Abstract
The microbiota of Anopheles mosquitoes interferes with mosquito infection by Plasmodium and influences mosquito fitness, therefore affecting vectorial capacity. This natural barrier to malaria transmission has been regarded with growing interest in the last 20 years, as it may be a source of new transmission-blocking strategies. The last decade has seen tremendous progress in the functional characterisation of the tripartite interactions between the mosquito, its microbiota and Plasmodium parasites. In this review, we provide insights into the effects of the mosquito microbiota on Plasmodium infection and on mosquito physiology, and on how these aspects together influence vectorial capacity. We also discuss three current challenges in the field, namely the need for a more relevant microbiota composition in experimental mosquitoes involved in vector biology studies, for a better characterisation of the non-bacterial microbiota, and for further functional studies of the microbiota present outside the gut.
Keywords: Microbiota, Anopheles, Plasmodium, Experimental models, Vectorial transmission, Colonisation resistance
Background
Plasmodium malaria parasites are transmitted by Anopheles mosquitoes. Female mosquitoes become infected after taking a blood meal on humans carrying Plasmodium pre-sexual stages. Upon entering the gut lumen, parasites undergo sexual reproduction and differentiate into motile forms, ookinetes, within 24 hours. After crossing the gut epithelium, the ookinete develops into an oocyst which undergoes mitoses in the following week and releases sporozoites in the hemolymph. About 10–14 days after the infected blood meal, sporozoites reach the salivary glands. The mosquito then becomes infectious and will inject parasites to humans with its saliva during subsequent bites for the rest of its life.
Ingestion of an infectious blood meal will only result in malaria transmission if the parasite makes it through bottlenecks in the gut and salivary glands and if the mosquito bites humans after the extrinsic incubation period, the time needed for parasites to become infectious. The gut, salivary glands and reproductive organs are colonised by a dynamic microbial community composed by bacteria, viruses and fungi, of which the bacterial part is the best characterised [1–3]. This microbiota impacts disease transmission by interfering with Plasmodium colonisation in the gut and by affecting different aspects of mosquito physiology, notably its lifespan.
As a consequence, microorganisms that colonise the mosquito are regarded as potential tools to reduce malaria transmission. They may be used to shorten the mosquito lifespan or to decrease Plasmodium infection rates, either via natural competition mechanisms [4, 5] or via the production of genetically introduced anti-Plasmodium molecules, so called paratransgenesis [6–8]. One of the main advantages of a strategy based on microbial colonisation is the potential for targeting several species of mosquitoes and Plasmodium at the same time.
During development, the mosquito acquires its microbiota from its mother’s genitalia and from its larval and pupal breeding site. Some of these microorganisms are trans-stadially transmitted to the adult [9, 10], while others are acquired by adults when feeding on different substrates or during mating [11]. The microbiota population is particularly dynamic in the mosquito gut, where it drastically expands after a blood meal. More specifically, this proliferation is observed in the middle region of the gut, the midgut, where the blood is stored during digestion over a 2-day period and where the early stages of Plasmodium development take place. Here, we review the current knowledge on tripartite interactions between the mosquito, its microbiota and Plasmodium and we discuss current challenges of the field.
Effects of the microbiota on vectorial capacity
Since the midgut represents the first and main bottleneck of parasite development, the microbial community present in its lumen has a strong role in the first stages of Plasmodium infection. Several functional studies have investigated the specific role of the midgut microbiota on Plasmodium infection. These studies, performed on five Anopheles species and four Plasmodium species, point to an overall inhibitory effect of the microbiota on the parasite, independent of the species of Anopheles and Plasmodium (Table 1). The effect is, however, highly bacterial strain specific [4, 12–14]. Most bacteria showing an anti-parasitic effect are Gram-negative [12, 14].
Table 1.
Effects of the microbiota or of specific bacterial species/strains on Plasmodium infection in different Anopheles mosquitoes in chronological order
| Mosquito | Parasite | Inhibition | No inhibition | Reference |
|---|---|---|---|---|
| An. stephensi (colony) | P. falciparum | Escherichia coli H243a; Pseudomonas aeruginosaa; Ewingella americanaa | Staphylococcus aureusb; S. epidermidisb; E. coli HB101a | [12] |
| An. stephensi (colony) | P. falciparum | E. coli HS5a; P. aeruginosaa; Serratia marcescensa; Xanthomonas malthophilaa; Cedecea lapageia | [67] | |
| An. albimanus (colony) | P. vivax | S. marcescensa; Enterobacter cloacaea; Enterobacter amnigenus 2a | [4] | |
| An. gambiaec (colony) | P. falciparum | Microbiota; Live and heat-inactivated mixture of S. aureusb + E. colia | [5] | |
| An. gambiaed, An. coluzzii (colonies) | P. berghei | S. aureusb; E. colia; E. cloacaea | [15] | |
| An. gambiaec, An. stephensi (colonies) | P. falciparum | Enterobacter sp. Zambiaa | Bacillus pumilus b | [13] |
| An. gambiaed (colony) | P. yoelii | Microbiota | [68] | |
| An. gambiaed (colony) | P. falciparum | Microbiota | ||
| An. coluzzii (colony) | P. falciparum | E. colia; S. marcescensa; Pseudomonas sutzeria; Comamonas spp.a; Enterobacter spp.a; B. pumilusb | Acinetobacter septicus a | [69] |
| An. stephensi (colony) | P. berghei | S. marcescens HB3a | S. marcescens HB18a | [14] |
| An. dirus (colony) | P. yoelii | Microbiota | [19] | |
| An. gambiaec (colony) | P. falciparum | Chromobacterium sp. Csp_Pa | [22] | |
| An. gambiaec (colony) | P. falciparum | P. putidaa; Pantoea sp.a; S. marcescensa | [16] | |
| An. gambiaee (from field larvae) | P. falciparum | Serratiaa; Methylobacteriuma | [3] |
aGram-negative bacteria
bGram-positive bacteria
cKeele strain, hybrid of An. gambiae and An. coluzzii
dG3 strain, hybrid of An. gambiae and An. coluzzii
eField-collected mosquitoes, not specified if An. gambiae or An. coluzzii
Antiparasitic effects of the microbiota on Plasmodium
The microbiota interferes with Plasmodium colonisation of the mosquito gut through at least two mechanisms: (i) stimulation of the mosquito immune response; and (ii) production of metabolites directly impairing parasite survival (Fig. 1).
