SUMMARY
Phosphatidylinositol (PtdIns) transfer proteins (PITPs) stimulate PtdIns-4-P synthesis and signaling in eukaryotic cells, but to what biological outcomes such signaling circuits are coupled remains unclear. Herein, we show that two highly related StART-like PITPs, PITPNA and PITPNB, act in a redundant fashion to support development of the embryonic mammalian neocortex. PITPNA/PITPNB do so by driving PtdIns-4-P-dependent recruitment of GOLPH3, and likely ceramide transfer protein (CERT), to Golgi membranes with GOLPH3 recruitment serving to promote MYO18A- and F-actin directed loading of the Golgi network to apical processes of NSCs. We propose the primary role for PITP/PtdIns-4-P/GOLPH3/CERT signaling in NSC Golgi is not in regulating bulk membrane trafficking, but in optimizing apically-directed membrane trafficking and/or apical membrane signaling during neurogenesis.
Keywords: brain development, neural stem cells, phosphatidylinositol transfer proteins, phosphoinositides, cell polarity, Golgi complex
eTOC Paragraph

Xie et al. describe a cell-autonomous phosphatidylinositol-4-phosphate-dependent pathway essential for neocortex development that regulates neural stem cell function by loading the Golgi network into the cellular apical compartment. The circuit involves two lipid transfer proteins (PITPNA/PITPNB) that potentiate phosphatidylinositol-4-phosphate synthesis, and GOLPH3 and CERT as effectors of phosphatidylinositol-4-phosphate signaling.
INTRODUCTION
The mammalian neocortex is the most recently evolved brain region. This six-layered structure houses the machinery that enables superior cognitive capabilities such as highly developed sensory perception and motor control, spatial reasoning, and language (Rakic, 2009). As such, it is a delicate anatomical unit in which even subtle derangements manifest themselves as pathologies that come with staggering social costs – e.g. neuropsychiatric disorders (Gage and Temple, 2013; Sun and Hevner, 2014; Taverna et al., 2014). In mice, the neocortex is built during embryonic stages E11.5 to E17.5, and the major classes of cells that populate the structure during this period of neurogenesis include neural stem cells (NSCs), intermediate progenitor cells (IPCs), and neurons. NSCs are bipolar cells that span the width of the developing neocortex with an apical process contacting the ventricular surface and a basal process contacting the pial surface. NSC somata form a compact pseudostratified layer termed the ventricular zone, these cells directly contact the lateral ventricle of the forebrain via their apical processes, and the ventricular zone represents the deepest cell layer in the neocortex. IPCs are multipolar and lineage-restricted progenitor cells that reside in the subventricular zone which itself overlays the ventricular zone. Most neurons within the neocortex are excitatory projection neurons. Some 90% of those cells are generated from NSCs via intervening IPC lineages. Projection neurons are born near the ventricular zone/subventricular zone and migrate radially to the most superficial layer of the neocortex.
NSCs are sophisticated cells that reside in complex 3D-microenvironments and exhibit complex behaviors in terms of interkinetic nuclear migrations, cell cycle regulation, and asymmetric cell divisions that govern the balance between stem cell renewal and differentiation (Gage and Temple, 2013; Taverna et al., 2014; Bjornsson et al., 2015). The striking apical-basal polarization of NSC architecture fosters unequal partitioning of cellular constituents during mitosis and thereby promotes asymmetric cell divisions. Although multiple aspects of NSC cell polarity establishment/maintenance have been studied (Ghosh et al., 2008; Mora-Bermúdez and Huttner, 2015), the process remains poorly understood. This is particularly the case for how lipid-signaling regulates NSC apical-basal cell polarity.
Phosphatidylinositol (PtdIns)-transfer proteins (PITPs) are evolutionarily conserved proteins that fall into two structural classes – the Sec14-like PITPs and PITPs of the StAR-related lipid transfer (StART) protein superfamily (Schaaf et al., 2008: Yoder et al., 2001). In cells, PITPs regulate the interface between phosphoinositide metabolism/signaling and discrete cohorts of effectors -- thereby diversifying the biological outcomes of phosphoinositide signaling by functionally specifying PtdIns-4-phosphate signaling pools (Bankaitis et al., 2010). Among the demonstrated Sec14-like PITP-dependent functions are establishment and/or maintenance of extreme cell polarity programs (Lopez et al., 1994; Vincent et al., 2005; Huang et al., 2016).
Much less is known regarding physiological functions of StART-like PITPs. Mammalian PITPNA and PITPNB are an enigmatic pair of highly related StART-like PITPs that share 77% primary sequence identity, exhibit phosphatidylinositol (PtdIns) and phosphatidylcholine (PtdCho) binding and transfer activities in vitro (De Vries et al., 1996; Yoder et al., 2001), and stimulate PtdIns-4-P production in vivo (Ile et al., 2010). Herein, we report that these two PITPs act in a redundant fashion to support development of the embryonic mammalian neocortex. PITPNA/PITPNB cooperate in maintaining radial polarity of NSCs via a PtdIns-4-P-dependent mechanism that employs the PtdIns-4-P and MYO-18A-binding protein GOLPH3 as downstream effector in promoting distribution of the NSC Golgi system throughout the apical process. We propose a mechanism where PITPNA/PITPNB drive PtdIns-4-P-dependent recruitment of GOLPH3 to Golgi membranes so as to promote MYO18A- and F-actin directed loading of the Golgi network throughout the NSC apical compartment. We further demonstrate at least one other PtdIns-4-P effector also operates in loading of the Golgi network into the NSC apical process. These collective findings argue that the primary function of NSC Golgi PITP/PtdIns-4-P signaling pathway(s) is to optimize apical targeting of membrane trafficking and/or apical membrane signaling during neurogenesis.
RESULTS
Murine PITPNA and PITPNB exhibit functional redundancy in embryonic development
The 77% primary sequence identity shared by mammalian PITPNA and PITPNB notwithstanding, it remains unclear as to how these proteins are functionally diversified in mammals, or whether these proteins share functional redundancy. At the least, PITPNA plays a critical role in mice independent of PITPNB activity. Pitpna null mice are born alive but die within the first few days of postnatal life from multiple systemic failures (Alb et al., 2003). Moreover, a hypomorphic Pitpna allele that reduces wild-type protein expression by ~5-fold is the genetic basis for the vibrator (vb) neurodegenerative mouse (Hamilton et al., 1997). Reliable information regarding Pitpnb null phenotypes in the mouse is lacking. Early unsuccessful attempts to generate Pitpnb null mice, or null embryonic stem cells, prompted the suggestion that PITPNB might execute an essential function in mammals (Alb et al., 2002).
To clarify the matter, TALEN mutagenesis was used to produce Pitpnb mice homozygous for a frameshift mutation that disqualifies production of PITPNB in all tissues (Suppl. Figure S1A). Like Pitpna null mice, the Pitpnb null progeny were born alive. However, unlike Pitpna null mice, Pitpnb null animals have yet to manifest any obvious pathologies at 12 months of age, and Pitpnb null mice are fertile. To address potential issues of functional redundancy, standard breeding strategies were followed to generate offspring where both PITPNA and PITPNB were functionally ablated in all tissues. Of the 60 P0 neonates and 69 E12.5 embryos analyzed from intercross of Pitpna+/− Pitpnb+/− parents, none exhibited Pitpna−/− Pitpnb−/− or Pitpna+/− Pitpnb−/− genotypes. This non-Mendelian ratio indicated whole body PITPNA/PITPNB deficiencies were nonpermissive for early embryonic development, and that a single functional Pitpna allele was insufficient to rescue developmental failure. Characterization of the critical failure point for the Pitpna/Pitpnb double null mice will be presented elsewhere.
PITPNA and PITPNB are expressed in embryonic mouse neocortical cells
Immunostaining analyses reported robust expression of both PITPNA and PITPNB in developing mouse neocortex at E12.5, and cytoplasmic immunoreactivity for both PITPNA and PITPNB was obvious in all cells of the neocortex (Figures 1A,B). Specificity of staining was confirmed by its strongly diminished intensity in corresponding brain regions of littermate Pitpna or Pitpnb null embryos (Figures 1A,B), and by the cytoplasmic distributions of EGFP-tagged PITPNA and PITPNB ectopically expressed in neocortical cells (Figures 1C,D). As neocortical cells at stage E12.5 are comprised of NSCs, IPCs, and newly generated deep-layer projection neurons (Suppl. Figure S1B; Greig et al., 2013), these data suggested that PITPNA and PITPNB are expressed in both NSCs and their more differentiated progenies. Moreover, these data identified embryonic neocortex as a tissue suitable for dissection of both unique and functionally redundant roles for PITPNA and PITPNB in brain development.
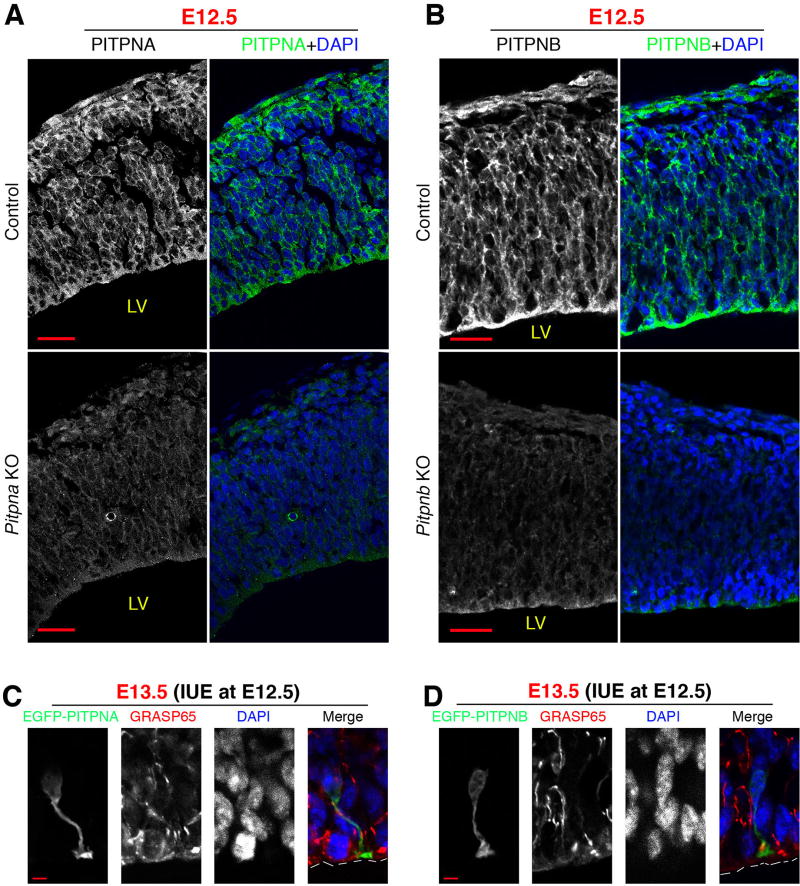
Figure 1. Expression and localization of PITPNA and PITPNB in the neocortex of mouse embryos.
(A and B) PITPNA and PITPNB immunoreactivity in the cytoplasm of neocortical cells of E12.5 mouse embryos. Immunoreactivity is diminished in the neocortex of knockout littermate embryos, demonstrating specificity. (C and D) Localization of EGFP-PITPNA and EGFP-PITPNB in NSCs in the neocortex of mouse embryos. Plasmids for expressing EGFP-PITPNA or EGFP-PITPNB were electroporated into the neocortex of E12.5 mouse embryos, and immunofluorescence analysis was performed at E13.5. GRASP65 immunostaining marked the Golgi apparatus. Both EGFP-PITPNA and EGFP-PITPNB were localized throughout the cytoplasmic compartment. LV: lateral ventricle. Scale bars: 20µm in (A) and (B), 5µm in (C) and (D). See also Suppl. Figure S1.
