Abstract
Renal repair after injury is dependent on clonal expansion of proliferation‐competent cells. In the human kidney, the expression of CD133 characterizes a population of resident scattered cells with resistance to damage and ability to proliferate. However, the biological function of the CD133 molecule is unknown. By RNA sequencing, we found that cells undergoing cisplatin damage lost the CD133 signature and acquired metanephric mesenchymal and regenerative genes such as SNAIL1, KLF4, SOX9, and WNT3. CD133 was reacquired in the recovery phase. In CD133‐Kd cells, lack of CD133 limited cell proliferation after injury and was specifically correlated with deregulation of Wnt signaling and E‐cadherin pathway. By immunoprecipitation, CD133 appeared to form a complex with E‐cadherin and β‐catenin. In parallel, CD133‐Kd cells showed lower β‐catenin levels in basal condition and after Wnt pathway activation and reduced TCF/LEF promoter activation in respect to CD133+ cells. Finally, the lack of CD133 impaired generation of nephrospheres while favoring senescence. These data indicate that CD133 may act as a permissive factor for β‐catenin signaling, preventing its degradation in the cytoplasm. Therefore, CD133 itself appears to play a functional role in renal tubular repair through maintenance of proliferative response and control of senescence. Stem Cells Translational Medicine 2018;7:283–294
Keywords: Regeneration, Acute kidney injury, Progenitor cells, Senescence, Wnt signaling pathway
Significance Statement.
In the human kidney, the expression of CD133 characterizes a population of resident scattered cells with resistance to damage and ability to proliferate. However, the biological function of the CD133 molecule is unknown. It was found that the CD133 molecule, expressed on human tubular cells, may act as a permissive factor for Wnt/beta‐catenin signaling, regulating cell proliferative response after damage, together with limiting cell senescence. These data may contribute to the understanding of the mechanisms involved in renal tissue repair.
Introduction
Acute kidney injury, defined as an abrupt loss of proper kidney function, currently affects up to 7% of hospitalized patients, with those treated in intensive care facing the highest risk 1. An increasing amount of evidence indicates that, in spite of an apparent functional repair, renal tissue is permanently damaged after injury and does not completely return to its original state. This may favor the development of fibrosis and the susceptibility to progression toward chronic kidney disease 2, 3.
Studies aimed to elucidate the process of renal tissue repair following an acute damage identified that it is mediated by proliferation of resident tubular cells 4, 5. The nature of the regenerating cells has been attributed, in mice, to both de‐differentiation of tubular mature cells 6 and/or activation of resident cells with progenitor characteristics 7. Indeed, genetic studies in mice recently showed that renal repair is dependent on clonal expansion of segment restricted unipotent cells, identified as proliferation‐competent cells 8. In mice, markers such as Sox‐9, vimentin, and CD24 have been used to label scattered cells along the nephron 9, 10, 11. In the human kidney, CD24+/CD133+ cells within different nephron segments appear as a population of resident renal progenitor cells (RPCs) with the ability of expansion, self‐renewal, and epithelial differentiation both in vitro and in vivo 11, 12, 13, 14. Indeed, the CD133+ population appears to be the proliferating one in human biopsies of patients undergoing renal insults, further implying its role in renal repair 14, 15.
The CD133 molecule is a pentaspan‐trans‐membrane glycoprotein 16, 17, recognized by a specific glycosylation dependent epitope using the AC133 Ab. It has been found to be expressed by immature cells such as hematopoietic stem/progenitors, tissue‐specific progenitors, and developing intestinal and neuronal cells 18, 19, 20. In cultured CD133+ renal cells, CD133 has been shown to be upregulated by hypoxia 21. However, the molecule's biological function along with its possible modulation during kidney damage is currently unknown.
In the present study, we aimed to evaluate the biological function of CD133 expressed by tubular cells during damage, and its possible implication in the repairing process. Our data showed that CD133 is involved in the early response to Wnt signaling, through regulation of β‐catenin levels. Moreover, CD133 favored cell proliferation in the regenerative phase and limited cell senescence.
Materials and Methods
Isolation of Human RPCs
CD133+ RPCs were obtained from biopsies of normal tissue of a human kidney surgically removed for polar carcinoma performed (after the approval of ethical committee for the use of human tissue of Molinette Hospital; n. 168/2004), as previously described 13. In particular, the outer medullary tissue at the opposite pole of the tumor was used and absence of tumor infiltration was evaluated by a pathologist. Tissue samples were cut to obtain 3–5 mm3 fragments, digested in 0.1% collagenase type I (Sigma‐Aldrich, St Louis, MO) for 30 minutes at 37°C and subsequently forced through a graded series of meshes for the separation of cell components from stroma and aggregates. The filtrate was then pelleted by centrifugation. To recover CD133+ cells, the single cell suspension underwent magnetic separation for CD133 (Mylteni, Biotec, Bergisch Gladbach, Germany, CD133 Cell Isolation Kit, containing the anti‐CD133/1 mAb, clone AC133). CD133+ cells (>98% as evaluated by cytofluorimetric analysis) were resuspended in expansion medium (Endothelial Basal Medium plus supplement kit; CambrexBioScience, East Rutherford, NJ, USA) and plated at density 1 × 104 viable cells per cm2. CD133 expression in the 12 cell lines in study tested before experiments of transfection and RNA sequencing was 90.5% ± 8.8%. Cells were used until passage six and maintained their phenotype during the passages.
Generation of CD133 Knockdown (CD133‐Kd) RPCs
For CD133 knockdown pGIPZlentiviral Vectors (Open Biosystems, Dharmacon, Lafayette, CO, USA) carrying two different shRNA against CD133 (shPROM1 and shPROM2), or a scrambled sequence (GFP) were used.
shPROM1: oligo ID: V2LHS_71819 sequence: GAGTCCTTCCTATAGAACA
shPROM2: oligo ID: V2LHS_71820 sequence: CTGTTGGTGATTTGTATAA
The cell line 293T was transfected with constructs using ViraPower Packaging Mix (Life Technologies, ThermoFischer, Waltham, MA, USA) for lentiviruses production according to manufacturer's instructions. After titering the lentiviral stock, CD133+ RPCs, isolated from 12 different nephrectomy specimens, were infected with lentiviral particles following the manufacturer's instructions. Cells were selected by puromycin (Gibco, ThermoFischer) (1 μg/ml) and, after 6 days, antibiotic‐resistant cells were expanded. Cell infection was considered effective when the percentage of GFP+ cells was >95%, and the reduction in CD133 protein expression was >70%, as assessed by FACS and Western blot analysis.
RNA Sequencing
For RNA sequencing library preparation, approximately 3 μg of total RNA isolated from CD133+ (GFP) and CD133‐Kd RPCs (n = 3), all at passage #4, were subjected to poly(A) selection, and libraries were prepared using the TruSeq RNA Sample Prep Kit (Illumina, San Diego, CA) following the manufacturer's instructions. Sequencing was performed on the Illumina HiScan SQ platform. Reads were mapped to the Homo sapiens hg19 reference assembly using TopHat v2.0.10,46 and RPKMs (reads per kilobase per million mapped reads) for transcripts were calculated with Cufflinks v2.1.1.47. Transcripts with RPKM ≥1 were considered expressed and further investigated for differential expression analysis between cisplatin treated (CIS GFP) or untreated cells (GFP). Genes with log2(CIS GFP/GFP) ≥1 or ≤–1 were considered modulated by cisplatin treatment. Modulated genes were subsequently analyzed in shPROM1/2 cells, at passage #4. Genes differentially modulated by cisplatin treatment in shPROM1/2 cells versus GFP cells (Fold change: log2(CIS shPROM1/2/CIS GFP) ≤–1 or ≥1) were defined as upregulated or downregulated by CD133‐Kd in cisplatin treated cells. Functional Enrichment analysis tool (FunRich, http://www.funrich.org/) 22 and PANTHER software (http://pantherdb.org/) were used for pathways enrichment analysis. Gene Ontology analysis for biological processes was conducted using BINGO plug‐in from Cytoscape software 23.