Fig. 1.
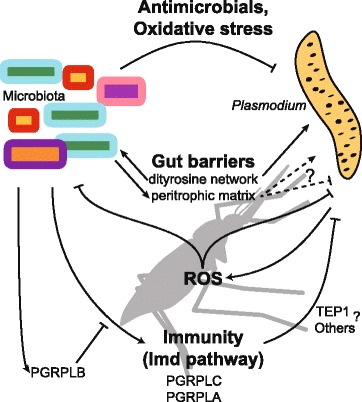
Interactions between the microbiota and Plasmodium in the mosquito midgut. The microbiota affects Plasmodium infection by several mechanisms: (i) Direct impact on parasites via inhibition of its oxidative defence system [13, 20] or by production of uncharacterised antimicrobials [16, 22]. (ii) Stimulation of the NF-κB dependent Immune-deficiency (Imd) pathway, which is regulated by Peptidoglycan Recognition Proteins (PGRPs) and restrains parasite infection [15, 30]. The mechanisms of action of the Imd pathway on parasites are still unclear, they probably include TEP1-dependent and independent components [5, 18, 19]. (iii) Blood meal inducible physical barriers affect gut microbes: a dityrosine network reduces the diffusion of elicitors, thus protecting the microbiota and Plasmodium from immune activation [32] and the microbiota-dependent induction of the peritrophic matrix [24] may have positive and/or negative impacts on parasite infection. Reciprocally, Plasmodium infection inhibits antioxidant enzymes in the mosquito gut, which has been suggested to help parasite infection via a reduction of the mosquito microbiota [21]
The fast multiplication of bacteria in the gut following a blood meal stimulates an immune response which is widely antimicrobial. At the level of the midgut epithelium, this immune response is largely due to the Immune-deficiency (Imd) pathway, which is induced upon detection of the bacterial cell wall peptidoglycan by Peptidoglycan Recognition Protein (PGRP) LC and positively regulated by PGRPLA [15–17]. How the Imd pathway affects the parasite has not yet been characterised, but this pathway has been shown to induce TEP1 expression, and TEP1 has been reported to participate in the microbiota-dependent control of Plasmodium infection [18, 19]. Ookinete numbers are, however, already reduced in the gut epithelium in a microbiota-dependent manner, suggesting that microbiota-induced immunity also affects parasites before the action of the complement system [5].
Some microorganisms isolated from the mosquito gut also produce metabolites that directly affect Plasmodium and reduce its ability to infect the mosquito. Enterobacter Esp_Z is a Gram-negative bacterium isolated from the gut of An. arabiensis mosquitoes that was found to reduce P. falciparum ookinete, oocyst and sporozoite loads in An. gambiae via production of reactive oxygen species and/or inhibition of the oxidative defence system of the parasite [13, 20]. Interestingly, Plasmodium infection has been reported to reciprocally inhibit antioxidant enzymes in the mosquito gut. This may promote parasite infection via a reduction of the mosquito microbiota [21]. Some Serratia marcescens strains and Chromobacterium Csp_P initially isolated from the mosquito midgut reduce P. falciparum infection in An. gambiae and show anti-parasitic activity in vitro by producing one or several uncharacterised metabolites [16, 22]. Moreover, the yeast Wickerhamomyces anomalus was isolated from the midgut of An. stephensi and found to produce a killing toxin with β-1,3-glucanase activity, which inhibits P. berghei ookinetes in vitro [23].
Besides these well-characterised effects of the microbiota on Plasmodium, some effects are indirectly suggested in the literature. For instance, the microbiota is also involved in the synthesis of the peritrophic matrix, a layer composed of chitin and proteins which surrounds the midgut epithelium after blood-feeding and protects the mosquito from the dissemination of midgut bacteria into the body cavity [24]. Plasmodium ookinetes secrete a chitinase required to cross the peritrophic matrix [25, 26], indicating that this barrier, if not impenetrable for malaria parasites, exerts at least some selection pressure on them. From this point-of-view, it may be a colonisation resistance mechanism induced by the microbiota against Plasmodium.
The microbiota may also nutritionally affect Plasmodium in the mosquito gut. Whole genome sequencing of bacterial strains of the mosquito microbiota identified genes involved in the digestion of macromolecules [27]. However, it is yet unknown whether this potential digestive role of the microbiota results in a benefit for the mosquito and for Plasmodium or in nutrient competition. In Drosophila, resistance to virus infection induced by endosymbionts is at least partly due to competition for cholesterol [28]. Nutrient availability is manipulated by Plasmodium in the mosquito gut through overexpression of digestive enzymes, suggesting that the high nutritional requirement of the parasite at its early developmental stages is a limiting factor for host colonisation [29].
Positive effects of the microbiota on Plasmodium
Several tolerance mechanisms avoid chronic activation of the immune system by gut microorganisms and participate in maintaining gut homeostasis, but they may also impair the mosquito immune defence against the parasite. Notably, PGRPLB is a negative regulator of the Imd pathway preventing systemic antimicrobial responses to the microbiota, which participates in a higher tolerance to Plasmodium infection [30]. Its expression is induced by the microbiota [24, 31]. Immunomodulatory Peroxidase (IMPer) and Dual oxidase (Duox) enzymes, induced by the blood meal in a microbiota-independent manner [24, 31], are involved in the formation of a dityrosine network between the gut epithelium and the peritrophic matrix [32]. This reduction in permeability protects both the microbiota and Plasmodium by preventing the activation of gut immunity. Whether the microbiota-induced peritrophic matrix [24] further reduces the diffusion of immune elicitors and is also involved in tolerance mechanisms has not yet been characterised. Along these lines, antibiotic treatment leads to an increase in the diffusion of 4kD dextran molecules from the gut to the body cavity even in IMPer-silenced mosquitoes [32].