Pitpna and Pitpnb are required for dorsal forebrain development
To investigate functional roles for PITPNA/PITPNB in developing embryonic neocortex, mouse lines carrying floxed Pitpna and Pitpnb alleles were created (Suppl. Figures S1C,D). Those animals were subsequently crossed to an Emx1Cre knock-in mouse line in which Emx1Cre prosecutes efficient deletion of floxed genes specifically in the dorsal forebrain, a structure largely comprised by the neocortex, by E12.5 (Gorski et al., 2002; Liang et al., 2012; Insolera et al. 2014). Tissue-specific double knockout (DKO) progeny characterized by Pitpnafl/fl Pitpnbfl/fl Emx1Cre/+ or Cre/Cre genotypes (designated as DKO in figures) were born alive and did not display overt phenotypes at birth (Figure 2A). However, DKO mice were devoid of a dorsal forebrain (Figure 2B). By contrast, progeny of all other genotypes (designated as Control in figures), including those carrying only a single copy of either Pitpna or Pitpnb (i.e. Pitpnafl/+ Pitpnbfl/fl Emx1Cre/+ or Cre/Cre or Pitpnafl/fl Pitpnbfl/+ Emx1Cre/+ or Cre/Cre), exhibited anatomically normal brains at this level of resolution. These data demonstrated that: (i) PITPNA and PITPNB execute a redundant function(s) essential for development of a dorsal forebrain in mammals, and (ii) that a single copy of Pitpna or Pitpnb was sufficient to restore development of a grossly normal brain.
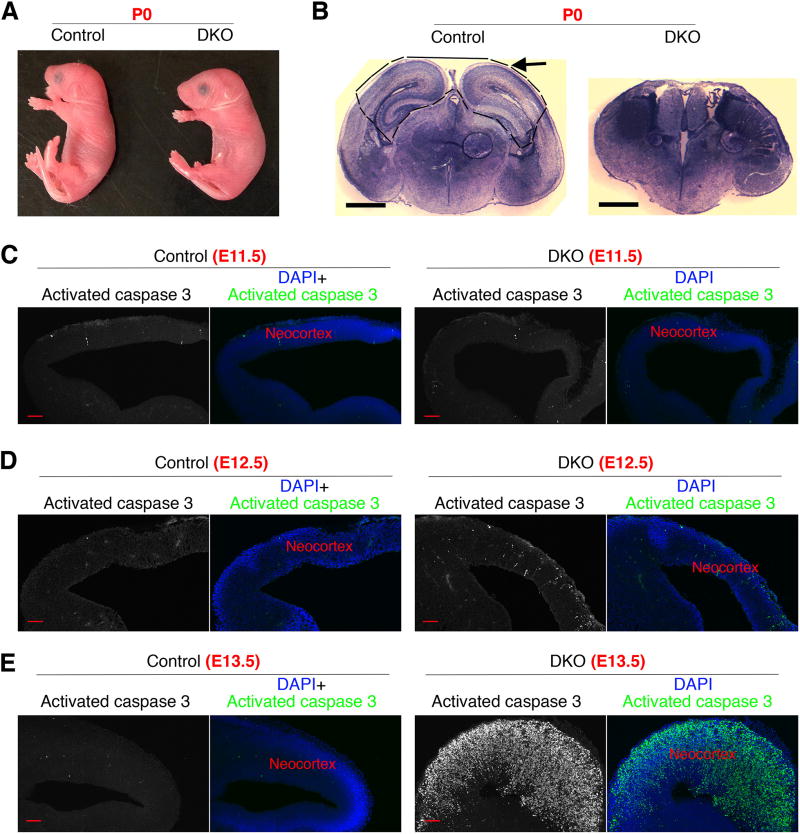
Figure 2. Simultaneous loss of PITPNA and PITPNB in the neocortex leads to absence of the dorsal forebrain at birth and pronounced apoptosis during embryonic stages.
(A) Mouse pups with Emxcre-mediated deletion of Pitpna and Pitpnb did not show overt phenotypes compared to littermates. The genotypes were Pitpnafl/+ Pitpnbfl/fl EmxCre/+ for control and Pitpnafl/fl Pitpnbfl/fl EmxCre/+ for double knockout (DKO). (B) The dorsal forebrain is absent in mouse pups with Emxcre-mediated deletion of Pitpna and Pitpnb. Dashed lines outline control dorsal forebrain structure that was absent in DKO pups. No anatomical abnormalities in the dorsal forebrain were detected for any of the other genotypes. (C–E) Rapid onset of apoptosis in PITPNA/PITPNB-depleted neocortex. The extent of apoptosis in the neocortex of DKO mouse embryos was normal at E11.5, moderately increased at E12.5, and drastically elevated at E13.5. The medial neocortex of DKO embryos was severely disorganized at E13.5. Scale bars: 1mm in (B), 50µm in (C–E). See also Suppl. Figure S2.
PITP-deficiencies and catastrophic derangements of embryonic neocortex
To examine the course of developmental failure in PITPNA/PITPNB-deficient dorsal forebrain in detail, the neocortex was visualized at various embryonic stages. At E16.5, the dorsal forebrain was already absent as an anatomical structure (Suppl. Figure S2A). At earlier stages from E11.5 to E14.5, however, the dorsal forebrain was recognizable (Suppl. Figures S2B–E). Nonetheless, gross disorganization of the structure was scored by E13.5 and became more pronounced at E14.5, when medial regions of the structure were completely missing (Suppl. Figures S2B,C). While apoptosis was not evident in the dorsal forebrain of DKO embryos at E11.5 (Figure 2C), a moderate increase was observed at E12.5 (Figure 2D), and apoptosis was evident over the entire dorsal forebrain at E13.5 and E14.5 (Figures 2E; Suppl. Figure S2F). No such apoptosis was recorded in developing neocortices of embryos of any other genotype.
By E14.5, the signature layered organization of the neocortex was grossly disorganized in the face of PITPNA/PITPNB-deficiency. In control embryos, distinct cell layers were on clear display in developing neocortex (Suppl. Figure S2G). Those layers included the ventricular zone (marked by Pax6), the subventricular zone (marked by Tbr2), the intermediate zone (i.e. the layer above the subventricular zone with relatively low cell density), and the cortical plate (upper neocortical layer with relatively high cell density). In matched PITPNA/PITPNB-deficient embryos, however, those layers were completely deranged (Suppl. Figure S2G). While PITPNA/PITPNB-deficient cells in deep neocortical layers expressed the NSC marker Pax6, as in control embryos, most of these cells also co-expressed the IPC marker Tbr2 (Suppl. Figure S2G) -- indicating abnormalities in the NSC differentiation program. The Pax6 and Tbr2 layers were partially preserved at E13.5 where widespread onset of apoptosis was observed (Figure 2E; Suppl. Figure S2H).
Deranged radial alignments of NSCs and their nuclei
To gain further insight into the base causes of neocortical deficits in PITPNA/PITPNB-deleted embryonic neocortex, the tissue was analyzed at developmental stages preceding onset of rampant apoptosis. At E12.5, close examination of the nuclear morphology and nestin immunostaining revealed loss of radial alignment of NSCs in the neocortex of double knockout embryos. At this stage, the vast majority of cells within the neocortex were Pax6+Tbr2− NSCs (Suppl. Figure S1B), which are comprised of two types of radially aligned cells, i.e. radial glial cells and neuroepithelial cells. Consistent with this, most nuclei in control neocortices were elongated, with the length axis showing a clear radial orientation (Figure 3A). Moreover, nestin immunoreactivity, which labeled the apical and basal processes of NSCs, also displayed a general radial orientation. In DKO neocortex, most nuclei were rounded and failed to exhibit discernible radial orientation. Moreover, nestin fibers in DKO neocortex were badly disorganized, and their radial alignment was dramatically perturbed (Figure 3A). Importantly, radial alignment defects were obvious in all DKO neocortical regions at E12.5, and were on prominent display in regions of developing neocortex that showed no apoptosis at all (Figure 3A). Thus, loss of NSC radial alignment preceded engagement of apoptosis.
Figure 3. Disrupted radial alignment of NSCs and altered Golgi localization are early defects in the neocortex of double knockout embryos.
(A) DAPI and nestin staining revealed disrupted radial alignment of NSCs in the neocortex of E12.5 double knockout (DKO) mouse embryos. Most nuclei in control neocortex were elongated with the length axis radially aligned. Moreover, nestin fibers in control neocortex showed a radial orientation. Both the radial orientation of nuclei and that of nestin fibers were lost throughout double knockout neocortex, including regions with no or limited apoptosis. (B) Immunoreactivity of GRASP65 revealed altered Golgi positioning in the neocortex of DKO embryos. In control neocortex, the Golgi apparatus was distributed as radially aligned short segments in the ventricular zone, reflecting localization of Golgi apparatus within the apical process of NSCs. In DKO neocortex, however, the pattern of Golgi staining was predominantly punctate and in a perinuclear location. (C) Electron microscopy analysis revealed no defects in general organization of Golgi apparatus in the neocortex of E12.5 DKO mouse embryos. (D–F) Loss of radial alignment of NSCs and altered Golgi localization were visualized in an IUE experiment. Plasmids for expressing mCherry and EGFP-GM130 were electroporated into the neocortex of E11.5 mouse embryos, and transfected cells were analyzed at E12.5 by confocal microscopy. (D) Representative confocal images. Arrow highlights an NSC that shows perinuclear Golgi distribution and retains apical contact with the ventricular surface (outlined by dashed lines). (E) Quantification of Golgi-nucleus distance in transfected cells. Distance between the center of EGFP-GM130 puncta/segments and the proximal edge of the nucleus was measured. If multiple EGFP-GM130 puncta/segments were present in a single NSC, average distance for these puncta/segments was used to represent the data point for that NSC. (F) Quantification of the length/width ratio of transected cells. Confocal images were used to measure the ratio of the total length of the soma and apical process to the width of the soma. A decrease in this ratio should reflect impaired radial alignment of NSCs. This ratio took into account two features characteristic of PITPNA/PITPNB-deficient NSCs: a rounded nucleus, which corresponds to increased width of soma, and a shorter apical process. ** p<0.01, one-way ANOVA. Scale bars: 20µm in (A), 10µm in (B) and (D), and 0.5µm in (C). See also Suppl. Figure S3.
Apical-to-perinuclear Golgi redistribution is an early event in loss of NSC polarity
NSC radial alignment defects were coincident with marked redistribution of the Golgi network in these cells. Whereas the Golgi system was arranged as perinuclear puncta in upper neocortical layers, the organelle adopted short, radially-aligned profiles distributed throughout the apical prcesses of NSCs in deep neocortical layers adjacent to the lateral ventricle – i.e. where NSCs reside (Figures 3B; Suppl. Figure S3A). These apically-distributed profiles were consistent with the observations of Taverna et al. (2016) who documented restriction of NSC Golgi distribution to the radially aligned apical processes. In E12.5 DKO neocortex, however, while the pattern of Golgi staining in superficial layers was similar to that of controls, the short, radially-aligned and apically distributed Golgi profiles in deep neocortical layers were strongly diminished in favor of perinuclear punctate patterns (Figure 3B; Suppl. Figure S3A). These Golgi puncta were juxtaposed with the apical aspect of the NSC nuclear envelope and were not distributed throughout the apical compartment. Redistribution of the Golgi network was most pronounced in regions with more severe defects in NSC radial alignment. Moreover, Golgi redistribution was observed in all regions of E12.5 neocortex -- including regions devoid of apoptotic foci. Both radial alignment and Golgi distribution profiles were similar in DKO and control embryos at E11.5 (Suppl. Figure S3B), indicating defects in those processes coincided with expression of the Emx1Cre driver in the E11.5 – E12.5 window.
Coincident loss of NSC radial alignment with Golgi redistribution from apical processes to perinuclear locations were the earliest detectable derangements recorded in PITPNA/PITPNB-deficient NSCs. Moreover, these defects did not appear to be accompanied by impaired Golgi secretory function. High resolution imaging of PITPNA/PITPNB-depleted NSC Golgi apparati by electron microscopy did not report unusual accumulations of secretory vesicles, or any other obvious morphological distentions of Golgi cisternae that typically accompany strong membrane trafficking defects (Figure 3C). Other behaviors of PITPNA/PITPNB-depleted NSCs were also consistent with robust bulk membrane trafficking capability (see below).