Real Time PCR
Total RNA was isolated from different cell preparations using Trizol reagent (Ambion, ThermoFischer) according to manufacturer's protocol. RNA was quantified spectrophotometrically (Nanodrop ND‐1000, Wilmington, DE). First‐strand cDNA was produced from 200 ng of total RNA using the High Capacity cDNA Reverse Transcription Kit (Applied Biosystems, ThermoFischer). For gene expression analysis, quantitative real‐time PCR (qRT‐PCR) was performed in 20‐µl reaction mixtures containing 5 ng of cDNA template, the sequence‐specific oligonucleotide primers (purchased from MWG‐Biotech, Eurofins Scientific, Bruxelles, Belgium) and the Power SYBR Green PCR Master Mix (Applied Biosystems). GAPDH mRNA was used as housekeeping normalizer. Fold change expression respect to control was calculated for all samples. The sequence‐specific oligonucleotide primers used are shown in the Supporting Information Table S1.
Protein Extraction and Western Blot
For protein analysis, cells were lysed at 4°C for 30 minutes in RIPA buffer (20 nM Tris·HCl, 150 nM NaCl, 1% deoxycholate, 0.1% SDS, 1% Triton X‐100, pH 7.8) supplemented with protease and phosphatase inhibitors cocktail and PMSF (Sigma‐Aldrich). Aliquots of cell lysates containing 20–30 µg of proteins, as evaluated by Bradford, run on 4%–20% Mini‐Protean TGX Stain–Free Gels (BIO‐RAD, Hercules, CA, USA) under reducing conditions and blotted onto PVDF membrane filters using the iBLOT system (Life Technologies). For Western blot analysis, anti‐CD133 (AC133 epitope) (Miltenyi Biotech), anti‐actin (Santa Cruz Biotechnology, Dallas, TX, USA), anti‐β‐catenin (R&D Systems, Minneapolis, MN, USA), anti‐E‐cadherin (BD Biosciences, Franklin Lakes, NJ, USA) and anti GSK3 alpha and beta (Abcam, Cambridge, UK) Abs were used. Data are expressed as mean ± SD of band intensity normalized to actin of six independent experiments.
Cytofluorimetric Analysis
For cytofluorimetric analysis, CD133+ and CD133‐Kd RPCs were detached from plates using the non‐enzymatic cell dissociation solution (Sigma‐Aldrich), and stained (30 minutes at 4°C) with allophycocyanin (APC)‐conjugated CD133 antibody (AC133 epitope) (MiltenyiBiotec). Isotype (MiltenyiBiotec) was used as negative control. Cells were then subjected to cytofluorimetric analysis (FACScan, Becton Dickinson), and the geometric mean of CD133‐APC fluorescence intensity was measured.
Cisplatin Induced Damage
Toxic damage induced by cell treatment with 5 μg/ml of cisplatin. Cisplatin (Vinci Biochem, Vinci, FI, Italy) was dissolved in DMSO, suspended in 0.9% NaCl to a final concentration of 1 mg/ml and stored at 4°C following manufacturer's instructions. Cells were treated with cisplatin in RPMI plus 2% Fetal Calf Serum for the evaluation of the early phases of damage. During cell recovery cultured medium was changed every 2 days until day 8 (recovery phase).
Wnt/β‐Catenin Signaling Stimulation
The Wnt signaling pathway was stimulated using a Wnt activator CHIR99021 (Sigma‐Aldrich), dissolved in DMSO up to a final concentration of 3 mM and stored at −20°C according to manufacturer's instructions. For Wnt signaling activation, CD133+ and CD133‐Kd RPCs were treated with 6 μM of CHIR99021 in RPMI 2% Fetal Calf Serum for 1, 3, or 6 hours. Data are normalized to CD133+ (GFP) not treated cells (CTL) and to one and expressed as mean ± SD of at least three different experiments.
Luciferase Reporter Assay
For the evaluation of Wnt/β‐catenin signaling, 8 × 103 cells per well were seeded in a 96‐well plate. The day after, cells were transfected in serum and antibiotic free DMEM with construct carrying the TCF/LEF promoter for β‐catenin. 48 hours later, cells were lysed with GloLysis Buffer (Promega, Fitchburg, WI, USA) for 15 minutes. Luciferin was then added and the luminescent signal was measured according to manufacturer's instructions. Data are expressed as mean ± SD of at least three independent experiments performed in triplicates normalized to GFP.
Immunoprecipitation
For protein immunoprecipitation 10 μg of anti‐E‐cadherin Ab (BD Biosciences) were added to 1 mg of Magnetic Beads (SureBeadsMegnatic Beads, BIO‐RAD) and the mix was left rotating at room temperature for 1 hour according to manufacturer's protocol. After three washes with PBS‐Tween 0.1% for the removal of unbound antibodies, 1 mg of CD133+ cell lysates was added to Beads‐Ab complex and left for 1 hour rotating at room temperature. For the elution, the new complex formed (Beads‐Ab‐Proteins), was suspended in 100 μl of Laemmli buffer (BIO‐RAD) with 10% β‐mercaptoethanol and incubated for 10 minutes at 70°C. Beads were then magnetically removed and 30 μl of the eluted complex were then used for protein analysis by Western blot.
Cell Proliferation
Cell proliferation was assessed by the incorporation of 5‐bromo‐2‐deoxyuridine (BrdU) into cellular DNA after 24 hour culture in expansion medium, according to the manufacturer's instructions (Roche Applied Science, Mannheim, Germany). Optical density was measured with an ELISA reader (BioRad) at 415 nm (reference 490 nm). Experiments were performed in triplicates; data are expressed as mean ± SD of the media of absorbance of at least three independent experiments, normalized to CD133+ cisplatin‐treated cells.
Spheroid Formation
Renal CD133+ and CD133‐Kd cells were plated in 6‐well culture plates at a density of 104 cells per milliliter in non‐adherent medium consisting of serum free DMEM F12 supplemented with 10 ng/ml basic fibroblast growth factor, 20 ng/ml epidermal growth factor, 5 µg/ml insulin and 0.4% bovine serum albumin (Sigma‐Aldrich), as previously described 24 and maintained in hypoxia chambers (1% O2 and 5% CO2) (Stem Cells Technologies, Vancouver, Canada) for 72 hours. Cells were then observed with a Nikon inverted microscope. For each experimental condition, at least 30 images with ×4 magnification for spheroid number evaluation and 10 images with ×20 magnification for spheroid area measurement were taken. For the assessment of spheroid forming capacity, images were analyzed with imageJ software. The number and area of spheroids were evaluated in each ×4 and ×20 image, respectively.
MTT Assay
For the evaluation of sphere vitality MTT (3‐(4,5‐dimethylthiazol‐2‐yl)‐2,5‐diphenyltetrazolium bromide) assay was performed. Briefly, spheroids deriving from 100,000 cells after 72 hours of culture in hypoxic conditions were centrifuged and resuspended in 300 μl of PBS solution for efficient disaggregation. Spheroid‐derived cells were then plated in 96‐well plates in triplicates and analyzed according to the manufacturer's instructions (Merck‐Millipore). Data are expressed as mean ± SD of the media absorbance of three different experiments performed in triplicates and normalized to GFP.
