Abstract
Genetic variants conferring risk for autism spectrum disorder (ASD) have been identified, but the role of post-transcriptional mechanisms in ASD is not well understood. We performed genome-wide microRNA (miRNA) expression profiling in post-mortem brains from ASD patients and controls, and identified miRNAs and co-regulated modules perturbed in ASD. Putative targets of these ASD-affected miRNAs were enriched for genes previously implicated in ASD risk. We confirmed causal regulatory relationships between several miRNAs and their putative target mRNAs in primary human neural progenitors. These include hsa-miR-21-3p, a miRNA up-regulated in ASD of unknown CNS function, which targets neuronal genes down-regulated in ASD, and a novel, primate-specific miRNA down-regulated in ASD, hsa_can_1002-m, which regulates the EGFR and FGFR signaling pathways involved in neural development and immune function. Our findings support a role for miRNA dysregulation in ASD pathophysiology and provide a rich dataset and framework for future analyses of miRNAs in neuropsychiatric diseases.
INTRODUCTION
Autism spectrum disorder (ASD) is a group of clinically heterogeneous neurodevelopmental disorders with deficits in social functioning and the presence of repetitive and restricted behaviors or interests1,2. ASD also manifests significant genetic heterogeneity; hundreds of genomic loci are implicated1,2. Dozens of rare Mendelian disorders, including fragile X syndrome, neurofibromatosis, Rett syndrome, tuberous sclerosis complex, and structural chromosomal variants, confer a high risk for ASD1,2. Recent studies have also revealed the contribution of rare, de novo single nucleotide mutations, none of which account for more than a small fraction of ASD cases1,2. Thousands of common variants are also estimated to contribute to ASD, although the effect size of individual loci is small1–4. Despite this remarkable heterogeneity, ASD-associated mutations have been suggested to target a few convergent biological processes, including synaptic function and neuronal activity, postsynaptic density protein metabolism, neuronal cell adhesion, WNT signaling, and chromatin remodeling during neurogenesis1,2.
In contrast, much less is known about the contribution of post-transcriptional regulatory mechanisms to ASD. MicroRNAs (miRNAs), which are small non-coding regulatory RNAs mediating mRNA destabilization and/or translational repression5, represent a sparsely studied class of putative contributors to ASD pathophysiology. Each miRNA can target up to hundreds of genes; collectively miRNAs are predicted to target > 60% of the transcriptome, establishing them as potential regulators of complex gene networks5,6. miRNAs have been shown to regulate processes pivotal to brain development and function, including neurogenesis, neuronal maturation, and synaptic plasticity7. To assess the potential role of miRNAs in ASD, we performed genome-wide miRNA expression profiling in post-mortem brains from ASD patients and controls. We found a shared pattern of miRNA dysregulation in a majority of ASD brains. The targets of ASD-affected miRNAs were enriched in ASD risk genes. Using bioinformatic and gene network analyses we were able to link these perturbations with transcriptomic changes in ASD brain.
RESULTS
Differential expression of miRNAs in ASD brain
We profiled miRNAs in 242 post-mortem brain tissue samples from 55 ASD cases and 42 controls (CTL) (Fig. 1a and Supplementary Table 1) using Illumina small RNA sequencing (sRNA-Seq; Methods). Up to three brain regions from each individual were assessed: frontal cortex (FC, Brodmann area [BA] 9), temporal cortex (TC, BA41/42/22), and cerebellar vermis (Fig. 1a), all of which have been implicated in ASD8. After quality control (Supplementary Fig. 1a–1b; Methods), mature miRNAs documented in miRBase release 20 (www.mirbase.org) were identified and quantified using the miRDeep2 algorithm9 (Methods). We also included in our analysis novel miRNAs identified in a recent study based on 94 human sRNA-Seq datasets and supported by experimental evidence10, as well as high-confidence novel miRNAs predicted from our sequencing data using two different methods (Methods; Supplementary Table 2). Six hundred and ninety-nine miRNAs (552 in miRBase 20 and 147 novel) were detected (Methods; Supplementary Table 2).
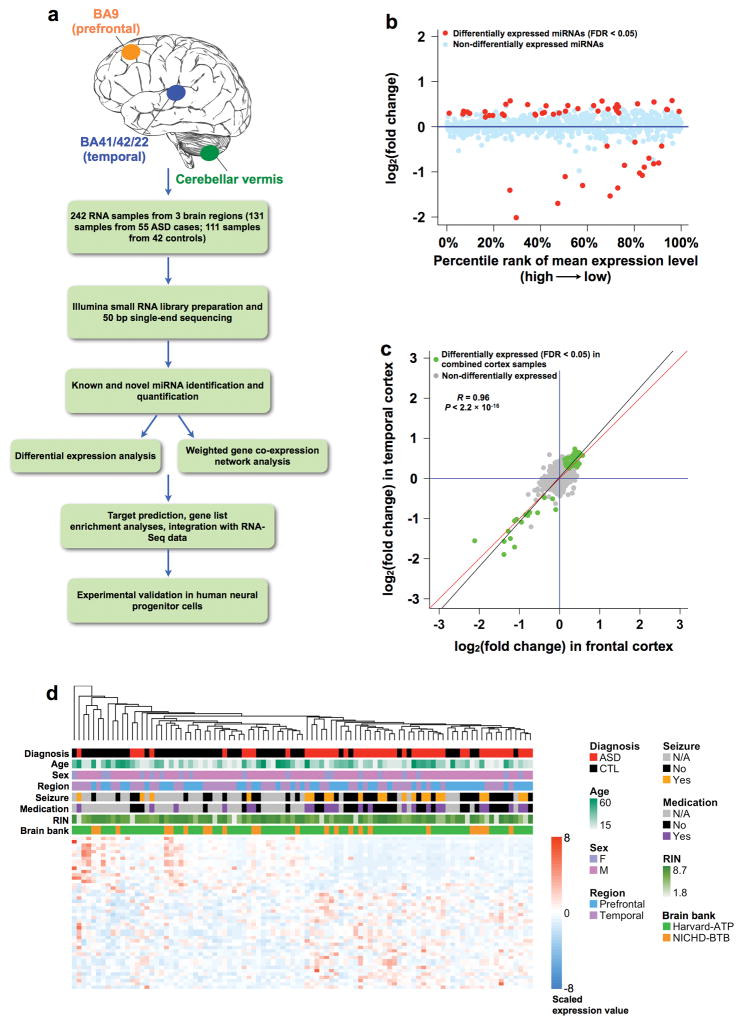
Figure 1. miRNA expression changes in post-mortem ASD cortex.
(a) Flow chart of the overall approach. (b) miRNA expression fold changes (> 0 if higher in ASD, < 0 if lower in ASD) between ASD and control cerebral cortex, plotted against the percentile rank of mean expression levels across 95 cortex samples (47 samples from 28 ASD cases and 48 samples from 28 controls) used for DGE analysis. Differentially expressed (linear mixed-effects model, FDR < 0.05) miRNAs are highlighted in red. (c) Comparison of miRNA expression fold changes in the temporal and the frontal cortex. Green dots, 58 miRNAs differentially expressed (FDR < 0.05) in 95 combined cortex samples; grey dots, non-differentially expressed miRNAs; black line, regression line between fold changes in the temporal and the frontal cortex for the differentially expressed miRNAs; red line, y=x. The Pearson correlation coefficient (R) and P value are also shown. (d) Dendrogram showing hierarchical clustering of 95 cortex samples based on top differentially expressed (FDR < 0.05, |log2(fold change)| ≥ 0.3) miRNAs. Information on diagnosis, age, sex, brain region, co-morbidity of seizures, psychiatric medication, RIN, and brain bank is indicated with color bars below the dendrogram according to the legend on the right. Heatmap on the bottom shows scaled (mean extracted and divided by standard deviation) expression values (color-coded according to the legend on the right) for miRNAs used for clustering.
The miRNA expression profiles were very similar between the frontal and temporal cortex, but distinct in the cerebellum (Supplementary Fig. 2d–2f), consistent with previous observations for mRNAs11,12. We therefore combined 95 covariate-matched samples (47 samples from 28 ASD cases and 48 samples from 28 controls; Supplementary Fig. 1c; Supplementary Table 1) from the FC and TC for differential gene expression (DGE) analysis, comparing ASD vs. CTL using a linear mixed-effects regression framework to control for potential confounders (Methods). We identified 58 miRNAs showing significant (FDR < 0.05) expression changes between ASD and CTL, 17 down-regulated and 41 up-regulated in ASD cortex (Fig. 1b; Supplementary Table 2). The fold changes for the differentially expressed miRNAs were highly concordant between the FC and TC (Pearson correlation coefficient R = 0.96, P < 2.2 × 10−16; Fig. 1c).
To ensure the robustness of the signal, we performed resampling (Methods), finding that the fold changes for all miRNAs were highly concordant between the resampled and the original sample sets (Pearson’s R = 0.93 – 0.97, P < 2.2 × 10−16; Supplementary Fig. 3a). To confirm that the result was not biased by a small number of samples with low RNA Integrity (RIN) or high post-mortem interval (PMI), or from ASD cases with chromosome 15q11-13 duplication syndrome, we also performed DGE analysis after removing these samples, and found that the expression changes were concordant with those observed in all samples combined (Pearson’s R = 0.99, 0.98, and 0.99, P < 2.2 × 10−16, for samples with RIN ≥ 5, PMI ≤ 30 hrs, and after removal of 15q11-13 duplication samples, respectively; Supplementary Fig. 3b–3d).
Hierarchical clustering based on the top differentially expressed miRNAs (FDR < 0.05, |log2(fold change)| ≥ 0.3) showed distinct clustering for the majority of ASD cortex samples; 37/47 ASD samples from 23/28 cases grouped together, and confounders such as age, sex, brain region, seizures, medication, RIN, and brain bank did not drive the clustering (Fig. 1d), suggesting a shared miRNA dysregulation signature among the majority of ASD samples.
We further validated ten differentially expressed miRNAs using quantitative reverse transcription PCR (qRT-PCR) and confirmed sequencing-detected changes (Supplementary Fig. 3f–3g). Together, these results support the robustness and reproducibility of our data.
Additionally, we performed DGE analysis for 47 covariate-matched cerebellum samples (21 samples from ASD cases and 26 samples from controls; Supplementary Fig. 1e; Supplementary Table 1) (Methods), and observed a similar trends in differential expression compared with the cortex (Pearson’s R = 0.86, P < 2.2 × 10−16 for miRNAs differentially expressed in the cortex; Supplementary Fig. 3e; 16 miRNAs differentially expressed at FDR < 0.05 in both cortex and cerebellum).
Perturbation of miRNA co-expression modules in ASD brain
To further gain a systems-level understanding of the relationship between miRNA expression changes and disease status, we next applied weighted gene co-expression network analysis (WGCNA) using 109 cortex samples and a method that is robust to outliers (Supplementary Fig. 1d, 4a; Methods) to assign individual miRNAs to co-expression modules13–15. We identified 11 modules (Fig. 2a), summarized by their first principal component (PC1), or module eigengene (ME)14. We then assessed ME relationship to ASD status, and identified four modules that were significantly correlated with disease status (Pearson’s correlation, FDR < 0.05) and not any of the technical confounders, two down-regulated (brown and salmon) and two up-regulated (yellow and magenta) in ASD samples (Fig. 2b–2j).
Figure 2. miRNA co-expression modules dysregulated in post-mortem ASD cortex.
