Abstract
Endoplasmic reticulum (ER) stress has been implicated in the development of maternal diabetes-induced neural tube defects (NTDs). ER stress-induced C/EBP homologous protein (CHOP) plays an important role in the pro-apoptotic execution pathways. However, the molecular mechanism underlying ER stress- and CHOP-induced neuroepithelium cell apoptosis in diabetic embryopathy is still unclear. Deletion of the Chop gene significantly reduced maternal diabetes-induced NTDs. CHOP deficiency abrogated maternal diabetes-induced mitochondrial dysfunction and neuroepithelium cell apoptosis. Further analysis demonstrated that CHOP repressed the expression of peroxisome-proliferator-activated receptor-γ coactivator-1α (PGC-1α), an essential regulator for mitochondrial biogenesis and function. Both CHOP deficiency in vivo and knockdown in vitro restore high glucose-suppressed PGC-1α expression. In contrast, CHOP overexpression mimicked inhibition of PGC-1α by high glucose. In response to the ER stress inducer tunicamycin, PGC-1α expression was decreased, whereas the ER stress inhibitor 4-phenylbutyric acid blocked high glucose-suppressed PGC-1α expression. Moreover, maternal diabetes in vivo and high glucose in vitro promoted the interaction between CHOP and the PGC-1α transcriptional regulator CCAAT/enhancer binding protein-β (C/EBPβ), and reduced C/EBPβ binding to the PGC-1α promoter leading to markedly decrease in PGC-1α expression. Together, our findings support the hypothesis that maternal diabetes-induced ER stress increases CHOP expression which represses PGC-1α through suppressing the C/EBPβ transcriptional activity, subsequently induces mitochondrial dysfunction and ultimately results in NTDs.
Keywords: neural tube defects, ER stress, CHOP, C/EBPβ, PGC-1α, diabetic embryopathy, mitochondrial dysfunction
Pre-gestational diabetes triggers failure of neural tube closure during embryonic development leading to neural tube defects (NTDs), also known as diabetic embryopathy (Dong et al., 2016a,b; Gabbay-Benziv et al., 2015; Li et al., 2012, 2013; Salbaum and Kappen, 2010; Wang et al., 2013, 2015a, 2017; Wu et al., 2015; Yang et al., 2013, 2015; Zhong et al., 2016a,b). Studies from our group and others have consistently demonstrated that endoplasmic reticulum (ER) stress and its resultant apoptosis in neuroepithelial cells play a critical role in the pathogenesis of maternal diabetes-induced NTDs (Dong et al., 2016a; Gu et al., 2016; Li et al., 2013; Wang et al., 2015b; Wu et al., 2015; Xu et al., 2013; Zhong et al., 2016a). However, the molecular events downstream of ER stress that mediate the teratogenicity of maternal diabetes remain to be elucidated.
CCAAT/enhancer-binding proteins (C/EBPs), a family of transcription factors, regulate an array of cellular functions including cell differentiation and apoptosis (Ramji and Foka, 2002). There are 6 members in the C/EBPs family including C/EBPα, β, γ, δ, ε, and C/EBP homologous protein (CHOP), which is also known as growth arrest and DNA damage-inducible gene 153 (GADD153) (Ramji and Foka, 2002). CHOP has been defined as an ER chaperone gene that mediates the pro-apoptotic effect of prolonged ER stress (Iurlaro and Munoz-Pinedo, 2016; Sano and Reed, 2013). Several pro-apoptotic events downstream of CHOP have been identified. CHOP modulates the Bcl-2 family members by down-regulating the anti-apoptotic Bcl-2 family proteins such as Mcl-1 and Bcl-2, and up-regulating the pro-apoptotic protein Bim (Iurlaro and Munoz-Pinedo, 2016). CHOP activates the death receptor 5 (DR5) which induces apoptosis through caspase 8 activation (Iurlaro and Munoz-Pinedo, 2016). CHOP is expressed in a low level under non-stress conditions, whereas its expression is robustly increased by the 3 arms of the unfolded protein response (UPR): IRE1α, PERK, and ATF6α, upon ER stress (Senft and Ronai, 2015). It has been shown that maternal diabetes induces ER stress and UPR activation leading to a robust increase of apoptosis in the developing neuroepithelium (Dong et al., 2016a; Gu et al., 2016; Li et al., 2013). However, the role CHOP in diabetic embryopathy is still elusive.
Consistent with the adverse impact of CHOP on Bcl-2 family members, critical regulators of mitochondrial function, defective mitochondrial, and mitochondrial dysfunction are manifested in neuroepithelial cells exposed to maternal diabetes (Wang et al., 2017; Xu et al., 2013), suggesting that CHOP may alter mitochondrial function by modulating key mitochondrial regulators. Peroxisome-proliferator-activated receptor-γ coactivator-1 alpha (PGC-1α) is a master regulator of mitochondrial biogenesis (Austin and St-Pierre, 2012; Scarpulla, 2011), and its activity is regulated by transcriptional and post-transcriptional mechanisms (Karamanlidis et al., 2007; Karamitri et al., 2009; Wang et al., 2008, 2017). Maternal diabetes suppresses PGC-1α expression (Wang et al., 2017). Previous studies have demonstrated that C/EBPβ is a positive transcriptional regulator for PGC-1α in the liver (Wang et al., 2008). C/EBPβ-induced PGC-1α expression is hindered by a transcription repressor, histone deacetylase 1 (Jin et al., 2013). Because CHOP represses gene transcription through binding with other C/EBP proteins and, thus, preventing them binding to DNA (Ron and Habener, 1992), CHOP may participate in maternal diabetes-repressed PGC-1α expression by interfering with the DNA binding activity of other C/EBP proteins. In addition, CHOP suppresses metabolic gene expression during ER stress in the liver (Chikka et al., 2013). Thus, CHOP may contribute to ER stress-induced neuroepithelial cell apoptosis and NTD formation through suppression of PGC-1α expression in diabetic embryopathy.
In the present study, we report that ER stress-increased CHOP is responsible for maternal diabetes-induced neuroepithelial cell apoptosis and NTD formation using CHOP knockout mice. We further reveal that deletion of the Chop gene reverses maternal diabetes-induced mitochondrial dysfunction and restores PGC-1α expression. C/EBPβ is a transcriptional activator for PGC-1α in the developing embryo, whereas CHOP blocks the DNA-binding ability of C/EBPβ through forming CHOP/C/EBP heterodimer and represses PGC-1α expression. These findings support the important role of CHOP-suppressed PGC-1α in ER stress-induced diabetic embryopathy.
MATERIALS AND METHODS
Mice
All procedures for animal use were approved by the Institutional Animal Care and Use Committee of University of Maryland School of Medicine. Wild-type (WT) C57BL/6J mice and CHOP knockout mice were purchased from the Jackson Laboratory (Bar Harbor, ME). The male CHOP+/− mice was mated with non-diabetic or diabetic female CHOP+/− to generate WT, CHOP+/−, CHOP−/− embryos in 1 l.
