Abstract
Ion channels form the basis for cellular electrical signaling. Despite the scores of genetically identified ion channels selective for other monatomic ions, only one type of proton-selective ion channel has been found in eukaryotic cells. By comparative transcriptome analysis of mouse taste receptor cells, we identified Otopetrin1 (OTOP1), a protein required for development of gravity-sensing otoconia in the vestibular system, as forming a proton-selective ion channel. We found that murine OTOP1 is enriched in acid-detecting taste receptor cells and is required for their Zn2+-sensitive proton conductance. Two related murine genes, Otop2 and Otop3, and a Drosophila ortholog also encode proton channels. Evolutionary conservation of the gene family and its widespread tissue distribution suggest a broad role for proton channels in physiology and pathophysiology.
Ion channels include a large and diverse group of membrane proteins that rapidly, and with great selectivity, move ions across the cell membrane, performing crucial roles in cell signaling and homeostasis (1). Ion channels selective for each of the physiologically relevant ions, Na+, K+, Ca2+ and Cl-, have been described at the molecular and structural levels (2, 3), but only a few types of proton-selective ion channels (proton channels) have been described (4). One is the 96-amino acid M2 protein of influenza A, which conducts protons into the virion interior, an essential step in the replication of the virus (5). The only one identified in eukaryotes is the voltage-gated Hv1 (6-8), which is present in immune cells, where it extrudes protons into the phagosome to inactivate infectious agents (9). Functional evidence indicates that ion channels that selectively transport protons into eukaryotic cells must also exist. For example, in acid-sensing taste receptor cells (TRCs), an inward-conducting Zn2+-sensitive proton current that is biophysically distinct from currents carried by Hv1 has been described (10, 11).
To identify candidates encoding such a proton channel, we compared the transcriptome of mouse TRCs positive for the inward-conducting Zn2+-sensitive proton current (PKD2L1 cells) with that of TRCs that lack the current (TRPM5 cells; Fig 1A). We selected genes that were enriched in PKD2L1 cells and that encoded poorly or uncharacterized transmembrane proteins (Fig. 1A and Table S1) (see methods). We expressed the candidates in human embryonic kidney (HEK-293) cells or Xenopus oocytes and measured ionic currents in response to lowering the extracellular pH (pHo) in the absence of extracellular Na+. Of the forty-one cDNAs tested, only Otopetrin1 (Otop1), which encodes a protein with 12 predicted transmembrane domains (12), generated large Zn2+-sensitive inward currents in response to extracellular acidification (Fig 1B).
Fig. 1.
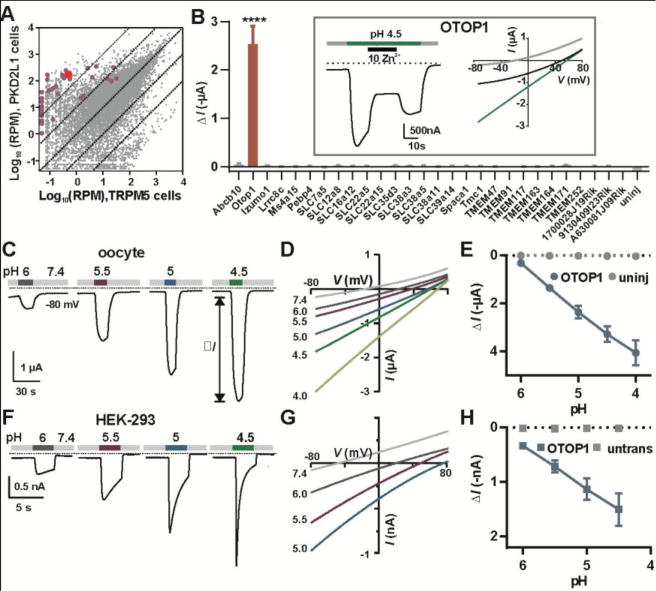