Second, these defects were manifested in the face of normal proliferation and normal distribution of other polarity markers. For instance, Ser10-phosphorylated histone H3 immunostaining demonstrated that mitotic NSCs were detected at normal frequencies in E12.5 DKO neocortex (fraction of nuclei that are at M-phase: 4.16 ± 0.14%, mean ± SD, n = 3 embryos, for control, and 4.17 ± 0.38%, mean ± SD, n = 3 embryos, for DKO; p = 0.96, Student’s t-test) and normal locations (fraction of mitotic nuclei that are at the ventricular surface: 74.92 ± 7.39%, mean ± SD, n = 3 embryos, for control, and 72.56 ± 3.02%, mean ± SD, n = 3 embryos, for DKO; p = 0.64, Student’s t-test; Suppl. Figure S3C). Pax6 and Tbr2 expression also maintained an appropriate anatomical segregation in PITPNA/PITPNB-deficient neocortex, suggesting that the NSC-IPC differentiation program was not strongly deranged at this stage (Suppl. Figure S3D). Most strikingly, both the PAR3 core polarity protein and the centrosomal marker pericentrin were correctly localized to the NSC apical surface (i.e. where NSCs interface with ventricular fluid), and the F-actin cytoskeleton was also appropriately organized at the ventricular surfaces of PITPNA/PITPNB-depleted NSCs (Suppl. Figure S3E). Those localization profiles reported that adherence junctions were unaffected at stage E12.5.
Apical-to-perinuclear shift in Golgi distribution is independent of loss of apical contact with the ventricular surface
One potential stimulus for the apical-to-perinuclear rearrangement of Golgi distribution in PITP-less NSCs was loss of apical contact with the ventricular surface. To examine this possibility directly, in utero electroporation (IUE) experiments were performed to visualize Golgi distribution in individual transfected NSCs in DKO neocortex. In those experiments, control or DKO mouse neocortex was electroporated with an mCherry expression plasmid to label all subcellular compartments of transfected cells, and an EGFP-GM130 expression plasmid to specifically label the Golgi network (Suppl. Figure S3F). The IUE approach was superior to standard staining approaches (e.g. nestin or F-actin labeling) in identifying individual NSCs and their Golgi bodies in the ventricular zone (Figure 3A, Suppl. Figure S3G). Electroporations were performed at E11–E11.5 -- i.e. stages at which PITPNA/PITPNB-depleted neocortex exhibited no obvious cellular defects. The embryos were subsequently harvested for analysis 20–24h after electroporation.
As expected, EGFP-GM130 profiles in PITPNA/PITPNB-deficient NSCs that maintained apical contact adopted prominent perinuclear punctate profiles at markedly increased frequencies relative to control transfected NSCs (Figure 3D). The Golgi-nucleus distance was significantly shortened in transfected PITPNA/PITPNB-deficient NSCs (Figure 3E), and the radial alignment of PITPNA/PITPNB-deficient NSCs was obviously disturbed. To quantify the radial alignment defects, the length-to-width ratios of NSC soma and apical processes were also measured (total length of soma and apical process divided by width of soma). A lower ratio reported a rounded nucleus and a shorter apical process, and reflected impaired radial alignment of NSCs. Indeed, this ratio was significantly decreased in transfected PITPNA/PITPNB-deficient NSCs (Figure 3F). Thus, NSC Golgi positioning defects and impaired radial alignment did not result from loss of apical contact with the ventricular surface.
PITP-mediated control of Golgi distribution is a cell autonomous function
The developing neocortex presents a highly complex 3-dimensional neurogenic niche to cells that populate it (Bjornsson et al, 2015). We therefore assessed whether PITPNA/PITPNB controlled NSC Golgi distribution and radial alignment in a cell autonomous fashion. To that end, IUE experiments were performed to evict Pitpna and Pitpnb from individual NSCs while preserving the complex neocortical environment in which such cells would normally reside. Those experiments took advantage of a chicken beta-actin (CAG) promoter-based expression plasmid that supported robust Cre expression in the neocortex. Thus, E12.5 Pitpnafl/fl Pitpnbfl/fl embryos were electroporated with mCherry, EGFP-GM130, and Cre expression plasmids, returned to the mother for an additional 90h incubation, harvested, and subsequently analyzed by immunostaining and confocal imaging.
Both the Golgi positioning and radial alignment defects observed in Emx1Cre-mediated PITPNA/PITPNB gene eviction embryos were recapitulated in individual NSCs for which those activities were functionally ablated. First, EGFP-GM130 localized predominantly to perinuclear puncta in transfected Pitpnafl/fl Pitpnbfl/fl NSCs as compared to control NSCs carrying at least one non-floxed allele of either Pitpna or Pitpnb where the reporter adopted short, radially-aligned profiles that were distributed throughout the apical compartment of these cells (Figures 4A,B). Moreover, radial alignment was compromised in electroporated Pitpnafl/fl Pitpnbfl/fl NSCs (Figures 4A,C). Those data established that PITPNA/PITPNB promote NSC radial alignment and loading of the Golgi network throughout the NSC apical process in a cell autonomous manner.
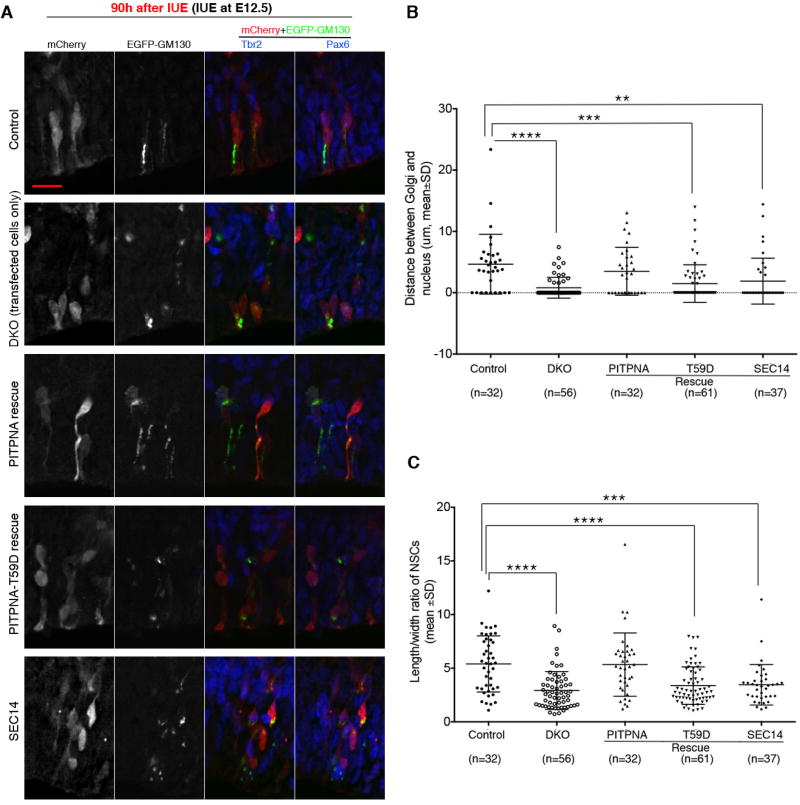
Figure 4. PITPNA/PITPNB regulate NSC radial alignment and Golgi position in a cell autonomous manner.
Mouse embryos (genotype: Pitpnafl/fl Pitpnbfl/fl for DKO and Pitpnafl/+ Pitpnbfl/fl for control) were electroporated with plasmids for expressing mCherry, EGFP-GM130, and Cre at E12.5, and sacrificed 90h after electroporation. In Pitpnafl/fl Pitpnbfl/fl embryos (DKO), transfected cells expressed Cre (from the Cre plasmid) and were therefore PITPNA/PITPNB-deficient, whereas non-transfected cells did not express Cre and therefore retained the expression of PITPNA/PITPNB. In rescue experiments, a plasmid for expressing wild-type PITPNA, PITPNAT59D (PtdIns-binding mutant), or Sec14 (a yeast PITP) was added to the plasmid mixture and Pitpnafl/fl Pitpnbfl/fl embryos were used. Confocal z-series images were used for analysis of NSC morphology and Golgi localization. Representative images are shown in (A). Pax6 and Tbr2 immunofluorescence (Cy5 and Dylight 405, respectively, both pseudocolored to blue) was used to determine NSC identity (Pax6+Tbr2−). NSCs from three or more embryos in each experimental group were pooled together for analysis. Note that not all cells co-expressed mCherry and EGFP-GM130, at least partially because of the detection limit of our methods. Quantifications of Golgi-nucleus distance and length/width ratio are shown in (B) and (C), respectively. **P<0.01, ***P<0.001, **** P<0.0001 compared to control, one-way ANOVA. Scale bars: 10µm. See also Suppl. Figure S4.
Two behaviors of the electroporated PITPNA/PITPNB-deficient NSCs were of particular note. First, transfected cells in electroporated Pitpnafl/fl Pitpnbfl/fl embryos did not show increased propensities to apoptose relative to control NSCs at different time points following electroporation (Suppl. Figure S4A–S4F). Those data identified the radial polarity and Golgi positioning derangements as primary cell-autonomous phenotypes that were not coupled to apoptosis. Second, PITPNA/PITPNB-deficient NSCs produced neurons that efficiently migrated from their birthplace (subventricular zone) to the most superficial layers of developing neocortex. That is, directional migration over distances of hundreds of micrometers (800–1000 µm in some cortical areas) was not at all affected by PITP-deficiencies (data not shown).
PITPNA/PITPNB specify a PtdIns-4-P pool that controls NSC radial alignment and Golgi distribution
PITPNA and PITPNB stimulate PtdIns-4-P synthesis in cells (Ile et al., 2010). Given PtdIns-4-P is a functionally important phosphoinositide of late Golgi membranes (Bard and Malhotra, 2006; Liu et al., 2008; Glick and Nakano, 2009; Graham and Burd, 2011), we first adopted standard PtdIns-4-P fluorescent biosensor approaches to address whether PITPNA/PITPNB deficiencies evoked Golgi PtdIns-4-P deficits. Unexpectedly, we found that biosensors routinely used to image late Golgi PtdIns-4-P pools (e.g. FAPP1-GFP and SidM-GFP) exhibited cytoplasmic fluorescence profiles when expressed in WT NSCs (data not shown). The failure of these biosensors to target to NSC Golgi membranes indicated that PtdIns-4-P pools are either less accessible in NSC Golgi than in the Golgi systems of cultured cells, or that NSC Golgi present relatively low levels of PtdIns-4-P in vivo.
Thus, two independent approaches were employed to determine whether PITPNA/PITPNB operated via a PtdIns-4-P-dependent pathway to regulate NSC radial alignment and apical Golgi distribution. In the first assay, an IUE approach was deployed with the strategy of interfering with PtdIns-4-P signaling in embryonic NSCs by ectopically expressing the SACM1L PtdIns-4-P phosphatase to enhance degradation of this phosphoinositide. In those experiments, mouse neocortex was electroporated with expression plasmids for mCherry, EGFP-GM130, and either the wild-type SACM1L or the phosphatase-deficient mutant SACM1LD391N. For embryos carrying two alleles each of wild-type Pitpna and Pitpnb, ectopic SACM1L expression had no effect on either Golgi localization or NSC radial alignment (Suppl. Figure S4G). However, in sensitized Pitpnafl/+ Pitpnbfl/fl EmxCre/+ embryos, where PITP function was reduced to the threshold of sufficiency for normal NSC behavior, elevated SACM1L expression not only induced redistribution of the Golgi apparatus from the apical process to perinuclear locations, but also disrupted the radial alignment of NSCs (Figures S4H–S4J). Neither of those defects was observed upon ectopic expression of the catalytic-dead SACM1LD391N (Figures S4H–S4J).
To independently interrogate the role of PtdIns-4-P in PITP-dependent regulation of Golgi positioning and radial alignment, we tested whether the PITPNAT59D mutant was able to correct these phenotypes in PITPNA/PITPNB-depleted NSCs. PITPNAT59D is specifically unable to bind PtdIns, cannot simulate PtdIns-4-P synthesis, and scores as a functional null when reconstituted as sole source of PITPNA in the mouse (Alb et al., 2007). Thus, E12.5 Pitpnafl/fl Pitpnbfl/fl embryonic neocortex was electroporated with mCherry, EGFP-GM130 and Cre expression plasmids, and a rescue plasmid for expressing either wild-type PITPNA or PITPNAT59D. Embryos were returned to the mother for an additional 90h incubation, harvested, and analyzed. Whereas ectopic re-expression of wild-type PITPNA fully rescued both Golgi positioning and NSC radial alignment defects in individual PITPNA/PITPNB-depleted NSCs, PITPNAT59D expression failed to do so (Figure 4). These collective SAC1ML and PITPNAT59D data supported a mechanism where PITPNA/PITPNB regulate NSC Golgi distribution and radial alignment in a PtdIns-4-P-dependent manner.