Model of diabetic embryopathy and morphological assessment of NTDs
We (Gu et al., 2016; Li, et al., 2012, 2013; Wang et al., 2015b; Yang et al., 2013; Zhong et al., 2016a) and others (Kamimoto et al., 2010; Salbaum and Kappen, 2010; Sugimura et al., 2009) have used a rodent model of Streptozotocin (STZ)-induced diabetes in research of diabetic embryopathy. Briefly, 10-week-old WT or knockout female mice were intravenously injected daily with 75 mg/kg Streptozotocin (STZ) for 2 days to induce diabetes. STZ from Sigma (St. Louis, MO) was dissolved in sterile 0.1 M citrate buffer (pH4.5). We used the U-100 Insulin Syringe (Becton Dickinson) with 28G1/2 needles for injections. Approximately 140 µl of STZ solution was injected per mouse. Diabetes was defined as a 12-h fasting blood glucose level of ≥16.7 mM. Male and female mice were paired at 3:00 P.M., and day 0.5 (E0.5) of pregnancy was established at noon of the day when a vaginal plug was present. Embryos were harvested for subsequence analysis on different developing stage. Embryos were harvested at E8.75 (2:00 PM at E8.5) for biochemical and molecular analyses. At E10.5, embryos were examined under a Leica MZ16F stereo microscope (Bannockburn, IL) to identify NTDs. Images of embryos were captured by a DFC420 5 MPix digital camera with software (Leica, Bannockburn, IL) and processed with Adobe Photoshop CS2. Normal embryos were classified as possessing completely closed neural tube and no evidence of other malformations. Malformed embryos were classified as showing evidence of failed closure of the anterior neural tubes resulting in exencephaly, a lethal type of NTDs with the absence of a major portion of the brain, skull, and scalp. Because our model induces NTD at E10.5, we did not examine other major structural malformations such as cardiovascular defects, which do not occur until later embryonic stages (E15.5).
RT-qPCR
Total RNA was isolated from embryos and cells using the TRIzol reagent (Ambion) and reverse transcribed using the QuantiTect Reverse Trancription Kit (Qiagen). RT-qPCR of the internal control β-actin (Forward: 5′GTGACGTTGACATCCGTAAAGA3′, Reverse: 5′GCCGGACTCATCGTACTCC3′), CHOP (Forward: 5′CGGAACCTGAGGAGAGAGTG3′, Reverse: 5′CTGTCAGCCAAGCTAGGGAC3′) and PGC-1α (Forward: 5′ TATGGAGTGACATAGAGTGTGCT3′, Reverse: 5′ CCACTTCAATCCACCCAGAAAG3′) was performed using the Maxima SYBR Green/ROX qPCR Master Mix assay Kit (Thermo Scientific). RT-qPCR and subsequent calculations were performed by the StepOnePlus™ Real-Time PCR System (Applied Biosystem).
Hematoxylin–eosin staining
E10.5 embryos were fixed in methacarn (60% methanol, 30% chloroform, and 10% glacial acetic acid), embedded in paraffin, and cut into 5 μm sections. After deparaffinization and rehydration, all specimens then underwent hematoxylin and eosin (H & E) staining in a standard procedure. All sections were photographed and examined for neural tube defects.
TUNEL staining
The TUNEL assay was performed using the In Situ Cell Death Detection Kit (Millipore) as previously described (Gu et al., 2016; Yang et al., 2013). Briefly, 10 μm frozen embryo sections were fixed with 1% paraformaldehyde in PBS, incubated with TUNEL reagents, counterstained with 4′,6-diamidino-2-phenylindole (DAPI) and mounted with aqueous mounting medium (Sigma). TUNEL-positive cells in each well were counted. The percentage of apoptotic cells was calculated as the number of TUNEL-positive (apoptotic) cells divided by the total number of cells in a microscopic field from 3 separate experiments.
Cell culture and treatment
C17.2 mouse neural stem cells, originally obtained from ECACC (European Collection of Cell Culture), were maintained in DMEM (5 mM glucose) supplemented with 10% fetal bovine serum, 100 U/ml penicillin and 100 µg/ml streptomycin at 37 °C in a humidified atmosphere of 5% CO2. The C17.2 cells are newborn mouse cerebellar progenitor cells transformed with retroviral v-myc (Snyder et al., 1992). Lipofectamine 2000 (Invitrogen) was used according to the manufacturer’s protocol for transfection of CHOP siRNA or scramble control siRNA (Thermo Scientific) and C/EBPβ expression vector into the cells using 1% fetal bovine serum culture conditions. C/EBPβ expression vector was a gift from Jed Friedman (addgene # #49198). In order to investigate the possible effect of ER stress on PGC-1α expression, ER stress inducer tunicamycin (1 μg/ml) (Sigma-Aldrich) was added to growth medium for indicated time in figures and then cells were harvested. To determine the effect of the ER stress inhibitor 4-phenylbutyric acid (4-PBA) on tunicamcin-suppressed PGC-1α expression, cells were cultured in tunicamycin for 6 h and then treated with 4-PBA (2 mmol/L) for another 48 h.
Immunoblotting
To extract protein, the embryos or cells were sonicated in ice-cold lysis buffer (Cell Signaling Technology, Beverly, MA) with protease inhibitor cocktail (Sigma-Aldrich). The mitochondria were isolated from the embryos using the Pierce mitochondria isolation kit. Equal amounts of protein from different experimental groups were resolved by the SDS-PAGE gel electrophoresis and transferred onto Immunobilon-P membranes (Millipore). Membranes were incubated in 5% non-fat milk for 45 min and then were incubated for 18 h at 4 °C with the following primary antibodies in 5% non-fat milk: protein Bcl-2 homologous antagonist killer (Bak) (Cell Signaling Technologies, dilution 1:1000), Bax (Cell Signaling Technologies, dilution 1:1000), p53 upregulated modulator of apoptosis (Puma) (Cell Signaling Technologies, dilution 1:1000), Bim (Cell Signaling Technologies, dilution 1:1000), tBid (Cell Signaling Technologies, dilution 1:1000), phosphorylated Bcl-2-associated death promoter (pBad), caspase 3 (Millipore, dilution 1:1000), caspase 8 (Enzo Technologies, dilution 1:1000), CHOP (Cell Signaling Technologies, dilution 1:1000), PGC-1a (Santa Cruz, dilution 1:1000), and C/EBPβ (Cell Signaling Technologies, dilution 1:1000). Membranes were then exposed to goat anti-rabbit or anti-mouse secondary antibodies (KPL, dilution 1:10000). To confirm that equivalent amounts of protein were loaded among samples, membranes were stripped and probed with a mouse antibody against β-actin (Abcam, dilution 1:5000). Signals were detected using the SuperSignal West Femto Maximum Sensitivity Substrate kit (Thermo Scientific). All experiments were repeated 3 times with the use of independently prepared tissue or cell lysates.
Electron microscopy
Mitochondrial structures were examined by transmission electron microscopy (EM) in our university’s EM core facility. Thick sections (1 μm) were cut and visualized at 100 × magnification to identify the neuroepithelia of the E8.75 embryos. Thin sections (80 nm) of identified neuroepithelia were cut and viewed with an electron microscope (Joel JEM-1200EX; Tokyo, Japan) at high resolution (10, 12, and 25 K) to identify the cellular organelle structures.
Neural stem cell isolation and flow cytometry analysis of mitochondrial potential
E8.75 embryos were dissected out of the uterus and then yolk sacs were removed from the embryos. The whole embryos were digested by Dispase II (3 mg/ml in PBS pH = 7.4) for 15 min at room temperature in order to remove non-neural tissue (De Bellard et al., 2002; Etchevers, 2011). After digestion of Dispase II, the whole neural tubes were washed with PBS for 3 times, cut into pieces and digested with 0.25% trypsin-EDTA (Thermo Scientific) for 5 min at 37 °C with shaking. The cell suspensions were centrifuged at 500 × g for 5 min and resuspended in DMEM medium (Thermo Scientific) with 10%FBS (Thermo Scientific) following by filtration using 70 μm cell strainer. The isolated cells were seeded into 6-well plate. The mitochondrial potential was analyzed using JC-1 dye (Invitrogen). Briefly, after adhesion, the isolated neural stem cells were incubated with JC-1 dye-contained culture medium (1 µg/ml) for 15 min at 37 °C. After dye staining, the cells were harvested and washed with PBS for 3 times. The stained cells were analyzed by a flow cytometer.