Expression analysis of taste-cell enriched genes identifies OTOP1 as a novel proton channel. (A) Transcriptome profiling of PKD2L1 and TRPM5 taste receptor cells (Each data point represents the average of 5 replicates). Genes tested by electrophysiology are highlighted in magenta or red (OTOP1). RPM, reads per million. (B) Magnitude of currents evoked in response to pH 4.5 Na+-free solution in Xenopus oocytes expressing the genes indicated (Vm = −80mV; data are mean ± SEM, n = 3 – 37; for OTOP1, n = 5). ****, P < 0.0001 compared to uninjected oocytes (n = 3). One-way ANOVA with Bonferroni correction. Inset: currents evoked in an OTOP1-expressing oocyte to the acid stimulus at Vm = −80mV (left) and the current-voltage relationship before (gray), during acid application (green) and during Zn2+ application (black). (C) Current measured by two-electrode voltage clamp in a Xenopus oocyte expressing OTOP1 in response to Na+-free extracellular solutions with pHo as indicated (Vm = −80 mV). (D) I-V relation of the current in (A) from voltage ramps (1V/s). (E) Evoked current (ΔI; mean ± SEM) as a function of pH in Xenopus oocytes expressing OTOP1 (blue circle; n = 4) and uninjected oocytes (grey circles; n = 4). (F) Currents measured by whole-cell patch clamp recording in a HEK-293 cell expressing OTOP1 in Na+-free extracellular solutions (pHi = 7.3, Vm = −80 mV). (G) I-V relation of currents in OTOP1-expressing HEK-293 cell from experiments as in (G) with voltage ramps (1V/s). (H) Evoked currents (ΔI; mean ± SEM) as a function of pH in HEK-293 cells expressing OTOP1 (blue squares; n = 5) and untransfected cells (grey squares; n = 3).
We characterized functional properties of OTOP1 expressed in Xenopus oocytes. Unless otherwise noted, the extracellular solution used in recordings was Na+-free [N-methyl-D-glucamine (NMDG+)-based]. OTOP1 currents increased monotonically as pHo was lowered (Fig 1, C through E) and the reversal potential (Erev) shifted toward more positive voltages (Fig. S1A). The currents showed a small time-dependent change in amplitude in response to hyperpolarizing voltage steps, indicating that gating of OTOP1 is mildly voltage-sensitive (Fig. S1, B and C).
OTOP1 also generated an ionic current in HEK-293 cells (Fig 1, F through H). An N-terminal YFP-tagged protein confirmed the presence of OTOP1 at the cell surface (Fig. S2A). Lowering pHo elicited large inward currents in OTOP1-expressing HEK-293 cells and, as in oocytes, the current magnitude increased monotonically with pHo (Fig. 1, F through H). OTOP1 currents in HEK-293 cells decayed within seconds, with faster kinetics observed in response to more acidic stimuli (Fig. 1F). The decay of the currents is likely to be due, in part, to a reduction in the driving force as protons accumulate in the cytosol. For a 15-μm diameter cell (1767 fL volume), a H+ current of 1000 pA flowing for 1 second will increase the total (bound + free) intracellular concentration of H+ by ~6 mM (4). We confirmed that OTOP1 mediated flux of protons into the cell cytosol with the membrane-permeant pH indicator pHrodo Red. In OTOP1-expressing HEK-293 cells, but not in mock-transfected cells, lowering extracellular pH from 7.4 to 5.0 caused a large increase in emission of pHrodo Red (Fig 2, A and B), corresponding to a large change in intracellular pH (Fig. S2, B and C).
Fig 2.