Evaluation of Senescence
Cells were infected and cultured for five passages, up to a total of 10 cell divisions of the starting population. GFP, shPROM1, shPROM2 cells were then plated in 6‐well culture plates at density of 1 × 104 cell per cm2 for β‐galactosidase assay (β‐gal staining kit, Invitrogen, Carlsbad, CA, USA), or in 58 cm2 dishes for DNA extraction and telomere length evaluation. Cells were washed with PBS, fixed and incubated with β‐galactosidase substrate staining solution for 16 hours at 37°C, according to the manufacturer's instructions. Cells were then observed with a Nikon inverted microscope and for each experimental condition at least 20 images were taken with ×10 magnification. The number of β‐galactosidase positive cells was then evaluated for each image.
For the evaluation of telomere length, genomic DNA was isolated from GFP and CD133‐Kd cells of five different lines with a DNeasy Blood & Tissue Kit (Qiagen, Venlo, The Netherlands) and quantified spectrophotometrically with Nanodrop (Nanodrop ND‐1000, Wilmington, DE). Relative telomere length was determined using an optimized assay 25, 26 originally described by Cawthon 27. Separate PCR experiments were performed for telomere (TEL) and 36B4, a single‐copy gene (S), in 96‐well optical reaction plates (Applied Biosystems). Briefly, 100 ng of DNA sample were transferred in triplicate to identical positions for TEL as well as for 36B4 experiments, together with 10 μl of Power SYBR Green PCR master mix (Applied Biosystems) and the sequence specific oligonucleotide primers. The 36B4 experiments were performed using forward and reverse primers 300 nM with a thermal cycling profile started with 10 minutes at 95°C to activate polymerase for the PCR. Repeating cycles were performed 30 times at 95°C for 15 seconds followed by 56°C for 20 seconds and 72°C for 20 seconds. The TEL experiments were performed using tel1 270 nM and tel2 900 nM with a thermal cycling profile started with 10 minutes at 95°C to activate polymerase for the PCR. Repeated cycles were performed 30 times at 95°C for 15 seconds followed by 54°C for 2 minutes. The T/S ratio was calculated as described 27, setting the control sample equal to 1, while the changes in telomere length (amount of telomeric DNA) were normalized to control. The primer pair sequences for telomere PCR are reported in Supporting Information Table S1.
Statistical Analysis
All data are expressed as mean ± SD. Two‐tailed Student's t test was used for comparison between two groups. One‐way analysis of variance was used for comparison of three or more groups. All statistical analyses were done with GraphPad Prism software version 7.0 (GraphPad Software, Inc.). p values of < .05 were considered significant.
Data Availability
FastQ files for RNA‐seq experiments are deposited on the Gene Expression Omnibus database, under the accession code GSE107273.
Results
Characterization of Adult Human RPCs
CD133 has been widely used as a marker for the isolation of renal human cells with the phenotype of undifferentiated progenitors and the ability to proliferate after damage 13, 14. In the current study, we aimed to elucidate the function of CD133 in renal tubular cells and its possible modulation during damage. To better characterize the phenotype of CD133+ RPCs we first assessed their transcriptional profile by RNA sequencing. The cultured cells expressed highly genes previously reported both by in vivo and ex vivo studies, as characteristics of RPCs 28. In particular, in our CD133+ RPCs we confirmed the expression of the progenitor markers CD24 and PAX2, as well as of vimentin and cytokeratins 18 and 19 (Table 1). The stem cell marker aldehyde dehydrogenase 1, the adhesion molecule VCAM1, claudin, decorin and S100 calcium bind protein A6 (Table 1), all described as characteristic of scattered tubular cells 11, 12, 15, 29, were found highly expressed in our CD133+ RPCs. In addition, these cells expressed the epithelial cell adhesion molecule, known to be expressed by adult tubular CD133+ cells 30, while genes characteristic of metanephric mesenchyme (such as FOXD1, SIX2, CITED1, OSR1, and LGR5) showed low expression or were completely absent (Table 1).
Table 1.
CD133+ cell phenotype
| Gene name | RKPM | |||
|---|---|---|---|---|
| S100A6 | S100 calcium binding protein A6 | 4027,670 | 4148,790 | 3993,890 |
| VIM | Vimentin | 1789,460 | 2013,330 | 1808,040 |
| CD24 | CD24 | 826,579 | 1698,620 | 1019,230 |
| DCN | Decorin | 1250,520 | 260,681 | 257,718 |
| VCAM1 | Vascular cell adhesion molecule | 357,469 | 457,619 | 394,905 |
| KRT18 | Cytokeratin 18 | 138,986 | 346,030 | 323,237 |
| CLDN1 | Claudin 1 | 365,699 | 169,325 | 149,874 |
| KRT19 | Cytokeratin 19 | 350,707 | 162,130 | 71,603 |
| CDH6 | Cadherin 6 | 201,737 | 157,222 | 155,958 |
| ALDH1A1 | Aldehyde dehydrogenase 1 | 43,699 | 216,921 | 243,215 |
| PROM1 | Prominin 1/CD133 | 114,613 | 55,030 | 90,327 |
| SALL1 | Spalt like transcription factor 1 | 30,503 | 18,535 | 16,688 |
| EPCAM | Epithelial cell adhesion molecule | 34,254 | 24,575 | 6,312 |
| PAX2 | Paired Box 2 | 10,375 | 14,563 | 10,320 |
| RARA | Retinoic acid receptor alpha | 7,101 | 10,539 | 8,702 |
| FOXD1 | Forkhead Box D1 | 7,744 | 4,209 | 3,987 |
| NOTCH2 | Notch 2 | 3,886 | 4,028 | 3,717 |
| OSR1 | Odd skipped related transcription factor 1 | 0,362 | 2,006 | 1,323 |
| NCAM1 | Neural cell adhesion molecule 1 | 0,779 | 1,218 | 1.265 |
| WT1 | Wilms tumor 1 | 0,092 | 0,586 | 0,457 |
| EYA1 | Eyes absent homolog 1 | 0,112 | 0,024 | 0,074 |
| CITED1 | Cbp/p300 interacting transactivator 1 | 0 | 0 | 0 |
| LGR5 |
Leucine‐rich repeat‐containing G‐protein coupled receptor 5 |
0 | 0 | 0 |
| SIX2 | SIX homeobox 2 | 0 | 0 | 0 |
| DLL1 | Delta like canonic Notch ligand 1 | 0 | 0 | 0 |
| ACTB | Actin‐B | 1802,060 | 2026,870 | 2783,190 |
| B2M | Beta 2 microglobulin | 1071,690 | 1634,860 | 1376,640 |
Expression of progenitor genes (green) and metanephric mesenchymal genes (red) by RNA sequencing of CD133+ cells presented as descending expression intensity. Data show the expression of three different lines.
Abbreviation: RKPM, reads per kilobase per million mapped reads.