(a) Dendrogram showing miRNA co-expression modules defined in 109 cortex samples. Color bars below indicate original module assignment, consensus module assignment based on 200 rounds of bootstrapping, Pearson correlation coefficients with diagnosis and other potential confounders or covariates (all treated as numeric variables), and expression level for each miRNA. Arrows indicate three modules (brown, magenta, and yellow) significantly correlated with diagnosis. (b–d) Pearson correlation between module eigengenes and different covariates in 109 cortex samples (b), 47 cortex samples from patients aged 15 to 30 years (c), and 42 cortex samples from patients > 30 years (d). Correlation coefficients (R) and FDRs (Methods) are shown where FDR < 0.05. (e–j) Scaled module eigengene values across 109 cortex samples (e, g, i) and network plots (f, h, j) for the brown (e, f), magenta (g, h), and yellow (i, j) modules. In (e, g, i), samples are plotted in groups according to disease status and sex and color-coded as indicated above the graphs, and ordered by age as indicated below the graphs. In (f, h, j), miRNAs with kME (Methods) ≥ 0.5 are plotted according to multidimensional scaling (MDS) of miRNA correlations. Edge thickness is proportional to the positive correlation between the two connected miRNAs and node size is proportional to node connectivity. Enriched transcription factors (TF) and chromatin regulators (CR) (Fisher’s exact test, FDR < 0.05) are listed below the plot. Underlined, TF/CR differentially expressed (P < 0.05) in ASD cortex.
WGCNA permits direct assessment of the relationship of modules to important experimental variables or confounders, such as age, sex, brain region, and technical covariates including RIN, PMI, and brain bank13,15. The salmon and yellow modules showed significant correlations with age (Fig. 2b). Further inspection of the salmon module showed that it could be driven by younger non-aged-matched samples in cases compared with controls (Fig. 2c–2d, Supplementary Fig. 1d; Methods), so it was excluded from subsequent analysis. The yellow and magenta modules showed a different pattern of age dependence, whereby their disease association was more prominent in the younger age-matched set (15–30 years) relative to the older set (> 30 years) (Fig. 2c–2d, 2g, 2i, Supplementary Fig. 4b–4c). In contrast, the down-regulation of the brown module persisted throughout age (Fig. 2c–2e).
To test whether the correlation structure is similar between ASD and CTL samples, we constructed networks using ASD or CTL samples only and performed module preservation tests16, observing that all three ASD-associated modules were preserved across the ASD and CTL networks (Supplementary Fig. 4d–4e). Similar analysis revealed that the ASD-associated modules were also preserved across the frontal and temporal cortex (Supplementary Fig. 4f–4g). Furthermore, these co-expression relationships were also observed in two independent datasets from human brain (Methods; Supplementary Fig. 4h–4i). These results support the robustness and reproducibility of the three ASD-associated modules.
Co-expression among miRNAs may arise from co-regulation by common transcription factors (TFs) and/or chromatin regulators (CRs). To test this possibility, we obtained genome-wide high-confidence binding sites for 61 human TFs/CRs expressed in the cerebral cortex17, and identified miRNAs potentially regulated by each TF/CR (Methods). We then assessed enrichment of these potential targets in the three ASD-associated miRNA modules, observing distinct patterns of TF/CR enrichment in each module, consistent with their differential co-regulation (Supplementary Table 3). Interestingly, the magenta module was enriched (Fisher’s exact test, FDR < 0.05) for potential targets of SMARCC1, a core component of the BAF complex that has been implicated in ASD via mutational and network analysis of brain gene expression data13,18 (Fig. 2h; Supplementary Table 3).
Enrichment for ASD risk genes among miRNA targets
Previous transcriptomic analyses have revealed convergent molecular pathology in ASD, characterized by down-regulation of genes involved in neuronal and synaptic function, concomitant with up-regulation of genes involved in immune/inflammatory response11. We hypothesized that miRNAs differentially expressed in ASD may contribute to these perturbations repressing their mRNA targets. Alternatively, they might function as compensatory mechanisms to mitigate the existing mRNA dysregulation.
We bioinformatically predicted the mRNA targets of top differentially expressed miRNAs and hub miRNAs in the ASD-related modules (Methods), applying a well-established and widely used algorithm, TargetScan, which searches for miRNA target sites in mRNA 3′ UTRs and evaluates the targeting efficacy and evolutionary conservation (while controlling for background conservation) of the target sites6,19–21 (Methods). We then selected the top targets expressed in the temporal and frontal cortex (Parikshak et al., under revision) using two different criteria: (1) “the strongest” targets, which have the highest predicted targeting efficacy and are shared by two or more miRNAs (Methods), and (2) the most conserved target sites, which are more likely to have conserved physiological roles, but may not include newly evolved targets with species-specific functions (Methods). Overall, these two methods identified comparable numbers of targets, many overlapping, and some that were unique (Supplementary Table 4; Methods).
To test the validity of the bioinformatic predictions, we over-expressed a well-studied miRNA, hsa-miR-21-5p, in human neural progenitor cells (hNPCs; Methods). We observed significant (one-sided t-test, P < 2.2 × 10−16) down-regulation of hsa-miR-21-5p mRNA targets predicted by our methods, with a magnitude comparable to those supported by previous experimental evidence (miRTarBase22 release 4.5; http://mirtarbase.mbc.nctu.edu.tw) (Supplementary Fig. 5; Supplementary Table 5).
To explore the relationship of the ASD-affected miRNAs to genes that have been previously implicated in ASD, we systematically assessed whether targets of the differentially expressed miRNAs or ASD-associated miRNA modules are enriched for these genes. We first tested enrichment for a set of ASD risk genes from the Simons Foundation Autism Research initiative (SFARI) AutDB database23 (ASD SFARI; Methods), which have been implicated via common variant association, candidate gene studies, copy number variation (CNV), and genetic syndromes. For comparison, we also examined genes implicated in monogenic forms of intellectual disability13. We found that top targets of the differentially expressed miRNAs as well as the brown and magenta modules showed significant (Fisher’s exact test, FDR < 0.05; Methods) enrichment for SFARI ASD genes, but not for ID genes (Fig. 3a; Supplementary Table 6), suggesting that targets of ASD-affected miRNAs are enriched for genes causally connected with ASD, but less so for genes related solely to intellectual disability.
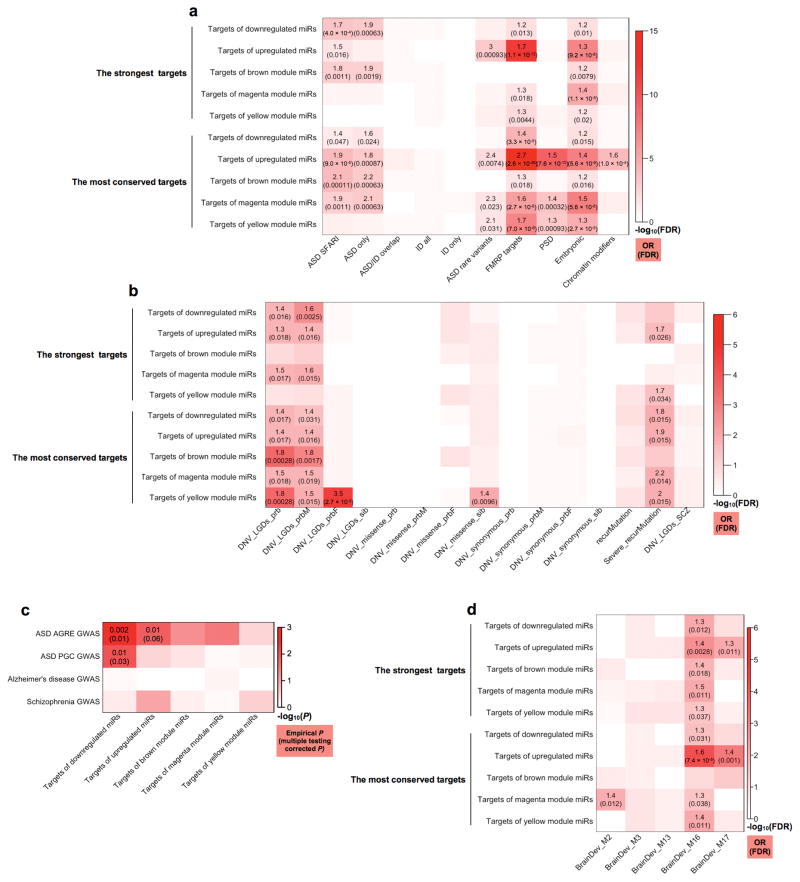
Figure 3. Enrichment of ASD risk genes among the top targets of ASD-affected miRNAs and miRNA modules.
(a) Heatmap showing enrichment (Fisher’s exact test) of ASD risk genes from SFARI (ASD SFARI) or implicated by rare variants (ASD rare variants), intellectual disability genes (ID all), genes encoding transcripts bound by FMRP (FMRP targets), genes encoding proteins in the postsynaptic density (PSD), genes expressed preferentially in human embryonic brains (Embryonic), and genes encoding chromatin modifiers. “ASD/ID overlap”, the overlap between the “ASD SFARI” and “ID all” sets. “ASD only” and “ID only”, non-overlapping ASD SFARI and ID genes, respectively. (b) Heatmap showing enrichment (logistic regression) of genes affected by de novo variants (DNVs), including likely gene-disrupting (LGD), missense, synonymous, and recurrent (recurMutation) mutations, in ASD-affected probands (prb, all probands; prbM, male probands; prbF, female probands) and unaffected siblings (sib). “Severe_recurMutation”, genes targeted by protein-disrupting recurrent mutations. “DNV_LGDs_SCZ”, LGD DNVs in individuals with schizophrenia. (c) Enrichment for overlap with linkage disequilibrium-based independent genomic regions associated with ASD (from Autism Genetic Resource Exchange [AGRE] or Psychiatric Genomics Consortium [PGC]), Alzheimer’s disease, or schizophrenia in GWAS among the strongest miRNA targets. The empirical and multiple-testing corrected P values calculated using the INRICH program are shown where the corrected P < 0.10. (d) Heatmap showing enrichment (Fisher’s exact test) for ASD-associated developmental gene co-expression modules in human cortex. In (a, b, d), enrichment odds ratios (OR) and FDR corrected P values (Methods) are shown for enrichments with FDR < 0.05.
We further examined additional classes of ASD-relevant genes: (1) genes whose transcripts are bound by the fragile X mental retardation protein (FMRP) 24,25, (2) genes encoding postsynaptic density (PSD) proteins24,26, (3) genes encoding chromatin modifiers24, and (4) genes expressed preferentially during embryonic brain development24,27. Enrichment for FMRP targets and embryonically expressed genes (Fisher’s exact test, FDR < 0.05; Methods) was observed for most target groups (Fig. 3a; Supplementary Table 6). The most conserved targets of the up-regulated miRNAs and miRNA modules significantly (Fisher’s exact test, FDR < 0.05; Methods) overlapped with PSD genes (Fig. 3a; Supplementary Table 6). The most conserved targets of the up-regulated miRNAs were also enriched (Fisher’s exact test, FDR < 0.05; Methods) for chromatin modifiers (Fig. 3a; Supplementary Table 6).
