Abstract
Influenza A virus (IAV) is an RNA virus with a segmented genome. These viral properties allow for the rapid evolution of IAV under selective pressure, due to mutation occurring from error-prone replication and the exchange of gene segments within a co-infected cell, termed reassortment. Both mutation and reassortment give rise to genetic diversity, but constraints shape their impact on viral evolution: just as most mutations are deleterious, most reassortment events result in genetic incompatibilities. The phenomenon of segment mismatch encompasses both RNA- and protein-based incompatibilities between co-infecting viruses and results in the production of progeny viruses with fitness defects. Segment mismatch is an important determining factor of the outcomes of mixed IAV infections and has been addressed in multiple risk assessment studies undertaken to date. However, due to the complexity of genetic interactions among the eight viral gene segments, our understanding of segment mismatch and its underlying mechanisms remain incomplete. Here, we summarize current knowledge regarding segment mismatch and discuss the implications of this phenomenon for IAV reassortment and diversity.
Keywords: influenza A virus, reassortment, segment mismatch, evolution, packaging, evolutionary constraint
Introduction
Influenza A virus (IAV) is an enveloped, negative-sense, single-stranded RNA virus with eight gene segments [1]. The segmented nature of the IAV genome allows the exchange of gene segments between viruses that co-infect the same cell, which can result in the formation of progeny viruses that are genetically distinct from both parental viruses. This process of genetic exchange is termed reassortment. Reassortment has a major impact on IAV evolution and therefore plays a substantial role in the ecology and epidemiology of IAV.
The principal reservoir of IAV resides in aquatic birds, although IAV can infect a diverse range of hosts, including humans, poultry, swine, horses and other mammals. IAV is grouped into distinct subtypes according to the antigenicity of two of its surface glycoproteins, haemagglutinin (HA) and neuraminidase (NA), of which there are currently 18 known HA subtypes and 11 known NA subtypes. Viruses of the H17N10 and H18N11 subtypes are confined to bats, while viruses of the remaining HA and NA subtypes circulate in wild birds [2–5].
In humans, IAV causes both epidemics and pandemics. IAV epidemics occur yearly during the winter months in both the northern and southern hemisphere. Reassortment among co-circulating seasonal IAV strains has been detected repeatedly and found to give rise to epidemiologically important strains [6–10]. Despite the evolutionary success of certain reassortant genotypes, however, comprehensive investigation of the fate of intra-subtype reassortants derived from the human H3N2 lineage revealed that reassortment is usually deleterious, resulting in negative selection of reassortant progeny [11]. This finding highlights the importance of segment mismatch in the evolution of seasonal IAV and indicates that inter-segment epistasis often functionally constrains reassortment even when co-infecting viruses are closely related.
Pandemics arise when an IAV strain acquires both antigenic novelty and the ability to spread efficiently through a human population. In the past century, there have been four IAV pandemics [12–15]. The origins of the 1918 H1N1 pandemic strain and the contribution of reassortment to its emergence remain under debate [16–18]. In 1957, the PB1, HA and NA genes from an avian H2N2 virus entered the human H1N1 background to form the H2N2 pandemic strain of that year. In 1968, the PB1 and HA genes from an avian H3 virus entered the human H2N2 background to form the 1968 H3N2 subtype pandemic strain [19]. A notable feature of both the 1957 and the 1968 pandemic strains is the transmission of an avian PB1 gene to humans, accompanying its cognate HA [20]. Lastly, extensive reassortment in swine brought together genes from avian, swine and human IAV lineages to produce the 2009 pandemic strain [21–23].
In addition to epidemics and pandemics, over the past 20 years, multiple zoonotic IAV outbreaks have occurred when viruses capable of causing severe disease in humans emerged from birds [24–29]. According to the World Health Organization (WHO), from 2003 to September 2017 there have been 860 human cases of H5N1 virus infection reported from 16 countries, with 454 of these cases resulting in death [30]. Furthermore, as of September 2017, 1564 laboratory-confirmed human cases of H7N9 virus infection have been reported in China, including at least 612 deaths [31]. The H5N1 and H7N9 subtype viruses causing these zoonotic outbreaks arose through reassortment of multiple avian-adapted IAVs, including strains circulating in both wild and domestic birds [32–34]. Interestingly, both the H5N1 and H7N9 zoonotic lineages derive their six ‘internal gene’ (non-HA, non-NA) segments from an H9N2 subtype lineage that circulates widely in the poultry of Southeast Asia. This internal gene cassette appears to be conducive to reassortment that pairs it with multiple different HA/NA subtypes [32, 33, 35–37]. In addition to the ongoing H5N1 and H7N9 IAV outbreaks, avian influenza viruses of other subtypes, such as H5N6, H9N2, H10N8 and H6N1, have sporadically caused disease in humans in China and Taiwan [37–40].
There is concern regarding the potential for viruses of these zoonotic subtypes to cause the next influenza pandemic. While the human population has little immunity against these avian IAVs, these viruses to date have not acquired effective human-to-human transmission, a key feature of a pandemic strain. Reassortment of zoonotic and human seasonal IAVs can facilitate the formation of a virus with both antigenic novelty and the ability to successfully transmit in humans. However, the fitness of reassortant progeny derived from divergent IAV strains is often limited by incompatibilities occurring at the RNA or protein level [41, 42]. Indeed, segment mismatch may be a major underlying reason that reassortment events involving H5N1 or H7N9 IAVs and seasonal human strains have not been detected to date.
As summarized above, IAVs are genetically diverse and circulate within a diverse range of host species. Under these conditions, reassortment can result in rapid viral evolution, potentially contributing to epidemics, pandemics and cross-species transfers of the virus. In this review, we will explore reassortment, the constraints imposed on reassortment by segment mismatch, and the implications of segment mismatch for IAV evolution.
Mutation and reassortment in IAV evolution
Fixation of changes in the viral genome requires that a virus population is genetically diverse and that it is subject to either a stochastic bottleneck or a selective pressure. For IAV, two distinct mechanisms give rise to genetic variation: mutation and reassortment. The diversity of genotypes generated through these two processes is the raw material on which selection acts. IAV evolution in humans is driven in large part by antibody-mediated selection and takes two forms: (i) antigenic drift, which is characterized by small changes and is a major feature of seasonal IAV, and (ii) antigenic shift, which is characterized by large changes and is a major feature of pandemic IAV. Drivers of IAV evolution in non-human hosts are less well characterized than in humans, and pressures unrelated to immunity, as well as stochastic processes, may play a larger role in these contexts.
