulation of genes involved in stem-cell maintenance. Blood. 2005;106:899–902. doi: 10.1182/blood-2005-02-0560. [DOI] [PubMed] [Google Scholar]
- 25.Huang L, Zhou K, Yang Y, Shang Z, Wang J, Wang D, Wang N, Xu D, Zhou J. FLT3-ITD-associated gene-expression signatures in NPM1-mutated cytogenetically normal acute myeloid leukemia. Int J Hematol. 2012;96:234–240. doi: 10.1007/s12185-012-1115-9. [DOI] [PubMed] [Google Scholar]
- 26.Radmacher MD, Marcucci G, Ruppert AS, Mrózek K, Whitman SP, Vardiman JW, Paschka P, Vukosavljevic T, Baldus CD, Kolitz JE, et al. Cancer and Leukemia Group B Independent confirmation of a prognostic gene-expression signature in adult acute myeloid leukemia with a normal karyotype: A Cancer and Leukemia Group B study. Blood. 2006;108:1677–1683. doi: 10.1182/blood-2006-02-005538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Wilson CS, Davidson GS, Martin SB, Andries E, Potter J, Harvey R, Ar K, Xu Y, Kopecky KJ, Ankerst DP, et al. Gene expression profiling of adult acute myeloid leukemia identifies novel biologic clusters for risk classification and outcome prediction. Blood. 2006;108:685–696. doi: 10.1182/blood-2004-12-4633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Marcinkowska-Swojak M, Handschuh L, Wojciechowski P, Goralski M, Tomaszewski K, Kazmierczak M, Lewandowski K, Komarnicki M, Blazewicz J, Figlerowicz M, et al. Simultaneous detection of mutations and copy number variation of NPM1 in the acute myeloid leukemia using multiplex ligation-dependent probe amplification. Mutat Res. 2016;786:14–26. doi: 10.1016/j.mrfmmm.2016.02.001. [DOI] [PubMed] [Google Scholar]
- 29.Oshlack A, Emslie D, Corcoran LM, Smyth GK. Normalization of boutique two-color microarrays with a high proportion of differentially expressed probes. Genome Biol. 2007;8:R2. doi: 10.1186/gb-2007-8-1-r2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Uszczyńska B, Zyprych-Walczak J, Handschuh L, Szabelska A, Kaźmierczak M, Woronowicz W, Kozłowski P, Sikorski MM, Komarnicki M, Siatkowski I, et al. Analysis of boutique arrays: A universal method for the selection of the optimal data normalization procedure. Int J Mol Med. 2013;32:668–684. doi: 10.3892/ijmm.2013.1443. [DOI] [PubMed] [Google Scholar]
- 31.Wenne R, Handschuh L, Pocwierz-Kotus A, Urbaniak R, Formanowicz P, Całkiewicz J, Brzozowska K, Figlerowicz M, Węgrzyn G, Wróbel B. The application of microarray technology to the identification of Tc1-like element sequences in fish genomes. Mar Biol Res. 2011;7:466–477. doi: 10.1080/17451000.2010.527990. [DOI] [Google Scholar]
- 32.Zmieńko A, Guzowska-Nowowiejska M, Urbaniak R, Pląder W, Formanowicz P, Figlerowicz M. A tiling microarray for global analysis of chloroplast genome expression in cucumber and other plants. Plant Methods. 2011;7:29. doi: 10.1186/1746-4811-7-29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Luczak M, Kaźmierczak M, Handschuh L, Lewandowski K, Komarnicki M, Figlerowicz M. Comparative proteome analysis of acute myeloid leukemia with and without maturation. J Proteomics. 2012;75:5734–5748. doi: 10.1016/j.jprot.2012.07.030. [DOI] [PubMed] [Google Scholar]
