Abstract
Glutathione S-transferase Pi (GSTP) is a thiolase that catalyzes the addition of glutathione (GSH) to receptive cysteines in target proteins, producing an S-glutathionylated residue. Accordingly, previous studies have reported that S-glutathionylation is constitutively decreased in cells from mice lacking GSTP (Gstp1/p2−/−). Here, we found that bone marrow–derived dendritic cells (BMDDCs) from Gstp1/p2−/− mice have proliferation rates that are greater than those in their WT counterparts (Gstp1/p2+/+). Moreover, Gstp1/p2−/− BMDDCs had increased reactive oxygen species (ROS) levels and decreased GSH:glutathione disulfide (GSSG) ratios. Estrogen receptor α (ERα) is linked to myeloproliferation and differentiation, and we observed that its steady-state levels are elevated in Gstp1/p2−/− BMDDCs, indicating a link between GSTP and ERα activities. BMDDCs differentiated by granulocyte–macrophage colony-stimulating factor had elevated ERα levels, which were more pronounced in Gstp1/p2−/− than WT mice. When stimulated with lipopolysaccharide for maturation, Gstp1/p2−/− BMDDCs exhibited augmented endocytosis, maturation rate, cytokine secretion, and T-cell activation; heightened glucose uptake and glycolysis; increased Akt signaling (in the mTOR pathway); and decreased AMPK-mediated phosphorylation of proteins. Of note, GSTP formed a complex with ERα, stimulating ERα S-glutathionylation at cysteines 221, 245, 417, and 447; altering ERα's binding affinity for estradiol; and reducing overall binding potential (receptor density and affinity) 3-fold. Moreover, in Gstp1/p2−/− BMDDCs, ERα S-glutathionylation was constitutively decreased. Taken together, these findings suggest that GSTP-mediated S-glutathionylation of ERα controls BMDDC differentiation and affects metabolic function in dendritic cells.
Keywords: dendritic cell, energy metabolism, estrogen, estrogen receptor, redox signaling, S-glutathionylation, glutathione, glutathione S-transferase, myeloproliferation
Introduction
Dendritic cells (DCs)4 are antigen-presenting cells that bridge innate and adaptive immunity and can be activated by a variety of antigenic threats. Agents such as GM-CSF can facilitate enhanced differentiation of bone marrow derived–dendritic cells (BMDDCs) from hematopoietic stem cells (HSCs), and newly developed BMDDCs are often critical in mounting an effective lymphocyte-mediated immune response during infection (1, 2). The majority of mature DCs have a short half-life in peripheral blood circulation (3), and partly as a consequence regulation of BMDDC developmental pathways is a critical determinant of the total numbers of mature DCs.
There is significant documentation that drugs that influence redox conditions, and particularly thiol levels, influence bone marrow function, T-cell activation, and generally adaptive immune responses (4, 5). However, there has been no consensus as to how regulating thiol metabolism causes such adaptations. Within the marrow compartment, there exist gradient niches of calcium, oxygen, and glutathione (GSH) (6–9), each of which can influence the localization of progenitor cells and their pathways to maturity/release into the peripheral circulation. A number of redox-active drugs have been used clinically to treat various human myeloproliferative pathologies, including N-acetylcysteine, amifostine, and NOV-002 (10–12), each of which influences thiol homeostasis in the bone marrow compartment (8). Glutathione S-transferase Pi (GSTP) is a thiolase that can catalyze the addition of GSH to receptive cysteines in target proteins, producing an S-glutathionylated residue (13). Telintra® is a peptidomimetic inhibitor of GSTP (14) and has proven clinical activity in patients with myelodysplastic syndrome (15, 16). Conversely, mice lacking glutathione S-transferase Pi (Gstp1/p2−/−) have a more highly proliferative bone marrow, resulting in higher levels of circulating blood cells of all lineages (14, 17). These findings collectively suggest that GSTP-controlled protein S-glutathionylation regulates bone marrow progenitor cell function.
Hematopoietic progenitor cells also express estrogen receptor α (ERα), and ERα signaling is known to influence both development and maturation of BMDDCs and the general sustainability of an immune response (18). ERα broadly impacts glucose metabolism and metabolic gene expression patterns that stimulate resting T cells toward rapid proliferation and differentiation into mature T-effector populations (19). Although a number of studies suggest that downstream transcriptional events triggered by higher levels of ERα and estrogens contribute to enhanced proliferation and functions of BMDDCs, it remains unclear how redox conditions and hormone-based pathways might bisect.
The goal of the present study was to elucidate whether GSTP regulates ERα and whether this in turn regulates proliferation, differentiation, and effector function of BMDDCs. We demonstrate that GSTP and ERα interact and that GSTP induces S-glutathionylation of critical cysteines within ERα. S-Glutathionylation of ERα decreases affinity of ERα for estradiol. We furthermore demonstrate that the absence of GSTP enhanced energy metabolism (glucose use and transition to glycolysis) and proliferative and differentiation rates and increased DC effector function in association with decreased ERα-SSG.
Results
A principle rationale for the initiation of the present studies was that analysis of an estrogen receptor signaling primer array comparing wildtype and knockout BMDDCs (Table 1 and Fig. S1) quantitatively demonstrated the existence of a nexus between GSTP and ERα: the most affected change caused by ablation of GSTP was increased expression of ERα, and this was accompanied by a decrease in expression of ERβ. A number of transcription and growth factors and accessory proteins were also increased (Table 1), including tyrosine kinase receptors of the ERB family and PELP1, a coactivator of ERα. Such data encouraged us to interrogate those factors that provided a link between GSTP-mediated redox regulation and estrogen receptor function, particularly as existing evidence already implicates each in bone marrow proliferation.
Table 1.
Relative expression of genes for estrogen receptor signaling pathway in BMDDCs (Gstp1p2−/−versus Gstp1p2+/+)
mRNAs from WT or Gstp1/p2−/− BMDDCs were extracted and quantified using the estrogen receptor signaling PCR array. Quantitative analyses of each gene were performed using Bio-Rad PrimePCR Analysis software (v.1.0.030.1023). MMTV, murine mammary tumor virus.
| Gene | Symbol | Expression |
|---|---|---|
| Estrogen receptor 2 (β) | Esr2 | 0.135 |
| Prostaglandin-endoperoxide synthase 2 | Ptgs2 | 0.144 |
| Trefoil factor 1 | Tff1 | 0.165 |
| WNT1-inducible signaling pathway protein 2 | Wisp2 | 0.188 |
| Adenosine A1 receptor | Adora1 | 0.254 |
| Insulin-like growth factor–binding protein 4 | Igfbp4 | 0.447 |
| Latent transforming growth factor β–binding protein 1 | Ltbp1 | 0.466 |
| Caveolin 1, caveolae protein | Cav1 | 0.497 |
| Heat shock protein 90, α (cytosolic), class A member 1 | Hsp90aa1 | 0.499 |
| Complement component 3 | C3 | 0.517 |
| Glucuronidase, β | Gusb | 0.539 |
| Nuclear receptor subfamily 0, group B, member 2 | Nr0b2 | 0.543 |
| Creatine kinase, brain | Ckb | 0.549 |
| Early growth response 3 | Egr3 | 0.587 |
| Nuclear receptor subfamily 5, group A, member 2 | Nr5a2 | 0.613 |
| Insulin receptor substrate 1 | Irs1 | 0.646 |
| Jun-B oncogene | Junb | 0.674 |
| Glucose-6-phosphate dehydrogenase X-linked | G6pdx | 0.680 |
| Thrombospondin 1 | Thbs1 | 0.682 |
| FBJ osteosarcoma oncogene | Fos | 0.696 |
| Lipoprotein lipase | Lpl | 0.706 |
| Cbp/p300–interacting transactivator | Cited2 | 0.725 |
| Breast cancer antiestrogen resistance 1 | Bcar1 | 0.730 |
| Insulin-like growth factor 1 | Igf1 | 0.761 |
| Heat shock protein 90, α (cytosolic), class B member 1 | Hsp90ab1 | 0.763 |
| Cathepsin D | Ctsd | 0.782 |
| v-maf musculoaponeurotic fibrosarcoma oncogene family | Maff | 0.795 |
| PrimePCR RNA Quality Assay | RQ1 | 0.805 |
| Secreted phosphoprotein 1 | Spp1 | 0.816 |
| Cytochrome P450, family 1, subfamily a, polypeptide 1 | Cyp1a1 | 0.816 |
| β2-Microglobulin | B2m | 0.816 |
| Connective tissue growth factor | Ctgf | 0.825 |
| Follistatin | Fst | 0.825 |
| L1 cell adhesion molecule | L1cam | 0.828 |
| Kallikrein B, plasma 1 | Klkb1 | 0.854 |
| Actin, β | Actb | 0.866 |
| BCL2-like 1 | Bcl2l1 | 0.878 |
| Vitamin D receptor | Vdr | 0.883 |
| Cyclin D1 | Ccnd1 | 0.884 |
| Lectin, galactose-binding, soluble 1 | Lgals1 | 0.905 |
| Breast cancer 1 | Brca1 | 0.922 |
| S100 calcium-binding protein A6 (calcyclin) | S100a6 | 0.922 |
| Forkhead box A1 | Foxa1 | 0.940 |
| Myelocytomatosis oncogene | Myc | 0.971 |
| Estrogen receptor–binding fragment-associated gene 9 | Ebag9 | 0.975 |
| Nuclear receptor corepressor 2 | Ncor2 | 1.009 |
| Nuclear receptor coactivator 3 | Ncoa3 | 1.021 |
| Vascular endothelial growth factor A | Vegfa | 1.031 |