Lipid transfer activity is insufficient for NSC Golgi positioning and radial alignment
Sec14 is the major PITP of yeast that controls PtdIns-4-P signaling in the yeast Golgi/endosomal system (Schaaf et al., 2008; Bankaitis et al., 2010). While structurally unrelated to PITPNA/PITPNB, Sec14 is very similar to these mammalian PITPs in its PtdIns/PtdCho-binding specificities and differential PtdIns/PtdCho-binding affinities. Interestingly, ectopic expression of Sec14 utterly failed to rescue apical Golgi distribution and radial alignment in NSCs depleted for PITPNA/PITPNB activity (Figure 4). High level Sec14 expression was no more effective in rescue than was expression of PITPNA/PITPNB functional nulls. Thus, PITP-lipid transfer/exchange activities are grossly insufficient for rescue of NSC apical Golgi positioning and radial alignment defects. The significance of this result is further discussed below.
GOLPH3 recruitment to NSC Golgi is PITP- and PtdIns-4-P-dependent
GOLPH3 is a peripheral Golgi PtdIns-4-P binding protein that controls Golgi shape in cultured mammalian cells in a manner that involves its direct physical interaction with the unconventional F-actin myosin MYO-18A (Dippold et al., 2009; Taft et al., 2013; Xing et al., 2016). Western blot and immunostaining experiments demonstrated GOLPH3 to be expressed in the neocortex of mouse embryos, and to exhibit specific Golgi localization in essentially all neocortical cells -- including NSCs (Suppl. Figure S5A,B). Interestingly, the GOLPH3 immunoreactivity profile was dramatically altered in PITPNA/PITPNB-depleted neocortex (Figure 5A). At E12.5, while GOLPH3 labeled the short, radially-aligned and apically-distributed Golgi structures in control embryonic neocortex, GOLPH3 immunoreactivity was diffusely distributed throughout the cytoplasm (i.e. as a thin layer surrounding the nucleus) in Pitpnafl/fl Pitpnbfl/fl Emx1Cre/+ DKO neocortical cells (Figure 5A). This delocalization was obvious in the face of unperturbed localization of other Golgi proteins in DKO neocortex (e.g. GRASP65, GM130, GOLGA1; Figure 3B, Suppl. Figures S5C, S5D).
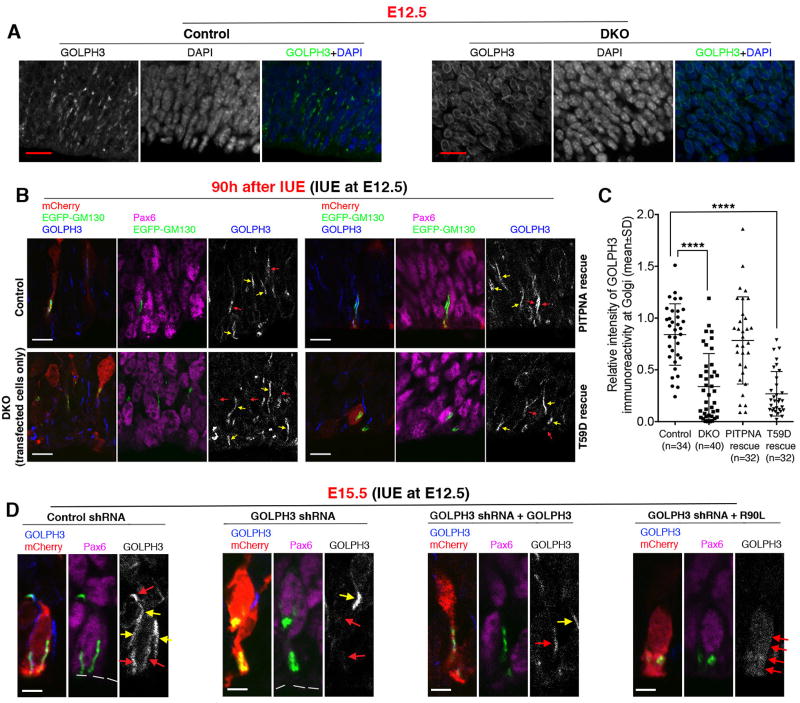
Figure 5. Golgi localization of GOLPH3 is PITP- and PtdIns-4-P-dependent.
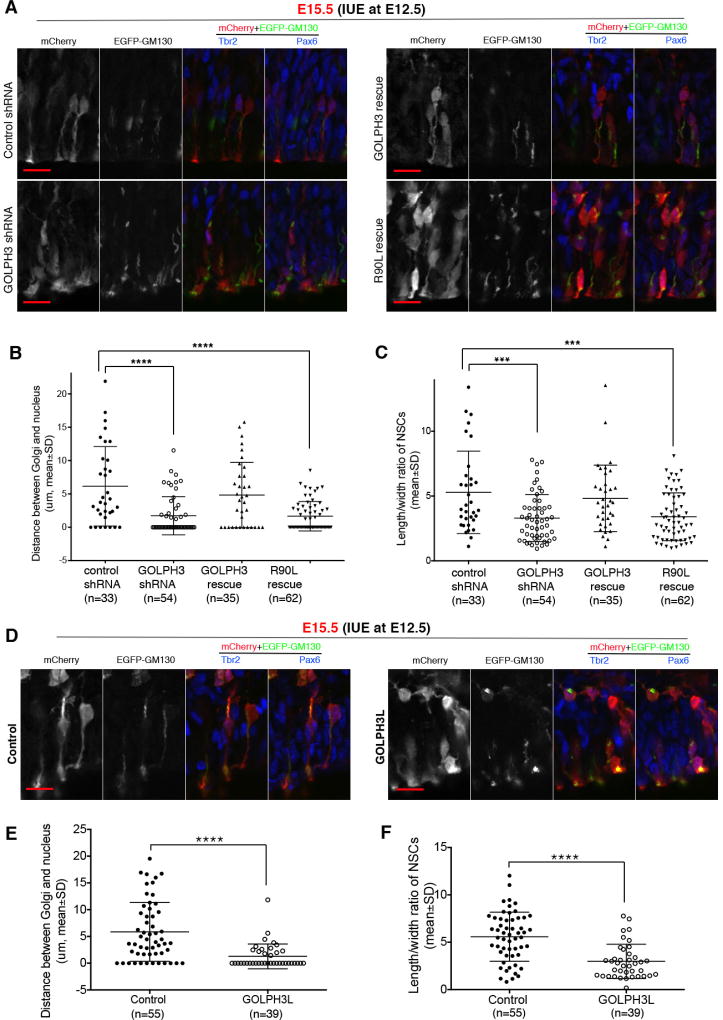
(A) GOLPH3 is mislocalized from the Golgi complex in the neocortex of E12.5 double knockout (DKO) mouse embryos. In the ventricular zone of the neocortex of control embryos, GOLPH3 staining showed typical labeling of radially aligned short segments, which were apically localized Golgi segments. In DKO embryos, however, GOLPH3 staining showed predominantly diffuse cytoplasmic labeling, which typically appears as a thin layer surrounding the nucleus in the neocortex. (B and C) Electroporation-mediated deletion of Pitpna and Pitpnb results in GOLPH3 mislocalization from the NSC Golgi complex. Mouse embryos (genotype: Pitpnafl/fl Pitpnbfl/fl for DKO and Pitpnafl/+ Pitpnbfl/fl for control) were electroporated with plasmids for expressing mCherry, EGFP-GM130, and Cre at E12.5, and sacrificed 90h after electroporation. In the DKO group, transfected cells expressed Cre (from the plasmid) and therefore were PITPNA/PITPNB-deficient, whereas non-transfected cells did not express Cre and therefore retained the expression of PITPNA/PITPNB. In rescue experiments, a plasmid for expressing wild-type PITPNA or the PITPNAT59D PtdIns-binding mutant was added to the plasmid mixture and Pitpnafl/fl Pitpnbfl/fl embryos were used. Representative images are shown in (B), and quantifications are shown in (C). The ratio of the intensity of GOLPH3 labeling at Golgi (indicated by EGFP-GM130 immunofluorescence) in transfected NSCs to that in bystander non-transfected NSCs was quantified as relative intensity of GOLPH3 immunoreactivity at Golgi. The relative intensity was significantly reduced in DKO NSCs (i.e. NSCs in Pitpnafl/fl Pitpnbfl/fl embryos electroporated with Cre plasmid) compared to control (i.e. NSCs in Pitpnafl/+ Pitpnbfl/fl embryos electroporated with Cre plasmid). Red and yellow arrows indicate GOLPH3 staining at Golgi in transfected and non-transfected bystander NSCs, respectively. **** P<0.0001 compared to control, one-way ANOVA. (D) PtdIns-4-P-binding activity of GOLPH3 is required for Golgi-specific localization of GOLPH3. Plasmids for expressing mCherry and EGFP-GM130 were co-electroporated with control shRNA or GOLPH3 shRNA into the neocortex of wild-type C57BL6 mouse embryos at E12.5, and electroporated embryos were harvested 72h later. In rescue experiments, a plasmid for expressing shRNA resistant wild-type GOLPH3 or GOLPH3R90L was added to the plasmid mixture for electroporation. GOLPH3 shRNA diminished Golgi-specific immunoreactivity of endogenous GOLPH3 in transfected cells. Thus, in the rescue group, GOLPH3 immunoreactivity reflected the localization of exogenous GOLPH3 proteins (wild-type or the R90L mutant expressed from the plasmid). Red and yellow arrows indicate GOLPH3 staining in transfected and non-transfected NSCs, respectively. Scale bars: 10µm in (A) and (B), 2µm in (D). See also Suppl. Figure S5.
IUE analyses demonstrated that failure to recruit GOLPH3 to PITP-depleted NSC Golgi bodies was a cell-autonomous defect. When Pitpnafl/fl Pitpnbfl/fl NSCs were electroporated with pCAG-Cre to evict these floxed alleles, GOLPH3 targeting to Golgi membranes was abolished in transfected cells (i.e. NSCs in which Pitpna and Pitpnb were deleted by Cre recombinase expressed from the plasmid). GOLPH3 localization to the Golgi system remained unperturbed in neighboring non-transfected NSCs (i.e. cells in which Pitpna and Pitpnb were not deleted; Figures 5B,C). Appropriate localization of GOLPH3 to the Golgi network was restored in PITP-depleted NSCs by ectopic re-expression of wild-type PITPNA, but not by PITPNAT59D expression (Figures 5B,C) -- indicating Golgi targeting of GOLPH3 was dependent on the ability of PITPNA/PITPNB to potentiate PtdIns-4-P synthesis.
To confirm that Golgi targeting of GOLPH3 in NSCs was PtdIns-4-P-dependent, plasmids for expressing mCherry, EGFP-GM130, control or GOLPH3 shRNA, and silencing-resistant rescue plasmids for expressing wild-type GOLPH3 or GOLPH3R90L (a PtdIns-4-P-binding deficient mutant) were co-electroporated into the neocortex of mouse embryos at E12.5. The electroporated embryos were subsequently harvested after 72h and processed for GOLPH3-immunostaining. As evidence that endogenous GOLPH3 expression was suitably silenced by GOLPH3 shRNA, the Golgi-specific GOLPH3 signal evident in control shRNA-transfected NSCs was ablated in GOLPH3 shRNA-transfected NSCs (Figure 5D). When NSCs were co-transfected with GOLPH3 shRNA and wild-type GOLPH3 rescue plasmids, Golgi-localized GOLPH3 immunoreactivity was restored (Figure 5D). By contrast, NSCs co-transfected with GOLPH3 shRNA and GOLPH3R90L rescue plasmids showed only diffuse GOLPH3 immunoreactivity distributed throughout the cell (Figure 5D). As the respective GOLPH3 signals reflected those of ectopically expressed GOLPH3 and GOLPH3R90L, these results established that GOLPH3 targeting to the NSC Golgi system requires a PITPNA/PITPNB-regulated Golgi PtdIns-4-P pool.