Antigenic drift
Like all RNA viruses, IAVs are constantly mutating. These mutations are the result of the absence of proofreading capability of the viral RNA-dependent RNA polymerase. IAV polymerase error translates into approximately two mutations per replicated genome [43]. The fixation of mutations that change the antigenicity of IAVs encompasses antigenic drift. Antibodies directed against the HA glycoprotein can neutralize the virus by blocking viral attachment to and/or fusion with the host cell [44–47]. These anti-HA antibodies therefore drive antigenic drift of the virus towards antibody escape. Antigenic drift, in turn, has a major impact on the efficacy of vaccination; the mutations that the virus acquires over time lead to vaccine escape, therefore necessitating periodic reformulation of the vaccine in order to match the currently circulating strains [48–50].
In addition to mutation, reassortment can also contribute to antigenic drift [9]. Since multiple sub-clades of seasonal H3N2 or H1N1 viruses co-circulate, there is the potential for reassortment between viruses of closely related lineages. Such intra-subtype reassortment events increase genomic diversity and contribute to the evolution and epidemiology of seasonal influenza [6, 7, 51, 52]. Most notably, reassortment within H1N1 or H3N2 subtypes can place a drifted HA onto a more fit background, which can give rise to unusually severe epidemics, as seen in 1946–1947 and 2003–2004 [6, 51]. Reassortment within IAV subtypes can also affect sensitivity to antiviral drugs. The global spread of adamantane-resistant IAVs was mapped to an H3N2 lineage that arose from an intra-subtype reassortment event [53, 54].
IAVs of the H1N1 and H3N2 subtypes have been co-circulating in humans since 1977 [55]. Their reassortment is detected only sporadically, however, and sustained transmission of a human H1N1/H3N2 reassortant virus has occurred only in one instance. Namely, in 2001, reassortant H1N2 viruses were detected in humans in the UK [56]. During the 2001–2002 season, H1N2 viruses were identified in 41 countries on four continents, causing various outbreaks across the globe and predominating over H3N2 and H1N1 viruses in several countries [56–59]. However, in subsequent seasons, these reassortant H1N2 viruses failed to predominate. In addition to this outbreak, co-infection and reassortment between human H1N1 and H3N2 lineages were occasionally detected in humans prior to 2009 and, since the 2009 H1N1 pandemic strain (pH1N1) replaced the H1N1 seasonal lineage, a handful of additional co-infection and reassortment events between these two subtypes have been reported [60–68]. Despite apparently ample opportunity for inter-subtype reassortment among human IAV lineages, the relatively low frequency with which reassortant viruses achieve sustained circulation suggests that evolutionary constraints lead to their negative selection. Indeed, analysis of reassortment between seasonal H3N2 and pH1N1 viruses in the laboratory supports this conclusion [69, 70].
Antigenic shift
Antigenic shift involves a large change in the antigenicity of influenza viruses circulating in the human population, and can result from introduction into humans of an IAV that circulates in non-human hosts, or introduction of the HA segment of a non-human IAV into a seasonal IAV background through reassortment [4, 5]. As noted above, reassortment between human and avian IAVs led to the formation of the 1957 and 1968 pandemic strains [20]. Reassortment was also critical in the emergence of the 2009 pandemic strain. In contrast to the 1957 and 1968 pandemics, however, this reassortment did not directly involve currently circulating human IAVs. In the years prior to 2009, reassortment brought together gene segments from two distinct swine IAV gene pools, that of the Eurasian avian-like and the North American triple-reassortant swine lineages. The triple-reassortant lineage, in turn, incorporated viral genes of human, avian and classical swine IAV origin. Thus, although the 2009 pandemic was caused by an intact swine IAV, the emergence of this swine virus resulted from extensive reassortment in pigs [21, 22, 71]. Owing to a lack of IAV isolates in the years prior to 1918, the origins of the 1918 pandemic strain remain relatively unclear. Following sequencing of the reconstructed genome of the 1918 virus, it was suggested to have been introduced into humans in toto from an avian reservoir [17]. A more recent analysis suggests, however, that reassortment between a previously circulating human IAV of the H1 subtype and an avian IAV may have placed the human H1 HA into a novel background and given rise to the 1918 pandemic [16].
Although reassortment is often associated with the emergence of IAV pandemic strains, it is likely not sufficient to give rise to highly fit, antigenically shifted variants. Indeed, experimental co-infection of pigs with swine IAVs of the precursor lineages to the 2009 pandemic strain did not yield a pandemic-like genotype at detectable levels [72]. In other experimental settings, mutations selected following reassortment have been shown to compensate for genetic incompatibilities and thereby increase fitness of reassortant viruses. For example, amino acid substitutions in HA were found to reduce HA–NA mismatch in human–avian reassortant viruses [73, 74]. Evidence of this phenomenon occurring in nature has also been reported in the context of intra-subtype reassortment of seasonal IAV [75]. Owing to the difficulty of detecting low-fitness precursors, and of differentiating adaptation to a new genetic context from adaptation to a new host, it is more challenging to demonstrate this phenomenon for pandemic IAV. Nevertheless, for a shifted virus to be evolutionarily successful, mutations are likely needed to ease segment mismatch that arises from the reassortment event.
Before discussing segment mismatch and its causes in more detail, we will briefly consider other factors that impact the frequency of reassortment. In addition to viral genetic compatibility, virus extrinsic factors, such as the timing and dose of infection, have been shown to impact reassortment outcomes by altering the likelihood of co-infection.