Finally, the microbiota is also reported to contribute to the nutrition of the mosquito and Plasmodium. After feeding An. stephensi with [14C]-glycine radiolabelled Pseudomonas isolated from the Anopheles microbiota, radioactivity is detectable not only in the whole mosquito body, but also in P. berghei oocysts and in disseminated sporozoites [33]. This may reflect a positive effect of the microbiota on proliferation within the oocyst.
So far, antibiotic treatments and bacterial feeding treatment have reproducibly shown that the microbiota reduces infection by Plasmodium (Table 1), suggesting that even though some tolerance pathways and nutritional benefits exist, they only lessen the overall negative impact of the microbiota on Plasmodium.
Impact of the microbiota on mosquito fitness
A variety of factors influence parasite transmission by the mosquito. An estimation of their relative contribution to vectorial capacity is provided by the Ross-Macdonald model, which quantifies the basic reproductive number R0, i.e. the expected number of secondary infections from a single infected individual in a susceptible population (Fig. 2; [34]). It depends on the population sizes of mosquitoes and humans, the biting rate on humans, the success of parasite infection, the incubation period of the parasite in the mosquito and the mosquito lifespan.
Fig. 2.
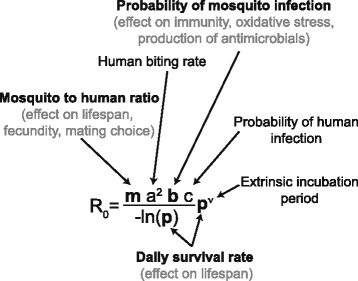
The microbiota impacts several parameters of the Ross-MacDonald model of vectorial transmission. R0 represents the basic reproductive number, the number of individuals that are expected to get infected via mosquito transmission when a single infected individual is present in a susceptible population. The variables indicated in bold are known to be microbiota-dependent. In grey, the specific effects of the microbiota on mosquito physiology and immunity are specified [13, 15–17, 20, 22, 23, 30, 35]. Potential roles of the mosquito microbiota on biting rate, anthropophily, incubation period and probability of human infection have not yet been investigated
As shown in Fig. 2, the microbiota impacts R0 via different aspects of mosquito fitness such as insect development, lifespan, fecundity and mating behaviour [10, 17, 35]. More specifically, the microbiota is fundamental for mosquito development: Ae. aegypti and An. gambiae larvae depleted of their gut microbes are unable to develop into adults [10]. A reduction in the proliferation of gut bacteria via a mild antibiotic treatment of the blood meal in An. coluzzii increases the mosquito fecundity and survival, meaning that the natural microbiota proliferation potentially decreases mosquito population size and time available for parasite transmission [17]. Some bacteria isolated from the midgut of Anopheles were shown to decrease mosquito lifespan when individually administered through sugar or blood meal while others have no effect [16, 20, 22]. Although the abundance and the effect of these single bacteria likely differ in physiological conditions, these studies suggest a composition-dependent negative impact of the gut microbiota on lifespan. A recent study reported a modification of the An. stephensi microbiota following manipulation of mosquito immunity and showed that this in turn influences reproduction behaviour, by increasing the chance of mating between mosquitoes harbouring a different microbiota [35]. All of these aspects may alter the probability of transmitting the Plasmodium parasite and thus have a strong impact on vector control.
Current challenges in the study of the mosquito microbiota
Working on mosquitoes with a representative microbiota composition
The Anopheles midgut hosts a simple, variable and dynamic microbial community whose composition mostly depends on environmental factors and individual history. More particularly, seasonality, diet, larval breeding site and blood-feeding history, but also host genetic identity, have a strong influence in shaping the midgut bacterial content [1, 3, 36–41]. The high variability of those factors in the field probably explains the high diversity of the mosquito microbiota composition between individuals, almost independently of the mosquito species [10, 36, 42–46]. Different studies suggest that the mosquito microbiota composition is not random, but whether a core bacterial community exists is still not clear. On one side, some bacterial genera are frequently found in Anopheles midguts. They are Gram-negative aerobic or facultative aerobic bacteria, mostly belonging to the families Enterobacteriaceae (Serratia, Ewingella, Enterobacter and Klebsiella), Acetobacteraceae (Acetobacter and Asaia) and Flavobacteriaceae (Elizabethkingia and Chryseobacterium) (reviewed in [47]). On the other side, the bacterial diversity is relatively low, with four operational taxonomic units (OTUs) representing more than 90% of the midgut community and a poor composition overlap between individual mosquitoes [45]. This suggests that, rather than a core microbiota, several typical microbiota compositions, or enterotypes, may be defined in Anopheles mosquitoes. Such types may derive from individual history, environment and host genetics.
Remarkably, Anopheles reared in laboratory conditions showed a reduced bacterial diversity in their midgut with respect to field-collected ones, although the majority of bacteria found in laboratory mosquitoes were present in wild mosquitoes [1, 36, 43]. Functional studies on the interactions between the microbiota and Plasmodium have mostly been conducted on mosquitoes reared under controlled laboratory conditions (Table 1). The microbiota composition of laboratory-reared mosquitoes varies between insectaries due to local variations and differences in husbandry [48]. This variability could be one of the explanations for some discrepancies of results between laboratories in vector biology. Meanwhile, microbiota studies on field-caught mosquitoes identified correlations, with Plasmodium infection for instance, but are elusive on the causes and consequences of these correlations. Notably, Straif et al. [49] found an association between Gram-positive bacteria and P. falciparum infection in field-collected An. funestus, while Boissière et al. [36] and Tchioffo et al. [3] found a positive correlation between parasite infection and Enterobacteriaceae or S. marcescens (Gram-negatives) abundance in An. gambiae after experimental infection of adults derived from field-collected larvae. These data point out the need for more appropriate microbiota models to perform functional studies on the tripartite relationship between the mosquito, the microbiota and Plasmodium. Several experimental setups may be favoured and are shown in Fig. 3 with their pros and cons. Choosing the most appropriate model to answer a scientific question, or combining the use of several models, may strengthen vector biology studies.