GOLPH3 is an effector of PITPNA/PITPNB-responsive PtdIns-4-P signaling
The PITPNA/PITPNB-and PtdIns-4-P-dependence of GOLPH3 targeting to the NSC Golgi network identified GOLPH3 as an attractive candidate for an effector of PITP and PtdIns-4-P signaling in NSCs. This hypothesis was supported by IUE experiments where, similar to Pitpna/Pitpnb deletion, GOLPH3 shRNA expression induced both a significant redistribution of the NSC Golgi system to a perinuclear location, and a compromised radial alignment, in individually transfected NSCs housed in an otherwise wild-type neurogenic niche (Figures 6A–C). Both phenotypic derangements were rescued by ectopic re-expression of wild-type GOLPH3, but not of the mutant GOLPH3R90L defective in PtdIns-4-P binding (Figure 6A–C).
Figure 6. Important role of GOLPH3 in Golgi localization and radial alignment of NSCs.
(A–C) Wild-type C57BL6 mouse embryos were electroporated with plasmids for expressing mCherry, EGFP-GM130, and control or GOLPH3 shRNA at E12.5, and sacrificed 72h after electroporation. In rescue experiments, a plasmid for expressing shRNA-resistant wild-type GOLPH3 or GOLPH3R90L, a mutant deficient in PtdIns-4-P binding, was added to the plasmid mixture. Confocal z-series images were used for analysis of NSC morphology and Golgi localization. Representative images are shown in (A), and quantifications are shown in (B) and (C). ***P<0.001, **** P<0.0001 compared to control, one-way ANOVA. (D–F) Mouse embryos with just one allele of Pitpna (genotype: Pitpnafl/+ Pitpnbfl/fl EmxCre/+) were electroporated with plasmids for expressing mCherry, EGFP-GM130, and with or without a plasmid for expressing GOLPH3L at E12.5. Electroporated embryos were sacrificed 72h later. Confocal z-series images were used for analysis of NSC morphology and Golgi localization. Representative images are shown in (D), and quantifications are shown in (E) and (F). **** P<0.0001 compared to control, Student’s t-test. Scale bars: 10µm. See also Suppl. Figure S6.
As an independent approach to assess the role of GOLPH3 in apical Golgi positioning and radial alignment, we took advantage of GOLPH3L -- a dominant-negative GOLPH3 antagonist (Ng et al., 2013). To that end, a GOLPH3L high-expression plasmid was introduced into the neocortex of mouse embryos by IUE. As was the case for SACM1L, elevated GOLPH3L expression was inconsequential for apical Golgi distribution, or for radial alignment, in NSCs carrying two wild-type alleles each of Pitpna and Pitpnb (data not shown). By contrast, ectopic GOLPH3L expression induced a marked disruption in the radial alignment of sensitized Pitpnafl/+ Pitpnbfl/fl Emx1Cre/+ NSCs expressing PITP activity at functional threshold along with a dramatic apical-to-perinuclear shift in Golgi distribution in these cells (Figures 6D–F). Taken together, the GOLPH3 knockdown and GOLPH3L interference data consistently demonstrated: (i) a key GOLPH3 requirement for maintaining the radial alignment and distribution of the NSC Golgi network throughout the apical process, and (ii) that GOLPH3 function was responsive to a PITPNA/PITPNB-regulated Golgi PtdIns-4-P pool. Interestingly, GOLPH3-deficient NSCs also gave rise to neurons that migrated over hundreds of micrometers to the most superficial layers of developing neocortex (data not shown). Thus, like PITPNA/PITPNB-deficient cells, GOLPH3-depleted migratory neurons were also competent for bulk membrane trafficking and efficient directed migration across large distances.
MYO18A defects phenocopy GOLPH3 defects
GOLPH3 interfaces with MYO18A to regulate Golgi shape in cultured mammalian cells (Dippold et al., 2009; Taft et al., 2013; Xing et al., 2016) -- thereby raising the possibility that a GOLPH3/MYO18A coupling regulates Golgi positioning in NSCs. If so, MYO18A deficiencies are predicted to induce NSC Golgi positioning defects similar to those caused by GOLPH3 knockdown. To test this prediction, we used a MYO18A shRNA system to assess whether MYO18A silencing could affect NSC apical Golgi distribution and radial alignment. Due to the lack of suitable MYO18A antibodies, we first assessed whether the shRNA could effectively silence MYO18A expression using a co-electroporation approach. Control shRNA or MYO18A shRNA were co-electroporated with an EGFP-MYO18A plasmid and an mCherry plasmid (as an indicator of transfection efficiency) into the neocortex of mouse embryos at E12.5. EGFP-MYO18A expression was subsequently analyzed 40h after electroporation. Relative to control shRNA, the MYO18A shRNA strongly reduced expression of EGFP-MYO18A (Suppl. Figure S6A).
To assess MYO18A function in NSCs, the MYO18A shRNA vector was co-electroporated with mCherry- and EGFP-GM130 expression plasmids into the neocortex of mouse embryos at E12.5, and the embryos were harvested after an additional 72h of incubation. Satisfyingly, familiar NSC Golgi positioning and radial alignment defects were induced by MYO18A shRNA expression, and those defects were rescued by ectopic re-expression of shRNA-resistant MYO18A (Suppl. Figures S6B–S6D).
Roles of other PtdIns-4-P-binding proteins
In addition to GOLPH3, other proteins, such as CERT (ceramide transfer protein; Hanada et al., 2007) and FAPP1/2 (Godi et al., 2004), bind PtdIns-4-P and are proposed to regulate various aspects of Golgi activity. It was of interest to determine whether these proteins also met the criteria for downstream PITP effectors in regulation of PtdIns-4-P-dependent NSC Golgi positioning and radial alignment. After confirming CERT expression in embryonic neocortex RT-PCR (data not shown), a suitable shRNA for CERT-silencing was identified using the IUE assay detailed in Suppl. Figure S7A. The shRNA plasmid was then co-electroporated with mCherry and EGFP-GM130 expression plasmids into E12.5 neocortex, and embryos were harvested after 72h. CERT shRNA expression evoked both NSC Golgi positioning and radial alignment defects resembling those induced by PITPNA/PITPNB or GOLPH3-deficiencies (Figure 7). Again, those defects were rescued by ectopic re-expression of an shRNA-resistant wild-type CERT, but not CERTG67E (Figure 7) -- i.e. a mutant CERT deficient in PtdIns-4-P-binding (Hanada et al., 2003). Thus, CERT also plays an important role in Golgi positioning in NSCs via a PtdIns-4-P-dependent mechanism. Golgi localization of GOLPH3 in NSCs was not compromised by CERT deficiencies, however, suggesting CERT was not operating through GOLPH3 in this system (Suppl. Figures S7B,S7C).
Figure 7. CERT regulates Golgi distribution and NSC radial alignment by a PtdIns-4-P-dependent mechanism.
Wild-type C57BL6 mouse embryos were electroporated with plasmids for expressing mCherry, EGFP-GM130, and control or CERT shRNA at E12.5, and sacrificed 72h after electroporation. In rescue experiments, a plasmid for expressing shRNA-resistant wild-type CERT or CERTG67E (a mutant deficient in PtdIns-4-P binding) was co-electroporated with CERT shRNA. Representative images are shown in (A), and quantifications are shown in (B) and (C). CERT shRNA-induced Golgi distribution defects and NSC polarity defects were rescued by wild-type CERT, but not the PtdIns-4-P-binding mutant G67E. **P<0.01, ***P<0.001 compared to control, one-way ANOVA. Scale bars: 10µm. See also Suppl. Figure S7. (D) Signaling pathway for apical Golgi positioning in NSCs. PITPNA and PITPNB leverage their PtdIns/PtdCho exchange activities to stimulate production of an NSC Golgi PtdIns-4-P pool that recruits GOLPH3 to Golgi membranes. The interaction of GOLPH3 with MYO18A bridges the system to the F-actin cytoskeleton and enables apical targeting of the Golgi system via this actin-based motor. CERT is also an effector of the PITPNA/PITPNB-dependent PtdIns-4-P pool. It does so either by acting downstream of GOLPH3 in the same NSC Golgi apical targeting pathway, or in a parallel targeting pathway.
FAPP1/2 mRNA expression profiles report that FAPP1 is robustly expressed in the neocortex of mouse embryos (GenePaint ID#: EH2708), whereas FAPP2 is not (GenePaint ID#: EN2065). IUE experiments indicated FAPP1 shRNA-mediated knockdown also induced NSC Golgi positioning and radial alignment defects. However, those defects were fully rescued by ectopic expression of the FAPP1R18L PtdIns-4-P binding mutant (data not shown; He et al., 2011). Thus, the FAPP1 involvement operates via a PtdIns-4-P-independent mechanism.
DISCUSSION
Both Sec14-like and StART-like PITPs stimulate PtdIns-4-P synthesis and signaling in eukaryotic cells, but to what biological outcomes such signaling circuits are coupled remains unclear. This is particularly true for the StART-like PITPs. Herein, we show that two highly related StART-like PtdIns/PtdCho-transfer proteins, PITPNA and PITPNB, cooperate in maintaining integrity of the neocortex during mammalian embryonic development. These findings outline five major conclusions: (1) PITPNA/PITPNB activity is critical for maintaining radial polarity of NSCs, (2) this function is executed via a cell-autonomous and PtdIns-4-P-dependent mechanism that controls efficient distribution of the NSC Golgi system throughout the apical compartment, (3) that the PITP involvement in NSC Golgi apical positioning exhibits a PITP-specificity independent of the lipid binding/exchange properties of the PITP, (4) the PtdIns-4-P and MYO18A-binding protein GOLPH3 is an effector of PITPNA/PITPNB-dependent PtdIns-4-P signaling in NSCs, and (5) that at least one other Golgi protein (CERT) also plays a key PtdIns-4-P-dependent role in apical distribution of the NSC Golgi complex. These results draw a mechanism where PITPNA/PITPNB chaperone PtdIns-4-P-dependent recruitment of GOLPH3 to Golgi membranes. This recruitment, in turn, promotes MYO18A- and F-actin-directed localization of the Golgi system throughout the apical compartment in a pathway that operates through, or in parallel with, CERT (Figure 7D). The collective data suggest segregation of the Golgi network to the apical compartment optimizes apical membrane trafficking and/or proper compartmentation of apical signaling during neurogenesis.
Functional redundancy for PITPNA and PITPNB in neurogenesis
PITPNA and PITPNB are highly similar at the primary sequence level. Both catalyze transfer of PtdIns and PtdCho between membrane bilayers in vitro, and both PITPs stimulate PtdIns-4-P synthesis by enhancing activities of PtdIns 4-OH kinases in vivo (Ile et al., 2010; Grabon et al., 2015). The functional relationship between these proteins in mammalian systems is not at all clear. The differential localization of the two proteins would suggest diversification of function in mammals. PITPNA adopts a cytoplasmic and nuclear localization, whereas PITPNB targets efficiently to the trans-Golgi network in cultured cells (De Vries et al., 1996; Phillips et al., 2006). In that regard, zebrafish PITPNB is required for maintenance and regeneration of retinal double cone cells and retinal integrity, but not for viability of new-born animals, whereas zebrafish PITPNA plays an essential role in early embryogenesis (Ile et al., 2010). The outcome is different in the mouse where PITPNA is not required for embryonic development but is essential for neonatal survival (Alb et al., 2003, 2007). By contrast, we find PITPNB null mice are viable and do not present obvious pathologies – at least up to 12 months of age. That PITPNA and PITPNB share functional redundancies during embryonic development is also clear. Whole-body DKO embryos failed to progress to E12.5, and a single wild-type allele of Pitpna was insufficient to rescue those defects.
PITPNA/PITPNB functional redundancies were apparent in embryonic neocortical development where we demonstrate that co-ablation of these two activities in NSCs resulted in mice lacking a dorsal forebrain. Functional ablation of PITPNA/PITPNB in embryonic neocortex evoked catastrophic disorganization of the tissue, and such global derangements ignited a cell non-autonomous train of apoptotic cell death of a scale incompatible with formation of even a rudimentary dorsal forebrain.