Cisplatin‐Induced Damage Causes Downregulation of CD133 and Expression of Mesenchymal Genes
We then evaluated the modulation of CD133+ RPC phenotype in a model of cisplatin‐induced cell damage. RNA sequencing analysis showed the upregulation of 964 genes and the downregulation of 1,521 genes in CD133+ RPC cells submitted to cisplatin treatment in respect to control untreated cells. Gene ontology analysis showed that cisplatin treatment induced upregulation of genes mainly associated with the following biological processes: nucleic acid metabolism, transcription regulation, cell cycle, proliferation, regulation of enzymatic activities, and cell death processes (Supporting Information Fig. S1). Downregulated genes were involved in multiple other functions, such as cell motility and Rho signal transduction, TGF‐β receptor signaling, chromatin remodeling, morphogenesis, regulation of gene expression, and cytoskeleton reorganization (Supporting Information Fig. S2). Moreover, RNA sequencing data showed that cisplatin treatment for 8 hours induced downregulation of the majority of genes characteristic of CD133+ RPCs, including CD133 itself, PAX2, VCAM1 and cadherin (Fig. 1A). At variance, genes characteristic of metanephric mesenchyme such as KLF4, SNAI2 and FOXD1, along with pro‐regenerative genes WNT3 and SOX9 were found upregulated in cisplatin treated RPCs (Fig. 1A). Cap mesenchyme genes (SIX2, CITED1, and EYA1) remained under the threshold of 1 RKPM. The modulation of PAX2, VCAM1, SOX9, and FOXD1 was confirmed by qRT‐PCR (Fig. 1B). CD133 expression in cisplatin treated RPCs was confirmed as downregulated at 48 hours, to be re‐acquired 1 week later (recovery phase) at RNA level (Fig. 1C). The observed downregulation was further detected at protein level by evaluation of both total CD133 by Western blot analysis and of the cell surface molecule by cytofluorimetric analysis (Fig. 1D, 1E). Moreover, the modulation of the surface CD133 molecule was confirmed on freshly sorted CD133+ cells (>98% positivity for CD133, Supporting Information Fig. S3).
Figure 1.
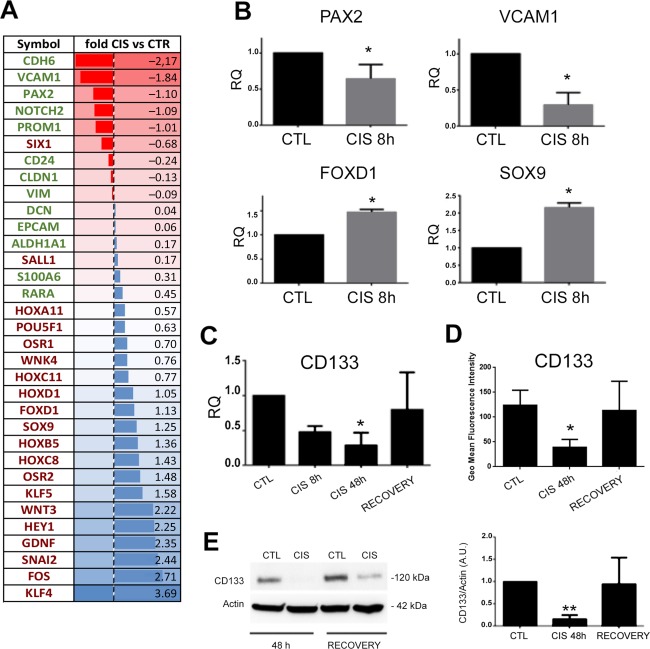
Effect of cisplatin on the phenotype of CD133+ cells. (A): Regulated progenitor genes (green) and mesenchymal genes (red) in CD133+ cells subjected to cisplatin damage, by RNA sequencing. Data show the mean log ratio of three different lines and are presented as descending expression intensity. (B, C): Validation of gene modulation (PAX2, VCAM, FOXD9, SOX9, and CD133), normalized to GAPDH and to CTL, as evaluated by quantitative real‐time PCR analysis. Data are expressed as mean ± SD of at least three different experiments normalized to CTL and to one, t test or One way analysis of variance (ANOVA) (for CD133) was performed: *, p < .05; **, p < .001 versus CTL. (D): CD133 fluorescence intensity, expressed as geometric mean, was evaluated by cytofluorimetric analysis in basal conditions (CTL), or after cisplatin treatment for 48 hours (CIS 48H) or 1 week (RECOVERY). Data are expressed as mean ± SD of four independent experiments. One way ANOVA was performed: *, p < .05 versus CTL. (E): Representative Western blot images and quantification of CD133 expression in cells subjected to cisplatin damage for 48 hours (CIS 48H) or 1 week (RECOVERY). Data are expressed as mean ± SD of three independent experiments, normalized to Actin expression and to one. One way ANOVA was performed: **, p < .001 versus CTL. Abbreviations: RQ, relative quantification; CIS, cisplatin; CTL, Control untreated cells.
Finally, to exclude that the observed modulation in CD133 expression was due to differential loss/expansion of the CD133+ fraction over the CD133− fraction present in the RPC lines (CD133− cells: 8.8 ± 6.1, n = 4), cells in culture were sorted on the basis of CD133 expression and the positive and negative populations labeled with green and red dyes, respectively. Cells were further mixed at 9:1 ratio, subjected to cisplatin damage, and their respective number followed, as described [14]. The ratio of CD133 positive and negative population did not significantly vary over time, indicating the absence of differential loss/expansion of the CD133+ fraction after damage and recovery (Supporting Information Fig. S3). Interestingly, the CD133− cells did not acquire positivity for CD133 (Supporting Information Fig. S3).
These data indicate that CD133+ RPCs lost the characteristic CD133 signature and upregulated genes needed for the reparative process during the early phase of cisplatin damage, modulating their phenotype toward a more mesenchymal state.
Gene Modulation upon Cisplatin Treatment in CD133‐Kd Cells
To further investigate the role of CD133 in cell response upon cisplatin damage, we generated CD133 knockdown RPCs (CD133‐Kd) by lentiviral infection, using two vectors silencing PROM1 gene (shPROM1 and shPROM2) and a scrambled sequence (GFP). The CD133‐Kd RPCs were silenced at high efficiency, as evaluated by Western blot, qRT‐PCR and cytofluorimetric analysis (Fig. 2). RNA sequencing analysis of CD133‐Kd RPCs showed only the specific downregulation of PROM1, indicating no effect of transfection on the cell phenotype (not shown). We then compared cisplatin‐induced gene modulations in both CD133+ (GFP) and CD133‐Kd RPCs. We firstly sorted only transcripts significantly modified in GFP cells by cisplatin. Subsequently, by comparative analysis, we found 102 genes differentially expressed in shPROM1 cells in respect to GFP cells after cisplatin damage. Enrichment analysis of pathways was then conducted using PANTHER bioinformatics tool. An over‐representation of genes related to Wnt and cadherin signaling pathways was observed (Fig. 3A). In addition, PDGF signaling, Alzheimer‐related and DNA replication pathways were also highlighted (Fig. 3A). Sixty‐nine of the 102 modulated transcripts, were confirmed in both shPROM1 and shPROM2 cells after cisplatin damage (mean shPROM1/2 vs. GFP) (Supporting Information Table S2). The analysis of the common genes, conducted using Funrich software, confirmed an enrichment in genes involved in Wnt pathway, along with the DNA repairing process and telomerase synthesis associated pathways (Fig. 3B), supporting the possible implication of these pathways in CD133‐mediated response of RPCs to cisplatin.
Figure 2.