Fig. 3.
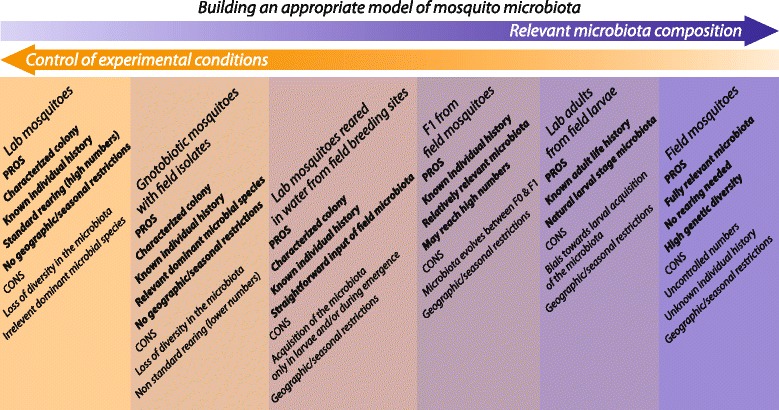

Alternative models for the study of the mosquito and its microbiota. Besides laboratory conventionally-reared mosquitoes and field-collected mosquitoes, several other microbiota set ups are or may be used to study the mosquito biology. Field bacterial isolates may be used to obtain gnotobiotic mosquitoes, i.e. mosquitoes colonised by known strains of bacteria. Laboratory larvae may be reared in water collected from natural breeding sites, or a first generation of mosquitoes (F1) may be obtained from field collected individuals. Finally, adult mosquitoes may be raised in the laboratory from field collected larvae. None of these models is perfect but each may be more appropriate depending on the aims and requirements of each study. Combining several models may also strengthen experimental work and participate in increasing the reproducibility of results between laboratories
Further identification of the role of the non-bacterial microbiota
Despite the high number of studies on the midgut microbiota, our knowledge of its non-bacterial components is still poor. Metagenomic data on total mosquito RNA detected several viruses associated with wild-caught Anopheles [50, 51]. Similar data from other mosquitoes also catalogued some viral, bacterial and eukaryotic components of the microbiota [2], but cost and computational requirements limit the number of samples analysed.
The analysis of the eukaryotic microbiota by 18S rRNA sequencing is in part complicated by the amplification of DNA from the mosquito or from the blood source [52]. Several fungi belonging to the genera Candida, Pichia and Penicillium were isolated from Anopheles larvae and adults through standard microbiological techniques [53–55]. Two functional studies reported that specific fungi isolated from mosquitoes influence Plasmodium infectiveness either negatively [56] or positively [57]. However, we still lack an exhaustive investigation of the viral and eukaryotic components of the mosquito microbiota and of their interactions with Plasmodium.
Along these lines, the parasite itself may be considered as part of the microbial community, or pathobiome [58], harboured by the mosquito. In other insect-pathogen models, it has been shown that the invading microorganism can induce significant changes in the host microbiota composition to increase its infection success. For example, when the human pathogen Anaplasma phagocytophilum infects the tick Ixodes scapularis, it induces significant variations in the composition of the host gut microbial community. This correlates with pathogen-dependent induction of the Ixodes Antifreeze Glycoprotein IAFGP, which reduces biofilm formation and impairs the integrity of the peritrophic matrix, facilitating penetration of the pathogen into the tick body cavity [59]. Since similar interactions have not been described for the Anopheles-microbiota-Plasmodium system, further studies are required to clarify the role of the parasite in influencing the microbiota and host immunity.
Widen the knowledge on the microbiota in other tissues
The microbial communities colonising the mosquito salivary glands and reproductive organs are as relevant as those in the midgut for several aspects concerning Plasmodium transmission and mosquito ecology. Microorganisms present in the salivary glands are prone to interact with the parasite when the mosquito is becoming infectious and thus they might affect transmission efficiency, while microbes colonising the Anopheles genitalia may impact reproductive success, immunity, lifespan and microbiota composition of the offspring. The endosymbiont Wolbachia is found in half of the insect species, but had not been detected in any Anopheles species until recently. When artificially introduced in an An. stephensi strain, Wolbachia persists in mosquito ovaries, is vertically transmitted and increases the resistance against P. falciparum in laboratory conditions [60]. Wolbachia has now been found associated with around 10% of Anopheles in a wild population in Burkina Faso [61]. Its load was, however, much lower than that of other insect species, notably Ae. albopictus where Wolbachia was found to account for 99% of the 16S rRNA reads in whole mosquitoes [40, 61–63]. It was proposed that some members of the ovary microbiota, notably Asaia, interfere with Wolbachia colonisation and thus reduce its possible use against malaria [64, 65].
The microbiota of the ovaries and salivary glands has been investigated in Anopheles mosquitoes in few metagenomics studies [3, 37, 63, 66]. A recent study of the microbiota composition in midguts, salivary glands and ovaries suggests that individual history rather than the tissue shapes the mosquito microbiota, but still identified some differences in taxon abundance among the three tissues [3].
Functional studies on the role of the microorganisms colonising other tissues beyond the midgut will help to clarify several aspects of Plasmodium transmission and mosquito ecology. Moreover, the isolation of bacteria able to colonise the body of the mosquito without affecting its lifespan and to be efficiently vertically transmitted will identify new paratransgenesis candidates able to spread efficiently in the mosquito population. Recently, Serratia was identified in both ovaries and salivary glands of Anopheles mosquitoes and proposed for malaria control [3, 8].