PGC-1α promoter activity analysis
The PGC-1α promoter containing the transcription factor C/EBPβ binding site was subcloned and inserted into the pGL4.10 Luciferase Reporter Vectors (Promega). Transfection of pGL4.1-PGC-1α promoter vector into mouse neural stem cell line C17.2 was performed using Lipofectamine 2000 (Invitrogen) and then the cells were cultured for 48 h with or without high glucose (25 mM) combined with CHOP siRNA transfection or 6 h with or without tunicamycin (1 μg/ml) combined with 4-PBA (2 mM) for another 48 h. Cells were harvested and lysated using lysis buffer from the Dual luciferase Assay system (promega). The firefly luciferase activities were measured using the Dual luciferase Assay system (promega). The firefly luciferase activities were normalized to internal control Renilla luciferase.
Immunoprecipitation
Cells were lysed in the non-denaturing lysis buffer containing 20 mM of Tris-HCl (pH 7.5), 150 mM of NaCl, 1 mM of Na2EDTA, 1 mM of EGTA, 1% Triton, 2.5 mM of sodium pyrophosphate, 1 mM of β−glycerophosphate, 1 mM of Na3VO4, and 1 μg/ml of leupeptin (Cell Signaling technology). Protease inhibitor (Sigma) was added before use. After centrifugation at 12 000 × g for 10 min at 4 °C, the supernatant was pre-cleaned with protein A/G magnetic beads (Thermo Scientific) for 2 h at 4 °C. About 300 μg of protein extraction were incubated with an anti-CHOP primary antibody (Cell Signaling Technologies) at 4 °C overnight. Twenty-five microliters of protein A/G magnetic beads was added for immunoprecipitation at room temperature for 2 h. Precipitated complexes were cleansed in washing buffer (Thermo Scientific), and bound proteins were analyzed by immunoblotting.
Chromatin immunoprecipitation
Chromatin immunoprecipitation (ChIP) assay was assessed by the Magna CHIPTM and EZ-Magna CHIPTM kits (Millipore). In brief, fresh cells and embryos were crosslinked in 1% formaldehyde at room temperature for 10 min and then stopped by 1× Glycine. Cells and embryos were lysed by sonication to shear DNA on wet ice. Lysates were spun at 10 000 × g for 10 min at 4 °C to remove insoluble material. Each 25-µg DNA chromatin sample was adjusted to a total volume of 500 µl in 450 µl of the Dilution Buffer containing protease inhibitors. Chromatin samples were then incubated with anti-C/EBPβ (Η−7) X (Santa Cruz, CA) or anti-rabbit IgG antibody (Cell Signaling Technologies) and fully suspended in protein A/G magnetic beads overnight at 4 °C with gentle rotation. Magnetic beads were washed and immunoprecipitated DNAs were eluted with 100 µl CHIP Elution Buffer with Proteinase K at 62 °C for 2 h. DNAs were purified using the Spin Columns and DNAs were dissolved in the Elution Buffer C. Chromatin DNAs were analyzed by PCR and Real-time Quantitative PCR (qPCR) as previously described (Wang et al., 2008). Primers used for CHIP PCR and qPCR were as follows (designated from 5′ to 3′): position −2409 to −2398 forward-GGGGAACCCAAGAGTCT and reverse-CCCAAATCAGCTGTCTCCT; position −765 to −754 forward-GGGCTGCCTTGGAGTGACG and reverse-GTCCCCAGTCACATGACAAA; position −16 to −5 forward-CTGAGTCTGGGGCTACTTGG and reverse-TCCATCCAAAACAGGCAAAT.
Statistical analyses
All experiments were completely randomized designed and repeated in triplicate. Data are presented as means ± standard errors (SE). Student’s t-test was used for 2 group comparisons. One-way or two-way ANOVA was performed for more than 2 group comparisons using the SigmaStat 3.5 software. In ANOVA analysis, a Tukey test was used to estimate the significance. Significant differences between groups in NTD incidence expressed by number of embryos were analyzed by χ2 test or Fisher’s exact test using SigmaStat 3.5 software. Statistical significance was indicated when p < .05.
RESULTS
Chop Gene Deletion Ameliorates Apoptosis and NTDs in Diabetic Embryopathy
In the diabetic embryopathy model, the average blood glucose level was significantly higher in the diabetic group than that in the non-diabetic group (Figure 1A and Table 1). Maternal diabetes increased CHOP expression at the mRNA and protein levels, whereas Chop gene deletion completely abolished CHOP expression in the developing embryo (Figure 1B and C). To investigate the effect of Chop gene deletion on maternal diabetes-induced NTDs, a total of 157 embryos from 23 dams including 9 non-diabetic dams and 14 diabetic dams were harvested and examined for failed neural tube closure (Table 1). Under non-diabetic conditions, Chop gene deletion did not affect embryonic neurulation (Table 1 and Figure 1D and E). Under diabetic conditions, maternal diabetes significantly increased the NTD incidence to 32.0% in WT embryos (Table 1 and Figure 1D and E). However, Chop gene deletion significantly alleviated NTD formation with a NTD incidence of 7.7% in embryos from diabetic dams (Table 1 and Figure 1D and E). Heterozygous Chop deletion did not significantly reduced NTD incidence induced by maternal diabetes (Table 1 and Figure 1D and E).
Figure 1.
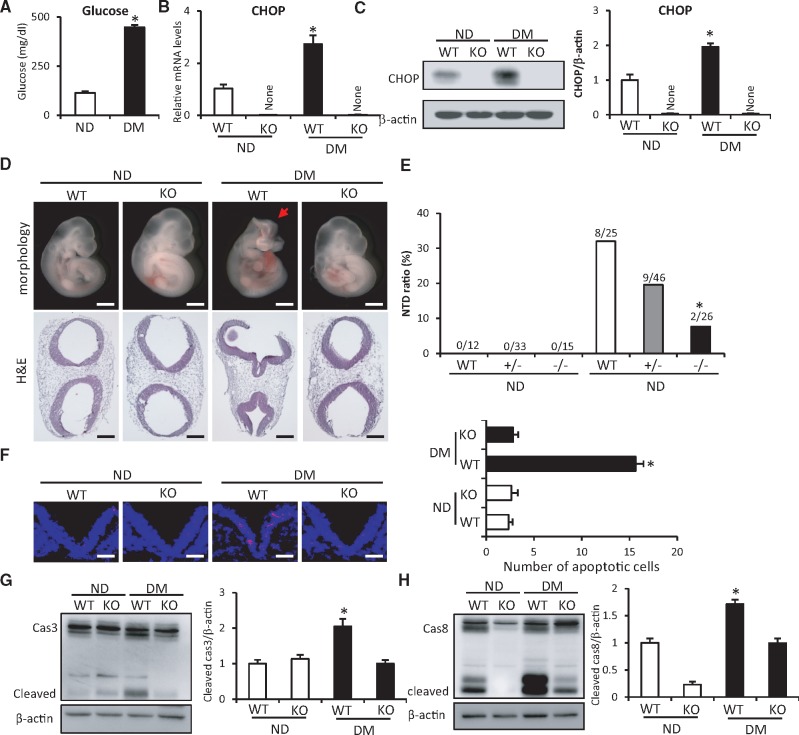
CHOP knockout ameliorates maternal diabetes-induced NTD formation and neuroepithelial cell apoptosis. A, The average blood glucose level in non-diabetic and diabetic groups. The mRNA abundance (B) and protein abundance of CHOP (C) in E8.5 embryos. D, Morphology and H&E staining images of E10.5 embryos. Arrow shows the NTD. Scale bars: 300 µm. E, NTD incidences of E10.5 embryos in each group. F, Representative images of the TUNEL assays in E8.5 embryos. Apoptotic cells were labeled and nuclei were labeled by DAPI. The dense blue V shape areas are the neural tubes. The bar graph showed the quantification of apoptotic cell number. Experiments were repeated using 3 embryos (N=3) from different dams and 3 images were obtained from each embryo. Scale bars: 70 µm. The protein abundance of cleaved caspase 3 (G) and caspase 8 (H) in E8.5 embryos. Bar graphs for protein abundance were quantitative data from 3 independent experiments. ND: non-diabetic dams; DM: diabetic dams; NTD: neural tube defect; KO: CHOP knockout. * indicate significant difference (p < .05) compared with other groups.