Reassortment efficiency in the absence of segment mismatch
Although multiple methods exist by which to evaluate IAV reassortment in the laboratory (reviewed in [76]), studies regarding the circumstances that allow for robust reassortment are often confounded by segment mismatch. In order to examine the conditions that favour IAV reassortment and to determine a baseline frequency for this process, we developed a system that allowed IAV reassortment in the absence of segment mismatch. This system employed two influenza A/Panama/2007/99 (H3N2) [Pan/99] viruses that were phenotypically similar but genotypically distinct [77]. Up to six silent mutations were introduced into the open reading frames (ORF) of each of the eight Pan/99 segments of one virus and it was designated as variant, or VAR. These mutations did not alter the fitness of the virus detectably, but allowed for differentiation of the VAR virus from the wild-type (WT) Pan/99 virus using the post-PCR method high-resolution melt (HRM) analysis [78]. In this way, progeny viruses produced from WT/VAR co-infected cells could be genotyped in a streamlined fashion using HRM analysis, without the need for partial or full sequencing of virus isolates. Two different epitope tags were also introduced into the HA proteins of these viruses, to allow enumeration of infected and co-infected cells by flow cytometry. Because these two viruses were of the same strain background, the virus pairing was devoid of segment mismatch. Using this system, we showed that reassortment was highly efficient in cell culture under high-multiplicity conditions, with an average of 88 % of the analysed virus isolates containing a reassortant genome [77]. Additionally, we found that the frequency of reassortment was dose-dependent, with higher levels of co-infection in vitro and higher inoculum dose in vivo resulting in increased reassortment. The observed dose dependency of reassortment was partly explained by Poisson statistics dictating the levels of co-infection; however, comparison to a computational simulation of co-infection and reassortment in cell culture indicated that reassortment was markedly more efficient than predicted based on random distribution of eight-segmented viruses into cells [79]. A combination of further modelling and experimentation revealed that the abundant reassortment observed could be explained by high levels of incomplete viral genomes within singly infected cells. In particular, the occurrence of incomplete genomes is thought to increase the proportion of productively infected cells that are co-infected [79]. With sequential infection, rather than co-infection, a delay of up to 8 h between the WT/VAR virus inoculations allowed for robust reassortment in vitro, while a delay of up to 12 h allowed for robust reassortment in vivo. However, further time delay between the WT/VAR virus inoculations led to markedly reduced levels of co-infected cells in cell culture, which prevented reassortment. In vivo, a delay of 24 h or more blocked productive infection by the second virus. Stripping of sialic acid receptors from infected cells by NA may contribute to the reduction in superinfection over time [80]. By excluding the limitations imposed by segment mismatch, this study demonstrated for the first time that reassortment is efficient both in vitro and in vivo, and that the timing and dose of infection strongly impact the likelihood of reassortment [77, 79, 81, 82]. Taken together, the above studies suggest that (i) virus extrinsic factors do not limit reassortment appreciably within a co-infected cell, and (ii) virus extrinsic factors such as timing and dose do, however, play a major role in determining the likelihood of co-infection at the levels of both the host and the individual cell.
Segment mismatch
As described above, reassortment occurs frequently in the absence of RNA or protein mismatch. Additionally, numerous studies to date have demonstrated that reassortment is prevalent in nature, in both avian and swine hosts [83–88]. However, early evidence suggested that reassortment of IAV genes was not random. In 1979, Lubeck et al. demonstrated that reassortment between influenza A/Puerto Rico/8/34 (H1N1) [PR8] and influenza A/Hong Kong/8/68 (H3N2) viruses resulted in some segments being randomly distributed, as expected, while other segments appeared to be linked, occurring together with a higher frequency than expected [41]. In particular, linkages between the polymerase gene segments PB2-PB1 and PB1-PA derived from the same parental virus were observed, suggesting that either (i) these genes interacted with each other during IAV packaging, or (ii) optimal protein–protein interactions occurred when these genes originated from the same parental strain, increasing the likelihood of detecting progeny viruses with these gene combinations [41]. Later, using statistical approaches, the non-random association of IAV genes during reassortment was confirmed [42, 89]. Similarly, multiple studies undertaken for risk assessment purposes have demonstrated that certain reassortants derived from (i) swine IAVs and human IAVs and (ii) avian IAVs and human IAVs have fitness defects when compared to the parental strains, suggesting incompatibilities between specific heterologous viral components [72, 90–96]. These incompatibilities underlie segment mismatch and can be categorized as either RNA mismatch or protein mismatch.
RNA mismatch
RNA mismatch comprises incompatibilities between the viral RNAs (vRNAs) of two different IAV strains, and manifests at the packaging signal regions of IAV segments. Packaging signals direct the incorporation of the eight vRNAs into new virus particles. During reassortment, mixing of gene segments occurs, thus juxtaposing heterologous viral packaging signals.
IAV packaging
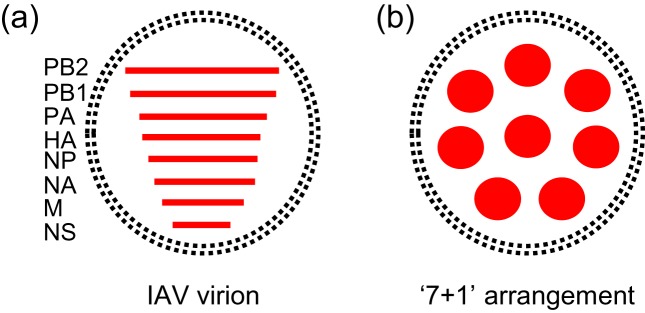
An IAV particle is only fully infectious if it contains one copy of each of the eight viral segments, as shown in Fig. 1. For many years, the mechanism by which packaging of eight distinct segments was achieved remained under debate. It was thought that either (i) the packaging of the IAV genome was random, or (ii) the packaging of the IAV genome was a selective process. Random assembly would require a full infectious genome to be generated purely by chance and, with eight segments, the probability of this occurring is 1/416 [97]. A major argument in favour of a random packaging model was that observed PFU per particle ratios of IAV were low, in the range of 1/10 to 1/100 [98–100]. Furthermore, if virus particles are able to package nine or more segments, this would bring the probability of packaging the eight distinct segments needed to make an infectious virus into the range of observed PFU/particle values [97]. Some studies offered support for packaging of greater than eight segments per virion. Results of one such study showed that 3–5 % of analysed virions incorporated two different reporter versions of the same vRNA segment, and the authors estimated that IAVs must package 9 to 11 segments for this to occur by chance [101]. In another study, reverse genetics techniques were used to produce an IAV that required delivery of nine segments to a cell to initiate productive infection [97]. For these reasons, a model in which IAV randomly incorporated 9 to 11 vRNPs per particle was plausible.
Fig. 1.
The influenza A virus genome. (a) Schematic diagram of the IAV genome, with the eight RNA genomic segments organized according to size from largest to smallest. (b) Arrangement of the IAV genome within a virus particle, demonstrating the ‘7+1’ pattern of the eight viral segments as observed via transmission electron microscopy performed by Noda et al. One central segment is surrounded by seven outer segments. However, the identity of the segments arranged in this pattern, as well as the precise interaction network between the segments, remains unknown. The glycoproteins present on the viral membrane have been removed for clarity.