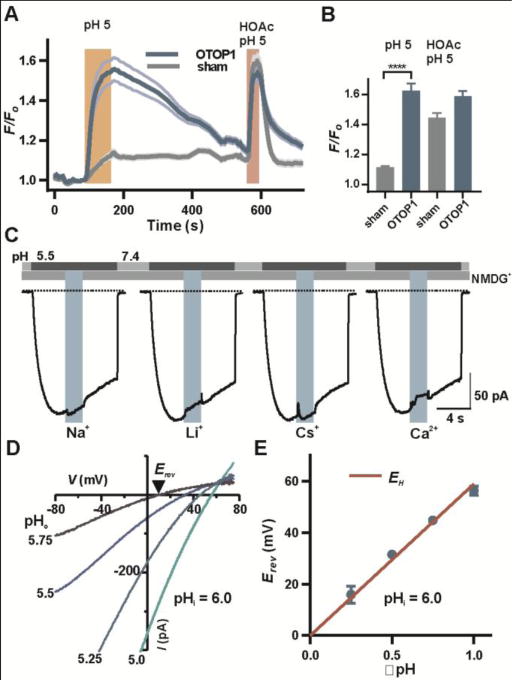
Selectivity of OTOP1 for protons. (A) Fluorescence emission of the pH indicator pHrodo Red in HEK-293 cells expressing OTOP1 (n = 9) and sham-transfected cells (n = 11; mean ± SEM) in response to the stimuli indicated. Similar results were obtained in 3 replicates. (B) Average data (mean ± SEM; n = 28-29 cells) were analyzed by 2-tailed t-test. ****: P < 0.0001. HOAc, acetic acid, which shuttles protons across membranes (4) served as a positive control. (C) OTOP1 currents in HEK-293 cells were evoked in response to a pH 5.5 solution with Na+, Li+ or Cs+ (160 mM each) or Ca2+ (40 mM) replacing NMDG+ in the extracellular solution as indicated (Vm = −80 mV). Percentage change in currents was 0.4 ± 0.7, n = 8, 2.7 ± 0.7, n = 8, 2.4 ± 0.5, n = 8, 3.6 ± 1.7, n = 7 for each ion replacement respectively. (D) Isolated OTOP1 currents in response to voltage ramps (1 V/s) at varying extracellular pH (pHi = 6.0; Zn2+-sensitive component is shown; see Fig. S5 and methods). (E) Erev as a function of ΔpH (pHi-pHo) from experiments as in (D). The red line is EH. The data were fit by linear regression with a slope of 53 mV/ΔpH and a Y intercept of 3.6 mV (R2 = 0.99).
Hv1 and M2 are highly selective for protons, present in high nanomolar concentrations, over other cations whose concentrations are a million times higher (4, 13). To determine if OTOP1 is similarly proton-selective, we evoked OTOP1 currents by lowering pHo from 7.4 to 5.5 and measured the effect of exchanging NMDG+ in the extracellular solution for equimolar concentrations of Na+, Cs+, or Li+ or isosmotic concentrations of Ca2+ (Fig 2C). In all cases, the observed change in current magnitude was less than 4%, indicating that OTOP1 is not appreciably permeable to these ions. Similar experiments showed that OTOP1 is not appreciably permeable to K+ (Fig. S3). To directly assess the selectivity of the channel for protons, we measured the potential at which the current reversed direction, the reversal potential (Erev), as a function of the H+ gradient (ΔpH = pHi – pHo). To study a predominantly OTOP1 current, we applied Zn2+ at a concentration that selectively and fully blocked the OTOP1 current in HEK-293 cells and focused on the Zn2+-sensitive component of the current (Fig. S4 and Fig. S5, A and B). We limited H+ accumulation by setting pHi at 6.0 and holding the membrane potential at Erev (Fig. S5A; see methods). Under these conditions, Erev closely followed the Nernst prediction for an H+-selective ion channel (Fig 2, D and E). To determine the selectivity of OTOP1 for H+ relative to Na+ and Cl-, we measured Erev upon replacement of NMDG+ by Na+ and with high and low concentrations of Cl- in the extracellular solution. In no case did we observe any change in Erev (Fig. S5C). Assuming a change of Erev of less than 5 mV, which would have been detectible, we used the Goldman-Hodgkin-Katz (GHK) equation to calculate the selectivity of OTOP1 for H+ relative to Na+ at greater than 2×105-fold and H+ to Cl- at greater than 1×105–fold (13).