| Aryl-hydrocarbon receptor | Ahr | 1.068 |
| Metastasis-associated 1 | Mta1 | 1.173 |
| X-box–binding protein 1 | Xbp1 | 1.177 |
| Suppressor of cytokine signaling 3 | Socs3 | 1.190 |
| Scaffold attachment factor B | Safb | 1.203 |
| Bone morphogenetic protein 4 | Bmp4 | 1.205 |
| PDZ domain–containing 1 | Pdzk1 | 1.214 |
| Chemokine (CC motif) ligand 12 | Ccl12 | 1.227 |
| Patched homolog 1 | Ptch1 | 1.227 |
| Wingless-related MMTV integration site 5A | Wnt5a | 1.266 |
| Bone morphogenetic protein 7 | Bmp7 | 1.291 |
| PrimePCR Reverse Transcription Control Assay | RT | 1.335 |
| Wingless-related MMTV integration site 4 | Wnt4 | 1.335 |
| Transforming growth factor, β3 | Tgfb3 | 1.345 |
| v-erb-b2 erythroblastic leukemia viral oncogene homolog 2 | Erbb2 | 1.372 |
| Nuclear receptor coactivator 1 | Ncoa1 | 1.391 |
| Nuclear receptor subfamily 3, group C, member 1 | Nr3c1 | 1.402 |
| Proline-, glutamic acid-, and leucine-rich protein 1 | Pelp1 | 1.403 |
| Nuclear receptor–interacting protein 1 | Nrip1 | 1.446 |
| Nuclear receptor coactivator 2 | Ncoa2 | 1.449 |
| Neuropilin 1 | Nrp1 | 1.460 |
| Hypoxanthine-guanine phosphoribosyltransferase | Hprt | 1.465 |
| Mediator complex subunit 1 | Med1 | 1.591 |
| Transforming growth factor, α | Tgfa | 1.591 |
| v-ral simian leukemia viral oncogene homolog A | Rala | 1.615 |
| PrimePCR Positive Control Assay | PCR | 1.622 |
| Ngfi-A–binding protein 2 | Nab2 | 1.628 |
| A kinase (PRKA) anchor protein 1 | Akap1 | 1.690 |
| Progesterone receptor | Pgr | 1.696 |
| Amyloid β (A4) precursor protein–binding, family B1 | Apbb1 | 1.812 |
| Insulin-like growth factor–binding protein 5 | Igfbp5 | 1.955 |
| Prohibitin 2 | Phb2 | 2.082 |
| Matrix metallopeptidase 9 | Mmp9 | 2.168 |
| PrimePCR RNA Quality Assay | RQ2 | 2.170 |
| Brain-derived neurotrophic factor | Bdnf | 2.255 |
| Nuclear receptor subfamily 2, group F, member 6 | Nr2f6 | 2.297 |
| Ephrin A5 | Efna5 | 2.598 |
| Nephroblastoma overexpressed gene | Nov | 2.717 |
| Retinoic acid receptor, α | Rara | 3.243 |
| Snail homolog 1 (Drosophila) | Snai1 | 4.669 |
| Estrogen receptor 1 (α) | Esr1 | 5.416 |
| v-erb-b2 erythroblastic leukemia viral oncogene homolog 3 | Erbb3 | 12.099 |
Gstp1/p2-null mice have higher levels of oxidative stress and ERα during BMDDC differentiation
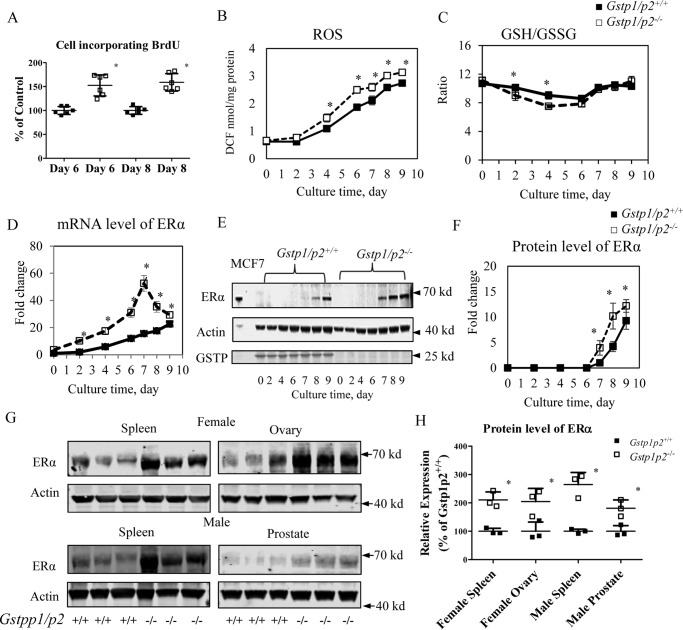
Bone marrow (BM) cells from GSTP-null mice proliferate faster (shorter doubling times) and produce more circulating blood cells of all lineages than the WT (17). Fig. 1A shows that enhanced proliferation rates in Gstp1/p2−/− cells were maintained when BMDDCs were differentiated from BM precursors in the presence of GM-CSF with incorporation of BrdU used as a marker. Differentiation of DCs is essentially complete at day 6 but can continue until day 9. In our present study, we focused on immature bone marrow–derived dendritic cells (imDCs), therefore we selected cells from days 6 and 8, each representing typical growth of imDCs. During the differentiation of BMDDCs from precursor cells, GM-CSF treatment caused a time-dependent increase in ROS in both GSTP WT and KO cells, and the change was more pronounced in cells from Gstp1/p2−/− mice (Fig. 1B). There was also a shift in the GSH/GSSG ratios toward a more oxidized state (Fig. 1C). We consistently observed that ROS levels were higher in the GSTP KO BMDDCs. It seems that GSH/GSSG ratios drop at the early days (mainly during HSC to DC differentiation from days 0 to 6) and then increase and stabilize during DC cell proliferation (days 6–9). However, the ROS levels consistently increased during the entire period of GM-CSF–induced differentiation and proliferation. In general, the data suggested that GM-CSF induced oxidative stress, causing a disruption in redox balance that was more pronounced in GSTP-deficient cells. In concordance with the comparative array data (Table 1), GM-CSF–induced differentiation of BMDDCs from BM precursors caused a gradual increase in expression of ERα, both at the transcript (Fig. 1D) and protein levels (Fig. 1, E and F). The up-regulation of mRNA is much greater in GSTP KO mice, and the highest levels occur on day 7. However, there is a lag of protein expression versus gene up-regulation, most pronounced after day 7. At early time points, low ERα expression and limited detection capacities of the ERα antibodies meant that we could not detect a band. ERα levels were higher in the Gstp1/p2−/− cells and increased at a faster rate in the time after GM-CSF stimulation. Gstp1/p2−/− cells had detectable protein levels from day 7 onward, whereas Gstp1/p2+/+ cells had detectable protein levels only from day 8 (Fig. 1, E and F). This seemed to be a characteristic of the Gstp1/p2−/− mice because higher resting-state levels of ERα were found in spleen, ovary, and prostate (Fig. 1, G and H).
Figure 1.
ERα levels and oxidative stress are higher in Gstp1/p2−/− cells during GM-CSF–driven BMDDC differentiation and proliferation. BM cells were prepared from femora and tibiae from Gstp1/p2−/− or WT mice and cultured in the presence of GM-CSF as described under “Experimental procedures.” Non-adherent cells were collected at days 0, 2, 4, 6, 7, 8, and 9, respectively. A, Gstp1/p2−/− BMDDCs differentiated with GM-CSF had higher proliferation rates. On days 4 and 6, BM cells were seeded at 0.2 × 106 cells/ml in DC medium and cultured for an extra 2 days. Cell proliferation was then assessed by BrdU incorporation. B, cellular ROS levels were measured using a proprietary quenched fluorogenic probe, DCFH-DiOxyQ. C, intracellular GSH levels were measured by thiol fluorescent probe IV after protein precipitation. Intracellular GSSG levels were determined based on the reduction of GSSG in the presence of glutathione reductase and NADPH. Data are presented as GSH/GSSG ratio. D, ERα mRNA levels were determined by quantitative RT-PCR. E and F, ERα protein levels were evaluated by immunoblotting and quantified with Image Studio 4.0 software. The breast cancer cell line MCF7 was used as a positive control for ERα. G and H, Gstp1/p2−/− mice have higher resting tissue levels of ERα. Tissue samples (spleen, ovary, or prostate) were freshly prepared from Gstp1/p2−/− or WT mice. ERα protein levels were evaluated by immunoblotting and quantified with Image Studio 4.0 software. Actin served as a loading control. Data shown are means ± S.D. from three animals. Error bars represent S.D. *, significant differences between Gstp1/p2−/− and WT, p < 0.05. DCF, dichlorofluorescein.
Gstp1/p2 depletion in BMDDCs results in a proinflammatory phenotype
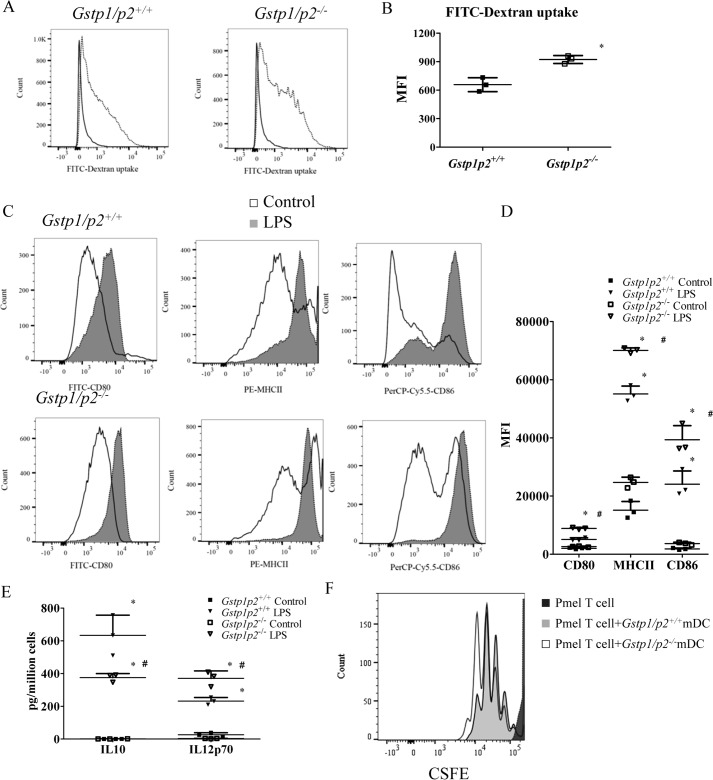
ERα has been suggested previously to play a role in regulating DC endocytosis, maturation, and cytokine secretion and priming T-cell proliferation (18, 19). We performed a series of experiments to measure comparative differences in BMDDC functions between WT and Gstp1/p2−/− mice. The data indicated that higher levels of DC functions occurred in the absence of GSTP expression. As a function specific to imDCs, we measured the capacity to endocytose FITC-dextran via the mannose receptor. Fig. 2, A and B, show that endocytotic functions were higher in Gstp1/p2−/− BMDDCs compared with WT. In response to an agonist of toll-like receptor 4, Gstp1/p2−/− BMDDCs matured normally following LPS treatment as demonstrated by their expression of CD80, MHCII, and CD86 (Fig. 2C). However, expression of these maturation markers was enhanced in LPS-treated Gstp1/p2−/− compared with WT BMDDCs (quantification (arbitrary units) of the MFI values for Gstp1/p2−/− versus WT: CD80, 8884 ± 308 versus 5068 ± 579; MHCII, 70,097 ± 891 versus 55,127 ± 2703; CD86, 39,359 ± 4872 versus 24,085 ± 4552; Fig. 2D). Even in the absence of LPS, Gstp1/p2−/− BMDDCs had higher levels of MHCII (Fig. 2D). Furthermore, following LPS activation, Gstp1/p2−/− BMDDCs produced significantly higher levels of the proinflammatory cytokine IL-12p70 and less of the anti-inflammatory cytokine IL-10 compared with their WT counterparts (Gstp1/p2−/− versus WT: IL-12p70, 371 ± 26 versus 232 ± 13; IL-10, 369 ± 22 versus 577 ± 53). Without LPS activation, IL-12p70 secretion was very low, and IL-10 secretion was undetectable in both WT and Gstp1/p2−/− BMDDCs (Fig. 2E). Finally, we investigated the efficiency with which BMDDCs primed naïve T cells by stimulating carboxyfluorescein diacetate succinimidyl ester (CFSE)–labeled naïve T cells (from Pmel-1 T-cell receptor (TCR) transgenic mice) with hgp100(25–33)-pulsed BMDDCs activated with LPS (mature bone marrow–derived dendritic cells (mDCs)). Fig. 2F shows that naïve CD8+ T cells undergo antigen-specific proliferation when cocultured with either WT or Gstp1/p2−/− mDCs, and higher levels of T-cell expansion were observed when cocultured with Gstp1/p2−/− mDCs. Together, these data show that Gstp1/p2 ablation resulted in a phenotype of enhanced DC endocytosis, maturation, proinflammatory cytokine secretion, and capacity to prime T-cell proliferation.