However, strong evidence now suggests that a selective IAV packaging mechanism is at work. This evidence has been reviewed recently [102]. Briefly, studies in the 1970s revealed that the eight IAV gene segments were present in equimolar ratios in a virus population [103] and that most IAV plaque-forming units contained a single full set of segments [41, 104]. In addition, virus populations released from cells that contained unequal copy numbers of the eight segments were nevertheless found to carry equal copy numbers of the vRNAs [105, 106]. In subsequent years, fluorescence in situ hybridization analysis of vRNA composition in individual virions revealed that a majority of IAV particles contained eight distinct segments and packaged a single copy of any given gene [107]. In addition, two virus-like RNAs developed from the same vRNA species were shown to compete for packaging [108], in agreement with previous studies demonstrating that defective interfering (DI) and standard segments of the same vRNA species compete for packaging [109–112]. Taken together, these studies offered substantial support for a selective packaging mechanism and the existence of elements present in IAV viral ribonucleoprotein complexes (vRNPs) that allow for segment differentiation during packaging.
Packaging signals and segment interactions
IAV packaging signals direct the incorporation of each IAV segment into assembling virus particles. These packaging signals have been found to be segment and strain specific and comprised of both untranslated region (UTR) and coding sequence at the 3′ and 5′ end of each segment. A study reported in 2003 represented the first description of a defined segment-specific packaging signal for IAV [113]. An influenza A/WSN/33 (H1N1) [WSN] NA construct was designed that contained GFP inserted in place of the NA coding region, flanked by 202 nt of the wild-type NA 3′ end (UTR +183 nt coding region) and 185 nt of the wild-type NA 5′ end (UTR +157 nt coding region). After serially passaging a virus containing this NA construct, it was demonstrated that this model NA segment was efficiently incorporated into virus particles over time. Subsequently, another construct identical to the first was designed except with the coding regions removed. In the absence of these coding nucleotides, the NA segment was no longer efficiently packaged, although it was still replicated [113]. Packaging signals have since been defined for the remaining seven WSN IAV segments and found to comprise similarly broad regions of the 3′ and 5′ termini [114–118]. A handful of studies also suggest that internal nucleotides outside the mapped termini may also contribute to packaging [119–121]. Additionally, the amino acid sequence of the viral nucleoprotein (NP) might play an important role in directing packaging of the IAV segments [122]. This NP amino acid code does not alter the role of packaging signals as previously described; rather, it may act in concert with the RNA-based packaging signals to instruct IAV segment bundling.
Elegant transmission electron microscopy studies have helped to visualize how the eight segments are arranged within a virus particle. By transversely sectioning viruses as they were budding off host cell membranes, Kawaoka and colleagues showed that IAV gene segments were arranged in a distinct ‘7+1’ pattern, with seven segments in a circle surrounding one central segment as shown in Fig. 1 [123, 124]. An array of IAVs were tested in this manner, including human, swine and avian strains, and the same arrangement was observed for all strains tested. IAVs with spherical particles as well as filamentous particles were subsequently examined, with the same result [123]. Therefore, it is likely that this arrangement is consistent across all budding IAVs. To date, the identity of the segments in this ‘7+1’ pattern remains unknown; however, using electron tomography and characterizing the segments based on length, it was shown via sectioning of IAV particles that the identity of the central segment was probably PB2, PB1, PA or HA [125].
The observation that the location (but not necessarily segment order or identity) of vRNAs within a virion is consistent, at least during budding, raises the question: how are the segments being held in this position? The presence of electron density between vRNPs suggested the existence of direct contacts, both between segments in the outer ‘7+1’ circle and between the central segment and outer segments [125]. Fournier et al. confirmed the presence of interactions between the segments via electron tomography, and also showed that these interactions are probably occurring between the packaging signal regions of the IAV segments as previously mapped, as the locations of the interactions overlapped with the earlier described terminal packaging signal locations for each segment [126].
Packaging signal mismatch
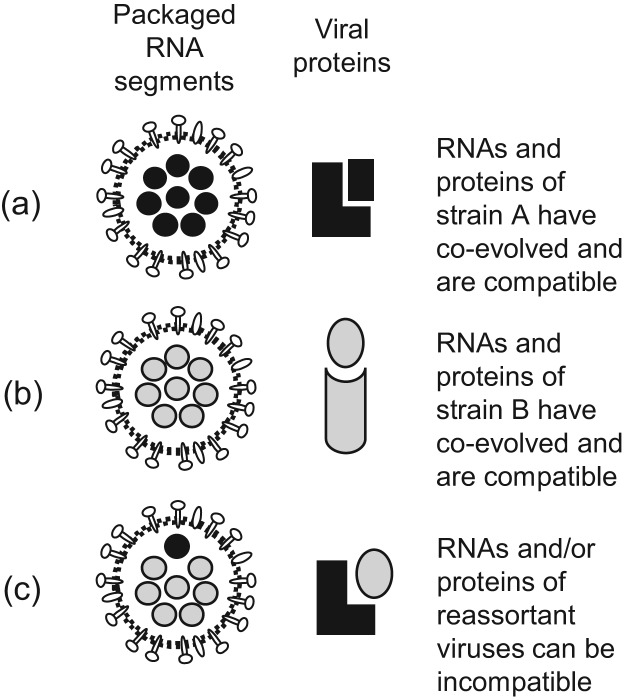
Based on the majority of evidence gathered to date, a leading model of IAV genome packaging is that the vRNAs are arranged in a particular pattern during virion assembly and interact with each other via base-pairing interactions between adjacent packaging signal sequences. If sequence-specific RNA-RNA interactions mediate selective genome assembly as this model suggests, it follows that segment mismatch at the RNA level would result from nucleotide differences among IAV strains. During IAV evolution, the RNAs of a given virus strain evolve coordinately, leading to optimal interactions among segments (Fig. 2a, b). When two divergent IAV strains undergo reassortment, the interactions between heterologous segments are expected to be less efficient since the precise sequences mediating these interactions might not be conserved (Fig. 2c). These inefficient interactions between vRNAs could, in turn, disfavour the production of certain reassortant genotypes. Indeed, packaging signal mismatch has been shown to dictate reassortment outcomes [127–129].
Fig. 2.
Reassortant influenza A viruses can have suboptimal RNA and/or protein interactions. During IAV evolution, the genomes and proteins of an individual strain co-evolve alongside one another, resulting in optimal interactions (a and b). During reassortment, the genomes of the two co-infecting strains are mixed. This can result in suboptimal interactions between the genomic segments and/or encoded proteins, leading to viruses that are less fit compared to the parental strains (c).
In an elegant series of experiments, Essere et al. showed that reassortment between influenza A/Moscow/10/99 (H3N2) and influenza A/Finch/England/2051/91 (H5N2) viruses resulted in strongly biased reassortment, with the H5 HA segment being unable to enter a H3N2 background [129]. However, when packaging signals from the H3 HA segment were grafted onto the H5 HA segment, this allowed for efficient incorporation of the H5 HA segment into a H3N2 background, demonstrating that packaging signal mismatch was inhibiting the formation of these reassortant progeny. Additionally, the incorporation of the H5 HA segment (with H5 packaging signals) could be achieved if accompanied by the H5N2 M segment, suggesting that the HA and M segments of this IAV strain interact at the level of genome assembly [129].