TABLE 1.
Targeted Deletion of the Chop Gene Ameliorates Maternal Diabetes-Induced Neural Tube Defects
| Groups | Number of NTD Embryos | Total Embryos | Blood Glucose Levels (mg/dl) | |
|---|---|---|---|---|
| Non-diabetic | WT | 0 | 12 | 114.2 ± 7.2 |
| (CHOP+/−♀ × CHOP+/−♂) | CHOP+/− | 0 | 33 | |
| n = 9 | CHOP−/− | 0 | 15 | |
| Diabetic | WT | 8 (32.0%) | 25 | 448.1 ± 11.6 |
| (CHOP+/−♀ × CHOP+/−♂) | CHOP+/− | 9 (19.6%) | 46 | |
| n = 14 | CHOP−/− | 2 (7.7%)* | 26 | |
indicates significant difference (p < .05) compared with the other groups in a Fisher Exact Test. NTD: neural tube defects; WT: Wild-type.
Neuroepithelial cell apoptosis is a causal factor for maternal diabetes-induced NTDs (Dong et al., 2016a, b; Gu et al., 2015, 2016; Li et al., 2012, 2013; Yang et al., 2013, 2015). Chop gene deletion significantly reduced the number of apoptotic cells in the developing neuroepithelium compared to that in the neuroepithelium of WT embryos under diabetic conditions (Figure 1F). Maternal diabetes triggered caspase 3 and caspase 8 cleavage in WT embryos, whereas Chop gene deletion abrogated maternal diabetes-induced caspase 3 and caspase 8 cleavage in neurulation stage embryos (Figure 1G and H). The findings collectively demonstrate that Chop gene deletion ameliorates diabetic embryopathy by suppressing neuroepithelial cell apoptosis.
Chop Gene Deletion Preserves Mitochondrial Function in Diabetic Embryopathy
Previous studies have shown that maternal diabetes causes mitochondrial defects such as disarrayed or disruptive cristae and decreased electronic density of the matrix (Wang et al., 2017; Xu et al., 2013) . Indeed, maternal diabetes significantly increased the number of defective mitochondria in neuroepithelial cells of WT embryos (Figure 2A). Chop gene deletion significantly reduced the number of defective mitochondria in neuroepithelial cells of embryos exposed to diabetes comparable to that in the non-diabetic group (Figure 2A). The mitochondrial membrane potential as an indicator of mitochondrial function was assessed by the JC-1 dye staining. Maternal diabetes triggers mitochondrial depolarization indicated by the increase of the green/red fluorescence intensity ratio (Figure 2B). Chop gene deletion abolished maternal diabetes-induced mitochondrial depolarization (Figure 2B).
Figure 2.
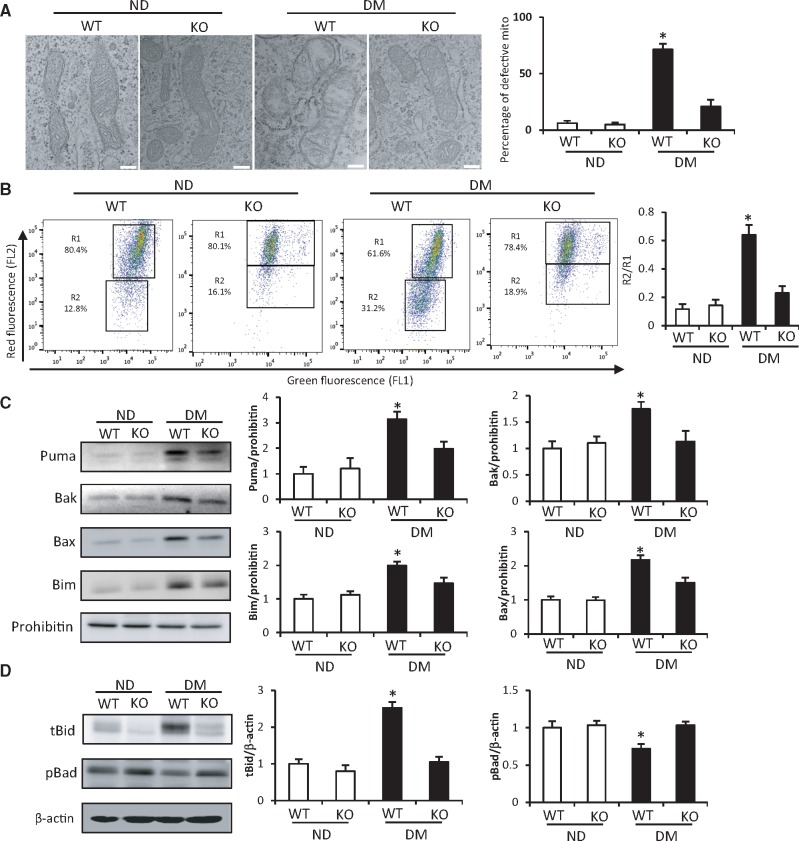
CHOP knockout restores maternal diabetes-disturbed mitochondrial function. A, Defective mitochondria (mito) rates (%). Defective mitochondria rate = the number of defective mitochondria divided by total number of mitochondria per image area (image size: 9.43 μm2) in neuroepithelial cells. Neuroepithelia from 3 embryos (N = 3) derived from different dams were used. Three serial sections per embryo were analyzed. Scale bars: 200 nm. B, JC-1 dye-stained isolated neuroepithelial cells were analyzed for the mitochondrial potential by a flow cytometer. Bar graphs for rate of fluorescence were quantitative data from 3 independent experiments. R1 indicates the membrane intact mitochondria. R2 indicates the membrane detective mitochondrial. C, The protein abundance of Puma, Bak, Bax, and Bim in isolated mitochondria from E8.5 embryos. D, The protein abundance of tBid (cleaved), pBad (phosphorylated) in E8.5 embryos. Bar graphs for protein abundance were quantitative data from 3 independent experiments. *indicate significant difference compared with other groups. ND: non-diabetic dams; DM: diabetic dams; KO: CHOP knockout. * indicate significant difference (p < .05) compared with other groups.
Furthermore, Chop gene deletion suppressed maternal diabetes-induced mitochondrial translocation of several pro-apoptotic Bcl-2 members including Bax, Puma, Bak, and Bim (Figure 2C). The active form of Bid, cleaved Bid, was significantly increased, whereas the inactive form of Bad, phosphorylated Bad, was significantly decreased in embryos from diabetic dams compared to those in embryos from non-diabetic dams (Figure 2D). Chop gene deletion reversed maternal diabetes-induced Bid cleavage and Bad dephosphorylation in the developing embryo (Figure 2D). These findings support the hypothesis that Chop gene deletion restores mitochondrial function in embryos exposed to maternal diabetes.