Conclusions
In the past 20 years, the pivotal role of the mosquito microbiota in shaping Plasmodium infection and transmission has gradually emerged. However, the tripartite interaction between the mosquito, its microbiota and the parasite is a complex relationship that still needs further investigation. In general, the microbiota was found to reduce Plasmodium infection and to impact several physiological aspects of the mosquito, notably affecting its lifespan. Surprisingly, these effects induced by the microbiota were consistent almost irrespective of the Anopheles and Plasmodium species, suggesting that this tripartite interaction is a stable system in which each component plays a role. Although our knowledge on the mosquito microbiota is continuously expanding, several aspects have not been completely elucidated yet and represent the current challenges of this field. In particular, the non-bacterial component of the mosquito microbiota has not been investigated as extensively as the bacterial one, although viruses and eukaryotes might be as relevant as prokaryotes in limiting Plasmodium infection. Moreover, it is not clear whether the microbiota of the reproductive track or salivary glands impacts parasite transmission or mosquito fitness. Finally, most of the functional studies conducted on the mosquito microbiota have been carried out on laboratory-reared insects, which are known to possess a different microbial community from that of field mosquitoes. Depending on the desired levels of control on experimental conditions and of relevance of microbiota composition, several experimental set ups may be used to improve the study of the mosquito microbiota.
Acknowledgements
The authors thank Faye Rodgers for proofreading of the manuscript.
Funding
This work is funded by the French Government’s Investissement d’Avenir program, Laboratoire d’Excellence “Integrative Biology of Emerging Infectious Diseases” (grant no. ANR-10-LABX-62-IBEID).
Availability of data and materials
Not applicable.
Abbreviations
- Duox
Dual oxidase
- IAFGP
Ixodes scapularis antifreeze glycoprotein
- Imd
Immune-deficiency
- IMPer
Immunomodulatory peroxidase
- OTU
Operational taxonomic unit
- PGRP
Peptidoglycan recognition protein
- ROS
Reactive oxygen species
- TEP1
Thioester-containing Protein 1
Authors’ contributions
MG conceived the paper. MG and OR revised the current literature and wrote the paper. Both authors read and approved the final manuscript.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Ottavia Romoli, Email: oromoli@pasteur-cayenne.fr.
Mathilde Gendrin, Email: mathilde.gendrin@pasteur.fr.
References
- 1.Wang Y, Gilbreath TM 3rd, Kukutla P, Yan G, Xu J. Dynamic gut microbiome across life history of the malaria mosquito Anopheles gambiae in Kenya. PLoS One. 2011;6(9):e24767. [DOI] [PMC free article] [PubMed]
- 2.Chandler JA, Liu RM, Bennett SN. RNA shotgun metagenomic sequencing of northern California (USA) mosquitoes uncovers viruses, bacteria, and fungi. Front Microbiol. 2015;6:185. doi: 10.3389/fmicb.2015.00185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Tchioffo MT, Boissiere A, Abate L, Nsango SE, Bayibeki AN, Awono-Ambene PH, et al. Dynamics of bacterial community composition in the malaria mosquito’s epithelia. Front Microbiol. 2015;6:1500. [DOI] [PMC free article] [PubMed]
- 4.Gonzalez-Ceron L, Santillan F, Rodriguez MH, Mendez D, Hernandez-Avila JE. Bacteria in midguts of field-collected Anopheles albimanus block Plasmodium vivax sporogonic development. J Med Entomol. 2003;40(3):371–374. doi: 10.1603/0022-2585-40.3.371. [DOI] [PubMed] [Google Scholar]
- 5.Dong Y, Manfredini F, Dimopoulos G. Implication of the mosquito midgut microbiota in the defense against malaria parasites. PLoS Pathog. 2009;5(5):e1000423. doi: 10.1371/journal.ppat.1000423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Yoshida S, Ioka D, Matsuoka H, Endo H, Ishii A. Bacteria expressing single-chain immunotoxin inhibit malaria parasite development in mosquitoes. Mol Biochem Parasitol. 2001;113(1):89–96. doi: 10.1016/S0166-6851(00)00387-X. [DOI] [PubMed] [Google Scholar]
- 7.Riehle MA, Moreira CK, Lampe D, Lauzon C, Jacobs-Lorena M. Using bacteria to express and display anti-Plasmodium molecules in the mosquito midgut. Int J Parasitol. 2007;37(6):595–603. doi: 10.1016/j.ijpara.2006.12.002. [DOI] [PubMed] [Google Scholar]
- 8.Wang S, Dos-Santos ALA, Huang W, Liu KC, Oshaghi MA, Wei G, et al. Driving mosquito refractoriness to Plasmodium falciparum with engineered symbiotic bacteria. Science. 2017;357(6358):1399–1402. doi: 10.1126/science.aan5478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Lindh JM, Borg-Karlson AK, Faye I. Transstadial and horizontal transfer of bacteria within a colony of Anopheles gambiae (Diptera: Culicidae) and oviposition response to bacteria-containing water. Acta Trop. 2008;107(3):242–50. [DOI] [PubMed]
- 10.Coon KL, Vogel KJ, Brown MR, Strand MR. Mosquitoes rely on their gut microbiota for development. Mol Ecol. 2014;23(11):2727–2739. doi: 10.1111/mec.12771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Favia G, Ricci I, Damiani C, Raddadi N, Crotti E, Marzorati M, et al. Bacteria of the genus Asaia stably associate with Anopheles stephensi, an Asian malarial mosquito vector. Proc Natl Acad Sci USA. 2007;104(21):9047–51. [DOI] [PMC free article] [PubMed]