Influenza A and B viruses co-circulate in humans but never undergo reassortment. The reason for this could be multifactorial, but Baker et al. showed that by grafting IAV packaging signals onto full-length influenza B virus HA or NA glycoproteins, reassortant viruses could be rescued that contained either the influenza B virus HA, NA or both in an IAV background [128]. Their study demonstrated that packaging signal divergence between influenza A and B viruses is at least partly responsible for the lack of intertypic reassortment in nature. Interestingly, this study also serves as an example of RNA incompatibility with protein compatibility, as full-length influenza B virus HA protein was able to functionally complement an HA-deficient IAV [128].
Using a co-infection system based on Pan/99 WT and VAR viruses (described above), we recently examined the potential for packaging signal differences between seasonal H3N2 and pH1N1 viruses to shape reassortment outcomes. The advantage of this approach is that the co-infecting WT and VAR viruses are identical at the protein level, ensuring that biases in reassortment can be fully attributed to RNA sequence. Our data revealed that the presence of pH1N1 virus-derived packaging signals on the WT HA segment led to it being significantly under-represented in reassortant progeny viruses compared to a control virus that carried homologous Pan/99 packaging signals. These results confirmed the findings of Essere et al. that heterologous packaging signals on the HA segment can restrict reassortment and extended their observation to an H3N2/H1N1 virus pairing. However, when the NA and NS segments were analyzed in a similar manner, the presence of homologous versus heterologous packaging signals had no significant effect on packaging efficiency [127]. Therefore, this study suggests that the sensitivity to changes in packaging signal sequence may vary with segment.
Additionally, Gao and Palese successfully prevented IAV reassortment by rewiring the packaging signals of the HA and NS segments. Using the PR8 (H1N1) strain background, they grafted HA-specific packaging signals onto the NS segment and grafted NS-specific packaging signals onto the HA segment [130]. This first-generation chimeric virus was able to undergo reassortment with the wild-type virus. However, after the original packaging signal sequences in terminal regions of the ORFs were disrupted via silent mutation, the chimeric virus lost its ability to reassort with the wild-type virus. This study provides further evidence for the role of packaging signals in IAV reassortment.
Protein mismatch
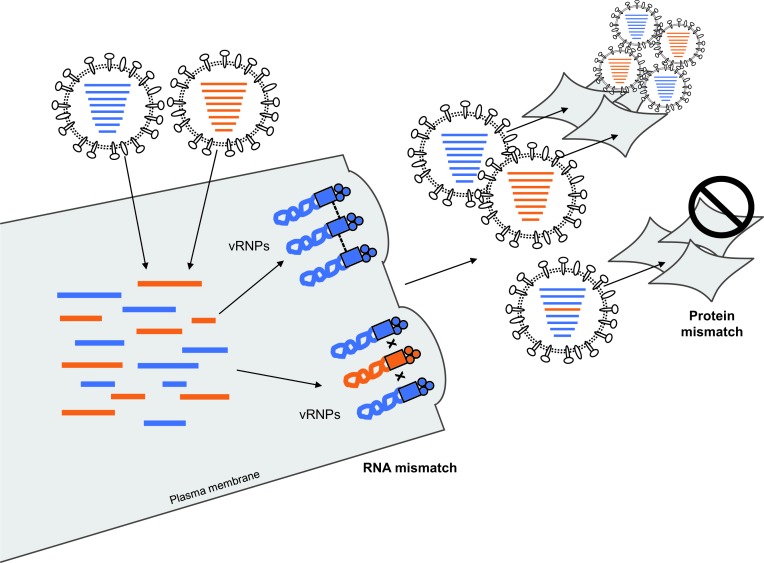
Similar to vRNAs, the proteins of the same virus strain co-evolve so that the interactions between them are optimal (Fig. 2a, b). During reassortment, progeny viruses can form that are not severely affected by packaging signal mismatch, but encode proteins that are not compatible (Fig. 2c). While RNA mismatch can limit the diversity of IAVs formed after a co-infection event, protein mismatch manifests only after the formation of progeny viruses, when the newly formed reassortants spread to new host cells (Fig. 3).
Fig. 3.
Manifestations of segment mismatch during the influenza A virus life cycle. When two different IAVs co-infect the same cell, their genomes mix, resulting in the genomic assembly of viral ribonucleoprotein complexes (vRNPs) from either the same virus (as shown in blue) or from different viruses (as shown in blue and orange). The interactions that occur between segments, most likely at the packaging signal regions (dotted lines between coloured rectangles), might be optimal when segments originate from the same parental virus, thus facilitating packaging. Conversely, these interactions might be suboptimal or non-existent (black x’s) between heterologous segments due to nucleotide differences, and thus formation of these progeny genotypes would be restricted due to RNA mismatch. Protein mismatch manifests later, after reassortant viruses have budded from the host cell and go on to infect new cells. Incompatibilities between polymerase subunits or HA-NA proteins, among others, can prevent the successful formation of progeny viruses, resulting in an abortive infection. Proteins and RNAs are shown with the same colour for clarity; however, in co-infected cells, proteins derived from co-infecting viruses are thought to mix.
PB2-PB1-PA-NP mismatch
Mismatch among the polymerase components is a common form of protein mismatch. Reassortment between influenza A/equine/Prague/1/56 (H7N7) [Prague] and influenza A/Yokohama/2017/03 (H3N2) [Yokohama] viruses was shown to be restricted by incompatibilities between the subunits comprising the vRNP complex (PB2, PB1, PA and NP) [131]. The combination of Prague PB2, Prague PB1 and Yokohama PA resulted in the inability of the three subunits to form the heterotrimer necessary for IAV transcription and replication. Compensatory mutations in PB2 and/or PA allowed for increased replicative activity, suggesting that low activity of chimeric RNP complexes can be a limiting factor in IAV reassortment. Hatta et al. made a similar observation using influenza A/Memphis/8/88 (H3N2) [Mem/88] and influenza A/Mallard/New York/6750/78 (H2N2) [Mal/NY] viruses. They were able to generate reassortant viruses containing either PB2/PB1/PA or PB2/PB1/PA/NP from Mem/88 in a Mal/NY background, but were unable to generate viruses containing only the PB1 and PA genes of Mem/88 in a Mal/NY background, demonstrating the incompatibility of polymerase subunits between these two virus strains [132].