Recent whole-exome sequencing studies have identified high-confidence, likely-gene-disrupting (LGD, including nonsense, splice site, and frameshift) de novo variants (DNVs) in ASD1,2. We next asked whether these independently identified ASD risk genes are over-represented in the targets of ASD-affected miRNAs. For comparison, we also examined LGD DNVs in unaffected siblings, missense and synonymous DNVs, and LGD DNVs in individuals with schizophrenia24 (Methods). As DNV frequency has been shown to be linearly correlated with gene length24, we applied logistic regression that incorporates gene coding region length covered in exome sequencing24 to assess enrichment (Methods). Strikingly, the top targets of the ASD-affected miRNAs and miRNA modules showed specific enrichment (FDR < 0.05; Methods) for genes harboring LGD DNVs in ASD probands, but little enrichment for genes with LGD DNVs in unaffected siblings or individuals with schizophrenia, or missense/synonymous DNVs (Fig. 3b; Supplementary Table 6). We also observed enrichment (FDR < 0.05; Methods) for a shorter list of genes hit by severe recurrent mutations (two or more LGD mutations, small indels, and mutations that remove start or stop codons)24 (Fig. 3b; Supplementary Table 6). We also found that the top targets of the up-regulated miRNAs and miRNA modules were enriched (Fisher’s exact test, FDR < 0.05; Methods) for a list of 65 ASD risk genes implicated through rare and de novo variations (ASD rare variants) (Fig. 3a)28. The observation that the targets of the down-regulated miRNAs were enriched for ASD risk genes affected by likely-gene-disrupting DNVs (Fig. 3b), which are expected to disrupt protein function, suggests that these miRNAs might be part of a compensatory, or adaptive mechanism, since their down-regulation would favor target mRNA up-regulation.
We next asked whether targets of the ASD-affected miRNAs are enriched for common genetic variants associated with ASD in genome-wide association studies (GWAS)29–31. We applied the INRICH method32,33 to assess the overlap between ASD-associated, linkage-disequilibrium-based independent genomic intervals and the miRNA target genes, observing enrichment (multiple testing-corrected P < 0.10 via nested permutation) for ASD GWAS signals near the strongest targets of the differentially expressed miRNAs (Fig. 3c; Methods). Notably, the most substantial enrichment was in the targets of the down-regulated miRNAs, further suggesting that these might be compensatory changes. As a control, we performed the same analysis using GWAS data for Alzheimer’s disease34 and schizophrenia33 and did not observe significant enrichment (Fig. 3c). Importantly, the lack of enrichment in the control datasets was unlikely to be due to lower SNP density or interval number, as all datasets included comparable numbers of SNPs and intervals (Supplementary Table 6).
A previous study examining the temporal expression trajectories of ASD genes in the developing human cortex identified gene co-expression modules that are enriched for ASD risk genes13. We therefore tested for over-representation of these modules in the targets of ASD-affected miRNAs. This analysis highlighted module M16, which is up-regulated during early cortical development, and enriched for genes implicated in neural development and synaptic function13; genes in M16 were over-represented (Fisher’s exact test, FDR < 0.05; Methods) in most miRNA target groups (Fig. 3d; Supplementary Table 6).
Because the TargetScan algorithm searches for miRNA target sites in the 3′ UTR regions of mRNAs, we also used logistic regression to assess enrichment while controlling for 3′ UTR length, and observed a similar enrichment for ASD-related genes (Supplementary Fig. 6a–6c). Together, these results suggest functional involvement of miRNAs perturbed in ASD in its molecular pathology.
We also assessed shared functions among the targets of ASD-affected miRNAs by enrichment for Gene Ontology (GO) terms (GO-Elite software; Methods). Targets of the up-regulated miRNAs and the magenta module were enriched (FDR < 0.10) for genes related to neural development and several signaling pathways (Supplementary Fig. 7a–7b). The top GO terms (P < 0.01) for the targets of the down-regulated miRNAs and miRNA module concern both neuronal processes and immune function (Supplementary Fig. 7a–7b).
Relationship between miRNA and mRNA expression changes
To directly assess the role of miRNA dysregulation in ASD-associated mRNA level alterations, we next examined the relationship between miRNA and mRNA expression changes. We utilized mRNA expression data generated in a separate study that investigated mRNA expression changes in post-mortem ASD brain using RNA-Seq in 163 cortex (frontal and temporal) samples from ASD cases and controls (Parikshak et al., under revision), 101 of which overlapped with our study. We evaluated the relationship between miRNA and mRNA expression changes in the same set of individuals and samples, while at the same time recognizing that ASD-associated mRNA level perturbations are likely driven by several regulatory mechanisms, including transcription factors and epigenetic changes.
DGE analysis for mRNAs using 106 covariate-matched samples identified 1156 genes differentially expressed (FDR ≤ 0.05) between ASD and CTL cortex, 574 decreased and 582 increased (Parikshak et al., under revision). Consistent with previous findings, the down-regulated set was enriched for genes involved in neuronal and synaptic function, and the up-regulated set was enriched for microglia- and astrocyte-related genes involved in immune-inflammatory function11. We first compared the fold changes of differentially expressed mRNAs that are predicted targets of the differentially expressed miRNAs or miRNA modules to those of non-targets. Overall, we observed a trend in accordance with a negative effect of miRNAs on target mRNA level; mRNAs targeted by the up-regulated miRNAs or modules showed lower (P < 0.05) fold changes compared to the non-targets, whereas mRNAs targeted by the down-regulated miRNAs or module showed higher (P < 0.05) fold changes compared to the non-targets (Supplementary Fig. 8a–8h). We further assessed the relationship between mRNA and miRNA differential expression signatures by examining the correlations between the PC1s of differentially expressed miRNAs and differentially expressed mRNAs that were predicted targets. We observed significant (Pearson’s correlation, P < 0.005) negative correlations (Fig. 4a–4c), suggestive of negative regulation of the ASD-affected mRNAs by the ASD-associated miRNAs in the brain. Notably, the result was unlikely driven by separate association of the miRNAs and mRNAs to ASD status, as we regressed out disease status from the expression data when deriving the PC1s (Fig. 4a–4c), and computed the correlations in CTL samples or ASD samples alone (Supplementary Fig. 9a–9c).
Figure 4. Relationship between miRNA and mRNA expression changes in post-mortem ASD cortex.
(a–c) Correlations between the PC1s of differentially expressed miRNAs (FDR < 0.05, |log2(fold change)| ≥ 0.3) and differentially expressed mRNAs (FDR < 0.05) that are predicted targets (after regressing out disease status) in 101 cortex samples. (a) All differentially expressed miRNAs vs. all differentially expressed mRNAs; (b) up-regulated miRNAs vs. down-regulated mRNAs; (c) down-regulated miRNAs vs. up-regulated mRNAs. Pearson correlation coefficients (R) and P values are shown below the plots. (d) Negative correlations between the PC1s of top miRNAs in the ASD-associated miRNA modules and their predicted targets in the ASD-associated mRNA models. mRNA modules are represented with network plots showing the top 20 most connected module genes. Pearson correlation coefficients (R) and P values are indicated. Correlations for the magenta and yellow miRNA modules were calculated using 45 younger samples (ages between 15 and 30 years), given their stronger disease association at younger ages relative to older ages (> 30 years).
In a separate study (Parikshak et al., under revision), we identified gene co-expression modules significantly correlated with ASD status that were enriched for genes differentially expressed between ASD and control. Consistent with previous findings11, three modules (M4/10/16) down-regulated in ASD were related to neuronal and synaptic function as revealed by GO analysis, while two modules (M9/19) up-regulated in ASD were related to immune/inflammatory response and enriched for genes highly expressed in astrocytes and microglia (Parikshak et al., under revision). To explore the potential regulatory relationship between ASD-affected miRNA and mRNA modules, we first asked whether the top miRNAs in the ASD-associated miRNA module were negatively correlated with their predicted targets in the ASD-associated mRNA modules. We observed significant (Pearson’s correlation, P < 0.05) negative correlations between the up-regulated magenta and yellow miRNAs modules and their predicted targets in the down-regulated mRNA modules, and between the down-regulated brown miRNA module and its predicted targets in the up-regulated M9 and M19 mRNA modules (Fig. 4d). We further assessed the enrichment of the predicted targets of the ASD-affected miRNA modules for ASD-affected mRNAs and mRNA modules, and observed that the targets of the yellow and magenta miRNA modules were enriched (Fisher’s exact test, FDR < 0.05; Methods) for the down-regulated mRNAs and M16 mRNA module, whereas targets of the brown miRNA module were enriched (Fisher’s exact test, FDR < 0.05; Methods) for the up-regulated mRNAs and M9 mRNA module (Fig. 5a; Supplementary Table 6). These data, combined with the enrichment for ASD risk genes within the predicted miRNA targets, are consistent a functional involvement of the ASD-affected miRNAs in the molecular pathology of ASD; the up-regulated miRNAs and miRNA modules may contribute to the down-regulation of neuronal and synaptic genes in ASD brain, whereas the down-regulated miRNAs and miRNA module may contribute to the up-regulation of immune/inflammatory genes. However, the down-regulated miRNAs may also play a compensatory role given the enrichment of their targets for rare likely-gene-disrupting and common genetic variants associated with ASD (Fig. 5b).
Figure 5. Enrichment for ASD-affected mRNAs and mRNA modules within the top targets of ASD-affected miRNAs.
(a) Heatmap showing enrichment (Fisher’s exact test) for ASD-affected mRNAs and mRNA modules within the top targets of ASD-related miRNA modules. P values were FDR corrected across 6 target groups for each mRNA group. Odds ratios and FDRs are shown for enrichments with FDR < 0.05. (b) Model for the role of miRNA dysregulation in ASD molecular pathology. The up-regulated miRNAs and miRNA modules may play a contributory role by repressing ASD risk genes and neuronal/synaptic genes down-regulated in ASD. The down-regulated miRNAs and miRNA module may contribute to the up-regulation of immune/inflammatory genes in ASD, but might also play a compensatory role given the enrichment of their targets for rare protein-disrupting and common genetic variants associated with ASD. (c–d) Enrichment (Fisher’s exact test) for ASD-affected mRNAs and mRNA modules within the strongest (c) or the most conserved (d) targets of individual candidate miRNAs.
To prioritize candidates for further characterization, we next examined whether the targets of individual candidate miRNAs were enriched for mRNAs or mRNA modules that were anti-correlated with the miRNAs in ASD. We found that the targets of several down-regulated miRNAs were enriched (P < 0.05) for the up-regulated mRNAs and M9 mRNA module, whereas targets of several up-regulated miRNAs showed enrichments (Fisher’s exact test, P < 0.05) for the down-regulated mRNAs and M4/M16 mRNA modules (Fig. 5c–5d; Supplementary Table 6).
hsa-miR-21-3p targets neuronal genes down-regulated in ASD
We next experimentally tested the predicted down-regulation of target mRNAs by miRNAs in ASD via in vitro perturbation of several dysregulated miRNAs in hNPCs. We first focused on hsa-miR-21-3p, which was the second most up-regulated miRNA in ASD cortex (1.5-fold increase, FDR < 0.05) (Supplementary Fig. 10a) and whose predicted targets exhibited prominent enrichment (Fisher’s exact test, P < 0.01) for down-regulated mRNAs and the M16 mRNA module (Fig. 5c–5d; Supplementary Table 6). hsa-miR-21-3p is conserved across vertebrates (Supplementary Fig. 10b) and widely expressed in different human brain regions throughout development (Supplementary Fig. 10d–10e). However, its role in the CNS has not been explored. We over-expressed hsa-miR-21-3p in hNPCs and examined the consequent mRNA changes by RNA-Seq (Supplementary Table 5). We found that predicted hsa-miR-21-3p target genes showed significant down-regulation (one-sided t-test, P < 2.2 × 10−16) as a group compared to non-targets (Fig. 6a), again validating the bioinformatic predictions.
Figure 6. hsa-miR-21-3p targets neuronal genes down-regulated in ASD.