Figure 2.
BMDDCs from Gstp1/p2−/− mice have enhanced antigen presentation functions. A, endocytosis was evaluated by FITC-dextran uptake. BMDDCs from Gstp1/p2−/− or WT mice were pulsed with 1 mg/ml FITC-dextran for 4 h at either 37 or 4 °C and counterstained for markers for DCs (CD11c), T cells (CD3e), B cells (CD45R/B220), monocytes/macrophages (CD14), and NK cells (NK1.1 molecules) before analysis by flow cytometry. Plots are gated on CD11c+, CD3e−, CD45R/B220−, CD14−, and NK1.1− cells. Representative histograms of three experiments are shown. B, FITC-dextran uptake at 37 °C compared with 4 °C (control) shows the mean MFI ± S.D. from three experiments. Error bars represent S.D. *, significant differences between Gstp1/p2−/− and WT imDCs, p < 0.05. C, BMDDCs were cultured in DC medium with or without 200 ng/ml LPS overnight and then analyzed for the expression of maturation markers by flow cytometry. Plots are gated on CD11c+ cells. Data are representative of three experiments. D, quantitative graphical representation of C. Data shown are mean MFI ± S.D. from three experiments. Error bars represent S.D. *, significant differences between control and LPS treatment, p < 0.05; #, significant differences between Gstp1/p2−/− and WT DCs after LPS treatment, p < 0.05. E, BMDDCs from Gstp1/p2−/− or WT mice were stimulated with 200 ng/ml LPS overnight, and medium was removed and examined by ELISA for the proinflammatory cytokine IL-12p70 and the anti-inflammatory cytokine IL-10. Data shown are means ± S.D. of three experiments. Error bars represent S.D. *, significant differences between control and LPS treatment, p < 0.05; #, significant differences between Gstp1/p2−/− and WT DCs after LPS treatment, p < 0.05. F, BMDDCs treated with 200 ng/ml LPS overnight were pulsed with 1 μg/ml hgp100(25–33) peptide in T-cell medium for 2 h and then cocultured with CFSE-labeled, hgp100(25–33)-specific Pmel-1 TCR transgenic T cells (DC/T cell ratio = 1:5) for 3 days. T-cell proliferation by CFSE dilution was measured by flow cytometry. Plots are gated on CD8+ cells. The proliferation profiles shown are representative of three experiments.
Gstp1/p2 depletion in BMDDCs results in increased glycolysis
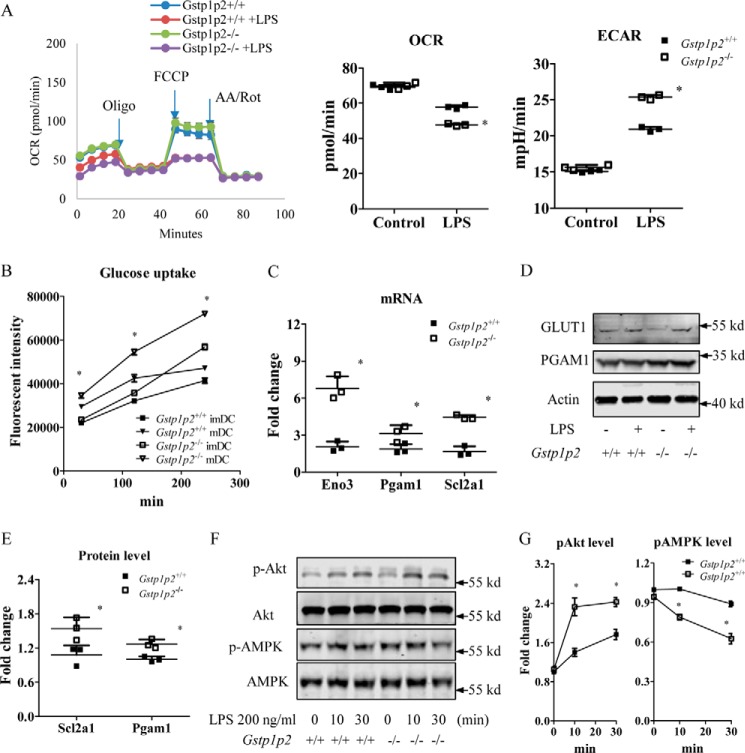
It seemed reasonable to expect that enhanced proliferation rates and DC activation by LPS should be accompanied by changes in cellular metabolism and bioenergetics. To support increased demands for synthesis and transport of proteins required for BMDDC maturation, recent evidence suggests that LPS activation of DCs drives a decline in oxidative phosphorylation (OXPHOS) and commitment to glycolysis (provides ATP as well as generates lipids for membrane synthesis, including endoplasmic reticulum and Golgi (20–22)). Because ERα has been shown to affect glucose metabolism (19), we reasoned that Gstp1/p2−/− BMDDCs should be metabolically different from WT. To obtain insight into functional differences between WT and Gstp1/p2−/− BMDDC mitochondria, we first mapped mitochondrial function by using a Seahorse Metabolic Analyzer to measure oxygen consumption rate (OCR), an indicator of oxidative phosphorylation, and extracellular acidification rate (ECAR), an indicator of glycolysis. For both WT and Gstp1/p2−/− BMDDCs, stimulation with LPS resulted in decreased OCR and an accompanying increased ECAR as compared with control (Fig. 3A). This indicated attenuated rates of OXPHOS and enhanced glycolysis. In addition, following LPS activation, Gstp1/p2−/− BMDDCs had significantly higher ECAR values when compared with WT cells, indicating their higher glycolytic capacity (Fig. 3A). Meanwhile, OCR values were significantly lower in Gstp1/p2−/− BMDDCs compared with WT cells (Fig. 3A). Furthermore, Fig. 3B shows the time-dependent uptake of glucose in BMDDCs. Stimulation with LPS resulted in increased glucose uptake in both WT and Gstp1/p2−/− mDCs. A comparison of glucose uptake rates revealed that Gstp1/p2−/− mDCs were more glycolytic than WT cells. For additional clarification, we used quantitative RT-PCR to measure mRNA expression levels of various glycolysis-associated genes, including Aldoa, Aldoc, Eno1, Eno3, Gck, Gpi, Hk2, Ldha, Ldhb, Ldhc, Pfk, Pgam1, Pgk1, pkm, Slc2a1, and Slc2a4. Among these, our data showed that the mRNA expression levels of several glycolytic genes were severalfold higher in Gstp1/p2−/− BMDDCs (Fig. 3C), including enolase (Eno3), phosphoglycerate isomerase (Pgam1), and glucose transporter 1 (Slc2a1). In addition, the more pronounced up-regulation of protein levels of phosphoglycerate isomerase (PGAM1) and glucose transporter 1 (GLUT1) was confirmed in Gstp1/p2−/− BMDDCs. Due to the availability and sensitivity of antibodies, β-enolase was not detected in this study (Fig. 3, D and E).
Figure 3.
Gstp1/p2−/− BMDDCs are more glycolytic after LPS activation. A, BMDDCs were seeded in a Seahorse XF96 microplate and stimulated overnight with DC medium (as a control) or 200 ng/ml LPS, and real-time OCR during sequential treatments with oligomycin (Oligo; ATP synthase inhibitor), carbonyl cyanide p-trifluoromethoxyphenylhydrazone (FCCP), and antimycin A/rotenone (AA/Rot; electron transport chain inhibitors) and basal OCR and ECAR were determined by the Seahorse XF96 analyzer. Data shown are means ± S.D. of three experiments. Error bars represent S.D. *, significant differences between Gstp1/p2−/− and WT mDCs, p < 0.05. mpH, milliunit of pH value. B, glucose uptake levels were evaluated using the fluorescently labeled deoxyglucose analog 2-NBDG. Data shown are means ± S.D. of three experiments. Error bars represent S.D. *, significant differences between Gstp1/p2−/− and WT mDCs, p < 0.05. C, quantitative RT-PCR analysis of the expression of key genes associated with glycolysis. Data shown are means ± S.D. of three experiments. Error bars represent S.D. *, significant differences between Gstp1/p2−/− and WT mDCs, p < 0.05. D and E, the protein levels of associated genes were further evaluated by immunoblotting with GLUT1, PGAM1, and β-enolase antibodies and quantified with Image Studio 4.0 software. β-Enolase was not detectable in BMDDCs. Results are representative of three experiments. *, significant differences between Gstp1/p2−/− and WT mDCs, p < 0.05. F and G, phosphorylation (p) levels of Akt and AMPK were evaluated by immunoblotting and quantified with Image Studio 4.0 software. Data shown are means ± S.D. of three experiments. Error bars represent S.D. *, significant differences between Gstp1/p2−/− and WT DCs, p < 0.05.
Increased glycolysis in Gstp1/p2−/− BMDDCs was accompanied by an increased activation of the mTOR pathway. This was assessed by phosphorylation of Akt and decreased activation of AMPK as determined by phosphorylation of AMPK (Fig. 3, F and G). Akt activation promotes the shift to glycolysis and anabolic metabolism required for DC activation and immunogenicity, whereas AMPK activation antagonizes these responses (21). These results support the concept that ablation of GSTP contributes directly to the shift toward these pathways.
Gstp1/p2−/− BMDDCs show different redox responses after LPS stimulation
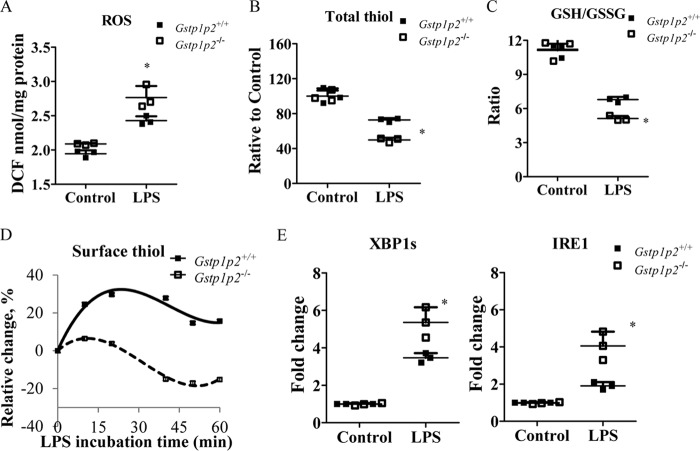
Compared with WT, Gstp1/p2−/− BMDDCs had higher endogenous levels of ROS (Fig. 4A) even before LPS treatment. When stimulated with LPS, there was a further increase in ROS coincident with a decrease in both total cellular thiols (Fig. 4B) and cell surface thiols (Fig. 4D), and the GSH/GSSG ratio was shifted toward a more oxidized state (Fig. 4C). Thus, LPS caused a general shift in oxidative conditions, but each measure was more pronounced in the Gstp1/p2−/− cells. Furthermore, LPS also induced an unfolded protein response (UPR) in BMDDCs. This is exemplified by the IRE1/XBP1 pathway response where Gstp1/p2−/− BMDDCs exhibited a more marked activation compared with WT (Fig. 4E). These results were similar to our earlier studies (24) where both thapsigargin and tunicamycin caused a more pronounced UPR in GSTp1/p2−/− BMDDCs compared with wildtype. IRE1/XBP1 activation has been shown to be essential for the development of DCs and regulation of innate immune responses in macrophages (39, 42). Therefore, the higher UPR activation with LPS in Gstp1/p2−/− cells might be correlated to its enhanced functionality as well.