OTOP1 is a member of the otopetrin family, which appears to be evolutionarily conserved from nematodes to humans (12, 14) (Fig. 3A). We confirmed that human OTOP1 (hOTOP1) forms a channel with similar properties to murine OTOP1 (Fig. S6). Murine OTOP2 and OTOP3 share 30% to 34% amino acid identity with murine OTOP1 (Fig. S7). Each shows a distinctive pattern of expression. Otop1 is expressed in vestibular and taste cells, brown adipose tissue (15), heart, uterus, dorsal root ganglion, adrenal gland, mammary gland, and stimulated mast cells, whereas Otop2 expression is highest in stomach, testis, and olfactory bulb, and Otop3 is expressed in epidermis, small intestine, stomach and retina (Fig 3B; (16)). When expressed in Xenopus oocytes OTOP2 and OTOP3 both generated large currents in response to lowering pHo in a Na+-free solution. Compared with OTOP1 and OTOP3, OTOP2 currents behaved anomalously; currents saturated at ~pH 5 and Erev shifted little over a range of pH 4 to 6 (Fig 3, C through E, Fig. S8A). OTOP2 currents measured in HEK-293 cells had similar properties (Fig. S9). Like OTOP1, OTOP3 showed evidence of selectivity for H+; the magnitude of OTOP3 currents increased linearly as a function of pHo over the entire pH range tested (Fig 3, F through H) and Erev shifted 46.3 mV/log[H+], close to the value of 58 mV/log[H+] expected for a proton-selective ion channel (Fig. S8C). In response to hyperpolarizing voltage steps, OTOP2 and OTOP3 currents showed evidence of mild (OTOP3) or no (OTOP2) voltage-dependence (Fig. S8, B and D). When expressed in HEK-293 cells and assessed with microfluorimetry, both OTOP2 and OTOP3 conducted protons into the cell cytosol in response to lowering pHo (Fig. S10), providing evidence that, like OTOP1, they form proton channels.
Fig 3.
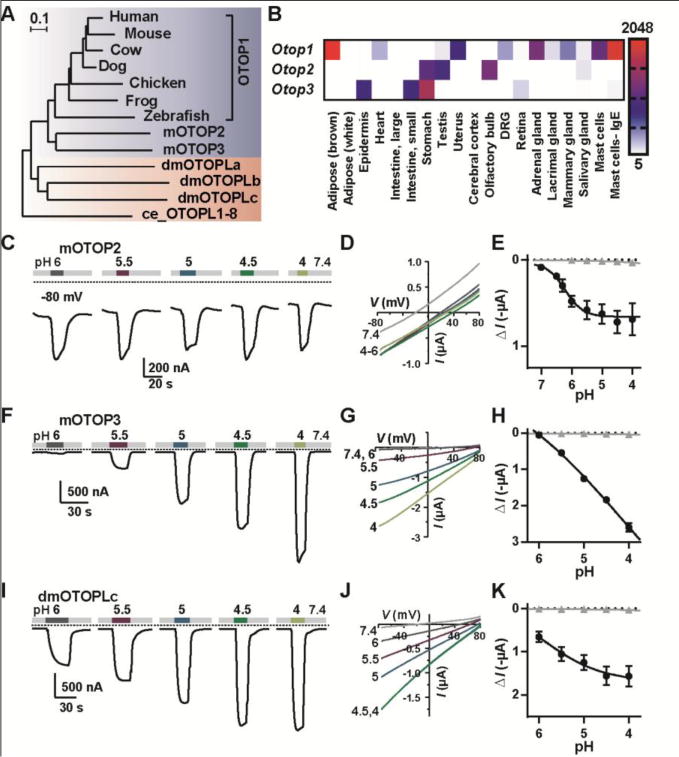
An evolutionarily conserved family of genes, expressed in diverse tissues and encoding proton channels. (A) Maximum-likelihood phylogenetic tree from the multi-sequence alignment of 13 otopetrin domain proteins. Scale bar is amino acid substitutions per site. (B) Distribution of Otops in selected murine tissues from microarray data (16). Scale represents expression level in arbitrary units (mean ± SEM, n = 2). (C, F, I) Representative traces (Vm = −80 mV) showing currents evoked in Xenopus oocytes expressing OTOP2, OTOP3 or dmOTOPLc in response to varying pHo of the Na+-free extracellular solution. (D, G, J) I-V relationship (from voltage ramps at 1 V/s) from experiments as in (C, F, I). (E, H, K) The average current induced at Vm = −80 mV (ΔI) as a function of pH for oocytes expressing each of the channels (black circles; mean ± SEM, n = 3–7) and for uninjected oocytes (gray triangles, mean ± SEM, n = 3).