- 34.Kaźmierczak M, Luczak M, Lewandowski K, Handschuh L, Czyż A, Jarmuż M, Gniot M, Michalak M, Figlerowicz M, Komarnicki M. Esterase D and gamma 1 actin level might predict results of induction therapy in patients with acute myeloid leukemia without and with maturation. Med Oncol. 2013;30:725. doi: 10.1007/s12032-013-0725-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Ley TJ, Miller C, Ding L, Raphael BJ, Mungall AJ, Robertson A, Hoadley K, Triche TJ, Jr, Laird PW, Baty JD, et al. Cancer Genome Atlas Research Network Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med. 2013;368:2059–2074. doi: 10.1056/NEJMoa1301689. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Smyth GK. Limma: linear models for microarray data. In: Gentleman R, Carey V, Dudoit S, Irizarry R, Huber W, editors. Bioinformatics and Computational Biology Solutions using R and Bioconductor. Springer; New York, NY: 2005. pp. 397–420. [DOI] [Google Scholar]
- 37.Benjamini Y, Hochberg Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J R Stat Soc B. 1995;57:289–300. [Google Scholar]
- 38.Huang W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57. doi: 10.1038/nprot.2008.211. [DOI] [PubMed] [Google Scholar]
- 39.Bergmann L, Miething C, Maurer U, Brieger J, Karakas T, Weidmann E, Hoelzer D. High levels of Wilms’ tumor gene (wt1) mRNA in acute myeloid leukemias are associated with a worse long-term outcome. Blood. 1997;90:1217–1225. [PubMed] [Google Scholar]
- 40.Lyu X, Xin Y, Mi R, Ding J, Wang X, Hu J, Fan R, Wei X, Song Y, Zhao RY. Overexpression of Wilms tumor 1 gene as a negative prognostic indicator in acute myeloid leukemia. PLoS One. 2014;9:e92470. doi: 10.1371/journal.pone.0092470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Wouters BJ, Löwenberg B, Delwel R. A decade of genome-wide gene expression profiling in acute myeloid leukemia: Flashback and prospects. Blood. 2009;113:291–298. doi: 10.1182/blood-2008-04-153239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Haferlach T, Kohlmann A, Schnittger S, Dugas M, Hiddemann W, Kern W, Schoch C. Global approach to the diagnosis of leukemia using gene expression profiling. Blood. 2005;106:1189–1198. doi: 10.1182/blood-2004-12-4938. [DOI] [PubMed] [Google Scholar]
- 43.Payton JE, Grieselhuber NR, Chang L-W, Murakami M, Geiss GK, Link DC, Nagarajan R, Watson MA, Ley TJ. High throughput digital quantification of mRNA abundance in primary human acute myeloid leukemia samples. J Clin Invest. 2009;119:1714–1726. doi: 10.1172/JCI38248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Gutiérrez NC, López-Pérez R, Hernández JM, Isidro I, González B, Delgado M, Fermiñán E, García JL, Vázquez L, González M, et al. Gene expression profile reveals deregulation of genes with relevant functions in the different subclasses of acute myeloid leukemia. Leukemia. 2005;19:402–409. doi: 10.1038/sj.leu.2403625. [DOI] [PubMed] [Google Scholar]
- 45.Andersson A, Edén P, Lindgren D, Nilsson J, Lassen C, Heldrup J, Fontes M, Borg A, Mitelman F, Johansson B, et al. Gene expression profiling of leukemic cell lines reveals conserved molecular signatures among subtypes with specific genetic aberrations. Leukemia. 2005;19:1042–1050. doi: 10.1038/sj.leu.2403749. [DOI] [PubMed] [Google Scholar]
- 46.Park MH, Cho SA, Yoo KH, Yang MH, Ahn JY, Lee HS, Lee KE, Mun YC, Cho DH, Seong CM, et al. Gene expression profile related to prognosis of acute myeloid leukemia. Oncol Rep. 2007;18:1395–1402. [PubMed] [Google Scholar]