Figure 4.
BMDDCs from Gstp1/p2−/− mice have higher levels of oxidative stress and UPR during LPS activation. A, cellular ROS levels were measured using a proprietary quenched fluorogenic probe, DCFH-DiOxyQ. B, intracellular reduced thiol levels were measured by thiol fluorescent probe IV. C, intracellular GSH levels were measured by thiol fluorescent probe IV after protein precipitation. Intracellular GSSG levels were determined based on the reduction of GSSG in the presence of glutathione reductase and NADPH. Data are presented as GSH/GSSG ratio. D, total surface reduced thiol levels were determined by an immediate fluorescence increase upon addition of ThioGlo-1. E, UPR genes IRE1 and XPB1s were quantified by quantitative RT-PCR. Data shown are means ± S.D. of three experiments. Error bars represent S.D. *, significant differences between Gstp1/p2−/− and WT after LPS treatment, p < 0.05. DCF, dichlorofluorescein.
ERα is subject to S-glutathionylation under oxidative stress
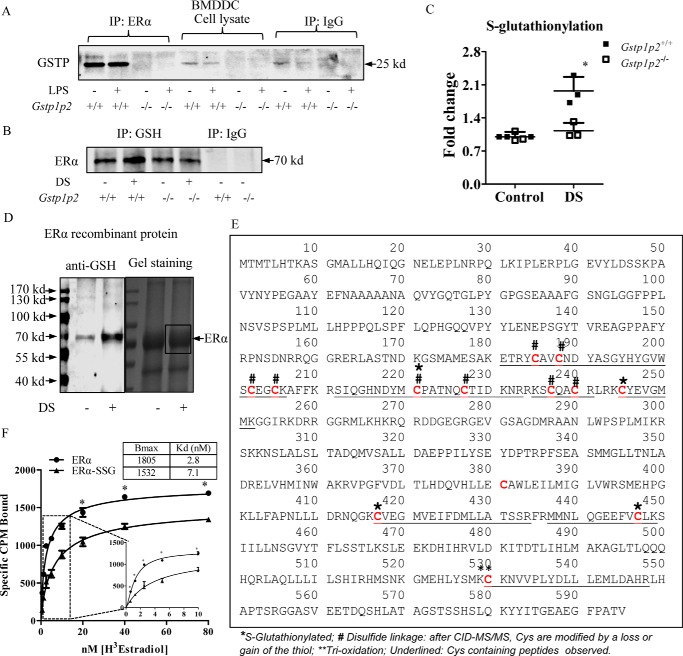
Fig. 5A shows that, in WT BMDDCs, GSTP and ERα coimmunoprecipitated with antibodies to ERα, indicating that they form part of a protein complex. Presumably as a consequence of this interaction, further immunoprecipitation with anti-GSH antibodies showed that, following ROS generation by disulfiram, ERα was a substrate for S-glutathionylation (Fig. 5, B and C). In the Gstp1/p2−/− cells, there was no complex formation (Fig. 5A), and levels of ERα S-glutathionylation were significantly lower (Fig. 5, B and C).
Figure 5.
GSTP catalyzes ERα S-glutathionylation through protein-protein interaction and regulates ERα binding affinity for estradiol. A, 500 μg of BMDDC lysates, untreated or treated with 200 ng/ml LPS (overnight), from WT and Gstp1/p2−/− mice were used for immunoprecipitation (IP) with anti-ERα antibodies. Samples were analyzed by SDS-PAGE with the IgG controls and blotted for GSTP. B and C, 500 μg of BMDDC lysates, untreated or treated with disulfiram (DS) to promote S-glutathionylation (10 μm for 30 min at 37 °C), from WT and Gstp1/p2−/− mice were used for immunoprecipitation with anti-GSH antibodies. Samples were analyzed by SDS-PAGE with the IgG controls, blotted for ERα, and quantified with Image Studio 4.0 software. Results are representative of three experiments. *, significant differences between Gstp1/p2−/− and WT, p < 0.05. D, recombinant ERα proteins, untreated and S-glutathionylated (10 μm disulfiram and 10 mm GSH for 30 min at 37 °C), were separated on a non-reducing gel. One part of the gel was transferred to a PVDF membrane and blotted with anti-GSH antibody (left panel), and the other part of the gel was stained by colloidal Coomassie stain (right panel). Bands (as indicated in the box) were excised, destained, enzymatically digested, and subjected to LC-MS/MS identification. E, proteomic identification of sites of cysteine modification on ERα (*, S-glutathionylated; #, disulfide linkage; **, trioxidation). F, saturation binding assays of estradiol. Recombinant ERα protein (untreated or S-glutathionylated) was incubated with [3H]estradiol (from 0.625 to 80 nm) at room temperature for 90 min, and bound estradiol was measured by liquid scintillation counting. Data were analyzed using a specific binding module from Prism 5.0. Maximum amounts of estradiol bound receptor (Bmax) and binding affinities (Kd) were calculated. Data shown are means ± S.D. of three experiments. Error bars represent S.D. *, significant differences between ERα and ERα-SSG, p < 0.05.
Proteomic identification of cysteine S-glutathionylation in ERα
Of the 595 amino acids in human ERα, there are 13 cysteines, all of which are conserved between mouse and human. Human recombinant ERα proteins were treated with disulfiram and separated on a non-reducing gel, and S-glutathionylation of ERα was confirmed by Western blotting using anti-GSH antibodies (Fig. 5D, left panel). The gel was stained with colloidal Coomassie stain, and the result confirmed even loading of ERα under both control and disulfiram-treated conditions (Fig. 5D, right panel). Protein bands (as indicated in the box) were excised, destained, enzymatically digested, and subjected to LC-MS/MS identification. Mass spectrometric analysis of recombinant ERα revealed that cysteines 185, 188, 202, 205, 221, 227, 237, and 240 were involved in intra- or intermolecular disulfide linkages. Following fragmentation, these cysteines were observed with either a sulfide or had lost the thiol group. Cys-381 was not detected, and Cys-530 was oxidized. Cysteines 221, 245, 417, and 447 were S-glutathionylated and yielded a diagnostic glutathione ion at m/z 308 following electron transfer dissociation (Fig. 5E and Table 2). Characteristic fragmentation patterns of S-glutathionylated peptides, details of the observed sites of cysteine modification, and tandem mass spectra are provided in Figs. S2 and S3.
Table 2.
Cysteine modifications observed in human ERα (P03372)
Tandem mass spectra are provided in the supporting information. Dha, dehydroalanine.
| Residues | Cys positions | Site of S–GSH | All modifications | Peptide sequence | MS/MS | Precursor m/z (MS/MS) | z | Enzyme |
|---|---|---|---|---|---|---|---|---|
| 184–206 | 185, 188, 202, 205 | Sulfide (Cys); Cys to Dha (Cys) | YCAVCNDYASGYHYGVWSCEGCK | CID | 1288.48 | 2 | Trypsin | |
| 190–206 | 202, 205 | Sulfide (Cys); Cys to Dha (Cys) | DYASGYHYGVWSCEGCK | ETD | 641.59 | 3 | Lys-C, Asp-N | |
| 212–231 | 221, 227 | Sulfide (Cys); Cys to Dha (Cys); trioxidation (Cys) | SIQGHNDYMCPATNQCTIDK | CID | 1127.47 | 2 | Trypsin | |
| 212–233 | 221, 227 | 221 | S-Glutathione (Cys); Cys to Dha (Cys) | SIQGHNDYMCPATNQCTIDKNR | ETD | 696.05 | 3 | Trypsin |
| 235–241 | 237, 240 | Sulfide (Cys); Cys to Dha (Cys) | KSCQACR | CID | 397.17 | 2 | Trypsin | |
| 245–252 | 245 | 245 | S-Glutathione (Cys) | CYEVGMMK | ETD | 422.49 | 2 | Trypsin |
| 245–252 | 245 | 245 | S-Glutathione (Cys); oxidation (Met) | CYEVGMMK | ETD | 427.82 | 2 | Trypsin |
| 417–434 | 417 | 417 | S-Glutathione (Cys) | CVEGMVEIFDMLLATSSR | CID | 1154.01 | 2 | Trypsin |
| 417–434 | 417 | 417 | S-Glutathione (Cys); oxidation (Met) | CVEGMVEIFDMLLATSSR | ETD | 769.68 | 3 | Trypsin |
| 437–449 | 447 | 447 | S-Glutathione | MMNLQGEEFVCLK | CID | 923.89 | 2 | Trypsin |
| 437–449 | 447 | 447 | S-Glutathione (Cys); oxidation (Met) | MMNLQGEEFVCLK | ETD | 621.6 | 3 | Trypsin |
| 530–548 | 530 | Trioxidation (Cys); oxidation (Met) | CKNVVPLYDLLLEMLDAHR | CID | 770.06 | 3 | Lys-C, trypsin |
Cysteine S-glutathionylation causes functional changes in ERα
To determine whether this post-translational modification caused functional changes in ERα function, we performed estrogen binding assays. Fig. 5F shows the radioligand binding results, establishing that S-glutathionylation reduced the total density of receptors (Bmax, 1805 versus 1532) and altered the equilibrium dissociation constant (Kd, 2.8 versus 7.1 nm). From these results, we calculated that S-glutathionylation of ERα reduced the overall binding potential for estrogen (receptor density × affinity) 3-fold, from 645 to 216. Overall, these findings suggest that GSTP-mediated S-glutathionylation of ERα decreases ERα activity.
Discussion
A primary rationale for the present study was derived from the original observation that genetic ablation of GSTP caused increased expression of ERα and implied the existence of a significant link between the two. In this context, mice deficient in either GSTP or ERα are known to be more susceptible to infections, suggesting that innate immunity is impaired (25, 26). Moreover, ERα signaling controls the development of dendritic cells, initiating the processes of innate and adaptive immunity (18), and regulates the expression of genes that influence lymphocyte metabolism and activation (27). In both human and mouse bone marrow, HSCs express ERα with levels coincident with the processes that lead to development of a mature immune system in neonatal life (28, 29). Greater levels of ERα signaling can regulate HSC numbers and BMDDC development and result in greater numbers of dendritic cells that will modulate the scale and/or quality of either adaptive or innate immunity (29). Our present results suggest linkages for these diverse observations and illustrate how altered redox homeostasis influences ERα and the processes that lead to BMDDC differentiation.