There are three genes in the genome of Drosophila melanogaster that encode proteins that appear to be evolutionarily related to mOTOP1 (12, 14) (Fig 3A). The transcript CG42265 encodes dmOTOPLc, a protein of 1576 amino acids that over the region of similarity bears 14.1% amino acid identity with OTOP1. Despite the modest level of conservation, when expressed in Xenopus oocytes, dmOTOPLc conducted large currents in response to decreasing extracellular pH, indicating that it too forms a proton channel (Fig 3, I through K). The shallow relation between the current amplitude and pH may endow the channel with a broader dynamic range.
OTOP1 is required for the development of otoconia, calcium carbonate-based structures that sense gravity and acceleration in the vestibular system. Two mutations of OTOP1, tilted (tlt) and mergulhador (mlh; Fig. S11A), lead to vestibular dysfunction in mice (14). These mutations affect trafficking of the protein to the cell surface in vestibular supporting cells (17). Mutant channels expressed in Xenopus oocytes, produced smaller currents but otherwise had similar functional properties, such as sensitivity to Zn2+ (Fig. S11, B and C), to wild-type OTOP1. This reduction in current magnitude may contribute to the vestibular dysfunction.
Finally, we sought to determine if OTOP1 constitutes the proton current in acid-sensing taste receptor cells (10, 11). We confirmed that in single cell transcriptome data, Otop1 was expressed in all PKD2L1 cells (19/19) implicated in sour transduction (18), whereas Otop2 and Otop3 were expressed in much lower amounts, and none of the three Otop transcripts were detected in TRPM5 cells, which lack proton currents (Fig. 4A). By immunocytochemistry, we confirmed that OTOP1 was present in taste cells in mouse circumvallate papillae that express Pkd2l1 (Fig. 4B). To directly determine if OTOP1 contributes to the proton current in taste cells, we measured currents in taste cells from either wild-type mice or mice that were homozygous for the tlt mutation of OTOP1. Mutation of OTOP1 resulted in significantly smaller currents than those measured in taste cells from wildtype mice (Fig 4C and D), over a range of H+ concentrations (Fig 4E and F), indicating that OTOP1 is a component of the proton channel in taste cells. While the contribution of proton currents to acid-sensing/sour taste behavior by mice is still speculative and complicated by contributions from multiple sensory organs and sensory receptors (19), the identification of OTOP1 provides a tool with which to start dissecting this system.
Fig 4.