- 47.Tominaga-Sato S, Tsushima H, Ando K, Itonaga H, Imaizumi Y, Imanishi D, Iwanaga M, Taguchi J, Fukushima T, Yoshida S, et al. Expression of myeloperoxidase and gene mutations in AML patients with normal karyotype: Double CEBPA mutations are associated with high percentage of MPO positivity in leukemic blasts. Int J Hematol. 2011;94:81–89. doi: 10.1007/s12185-011-0883-y. [DOI] [PubMed] [Google Scholar]
- 48.Cristóbal I, Garcia-Orti L, Cirauqui C, Cortes-Lavaud X, García-Sánchez MA, Calasanz MJ, Odero MD. Overexpression of SET is a recurrent event associated with poor outcome and contributes to protein phosphatase 2A inhibition in acute myeloid leukemia. Haematologica. 2012;97:543–550. doi: 10.3324/haematol.2011.050542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Artero-Castro A, Castellvi J, García A, Hernández J, Ramón y Cajal S, Lleonart ME. Expression of the ribosomal proteins Rplp0, Rplp1, and Rplp2 in gynecologic tumors. Hum Pathol. 2011;42:194–203. doi: 10.1016/j.humpath.2010.04.020. [DOI] [PubMed] [Google Scholar]
- 50.Tsai ST, Chien IH, Shen WH, Kuo YZ, Jin YT, Wong TY, Hsiao JR, Wang HP, Shih NY, Wu LW. ENO1, a potential prognostic head and neck cancer marker, promotes transformation partly via chemokine CCL20 induction. Eur J Cancer. 2010;46:1712–1723. doi: 10.1016/j.ejca.2010.03.018. [DOI] [PubMed] [Google Scholar]
- 51.Rowlands DC, Williams A, Jones NA, Guest SS, Reynolds GM, Barber PC, Brown G. Stathmin expression is a feature of proliferating cells of most, if not all, cell lineages. Lab Invest. 1995;72:100–113. [PubMed] [Google Scholar]
- 52.Roos G, Brattsand G, Landberg G, Marklund U, Gullberg M. Expression of oncoprotein 18 in human leukemias and lymphomas. Leukemia. 1993;7:1538–1546. [PubMed] [Google Scholar]
- 53.Johnson WE, Jones NA, Rowlands DC, Williams A, Guest SS, Brown G. Down-regulation but not phosphorylation of stathmin is associated with induction of HL60 cell growth arrest and differentiation by physiological agents. FEBS Lett. 1995;364:309–313. doi: 10.1016/0014-5793(95)00416-7. [DOI] [PubMed] [Google Scholar]
- 54.Matushansky I, Radparvar F, Skoultchi AI. Reprogramming leukemic cells to terminal differentiation by inhibiting specific cyclin-dependent kinases in G1. Proc Natl Acad Sci USA. 2000;97:14317–14322. doi: 10.1073/pnas.250488697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Müller A, Lange K, Gaiser T, Hofmann M, Bartels H, Feller AC, Merz H. Expression of angiopoietin-1 and its receptor TEK in hematopoietic cells from patients with myeloid leukemia. Leuk Res. 2002;26:163–168. doi: 10.1016/S0145-2126(01)00110-2. [DOI] [PubMed] [Google Scholar]
- 56.Tagliafico E, Tenedini E, Manfredini R, Grande A, Ferrari F, Roncaglia E, Bicciato S, Zini R, Salati S, Bianchi E, et al. Identification of a molecular signature predictive of sensitivity to differentiation induction in acute myeloid leukemia. Leukemia. 2006;20:1751–1758. doi: 10.1038/sj.leu.2404358. [DOI] [PubMed] [Google Scholar]
- 57.Qian Z, Fernald AA, Godley LA, Larson RA, Le Beau MM. Expression profiling of CD34+ hematopoietic stem/progenitor cells reveals distinct subtypes of therapy-related acute myeloid leukemia. Proc Natl Acad Sci USA. 2002;99:14925–14930. doi: 10.1073/pnas.222491799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Shimamura R, Kudo J, Kondo H, Dohmen K, Gondo H, Okamura S, Ishibashi H, Niho Y. Expression of the thymosin beta 4 gene during differentiation of hematopoietic cells. Blood. 1990;76:977–984. [PubMed] [Google Scholar]