- 12.Pumpuni CB, Beier MS, Nataro JP, Guers LD, Davis JR. Plasmodium falciparum: inhibition of sporogonic development in Anopheles stephensi by Gram-negative bacteria. Exp Parasitol. 1993;77(2):195–199. doi: 10.1006/expr.1993.1076. [DOI] [PubMed] [Google Scholar]
- 13.Cirimotich CM, Dong Y, Clayton AM, Sandiford SL, Souza-Neto JA, Mulenga M, et al. Natural microbe-mediated refractoriness to Plasmodium infection in Anopheles gambiae. Science. 2011;332(6031):855–858. doi: 10.1126/science.1201618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Bando H, Okado K, Guelbeogo WM, Badolo A, Aonuma H, Nelson B, et al. Intra-specific diversity of Serratia marcescens in Anopheles mosquito midgut defines Plasmodium transmission capacity. Sci Rep. 2013;3:1641. doi: 10.1038/srep01641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Meister S, Agianian B, Turlure F, Relogio A, Morlais I, Kafatos FC, et al. Anopheles gambiae PGRPLC-mediated defense against bacteria modulates infections with malaria parasites. PLoS Pathog. 2009;5(8):e1000542. doi: 10.1371/journal.ppat.1000542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Bahia AC, Dong Y, Blumberg BJ, Mlambo G, Tripathi A, BenMarzouk-Hidalgo OJ, et al. Exploring Anopheles gut bacteria for Plasmodium blocking activity. Environ Microbiol. 2014;16(9):2980–2994. doi: 10.1111/1462-2920.12381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Gendrin M, Rodgers FH, Yerbanga RS, Ouedraogo JB, Basanez MG, Cohuet A, et al. Antibiotics in ingested human blood affect the mosquito microbiota and capacity to transmit malaria. Nat Commun. 2015;6:5921. doi: 10.1038/ncomms6921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Dong Y, Das S, Cirimotich C, Souza-Neto JA, McLean KJ, Dimopoulos G. Engineered Anopheles immunity to Plasmodium infection. PLoS Pathog. 2011;7(12):e1002458. doi: 10.1371/journal.ppat.1002458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Wang Y, Wang Y, Zhang J, Xu W, Zhang J, Huang FS. Ability of TEP1 in intestinal flora to modulate natural resistance of Anopheles dirus. Exp Parasitol. 2013;134(4):460–5. [DOI] [PubMed]
- 20.Dennison NJ, Saraiva RG, Cirimotich CM, Mlambo G, Mongodin EF, Dimopoulos G. Functional genomic analyses of Enterobacter, Anopheles and Plasmodium reciprocal interactions that impact vector competence. Malar J. 2016;15(1):425. doi: 10.1186/s12936-016-1468-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Bahia AC, Oliveira JH, Kubota MS, Araujo HR, Lima JB, Rios-Velasquez CM, et al. The role of reactive oxygen species in Anopheles aquasalis response to Plasmodium vivax infection. PLoS One. 2013;8(2):e57014. doi: 10.1371/journal.pone.0057014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ramirez JL, Short SM, Bahia AC, Saraiva RG, Dong Y, Kang S, et al. Chromobacterium Csp_P reduces malaria and dengue infection in vector mosquitoes and has entomopathogenic and in vitro anti-pathogen activities. PLoS Pathog. 2014;10(10):e1004398. doi: 10.1371/journal.ppat.1004398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Valzano M, Cecarini V, Cappelli A, Capone A, Bozic J, Cuccioloni M, et al. A yeast strain associated to Anopheles mosquitoes produces a toxin able to kill malaria parasites. Malar J. 2016;15:21. doi: 10.1186/s12936-015-1059-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Rodgers FH, Gendrin M, Wyer CAS, Christophides GK. Microbiota-induced peritrophic matrix regulates midgut homeostasis and prevents systemic infection of malaria vector mosquitoes. PLoS Pathog. 2017;13(5):e1006391. doi: 10.1371/journal.ppat.1006391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Huber M, Cabib E, Miller LH. Malaria parasite chitinase and penetration of the mosquito peritrophic membrane. Proc Natl Acad Sci USA. 1991;88(7):2807–10. [DOI] [PMC free article] [PubMed]
- 26.Tsai YL, Hayward RE, Langer RC, Fidock DA, Vinetz JM. Disruption of Plasmodium falciparum chitinase markedly impairs parasite invasion of mosquito midgut. Infect Immun. 2001;69(6):4048–4054. doi: 10.1128/IAI.69.6.4048-4054.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Kukutla P, Lindberg BG, Pei D, Rayl M, Yu W, Steritz M, et al. Insights from the genome annotation of Elizabethkingia anophelis from the malaria vector Anopheles gambiae. PLoS One. 2014;9(5):e97715. [DOI] [PMC free article] [PubMed]
- 28.Caragata EP, Rances E, Hedges LM, Gofton AW, Johnson KN, O'Neill SL, et al. Dietary cholesterol modulates pathogen blocking by Wolbachia. PLoS Pathog. 2013;9(6):e1003459. doi: 10.1371/journal.ppat.1003459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Lavazec C, Boudin C, Lacroix R, Bonnet S, Diop A, Thiberge S, et al. Carboxypeptidases B of Anopheles gambiae as targets for a Plasmodium falciparum transmission-blocking vaccine. Infect Immun. 2007;75(4):1635–1642. doi: 10.1128/IAI.00864-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Gendrin M, Turlure F, Rodgers FH, Cohuet A, Morlais I, Christophides GK. The peptidoglycan recognition proteins PGRPLA and PGRPLB regulate Anopheles immunity to bacteria and affect infection by Plasmodium. J Innate Immun. 2017;9:333–342. doi: 10.1159/000452797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Vectorbase. RNA-Seq Experiment - Antibiotic treatment of midgut microbiota during blood feeding (Rodgers et al. 2017); 2017. https://www.vectorbase.org/expression-browser/experiment/SRP106793. Accessed 16 Nov 2017.