PtdIns-4-P and GOLPH3 control NSC Golgi positioning and cell polarity
Taken together, our data draw a mechanism where PITPNA/PITPNB and GOLPH3 act cooperatively in PtdIns-4-P-dependent control of NSC radial alignment and apical Golgi distribution, and that GOLPH3 is a key effector of PITPA/PITPNB-dependent PtdIns-4-P signaling in NSCs (Figure 7D). The case is built on the fact that: (i) the earliest derangements scored in PITPNA/PITPNB-deprived NSCs were cell-autonomous derangements in NSC cell polarity and the redistribution of the Golgi network from the apical cytoplasm to a perinuclear location at E12.5, (ii) PITPNAT59D expression (defective in stimulating PtdIns-4-P synthesis) failed to rescue PITPNA/PITPNB deficiency phenotypes, and (iii) that elevated expression of a catalytically active SACM1L PtdIns-4-P phosphatase in sensitized NSCs (i.e. expressing PITPNA at functional threshold) recapitulated those same Golgi and NSC cell polarity phenotypes whereas expression of the catalytic dead SACM1LD391N did not. These results demonstrated that PtdIns-4-P depletion defined the biochemical basis for the observed PITPNA/PITPNB-deficiency phenotypes. The case for GOLPH3 as effector of PtdIns-4-P signaling in NSC Golgi membranes was made by demonstrations that PITPNA/PITPNB deficiencies were phenocopied by: (i) knock-down of the PtdIns-4-P binding protein GOLPH3 in a manner that could not be complemented by a GOLPH3R90L defective in PtdIns-4-P binding, or (ii) expression of a dominant-negative GOLPH3L in sensitized NSCs expressing PITP activity at functional threshold.
GOLPH3 is the mammalian counterpart to the yeast Vps74, and these proteins both bind PtdIns-4-P and regulate protein trafficking through late Golgi compartments in their respective systems (Dippold et al., 2009; Wood et al., 2009; Wood et al., 2012). The relevance of GOLPH3 for human disease is well-established as the structural gene is amplified in a number of aggressive cancers (Scott et al., 2009). At the cellular level, GOLPH3 links the Golgi network to the actin cytoskeleton via its interactions with the non-conventional myosin MYO18A. In so doing, the GOLPH3/MYO18A/F-actin axis is proposed to constitute a core component of the trafficking machinery that applies a tensile force onto Golgi membranes and drives secretory vesicle release from this organelle (Dippold et al., 2009; Taft et al., 2013). Thus, one interpretation of our data is that PITPNA/PITPNB-deficiencies compromise Golgi secretory capacity and that a bulk membrane trafficking inefficiency sets the stage for wholesale NSC derangement and catastrophic failure of neocortical development.
While the possibility that PITPNA/PITPNB/GOLPH3 signaling promotes efficient secretion of a specific set of cargoes cannot yet be excluded, a role for GOLPH3 as a core component of the bulk vesicle trafficking machinery is not supported by our data. EM micrographs showed the Golgi network of PITPNA/PITPNB-deficient NSCs to be morphologically unperturbed -- arguing that bulk membrane trafficking from the Golgi complex was not strongly affected in mutant NSCs. PITPNA/PITPNB/GOLPH3-dependent PtdIns-4-P signaling executes a more nuanced Golgi activity. Another striking finding from the IUE studies was that neither PITPNA/PITPNB- nor GOLPH3-deficient NSCs were prone to apoptosis, nor were differentiated cells derived from those mutant NSCs defective in neuronal migration over significant distances in the developing neocortex. While we cannot at this time formally exclude the possibility that timing of onset of PITPNA/PITPNB depletion (E11.5 for DKO neocortex and E12.5 for the IUE experiments) determines whether PITP-depleted NSCs will apoptose, or not, we favor the interpretation that PITP-deficient NSCs are viable cells when housed in an otherwise appropriate neurogenic niche.
Substantial evidence indicates that the Golgi network orients itself towards the leading edge of cells undergoing directional migration (Kupfer et al., 1982; Yadav et al., 2009; Krause and Gautreau, 2014). Indeed, GOLPH3 coordinates Golgi reorientation with directed cell migration in cultured cells (Xing et al., 2016). However, PITP- and GOLPH3-deficient neurons efficiently negotiated directional migration over large distances (800–1000 micrometers in some cortical areas). Thus, neither PITPNA/PITPNB nor GOLPH3 appear to serve as essential regulators of neuronal directional migration. Rather, we posit that a PITPNA/PITPNB-regulated PtdIns-4-P pool recruits GOLPH3 to late Golgi membranes where it leverages its MYO18A-binding activity to engage the Golgi system with the F-actin cytoskeleton -- thereby facilitating loading of the Golgi network throughout the NSC apical compartment (Figure 7D). Such an apically-disposed Golgi distribution would facilitate apical sorting of protein and lipid secretory cargo and thereby optimize Golgi responses to apical plasma membrane signaling.
Broader implications for PtdIns-4-P signaling in NSC homeostasis and neocortical development
Data from this work, and from a companion study (Koe et al, 2018), support the emerging understanding that PITP involvement in NSC polarity and NSC homeostasis will be a general concept. In that study, PITP activity in Drosophila neuroblasts (NSC equivalents) is shown to stimulate production of a PtdIns-4-P pool that promotes proper neuroblast homeostasis. It does so by ensuring non-muscle myosin II-dependent asymmetric cell division into a self-renewing neuroblast and a differentiating daughter cell. These concepts also extend to the NSC progeny. We find the PITP/PtdIns-4-P/GOLPH3 signaling axis also plays a critical role in the behavior of differentiating NSC progeny that have reached the cortical plate – i.e. projection neurons. Failures in this signaling axis result in projection neuron accumulation at the pial surface where defects in apical processes of these neurons derange stratification of the growing neocortex into its anatomically correct six-layered structure (data not shown).
Implications for mechanisms of mammalian PITP function
Finally, this study holds interesting mechanistic implications for how PITPNA/PITPNB operate in potentiating PtdIns-4-P signaling in the NSC Golgi system. Historically, PITPs are considered to drive PtdIns transport to membranes engaged in phosphoinositide signaling that become deficient in this lipid due to its consumption by high-performance phosphoinositide metabolism. The rationale has been that the differential PtdIns/PtdCho-binding activities of these PITPs drives vectorial transfer of PtdIns from the endoplasmic reticulum (ER) to a signaling membrane system, and retrograde transport of PtdCho from that system back to the ER, on the basis of differential ER/signaling membrane lipid compositions (Wirtz et al., 1991). The acid test of this model, not previously performed in mammalian systems, is to express a structurally unrelated PITP with essentially the same PtdIns/PtdCho-binding properties in NSCs, and assess the ability of that PITP to substitute for PITPNA/PITPNB in apical Golgi positioning. Our demonstration that yeast Sec14 is completely unable to do so does not conform to the predictions of simple lipid transfer models that it should be an effective surrogate. Rather, the data support the concept that PITPs define highly specified PtdIns-4-P pools generated by instructive regulation of PtdIns 4-OH kinases (Schaaf et al., 2008; Bankaitis et al., 2010; Grabon et al., 2015).
STAR METHODS
CONTACT FOR RESOURCE AND REAGENT SHARING
Requests for resources, reagents or further information should be directed to and will be fulfilled by Vytas A. Bankaitis (vytas@tamhsc.edu) or Zhigang Xie (zxie@tamhsc.edu).
EXPERIMENTAL MODEL AND SUBJECT DETAILS
Mouse Models
Mice used in this study, including those for IUE experiments, were on the background of C57BL6 and maintained in the animal facility of Texas A&M University Health Science Center. Mice were handled in accordance with National Institute of Health and institutional guidelines on the care and use of animals. Pitpna knockout mice have been previously described (Alb et al., 2003). Pitpnb knockout, Pitpnafl/fl, and Pitpnbfl/fl mice were generated in this study. Emx1Cre mice were procured from The Jackson Laboratory (Stock #005628, B6.129S2-Emx1tm1(cre)Krj/J. For all experiments involving floxed alleles of Pitpna and/or Pitpnb, littermate embryos with at least one non-floxed allele of Pitpna or Pitpnb was used as controls. For experiments involving whole-body Pitpna knockout or Pitpnb knockout, wild-type littermate embryos were used as controls. Embryonic stages for each experiment are indicated in the figures and figure legends. The gender of embryos was unknown at the time of harvest.
METHOD DETAILS
Plasmids
pCAX-EGFP and pCAX-mCherry, which directs the expression of EGFP and mCherry, respectively, under the CAG promoter, were previously described (Xie et al., 2016). pCAG-Cre was constructed by replacing the GFP-Cre coding sequence of pCAG-Cre:GFP (Matsuda and Cepko, 2007) with Cre coding sequence derived from the same plasmid via polymerase chain reaction (PCR). To generate plasmids for the expression of Pitpna, Pitpnb (isoform 2), Sacm1l, Golph3, Golph3l, Myo18a, Cert (Col4a3bp), and Fapp1 (plekha3), the coding sequences of these genes were amplified from neocortical tissues of E14.5 mouse embryos by reverse transcription (RT)-PCR using iScript Select cDNA Synthesis Kit (BIO-RAD) and Phusion high-fidelity DNA polymerase (New England Biolabs). These coding sequences were inserted into the pCAX vector using the restriction sites ECoRI, NotI (engineered into pCAX immediately downstream of EcoRI), BamHI, and NheI. The EcoRI site in the Sacm1l coding sequence was removed by site-directed mutagenesis that did not alter amino acid coding at the mutated site. Nucleotide changes encoding T59D, D391N, R90L, G67E, and R18L mutations were introduced into pCAX-PITPNA, pCAX-SACM1L, pCAX-GOLPH3, pCAX-CERT, and pCAX-FAPP1, respectively, by site-directed mutagenesis to generate pCAX-PITPNA-T59D, pCAX-SACM1L-D391N, pCAX-GOLPH3-R90L, pCAX-CERT-G67E, and pCAX-FAPP1-R18L. The plasmids for expression of EGFP-tagged PITPNA and PITPNB were generated by replacing the EGFP coding sequence of pCAX-EGFP with a fragment comprised of EGFP-coding sequence fused in-frame with mouse Pitpna and Pitpnb (isoform 2) coding sequences, respectively. For expression of EGFP-GM130, a fluorescent Golgi marker, the EGFP coding sequence of pCAX-EGFP was replaced with a fragment comprised of EGFP coding sequence fused in-frame with mouse Gm130 coding sequence (derived from E14.5 mouse neocortical tissues via RT-PCR). For expression of FAPP1PH, a PtdIns-4-P biosensor, the EGFP coding sequence of pCAX-EGFP was replaced with a fragment comprised of human FAPP1 PH domain fused in-frame with EGFP. For expression of EGFP-SidMP4M, another PtdIns-4-P biosensor, the EGFP coding sequence of pCAX-EGFP was replaced by a fragment comprised of EGFP coding sequence fused in-frame with the coding sequence for the P4M domain (residues 546–647) of L. Pneumophila SidM protein. For expression of EGFP-MYO18A, EGFP-CERT, and EGFP-FAPP1, the PITPNA coding sequence of pCAX-EGFP-PITPNA was replaced with coding sequences of MYO18A, CERT, or FAPP1 (the new coding sequence was in-frame with the upstream EGFP coding sequence). The shRNA constructs for silencing mouse Golph3, Myo18a, Cert, and Fapp1 were generated by inserting annealed oligonucleotides, which contained small hairpin sequences targeting specific regions in mouse Golph3, Myo18a, Cert, or Fapp1, into the BamHI/Hind III sites of pSilencer 2.0-U6. The targeting sequences were 5’-gaaacgtacgggaacgattag-3’ for golph3 shRNA, 5’-gctgagcttcaggacatttct-3’ for Myo18a shRNA, 5’-gccacagtttacgtgagaaac-3’ for Cert shRNA, and 5’-ggaattcgtccaccgtgatga-3’ for Fapp1 shRNA. For shRNA rescue experiments, silent mutations were introduced into pCAX-GOLPH3 (5’-gaaacgtacggg aacgattag-3’ was mutated to 5’-gaaaTgtCCGTgaGcgTttag-3’), pCAX-MYO18A (5’-gctgagcttcaggacatttct-3’ was mutated to 5’-gctgaATtAcaAgaTatttct-3’), pCAX-CERT (5’-gccacagtttacgtgagaaac-3’ was mutated to 5’-gccacTCACtGcgtgagaaac-3’), and pCAX-FAPP1 (5’-ggaattcgtccaccgtgatga-3’ was mutated to ggaGttTgtGcaGcgAGatga) by site-directed mutagenesis. All plasmids were verified by sequencing. For shRNA rescue experiments, silent mutations were engineered into both plasmids for expressing wild-type proteins and plasmids for expressing their corresponding mutants.