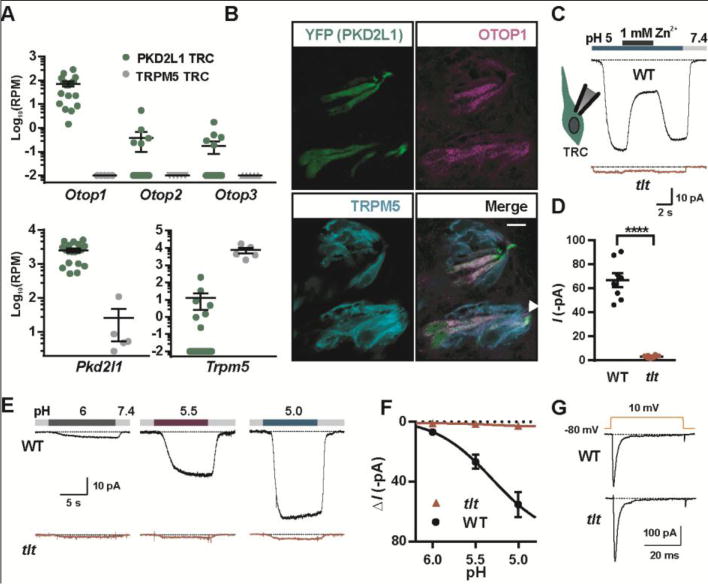
Requirement of OTOP1 for the proton current in taste receptor cells. (A) Read counts per million (RPM) for the genes indicated from RNA-seq data obtained from single PKD2L1 (n = 19) or TRPM5 taste cells (n = 5). 0 RPM was adjusted to 0.01 RPM. (B) Confocal images showing taste buds in the circumvallate papillae from a mouse in which Pkd2l1 drives expression of YFP immunostained with antibodies against YFP (green), OTOP1 (magenta) and TRPM5 (cyan). Scale bar is 10 μM. Arrow indicates taste pore. (C) Current in response to pH 5.0 stimulus in isolated PKD2L1 TRCs from tlt mutant or wildtype mice in NMDG+-based solution (Vm = −80 mV). (D) Average data from experiments as in (C) (****: P < 0.0001 by two tailed t-test, n = 8 cells/genotype). (E) Response of PKD2L1 TRCs to NMDG+-based extracellular solution of varying pH (Vm = −80 mV). (F) Average data from experiments as in (E). (G) Voltage-gated Na+ currents in TRCs from tlt and wildtype mice were indistinguishable (P > 0.05, two-tailed t-test).
Our data show that the Otop gene family encodes a family of ion channels that are unrelated structurally to previously identified ion channels and are highly selective for protons. Unlike Hv1, OTOP1 is only weakly sensitive to voltage. Whether, like the viral proton channel M2 (13), low pH gates OTOP1 is not clear. OTOP channels conduct protons at normal resting potentials and can mediate the entry of protons into cells. Most cells guard against proton entry, which is generally cytotoxic. Thus, we expect OTOP channels are restricted to cell types that use changes in intracellular pH for cell signaling or to regulate biochemical or developmental processes. Along with a role in formation of vestibular otoconia (14), OTOP1 has been shown to protect mice from obesity-induced metabolic dysfunction (15), and it is upregulated in dorsal root ganglion cells in response to cell damage (20). The knowledge that this gene family encodes proton channels can be used to understand its contribution to physiology and disease.
Supplementary Material
One Sentence Summary.
A conserved gene family encodes ion channels that selectively transport H+ across cellular membranes and regulate pH.
Acknowledgments
E.L. and USC have filed a provisional patent application no. 62/537,900 that claims methods of screening molecules that modulate Otopetrin-dependent ion channel activities. We thank S. Rao, L. Goggins and A. Bernanke for technical assistance, J. Bushman for assistance with electrophysiology and D. Arnold, B. Bean, B. Herring and R. Kramer for careful reading of the manuscript. Funding was provided by the National Institutes of Health grants R01DC013741 and R21DC012747 to E.R.L. and R01HG006015 to A.D.S.