Although we found that resting levels of ERα were elevated in all tissues of the Gstp1/p2−/− mouse, our results also showed that GSTP-deficient mice were more sensitive to the effects of LPS at amplifying pathways leading to BMDDC activation. Although GSTP has some detoxification enzyme properties, LPS is not a substrate. Instead, its capacity to act as a thiolase and catalyze S-glutathionylation of ERα can provide an explanation for the observed effects. We showed that GSTP forms a complex with ERα, but in perspective, ERα interacts with ∼700 proteins, so in and of itself, its interaction with GSTP may not be remarkable. In the same vein, GSTP also has a tendency to form protein complexes with a large number of proteins, primarily because such proteins tend to be substrates for S-glutathionylation. In this regard, the donation of the –SG moiety to an activated cysteine thiolate anion would imply that part of ERα is recognized by the designated H-site on GSTP. This catalysis is facilitated by GSTP forming a temporal, reversible association with ERα, allowing for the transfer of the –SG moiety from the enzyme's “G” site (30). Whether GSTP and ERα form a direct or indirect (part of a larger protein cluster) complex (24, 31), it is apparent that the latter is a substrate for S-glutathionylation. Because S-glutathionylation has a profound impact on the structure, function, and subcellular distribution of most proteins (13, 32), it seems reasonable to suggest that ERα will be similarly impacted. Of the 595 amino acids in human ERα, there are 13 cysteines, all of which are conserved between mouse and human. In this study, we identified four cysteines that were susceptible to S-glutathionylation, residues 221, 245, 417, and 447. Cysteine 530 of the human ERα is the main covalent attachment site of photoaffinity-labeled estradiol and is in close proximity to Cys-417 and Cys-447, each located at the border (or in structural elements) delineating the hormone-binding pocket (33). S-Glutathionylation of any of these residues would also serve to impact the conformation of the binding pocket, influencing steroid recognition and binding as well as subcellular localization. Cysteines 221 and 245 are located in the DNA-binding region of the protein, a section that generally encompasses residues 185–250. Why does the absence of GSTP cause such fundamental changes? Our results suggest a link between estrogen receptor–related effects and altered capabilities to maintain redox characteristics provided by GSTP. For example, we found that S-glutathionylation of ERα caused a mild reduction in receptor density (Bmax) and reduced the affinity for 17β-estradiol. In contradistinction, palmitoylation of cysteine 447 (34, 35) of ERα enhances the binding affinity, indicating that the effects are dependent upon both the site and type of post-translational modification (36). Moreover, although there are somewhat limited results indicating that enhanced transcriptional activities correlate with increased binding affinity values, this relationship may not be universal (37), and other factors such as ERα dimer formation, translocation to cell compartments, altered half-life, or altered recognition of protein binding partners may play a role in regulating ERα functions. The fact that ERα has a comparatively large number of potential binding partners might also influence interpretation.
Dendritic cells are critical regulators of innate and acquired immunities, responding to stimuli, frequently downstream of toll-like receptors, that program them to undergo metabolic changes. Earlier results showed that toll-like receptor agonists caused BMDDCs to transition from oxidative phosphorylation to aerobic glycolysis, an event promoted by Akt signaling and inhibited by AMPK phosphorylation (38). Our present results suggest that ablation of GSTP is sufficient on its own to initiate similar kinase pathway responses and substantiate the involvement of mTOR signaling pathways as mediators of augmented energy production, particularly aerobic glycolysis, enhanced proliferation, and differentiation in BMDDCs. These pathway changes also establish a linear relationship with commensurate diminishment of S-glutathionylation of ERα.
Thus, the higher levels of S-glutathionylation of ERα in the wildtype BMDDCs alter its transcriptional activities and adjust gene expression patterns to favor proliferation and differentiation. In turn, this would provide an explanation of why Gstp1/p2−/− BMDDCs exhibit a glycolytic phenotype. Enhanced energy production can then give rise to an increased proinflammatory phenotype in these cells. Induction of glycolysis plays a central role for BMDDCs to acquire immunogenic properties following activation (21). Under non-inflammatory conditions, most BMDDCs exist in a quiescent, immature state and are poorly immunogenic. However, upon triggering of toll-like receptor by pathogen-derived products or inflammatory stimuli, BMDDCs undergo activation/maturation and are highly immunogenic. OXPHOS is associated with BMDDC quiescence, whereas BMDDC activation and immunogenicity rely heavily on glycolysis to provide carbon sources for de novo fatty acid synthesis to permit increased production and secretion of mediators. Comparing the quantitative RT-PCR data for the GSTP knockout and wildtype BMDDCs, a series of expression changes are consistent with the advancement of glycolysis. Eno3 is an enolase glycolytic enzyme that catalyzes the reversible conversion of 2-phosphoglycerate to phosphoenolpyruvate. Pgam1 is an isomerase that transfers a phosphate group from the C3 carbon of 3-phosphoglycerate to the C2 carbon, forming 2-phosphoglycerate. Slc2a1 is the gene for GLUT1, a glucose transporter, highly conserved in humans and mice and is one of a family of 14 genes encoding GLUT proteins. It functions through maintenance of the low levels of basal glucose uptake required to sustain respiration. In cell membranes, GLUT1 levels can increase or decrease, respectively, in response to low or high glucose availability. It is important to remember that these alterations are found only as a consequence of ablation of GSTP. There is evidence that BMDDC functions may also be influenced by endoplasmic reticulum–induced stress, particularly as they relate as precursors of the unfolded protein response (39–42). We previously showed that markers for UPR, including IRE1 and ATF6, are constitutively higher in cells from Gstp1/p2−/− mice and that these animals have a narrower threshold of tolerance to oxidative stress compared with wildtype (24). In this regard, such data indicate that Gstp1/p2−/− mice are under higher levels of constitutive oxidative stress. Our present results indicate that LPS induces in BMDDCs the IRE1/XBP1 pathways linked with UPR and that the effects are more marked in Gstp1/p2−/− BMDDCs compared with wildtype (Fig. 4E).
Manipulation of metabolism of BMDDCs is quite critical as a determinant factor in mounting an immune response. The results that our Gstp1/p2−/− BMDDCs are more glycolytic and have a more proinflammatory phenotype suggest a potential for translational applications to assess whether pharmacological inhibition of GSTP can be used as a strategy to enhance the immunogenicity of BMDDCs in therapeutic settings. As such, the fact that redox intercepts estrogen-mediated signaling pathways provides a framework for linking those events that underlie immunity and bone marrow pathologies. For example, clinical results indicate that, in myeloma, patients homozygous for a variant allele of GSTP (one that we have shown to be less effective at catalyzing S-glutathionylation (43)) have greater progression-free survival (44). High levels of ERα are also present in the majority of multiple myeloma patient samples (45). Consequently, there may be a clinical correlate through lowered ERα S-glutathionylation levels in this patient population. Previously, the pharmaceutical inhibition of GSTP through the use of Telintra has proved effective in the management of myelodysplastic syndrome patients, in some cases causing the differentiation of precursor cancer cells, prior to establishment of a more malignant phenotype (15, 16). There are numerous other examples where drugs that alter redox homeostasis cause enhanced proliferation and functional changes in hematopoietic progenitor cells. At the simplest level, N-acetylcysteine (a precursor of cysteine) can enhance intracellular concentrations of GSH, reverse ROS suppression of N-cadherin–mediated hematopoietic stem cell adhesion to osteoblasts, and induce cell migration (46). Cysteine prodrugs can alter redox balance and enhance self-renewal of progenitor cells, whereas agents that diminish GSH, such as buthionine sulfoximine, can promote differentiation (4, 5). It appears that redox gradients in the marrow compartment are amenable to pharmacological manipulation, and altered redox gradients can influence HSC migration, differentiation, and myeloproliferation (9). Telintra continues to be developed in therapeutic settings, and its capacity to impair S-glutathionylation is a likely conduit to its clinical activities.
Experimental procedures
Mice
C57BL/6 wildtype mice were purchased from The Jackson Laboratory (Bar Harbor, ME). Gstp1/p2−/− mice were generated as described earlier (26). The mice were bred and kept in an Association for Assessment and Accreditation of Laboratory Animal Care –certified animal facility of the Medical University of South Carolina. All of the mice were used at ∼8–12 weeks of age. The Institutional Animal Care and Use Committee of Medical University of South Carolina approved all the experimental procedures used in this study.
Primary BMDDC culture conditions
Immature bone marrow–derived dendritic cells were generated according to procedures reported previously (9). Briefly, BM cells (4–5 × 105/ml, 10 ml/plate) were plated in RPMI 1640 medium (HyClone, Logan, UT) supplemented with 10% fetal bovine serum (FBS; Atlas Biologicals, Fort Collins, CO), 100 units/ml penicillin, 100 μg/ml streptomycin (all from Mediatech, Manassas, VA), and 20 ng/ml recombinant mouse GM-CSF (BioAbChem, Ladson, SC) (DC medium) into 100-mm culture dishes (Sarstedt, Newton, NC). Fresh DC medium was added on day 4 and gently replaced by fresh DC medium containing 10 ng/ml recombinant mouse GM-CSF on day 7. For mDCs, LPS (200 ng/ml; Sigma) was added to the culture described above, and the cells were cultured further overnight (∼20 h).
Antibodies
The following antibodies were used for immunoblotting: rabbit monoclonal anti-ERα (Millipore, Burlington, MA; 04-820, Lot 2135146; 1:500 dilution); rabbit polyclonal anti-GSTP (MBL International, Woburn, MA; 311, Lot 071; 1:1000 dilution); mouse monoclonal anti-GSH (Virogen, Watertown, MA; 101-A, Lot GS-D8/G2-68; 1:1000 dilution); rabbit polyclonal anti-β-actin (ab8227, Lot GR60359-1), rabbit polyclonal anti-glucose transporter GLUT1 (ab32551, Lot GR208248-1), and goat polyclonal anti-GSTP (ab53943, Lot GR121458-9) (all from Abcam, Cambridge, MA; 1:1000 dilution); rabbit polyclonal anti-Akt (9272, Lot 24), rabbit polyclonal anti-phospho-Akt (Ser-473; 9271, Lot 22), mouse monoclonal anti-AMPK (2793, Lot 4), and rabbit polyclonal anti-phospho-AMPK (Thr-172; 2535, Lot 16) (all from Cell Signaling Technology, Danvers, MA; 1:500 dilution); mouse monoclonal anti-PGAM1 (sc-130334, Lot G0317) and mouse monoclonal anti-β-enolase (sc-10081, Lot G2417) (both from Santa Cruz Biotechnology, Dallas, TX; 1:200 dilution); and IRDye 800CW goat anti-mouse IgG (926-32210, Lot C20510-05), IRDye 800CW goat anti-rabbit IgG (926-32211, Lot C21227-02), 800CW donkey anti-goat IgG (926-32214, Lot C60119-05), IRDye 680RD goat anti-mouse IgG (926-68070, Lot C70427-05), IRDye 680RD goat anti-rabbit IgG (926-68071, Lot C20530-01), and IRDye 680RD donkey anti-goat IgG (926-68074, Lot C20606-04) (all from LI-COR Biosciences, Lincoln, NE; 1:15,000 dilution).
Immunoblotting
Total soluble protein was quantitated by BCA protein assay (Pierce). Cell lysates were resolved in SDS loading buffer (80 mm Tris-HCl, pH 6.8, 2% SDS, 10% glycerol, 0.02% bromphenol blue, ±5 mm tris(2-carboxyethyl)phosphine) and heated at 95 °C for 5 min. Equal amounts of protein were electrophoretically separated by SDS-PAGE (Bio-Rad) and transferred onto Low Fluorescent PVDF membranes (Millipore) or nitrocellulose membranes (Bio-Rad) by a Trans-Blot Turbo Transfer System (Bio-Rad). PVDF or nitrocellulose membranes were incubated in Odyssey blocking buffer (LI-COR Biosciences) for 1 h to reduce nonspecific binding and then probed with appropriate primary antibodies (diluted in Odyssey blocking buffer) at 4 °C overnight. Immunoblots were then developed with infrared (IR) fluorescence IRDye secondary antibodies, imaged with a two-channel (red and green) IR fluorescence Odyssey CLx imaging system (LI-COR Biosciences), and quantified with Image Studio 4.0 software (LI-COR Biosciences).