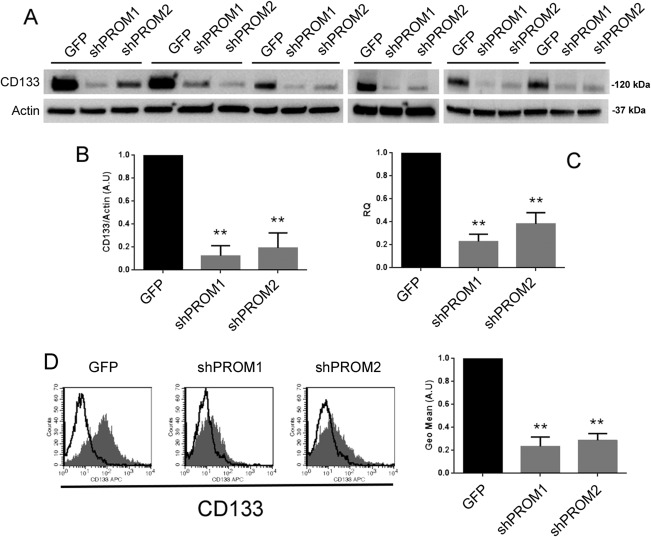
CD133‐Kd generation. The silencing of CD133 antigen in different cell lines was assessed by Western blot, quantitative real‐time PCR (qRT‐PCR) and cytofluorimetric analysis. (A): Representative Western blots of control GFP and different CD133‐Kd lines (shPROM1 and shPROM2). (B): Quantification of CD133 expression as evaluated by Western blot analysis. (C): Quantification of CD133 mRNA expression, normalized to GAPDH and to GFP, as evaluated by qRT‐PCR analysis. (D): Representative cytofluorimetric analysis and quantification as geometric mean of GFP and CD133‐Kd cell lines. The dark gray filled area represents the binding of CD133 mAb, while the overlaying black line represents the isotypic control. (B–D) Represent the mean ± SD of 12 different cell lines generated in the study and are normalized to GFP and to one. One way analysis of variance was performed: **, p < .001 versus GFP.
Figure 3.
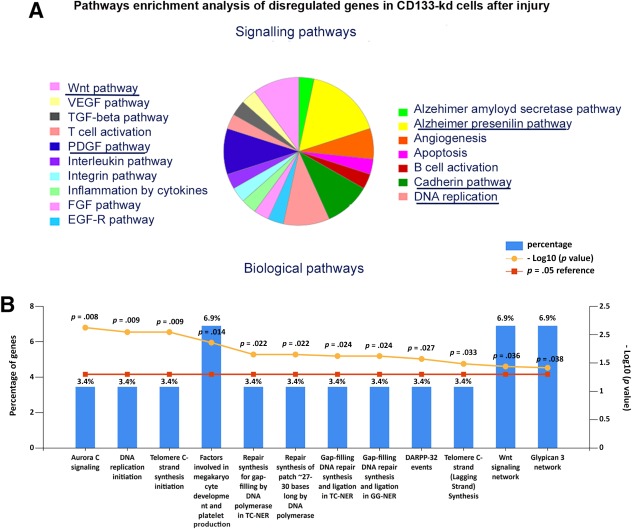
Molecular effects of cisplatin on CD133‐Kd cells. (A): Signaling pathways over‐represented by the 102 genes differentially expressed in shPROM1 in respect to CD133+ control cells after cisplatin treatment. (A) Panther analysis showed the enrichment of different pathways. The most represented pathways were PDGF signaling, Alzheimer, DNA replication, Wnt, and E‐cadherin related pathways (underlined in the figure). (B): Funrich confirmation of the biological pathways associated with genes disregulated in CD133‐Kd cisplatin treated cells (shPROM1 and shPROM2 cells). Columns represent the percentage of genes involved in each biological pathway.
CD133 Is Implicated in Wnt/β‐Catenin Signaling
We therefore focused our attention on the role of CD133 in the activation of Wnt pathway. Interestingly, CD133‐Kd cell lines showed a reduced expression of Wnt4 after cisplatin‐induced damage in respect to GFP cells (Fig. 4A). In addition, CD133‐Kd cells showed an impaired response to the pharmacological activation of Wnt signaling pathway using CHIR99021 31 (Fig. 4B). The role of CD133 in Wnt signaling activation was confirmed using a luciferase assay reporting the activation of the TCF/LEF promoter. As shown in Figure 4C, luciferase activity in CD133+ RPCs was significantly higher than the one in CD133‐Kd cell lines, indicating a reduced β‐catenin activation in CD133‐Kd cells. This was not related to differential levels of GSK3 alpha and beta in CD133‐Kd cell lines in respect to CD133+ RPCs, as assessed by Western blot analysis (not shown).
Figure 4.
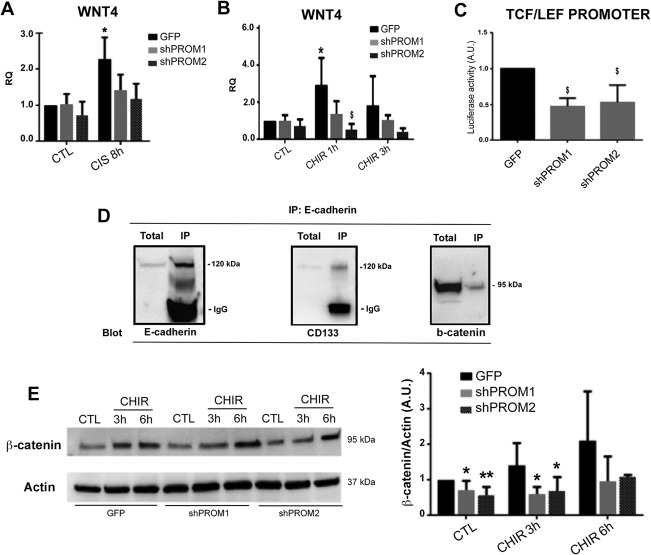
Impaired activation of Wnt signaling in CD133‐Kd lines. (A, B): Quantitative real‐time PCR showing the expression of WNT4 upon cisplatin induced damage (A) or stimulation with CHIR99021 (B) by CD133+ (GFP) and CD133‐Kd (shPROM1 and shPROM2) cell lines. Data represent the mean ± SD of five different experiments and are normalized to GAPDH and to GFP. One way analysis of variance (ANOVA) was performed: *, p < .05 versus CTL. (C): Luciferase activity in CD133+ and CD133‐Kd cells as evaluated by the TCF/LEF reporter assay. (A–C) Data represent the mean ± SD of three different experiments and are normalized to GFP and to one. One way ANOVA was performed: $, p < .05 versus GFP. (D): Western blot analysis of E‐cadherin, CD133, and β‐catenin after immunoprecipitation with E‐cadherin, showing the association of the molecules. (E): Expression of β‐catenin by CD133+ and CD133‐Kd cell lines, in basal condition (CTL) and upon stimulation with CHIR99021 (CHIR) for 3 and 6 hours respectively as assessed by Western blot. Data are normalized to GFP and to one and expressed as mean ± SD of at least five different experiments. One way ANOVA was performed: *, p < .05; **, p < .001 versus GFP. Abbreviation: RQ, relative quantification.
We then analyzed the possible interaction of CD133 with β‐catenin and E‐cadherin, as reported in other stem cell types 32. Western blot analysis after immunoprecipitation of E‐cadherin showed the concomitant presence of CD133 and β‐catenin, suggesting that CD133 and E‐cadherin may form a complex at the membrane level with β‐catenin, thus limiting its cytoplasmic degradation (Fig. 4D). Indeed, β‐catenin levels were significantly lower in CD133‐Kd cells in respect to CD133+ RPCs both in basal culture conditions and upon stimulation with β‐catenin stabilizer CHIR99021 (Fig. 4E). Taken together, these data confirm the results obtained by RNA sequencing analysis, involving CD133 molecule in the regulation of Wnt canonical pathway in RPCs.