Footnotes
References and Notes
- 1.Hille B. Ionic channels of excitable membranes. Sinauer Associates Inc.; Sunderland, MA: 2001. [Google Scholar]
- 2.Catterall WA, Wisedchaisri G, Zheng N. The chemical basis for electrical signaling. Nat Chem Biol. 2017;13:455–463. doi: 10.1038/nchembio.2353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Gouaux E, Mackinnon R. Principles of selective ion transport in channels and pumps. Science. 2005;310:1461–1465. doi: 10.1126/science.1113666. [DOI] [PubMed] [Google Scholar]
- 4.Decoursey TE. Voltage-gated proton channels and other proton transfer pathways. Physiol Rev. 2003;83:475–579. doi: 10.1152/physrev.00028.2002. [DOI] [PubMed] [Google Scholar]
- 5.Pinto LH, Holsinger LJ, Lamb RA. Influenza virus M2 protein has ion channel activity. Cell. 1992;69:517–528. doi: 10.1016/0092-8674(92)90452-i. [DOI] [PubMed] [Google Scholar]
- 6.DeCoursey TE. Hydrogen ion currents in rat alveolar epithelial cells. Biophys J. 1991;60:1243–1253. doi: 10.1016/S0006-3495(91)82158-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ramsey IS, Moran MM, Chong JA, Clapham DE. A voltage-gated proton-selective channel lacking the pore domain. Nature. 2006;440:1213–1216. doi: 10.1038/nature04700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Sasaki M, Takagi M, Okamura Y. A voltage sensor-domain protein is a voltage-gated proton channel. Science. 2006;312:589–592. doi: 10.1126/science.1122352. [DOI] [PubMed] [Google Scholar]
- 9.Morgan D, et al. Voltage-gated proton channels maintain pH in human neutrophils during phagocytosis. Proc Natl Acad Sci U S A. 2009;106:18022–18027. doi: 10.1073/pnas.0905565106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chang RB, Waters H, Liman ER. A proton current drives action potentials in genetically identified sour taste cells. Proc Natl Acad Sci U S A. 2010;107:22320–22325. doi: 10.1073/pnas.1013664107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bushman JD, Ye W, Liman ER. A proton current associated with sour taste: distribution and functional properties. FASEB J. 2015;29:3014–3026. doi: 10.1096/fj.14-265694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hughes I, et al. Identification of the Otopetrin Domain, a conserved domain in vertebrate otopetrins and invertebrate otopetrin-like family members. BMC Evol Biol. 2008;8:41. doi: 10.1186/1471-2148-8-41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Chizhmakov IV, et al. Selective proton permeability and pH regulation of the influenza virus M2 channel expressed in mouse erythroleukaemia cells. J Physiol. 1996;494(Pt 2):329–336. doi: 10.1113/jphysiol.1996.sp021495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hurle B, et al. Non-syndromic vestibular disorder with otoconial agenesis in tilted/mergulhador mice caused by mutations in otopetrin 1. Hum Mol Genet. 2003;12:777–789. doi: 10.1093/hmg/ddg087. [DOI] [PubMed] [Google Scholar]
- 15.Wang GX, et al. Otopetrin 1 protects mice from obesity-associated metabolic dysfunction through attenuating adipose tissue inflammation. Diabetes. 2014;63:1340–1352. doi: 10.2337/db13-1139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Wu C, et al. BioGPS: an extensible and customizable portal for querying and organizing gene annotation resources. Genome Biol. 2009;10:R130. doi: 10.1186/gb-2009-10-11-r130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kim E, et al. Missense mutations in Otopetrin 1 affect subcellular localization and inhibition of purinergic signaling in vestibular supporting cells. Mol Cell Neurosci. 2011;46:655–661. doi: 10.1016/j.mcn.2011.01.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Huang AL, et al. The cells and logic for mammalian sour taste detection. Nature. 2006;442:934–938. doi: 10.1038/nature05084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Liman ER, Zhang YV, Montell C. Peripheral coding of taste. Neuron. 2014;81:984–1000. doi: 10.1016/j.neuron.2014.02.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Reinhold AK, et al. Differential transcriptional profiling of damaged and intact adjacent dorsal root ganglia neurons in neuropathic pain. PLoS One. 2015;10:e0123342. doi: 10.1371/journal.pone.0123342. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