Cell labeling and flow cytometry
Labeling was performed at room temperature using standard techniques. Cells were incubated with saturating concentrations of flow antibodies for 20 min, washed twice, and analyzed using a BD FACSVerse flow cytometer equipped with three lasers (407, 488, and 640 nm). 20,000 events per sample were acquired and analyzed using BD FACSuite software. The LIVE/DEAD Fixable Near-IR Dead Cell Stain kit (Invitrogen) was used to gate live cells. Data were analyzed with FlowJo software (Tree Star, Ashland, OR).
FITC-dextran uptake (endocytosis)
Immature dendritic cells were suspended in DC medium and incubated with 1 mg/ml FITC-dextran (molecular weight, 40,000; Sigma) for 4 h at 4 or 37 °C. The cells were washed two times with ice-cold PBS and 1% BSA and labeled with anti-mouse antibody mixture including APC-CD11c (from eBioscience, San Diego, CA) and PE-CD3e, PE-CD45R/B220, PE-CD14, and PE-NK1.1 (all from BD Pharmingen). The cells were washed two times, fixed with 2% paraformaldehyde, and analyzed by flow cytometry. The uptake was calculated as the change in MFI between cell samples incubated at 37 and 4 °C.
ELISAs
Measurements of IL-12p70 and IL-10 in imDC or mDC supernatants were performed using the ELISA Ready-Set-Go! Reagent set from eBioscience according to the manufacturer's protocol.
DC–T cell coculture
Splenocytes from Pmel-1 TCR transgenic C57BL/6 mice were depleted of erythrocytes by ammonium-chloride-potassium lysing buffer (Life Technologies). Isolation of T cells was achieved using the Dynabeads Untouched Mouse T Cells kit (Life Technologies) according to the manufacturer's protocol. Isolated T cells were labeled with 2.5 μm intracellular fluorescent dye CFSE (Life Technologies) and then cocultured (1 × 106 cells/well) with hgp100(25–33)-pulsed (1 μg/ml for 2 h) mDCs (2 × 105 cells/well) in 48-well plates (Sarstedt) in T-cell medium (RPMI 1640 medium supplemented with 10% FBS, 100 units/ml penicillin, 100 μg/ml streptomycin, 1 mm sodium pyruvate, 1× nonessential amino acids, 10 ng/ml recombinant mouse IL-2 (BioAbChem)). After 3 days, the cells were labeled with anti-mouse PE-CD8a (BD Pharmingen), and T-cell proliferation was analyzed by flow cytometry. As a control, splenocytes from Pmel-1 TCR transgenic mouse were labeled with CFSE, cultured in T-cell medium in the presence of 1 μg/ml hgp100(25–33) for 3 days, and examined for T-cell proliferation.
Measurement of intracellular ROS
Intracellular ROS was measured using the OxiSelect In Vitro ROS/RNS kit from Cell Biolabs (San Diego, CA) according to the manufacturer's protocol. Briefly, BMDDCs (1.0–2.0 × 106 cells/ml) were suspended in DC medium and incubated with 200 ng/ml LPS at 37 °C at the indicated times. Cells were harvested and solubilized by ice-cold lysis buffer (50 mm Tris-HCl pH 7.5, 150 mm NaCl, 1% Triton, 1 mm EDTA, 1 mm EGTA, protease inhibitor mixture (Roche Applied Science/Sigma)). Cell lysates were collected and immediately subjected to the ROS/RNS measurement. The fluorescence intensity of fluorophore dichlorofluorescein, which was formed by peroxide oxidation of the non-fluorescent precursor dichlorodihydrofluorescin (DCFH), was detected at 480-nm excitation/530-nm emission using a SpectraMax M5 Multi-Mode microplate reader (Molecular Devices, Sunnyvale, CA). DCFH with lysis buffer was used as a blank control.
Measurement of intracellular reduced thiol levels
Immature DCs (1.0–2.0 × 106cells/ml) were suspended in DC medium and incubated with 200 ng/ml LPS overnight at 37 °C. Cells were harvested and solubilized by ice-cold lysis buffer as mentioned above. Cell lysates were collected and immediately subjected to reduced thiol measurement using thiol fluorescent probe IV (Millipore). Fluorescence intensities were detected at 400-nm excitation/465-nm emission using a microplate reader. Thiol fluorescent probe IV with lysis buffer was used as a blank control.
Measurement of surface reduced thiol levels
ThioGlo-1 (TG-1; Calbiochem) was used to monitor the cell surface reduced thiols as reported earlier (11). Briefly, BMDDCs (1.0 × 106 cells/ml) were suspended in PBS and 100 μm CaCl2 and stimulated with 200 ng/ml LPS for the indicated times. Total surface reduced thiol levels were then determined by an immediate fluorescence increase upon addition of TG-1 (final concentration, 5 μm) at 379-nm excitation/513-nm emission, measured by a QuantaMaster spectrofluorometer equipped with an A-1010B continuous xenon arc lamp and an 814 Photomultiplier Detection System (Photon Technology International, Birmingham, NJ) in a quartz cuvette under constant stirring at 37 °C, controlled by a TC 125 temperature controller (Quantum Northwest, Shoreline, WA). The emission of each sample was recorded for 30 s (background) before and until 250 s after the addition of TG-1 (resolution, 0.1 s). Saturated TG-1 fluorescence values were corrected for background emissions, normalized for cell numbers, and averaged using SigmaPlot 10.0 software (Systat Software, San Jose, CA).
Measurement of GSH and GSSG levels
Quantitative determinations of GSH and GSSG levels were performed using the enzymatic recycling method (48). Briefly, protein in the cell extracts was precipitated by sulfosalicylic acid, and the supernatant was then divided into two parts. For reduced GSH, the supernatant was incubated with thiol fluorescent probe IV, and fluorescence intensities were measured at 400-nm excitation/465-nm emission. For total GSH (GSH + GSSG), the supernatant was neutralized by triethanolamine and incubated with the reduction system (containing NADPH and glutathione reductase) at 37 °C for 20 min. GSSG was calculated based on the results from reduced GSH and total GSH; the ratio of GSH/GSSG = [GSH]/(([Total GSH] − [GSH])/2).
Glucose uptake
Glucose uptake was determined by 2-NBDG (Cayman Chemical, Ann Arbor, MI) according to the manufacture's protocol. 2-NBDG is a fluorescently labeled deoxyglucose analog. Briefly, imDCs or mDCs (0.2 × 106 cells/well in 100 μl) were seeded in a 96-well black clear-bottom plate (Greiner) in glucose- and pyruvate-free RPMI 1640 medium and incubated with 100 μg/ml 2-NBDG at 37 °C at the indicated times. Cells were washed with PBS and resuspended in 200 μl of PBS, and the fluorescence intensities were measured at 485-nm excitation/535-nm emission using a microplate reader.
Proteomic analyses: Identification of S-glutathionylated cysteine
Recombinant human ERα protein (Thermo Fisher Scientific, Waltham, MA) (UniProtKB P03372) was S-glutathionylated and separated on a non-reducing gel. Following colloidal Coomassie staining with ProtoBlue Safe (National Diagnostics, Atlanta, GA), bands were excised, destained, and enzymatically digested with porcine trypsin (Sigma) at a 1:20 enzyme/protein ratio sequentially with lysyl endopeptidase (Wako, Richmond, VA) and Asp-N (Roche Applied Science/Sigma) at a 1:20 enzyme/protein ratio or with Glu-C (Thermo Fisher Scientific) at a 1:15 enzyme/protein ratio. Digestions were performed at 37 °C for 15 h with agitation in 100 mm ammonium bicarbonate. The digested peptides were extracted with 5% formic acid in 50% acetonitrile or 85% acetonitrile sequentially and dried under vacuum, and particulates were removed using a μC18 ZipTip (Millipore).
LC-MS/MS
Peptides dissolved in 2% formic acid, 2% acetonitrile were loaded onto a C18 PepMap 100 (300-μm × 5-mm) trap column (Thermo Fisher Scientific) for 10 min at 30 μl/min and separated on a 75-μm internal diameter × 15-cm fused-silica column packed in house with ReproSil-Pur 120 C18-AQ (1.9 μm; Dr. Maisch GmbH, Germany) at 180 nl/min using a gradient of 5–50% solvent B in 180 min on an LC Packings U3000 Nano LC system. Solvent A was 0.2% formic acid in 5% acetonitrile; solvent B was 0.2% formic acid in 80% acetonitrile. Peptides were mass-analyzed on a Thermo Orbitrap Elite with alternating collision-induced dissociation (CID) and electron transfer dissociation (ETD) fragmentation. Mass spectra were acquired in data-dependent mode using a top 7 method. Each FTMS survey scan was acquired with a mass range of m/z 400–1700 in the Orbitrap followed by acquisition of the tandem mass spectra of the seven most intense ions in the dual-pressure ion trap. The automatic gain control (AGC) target value in the Orbitrap was 106 for the survey FTMS scan at a resolution of 60,000 at m/z 400. A normalized collision energy of 35% was used for CID with an AGC of 1000. The ETD reaction time was adjusted based on the charge state of the ion, AGC was set to 104 ions, and supplemental activation was enabled. The threshold for triggering CID and ETD was 500. Dynamic exclusion was enabled with a repeat count of 1 (or 2 depending on the run), repeat duration of 30 s, and exclusion duration of 180 s. The chromatography feature was enabled for all runs. Lock mass was not utilized. Targeted LC-MS/MS analyses were also performed using inclusion masses for predicted S-glutathionylated peptides.
Database searching
The CID and ETD MS/MS spectra were searched using Mascot (version 2.4.01) and SEQUEST HT search algorithms within Proteome Discoverer 1.4 (Thermo Fisher Scientific) against a UniProtKB human database (Swiss-Prot + TrEMBL downloaded on August 23, 2016) containing 154,527 entries plus digestion enzymes. For both algorithms, the search parameters allowed for three missed cleavages, precursor mass tolerances of ±25 ppm, and fragment mass tolerances ±0.8 Da. Dynamic modifications on cysteines included glutathionylation (+305.0682 Da with a +129.0426-Da neutral loss), conversion of cysteine to dehydroalanine (−33.9877 Da), sulfide (+31.9721 Da), and trioxidation (+47.9847 Da). Methionine oxidation was also used as a variable modification. Tandem mass spectra were also searched using the Andromeda algorithm within MaxQuant version 1.5.5.1 (23) using the same database and modifications as above. Following recalibration, peptides were filtered using a 4.5-ppm precursor mass tolerance. CID and ETD spectra were also converted to “Mascot generic format (mgf)” and searched using Protein Prospector version 5.19.1 (47) against the estrogen receptor sequence (P03372) with the same variable modifications as above, including searching for disulfide linkages.
All identified spectra were manually inspected. The ETD spectra of S-glutathionylated peptides with charge states greater than +2 were inspected for the presence of the diagnostic glutathione ion at m/z 308. Both the CID and ETD spectra were also inspected for the presence of fragments originated by the loss of glutathione (−305 Da) or the neutral loss of glutamic acid (−129 Da). The characteristic fragmentation pattern and expected neutral losses of S-glutathionylated peptides are shown in the supporting information.