High Glucose Inhibits PGC-1α Expression Through CHOP
PGC-1α is a master regulator of mitochondrial biogenesis and respiration (Austin and St-Pierre, 2012; Scarpulla, 2011). Tissue from Ppargc1a-null mice display reduced expression of mitochondrial genes, decreased mitochondrial respiration and impaired mitochondrial function (Arany et al., 2005; Lin et al., 2004). Maternal diabetes suppressed PGC-1α expression, whereas Chop gene deletion restored PGC-1α expression in embryos exposed to maternal diabetes (Figure 3A). High glucose (25 mM) in vitro significantly increased CHOP expression (Figure 3B) and decreased PGC-1α expression (Figure 3C) compared to the normal glucose group (5 mM). CHOP siRNA knockdown repressed high glucose-increased CHOP expression (Figure 3B and D) and abrogated high glucose-inhibited PGC-1α expression (Figure 3C and D), indicating maternal diabetes in vivo- or high glucose in vitro-increased CHOP inhibits PGC-1α expression. CHOP overexpression mimicked high glucose in suppressing PGC-1α expression at both mRNA and protein levels (Figure 3E and F).
Figure 3.
Maternal diabetes or high glucose-increased CHOP inhibits PGC-1α expression. A, The mRNA abundance of PGC-1α in E8.5 embryos. B, The mRNA abundance of CHOP in C17.2 cells transfected with or without CHOP siRNA under normal or high glucose conditions. C, The mRNA abundance of PGC-1α in C17.2 cells transfected with or without CHOP siRNA under normal or high glucose conditions. D, The protein abundance of PGC-1α and CHOP in C17.2 cells transfected with or without CHOP siRNA under normal or high glucose conditions. Bar graphs for protein abundance were quantitative data from 3 independent experiments. E, The mRNA abundance of PGC-1α and CHOP in C17.2 cells transfected with or without CHOP expression vector under normal or high glucose conditions. F, The protein abundance of PGC-1α and CHOP in C17.2 cells transfected with or without CHOP expression vector under normal or high glucose conditions. Bar graphs for protein abundance were quantitative data from 3 independent experiments. G, PGC-1α promoter activity analyzed by relative luciferase reporter activities in C17.2 cells. Experiments were repeated 3 times. ND: non-diabetic dams; DM: diabetic dams; KO: CHOP knockout; NG: normal glucose; HG: high glucose. Control: blank pcDNA3 vector. *indicate significant difference (p < .05) compared with other groups.
High glucose significantly decreased PGC-1α promoter activity and CHOP siRNA knockdown restored PGC-1α promoter activity (Figure 3G). CHOP overexpression mimicked the inhibitory effect of high glucose on PGC-1α promoter activity (Figure 3G). Thus, maternal diabetes in vivo or high glucose in vitro inhibits PGC-1α expression through CHOP at the transcriptional level.
The ER Stress Inducer Tunicamycin Represses PGC-1α Through CHOP
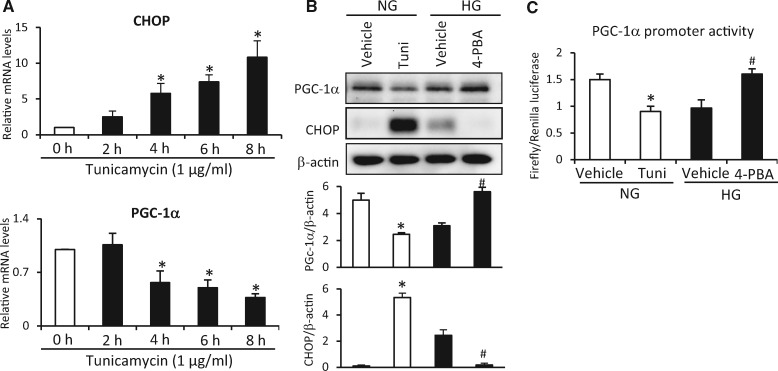
To further elucidate the effect of CHOP on PGC-1α expression, the ER stress inducer tunicamycin was used to increase CHOP expression (Wang et al., 2015b). Tunicamycin robustly increased CHOP expression along with the decrease of PGC-1α mRNA and protein expression in a dose-dependent manner (Figure 4A and B). The ER stress inhibitor 4-PBA blocked high glucose-increased CHOP expression and reversed high glucose-suppressed PGC-1α expression (Figure 4B). Tunicamycin repressed PGC-1α promoter activity and 4-PBA abolished high glucose-suppressed PGC-1α promoter activity (Figure 4C). These data indicate that ER stress-induced CHOP negatively regulates PGC-1α expression.
Figure 4.
ER stress inducer Tunicamycin-increased CHOP inhibits PGC-1α expression. A, The mRNA abundance of CHOP and PGC-1α in C17.2 cells treated with tunicamycin under different time. B, The protein abundance of CHOP and PGC-1α in C17.2 cells treated with tunicamycin or high glucose combined with 4-PBA. Bar graphs for protein abundance were quantitative data from 3 independent experiments. C, PGC-1α promoter activity analyzed by relative luciferase reporter activities in C17.2 cells treated with tunicamycin or high glucose combined with 4-PBA. Experiments were repeated 3 times. NG: normal glucose; HG: high glucose. *indicate significant difference (p < .05) compared with 0 h group in A and NG-Vehicle group in B and C. # indicate significant difference (p < .05) compared with HG-vehicle group in B and C.
C/EBPβ Abolishes High Glucose- and Tunicamycin-Suppressed PGC-1α Expression
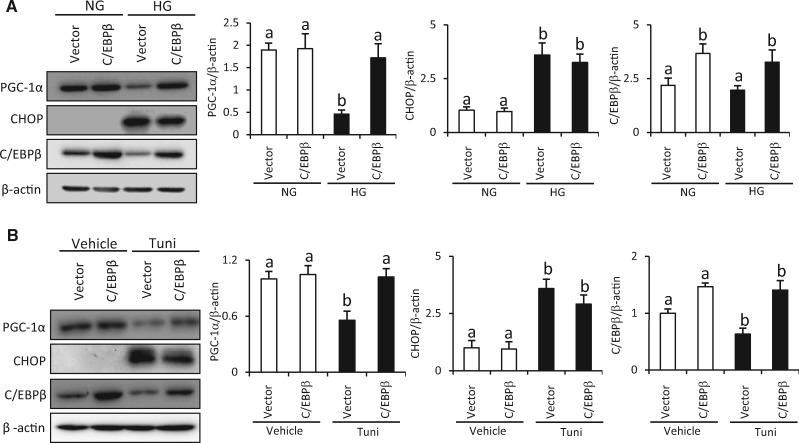
It has been shown that CCAAT/enhancer binding protein beta (C/EBPβ) is a transcriptional regulator of PGC-1α expression in adipocytes and hepatocytes (Karamanlidis et al., 2007; Karamitri et al., 2009; Wang et al., 2008). Chromatin immunoprecipitation reveals that C/EBPβ binds to the cAMP response element of the PGC-1α promoter and positively regulates PGC-1α expression (Karamitri et al., 2009). To investigate the potential role of C/EBPβ on the CHOP-suppressed PGC-1α expression, ectopic C/EBPβ overexpression was applied. C/EBPβ overexpression did not affect high glucose-increased CHOP expression, whereas it reversed high glucose-inhibited PGC-1α expression (Figure 5A). These findings were confirmed in tunicamycin treatment. Tunicamycin treatment increased CHOP expression and decreased PGC-1α expression (Figure 5B). C/EBPβ overexpression did not change the stimulatory effect of tunicamycin on CHOP expression (Figure 5B). Nevertheless, C/EBPβ overexpression abolished tunicamycin-induced PGC-1α suppression (Figure 5B).