Antibodies
The antibodies used include: rabbit polyclonal anti-PITPNA (ProteinTech), rabbit polyclonal anti-PITPNB (Hamilton et al., 1997), rabbit polyclonal anti-GOLPH3 (Abcam), rabbit polyclonal anti-GRASP65 (Abcam), rabbit polyclonal anti-GM130 (Proteintech), rabbit polyclonal anti-GOLGA1 (Cell Signaling), chicken polyclonal anti-GFP (Aves Labs), goat polyclonal anti-GFP (Abcam), rabbit polyclonal anti-mCherry (Abcam), mouse monoclonal anti-Pax6 (Developmental Studies Hybridoma Bank), rabbit polyclonal anti-Tbr2 (Abcam), mouse monoclonal anti-Nestin (BD Biosciences), mouse monoclonal anti-ı-actin (GenScript), rabbit polyclonal anti-activated Caspase 3 (Cell Signaling), and rabbit anti-Ser10-phosphorylated histone H3 (Genscript). Secondary antibodies were from Jackson Immuno Research Laboratories, Inc (West Grove, PA) with minimal species cross reactivity.
IUE
The IUE procedure was performed in accordance with National Institute of Health and institutional guidelines on the care and use of animals. Briefly, timed-pregnant female mice were anesthetized, and laparotomy was performed to expose the uteri. Sterile plasmid solutions were injected into the brain of embryos through the uterine wall, and electric pulses were then delivered across the head of embryos. After electroporation, the uteri were returned to the abdominal cavity of the pregnant female, the incision was sutured, and the pregnant female was allowed to recover at a warm location. The following plasmid mass ratios were used in experiments where a mixture of plasmids were electroporated into the neocortex of mouse embryos: 1:3 for experiments using a mixture of pCAX-EGFP and Pitpnb shRNA (or control shRNA); 1:1:3 for experiments using a mixture of pCAX-mCherry, pCAX-EGFP-GM130, and Golph3 shRNA (or control shRNA); 1:1:3:3 for experiments using a mixture of pCAX-mCherry, pCAX-EGFP-GM130, Golph3 shRNA, and pCAX-GOLPH3 or pCAX-GOLPH3-R90L; 1:1:2 for experiments using a mixture of pCAX-mCherry, pCAX-EGFP-GM130, and pCAG-Cre; 1:1:2:2 for experiments using a mixture of pCAX-mCherry, pCAX-EGFP-GM130, pCAG-Cre, and pCAX-PITPNA or pCAX-PITPNA-T59D; 1:1:2 for experiments using pCAX-mCherry, pCAX-EGFP-GM130, and pCAX-SACM1L or pCAX-SACM1L-D391N; and 1:1:2 for experiments using pCAX-mCherry, pCAX-EGFP-GM130, and pCAX-GOLPH3L. For experiments involving shRNA plasmids targeting Myo18a, Cert, or Fapp1, plasmid ratios were similar to those used in GOLPH3 shRNA experiments.
Mouse Genotyping
Tissues obtained from ear punching of mice were used for genomic DNA preparation and PCR genotyping. Genotyping primers for mice with systemic deletion of Pitpna were previously described (Alb et al., 2003). Primers and the length of the PCR products for other alleles were: Delta13-F3 (5’-AGCCTACATCAATTATATGTAATGTATATACA-3’) and Delta13-R1 (5’-GCAAAAATACTTACCTCTTGAACAG-3’) for systemic Pitpnb deletion (expected PCR product sizes: 100bp for wild-type allele and 87bp for deleted allele). aLOX1 (5’-CTTCCTCTGCCTTGTAATCCTGAG-3’) and SDL1 (5’-GAACAAGAAACTATCCAGCAGACAGACT-3’) for Pitpna floxed allele (expected PCR product sizes: 303bp for non-floxed allele and 365bp for floxed allele; bLOX1 (5’-GAGGACTGCTGTGTCTGCTGC-3’) and SDL2 (5’-GTTTAGCTATGTAAGGGTTACTGT GCA-3’) for Pitpnb floxed allele (expected PCR product sizes: 347bp for non-floxed allele and 415bp for floxed allele); Emx1-1 (5’-AAGGTGTGGTTCCAGAATCG-3’) and Emx1-2 (5’-CTCTCCACCAGAAGGCTGAG-3’) for wild-type Emx1 allele (no Cre knockin; expected PCR product size: 378bp); and Cre1 (5’-GCGGTCTGGCAGTAAAAACTATC-3’) and Cre2 (5’-GTGAAACAGCATTGCTGT CACTT-3’) for Cre knock-in allele (expected PCR product size: 102bp).
Tissue Preparation, Immunostaining, and Confocal Microscopy
Forebrain hemispheres of electroporated mouse embryos were harvested as previously described (Xie et al., 2006). Briefly, the lateral ventricle was exposed by removal of the hippocampal tissue, the remaining forebrain hemisphere was then fixed in 2% paraformaldehyde (prepared in PBS) for 20–30min. Following fixation, the hemispheres were cryoprotected with 20% sucrose (prepared in PBS) and then embedded in Tissue-Tek OCT. Thirty-micron cryosections were prepared from hemispheres and used for immunostaining. Antibodies were diluted in 1×PBS containing 3% bovine serum albumin and 0.2% Triton-X-100. Generally, the primary antibody incubation step was performed at room temperature for overnight, and the secondary antibody incubation step was performed at room temperature for 1h. Secondary antibodies were conjugated with Cy2 (green), Cy3 (red), Cy5 (far red), or DyLight405 (blue). For some of the samples, DAPI was used to label the nucleus. Confocal images were acquired on a Nikon TiE confocal microscope using the NIS-Elements software.
Transmission Electron Microscopy
Forebrain tissues of E12.5 mouse embryos were fixed in 4% paraformaldehyde and 2.5% glutaraldehyde overnight, and subjected to transmission electron microscopy analysis at Image Analysis Laboratory in College of Veterinary Medicine & Biomedical Sciences at Texas A&M University. Images were acquired on comparable neocortical regions (medial to mediolateral in the mediolateral axis; middle part of ventricular zone in the radial axis) of Pitpna Pitpnb double knockout embryos and control embryos.
QUANTIFICATION AND STATISTICAL ANALYSIS
Confocal images obtained using the Nikon NIS Elements software were converted to TIF files, which were then used for quantification of gray value intensity or length/distance in Adobe Photoshop CS6. For all of the quantifications, cells from images obtained from at least three embryos were pooled together for each group. To quantify GOLPH3 immunostaining intensity for each cell, the background gray value (obtained in a nearby non-transfected area, outside of the Golgi apparatus) was subtracted from the mean gray value of GOLPH3 immunostaining in the Golgi area (indicated by EGFP-GM130 fluorescence in a different channel). The GOLPH3 immunostaining intensity in the Golgi area of a transfected cell was then compared to the average of GOLPH3 immunostaining intensity in the Golgi area of three nearby non-transfected cells (in which GOLPH3 exhibited Golgi-specific staining) to obtain a ratio that reflected the relative GOLPH3 immunostaining intensity in the Golgi area of that transfected cell. For Golgi-nucleus distance quantification, the distance between the proximal edge of the nucleus and the center of the Golgi segments (indicated by EGFP-GM130 fluorescence) in a transfected NSC was measured and compared between different groups.
For radial alignment quantification, the sum of the length (in the radial orientation) of the soma plus the length of the apical process was compared to the width (perpendicular to the radial orientation) of the soma in the region where the soma was the widest. This length to width ratio was then compared between different groups. To assess apoptosis in the in utero electroporation experiments, confocal images of transfected neocortices (from at least three embryos for each group) were used for quantification of the percentages of transfected cells (indicated by mCherry expression or EGFP expression) that were undergoing apoptosis (indicated by immunoreactivity of activated Caspase 3). For each embryo, about 100–1000 transfected cells were quantified for immunoreactivity of activated Caspase 3.
GraphPad Prism (Prism 6, Version 6.0b, GraphPad Software, Inc) was used for statistical analysis. Student’s t-test (unpaired, two-tailed) and one-way ANOVA were used to compare results from two different groups and compare results from more than two groups, respectively. In experiments where each cell represents one data point, cells from three or more mouse embryos were pooled together for each group. Values are given as mean ± standard deviation.
Supplementary Material
Highlights.
-
-
PITPNA/PITPNB regulate radial polarity of neural stem cells
-
-
PITPNA/PITPNB promote loading of the Golgi network into the apical process
-
-
PITPNA/PITPNB operate via a Golgi pool of phosphatidylinositol-4-phosphate
-
-
GOLPH3 and CERT are effectors of phosphatidylinositol-4-phosphate signaling
Acknowledgments
GRANT SUPPORT. This work was supported by grants RO1GM112591 from the National Institutes of Health and BE-0017 from the Robert A. Welch Foundation to VAB. LZ and CSA were supported by grants HL120846 and HL40387 from the National Institutes of Health to CSA. We thank the Image Analysis Laboratory in College of Veterinary Medicine & Biomedical Sciences at Texas A&M University for assistance with thin-section transmission electron microscopy, and Gerald Hammond (Univ. Pittsburgh) for a SidM PtdIns-4-P biosensor plasmid.
VAB dedicates this paper to the memory of Philip W. Majerus, a pioneer in the field of inositol-phosphate and phosphoinositide signaling whose giant intellect and exacting standards enriched the lives of those of us fortunate enough to have been included in his circle.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
DECLARATION OF INTERESTS. The authors declare no competing interests.
AUTHOR CONTRIBUTIONS. ZX, SKH, LZ, CSA and VAB designed the genetic, IUE, imaging, and cell biological experiments. ZX and SKH performed all the experiments reported herein, while LZ and CSA designed and generated the Pitpnbflox mouse line. ZX, SKH and VAB designed and produced the figures, and all authors contributed to writing of the manuscript.