Estrogen receptor binding assays
Saturation binding assays of estradiol with ERα were performed using [3H]estradiol (Moraver, Brea, CA). Briefly, recombinant human ERα protein (untreated or treated with disulfiram to induce S-glutathionylation) was incubated with [3H]estradiol (from 0.625 to 80 nm) in assay buffer (100 mm Tris-HCl, 1 mm EDTA, 1 mm EGTA, protease inhibitor mixtures, 10% glycerol, 10 mg/ml BSA, pH 7.4) at room temperature for 90 min; unbound [3H]estradiol was then removed by adding the same amount of 0.4% dextran-coated charcoal (Activated Charcoal Norit®, Sigma) suspension (freshly prepared). After incubating for 10 min on ice, suspensions were centrifuged at 13,000 rpm for 10 min, and supernatants were subjected to liquid scintillation counting. Data were analyzed using a specific binding module from Prism 5.0. Maximum amounts of estradiol bound receptor (Bmax) and binding affinities (Kd) were calculated.
OCR and ECAR analyses
OCR and ECAR analyses were performed using an XF-96 Extracellular Flux Analyzer (Seahorse Bioscience). Briefly, BMDDCs (70,000 cells/well in 70 μl) were seeded in an XF-96 cell culture plate and either left unstimulated or stimulated with 200 ng/ml LPS overnight. Cells were washed and analyzed in XF running buffer according to the manufacturer's instructions to obtain real-time measurement of OCR and ECAR. Where indicated, OCR and ECAR were analyzed in response to 1 μm oligomycin, 1.5 μm carbonyl cyanide p-trifluoromethoxyphenylhydrazone, and 100 nm rotenone plus 1 μm antimycin (all from Sigma).
RNA isolation and RT-PCR
Total RNA was prepared using the Isolate II RNA Mini kit (Bioline, Taunton, MA), and cDNA was then generated with the iScript cDNA synthesis kit (Bio-Rad) according to the manufacturers' protocols. Subsequently, quantification of gene expression was performed in duplicates using iQTM SYBR® Green Supermix (Bio-Rad) with detection on a MyiQTM Real-Time PCR System (Bio-Rad). The reaction cycles used were 95 °C for 5 min and then 40 cycles at 95 °C for 15 s and 58 °C for 1 min followed by melt curve analysis. The following primers were used: Aldoa: forward 5′-TCAGTGCTGGGTATGGGTG-3′, reverse 5′-GCTCCTTAGTCCTTTCGCCT-3′; Aldoc: forward, 5′-GCGGGCAGAGATGAACGGGC-3′; reverse, 5′-AGAGGGACTGTGCTGCCGCT-3′; Eno1: forward, 5′-AAAGATCTCTCTGGCGTGGAC-3′; reverse, 5′-CTTAACGCTCTCCTCGGTGT-3′; Eno3: forward, 5′-GGACTCCAGGGGCAACCCCA-3′; reverse, 5′-TGCTCGGAATCGACCCTTGGC-3′; Esr1: forward, 5′-CCTCCCGCCTTCTACAGGT-3′; reverse, 5′-CACACGGCACAGTAGCGAG-3′; Gapdh: forward,5′-CCCAGCAAGGACACTGAGCAA-3′; reverse, 5′-AGGCCCCTCCTGTTATTATGG-3′; Gck: forward, 5′-GCTCAGTGAACCCCGGTCAGC-3′; reverse, 5′-TGTGCGCAGCTGCTCTGAGG-3′; Gpi: forward, 5′-GCTGGCCAAGTCCAGAGGCG-3′; reverse, 5′-ACCGCCCGATCCTCGGTGTA-3′; Hk2: forward, 5′-GGAACCGCCTAGAAATCTCC-3′; reverse, 5′-GGAGCTCAACCAAAACCAAG-3′; Ldha: forward, 5′-TGTCTCCAGCAAAGACTACTGT-3′; reverse, 5′-GACTGTACTTGACAATGTTGGGA-3′; Ldhb: forward, 5′-CCCAGCAGCTGCCACGGATG-3′; reverse, 5′-CCGCTCCACACAGCCACACT-3′; Ldhc: forward, 5′-TGCGGAGTCAGCAGTAAGGCTC-3′; reverse, 5′-AGCACACGCCATGCCCACAT-3′; Nd4: forward, 5′-ATTATTATTACCCGATGAGGGAACC-3′; reverse, 5′-ATTAAGATGAGGGCAATTAGCAGT-3′; Pfk: forward, 5′-AGGAGGGCAAAGGAGTGTTT-3′; reverse,5′-TTGGCAGAAATCTTGGTTCC-3′; Pgam1: forward, 5′-TCTGCACAGAGGTGAAGCAG-3′; reverse, 5′-GTACGACGCCGACCTGAG-3′; Pgk1: forward, 5′-GGAGGCCCGGCATTCTGCAC-3′; reverse, 5′-AGTCCACCCTCATCACGACCCG-3′; pkm: forward, 5′-GTCTGAATGAAGGCAGTCCC-3′; reverse, 5′-GTCCGCTCTAGGTATCGCAG-3′; Slc2a1: forward, 5′-GGCCGCCTCATGTTGGCTGT-3′, reverse, 5′-TGGGCTCTCCGTAGCGGTGG-3′; Slc2a4: forward, 5′-GTGACTGGAACACTGGTCCTA-3′; reverse, 5′-CCAGCCACGTTGCATTGTAG-3′. A mouse estrogen receptor signaling PrimePCRTM pathway array plate was purchased from Bio-Rad, the genes were listed in supporting Table S1, and expression of genes was evaluated by determining mRNA levels in day 7 BMDDCs. Relative gene expression quantification was based on the comparative threshold cycle (CT) method (2−ΔΔCT) with normalization of the raw data to the included housekeeping gene (Gapdh). PrimePCR gene expression was analyzed using Bio-Rad PrimePCR Analysis software.
Statistical analysis
Student's t tests were used to analyze significant differences. p values <0.05 were regarded as statistically significant. Data were expressed as means ± S.D.
Author contributions
D. M. T., K. D. T., J. Z., Z.-w. Y., and S. M. designed the study. J. Z., Z.-w. Y., W. C., and Y. M. performed experiments and assisted D. M. T., K. D. T., and Y. J.-H. in analyzing data. L. B. performed proteomic analyses. D. M. T., K. D. T., J. Z., Z.-w. Y., and Y. J.-H. wrote the manuscript.
Supplementary Material
Acknowledgment
This work was conducted in a facility constructed with support from National Institutes of Health Grant C06 RR015455 from the Extramural Research Facilities Program of the National Center for Research Resources.
This work was supported in part by National Institutes of Health Grants CA085660 and CA117259; National Center for Research Resources Grant P20RR024485; the Center of Biomedical Research Excellence in Oxidants, Redox Balance, and Stress Signaling, particularly the Analytical Redox Biology Core; and the South Carolina Centers of Excellence program. The authors declare that they have no conflicts of interest with the contents of this article. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
This article contains Figs. S1–S3 and Table S1.
- DC
- dendritic cell
- AGC
- automatic gain control
- BM
- bone marrow
- BMDDC
- bone marrow–derived dendritic cell
- CFSE
- carboxyfluorescein diacetate succinimidyl ester
- CID
- collision-induced dissociation
- ECAR
- extracellular acidification rate
- ER
- estrogen receptor
- ETD
- electron transfer dissociation
- GLUT
- glucose transporter
- GSTP
- glutathione S-transferase Pi
- HSC
- hematopoietic stem cell
- imDC
- immature bone marrow–derived dendritic cell
- mDC
- mature bone marrow–derived dendritic cell
- OCR
- oxygen consumption rate
- OXPHOS
- oxidative phosphorylation
- ROS
- reactive oxygen species
- TCR
- T-cell receptor
- UPR
- unfolded protein response
- MFI
- mean fluorescence intensity
- AMPK
- AMP-activated protein kinase
- APC
- allophycocyanin
- PE
- phycoerythrin
- RNS
- reactive nitrogen species
- DCFH
- dichlorodihydrofluorescin
- TG-1
- ThioGlo-1
- 2-NBDG
- 2-(N-(7-nitrobenz-2-oxa-1,3-diazol-4-yl)amino)-2-deoxyglucose
- NK
- natural killer.
References
- 1. Ballesteros-Tato A., León B., Lund F. E., and Randall T. D. (2010) Temporal changes in dendritic cell subsets, cross-priming and costimulation via CD70 control CD8+ T cell responses to influenza. Nat. Immunol. 11, 216–224 10.1038/ni.1838 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Nakano H., Lin K. L., Yanagita M., Charbonneau C., Cook D. N., Kakiuchi T., and Gunn M. D. (2009) Blood-derived inflammatory dendritic cells in lymph nodes stimulate acute T helper type 1 immune responses. Nat. Immunol. 10, 394–402 10.1038/ni.1707 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Hashimoto D., Miller J., and Merad M. (2011) Dendritic cell and macrophage heterogeneity in vivo. Immunity 35, 323–335 10.1016/j.immuni.2011.09.007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Murata Y., Shimamura T., and Hamuro J. (2002) The polarization of Th1/Th2 balance is dependent on the intracellular thiol redox status of macrophages due to the distinctive cytokine production. Int. Immunol 14, 201–212 10.1093/intimm/14.2.201 [DOI] [PubMed] [Google Scholar]
- 5. Hadzic T., Li L., Cheng N., Walsh S. A., Spitz D. R., and Knudson C. M. (2005) The role of low molecular weight thiols in T lymphocyte proliferation and IL-2 secretion. J. Immunol. 175, 7965–7972 10.4049/jimmunol.175.12.7965 [DOI] [PubMed] [Google Scholar]
- 6. Arai F., Yoshihara H., Hosokawa K., Nakamura Y., Gomei Y., Iwasaki H., and Suda T. (2009) Niche regulation of hematopoietic stem cells in the endosteum. Ann. N.Y. Acad. Sci. 1176, 36–46 10.1111/j.1749-6632.2009.04561.x [DOI] [PubMed] [Google Scholar]
- 7. Calvi L. M., Adams G. B., Weibrecht K. W., Weber J. M., Olson D. P., Knight M. C., Martin R. P., Schipani E., Divieti P., Bringhurst F. R., Milner L. A., Kronenberg H. M., and Scadden D. T. (2003) Osteoblastic cells regulate the haematopoietic stem cell niche. Nature 425, 841–846 10.1038/nature02040 [DOI] [PubMed] [Google Scholar]
- 8. Grek C. L., Townsend D. M., and Tew K. D. (2011) The impact of redox and thiol status on the bone marrow: pharmacological intervention strategies. Pharmacol. Ther. 129, 172–184 10.1016/j.pharmthera.2010.09.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Zhang J., Ye Z. W., Gao P., Reyes L., Jones E. E., Branham-O'Connor M., Blumer J. B., Drake R. R., Manevich Y., Townsend D. M., and Tew K. D. (2014) Glutathione S-transferase P influences redox and migration pathways in bone marrow. PLoS One 9, e107478, 10.1371/journal.pone.0107478 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Eylar E., Rivera-Quinones C., Molina C., Báez I., Molina F., and Mercado C. M. (1993) N-Acetylcysteine enhances T cell functions and T cell growth in culture. Int. Immunol 5, 97–101 10.1093/intimm/5.1.97 [DOI] [PubMed] [Google Scholar]
- 11. Townsend D. M., He L., Hutchens S., Garrett T. E., Pazoles C. J., and Tew K. D. (2008) NOV-002, a glutathione disulfide mimetic, as a modulator of cellular redox balance. Cancer Res. 68, 2870–2877 10.1158/0008-5472.CAN-07-5957 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. List A. F., Brasfield F., Heaton R., Glinsmann-Gibson B., Crook L., Taetle R., and Capizzi R. (1997) Stimulation of hematopoiesis by amifostine in patients with myelodysplastic syndrome. Blood 90, 3364–3369 [PubMed] [Google Scholar]
- 13. Townsend D. M., Manevich Y., He L., Hutchens S., Pazoles C. J., and Tew K. D. (2009) Novel role for glutathione S-transferase Pi. Regulator of protein S-glutathionylation following oxidative and nitrosative stress. J. Biol. Chem. 284, 436–445 10.1074/jbc.M805586200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Ruscoe J. E., Rosario L. A., Wang T., Gaté L., Arifoglu P., Wolf C. R., Henderson C. J., Ronai Z., and Tew K. D. (2001) Pharmacologic or genetic manipulation of glutathione S-transferase P1-1 (GSTpi) influences cell proliferation pathways. J. Pharmacol. Exp. Ther. 298, 339–345 [PubMed] [Google Scholar]