(a) Distributions (left) and cumulative distributions (right) of mRNA log2(fold change) in response to over-expression of hsa-miR-21-3p in hNPCs. Statistical significance between target groups and non-targets was assessed using one-sided t-tests assuming unequal variance. (b–e) Heatmaps showing enrichment of validated hsa-miR-21-3p targets for ASD-related gene lists (b–d) as well as ASD-affected mRNAs and mRNA modules (e). (b) Logistic regression, (c–e) Fisher’s exact test. Odds ratios and P values are shown where P < 0.05. (f) A partial list of validated hsa-miR-21-3p target genes associated with ASD.
We next performed a series of enrichment analyses to characterize the overlap between validated hsa-miR-21-3p targets and ASD-related genes. hsa-miR-21-3p targets showed enrichment for genes harboring LGD and recurrent DNVs and rare variants in ASD-affected individuals (logistic regression, P < 0.05), but little enrichment for missense/synonymous DNVs, or LGD DNVs in unaffected siblings or schizophrenia-affected individuals (Fig. 6b–6c, 6f; Supplementary Table 6). ASD SFARI genes, FMRP targets, chromatin modifiers, embryonically expressed genes, and the ASD-related M3 and M16 brain developmental modules13 were also enriched (Fisher’s exact test, P < 0.05), but not genes associated with intellectual disability (Fig. 6b, 6d, and 6f; Supplementary Table 6). Enrichment (Fisher’s exact test, P < 0.005) was also observed for genes down-regulated in post-mortem ASD cortex, in particular the M16 mRNA module (Fig. 6e–6f; Supplementary Table 6), the ME of which was negatively correlated with hsa-miR-21-3p (Pearson’s R = −0.50, P = 8.5 × 10−8). Notably, hsa-miR-21-3p over-expression down-regulated (one-sided t-test, P < 0.05) several hub genes in M16, including PAFAH1B1/LIS1, which is critical for neuronal migration and causally linked to lissencephaly35, DLGAP1, a PSD scaffold protein that binds to SHANK3, an ASD risk gene1, and ATP2B1/PMCA1, a calcium transporter. In addition, hsa-miR-21-3p repressed the levels of ATP1B1, a PSD-localized Na+/K+ transporter haboring ASD-associated LGD DNVs24, DYNC1I1, which is critical for neuronal migration35, NEEP21, which is involved in synaptic transmission36, SV2B, a neurotransmitter release regulator37, and several genes in the ubiquitin-proteasome pathway (UCHL5, USP33, USP7, UBE2K, FBXO11, KIAA0368) (Fig. 6f; Supplementary Table 5). Also of interest, hsa-miR-21-3p over-expression led to a pronounced decrease (38%; one-sided t-test, P = 0.0001) in PCDH19, mutations in which cause epilepsy and ASD in females38. hsa-miR-21-3p and PCDH19 levels were also negatively correlated in post-mortem cortex (Pearson’s R = −0.48, P = 4.0 × 10−7), consistent with the in vitro observation. Together, these results implicate hsa-miR-21-3p in regulating the mRNA levels of neuronal and synaptic genes and suggest a role for hsa-miR-21-3p in the down-regulation of these genes in ASD.
hsa_can_1002-m targets genes in the EGFR and FGFR pathways
Another candidate of particular interest was a predicted novel miRNA, hsa_can_1002-m, one of the most down-regulated miRNAs in ASD cerebral cortex (2.5-fold decrease, FDR < 0.05; Supplementary Fig. 10a) and the top hub miRNA of the brown module (Fig. 2e; Supplementary Table 2). Over-expression of hsa_can_1002-m in hNPCs led to significant decreases(one-sided t-test, P < 10−7) in its predicted targets (Fig. 7a; Supplementary Table 5), supporting its function. Interestingly, the predicted hsa_can_1002-m precursor is located in a genomic region only present in primates, but not lower vertebrates (Supplementary Fig. 10c). Moreover, RNA-Seq in the mouse cortex did not detect the hsa_can_1002-m sequence. We further performed qRT-PCR using a primer for the human mature sequence, detecting robust expression in human, chimpanzee, and rhesus macaque cerebral cortices, but not the mouse cortex (Fig. 7b), confirming the data from the genome sequence that hsa_can_1002-m is a primate-specific miRNA. RNA-Seq data from the BrainSpan project showed that hsa_can_1002-m is broadly expressed, but developmentally regulated in the human brain from infancy to adulthood (Supplementary Fig. 10f–10g).
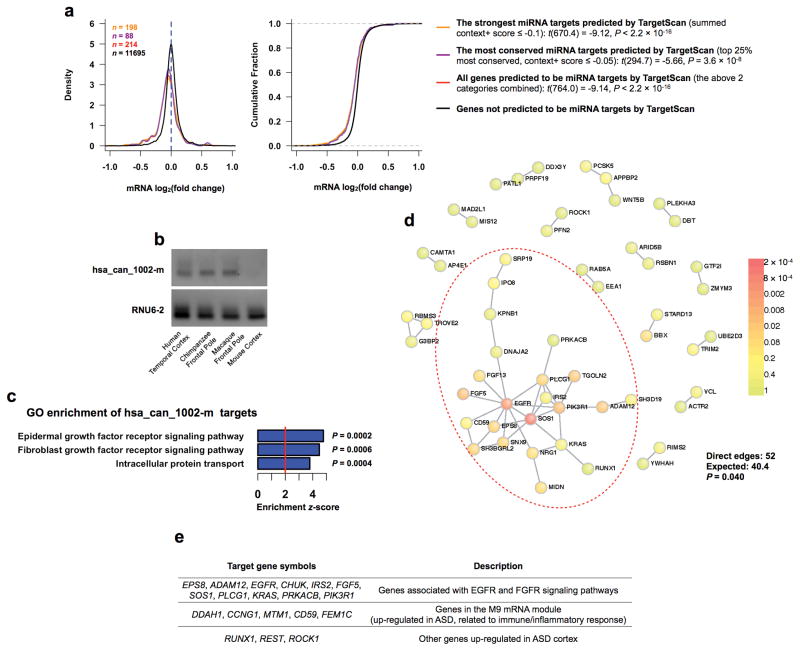
Figure 7. hsa_can_1002-m is primate-specific and targets genes in the EGFR and the FGFR signaling pathways.
(a) Distributions (left) and cumulative distributions (right) of mRNA log2(fold change) in response to over-expression of hsa_can_1002-m in hNPCs. Statistical significance between target groups and non-targets was assessed using ones-sided t-tests assuming unequal variance. (b) RT-PCR in human, chimpanzee, macaque, and mouse cortices using primers designed for human hsa_can_1002-m. RNU6-2 was used as control. The experiment was repeated twice and the result was reproducible. (c) Top GO terms for validated hsa_can_1002-m targets. Uncorrected P values are shown. (d) Direct protein-protein interaction network between validated hsa_can_1002-m targets. Nodes are colored based on the P values of the seed proteins (the probability that by chance the seed protein would be as connected as is observed) according to the legend on the right. (e) A partial list of validated hsa_can_1002-m target genes.
Validated hsa_can_1002-m targets did not show enrichment for known ASD risk genes. Nonetheless, the top GO categories (GO-Elite software, P < 0.001; Methods) implicated the epidermal growth factor receptor (EGFR) and the fibroblast growth factor receptor (FGFR) signaling pathways (Fig. 7c–7e; Supplementary Table 6), which have been importantly implicated in neuron/glia proliferation, differentiation, and survival in both the developing and adult brain, as well as inflammatory/immune processes39,40. Notably, all implicated targets are involved in the activation of these pathways, suggesting that hsa_can_1002-m down-regulation would lead to an increase in pathway activity. These targets form a more highly connected local protein-protein interaction network than expected by chance (DAPPLE software, P < 0.05; Methods), providing independent confirmation of co-regulation at the protein level (Fig. 7d). Of particular interest are EPS8 (a signaling adaptor protein regulating dendritic spine density and synaptic plasticity), ADAM12 (a metalloprotease required for EGFR ligand processing), CHUK (a kinase critical for NF-kappa-B activation), and RUNX1 (a transcription factor essential for immune cell development and activation), all of which were up-regulated (P < 0.05) in ASD post-mortem cortex (Parikshak et al., under revision) (Supplementary Table 5). In addition, the validated strongest hsa_can_1002-m targets were enriched for genes in the immune-related, up-regulated M9 mRNA module (Fisher’s exact test, Odds ratio = 1.9, P = 0.02; Fig. 7e), consistent with prediction (Fig. 5c). Together, these findings implicate a novel, primate-specific miRNA hsa_can_1002-m in regulating the EGFR and the FGFR signaling pathways, shedding light on its potential role in neuronal and glial development and function, as well as in ASD molecular pathology involving neural-immune interactions.
Experimental characterization of other candidate miRNAs
We also experimentally validated the targets of several other candidate miRNAs, including hsa-miR-103a-3p in the yellow module and hsa-miR-143-3p and hsa-miR-23a-3p in the magenta module, the predicted targets of which were enriched (Fisher’s exact test, P < 0.05) for down-regulated mRNAs and the M16 mRNA module (Fig. 5c–5d; Supplementary Table 6). Notably, hsa-miR-23a-3p has also been reported to be up-regulated in lymphoblasts in ASD patients41, and hsa-miR-143-3p has been recently shown to be regulated by a primate-specific long non-coding RNA and implicated in neural progenitor proliferation42. Over-expression of these miRNAs in hNPCs led to significant reductions (one-sided t-test, P < 2.2 × 10−16) in the predicted target mRNAs (Fig. 8a–8c; Supplementary Table 5). Consistent with bioinformatic predictions, enrichment analysis revealed that the validated targets of hsa-miR-103a-3p and hsa-miR-143-3p were enriched (Fisher’s exact test, P < 0.05) for down-regulated mRNAs and/or the M16 mRNA module (Fig. 8d, 8f; Supplementary Tables 6). Validated targets of hsa-miR-23a-3p were enriched (Fisher’s exact test, P < 0.05) for ASD SFARI genes and ASD rare variants (Fig. 8e–8f; Supplementary Tables 6). These data are consistent with a potential functional involvement of these miRNAs in ASD, making them interesting candidates for further functional manipulation in model systems.
Figure 8. Experimental validation of targets of other candidate miRNAs.
(a–c) Distributions (left) and cumulative distributions (right) of mRNA log2(fold change) in response to over-expression of hsa-miR-103a-3p (a), hsa-miR-143-3p (b), and hsa-miR-23a-3p (c) in hNPCs. Statistical significance between target groups and non-targets was assessed using ones-sided t-tests assuming unequal variance. (d) Enrichment (Fisher’s exact test) of validated targets of hsa-miR-103a-3p and hsa-miR-143-3p for down-regulated mRNAs and the down-regulated M16 mRNA module. (e) Enrichment (Fisher’s exact test) of validated targets of hsa-miR-23a-3p for ASD SFARI genes and ASD risk genes implicated by rare variants. (f) A partial list of validated target genes.
DISCUSSION
Our genome-wide, integrative analysis provides new insights into the role of miRNAs in ASD pathophysiology. We observe a miRNA differential expression signature shared by a majority of ASD cortex samples11. Within the targets of the ASD-affected miRNAs and miRNA co-expression modules, we observe enrichment for ASD risk genes implicated by multiple forms of genetic variation, and much less so for variants associated with intellectual disability, schizophrenia, or Alzheimer’s disease. This suggests that ASD risk genes are highly dosage sensitive; we surmise that miRNA dysregulation provides an alternative pathway to gene disrupting mutations for perturbing key transcript levels, thus potentially contributing to ASD susceptibility (Fig. 5b). This model is supported by the negative correlation between the expression changes of ASD-affected miRNAs and mRNAs, and our experimental validation showing regulation of mRNA targets by several top candidate miRNAs. Collectively, our findings suggest that ASD-associated transcriptomic changes may be partially attributable to miRNA dysregulation, with the up-regulated miRNAs potentially contributing to the down-regulation of neuronal and synaptic genes and the down-regulated miRNAs contributing to the up-regulation of immune/inflammatory genes, as well as possible compensatory changes (Fig. 5b).