- 59.Schwanhäusser B, Wolf J, Selbach M, Busse D. Synthesis and degradation jointly determine the responsiveness of the cellular proteome. BioEssays. 2013;35:597–601. doi: 10.1002/bies.201300017. [DOI] [PubMed] [Google Scholar]
- 60.Vogel C, Marcotte EM. Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat Rev Genet. 2012;13:227–232. doi: 10.1038/nrg3185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Chen G, Gharib TG, Huang CC, Taylor JM, Misek DE, Kardia SL, Giordano TJ, Iannettoni MD, Orringer MB, Hanash SM, et al. Discordant protein and mRNA expression in lung adenocarcinomas. Mol Cell Proteomics. 2002;1:304–313. doi: 10.1074/mcp.M200008-MCP200. [DOI] [PubMed] [Google Scholar]
- 62.Jansen R, Greenbaum D, Gerstein M. Relating whole-genome expression data with protein-protein interactions. Genome Res. 2002;12:37–46. doi: 10.1101/gr.205602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Miwa H, Beran M, Saunders GF. Expression of the Wilms’ tumor gene (WT1) in human leukemias. Leukemia. 1992;6:405–409. [PubMed] [Google Scholar]
- 64.El-Rifai W, Moskaluk CA, Abdrabbo MK, Harper J, Yoshida C, Riggins GJ, Frierson HF, Jr, Powell SM. Gastric cancers overexpress S100A calcium-binding proteins. Cancer Res. 2002;62:6823–6826. [PubMed] [Google Scholar]
- 65.Su YJ, Xu F, Yu JP, Yue DS, Ren XB, Wang CL. Up-regulation of the expression of S100A8 and S100A9 in lung adenocarcinoma and its correlation with inflammation and other clinical features. Chin Med J (Engl) 2010;123:2215–2220. [PubMed] [Google Scholar]
- 66.Whitman SP, Maharry K, Radmacher MD, Becker H, Mrózek K, Margeson D, Holland KB, Wu YZ, Schwind S, Metzeler KH, et al. FLT3 internal tandem duplication associates with adverse outcome and gene- and microRNA-expression signatures in patients 60 years of age or older with primary cytogenetically normal acute myeloid leukemia: A Cancer and Leukemia Group B study. Blood. 2010;116:3622–3626. doi: 10.1182/blood-2010-05-283648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Kim HS, Lee TB, Choi CH. Down-regulation of catalase gene expression in the doxorubicin-resistant AML subline AML-2/DX100. Biochem Biophys Res Commun. 2001;281:109–114. doi: 10.1006/bbrc.2001.4324. [DOI] [PubMed] [Google Scholar]
- 68.Bergmann L, Maurer U, Weidmann E. Wilms tumor gene expression in acute myeloid leukemias. Leuk Lymphoma. 1997;25:435–443. doi: 10.3109/10428199709039030. [DOI] [PubMed] [Google Scholar]
- 69.Lossos IS, Czerwinski DK, Wechser MA, Levy R. Optimization of quantitative real-time RT-PCR parameters for the study of lymphoid malignancies. Leukemia. 2003;17:789–795. doi: 10.1038/sj.leu.2402880. [DOI] [PubMed] [Google Scholar]
- 70.Hussein S, Michael P, Brabant D, Omri A, Narain R, Passi K, Ramana CV, Parrillo JE, Kumar A, Parissenti A, et al. Characterization of human septic sera induced gene expression modulation in human myocytes. Int J Clin Exp Med. 2009;2:131–148. [PMC free article] [PubMed] [Google Scholar]
- 71.Kreuzer KA, Saborowski A, Lupberger J, Appelt C, Na IK, le Coutre P, Schmidt CA. Fluorescent 5′-exonuclease assay for the absolute quantification of Wilms’ tumour gene (WT1) mRNA: Implications for monitoring human leukaemias. Br J Haematol. 2001;114:313–318. doi: 10.1046/j.1365-2141.2001.02912.x. [DOI] [PubMed] [Google Scholar]