CD133 Is Involved in the Regenerative Response After Damage
Tubular cell recovery after damage appears to be mediated by clonal proliferation of Wnt‐responsive cells 8. We therefore, evaluated whether CD133 molecule could play a role in the regenerative response, by studying proliferation of CD133‐Kd cells after cisplatin‐induced damage, and generation of spheroids under hypoxia (Fig. 5). When CD133+ and CD133‐Kd RPCs were subjected to cisplatin treatment, they similarly underwent reduction in cell number (Fig. 5A). However, CD133‐Kd cell lines exhibited a significantly lower proliferative capacity during the recovery phase compared to the CD133+ RPCs (Fig. 5A). In addition, evaluation of kidney spheroid generation in hypoxic conditions showed an impaired capacity of CD133‐Kd cells to generate spheres, which were less in number and significantly smaller in surface area (Fig. 5B, 5C). The viability of spheroids formed by CD133‐Kd cells appeared reduced as well (Fig. 5C). These data suggest a role of CD133 in the maintenance of clonal proliferation of RPCs, an important process for the regenerative capacity of the kidney.
Figure 5.
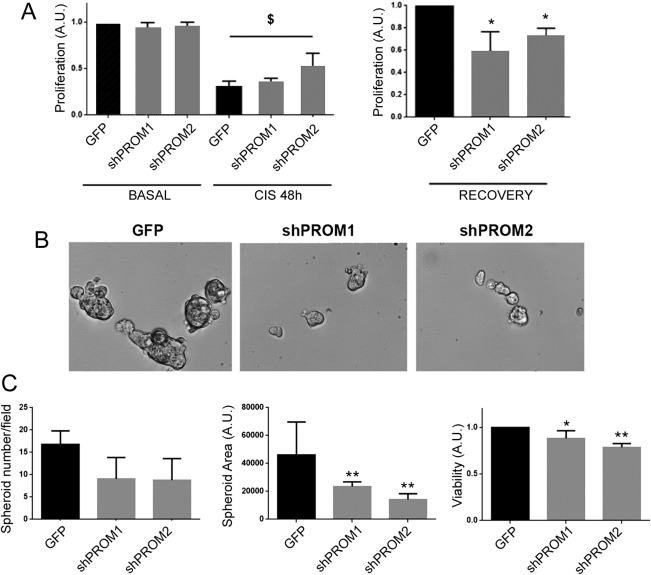
Impaired proliferation and spheroid generation of CD133‐Kd cells after damage. (A): Proliferation of CD133+ (GFP) and CD133‐Kd (shPROM1 and shPROM2) cells was evaluated in the early phase of damage (CIS 48H) and during recovery phase (RECOVERY) by the incorporation of BrdU. Data are expressed as mean ± SD of three different experiments normalized to CTL and to one. One way analysis of variance (ANOVA) was performed: $, p < .05 versus BASAL and *, p < .05 versus GFP. (B): Representative micrograph of GFP and CD133‐Kd (shPROM1 and shPROM2) cells growing as spheroids. Original magnification: ×200. (C): Evaluation of number, surface area and viability of spheroids. Data are expressed as mean ± SD of the number of spheroids/field or spheroid area/field in at least 20 fields per experiment (n = 3). Data on viability are expressed as mean ± SD of three experiments and normalized to GFP and to one. One way ANOVA was performed: *, p < .05; **, p < .001 versus GFP.
CD133 Expression Limits Cell Senescence
Finally, we evaluated the role of CD133 in cell senescence, a mechanism of exhaustion occurring after repeated proliferation cycles 33. CD133+ and CD133‐Kd RPCs were maintained in culture for five passages, and β‐galactosidase expression and telomere length were evaluated at passage six. The number of β‐galactosidase expressing cells undergoing senescence appeared significantly higher in the CD133‐Kd cells compared to their CD133+ control (Fig. 6A, 6B). To further confirm these data, we assessed the telomere length using an optimized assay originally described by Cawthon 27. The evaluation of T/S ratio revealed a significant reduction in telomere length in CD133‐Kd cells compared to their CD133+ controls (Fig. 6C), suggesting an implication of CD133 in the prevention of cell senescence.
Figure 6.
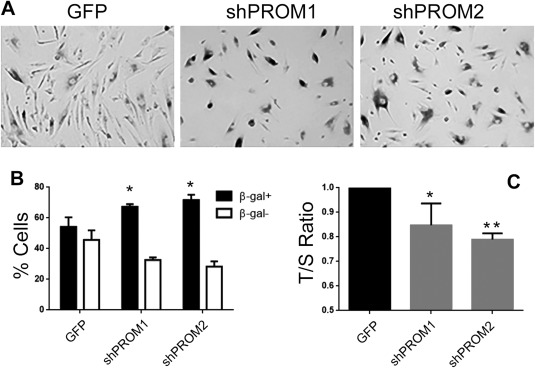
Involvement of CD133 in cell senescence. (A): Representative micrograph of CD133+ (GFP) and CD133‐Kd (shPROM1 and shPROM2) cells stained for β‐galactosidase (β‐gal). Original magnification: ×200. The number of β‐gal positive cells in CD133+ and CD133‐Kd lines was evaluated as the percentage of β‐gal positive and negative cells/field in at least 10 fields per experiment (B). Data are expressed as mean ± SD of three different experiments. (C): T/S ratio analysis expressed as mean ± SD of T/S ratio normalized to GFP and to one, of eight different experiments. One way analysis of variance was performed: *, p < .05; **, p < .001 versus GFP.
Discussion
The identification of mechanisms involved in proliferation and repair after injury are still unclear in human tubular cells. In the present study, we evaluated the role of stem cell marker CD133 in renal cellular repair at both molecular and functional level. We found that CD133 regulated β‐catenin and that its absence impaired cell proliferation after injury, clonal generation of spheres while favored senescence, indicating that CD133 may act as a permissive factor for Wnt/beta catenin signaling and may play a role in tissue repair.
During nephrogenesis, CD133 is expressed from the vesicle stage onwards 34, 35 and marks a population of epithelial differentiated embryonic renal cells 36, while in adult renal tissue is maintained in unevenly distributed cells, commonly referred to as scattered cells 11, 13, 14. Beside CD133, a number of different markers have been reported as characteristic of these scattered cells. We here validated the expression of genes characterizing the progenitor cell phenotype in cultured CD133+ RPCs. In particular, CD133+ RPCs expressed CD24, vimentin, cytokeratins, VCAM1 and decorin, as previously reported 11, 21, 29 as well as claudin 1, S100A6 [11, 15] and aldehyde dehydrogenase 1 [12]. These data indicate that the phenotype of cultured CD133+ RPCs reflects the one observed in vivo in human tissues, validating our in vitro model. The expression of CD133 has been shown to increase in biopsies of renal tissue after acute tubular necrosis 14. It was unclear, however, whether this increase could represent the expansion of CD133+ RPCs or a de novo acquisition of CD133. When analyzing the changes occurring in CD133+ RPCs upon cisplatin damage, we found a down‐regulation of the CD133 molecule that was reversed during recovery. This modulation was accompanied by loss of the characteristic signature reported for RPCs. In parallel, CD133+ RPCs activated a number of mesenchymal‐associated transcriptional factors including FOXD1 and SNAIL1 as well as SOX9 required for the reparative process. Indeed, mesenchymal genes are known to be expressed in vivo by renal tissue after injury in human and murine tissues 9, 30, 37. It can be therefore postulated that CD133+ RPCs possess a plastic phenotype, and that the CD133 molecule marks cells able to contribute to the regeneration processes of the kidney. However, RPCs did not acquire a phenotype comparable to embryonic nephron progenitors, characterized by the expression of SIX2, CITED1, OSR1, and EYA. Indeed, it has been reported that SNAIL1 is required but not sufficient to generate cells with the phenotype of nephron progenitors 38.