A few previous studies have also examined miRNA expression changes associated with ASD, but assessed a limited number of miRNAs (using qRT-PCR or microarray), had a small sample size, and/or used non-neuronal tissues/cells41,43,44. Our genome-wide analysis using the most relevant tissue and a better-powered sample, along with integration with multiple gene sets and expression data, provides the most robust and comprehensive assessment of miRNA dysregulation in ASD brain to date. Most of the differentially expressed miRNAs that we identified were not reported previously. Interestingly, several miRNAs, including hsa-miR-107, hsa-miR-106a-5p, hsa-miR-10a-5p, has-miR-136-5p, and has-miR-155-5p overlapped with previous studies and would be interesting candidates for further experimental investigation.
Besides providing a systems-level view of miRNA expression landscape in post-mortem ASD brains, we have also experimentally characterized the targets of several top candidate miRNAs in hNPCs. We found that transcripts regulated by hsa-miR-21-3p, an up-regulated miRNA of unknown function in the nervous system, showed enrichment for ASD candidate genes and genes down-regulated in ASD cortex. Its connection with the M16 mRNA module, which is enriched for neuronal and synaptic genes, suggests a role in regulating neuronal development and synaptic function and a link with the neuronal and synaptic defects in ASD.
We also show that hsa_can_1002-m, a novel, primate-specific miRNA down-regulated in ASD, regulates several transcripts involved in the EGFR and the FGFR signaling pathways. This is intriguing from an evolutionary point of view given the important roles of these pathways in regulating neural stem cell proliferation in the brain, as a rapid increase in brain size resulting from increased neural stem cell division has been suggested as a critical step in primate brain evolution45. Additionally, early postnatal brain overgrowth followed by relative growth normalization has been repeatedly observed in ASD-affected children and is thought to reflect abnormal early neurodevelopment46. It is possible that negative regulatory mechanisms, such as miRNAs, have evolved to restrict increased cell proliferation in the primate brain, as uncontrolled proliferation would disrupt brain development and function.
The observation that the cerebellum showed a similar trend of miRNA differential expression to the cortex was somewhat unexpected, given previous findings11 showing very few mRNAs differentially expressed in the cerebellum. Our recent RNA-Seq analysis (Parikshak et al., under revision) revealed that some mRNAs showed similar trends of differential expression in the cerebellum, albeit of substantially smaller magnitude, compared with the cortex (Parikshak et al., under revision). We speculate that this might be due to the differences in cell types or some other aspect of the molecular milieu of the cerebellum that renders it resilient to the mRNA changes observed in the cortex.
One limitation of gene expression studies using post-mortem brains is that while ASD likely arises from abnormalities during early brain development, the majority of available samples are from adults. Although there was no clear association of miRNA or mRNA11 (Parikshak et al., under revision) perturbations with medication history, some observed changes almost certainly reflect the consequences (rather than the causes) of having the disorder or compensatory responses, whereas some early gene expression perturbations that play a causal role may not be captured in adult brain. In this regard, it is interesting to note that the two miRNA modules up-regulated in ASD showed stronger disease association at younger ages than at older ages, suggesting that they might be more related to early pathogenic features. Future studies using brain samples from younger patients and patient-derived brain organoids that model early brain development47 can provide more insights into early transcriptomic perturbations. Another limitation is the cellular heterogeneity of post-mortem tissue; cell type-specific miRNA expression data in the human brain are still lacking. The observed expression changes could occur in a single or multiple cell types, or reflect changes in cell composition. Future studies using transcriptomic profiling of isolated cell types and single-cell sequencing48 could help resolve these possibilities.
Collectively, our genome-wide, integrative analysis provides a framework for assessing the functional involvement of miRNA in ASD. By integrating several ASD candidate gene sets and correlating with mRNA expression data, we provide multiple lines of evidence for a functional role of miRNA dysregulation in ASD, either as contributory or compensatory factors. These analyses also identify a rich set of ASD-associated candidate miRNAs for further study.
METHODS
Brain tissue samples
Brain tissue samples were acquired from the Autism Tissue Program (ATP) brain bank at the Harvard Brain and Tissue Bank, the National Institute for Child Health and Human Development (NICHD) Eunice Kennedy Shriver Brain and Tissue Bank for Developmental Disorders, the UK Brain Bank for Autism and Related Developmental Research (BBA), and the MRC London Neurodegenerative Diseases Brain Bank. Up to three brain regions from each individual were assessed: frontal cortex (FC, Brodmann area [BA] 9), temporal cortex (TC, BA41/42/22), and cerebellar vermis. For some individuals, not all three regions were included due to limited tissue availability. Metadata for all 242 samples, including age, sex, brain region, brain bank, medical history, and sample quality metrics are summarized in Supplementary Table 1. Individuals defined as autistic had either a confirmed ADI-R diagnosis, duplication 15q syndrome with confirmed ASD, or a diagnosis of autism supported by other evidence such as clinical history. Dissections of frozen samples were performed on dry ice in a dehydrated dissection chamber, and randomized for balance of age, sex, brain region, and diagnostic status.
RNA extractions, library preparation, and small RNA sequencing
Total RNA was extracted from approximately 100 mg of frozen tissue using the miRNeasy kit (Qiagen). RNA integrity number (RIN) of the extracted total RNA was measured using an Agilent Bioanalyzer. For 195 of the 242 samples, rRNAs were depleted from 2 μg total RNA with the Ribo-Zero Gold kit (Illumina). Remaining RNA was size selected with AMPure XP beads (Beckman Coulter). Small RNAs (including miRNAs) in the supernatant (250 μL) were precipitated with 2 μL glycogen (Roche), 2 μL 0.1x Pellet Paint NF Co-precipitant (Merck Millipore), 25 μL 3 M NaOAc (pH 5.2), and 700 μL 100% ethanol (−20 °C), and resuspended in 5 μL RNase-free water for subsequent library preparation. For the other 47 samples, 0.7 μg total RNA was directly used for library preparation, due to the lack of sufficient material for rRNA depletion. We compared sequencing results on 4 brain tissue samples for which both library preparation methods were used and found the results on the same sample to be highly positively correlated (Pearson correlation coefficients = 0.93–0.97, P < 2.2 × 10−16). Small RNA libraries were prepared in batches of 12 – 48 samples using the TruSeq Small RNA library Preparation Kits (Illumina) according to the manufacturer’s protocol. Library preparation was randomized for balance of diagnostic status (ASD vs. CTL), age, sex, and brain region. Libraries were then validated using an Agilent Bioanalyzer and quantified with the Qubit dsDNA HS assay (Life Technologies). 19–29 libraries barcoded with Illumina TruSeq small RNA indexes were pooled and sequenced in each lane on a Illumina HiSeq2500 instrument using the high output mode with 50 bp single-end reads. Investigators were blinded during dissection, RNA extraction, library preparation, and sequencing to all metadata information about the samples.
Quality control (QC)
Sequencing reads were demultiplexed using CASAVA v1.8.2 (Illumina) and sequencing adaptors were removed using the fastx_clipper function in the FASTX-Toolkit (http://hannonlab.cshl.edu/fastx_toolkit/index.html). Sequencing quality (including quality scores, GC content, sequence length distribution, duplication levels, overrepresented sequences, and Kmer content) was examined using FastQC v0.10.1 (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/). All 242 samples sequenced for small RNAs were also sequenced for mRNAs using RNA-Seq (Parikshak et al., under revision). Genotypes for sites that are heterozygous or homozygous for the minor allele relative to the reference genome were called from RNA-Seq data49 (Parikshak et al., under revision). Genotypes were then coded as NA (homozygous for the major allele or not enough depth to detect), 1 (detected heterozygous), or 2 (homozygous for the minor allele). Pairwise Spearman correlations between samples were calculated, based on which the samples were clustered. Any sample that did not cluster with other samples from the same individual was further examined for possible contamination or sample mix-up and excluded from the downstream analyses if sample mix-up was not resolvable. Twenty samples from 18 individuals were removed using this criterion. In addition, 6 samples from 2 individuals who were not diagnosed with ASD, but had other conditions (one with idiopathic epilepsy, one with copy number variation) were also removed to avoid confounding effects. This resulted in a total of 216 samples that passed QC.
miRNA quantification and prediction
For quantification of mature miRNAs documented in miRBase release 20, sequencing reads were first mapped to the hg19 reference genome using the mapper.pl script in the miRDeep2 package9,50 with reads < 18 nt discarded. The quantifier.pl script was then used with default settings to quantify the number of reads mapped to mature miRNAs, allowing 0 mismatch.
For the prediction of novel miRNAs, two methods were used. First, the miRDeep2.pl script in the miRDeep2 package was run with default settings and mature miRNAs in Pan troglodytes, Gorilla gorilla, Pans paniscus, and Pongo pygmaeus (miRBase release 20) provided as related species. miRDeep2 examines the position and frequency of reads aligned to the genome (“signature”) with respect to a putative RNA hairpin and scores miRNA candidates employing a probabilistic model based on miRNA biogenesis. The score reflects the energetic stability of the putative hairpin and the compatibility of the observed read distribution with miRNA cleavage. The higher the score, the more likely that the miRNA candidate is a true positive. We only kept predictions with a score ≥ 4 (corresponding to a true positive probability of 78 ± 2%; names of novel miRNAs predicted using this method start with “hsa_chr”). Second, we applied miRanalyzer51,52, which predicts putative mature miRNAs and precursors based on mapped reads and folding energy and employs 5 different Random Forest models to calculate the probability that a candidate is a true miRNA. We only kept candidates with positive predictions in at least 4 out of all 5 models (names of novel miRNAs predicted using this method start with “hsa-P”). In addition, we included novel miRNAs identified in a recent study, which were predicted using miRDeep2 based on 94 human sRNA-Seq datasets10 (names start with “hsa_can”). Many of these were supported by various levels of experimental evidence, including interaction with Ago1/2, interaction with DGCR8, response to silencing of the miRNA biogenesis machinery, and/or interaction with target mRNAs in CLASH experiments10. Duplicate predictions in the above 3 novel miRNA sets were collapsed. Predicted miRNAs > 25 nt were removed according to the normal size range of miRNAs. The quantifier.pl script was used to quantify the number of reads mapped to novel mature miRNAs with 0 mismatch.
miRNAs (from miRBase release 20 or predicted) with read counts ≥ 3 in at least 50% of samples in each region, sex, or diagnostic status (ASD vs. CTL) group were kept for further analysis. 699 miRNAs (552 in miRBase 20 and 147 predicted) in our dataset met this criterion. This step helps remove miRNAs that are supported by only a few reads and likely expressed at very low levels. Such low levels of expression are unlikely to provide reliable, statistically significant differential expression results.