- 32.Kumar S, Molina-Cruz A, Gupta L, Rodrigues J, Barillas-Mury C. A peroxidase/dual oxidase system modulates midgut epithelial immunity in Anopheles gambiae. Science. 2010;327(5973):1644–8. [DOI] [PMC free article] [PubMed]
- 33.Jadin J. Role of bacteria in the digestive tube of insects, vectors of plasmodidae and trypanosomidae. Ann Soc Belges Med Trop Parasitol Mycol. 1967;47(4):331–42. (In French). [PubMed]
- 34.Smith DL, Battle KE, Hay SI, Barker CM, Scott TW, FE MK. Ross, Macdonald, and a theory for the dynamics and control of mosquito-transmitted pathogens. PLoS Pathog. 2012;8(4):e1002588. doi: 10.1371/journal.ppat.1002588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Pike A, Dong Y, Dizaji NB, Gacita A, Mongodin EF, Dimopoulos G. Changes in the microbiota cause genetically modified Anopheles to spread in a population. Science. 2017;357(6358):1396–1399. doi: 10.1126/science.aak9691. [DOI] [PubMed] [Google Scholar]
- 36.Boissiere A, Tchioffo MT, Bachar D, Abate L, Marie A, Nsango SE, et al. Midgut microbiota of the malaria mosquito vector Anopheles gambiae and interactions with Plasmodium falciparum infection. PLoS Pathog. 2012;8(5):e1002742. doi: 10.1371/journal.ppat.1002742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Gimonneau G, Tchioffo MT, Abate L, Boissiere A, Awono-Ambene PH, Nsango SE, et al. Composition of Anopheles coluzzii and Anopheles gambiae microbiota from larval to adult stages. Infect Genet Evol. 2014;28:715–724. doi: 10.1016/j.meegid.2014.09.029. [DOI] [PubMed] [Google Scholar]
- 38.Stathopoulos S, Neafsey DE, Lawniczak MK, Muskavitch MA, Christophides GK. Genetic dissection of Anopheles gambiae gut epithelial responses to Serratia marcescens. PLoS Pathog. 2014;10(3):e1003897. doi: 10.1371/journal.ppat.1003897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Akorli J, Gendrin M, Pels NA, Yeboah-Manu D, Christophides GK, Wilson MD. Seasonality and locality affect the diversity of Anopheles gambiae and Anopheles coluzzii midgut microbiota from Ghana. PLoS One. 2016;11(6):e0157529. [DOI] [PMC free article] [PubMed]
- 40.Buck M, Nilsson LK, Brunius C, Dabire RK, Hopkins R, Terenius O. Bacterial associations reveal spatial population dynamics in Anopheles gambiae mosquitoes. Sci Rep. 2016;6:22806. doi: 10.1038/srep22806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Novakova E, Woodhams DC, Rodriguez-Ruano SM, Brucker RM, Leff JW, Maharaj A, et al. Mosquito microbiome dynamics, a background for prevalence and seasonality of West Nile Virus. Front Microbiol. 2017;8:526. doi: 10.3389/fmicb.2017.00526. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Terenius O, de Oliveira CD, Pinheiro WD, Tadei WP, James AA, Marinotti O. 16S rRNA gene sequences from bacteria associated with adult Anopheles darlingi (Diptera: Culicidae) mosquitoes. J Med Entomol. 2008;45(1):172–175. doi: 10.1093/jmedent/45.1.172. [DOI] [PubMed] [Google Scholar]
- 43.Rani A, Sharma A, Rajagopal R, Adak T, Bhatnagar RK. Bacterial diversity analysis of larvae and adult midgut microflora using culture-dependent and culture-independent methods in lab-reared and field-collected Anopheles stephensi - an Asian malarial vector. BMC Microbiol. 2009;9:96. doi: 10.1186/1471-2180-9-96. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Dinparast Djadid N, Jazayeri H, Raz A, Favia G, Ricci I, Zakeri S. Identification of the midgut microbiota of An. stephensi and An. maculipennis for their application as a paratransgenic tool against malaria. PLoS One. 2011;6(12):e28484. doi: 10.1371/journal.pone.0028484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Osei-Poku J, Mbogo CM, Palmer WJ, Jiggins FM. Deep sequencing reveals extensive variation in the gut microbiota of wild mosquitoes from Kenya. Mol Ecol. 2012;21(20):5138–5150. doi: 10.1111/j.1365-294X.2012.05759.x. [DOI] [PubMed] [Google Scholar]
- 46.Manguin S, Ngo CT, Tainchum KT, Juntarajumnong W, Chareonviriyaphap T, Michon AL, et al. Chapter 18. Bacterial biodiversity in midguts of Anopheles mosquitoes, malaria vectors in southeast Asia. In: Manguin S, et al., editors. Anopheles mosquitoes - New insights into malaria vectors. InTech: Rijeka; 2013. [Google Scholar]
- 47.Gendrin M, Christophides GK, Manguin S. Anopheles mosquitoes - New insights into malaria vectors. InTech: Rijeka; 2013. Chapter 17. The Anopheles mosquito microbiota and their impact on pathogen transmission. [Google Scholar]
- 48.Linenberg I, Christophides GK, Gendrin M. Larval diet affects mosquito development and permissiveness to Plasmodium infection. Sci Rep. 2016;6:38230. doi: 10.1038/srep38230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Straif SC, Mbogo CN, Toure AM, Walker ED, Kaufman M, Toure YT, et al. Midgut bacteria in Anopheles gambiae and An. funestus (Diptera: Culicidae) from Kenya and Mali. J Med Entomol. 1998;35(3):222–226. doi: 10.1093/jmedent/35.3.222. [DOI] [PubMed] [Google Scholar]
- 50.Zhuang L, Zhang Z, An X, Fan H, Ma M, Anderson BD, et al. An efficient strategy of screening for pathogens in wild-caught ticks and mosquitoes by reusing small RNA deep sequencing data. PLoS One. 2014;9(3):e90831. doi: 10.1371/journal.pone.0090831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Fauver JR, Grubaugh ND, Krajacich BJ, Weger-Lucarelli J, Lakin SM, Fakoli LS, 3rd, et al. West African Anopheles gambiae mosquitoes harbor a taxonomically diverse virome including new insect-specific flaviviruses, mononegaviruses, and totiviruses. Virology. 2016;498:288–299. doi: 10.1016/j.virol.2016.07.031. [DOI] [PubMed] [Google Scholar]