- 15. Raza A., Galili N., Callander N., Ochoa L., Piro L., Emanuel P., Williams S., Burris H. 3rd, Faderl S., Estrov Z., Curtin P., Larson R. A., Keck J. G., Jones M., Meng L., et al. (2009) Phase 1–2a multicenter dose-escalation study of ezatiostat hydrochloride liposomes for injection (Telintra, TLK199), a novel glutathione analog prodrug in patients with myelodysplastic syndrome. J. Hematol. Oncol. 2, 20 10.1186/1756-8722-2-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Lyons R. M., Wilks S. T., Young S., and Brown G. L. (2011) Oral ezatiostat HCl (Telintra(R), TLK199) and idiopathic chronic neutropenia (ICN): a case report of complete response of a patient with G-CSF resistant ICN following treatment with ezatiostat, a glutathione S-transferase P1-1 (GSTP1-1) inhibitor. J. Hematol. Oncol. 4, 43 10.1186/1756-8722-4-43 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Gate L., Majumdar R. S., Lunk A., and Tew K. D. (2004) Increased myeloproliferation in glutathione S-transferase Pi-deficient mice is associated with a deregulation of JNK and Janus kinase/STAT pathways. J. Biol. Chem. 279, 8608–8616 10.1074/jbc.M308613200 [DOI] [PubMed] [Google Scholar]
- 18. Kovats S. (2012) Estrogen receptors regulate an inflammatory pathway of dendritic cell differentiation: mechanisms and implications for immunity. Horm. Behav. 62, 254–262 10.1016/j.yhbeh.2012.04.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Michalek R. D., Gerriets V. A., Nichols A. G., Inoue M., Kazmin D., Chang C. Y., Dwyer M. A., Nelson E. R., Pollizzi K. N., Ilkayeva O., Giguere V., Zuercher W. J., Powell J. D., Shinohara M. L., McDonnell D. P., et al. (2011) Estrogen-related receptor-alpha is a metabolic regulator of effector T-cell activation and differentiation. Proc. Natl. Acad. Sci. U.S.A. 108, 18348–18353 10.1073/pnas.1108856108 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Everts B., Amiel E., van der Windt G. J., Freitas T. C., Chott R., Yarasheski K. E., Pearce E. L., and Pearce E. J. (2012) Commitment to glycolysis sustains survival of NO-producing inflammatory dendritic cells. Blood 120, 1422–1431 10.1182/blood-2012-03-419747 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Everts B., and Pearce E. J. (2014) Metabolic control of dendritic cell activation and function: recent advances and clinical implications. Front. Immunol. 5, 203 10.3389/fimmu.2014.00203 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Pearce E. J., and Everts B. (2015) Dendritic cell metabolism. Nat. Rev. Immunol 15, 18–29 10.1038/nri3771 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Cox J., and Mann M. (2008) MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 10.1038/nbt.1511 [DOI] [PubMed] [Google Scholar]
- 24. Ye Z. W., Zhang J., Ancrum T., Manevich Y., Townsend D. M., and Tew K. D. (2017) Glutathione S-transferase P-mediated protein S-glutathionylation of resident endoplasmic reticulum proteins influences sensitivity to drug-induced unfolded protein response. Antioxid. Redox Signal. 26, 247–261 10.1089/ars.2015.6486 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Sonoda J., Laganière J., Mehl I. R., Barish G. D., Chong L. W., Li X., Scheffler I. E., Mock D. C., Bataille A. R., Robert F., Lee C. H., Giguère V., and Evans R. M. (2007) Nuclear receptor ERRα and coactivator PGC-1β are effectors of IFN-γ-induced host defense. Genes Dev. 21, 1909–1920 10.1101/gad.1553007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Henderson C. J., Smith A. G., Ure J., Brown K., Bacon E. J., and Wolf C. R. (1998) Increased skin tumorigenesis in mice lacking pi class glutathione S-transferases. Proc. Natl. Acad. Sci. U.S.A. 95, 5275–5280 10.1073/pnas.95.9.5275 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Giguère V. (2008) Transcriptional control of energy homeostasis by the estrogen-related receptors. Endocr. Rev. 29, 677–696 10.1210/er.2008-0017 [DOI] [PubMed] [Google Scholar]
- 28. Igarashi H., Kouro T., Yokota T., Comp P. C., and Kincade P. W. (2001) Age and stage dependency of estrogen receptor expression by lymphocyte precursors. Proc. Natl. Acad. Sci. U.S.A. 98, 15131–15136 10.1073/pnas.011513098 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Carreras E., Turner S., Paharkova-Vatchkova V., Mao A., Dascher C., and Kovats S. (2008) Estradiol acts directly on bone marrow myeloid progenitors to differentially regulate GM-CSF or Flt3 ligand-mediated dendritic cell differentiation. J. Immunol. 180, 727–738 10.4049/jimmunol.180.2.727 [DOI] [PubMed] [Google Scholar]
- 30. Hayes J. D., Flanagan J. U., and Jowsey I. R. (2005) Glutathione transferases. Annu. Rev. Pharmacol. Toxicol. 45, 51–88 10.1146/annurev.pharmtox.45.120403.095857 [DOI] [PubMed] [Google Scholar]
- 31. Townsend D. M., Lushchak V. I., and Cooper A. J. (2014) A comparison of reversible versus irreversible protein glutathionylation. Adv. Cancer Res. 122, 177–198 10.1016/B978-0-12-420117-0.00005-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Townsend D. M., Manevich Y., He L., Xiong Y., Bowers R. R. Jr, Hutchens S., and Tew K. D. (2009) Nitrosative stress-induced S-glutathionylation of protein disulfide isomerase leads to activation of the unfolded protein response. Cancer Res. 69, 7626–7634 10.1158/0008-5472.CAN-09-0493 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Aliau S., El Garrouj D., Yasri A., Katzenellenbogen B. S., and Borgna J. L. (1997) 17α-(Haloacetamidoalkyl) estradiols alkylate the human estrogen receptor at cysteine residues 417 and 530. Biochemistry 36, 5861–5867 10.1021/bi963111c [DOI] [PubMed] [Google Scholar]
- 34. Acconcia F., Ascenzi P., Bocedi A., Spisni E., Tomasi V., Trentalance A., Visca P., and Marino M. (2005) Palmitoylation-dependent estrogen receptor alpha membrane localization: regulation by 17β-estradiol. Mol. Biol. Cell 16, 231–237 10.1091/mbc.E04-07-0547 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Marino M., Ascenzi P., and Acconcia F. (2006) S-Palmitoylation modulates estrogen receptor alpha localization and functions. Steroids 71, 298–303 10.1016/j.steroids.2005.09.011 [DOI] [PubMed] [Google Scholar]
- 36. Lin A. H., Li R. W., Ho E. Y., Leung G. P., Leung S. W., Vanhoutte P. M., and Man R. Y. (2013) Differential ligand binding affinities of human estrogen receptor-α isoforms. PLoS One 8, e63199 10.1371/journal.pone.0063199 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Klinge C. M. (2001) Estrogen receptor interaction with estrogen response elements. Nucleic Acids Res. 29, 2905–2919 10.1093/nar/29.14.2905 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Krawczyk C. M., Holowka T., Sun J., Blagih J., Amiel E., DeBerardinis R. J., Cross J. R., Jung E., Thompson C. B., Jones R. G., and Pearce E. J. (2010) Toll-like receptor-induced changes in glycolytic metabolism regulate dendritic cell activation. Blood 115, 4742–4749 10.1182/blood-2009-10-249540 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Martinon F., Chen X., Lee A. H., and Glimcher L. H. (2010) TLR activation of the transcription factor XBP1 regulates innate immune responses in macrophages. Nat. Immunol. 11, 411–418 10.1038/ni.1857 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Goodall J. C., Wu C., Zhang Y., McNeill L., Ellis L., Saudek V., and Gaston J. S. (2010) Endoplasmic reticulum stress-induced transcription factor, CHOP, is crucial for dendritic cell IL-23 expression. Proc. Natl. Acad. Sci. U.S.A. 107, 17698–17703 10.1073/pnas.1011736107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Osorio F., Tavernier S. J., Hoffmann E., Saeys Y., Martens L., Vetters J., Delrue I., De Rycke R., Parthoens E., Pouliot P., Iwawaki T., Janssens S., and Lambrecht B. N. (2014) The unfolded-protein-response sensor IRE-1α regulates the function of CD8α+ dendritic cells. Nat. Immunol. 15, 248–257 10.1038/ni.2808 [DOI] [PubMed] [Google Scholar]
- 42. Iwakoshi N. N., Pypaert M., and Glimcher L. H. (2007) The transcription factor XBP-1 is essential for the development and survival of dendritic cells. J. Exp. Med. 204, 2267–2275 10.1084/jem.20070525 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Manevich Y., Hutchens S., Tew K. D., and Townsend D. M. (2013) Allelic variants of glutathione S-transferase P1-1 differentially mediate the peroxidase function of peroxiredoxin VI and alter membrane lipid peroxidation. Free Radic. Biol. Med. 54, 62–70 10.1016/j.freeradbiomed.2012.10.556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Dasgupta R. K., Adamson P. J., Davies F. E., Rollinson S., Roddam P. L., Ashcroft A. J., Dring A. M., Fenton J. A., Child J. A., Allan J. M., and Morgan G. J. (2003) Polymorphic variation in GSTP1 modulates outcome following therapy for multiple myeloma. Blood 102, 2345–2350 10.1182/blood-2003-02-0444 [DOI] [PubMed] [Google Scholar]
- 45. Otsuki T., Yamada O., Kurebayashi J., Moriya T., Sakaguchi H., Kunisue H., Yata K., Uno M., Yawata Y., and Ueki A. (2000) Estrogen receptors in human myeloma cells. Cancer Res. 60, 1434–1441 [PubMed] [Google Scholar]
- 46. Jang Y. Y., and Sharkis S. J. (2007) A low level of reactive oxygen species selects for primitive hematopoietic stem cells that may reside in the low-oxygenic niche. Blood 110, 3056–3063 10.1182/blood-2007-05-087759 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Trnka M. J., Baker P. R., Robinson P. J., Burlingame A. L., and Chalkley R. J. (2014) Matching cross-linked peptide spectra: only as good as the worse identification. Mol. Cell. Proteomics 13, 420–434 10.1074/mcp.M113.034009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Rahman I., Kode A., and Biswas S. K. (2006) Assay for quantitative determination of glutathione and glutathione disulfide levels using enzymatic recycling method. Nat. Protoc. 1, 3159–3165 10.1038/nprot.2006.378 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.