Expression value normalization and adjustment of covariates
Raw expression data for all 242 samples were normalized for library size using the estimateSizeFactors function in the DESeq R package53. For each miRNA in each sample, a ratio is calculated by dividing the read count by the geometric mean across all samples. A scaling factor for each sample is then calculated as the median of this ratio for all miRNAs in the sample. The raw read counts in each sample are then divided by the scaling factor to generate the library size-normalized data. The effect of GC content was normalized using the CQN R package54. Library preparation batch effects were normalized using the ComBat function in the sva R package55. The effects of other technical covariates, including RIN, PMI, and brain bank, were normalized together using a linear model. Additionally, mRNAs prepared from the same total RNA samples used in this study were also analyzed using RNA-Seq in a separate study (Parikshak et al., under revision). We observed that the proportion of mRNA reads mapped to exons also showed significant correlation with the miRNA expression data, as it likely reflected the ratio between cytoplasmic (where miRNAs are in the processed, mature form) and nuclear (where miRNAs are in the precursor form) RNAs. Therefore we also included this technical variable in the linear regression model along with other technical variables. The log2 transformed expression data normalized for library size and technical covariates were used for subsequent differential gene expression analysis and weighted gene co-expression network analysis.
Definition of sample sets for analysis
In exploratory data analysis, we performed principle component analysis (PCA) and hierarchical sample clustering for 216 samples that passed QC (180 samples prepared using the rRNA depletion method and 36 samples prepared using total RNA) to examine the relationship between the expression data and different covariates. We observed that library preparation method (rRNA depletion vs. total RNA) and brain region (cortex vs. cerebellum) had major effects on the expression profile, both having strong correlations with PC1 and PC2 (Supplementary Fig. 2a–2b). Accordingly, samples prepared with the two different methods or from different brain regions (cortex vs. cerebellum) were clustered into distinct branches in hierarchical clustering (Supplementary Fig. 2c). In addition, brain bank, age, RIN, and PMI also showed significant correlations with PCs 1–5 (Supplementary Fig. 2a).
To avoid the confounding effects of library preparation method and brain region, we divided our samples into different subsets based on these two covariates, and first focused on cortex samples prepared using the rRNA depletion method (116 samples) for our main analysis. We performed outlier detection using a standard method12,13,20,21,56. Specifically, we calculated the connectivity between samples based on the biweight midcorrelation of miRNA expression using the fundamentalNetworkConcepts function In the WGCNA R package20,57, and removed samples with connectivity more than 3 standard deviations away from the mean as outliers. This process was then repeated on the remaining samples until no more outliers were detected. Using this method, we removed 2 samples as outliers. We also removed 5 samples (from 3 individuals) for which PMI information was not available, resulting in 109 samples, which were used for the main WGCNA analysis. For the main DGE analysis, to avoid the potential confounding effect of age, we further removed 14 ASD samples for which there were no age-matched control samples, including 13 samples with age ≤ 11 years and 1 sample with age of 67 years (ages of ASD and CTL groups were both between 15 – 60 years after removing these samples), resulting in 95 samples. The WGCNA analysis relies on correlation between gene expression levels across samples and is not as critically affected by unmatched covariates as the DGE analysis. It also permits direct assessment of the relationship of modules to experimental variables or confounders. Therefore we used all 109 cortex samples to maximize the robustness of the WGCNA analysis. We combined samples from the frontal cortex and the temporal cortex, as PCA analysis and hierarchical sample clustering indicated that miRNA expression profiles were very similar between these two regions, but distinct in the cerebellum (Supplementary Fig. 2a–2f).
The cortex samples prepared using the total RNA method (31 samples) were imbalanced in brain bank between ASD and CTL; 10 out of 16 ASD samples were from UK BBA while none of the 15 CTL samples was from this brain bank. To avoid potential confounding effects, this set was not used for DGE analysis, but instead for evaluating miRNA module preservation, which relies on miRNA co-expression relationships across samples and should not be critically affected by unmatched covariates.
For DGE analysis in the cerebellum, we selected 47 samples prepared using the rRNA depletion method as follows. From 64 samples that passed QC, 4 samples were removed as outliers as described above. We also removed 3 samples with no PMI information, 1 sample with very high PMI (50 hours), 3 young ASD samples (age ≤ 5 years) with no age-matched controls, and 6 ASD samples with RINs lower than the lowest RIN in CTL.
Differential gene expression (DGE) analysis
Differential expression between ASD and CTL was calculated for each miRNA in the cortex using a linear mixed-effects (LME) model using the R package nlme, as more than one brain region from the same individuals were included (Supplementary Software). The model treated diagnosis, age, sex, and region as fixed effects (numeric or factor variables) and individual brain ID as a random effect: lme(expression level ~ diagnosis + age + sex + region, rand = ~1|brainID). Effect sizes (Beta values) and P values for diagnosis were calculated from the model for all miRNAs, and P values were adjusted for multiple comparisons using Benjamini-Hochberg correction to assess false discovery rate (FDR). To test if the result is robust to resampling, we performed random resampling by dividing the samples into 10 age deciles and randomly picking 7 deciles in each round of resampling, thus ensuring that age is still matched between ASD and CTL. We found that miRNA fold changes were highly concordant between the resampled sets and the original set (Supplementary Fig. 3a). To ensure that the P values were not skewed by the distribution of the data, we also computed P values from 100 rounds of permutation (randomizing the ASD/CTL status of all samples from the same individual each time), and found that the P value rankings of genes were highly concordant with those obtained with the original sample set (Pearson’s R = 0.99, P < 2.2 × 10−16). For DGE analysis in the cerebellum, a linear model that included diagnosis, age, and sex was used (as only one region from each individual was included): lm(expression level ~ diagnosis + age + sex). Log2 transformed expression data that have been normalized for library size and technical covariates were used.
Weighted gene co-expression network analysis (WGCNA) and further network characterization
The WGCNA R package14,24,57 was used for building signed co-expression networks (Supplementary Software). Briefly, biweight midcorrelation was first used to calculate pair-wise correlations (values between −1 and 1) between miRNAs. Next, pair-wise topological overlap (TO, values between 0 and 1) between miRNAs was calculated with a power of 8 based on a fit to scale-free topology. To construct the network in a way that is robust to outliers, we performed 200 rounds of bootstrapping and computed the TO matrix for each of the resampled networks. We then took the medians of the TOs to generate a consensus TO matrix. Co-expression modules comprised of positively correlated miRNAs with high consensus TO were then identified using the cutreeDynamic function in the dynamicTreeCut R package, which employs a “dynamic hybrid” algorithm58,59, with the following parameters: method = “hybrid”, deepSplit = 3, pamStage=T, pamRespectsDendro = T, minClusterSize = 10. The algorithm first detected preliminary clusters based on the merging information of the dendrogram. miRNAs unassigned in the first step were next assigned to the preliminary clusters if they were sufficiently close to the clusters. In this step the dendrogram was ignored and only dissimilarity information is used. The expression of each module was summarized by the module eigengene (ME, defined as the first principle component of all miRNAs in a module). Modules whose eigengenes were highly correlated were further merged using the mergeCloseModules function in the WGCNA R package. The module membership of each miRNA in each module (kME) is defined as the biweight midcorrelation to the ME. Pearson correlations between MEs and diagnosis, age, sex, brain region, RIN, PMI, and brain bank were calculated. P values were FDR adjusted across 11 modules for each covariate using Benjamini-Hochberg correction. We identified four modules that were significantly (FDR < 0.05) correlated with disease status, two (brown and salmon) down-regulated and two (yellow and magenta) up-regulated in ASD samples. Further inspection of the salmon module revealed that it was not significantly (FDR < 0.05) correlated with disease status in subsets of younger (15 – 30 years) and older (> 30 years) age-matched samples (Fig. 2c–2d). Additionally, none of the miRNAs in this module was differentially expressed (FDR < 0.05) in our DGE analysis using the 95 age-matched cortex samples. These observations suggest that the disease association of the salmon module was likely driven by younger non-aged-matched samples in cases compared with controls (Supplementary Fig. 1d), so it was excluded from subsequent analysis. Network plots were generated with the igraph R package32,60.
To examine whether the co-expression structure is similar between ASD and CTL samples, between FC and TC samples, or in independent data sets, we performed module preservation analysis using the modulePreservation function in the WGCNA R package16,32, which calculates for each module the Zsummary statistic, a measure that combines module density and intramodular connectivity metrics. 2 < Z < 10 indicates weak to moderate preservation and Z > 10 indicates high preservation.
Transcription factor (TF)/chromatin regulator (CR) binding site analysis
Genome-wide high-confidence binding sites for a set of 61 human TFs/CRs expressed in the cortex were obtained from a previous study17,32. miRNAs located within 20 kb of the binding sites of each TF/CR were identified using bedtools61 and defined as potential targets. Fisher exact tests were then performed using the fisher.test R function to assess enrichment of the targets of each TF/CR for miRNAs with kME ≥ 0.5 in the brown, magenta, and yellow modules. All 699 expressed miRNAs were used as background. P values were FDR adjusted for 61 TFs/CRs tested for each module using the Benjamini-Hochberg correction.
Prediction of miRNA targets
For prediction of miRNA targets, the stand-alone version of TargetScan6,19–21 was used. 3′ UTR sequences of RefSeq genes were download from the TargetScan website (http://www.targetscan.org/vert_61/vert_61_data_download/UTR_Sequences.txt.zip). For each miRNA target site, a branch length score, which evaluates target site conservation while controlling for 3′ UTR conservation, and a context+ score, which measures targeting efficacy irrespective of conservation, were calculated. To select top differentially expressed miRNAs and top miRNAs in ASD-associated modules for target prediction, we used the following criteria: (1) for top differentially expressed miRNAs, FDR < 0.05 and |log2(fold change)| ≥ 0.3 in 95 cortex samples; (2) for top miRNAs in ASD-associated modules, kME ≥ the median for each module (kME ≥ 0.63, 0.60, and 0.51 for the brown, magenta, and yellow modules, respectively) and differentially expressed in 95 cortex samples (FDR < 0.05; for the brown module) and/or 47 younger (15 – 30 years) cortex samples (P < 0.05; for the yellow and magenta modules as they show stronger ASD-association in younger individuals). We also performed manual inspection to ensure relatively uniform 5′ termini and precise 5′ sequence prediction, as the seed region (nucleotides 2–8) is critical for target prediction by TargetScan. In a few cases where two miRNAs are 3′ isoforms, we used only one isoform for target prediction to prevent duplicate predictions. Using these criteria, 10 down-regulated miRNAs, 24 up-regulated miRNAs, 5 brown module miRNAs, 4 magenta module miRNAs, and 7 yellow module miRNAs were selected for target prediction.
We used two approaches to select top targets as recommended by the developers of the TargetScan algorithm6,20,21. First, we identified all predicted miRNA target sites in a given mRNA 3′ UTR and calculated the summed context+ score for all sites, as miRNA targeting at different sites has been shown to have non-cooperative effects in most cases20. We then selected the strongest targets that are hit by two or more miRNAs in each group (≥ 2 for down-regulated miRNAs or miRNAs in the brown, magenta, and yellow modules, ≥ 4 for up-regulated miRNAs due to the larger number of miRNAs in this group) and have a summed context+ score of ≤ − 0.2 (the more negative the context+ score, the stronger the predicted targeting efficacy). Second, we selected the top 25% most conserved target sites (based on branch length scores) with context+ score ≤ −0.1. For individual miRNAs, we define “the strongest” targets as those with a summed context+ score of ≤ −0.1, and “the most conserved” targets as those with a branch length score in the top 25% and a context+ score of ≤ −0.05.