Similar studies have been undertaken utilizing viruses of the pH1N1 lineage in combination with heterologous IAV strains. In two such studies, reassortant viral polymerase complexes were generated and polymerase activity was compared to that of the wild-type polymerase complex [133, 134]. Resulting data independently showed that (i) the combination of pH1N1 PB2 and seasonal H1N1 (sH1N1) PB1 significantly reduced polymerase activity, and that a compensatory mutation in the sH1N1 PB1 could help alleviate this mismatch; and (ii) replacing the pH1N1 PB2 or PB1 was less detrimental to polymerase activity than replacing the pH1N1 PA. Our own data on reassortment between pH1N1 and seasonal H3N2 viruses similarly revealed that replacement of the pH1N1 PA with that of the H3N2 virus was strongly disfavoured. In particular, mini-replicon studies and reassortment patterns indicated the pH1N1 PB2 required the cognate PA to support high polymerase activity and virus viability [69]. Co-assortment of PB2 and PA of pH1N1 was also noted by Schrauwen et al. [70]. Finally, an in-depth analysis of the heterologous PB2pH1N1:PAH3N2 pairing revealed that the defect lied in impaired replication-initiation relative to the full pH1N1 polymerase complex, and that this phenotype mapped largely to PAH3N2 residues 184N and 383N [135]. Overall, these data suggest that reassortment between various IAV lineages is limited by polymerase subunit mismatch.
HA–NA mismatch
Another common example of protein incompatibility is between the functions of the HA (receptor-binding) and NA (receptor-destroying) IAV glycoproteins. The function of these proteins must be balanced in order for incoming virus to attach to host cells and be successfully released at the end of the viral life cycle [136]. A substantial body of literature indicates that, when these proteins are in imbalance, viruses exhibit fitness defects. Some examples of this literature follow. Studies performed with influenza A/USSR/90/77 (H1N1) [USSR] showed that reassortant viruses containing the USSR NA gene and HA genes of H3, H4, H10 or H13 subtypes exhibited extensive virion aggregation, and that treatment with a bacterial neuraminidase was able to irreversibly dissociate these virion clumps [137, 138]. A subsequent study suggested that the USSR NA gene was essentially non-functional in both the reassortant and parental strains and, in the reassortant viruses, this functional defect could be compensated for by mutations in the HA segment which led to decreased sialic acid affinity [73]. In the same study discussed above in the context of polymerase mismatch, Hatta et al. were able to generate viruses containing the HA and NA genes from Mem/88 (H3N2) in a Mal/NY (H2N2) background, but were unable to generate viruses containing only the HA or the NA gene from Mem/88, demonstrating the importance of having a functional balance between HA and NA genes for successful IAV replication [132]. Castrucci and Kawaoka generated a mutant WSN (H1N1) virus that had a 24-amino acid deletion in the NA stalk, which rendered the virus unable to replicate in embryonated chicken eggs due to defective NA function [139]. However, passage of this mutant virus in eggs resulted in egg-adapted viruses which were able to replicate efficiently. Half of these adapted viruses displayed reduced affinity for sialic acid residues when compared to the parental virus, and analysis of the HA gene sequence of one of these viruses revealed mutations near the receptor binding site [140]. Thus, reduction in HA affinity was found to compensate for low NA activity. These findings underscore the importance of balanced HA–NA function for viral fitness and highlight the role of mutation in correcting functional imbalances.
NS–vRNP mismatch
A third example of protein mismatch involves gene products of the IAV NS segment in combination with the vRNP components. The NS segment encodes two proteins: the non-structural protein 1 (NS1) and the non-structural protein 2 (NS2). NS1 allows IAV to evade the host antiviral response by delaying the onset of interferon production [141]. NS2, also referred to as the nuclear export protein (NEP), interacts with nucleoporins in the host cell nuclear membrane, enabling IAV vRNPs to exit the nucleus [142]. In addition to these classical roles, it has also been shown that both NS proteins influence viral transcription and replication. For example, the addition of NS2 to vRNP reconstitution assays alters the relative accumulation of mRNA, cRNA and vRNA, resulting in a shift towards replication products (vRNA and cRNA) and away from transcription products (mRNA) [143]. Additionally, co-immunoprecipitation assays have revealed that NS1 interacts physically with vRNPs [144, 145], while genetic approaches suggest a role in the regulation of vRNA production [146]. NS1 also contributes to control of viral gene expression by enhancing the translation of viral proteins [147–149].
Using highly pathogenic avian influenza viruses, Wang et al. introduced NS segments from either influenza A/goose/Guangdong/1/96 (H5N1) [GD/1], influenza A/Vietnam/1203/2004 (H5N1) or influenza A/Mallard/NL/12/2000 (H7N3) viruses into an influenza A/FPV/Rostock/1934 (H7N1) background by reverse genetics. Cells infected with the virus containing the GD/1 NS segment accumulated more mRNA, cRNA and vRNA than cells infected with the parental H7N1 virus or the other NS reassortants [150], demonstrating how a heterologous NS segment can alter the efficiency of viral RNA synthesis. Subsequent RNP reconstitution assays suggested that the NS1 protein, in concert with other host or viral factors, was responsible for the observed differences in RNA accumulation.
NS1 can bind to cleavage and polyadenylation specificity factor 30 (CPSF30), a cellular factor involved in processing the 3’ ends of cellular pre-mRNAs [151–153]. The IAV polymerase complex (PB2, PB1, PA and NP) was shown to stabilize the binding of NS1 to CPSF30 and, furthermore, PA and NP were responsible for mediating this stability [154]. Interestingly, this stabilization could only be achieved with homologous NS1-polymerase components; the polymerase complex of influenza A/Udorn/307/72 (H3N2) virus was unable to stabilize a NS1-CPSF30 complex containing an influenza A/Hong Kong/483/97 (H5N1) virus NS1 protein, and vice versa [154, 155]. As successful binding of NS1 to CPSF30 is important for helping control the production of interferon by the host cell, mismatch between NS1 and specific vRNP components can lead to inadequate control of the host immune response in IAV-infected cells.