References
- Alb JG, Jr, Cortese JD, Phillips SE, Albin RL, Nagy TR, Hamilton BA, Bankaitis VA. Mice lacking phosphatidylinositol transfer protein alpha exhibit spinocerebellar degeneration, intestinal and hepatic steatosis, and hypoglycemia. J. Biol. Chem. 2003;278:33501–33518. doi: 10.1074/jbc.M303591200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alb JG, Jr, Phillips SE, Rostand K, Cotlin L, Pinxteren J, Manning T, Guo S, York JD, Sontheimer H, Collawn JF, Bankaitis VA. Ablation of phosphatidylinositol transfer protein function in murine cells. Mol. Biol. Cell. 2002;13:739–754. doi: 10.1091/mbc.01-09-0457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alb JG, Jr, Phillips SE, Wilfley LR, Philpot BD, Bankaitis VA. The pathologies associated with functional titration of phosphatidylinositol transfer protein α activity in mice. J. Lipid Res. 2007;48:1857–1872. doi: 10.1194/jlr.M700145-JLR200. [DOI] [PubMed] [Google Scholar]
- Bankaitis VA, Mousley CJ, Schaaf G. Sec14-superfamily proteins and the crosstalk between lipid signaling and membrane trafficking. Trends in Biochemical Sciences. 2010;35:150–160. doi: 10.1016/j.tibs.2009.10.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bard F, Malhotra V. The formation of TGN-to-plasma membrane transport carriers. Annu. Rev. Cell Dev. Biol. 2006;22:439–455. doi: 10.1146/annurev.cellbio.21.012704.133126. [DOI] [PubMed] [Google Scholar]
- Bjornsson CS, Apostolopolou M, Tian Y, Temple S. It takes a village: Constructing the neurogenic niche. Dev. Cell. 2015;32:435–446. doi: 10.1016/j.devcel.2015.01.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Vries KJ, Westerman J, Bastiaens PI, Jovin TM, Wirtz KW, Snoek GT. Fluorescently labeled phosphatidylinositol transfer protein isoforms (alpha and beta), microinjected into fetal bovine heart endothelial cells, are targeted to distinct intracellular sites. Exp. Cell Res. 1996;227:33–39. doi: 10.1006/excr.1996.0246. [DOI] [PubMed] [Google Scholar]
- Dippold HC, et al. GOLPH3 bridges phosphatidylinositol-4-phosphate and actomyosin to stretch and shape the Golgi to promote budding. Cell. 2009;139:337–351. doi: 10.1016/j.cell.2009.07.052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghosh S, Marquardt T, Thaler JP, Carter N, Andrews SE, Pfaff SL, Hunter T. Instructive role of a PKCζ subcellular localization in the assembly of adherens junctions in neural progenitors. Proc. Natl. Acad. Sci. U.S.A. 2008;105:335–340. doi: 10.1073/pnas.0705713105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Glick BS, Malhotra V. The curious status of the Golgi apparatus. Cell. 1998;95:883–889. doi: 10.1016/s0092-8674(00)81713-4. [DOI] [PubMed] [Google Scholar]
- Glick BS, Nakano A. Membrane traffic within the Golgi apparatus. Annu. Rev. Cell Dev. Biol. 2009;25:113–132. doi: 10.1146/annurev.cellbio.24.110707.175421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Godi A, Di Campli A, Konstantakopoulos A, Di Tullio G, Alessi DR, Kular GS, Daniele T, Marra P, Lucocq JM, De Matteis MA. FAPPs control Golgi-to-cell-surface membrane traffic by binding to ARF and PtdIns(4)P. Nat Cell Biol. 2004;6:393–404. doi: 10.1038/ncb1119. [DOI] [PubMed] [Google Scholar]
- Gorski JA, Talley T, Qiu M, Puelles L, Rubenstein JLR, Jones KR. Cortical excitatory neurons and glia, but not GABAergic neurons, are produced in the Emx1-expressing lineage. J. Neurosci. 2002;22:6309–6314. doi: 10.1523/JNEUROSCI.22-15-06309.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grabon A, Khan D, Bankaitis VA. Phosphatidylinositol transfer proteins and instructive regulation of lipid kinase biology. Biochim Biophys Acta. 2015;1851:724–735. doi: 10.1016/j.bbalip.2014.12.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graham TR, Burd CG. Coordination of Golgi functions by phosphatidylinositol 4-kinases. Trends Cell Biol. 2011;21:113–121. doi: 10.1016/j.tcb.2010.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Greig LC, Woodworth MB, Galazo MJ, Padmanabhan H, Macklis JD. Molecular logic of neocortical projection neuron specification, development and diversity. Nat Rev Neurosci. 2013;14:755–769. doi: 10.1038/nrn3586. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamilton BA, et al. The vibrator mutation causes neurodegeneration via reduced expression of PITPα: Positional complementation cloning and extragenic suppression. Neuron. 1997;18:711–722. doi: 10.1016/s0896-6273(00)80312-8. [DOI] [PubMed] [Google Scholar]
- Hanada K, Kumagai K, Yasuda S, Miura Y, Kawano M, Fukasawa M, Nishijima M. Molecular machinery for non-vesicular trafficking of ceramide. Nature. 2003;426:803–809. doi: 10.1038/nature02188. [DOI] [PubMed] [Google Scholar]
- Hanada K, Kumagai K, Tomishige N, Kawano M. CERT and intracellular trafficking of ceramide. Biochim Biophys Acta. 2007;1771:644–653. doi: 10.1016/j.bbalip.2007.01.009. [DOI] [PubMed] [Google Scholar]
- He J, Scott JL, Heroux A, Roy S, Lenoir M, Overduin M, Stahelin RV, Kutateladze TG. Molecular basis of phosphatidylinositol 4-phosphate and ARF1 GTPase recognition by the FAPP1 pleckstrin homology (PH) domain. J Biol Chem. 2011;286:18650–18657. doi: 10.1074/jbc.M111.233015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang J, Ghosh R, Tripathi A, Lonnfors M, Somerharju P, Bankaitis VA. Two-ligand priming mechanism for potentiated phosphoinositide synthesis is an evolutionarily conserved feature of Sec14-like phosphatidylinositol and phosphatidylcholine exchange proteins. Mol. Biol. Cell. 2016;27:2317–2330. doi: 10.1091/mbc.E16-04-0221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ile KE, Kassen S, Cao C, Vihtehlic T, Shah SD, Huijbregts RPH, Alb JG, Jr, Stearns GW, Brockerhoff SE, Hyde DR, Bankaitis VA. The zebrafish class 1 phosphatidylinositol transfer protein family: PITPβ isoforms and double cone cell outer segment integrity in retina. Traffic. 2010;11:1151–1167. doi: 10.1111/j.1600-0854.2010.01085.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Insolera R, Bazzi H, Shao W, Anderson KV, Shi SH. Cortical neurogenesis in the absence of centrioles. Nat Neurosci. 2014;17:1528–1535. doi: 10.1038/nn.3831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koe CT, Tan YS, Lonnfors M, Hur SK, Low CS, Zhang Y, Yu F, Kanchanawong P, Bankaitis VA, Wang H. Vibrator and PI4KIIIα govern neuroblast polarity by anchoring non-muscle myosin II. eLife. 2018 doi: 10.7554/eLife.33555. (In Press) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krause M, Gautreau A. Steering cell migration: lamellapodium dynamics and the regulation of directional persistence. Nat. Rev. Mol. Cell Biol. 2014;15:577–590. doi: 10.1038/nrm3861. [DOI] [PubMed] [Google Scholar]
- Kupfer A, Louvard D, Singer SJ. Polarization of the Golgi apparatus and the microtubule-organizing center in cultured fibroblasts at the edge of the experimental wound. Proc. Natl. Acad. Sci. U.S.A. 1982;79:2603–2607. doi: 10.1073/pnas.79.8.2603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liang H, Hippenmeyer S, Ghashghaei HT. A Nestin-cre transgenic mouse is insufficient for recombination in early embryonic neural progenitors. Biol Open. 2012;1:1200–1203. doi: 10.1242/bio.20122287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y, Boukhelifa M, Tribble E, Morin-Kensicki E, Uetrecht A, Bear JE, Bankaitis VA. The Sac1 phosphoinositide phosphatase regulates Golgi membrane morphology and mitotic spindle organization in mammals. Mol. Biol. Cell. 2008;19:3080–3096. doi: 10.1091/mbc.E07-12-1290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lopez MC, Nicaud J-M, Skinner HB, Vergnolle C, Kader JC, Bankaitis VA, Gaillardin C. A phosphatidylinositol/ phosphatidylcholine transfer protein is required for differentiation of the dimorphic yeast Yarrowia lipolytica from the yeast to the mycelial form. J. Cell Biol. 1994;125:113–127. doi: 10.1083/jcb.125.1.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsuki T, Matthews RT, Cooper JA, et al. Reelin and Stk25 have opposing roles in neuronal polarization and dendritic Golgi deployment. Cell. 2010;143:826–836. doi: 10.1016/j.cell.2010.10.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mora-Bermúdez F, Huttner WB. Novel insights into mammalian embryonic neural stem cell division: focus on microtubules. Mol. Biol. Cell. 2015;26:4302–4306. doi: 10.1091/mbc.E15-03-0152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ng MM, Dippold HC, Buschman MD, Noakes CJ, Field SJ. GOLPH3L antagonizes GOLPH3 to determine Golgi morphology. Mol. Biol. Cell. 2013;24:796–808. doi: 10.1091/mbc.E12-07-0525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Phillips SE, Ile K, Boukhelifa M, Huijbregts RPH, Bankaitis VA. Specific and nonspecific membrane binding determinants cooperate in targeting phosphatidylinositol transfer protein β-isoform to the murine trans-Golgi network. Mol. Biol. Cell. 2006;17:2498–2512. doi: 10.1091/mbc.E06-01-0089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rakic P. Evolution of the neocortex: Perspective from developmental biology. Nat Rev Neurosci. 2009;10:724–735. doi: 10.1038/nrn2719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schaaf G, Ortlund E, Tyeryar K, Mousley C, Ile K, Woolls M, Garrett T, Raetz CRH, Redinbo M, Bankaitis VA. The functional anatomy of PL binding and regulation of PIP homeostasis by proteins of the Sec14-superfamily. Molecular Cell. 2008;29:191–206. doi: 10.1016/j.molcel.2007.11.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scott KL, et al. GOLPH3 modulates mTOR signaling and rapamycin sensitivity in cancer. Nature. 2009;459:1085–1090. doi: 10.1038/nature08109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun T, Hevner RF. Growth and folding of the mammalian cerebral cortex: from molecules to malformations. Nat. Rev. Neurosci. 2014;15:217–232. doi: 10.1038/nrn3707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taft MH, Behrmann E, Munske-Weidemann L-C, Thiel C, Raunser S, Manstein DJ. Functional characterization of human myosin MYO-18A and its interaction with F-actin and GOLPH3. J. Biol. Chem. 2013;288:30029–30041. doi: 10.1074/jbc.M113.497180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taverna E, Götz M, Huttner WB. The cell biology of neurogenesis: toward an understanding of the development and evolution of the neocortex. Annu. Rev. Cell Dev. Biol. 2014;30:465–502. doi: 10.1146/annurev-cellbio-101011-155801. [DOI] [PubMed] [Google Scholar]
- Taverna E, Mora-Bermudez F, Strzyz PJ, Florio M, Icha J, Haffner C, Norden C, Wilsch-Brauninger, Huttner WB. Non-canonical features of the Golgi apparatus in bipolar epithelial neural stem cells. Scientific Reports. 2016;6 doi: 10.1038/srep21206. Article number 21206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vincent P, Chua M, Nogue F, Fairbrother A, Mekheel H, Xu Y, Allen N, Bibikova TN, Gilroy S, Bankaitis VA. A Sec14p-nodulin domain phosphatidylinositol transfer protein polarizes membrane growth of Arabidopsis root hairs. Journal of Cell Biology. 2005;168:801–812. doi: 10.1083/jcb.200412074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wirtz KWA. Phospholipid transfer proteins. Annu. Rev. Biochem. 1991;60:73–99. doi: 10.1146/annurev.bi.60.070191.000445. [DOI] [PubMed] [Google Scholar]
- Wood CS, Hung C-S, Huoh Y-S, Mousley CJ, Stefan CJ, Bankaitis V, Ferguson KM, Burd CG. Local control of PtdIns4P signaling in the Golgi apparatus by Vps74 and Sac1 phosphoinositide phosphatase. Mol. Biol. Cell. 2012;23:2527–2536. doi: 10.1091/mbc.E12-01-0077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wood CS, Schmitz KR, Bessman NJ, Setty TG, Ferguson KM, Burd CG. PtdIns4P recognition by Vps74/GOLPH3 links PtdIns 4-kinase signaling to retrograde Golgi trafficking. J. Cell Biol. 2009;187:967–975. doi: 10.1083/jcb.200909063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xie Z, Jones A, Deeney JT, Huh SK, Bankaitis VA. Inborn errors of long-chain fatty acid β-oxidation link neural stem cell homeostasis to autism. Cell Reports. 2016;14:991–999. doi: 10.1016/j.celrep.2016.01.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xie Z, Samuels BA, Tsai L-H. Cyclin-dependent kinase 5 permits efficient cytoskeletal remodeling—a hypothesis on neuronal migration. Cereb Cortex. 2006;16:i64–i68. doi: 10.1093/cercor/bhj170. [DOI] [PubMed] [Google Scholar]
- Xing M, Peterman MC, Davis RL, Oegema K, Siau AK, Field SJ. GOLPH3 drives cell migration by promoting Golgi reorientation and directional trafficking to the leading edge. Mol. Biol. Cell. 2016;27:3828–3840. doi: 10.1091/mbc.E16-01-0005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yadav S, Puri S, Lindstedt AD. A primary role for Golgi positioning in directed secretion, cell polarity and wound healing. Mol. Biol. Cell. 2009;20:1728–1736. doi: 10.1091/mbc.E08-10-1077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoder MD, Thomas LM, Tremblay JM, Oliver RL, Yarbrough LR, Helmkamp GM., Jr Structure of a multifunctional protein: Mammalian phosphatidylinositol transfer protein complexed with phosphatidylcholine. J. Biol. Chem. 2001;276:9246–9252. doi: 10.1074/jbc.M010131200. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.