Figure 5.
C/EBPβ overexpression restores high glucose/tunicamycin-suppressed PGC-1α expression. A, The protein abundance of PGC-1α, CHOP, and C/EBPβ in C17.2 cells transfected with C/EBPβ expression vector under normal or high glucose conditions. B, The protein abundance of PGC-1α, CHOP, and C/EBPβ in C17.2 cells transfected with C/EBPβ expression vector and treated with tunicamycin (1 μg/ml). Bar graphs for protein abundance were quantitative data from 3 independent experiments. NG: normal glucose; HG: high glucose; Tuni: tunicamycin. Different letter indicates significant difference (p < .05).
CHOP Inhibits C/EBPβ DNA-Binding Activity by Forming CHOP-C/EBPβ Heterodimers
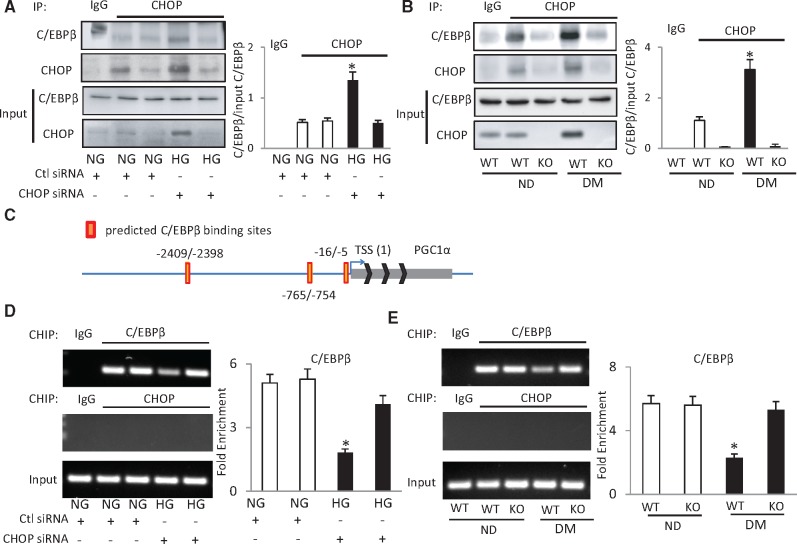
Because C/EBPβ overexpression did not affect CHOP expression, we hypothesized that high glucose- or maternal diabetes-increased CHOP inhibits C/EBPβ DNA-binding activity through forming the CHOP-C/EBPβ complex. Co-immunoprecipitation showed that high glucose in vitro and maternal diabetes in vivo enhanced the association between CHOP and C/EBPβ (Figure 6A and B 6Α,). Τo investigate whether increased CHOP blocks the binding ability of C/EBPβ, C/EBPβ consensus sites were identified in the PGC-1α promoter. There are 3 potential C/EBPβ binding sites in the PGC-1α promoter at nucleotide positions −2409 to −2398, −765 to −754, and −16 to −5 (Figure 6C). ChIP assays were performed to evaluate C/EBPβ binding to the PGC-1α promoter using primers spanning these 3 putative sites and demonstrated that C/EBPβ only bound to the PGC-1α promoter at the position −765 to −754 (Figure 6D and E). However, as a member of the C/EBP family, CHOP did not bind to the PGC-1α promoter (Figure 6D and E). Both high glucose and maternal diabetes decreased the binding of C/EBPβ to the promoter of PGC-1α, whereas CHOP siRNA knockdown in C17.2 cells under high glucose conditions and Chop gene deletion in embryos of diabetic dams restored the binding activity of C/EBPβ to the PGC-1α promoter (Figure 6D and E). Thus, elevated CHOP competitively interacts with C/EBPβ in preventing its binding to the PGC-1α promoter leading to inhibition PGC-1α expression in diabetic embryopathy.
Figure 6.
CHOP inhibits C/EBPβ DNA-binding activity by forming CHOP-C/EBPβ heterodimers. A, Representative images of Co-immunoprecipitation using an anti-CHOP antibody in C17.2 cells transfected with CHOP siRNA or scramble control siRNA under normal or high glucose conditions. B, Representative images of Co-immunoprecipitation using an anti-CHOP antibody in E8.5 embryos. In the bar graphs, the protein level of C/EBPβ was assessed in CHOP immunoprecipitates and was normalized by 10% input. Normal rabbit IgG was used as control. C, Potential binding sites for C/EBPβ in the PGC-1α promoter. D, Immunoprecipitated chromatin was analyzed by PCR for the C/EBPβ site and quantification of C/EBPβ occupancy on the C/EBPβ site in C17.2 cells transfected with CHOP siRNA or scramble control siRNA under normal or high glucose conditions. E, Immunoprecipitated chromatin was analyzed by PCR for the C/EBPβ site and quantification of C/EBPβ occupancy on the C/EBPβ site in E8.5 embryos. ND: non-diabetic dams; DM: diabetic dams; NG: normal glucose; HG: high glucose; KO: CHOP knockout. *indicate significant difference (p < .05) compared with other groups.
DISCUSSION
The purpose of this study was to investigate whether CHOP contributes to ER stress-induced neuroepithelial cell apoptosis in diabetic embryopathy, and to further identify the molecular mechanism underlying CHOP-mediated apoptosis in diabetic embryopathy. The results of the present study demonstrate that maternal diabetes-induced neuroepithelial cell apoptosis can be attributed to increased CHOP expression. Previous studies have shown that CHOP is involved in the mitochondria-dependent apoptotic pathway by suppressing the expression of 2 major anti-apoptotic proteins Bcl-2 and Mcl-1, and up-regulating the pro-apoptotic protein Bim (Iurlaro and Munoz-Pinedo, 2016). In addition, CHOP interacts with c-Jun and GCN5 to increase cell death receptor DR4 and DR5 expression leading to a widespread cleavage of caspases and cell death (Iurlaro and Munoz-Pinedo, 2016). In the present study, CHOP is responsible for maternal diabetes-impaired mitochondrial function and caspase 3/8 cleavage. Thus, in diabetic embryopathy, ER stress-induced and CHOP-mediated neuroepithelial cell apoptosis occurs through the mitochondrial apoptotic pathway.
To elucidate how CHOP induces the mitochondrial dysfunction pathway of cell death, the present study focuses on the master regulator of mitochondrial biogenesis and respiration PGC-1α. C/EBPβ is a positive regulator for PGC-1α, whereas CHOP antagonizes the transcriptional activity of C/EBPβ (Chikka et al., 2013; Karamitri et al., 2009; Ron and Habener, 1992; Tang and Lane, 2000). The current findings reveal that maternal diabetes- or high glucose-increased CHOP reduces PGC-1α expression, and deletion or knocking down of CHOP restores PGC-1α expression. In contrast, C/EBPβ overexpression reverses the effect of maternal diabetes or high glucose on PGC-1α expression. Previous studies suggest that both CHOP and C/EBPβ can bind together to the C/EBP-binding site in the promoters of target genes (Chikka et al., 2013; Karamitri et al., 2009). However, CHOP cannot bind to the PGC-1α promoter directly in fat tumor cells (Karamitri et al., 2009). In fact, CHOP lacks the DNA-binding activity for the C/EBP-binding consensus sequence in promoter regions because of 2 proline residues in its DNA-binding domain, and a unique DNA sequence termed as the CHOP-binding site is required for CHOP to bind to a specific promoter (Ohoka et al., 2007; Tang and Lane, 2000). The findings in this present study confirmed that CHOP could not directly bind to the PGC-1α promoter directly. Additionally, CHOP cannot form homodimers, but it inhibits the DNA binding activity of other C/EBP proteins such as C/EBPβ and exerts transcriptional regulatory activity through forming heterodimers with other C/EBP proteins (Ron and Habener, 1992). In this study, the direct interaction between CHOP and C/EBPβ is identified and enhanced by ER stress induced by high glucose. Chop gene deletion or knocking down CHOP restores high glucose-suppressed C/EBPβ binding to the PGC-1α promoter. Thus, CHOP suppresses PGC-1α expression by inhibiting the DNA-binding ability of C/EBPβ through forming CHOP-C/EBPβ heterodimers, ultimately leading to mitochondrial dysfunction.