To examine the fitness of viruses encoding heterologous vRNP and NS proteins, Shelton et al. rescued viruses containing PB2/PB1/PA/NP from a pH1N1 virus and an NS segment from either pH1N1, PR8 (H1N1) or influenza A/England/612/2003 (H3N2) viruses. Experiments in A549 cells showed that the viruses containing either heterologous NS produced less vRNA at 7 and 24 h post-infection compared to the virus containing the homologous NS segment [156]. The observed differences in vRNA production could potentially be mediated through either NS1 or NEP function (or both). The authors suggest NS1, since the homologous NS1 lacks the ability to bind to CPSF30. While this inability to bind CPSF30 would lessen control of interferon production by the host cell, the authors argue it could also lead to an increase in polymerase function and therefore an increase in vRNA levels. Additionally, in mice, the virus with the homologous NS produced the most weight loss, the highest viral load and greatest interferon-β production. These results suggest that NS-vRNP mismatch can lessen virulence by reducing replicative fitness [156]. Taken together, mismatch between NS and vRNP components can alter essential viral processes, control of host immune responses and ultimately decrease the fitness of reassortant progeny.
Additional remarks
Owing to the large number of physical and functional interactions among viral proteins that are critical for maintaining viral fitness, and the diversity of amino acid sequences represented in circulating IAVs, protein mismatch is a complex phenomenon. Although much progress has been made in characterizing mismatch between heterologous viral proteins, results are often seen to depend on the parental strain pairing. While this reality makes predictions of protein compatibility for a given pair of parental strains extremely challenging, documentation of mismatch in reassortant viruses remains a useful tool for deciphering functional interactions occurring throughout the viral life cycle. In other words, where incompatibilities occur, interactions can be detected. Although the same mismatch may not be apparent with a different pairing of parental viruses, the functional interaction identified is likely to be broadly conserved.
Advantages of reassortment between heterologous IAVs
Although often constrained by segment mismatch, reassortment can also facilitate host adaptation, an increase in host range or evasion from pre-existing immunity [157, 158]. Most notably, reassortant viruses that derive a subset of gene segments from non-human hosts were seen to cause the last three pandemics. In this natural situation, post-reassortment adaptive changes were likely needed to achieve high fitness. In addition, however, multiple experimental studies using avian plus human, swine plus human or human plus human IAV strains have demonstrated that, even though some reassortant viruses displayed relatively low fitness, others exhibited high levels of genetic compatibility and increased pathogenicity or transmissibility when compared to the parental strains in model systems [70, 159–163].
Multiple risk assessment studies to date in the mouse model have demonstrated an increase in IAV virulence following reassortment. For example, incorporation of the PB1 gene from an avian H5N1 virus into a background of human H3N2 internal genes led to increased pathogenicity [92]. Based on these results, and the derivation of the PB1 segment from avian IAVs in the 1957 and 1968 pandemic strains, the authors postulated that acquisition of an avian PB1 gene by a human IAV might allow for the reassortant to outcompete currently circulating IAVs [20, 92]. In a panel of H5N1 subtype reassortants where each contained one internal H9N2 gene, viruses containing the H9N2 PB2 gene were attenuated in mice compared to the parental virus strains, while viruses containing the H9N2 PA or NP genes were 100 % lethal in mice [164]. Strikingly, none of the other viruses tested, including the parental strains, caused any mortality. Furthermore, the presence of the H9N2 PA or NP gene conferred a wider tissue tropism in mice and an increase in polymerase activity, while the presence of the H9N2 PB2 gene resulted in a decrease in polymerase activity. Ma et al. rescued a H7N1 virus containing a H5N1 NS gene and showed that the reassortant virus could replicate more effectively in cell culture than the wild-type H7N1 virus [165]. Additionally, the reassortant virus caused morbidity and mortality in mice, while the wild-type virus was unable to successfully replicate in mice. These data highlight how segment mismatch can limit the fitness of certain reassortant genotypes while other combinations of heterologous segments can confer an advantage in replication or tissue tropism.
Two additional reverse genetics-based risk assessment studies examined the fitness of pH1N1 and H5N1 reassortant viruses. Cline et al. demonstrated that a pH1N1 virus containing a H5N1 HA gene was able to replicate more efficiently than both the parental viruses and all other reassortant viruses tested in MDCK and A549 cells [166]. In addition, mice infected with the H5 HA-containing virus exhibited increased pulmonary disease compared to mice infected with either parental strain, but this virus failed to transmit in a ferret model. Generating 127 reassortant viruses between a pH1N1 isolate and a highly pathogenic H5N1 strain, Zhang et al. identified single gene reassortants that conferred aerosol transmission in a guinea pig model to the contact-transmissible H5N1 virus [167]. Both the pH1N1 PA and NS genes were individually found to allow for respiratory droplet transmission of H5N1 subtype reassortants in guinea pigs.
Taken together, these data offer examples of the wide array of effects a single reassortment event can have on factors such as virus growth, cell tropism, host range, immune evasion and transmissibility. While segment mismatch often hinders the assembly or subsequent spread of certain reassortant genotypes (Fig. 3), these constraints can sometimes be minimal, enabling the successful evolution of IAV under selective pressure. Importantly, the impact of segment mismatch on IAV reassortment is determined not only by the degree of RNA- and protein-based incompatibilities present between two co-infecting strains, but also by the surrounding host and environmental conditions, which define the selective environment.
Conclusions
When two divergent IAVs co-infect the same cell, the genetic diversity and fitness of reassortant progeny are limited by segment mismatch, or the incompatibilities between heterologous components of parental virus strains. While segment mismatch imposes barriers to reassortment, in some genetic constellations the detrimental effects associated with segment mismatch might be offset by enhancements in immune evasion or host adaptation of the reassortant virus. Due to the complexity of reassortant virus fitness, the models and experimental conditions used successfully for one study might prove suboptimal for the next. Further studies are therefore needed to more systematically document the underlying mechanisms of segment mismatch. A point of particular interest is the relative contributions of RNA and protein mismatch in determining the outcomes of reassortment between divergent IAV strains. This distinction is important because, once greater mechanistic understanding of IAV genome packaging is achieved, mapping of mismatches among RNA-based packaging signals may allow sequence-based prediction of compatible gene constellations. These studies will yield important contributions towards IAV pandemic preparedness and strengthen efforts to predict and control zoonotic outbreaks.
Funding information
We acknowledge funding support from the NIAID Centers of Excellence in Influenza Research and Surveillance (CEIRS), contract number HHSN272201400004C, and R01 grants AI125268 and AI099000.
Conflicts of interest
The authors declare that there are no conflicts of interest.
Footnotes
Abbreviations: CPSF30, cleavage and polyadenylation specificity factor 30; DI, defective interfering; HA, hemagglutinin; HRM, high resolution melt; IAV, influenza A virus; NA, neuraminidase; NEP, nuclear export protein; NP, nucleoprotein; NS1, non-structural protein 1; NS2, non-structural protein 2; pH1N1, 2009 H1N1 pandemic strain; UTR, untranslated region; VAR, variant; vRNAs, viral RNAs; vRNPs, viral ribonucleoprotein complexes; WT, wild-type.