In our in vitro model, we found a rapid (24–48 hours) downregulation of CD133 after injury, followed by its re‐expression during the regeneration phase, characterized by cell proliferation. The observed loss of CD133 upon tissue damage (24–48 hours) was not reported in renal biopsies of AKI patients 15, 16. In particular, the description of AKI tissue reported the presence of long stretches of CD133+ cells in regenerating tubules 15, and association of CD133 expression with light microscopic signs of tubular regeneration such as nuclear prominence or flattening of tubular cells 16. It could be speculated that CD133 expression in renal biopsies of AKI patients may reflect a phase of tubular regeneration occurring in the days after the damage, rather than a phase of injury. As there is no indication on the timing of the biopsy in both papers, this cannot be precisely estimated. It is also conceivable that injuries of a different nature (toxic vs. ischemic) may have a differential impact on CD133 expression. At variance, our results are consistent with the analysis of human tissue deriving from patients undergoing kidney transplant. In fact, patients with delayed graft function, undergoing acute tubular damage, showed an early reduction in the number of CD133+ proliferating cells in respect to pre‐transplant biopsies and early graft function tissues 39. In this study, CD133 levels were restored within 4 weeks after transplant 39. The loss and re‐acquisition of CD133 upon cisplatin damage clearly limit the role of this molecule to the early or late phases of tissue regeneration. Indeed, a very limited number of genes showed an altered modulation in the early response to injury in CD133‐Kd cells respect to CD133+ cells. In particular, genes related to Wnt and E‐cadherin signaling were disregulated. The role of CD133 as inductor of Wnt/β‐catenin signaling has been previously reported in tumor stem cells. In particular, CD133 has been found implicated in Wnt canonical pathway through the stabilization of Wnt's primary target β‐catenin 40 and the consequent activation of TCF/LEF promoter 41. Accordingly, we found that CD133 is associated with E‐cadherin and β‐catenin, and that β‐catenin levels and transcriptional activity are decreased in CD133‐Kd cells, implying a role for CD133 in β‐catenin protection from degradation. The activation of Wnt/β‐catenin signaling is involved in kidney damage by promoting cellular repair 42, 43, and in particular by inducing the proliferation of regenerating clonal cells in tubules 8. Therefore, CD133+ cells appear a relevant population involved in kidney regeneration. In addition, these data parallel the role of CD133, expressed on human renal tissue, with the one of Lgr5, a Wnt target gene restricted to clusters of undifferentiated epithelial cells residing within tubular structures in the early postnatal kidneys 7. The Lgr5+ cells have been shown to selectively contribute to the development of Henle's loop and distal tubules 7.
The function of CD133 has been previously associated with glycolytic metabolism and invasion of tumor stem cells 44. In our study, we identified a functional role of CD133 in normal RPCs related to the proliferation and to prevention of cell senescence. In particular, our data showed that CD133 loss limited cell proliferation after injury, in the recovery phase, and cell ability to grow in spheres under hypoxia. In this last condition, kidney cells are known to enhance the expression of stemness related markers and tubulogenic potential 45, 46. Our data, suggesting an implication of CD133+ RPCs in the repairing process of renal tissue as well as an ability to survive an insult, are in line with data reported in an ex vivo explant model, in which CD133+ scattered cells selectively survived an ischemia–reperfusion damage 15. Finally, our study showed that the expression of CD133 is pivotal in the regulation of cell senescence, a characteristic of cell exhaustion after repeated proliferation cycles. Senescence, physiologically associated with aging, reduces the ability of tissue repair and may be correlated with maldifferentiation and fibrosis development after injury 33. Considering these implications of CD133 in the process of renal tissue repair, our data suggest that CD133 does not solely represent a stem cell marker, but rather a functional protein.
Author Contributions
A.B. and E.P.: study design, data collection, analysis and interpretation, manuscript writing; F.C. and S.O.: data analysis and interpretation; D.I.: data collection, analysis and interpretation; G.C.: manuscript writing; B.B.: conception and design, financial support, data analysis and interpretation, manuscript writing.
Disclosure of Potential Conflicts of Interest
The authors indicated no potential conflicts of interest.
Supporting information
Supplementary Figure 1
Supplementary Figure 2
Supplementary Figure 3
Supplementary Tables, Materials and Fig. Legends
Conclusion
Our data indicate that CD133 identifies a Wnt‐responding population, able to de‐differentiate and proliferate in response to injury. In particular, CD133 itself appears to play a role in the maintenance of cell homeostasis, clonal proliferation, and renal tubular repair, by controlling Wnt signaling while limiting cell senescence.
Acknowledgments
This work was supported by the FP7 Marie Curie NephroTools. E.P. was a Marie Curie fellow of the NephroTools project.
References
- 1. Eckardt KU, Coresh J, Devuyst O et al. Evolving importance of kidney disease: From subspecialty to global health burden. Lancet 2013;382:158–169. [DOI] [PubMed] [Google Scholar]
- 2. Bydash JR, Ishani A. Acute kidney injury and chronic kidney disease: A work in progress. Clin J Am Soc Nephrol 2011;6:2555–2557. [DOI] [PubMed] [Google Scholar]
- 3. Coca SG, Singanamala S, Parikh CR. Chronic kidney disease after acute kidney injury: A systematic review and meta‐analysis. Kidney Int 2012;81:442–448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Chang‐Panesso M, Humphreys BD. Cellular plasticity in kidney injury and repair. Nat Rev Nephrol 2017;13:39–46. [DOI] [PubMed] [Google Scholar]
- 5. Lombardi D, Becherucci F, Romagnani P. How much can the tubule regenerate and who does it? An open question. Nephrol Dial Transplant 2016;31:1243–1250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Kusaba T, Lalli M, Kramann R et al. Differentiated kidney epithelial cells repair injured proximal tubule. Proc Natl Acad Sci USA 2014;111:1527–1532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Barker N, Rookmaaker MB, Kujala P et al. Lgr5(+ve) stem/progenitor cells contribute to nephron formation during kidney development. Cell Rep 2012;2:540–552. [DOI] [PubMed] [Google Scholar]
- 8. Rinkevich Y, Montoro DT, Contreras‐Trujillo H et al. In vivo clonal analysis reveals lineage‐restricted progenitor characteristics in mammalian kidney development, maintenance, and regeneration. Cell Rep 2014;7:1270–1283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Kumar S, Liu J, Pang P et al. Sox9 activation highlights a cellular pathway of renal in the acutely injured mammalian kidney. Cell Rep 2015;12:1325–1338. [DOI] [PubMed] [Google Scholar]