The molecular mechanism underlying maternal diabetes or high glucose enhances the CHOP-C/EBPβ heterodimer formation still remains to be elucidated. It is possible that high glucose dramatically induces CHOP expression leading to the increased formation of CHOP-C/EBPβ heterodimers. In addition, modification of a transcription factor, such as phosphorylation, glycosylation and acetylation, may affect its DNA binding activity and function. Indeed, glycosylation of C/EBPβ prevents its phosphorylation and DNA binding activity (Li et al., 2009). Glycosylation of C/EBPβ may enhance CHOP-C/EBPβ heterodimers formation in inhibiting C/EBPβ DNA transcription activity.
Previous studies have demonstrated that PGC-1α activates autophagy both in vivo and in vitro (Wang et al., 2017), and PGC-1α overexpression in the developing neuroepithelium restores maternal diabetes-suppressed autophagy and alleviates NTDs in diabetic pregnancy (Wang et al., 2017). In the present study, we demonstrated that Chop gene deletion ameliorated maternal diabetes-induced NTD formation and CHOP is the negative regulator for PGC-1α expression at the transcriptional level. The previous study also reveals that miR-129-2 mediates the inhibitory effect of high glucose on PGC-1α expression (Wang et al., 2017). Thus, maternal diabetes represses PGC-1α expression at both transcriptional and post-transcriptional levels.
Because PGC-1α is an autophagy activator (Wang et al., 2017), CHOP may also contribute to maternal diabetes-induced neuroepithelial cell apoptosis and NTD formation through the autophagy pathway. ER stress-increased CHOP under maternal diabetic conditions suppresses PGC-1α expression, and, thus, PGC-1α-induced autophagy is inhibited and leads to accumulation of damaged mitochondria which causes mitochondrial dysfunction and the mitochondrial-dependent apoptosis in neuroepithelial cells. Recent studies have shown that CHOP has a dual role in both inducing apoptosis and limiting autophagy (Senft and Ronai, 2015). As a result, CHOP may induce neuroepithelial cell apoptosis through inducing mitochondrial dysfunction and inhibiting autophagy.
Consistent with early studies showing that either Chop gene deletion or CHOP deficiency alleviates renal fibrosis in kidney diseases (Liu et al., 2016), myocardial reperfusion injury and cardiac dysfunction induced by pressure overload (Miyazaki et al., 2011), the present study reveals that Chop gene deletion inhibits diabetic embryopathy. These studies suggest that CHOP and its downstream signals may be potential therapeutic targets for ER stress-induced human pathogenesis. In summary, this study demonstrated that maternal diabetes-increased CHOP contributes to ER stress-induced neuroepithelial cell death through the mitochondrial-dependent apoptosis pathway.
FUNDING
NIH (R01DK083243, R01DK101972, R01HL131737, and R01DK103024).
CONFLICT OF INTEREST
None of the authors has a conflict of interest.
REFERENCES
- Arany Z., He H., Lin J., Hoyer K., Handschin C., Toka O., Ahmad F., Matsui T., Chin S., Wu P. H., et al. (2005). Transcriptional coactivator PGC-1 alpha controls the energy state and contractile function of cardiac muscle. Cell Metab. 1, 259–271. [DOI] [PubMed] [Google Scholar]
- Austin S., St-Pierre J. (2012). PGC1alpha and mitochondrial metabolism–emerging concepts and relevance in ageing and neurodegenerative disorders. J. Cell Sci. 125, 4963–4971. [DOI] [PubMed] [Google Scholar]
- Chikka M. R., McCabe D. D., Tyra H. M., Rutkowski D. T. (2013). C/EBP homologous protein (CHOP) contributes to suppression of metabolic genes during endoplasmic reticulum stress in the liver. J. Biol. Chem. 288, 4405–4415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Bellard M. E., Ching W., Gossler A., Bronner-Fraser M. (2002). Disruption of segmental neural crest migration and ephrin expression in delta-1 null mice. Dev. Biol. 249, 121–130. [DOI] [PubMed] [Google Scholar]
- Dong D., Fu N., Yang P. (2016a). MiR-17 downregulation by high glucose stabilizes thioredoxin-interacting protein and removes thioredoxin inhibition on ASK1 leading to apoptosis. Toxicol. Sci. 150, 84–96. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dong D., Reece E. A., Lin X., Wu Y., AriasVillela N., Yang P. (2016b). New development of the yolk sac theory in diabetic embryopathy: Molecular mechanism and link to structural birth defects. Am. J. Obstet. Gynecol. 214, 192–202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Etchevers H. (2011). Primary culture of chick, mouse or human neural crest cells. Nat. Protoc. 6, 1568–1577. [DOI] [PubMed] [Google Scholar]
- Gabbay-Benziv R., Reece E. A., Wang F., Yang P. (2015). Birth defects in pregestational diabetes: Defect range, glycemic threshold and pathogenesis. World J. Diabetes 6, 481–488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gu H., Yu J., Dong D., Zhou Q., Wang J. Y., Fang S., Yang P. (2016). High glucose-repressed CITED2 expression through miR-200b triggers the unfolded protein response and endoplasmic reticulum stress. Diabetes 65, 149–163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gu H., Yu J., Dong D., Zhou Q., Wang J. Y., Yang P. (2015). The miR-322-TRAF3 circuit mediates the pro-apoptotic effect of high glucose on neural stem cells. Toxicol. Sci. 144, 186–196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iurlaro R., Munoz-Pinedo C. (2016). Cell death induced by endoplasmic reticulum stress. FEBS J. 283, 2640–2652. [DOI] [PubMed] [Google Scholar]
- Jin J., Iakova P., Jiang Y., Lewis K., Sullivan E., Jawanmardi N., Donehower L., Timchenko L., Timchenko N. A. (2013). Transcriptional and translational regulation of C/EBPbeta-HDAC1 protein complexes controls different levels of p53, SIRT1, and PGC1alpha proteins at the early and late stages of liver cancer. J. Biol. Chem. 288, 14451–14462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamimoto Y., Sugiyama T., Kihira T., Zhang L., Murabayashi N., Umekawa T., Nagao K., Ma N., Toyoda N., Yodoi J., et al. (2010). Transgenic mice overproducing human thioredoxin-1, an antioxidative and anti-apoptotic protein, prevents diabetic embryopathy. Diabetologia 53, 2046–2055. [DOI] [PubMed] [Google Scholar]
- Karamanlidis G., Karamitri A., Docherty K., Hazlerigg D. G., Lomax M. A. (2007). C/EBPbeta reprograms white 3T3-L1 preadipocytes to a Brown adipocyte pattern of gene expression. J. Biol. Chem. 282, 24660–24669. [DOI] [PubMed] [Google Scholar]