Gene set enrichment analysis
Gene set enrichment analyses were performed using two-sided Fisher exact tests with the fisher.test R function, except for enrichment for de novo variants (DNVs). For DNVs, a previous study showed a near linear relationship between gene length and de novo mutation frequency24. Therefore, for assessing enrichment of genes affected by DNVs in miRNA targets, we applied logistic regression, in which the probability of a gene being hit by a specific category of DNVs was coded as a function of gene coding region length covered in exome sequencing and whether the gene belongs to a certain miRNA target group. In addition, we also applied logistic regression to assess enrichment while controlling for gene 3′ UTR length (and also for gene coding region length for DNVs) in Supplementary Fig. 6. P values were FDR adjusted across 10 target groups for each gene list using Benjamini-Hochberg correction. The background gene lists were defined as follows: (1) for the strongest targets of differentially expressed miRNAs and ASD-associated miRNA modules, the intersection between (a) protein-coding genes expressed in the cortex and (b) the targets of all 699 miRNAs that are hit by two or more miRNAs and have a summed context+ score of ≤ − 0.2; (2) for the most conserved targets of differentially expressed miRNAs and ASD-associated miRNA modules, the intersection between (a) protein-coding genes expressed in the cortex and (b) the targets of all 699 miRNAs with a branch length ≥ the lowest branch length for the targets of the ASD-related miRNA groups and a summed context+ score of ≤ − 0.1; (3) for the strongest targets of individual candidate miRNAs, the intersection between (a) protein-coding genes expressed in the cortex and (b) the targets of all 699 miRNAs with a summed context+ score of ≤ − 0.1; (4) for the most conserved targets of individual candidate miRNAs, the intersection between (a) protein-coding genes expressed in the cortex and (b) the targets of all 699 miRNAs with a branch length in the top 25% for each miRNA and a context+ score of ≤ −0.05; (5) for the strongest or the most conserved targets of individual candidate miRNAs that were validated in the hNPCs, the background used in (3) or (4) intersected with all genes expressed in the hNPCs as detected by our RNA-Seq analysis.
Enrichment of common variants from genome-wide association studies
GWAS data for ASD, schizophrenia, and Alzheimer’s disease were obtained from the Autism Genome Resource Exchange/Children’s Hospital Philadelphia (AGRE/CHOP), the Psychiatry Genomics Consortium (PGC), and the International Genomics of Alzheimer’s Project (IGAP), respectively. Linkage disequilibrium (LD)-based SNP clumping was performed using PLINK58 (version 1.07) with the following parameters: --clump-p1 0.001 --clump-p2 0.05 --clump-r2 0.50 --clump-kb 250. LD information was obtained from AGRE or HapMap (release 23). Overlap between disease-associated SNP clumps and miRNA targets plus 20 kb flanking regions was then assessed using INRICH32 with the following parameters: -w 20 -r 10000 -q 5000 -d 0.1 -p 0.1. INRICH takes a genomic permutation approach that accounts for linkage disequilibrium, SNP number and density, and gene density to calculate empirical P-values for each gene set32. It performs multiple testing correction via a second, nested round of permutation to assess the null distribution of the minimum empirical P-value across all tested gene-sets32. We used the same background gene lists as in gene set enrichment analysis.
Gene ontology analysis
Gene ontology analysis was performed using GO-Elite (version 1.2.5), which uses a Z-score approximation of the hypergeometric distribution to assess term enrichment, with default settings and 5000 permutations62. False-discovery rate adjusted P-values were calculated using Benjamini-Hochberg correction. We used the same background gene lists as in gene set enrichment analysis.
Quantitative RT-PCR
200 ng total RNA treated with RNase-free DNase I (Qiagen) was reverse-transcribed using the miScript II RT Kit (Qiagen). Real-time PCR was performed using the miScript Primer Assays (Qiagen) and miScript SYBR Green PCR Kit (Qiagen) on a Roche LightCycler 480 instrument. Human RNU6B was used as internal control.
miRNA overexpression in hNPCs and RNA-Seq
Primary human neural progenitor cells (hNPCs) were generated in a previous study and cultured as described63. The cells were free of mycoplasma contamination based on DAPI staining. At the fourth passage, hNPCs were seeded in 6-well plates at 250,000 cells/well. 24 hrs later, mimics of mature miRNAs (GE Healthcare) were transfected at a final concentration of 50 nM using the HiPerFect Transfection Reagent (Qiagen). The miRDIAN microRNA Mimic Negative Control #1 (GE Healthcare), which is based on cel-miR-67 and has minimal sequence identity with human miRNAs, was used as negative control. Transfection for each miRNA mimic was performed in triplicate. 48 hrs after transfection, total RNA was extracted using the miRNeasy kit (Qiagen). RNA integrity number (RIN) was measured using an Agilent Bioanalyzer and all samples had a RIN > 9. Overexpression of miRNAs was confirmed with quantitative RT-PCR. 1.5 μg total RNA was then converted to mRNA libraries using the Illumina TruSeq Stranded mRNA Library Preparation Kit with poly-A selection. ERCC ExFold RNA Spike-In Mixes (Life Technologies) were added as internal controls. Libraries were then validated on an Agilent 2200 TapeStation system and quantified with the Quant-iT PicoGreen assay (Life Technologies). 12 libraries barcoded with Illumina TruSeq indexes were pooled into one lane and sequenced 3 times on an Illumina HiSeq2500 instrument using the rapid run mode with 69 bp paired-end reads.
After demultiplexing with CASAVA v1.8.2 (Illumina), reads were mapped to the GRCh37.75 reference genome using TopHat264. Sequencing quality was then examined using Picard Tools version 1.128 (commands CollectMultipleMetrics, CollectRnaSeqMetrics, and CollectGcBiasMetrics) and the flagstat command in SAMtools65 (version 1.2). 41.0 – 77.9 million QC-passed reads were mapped to the reference genome, with 84.8% – 87.6% mapped to mRNAs, for each sample. Gene expression levels were then quantified using HTSeq66 version 0.6.1 with a union exon model. Genes with 10 or more counts in at least 2 samples in any miRNA overexpression or negative control group were kept for further analysis. Gene expression levels in samples within the same group were highly correlated (R2 ≥ 0.99). The expression data were then normalized using the DESeq R package for library size53 and/or the CQN R package for GC content54. Differential gene expression analysis was performed using one-sided t-test assuming unequal variance. Uncorrected P values were used to define differentially expressed genes, as the sample size was small (n = 3 for each group).
Protein-protein interaction (PPI) analysis
PPI analysis was performed using DAPPLE67 v2.0 (http://www.broadinstitute.org/mpg/dapple/dappleTMP.php), which uses the InWeb database68 and applies a within-degree within node permutation methods. 10,000 permutations were used.
Summary of statistical methods
Blinding
Tissue dissection, RNA extraction, library preparation, and sequencing were performed blind to all metadata information about the samples. Data analysis was not performed blind to metadata information about the samples.
Randomization
Tissue dissection, library preparation, and sequencing were randomized for balance of diagnostic status, age, sex, and brain region.
Sample sizes
No statistical methods were used to pre-determine sample sizes as effect sizes were not known a priori, but our sample sizes are larger than those reported in previous publications that detected mRNA changes (ref. 41, 42, 43).
Parametric tests
For DGE analyses using linear mixed-effects and linear models (Fig. 1b–1c, Supplementary Fig. 3a–3e, Supplementary Table 2), normality was not formally tested for each miRNA. For the main DGE analysis of 95 cortex samples, we also computed permutation-based P values and found that the P value rankings of miRNAs were highly concordant with those observed in the original sample set (Pearson’s R = 0.99, P < 2.2 × 10−16). For calculation of Pearson correlations (Fig. 1c, 2b–2d, 4a–4d, Supplementary Fig. 2a, 2d, 3a–3e, 9a–9c), normality was not formally tested. One-sided t-tests were used for Fig. 6a, 7a, 8a–8c, and Supplementary Fig. 5 because the distributions are approximately normal and sample sizes were reasonably large (n = 88–11695). For Supplementary Fig. 3f–3g, normality was tested by the Shapiro-Wilk test. All groups except for the CTL group for hsa-miR-10a-5p (P = 0.005) and the ASD group for hsa_can_115-m (P = 0.03) meet the normality assumption (P > 0.05). Two-sided t-tests were used for all groups. For differential gene expression analysis after miRNA over-expression in hNPCs (Supplementary Table 6), one-sided t-tests were used but normality was not formally tested for each gene. For all t-tests, equal variances were not formally tested, and so all tests were performed assuming unequal variance. For DGE analyses using linear mixed-effects and linear models, equal variances were not formally tested for each miRNA.
Non-parametric tests
One-tailed Wilcoxon rank sum tests were performed in Supplementary Fig. 8a–8h because the distributions do not appear to be normal. A Supplementary Methods Checklist is available.
Code availability
The R code for the DGE analysis using a linear mixed-effects model and the WGCNA analysis is provided in Supplementary Software.
Data availability
Brain sample metadata, miRNA raw read counts, miRNA DGE analysis data, miRNA WGCNA data, and mRNA DGE analysis data following miRNA over-expression in hNPCs are provided in Supplementary Tables 1–2, 5. Raw small RNA-Seq data from brain samples have been deposited to the PsychENCODE Knowledge Portal (doi:10.7303/syn6133183). Raw RNA-Seq data from hNPCs following miRNA over-expression are available from the corresponding author upon request.
Supplementary Material
Acknowledgments
This work was supported by US National Institutes of Health grant R01MH094714 to D.H.G and is part of the PsychEncode Consortium. We would also like to acknowledge the BrainSpan Consortium principal investigator N. Sestan (Yale University) for providing BrainSpan microRNA data and Jing Ou for excellent technical assistance.
Footnotes
Accession codes
Raw small RNA-Seq data have been deposited to the PsychENCODE Knowledge Portal (doi:10.7303/syn6133183).
AUTHOR CONTRIBUTIONS
N.N.P. and T.G.B. performed brain sample dissections. Y.E.W. performed the other experiments and data analyses. N.N.P. provided code for differential gene expression, co-expression, and enrichment analyses. D.H.G. provided guidance and oversight on all experiments and analyses. Y.E.W. and D.H.G wrote the manuscript.
COMPETING FINANCIAL INTERESTS
The authors declare no competing financial interests.