- 10. Kang HM, Huang S, Reidy K et al. Sox9‐positive progenitor cells play a key role in renal tubule epithelial regeneration in mice. Cell Rep 2016;14:861–871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Smeets B, Boor P, Dijkman H et al. Proximal tubular cells contain a phenotypically distinct, scattered cell population involved in tubular regeneration. J Pathol 2013;229:645–659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Lindgren D, Boström AK, Nilsson K et al. Isolation and characterization of progenitor‐like cells from human renal proximal tubules. Am J Pathol 2011;178:828–837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Bussolati B, Bruno S, Grange C et al. Isolation of renal progenitor cells from adult human kidney. Am J Pathol 2005;166:545–555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Angelotti ML, Ronconi E, Ballerini L et al. Characterization of renal progenitors committed toward tubular lineage and their regenerative potential in renal tubular injury. Stem Cells 2012;30:1714–1725. [DOI] [PubMed] [Google Scholar]
- 15. Hansson J, Hultenby K, Cramnert C et al. Evidence for a morphologically distinct and functionally robust cell type in the proximal tubules of human kidney. Hum Pathol 2014;45:382–393. [DOI] [PubMed] [Google Scholar]
- 16. Miraglia S, Godfrey W, Yin AH et al. A novel five‐transmembrane hematopoietic stem cell antigen: Isolation, characterization, and molecular cloning. Blood 1997;90:5013–5021. [PubMed] [Google Scholar]
- 17. Corbeil D, Fargeas CA, Huttner WB. Rat prominin, like its mouse and human orthologues, is a pentaspan membrane glycoprotein. Biochem Biophys Res Commun 2001;285:939–944. [DOI] [PubMed] [Google Scholar]
- 18. Kemper K, Sprick MR, de Bree M et al. The AC133 epitope, but not the CD133 protein, is lost upon cancer stem cell differentiation. Cancer Res 2010;70:719–729. [DOI] [PubMed] [Google Scholar]
- 19. Yin AH, Miraglia S, Zanjani ED et al. AC133, a novel marker for human hematopoietic stem and progenitor cells. Blood 1997;90:5002–5012. [PubMed] [Google Scholar]
- 20. Corbeil D, Röper K, Hellwig A et al. The human AC133 hematopoietic stem cell antigen is also expressed in epithelial cells and targeted to plasma membrane protrusions. J Biol Chem 2000;275:5512–5520. [DOI] [PubMed] [Google Scholar]
- 21. Bussolati B, Moggio A, Collino F et al. Hypoxia modulates the undifferentiated phenotype of human renal inner medullary CD133+ progenitors through Oct4/miR‐145 balance. Am J Physiol Renal Physiol 2012;302:F116–F128. [DOI] [PubMed] [Google Scholar]
- 22. Pathan M, Keerthikumar S, Ang CS et al. An open access standalone functional enrichment and interaction network analysis tool. Proteomics 2015;15:2597–2601. [DOI] [PubMed] [Google Scholar]
- 23. Maere S, Heymans K, Kuiper M. BiNGO: A Cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. Bioinformatics 2005;21:3448–3449. [DOI] [PubMed] [Google Scholar]
- 24. Bombelli S, Zipeto MA, Torsello B et al. PKH(high) cells within clonal human nephrospheres provide a purified adult renal stem cell population. Stem Cell Res 2013;11:1163–1177. [DOI] [PubMed] [Google Scholar]
- 25. Koppelstaetter C, Jennings P, Hochegger K et al. Effect of tissue fixatives on telomere length determination by quantitative PCR. Mech Ageing Dev 2005;126:1331–1333. [DOI] [PubMed] [Google Scholar]
- 26. Pooley KA, Sandhu MS, Tyrer J et al. Telomere length in prospective and retrospective cancer case‐control studies. Cancer Res 2010;70:3170–3176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Cawthon RM. Telomere measurement by quantitative PCR. Nucleic Acids Res 2002;30:e47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Bussolati B, Camussi G. Therapeutic use of human renal progenitor cells for kidney regeneration. Nat Rev Nephrol 2015;11:695–706. [DOI] [PubMed] [Google Scholar]
- 29. Sallustio F, De Benedictis L, Castellano G et al. TLR2 plays a role in the activation of human resident renal stem/progenitor cells. FASEB J 2010;24:514–525. [DOI] [PubMed] [Google Scholar]
- 30. Buzhor E, Omer D, Harari‐Steinberg O et al. Reactivation of NCAM1 defines a subpopulation of human adult kidney epithelial cells with clonogenic and stem/progenitor properties. Am J Pathol 2013;183:1621–1633. [DOI] [PubMed] [Google Scholar]
- 31. Kim H, Wu J, Ye S et al. Modulation of β‐catenin function maintains mouse epiblast stem cell and human embryonic stem cell self‐renewal. Nat Commun 2013;4:2403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Gay DL, Yang CC, Plikus MV et al. CD133 expression correlates with membrane beta‐catenin and E‐cadherin loss from human hair follicle placodes during morphogenesis. J Invest Dermatol 2015;135:45–55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Ferenbach DA, Bonventre JV. Mechanisms of maladaptive repair after AKI leading to accelerated kidney ageing and CKD. Nat Rev Nephrol 2015;11:264–276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Lazzeri E, Crescioli C, Ronconi E et al. Regenerative potential of embryonic renal multipotent progenitors in acute renal failure. J Am Soc Nephrol 2007;18:3128–3138. [DOI] [PubMed] [Google Scholar]
- 35. Pode‐Shakked N, Pleniceanu O, Gershon R et al. Dissecting stages of human kidney development and tumorigenesis with surface markers affords simple prospective purification of nephron stem cells. Sci Rep 2016;6:23562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Metsuyanim S, Harari‐Steinberg O, Buzhor E et al. Expression of stem cell markers in the human fetal kidney. PLoS One 2009;4:e6709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Witzgall R, Brown D, Schwarz C et al. Localization of proliferating cell nuclear antigen, vimentin, c‐Fos, and clusterin in the postischemic kidney. Evidence for a heterogenous genetic response among nephron segments, and a large pool of mitotically active and dedifferentiated cells. J Clin Invest 1994;93:2175–2188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Hendry CE, Vanslambrouck JM, Ineson J et al. Direct transcriptional reprogramming of adult cells to embryonic nephron progenitors. J Am Soc Nephrol 2013;24:1424–1434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Loverre A, Capobianco C, Ditonno P et al. Increase of proliferating renal progenitor cells in acute tubular necrosis underlying delayed graft function. Transplantation 2008;85:1112–1119. [DOI] [PubMed] [Google Scholar]
- 40. Mak AB, Nixon AM, Kittanakom S et al. Regulation of CD133 by HDAC6 promotes β‐catenin signaling to suppress cancer cell differentiation. Cell Rep 2012;2:951–963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Rappa G, Mercapide J, Anzanello F et al. Wnt interaction and extracellular release of prominin‐1/CD133 in human malignant melanoma cells. Exp Cell Res 2013;319:810–819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Kawakami T, Ren S, Duffield JS. Wnt signalling in kidney diseases: Dual roles in renal injury and repair. J Pathol 2013;229:221–231. [DOI] [PubMed] [Google Scholar]
- 43. Terada Y, Tanaka H, Okado T et al. Expression and function of the developmental gene Wnt‐4 during experimental acute renal failure in rats. J Am Soc Nephrol 2003;14:1223–1233. [DOI] [PubMed] [Google Scholar]
- 44. Rappa G, Fodstad O, Lorico A. The stem cell‐associated antigen CD133 (Prominin‐1) is a molecular therapeutic target for metastatic melanoma. Stem Cells 2008;26:3008–3017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Buzhor E, Harari‐Steinberg O, Omer D et al. Kidney spheroids recapitulate tubular organoids leading to enhanced tubulogenic potency of human kidney‐derived cells. Tissue Eng Part A 2011;17:2305–2319. [DOI] [PubMed] [Google Scholar]
- 46. Little MH, Kairath P. Does renal repair recapitulate kidney development? J Am Soc Nephrol 2017;28:34–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Figure 1
Supplementary Figure 2
Supplementary Figure 3
Supplementary Tables, Materials and Fig. Legends
Data Availability Statement
FastQ files for RNA‐seq experiments are deposited on the Gene Expression Omnibus database, under the accession code GSE107273.