- Karamitri A., Shore A. M., Docherty K., Speakman J. R., Lomax M. A. (2009). Combinatorial transcription factor regulation of the cyclic AMP-response element on the Pgc-1alpha promoter in white 3T3-L1 and brown HIB-1B preadipocytes. J. Biol. Chem. 284, 20738–20752. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li X., Molina H., Huang H., Zhang Y. Y., Liu M., Qian S. W., Slawson C., Dias W. B., Pandey A., Hart G. W., et al. (2009). O-linked N-acetylglucosamine modification on CCAAT enhancer-binding protein beta: Role during adipocyte differentiation. J. Biol. Chem. 284, 19248–19254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li X., Weng H., Xu C., Reece E. A., Yang P. (2012). Oxidative stress-induced JNK1/2 activation triggers proapoptotic signaling and apoptosis that leads to diabetic embryopathy. Diabetes 61, 2084–2092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li X., Xu C., Yang P. (2013). c-Jun NH2-terminal kinase ½ and endoplasmic reticulum stress as interdependent and reciprocal causation in diabetic embryopathy. Diabetes 62, 599–608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin J., Wu P. H., Tarr P. T., Lindenberg K. S., St-Pierre J., Zhang C. Y., Mootha V. K., Jager S., Vianna C. R., Reznick R. M., et al. (2004). Defects in adaptive energy metabolism with CNS-linked hyperactivity in PGC-1alpha null mice. Cell 119, 121–135. [DOI] [PubMed] [Google Scholar]
- Liu S. H., Wu C. T., Huang K. H., Wang C. C., Guan S. S., Chen L. P., Chiang C. K. (2016). C/EBP homologous protein (CHOP) deficiency ameliorates renal fibrosis in unilateral ureteral obstructive kidney disease. Oncotarget 7, 21900–21912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyazaki Y., Kaikita K., Endo M., Horio E., Miura M., Tsujita K., Hokimoto S., Yamamuro M., Iwawaki T., Gotoh T., et al. (2011). C/EBP homologous protein deficiency attenuates myocardial reperfusion injury by inhibiting myocardial apoptosis and inflammation. Arterioscler. Thromb. Vasc. Biol. 31, 1124–1132. [DOI] [PubMed] [Google Scholar]
- Ohoka N., Hattori T., Kitagawa M., Onozaki K., Hayashi H. (2007). Critical and functional regulation of CHOP (C/EBP homologous protein) through the N-terminal portion. J. Biol. Chem. 282, 35687–35694. [DOI] [PubMed] [Google Scholar]
- Ramji D. P., Foka P. (2002). CCAAT/enhancer-binding proteins: structure, function and regulation. Biochem. J. 365, 561–575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ron D., Habener J. F. (1992). CHOP, a novel developmentally regulated nuclear protein that dimerizes with transcription factors C/EBP and LAP and functions as a dominant-negative inhibitor of gene transcription. Genes Dev. 6, 439–453. [DOI] [PubMed] [Google Scholar]
- Salbaum J. M., Kappen C. (2010). Neural tube defect genes and maternal diabetes during pregnancy. Birth Defects Res. A Clin. Mol. Teratol. 88, 601–611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sano R., Reed J. C. (2013). ER stress-induced cell death mechanisms. Biochim. Biophys. Acta 1833, 3460–3470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scarpulla R. C. (2011). Metabolic control of mitochondrial biogenesis through the PGC-1 family regulatory network. Biochim. Biophys. Acta 1813, 1269–1278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Senft D., Ronai Z. A. (2015). UPR, autophagy, and mitochondria crosstalk underlies the ER stress response. Trends Biochem. Sci. 40, 141–148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Snyder E. Y., Deitcher D. L., Walsh C., Arnold-Aldea S., Hartwieg E. A., Cepko C. L. (1992). Multipotent neural cell lines can engraft and participate in development of mouse cerebellum. Cell 68, 33–51. [DOI] [PubMed] [Google Scholar]
- Sugimura Y., Murase T., Oyama K., Uchida A., Sato N., Hayasaka S., Kano Y., Takagishi Y., Hayashi Y., Oiso Y., et al. (2009). Prevention of neural tube defects by loss of function of inducible nitric oxide synthase in fetuses of a mouse model of streptozotocin-induced diabetes. Diabetologia 52, 962–971. [DOI] [PubMed] [Google Scholar]
- Tang Q. Q., Lane M. D. (2000). Role of C/EBP homologous protein (CHOP-10) in the programmed activation of CCAAT/enhancer-binding protein-beta during adipogenesis. Proc. Natl. Acad. Sci. U. S. A. 97, 12446–12450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang F., Reece E. A., Yang P. (2013). Superoxide dismutase 1 overexpression in mice abolishes maternal diabetes-induced endoplasmic reticulum stress in diabetic embryopathy. Am. J. Obstet. Gynecol. 209, 345.e1–345.e7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang F., Reece E. A., Yang P. (2015a). Advances in revealing the molecular targets downstream of oxidative stress-induced proapoptotic kinase signaling in diabetic embryopathy. Am. J. Obstet. Gynecol. 213, 125–134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang F., Wu Y., Gu H., Reece E. A., Fang S., Gabbay-Benziv R., Aberdeen G., Yang P. (2015b). Ask1 gene deletion blocks maternal diabetes-induced endoplasmic reticulum stress in the developing embryo by disrupting the unfolded protein response signalosome. Diabetes 64, 973–988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang F., Xu C., Reece E. A., Li X., Wu Y., Harman C., Yu J., Dong D., Wang C., Yang P., et al. (2017). Protein kinase C-alpha suppresses autophagy and induces neural tube defects by stimulating miR-129-2 that targets PGC-1α in diabetic pregnancy. Nat. Commun. 8, 15182. doi: 10.1038/ncomms15182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang H., Peiris T. H., Mowery A., Le Lay J., Gao Y., Greenbaum L. E. (2008). CCAAT/enhancer binding protein-beta is a transcriptional regulator of peroxisome-proliferator-activated receptor-gamma coactivator-1alpha in the regenerating liver. Mol. Endocrinol. 22, 1596–1605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu Y., Wang F., Reece E. A., Yang P. (2015). Curcumin ameliorates high glucose-induced neural tube defects by suppressing cellular stress and apoptosis. Am. J. Obstet. Gynecol. 212, 802.e1–802.e8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu C., Li X. Z., Wang F., Weng H. B., Yang P. X. (2013). Trehalose prevents neural tube defects by correcting maternal diabetes-suppressed autophagy and neurogenesis. Am. J. Physiol.Endocrinol. Metab. 305, E667–E678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang P., Li X., Xu C., Eckert R. L., Reece E. A., Zielke H. R., Wang F. (2013). Maternal hyperglycemia activates an ASK1-FoxO3a-caspase 8 pathway that leads to embryonic neural tube defects. Sci. Signal. 6, ra74.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang P., Reece E. A., Wang F., Gabbay-Benziv R. (2015). Decoding the oxidative stress hypothesis in diabetic embryopathy through proapoptotic kinase signaling. Am. J. Obstet. Gynecol. 212, 569–579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhong J., Xu C., Gabbay-Benziv R., Lin X., Yang P. (2016a). Superoxide dismutase 2 overexpression alleviates maternal diabetes-induced neural tube defects, restores mitochondrial function and suppresses cellular stress in diabetic embryopathy. Free Radic. Biol. Med. 96, 234–244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhong J., Xu C., Reece E. A., Yang P. (2016b). The green tea polyphenol EGCG alleviates maternal diabetes-induced neural tube defects by inhibiting DNA hypermethylation. Am. J. Obstet. Gynecol. 215, 368.e1–368.e10. [DOI] [PMC free article] [PubMed] [Google Scholar]