References
- 1.Bourgeron T. From the genetic architecture to synaptic plasticity in autism spectrum disorder. Nat Rev Neurosci. 2015;16:551–563. doi: 10.1038/nrn3992. [DOI] [PubMed] [Google Scholar]
- 2.Geschwind DH, State MW. Gene hunting in autism spectrum disorder: on the path to precision medicine. Lancet Neurol. 2015;14:1109–1120. doi: 10.1016/S1474-4422(15)00044-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Gaugler T, et al. Most genetic risk for autism resides with common variation. Nat Genet. 2014;46:881–885. doi: 10.1038/ng.3039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Geschwind DH, Flint J. Genetics and genomics of psychiatric disease. Science. 2015;349:1489–1494. doi: 10.1126/science.aaa8954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ha M, Kim VN. Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol. 2014;15:509–524. doi: 10.1038/nrm3838. [DOI] [PubMed] [Google Scholar]
- 6.Friedman RC, Farh KKH, Burge CB, Bartel DP. Most mammalian mRNAs are conserved targets of microRNAs. Genome Research. 2008;19:92–105. doi: 10.1101/gr.082701.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Im HI, Kenny PJ. MicroRNAs in neuronal function and dysfunction. Trends Neurosci. 2012;35:325–334. doi: 10.1016/j.tins.2012.01.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Amaral DG, Schumann CM, Nordahl CW. Neuroanatomy of autism. Trends Neurosci. 2008;31:137–145. doi: 10.1016/j.tins.2007.12.005. [DOI] [PubMed] [Google Scholar]
- 9.Friedlander MR, Mackowiak SD, Li N, Chen W, Rajewsky N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Research. 2011;40:37–52. doi: 10.1093/nar/gkr688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Friedländer MR, et al. Evidence for the biogenesis of more than 1,000 novel human microRNAs. Genome Biol. 2014;15:R57. doi: 10.1186/gb-2014-15-4-r57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Voineagu I, et al. Transcriptomic analysis of autistic brain reveals convergent molecular pathology. Nature. 2011;474:380–384. doi: 10.1038/nature10110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Oldham MC, et al. Functional organization of the transcriptome in human brain. Nat Neurosci. 2008;11:1271–1282. doi: 10.1038/nn.2207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Parikshak NN, et al. Integrative functional genomic analyses implicate specific molecular pathways and circuits in autism. Cell. 2013;155:1008–1021. doi: 10.1016/j.cell.2013.10.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Zhang B, Horvath S. A general framework for weighted gene co-expression network analysis. Stat Appl Genet Mol Biol. 2005;4 doi: 10.2202/1544-6115.1128. Article17. [DOI] [PubMed] [Google Scholar]
- 15.Parikshak NN, Gandal MJ, Geschwind DH. Systems biology and gene networks in neurodevelopmental and neurodegenerative disorders. Nat Rev Genet. 2015;16:441–458. doi: 10.1038/nrg3934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Langfelder P, Luo R, Oldham MC, Horvath S. Is my network module preserved and reproducible? PLoS Comput Biol. 2011;7:e1001057. doi: 10.1371/journal.pcbi.1001057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Arbiza L, et al. Genome-wide inference of natural selection on human transcription factor binding sites. Nat Genet. 2013;45:723–729. doi: 10.1038/ng.2658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ronan JL, Wu W, Crabtree GR. From neural development to cognition: unexpected roles for chromatin. Nat Rev Genet. 2013;14:347–359. doi: 10.1038/nrg3413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Lewis BP, Burge CB, Bartel DP. Conserved Seed Pairing, Often Flanked by Adenosines, Indicates that Thousands of Human Genes are MicroRNA Targets. Cell. 2005;120:15–20. doi: 10.1016/j.cell.2004.12.035. [DOI] [PubMed] [Google Scholar]
- 20.Grimson A, et al. MicroRNA Targeting Specificity in Mammals: Determinants beyond Seed Pairing. Molecular Cell. 2007;27:91–105. doi: 10.1016/j.molcel.2007.06.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Garcia DM, et al. Weak seed-pairing stability and high target-site abundance decrease the proficiency of. Nature Structural & Molecular Biology. 2011;18:1139–1146. doi: 10.1038/nsmb.2115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hsu SD, et al. miRTarBase update 2014: an information resource for experimentally validated miRNA-target interactions. Nucleic Acids Research. 2014;42:D78–85. doi: 10.1093/nar/gkt1266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Basu SN, Kollu R, Banerjee-Basu S. AutDB: a gene reference resource for autism research. Nucleic Acids Research. 2009;37:D832–6. doi: 10.1093/nar/gkn835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Iossifov I, et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature. 2014;515:216–221. doi: 10.1038/nature13908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Darnell JC, et al. FMRP Stalls Ribosomal Translocation on mRNAs Linked to Synaptic Function and Autism. Cell. 2011;146:247–261. doi: 10.1016/j.cell.2011.06.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Bayés A, et al. Characterization of the proteome, diseases and evolution of the human postsynaptic density. Nat Neurosci. 2011;14:19–21. doi: 10.1038/nn.2719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Kang HJ, et al. Spatio-temporal transcriptome of the human brain. Nature. 2011;478:483–489. doi: 10.1038/nature10523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Sanders SJ, et al. Insights into Autism Spectrum Disorder Genomic Architecture and Biology from 71 Risk Loci. Neuron. 2015;87:1215–1233. doi: 10.1016/j.neuron.2015.09.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Anney R, et al. Individual common variants exert weak effects on the risk for autism spectrum disorders. Hum Mol Genet. 2012;21:4781–4792. doi: 10.1093/hmg/dds301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Wang K, et al. Common genetic variants on 5p14.1 associate with autism spectrum disorders. Nature. 2009;459:528–533. doi: 10.1038/nature07999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Cross-Disorder Group of the Psychiatric Genomics Consortium. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet. 2013;381:1371–1379. doi: 10.1016/S0140-6736(12)62129-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lee PH, O’Dushlaine C, Thomas B, Purcell SM. INRICH: interval-based enrichment analysis for genome-wide association studies. Bioinformatics. 2012;28:1797–1799. doi: 10.1093/bioinformatics/bts191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ripke S, et al. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014;511:421–427. doi: 10.1038/nature13595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Lambert JC, et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet. 2013;45:1452–1458. doi: 10.1038/ng.2802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Hirokawa N, Niwa S, Tanaka Y. Molecular Motors in Neurons: Transport Mechanisms and Roles in Brain Function, Development, and Disease. Neuron. 2010;68:610–638. doi: 10.1016/j.neuron.2010.09.039. [DOI] [PubMed] [Google Scholar]
- 36.Lasiecka ZM, Winckler B. Mechanisms of polarized membrane trafficking in neurons -- focusing in on endosomes. Mol Cell Neurosci. 2011;48:278–287. doi: 10.1016/j.mcn.2011.06.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wan QF, et al. SV2 acts via presynaptic calcium to regulate neurotransmitter release. Neuron. 2010;66:884–895. doi: 10.1016/j.neuron.2010.05.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Redies C, Hertel N, Hübner CA. Cadherins and neuropsychiatric disorders. Brain Research. 2012;1470:130–144. doi: 10.1016/j.brainres.2012.06.020. [DOI] [PubMed] [Google Scholar]
- 39.Wong RWC, Guillaud L. The role of epidermal growth factor and its receptors in mammalian CNS. Cytokine Growth Factor Rev. 2004;15:147–156. doi: 10.1016/j.cytogfr.2004.01.004. [DOI] [PubMed] [Google Scholar]
- 40.Turner CA, Akil H, Watson SJ, Evans SJ. The Fibroblast Growth Factor System and Mood Disorders. Biological Psychiatry. 2006;59:1128–1135. doi: 10.1016/j.biopsych.2006.02.026. [DOI] [PubMed] [Google Scholar]
- 41.Mellios N, Sur M. The Emerging Role of microRNAs in Schizophrenia and Autism Spectrum Disorders. Front Psychiatry. 2012;3:39. doi: 10.3389/fpsyt.2012.00039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Rani N, et al. A Primate lncRNA Mediates Notch Signaling during Neuronal Development by Sequestering miRNA. Neuron. 2016;90:1174–1188. doi: 10.1016/j.neuron.2016.05.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Mundalil Vasu M, et al. Serum microRNA profiles in children with autism. Mol Autism. 2014;5:40. doi: 10.1186/2040-2392-5-40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Mor M, Nardone S, Sams DS, Elliott E. Hypomethylation of miR-142 promoter and upregulation of microRNAs that target the oxytocin receptor gene in the autism prefrontal cortex. Mol Autism. 2015;6:46. doi: 10.1186/s13229-015-0040-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Geschwind DH, Rakic P. Cortical evolution: judge the brain by its cover. Neuron. 2013;80:633–647. doi: 10.1016/j.neuron.2013.10.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Geschwind DH. Advances in autism. Annu Rev Med. 2009;60:367–380. doi: 10.1146/annurev.med.60.053107.121225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Mariani J, et al. FOXG1-Dependent Dysregulation of GABA/Glutamate Neuron Differentiation in Autism Spectrum Disorders. Cell. 2015;162:375–390. doi: 10.1016/j.cell.2015.06.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Gawad C, Koh W, Quake SR. Single-cell genome sequencing: current state of the science. Nat Rev Genet. 2016;17:175–188. doi: 10.1038/nrg.2015.16. [DOI] [PubMed] [Google Scholar]
- 49.Quinn EM, et al. Development of strategies for SNP detection in RNA-seq data: application to lymphoblastoid cell lines and evaluation using 1000 Genomes data. PLoS ONE. 2013;8:e58815. doi: 10.1371/journal.pone.0058815. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Friedländer MR, et al. Discovering microRNAs from deep sequencing data using miRDeep. Nat Biotechnol. 2008;26:407–415. doi: 10.1038/nbt1394. [DOI] [PubMed] [Google Scholar]
- 51.Hackenberg M, Sturm M, Langenberger D, Falcon-Perez JM, Aransay AM. miRanalyzer: a microRNA detection and analysis tool for next-generation sequencing experiments. Nucleic Acids Research. 2009;37:W68–W76. doi: 10.1093/nar/gkp347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Hackenberg M, Rodriguez-Ezpeleta N, Aransay AM. miRanalyzer: an update on the detection and analysis of microRNAs in high-throughput sequencing experiments. Nucleic Acids Research. 2011;39:W132–W138. doi: 10.1093/nar/gkr247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Anders S, Huber W. Differential expression analysis for sequence count data. Genome Biol. 2010;11:R106. doi: 10.1186/gb-2010-11-10-r106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Hansen KD, Irizarry RA, Wu Z. Removing technical variability in RNA-seq data using conditional quantile normalization. Biostatistics. 2012;13:204–216. doi: 10.1093/biostatistics/kxr054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Leek JT, Johnson WE, Parker HS, Jaffe AE, Storey JD. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics. 2012;28:882–883. doi: 10.1093/bioinformatics/bts034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Miller JA, Horvath S, Geschwind DH. Divergence of human and mouse brain transcriptome highlights Alzheimer disease pathways. Proc Natl Acad Sci U S A. 2010;107:12698–12703. doi: 10.1073/pnas.0914257107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics. 2008;9:559. doi: 10.1186/1471-2105-9-559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Purcell S, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. The American Journal of Human Genetics. 2007;81:559–575. doi: 10.1086/519795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Langfelder P, Zhang B, Horvath S. Defining clusters from a hierarchical cluster tree: the Dynamic Tree Cut package for R. Bioinformatics. 2008;24:719–720. doi: 10.1093/bioinformatics/btm563. [DOI] [PubMed] [Google Scholar]
- 60.Csardi G, Nepusz T. The igraph software package for complex network research. InterJournal Complex Systems. 2006:1695. [Google Scholar]
- 61.Quinlan AR, Hall IM. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics. 2010;26:841–842. doi: 10.1093/bioinformatics/btq033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Zambon AC, et al. GO-Elite: a flexible solution for pathway and ontology over-representation. Bioinformatics. 2012;28:2209–2210. doi: 10.1093/bioinformatics/bts366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Stein JL, et al. A quantitative framework to evaluate modeling of cortical development by neural stem cells. Neuron. 2014;83:69–86. doi: 10.1016/j.neuron.2014.05.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Kim D, et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013;14:R36. doi: 10.1186/gb-2013-14-4-r36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Li H, et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics. 2009;25:2078–2079. doi: 10.1093/bioinformatics/btp352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Anders S, Pyl PT, Huber W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31:166–169. doi: 10.1093/bioinformatics/btu638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Rossin EJ, et al. Proteins Encoded in Genomic Regions Associated with Immune-Mediated Disease Physically Interact and Suggest Underlying Biology. PLoS Genet. 2011;7:e1001273. doi: 10.1371/journal.pgen.1001273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Lage K, et al. A human phenome-interactome network of protein complexes implicated in genetic disorders. Nat Biotechnol. 2007;25:309–316. doi: 10.1038/nbt1295. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Brain sample metadata, miRNA raw read counts, miRNA DGE analysis data, miRNA WGCNA data, and mRNA DGE analysis data following miRNA over-expression in hNPCs are provided in Supplementary Tables 1–2, 5. Raw small RNA-Seq data from brain samples have been deposited to the PsychENCODE Knowledge Portal (doi:10.7303/syn6133183). Raw RNA-Seq data from hNPCs following miRNA over-expression are available from the corresponding author upon request.