- 52.Belda E, Coulibaly B, Fofana A, Beavogui AH, Traore SF, Gohl DM, et al. Preferential suppression of Anopheles gambiae host sequences allows detection of the mosquito eukaryotic microbiome. Sci Rep. 2017;7(1):3241. doi: 10.1038/s41598-017-03487-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.da S Pereira E, de M Sarquis MI, Ferreira-Keppler RL, Hamada N, Alencar YB. Filamentous fungi associated with mosquito larvae (Diptera: Culicidae) in municipalities of the Brazilian Amazon. Neotrop Entomol. 2009;38(3):352–359. doi: 10.1590/S1519-566X2009000300009. [DOI] [PubMed] [Google Scholar]
- 54.Ricci I, Damiani C, Scuppa P, Mosca M, Crotti E, Rossi P, et al. The yeast Wickerhamomyces anomalus (Pichia anomala) inhabits the midgut and reproductive system of the Asian malaria vector Anopheles stephensi. Environ Microbiol. 2011;13(4):911–921. doi: 10.1111/j.1462-2920.2010.02395.x. [DOI] [PubMed] [Google Scholar]
- 55.Bozic J, Capone A, Pediconi D, Mensah P, Cappelli A, Valzano M, et al. Mosquitoes can harbour yeasts of clinical significance and contribute to their environmental dissemination. Environ Microbiol Rep. 2017;9(5):642–648. doi: 10.1111/1758-2229.12569. [DOI] [PubMed] [Google Scholar]
- 56.Bargielowski I, Koella JC. A possible mechanism for the suppression of Plasmodium berghei development in the mosquito Anopheles gambiae by the microsporidian Vavraia culicis. PLoS One. 2009;4(3):e4676. [DOI] [PMC free article] [PubMed]
- 57.Anglero-Rodriguez YI, Blumberg BJ, Dong Y, Sandiford SL, Pike A, Clayton AM, et al. A natural Anopheles-associated Penicillium chrysogenum enhances mosquito susceptibility to Plasmodium infection. Sci Rep. 2016;6:34084. doi: 10.1038/srep34084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Vayssier-Taussat M, Albina E, Citti C, Cosson JF, Jacques MA, Lebrun MH, et al. Shifting the paradigm from pathogens to pathobiome: new concepts in the light of meta-omics. Front Cell Infect Microbiol. 2014;4:29. doi: 10.3389/fcimb.2014.00029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Abraham NM, Liu L, Jutras BL, Yadav AK, Narasimhan S, Gopalakrishnan V, et al. Pathogen-mediated manipulation of arthropod microbiota to promote infection. Proc Natl Acad Sci USA. 2017;114(5):E781–90. [DOI] [PMC free article] [PubMed]
- 60.Bian G, Joshi D, Dong Y, Lu P, Zhou G, Pan X, et al. Wolbachia invades Anopheles stephensi populations and induces refractoriness to Plasmodium infection. Science. 2013;340(6133):748–751. doi: 10.1126/science.1236192. [DOI] [PubMed] [Google Scholar]
- 61.Baldini F, Segata N, Pompon J, Marcenac P, Shaw WR, Dabire RK, et al. Evidence of natural Wolbachia infections in field populations of Anopheles gambiae. Nat Commun. 2014;5:3985. doi: 10.1038/ncomms4985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Minard G, Tran FH, Dubost A, Tran-Van V, Mavingui P, Moro CV. Pyrosequencing 16S rRNA genes of bacteria associated with wild tiger mosquito Aedes albopictus: a pilot study. Front Cell Infect Microbiol. 2014;4:59. doi: 10.3389/fcimb.2014.00059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Segata N, Baldini F, Pompon J, Garrett WS, Truong DT, Dabire RK, et al. The reproductive tracts of two malaria vectors are populated by a core microbiome and by gender- and swarm-enriched microbial biomarkers. Sci Rep. 2016;6:24207. doi: 10.1038/srep24207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Hughes GL, Dodson BL, Johnson RM, Murdock CC, Tsujimoto H, Suzuki Y, et al. Native microbiome impedes vertical transmission of Wolbachia in Anopheles mosquitoes. Proc Natl Acad Sci USA. 2014;111(34):12498–503. [DOI] [PMC free article] [PubMed]
- 65.Rossi P, Ricci I, Cappelli A, Damiani C, Ulissi U, Mancini MV, et al. Mutual exclusion of Asaia and Wolbachia in the reproductive organs of mosquito vectors. Parasit Vectors. 2015;8:278. doi: 10.1186/s13071-015-0888-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Sharma P, Sharma S, Maurya RK, Das De T, Thomas T, Lata S, et al. Salivary glands harbor more diverse microbial communities than gut in Anopheles culicifacies. Parasit Vectors. 2014;7:235. doi: 10.1186/1756-3305-7-235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Pumpuni CB, Demaio J, Kent M, Davis JR, Beier JC. Bacterial population dynamics in three anopheline species: the impact on Plasmodium sporogonic development. Am J Trop Med Hyg. 1996;54(2):214–218. doi: 10.4269/ajtmh.1996.54.214. [DOI] [PubMed] [Google Scholar]
- 68.Noden BH, Vaughan JA, Pumpuni CB, Beier JC. Mosquito ingestion of antibodies against mosquito midgut microbiota improves conversion of ookinetes to oocysts for Plasmodium falciparum, but not P. yoelii. Parasitol Int. 2011;60(4):440–446. doi: 10.1016/j.parint.2011.07.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Tchioffo MT, Boissiere A, Churcher TS, Abate L, Gimonneau G, Nsango SE, et al. Modulation of malaria infection in Anopheles gambiae mosquitoes exposed to natural midgut bacteria. PLoS One. 2013;8(12):e81663. doi: 10.1371/journal.pone.0081663. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Not applicable.