References
- 1.Shaw ML, Palese P. Orthomyxoviridae. In: Knipe D, Howley P, editors. Fields Virology. Philadelphia, PA: Lippincott Williams & Wilkins; 2013. pp. 1151–1185. (editors) [Google Scholar]
- 2.Tong S, Li Y, Rivailler P, Conrardy C, Castillo DA, et al. A distinct lineage of influenza A virus from bats. Proc Natl Acad Sci USA. 2012;109:4269–4274. doi: 10.1073/pnas.1116200109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Tong S, Zhu X, Li Y, Shi M, Zhang J, et al. New world bats harbor diverse influenza A viruses. PLoS Pathog. 2013;9:e1003657. doi: 10.1371/journal.ppat.1003657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Munster VJ, Baas C, Lexmond P, Waldenström J, Wallensten A, et al. Spatial, temporal, and species variation in prevalence of influenza A viruses in wild migratory birds. PLoS Pathog. 2007;3:e61. doi: 10.1371/journal.ppat.0030061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Webster RG, Bean WJ, Gorman OT, Chambers TM, Kawaoka Y. Evolution and ecology of influenza A viruses. Microbiol Rev. 1992;56:152–179. doi: 10.1128/mr.56.1.152-179.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Holmes EC, Ghedin E, Miller N, Taylor J, Bao Y, et al. Whole-genome analysis of human influenza A virus reveals multiple persistent lineages and reassortment among recent H3N2 viruses. PLoS Biol. 2005;3:e300. doi: 10.1371/journal.pbio.0030300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Nelson MI, Simonsen L, Viboud C, Miller MA, Taylor J, et al. Stochastic processes are key determinants of short-term evolution in influenza a virus. PLoS Pathog. 2006;2:e125. doi: 10.1371/journal.ppat.0020125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Nelson MI, Edelman L, Spiro DJ, Boyne AR, Bera J, et al. Molecular epidemiology of A/H3N2 and A/H1N1 influenza virus during a single epidemic season in the United States. PLoS Pathog. 2008;4:e1000133. doi: 10.1371/journal.ppat.1000133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Westgeest KB, Russell CA, Lin X, Spronken MI, Bestebroer TM, et al. Genomewide analysis of reassortment and evolution of human influenza A(H3N2) viruses circulating between 1968 and 2011. J Virol. 2014;88:2844–2857. doi: 10.1128/JVI.02163-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Rambaut A, Pybus OG, Nelson MI, Viboud C, Taubenberger JK, et al. The genomic and epidemiological dynamics of human influenza A virus. Nature. 2008;453:615–619. doi: 10.1038/nature06945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Villa M, Lässig M. Fitness cost of reassortment in human influenza. PLoS Pathog. 2017;13:e1006685. doi: 10.1371/journal.ppat.1006685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Johnson NP, Mueller J. Updating the accounts: global mortality of the 1918-1920 "Spanish" influenza pandemic. Bull Hist Med. 2002;76:105–115. doi: 10.1353/bhm.2002.0022. [DOI] [PubMed] [Google Scholar]
- 13.Viboud C, Simonsen L, Fuentes R, Flores J, Miller MA, et al. Global mortality impact of the 1957-1959 influenza pandemic. J Infect Dis. 2016;213:738–745. doi: 10.1093/infdis/jiv534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Cockburn WC, Delon PJ, Ferreira W. Origin and progress of the 1968-69 Hong Kong influenza epidemic. Bull World Health Organ. 1969;41:345–348. [PMC free article] [PubMed] [Google Scholar]
- 15.Simonsen L, Spreeuwenberg P, Lustig R, Taylor RJ, Fleming DM, et al. Global mortality estimates for the 2009 Influenza Pandemic from the GLaMOR project: a modeling study. PLoS Med. 2013;10:e1001558. doi: 10.1371/journal.pmed.1001558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Worobey M, Han GZ, Rambaut A. Genesis and pathogenesis of the 1918 pandemic H1N1 influenza A virus. Proc Natl Acad Sci USA. 2014;111:8107–8112. doi: 10.1073/pnas.1324197111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Taubenberger JK, Reid AH, Lourens RM, Wang R, Jin G, et al. Characterization of the 1918 influenza virus polymerase genes. Nature. 2005;437:889–893. doi: 10.1038/nature04230. [DOI] [PubMed] [Google Scholar]
- 18.Gibbs MJ, Gibbs AJ. Molecular virology: was the 1918 pandemic caused by a bird flu? Nature. 2006;440:E8. doi: 10.1038/nature04823. [DOI] [PubMed] [Google Scholar]
- 19.Kilbourne ED. Influenza pandemics of the 20th century. Emerg Infect Dis. 2006;12:9–14. doi: 10.3201/eid1201.051254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kawaoka Y, Krauss S, Webster RG. Avian-to-human transmission of the PB1 gene of influenza A viruses in the 1957 and 1968 pandemics. J Virol. 1989;63:4603–4608. doi: 10.1128/jvi.63.11.4603-4608.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Garten RJ, Davis CT, Russell CA, Shu B, Lindstrom S, et al. Antigenic and genetic characteristics of swine-origin 2009 A(H1N1) influenza viruses circulating in humans. Science. 2009;325:197–201. doi: 10.1126/science.1176225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Smith GJ, Vijaykrishna D, Bahl J, Lycett SJ, Worobey M, et al. Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature. 2009;459:1122–1125. doi: 10.1038/nature08182. [DOI] [PubMed] [Google Scholar]
- 23.York I, Donis RO. The 2009 pandemic influenza virus: where did it come from, where is it now, and where is it going? Curr Top Microbiol Immunol. 2013;370:241–257. doi: 10.1007/82_2012_221. [DOI] [PubMed] [Google Scholar]
- 24.Ku AS, Chan LT. The first case of H5N1 avian influenza infection in a human with complications of adult respiratory distress syndrome and Reye's syndrome. J Paediatr Child Health. 1999;35:207–209. doi: 10.1046/j.1440-1754.1999.t01-1-00329.x. [DOI] [PubMed] [Google Scholar]
- 25.Chan PK. Outbreak of avian influenza A(H5N1) virus infection in Hong Kong in 1997. Clin Infect Dis. 2002;34 Suppl 2:S58–S64. doi: 10.1086/338820. [DOI] [PubMed] [Google Scholar]
- 26.Tran TH, Nguyen TL, Nguyen TD, Luong TS, Pham PM, et al. Avian influenza A (H5N1) in 10 patients in Vietnam. N Engl J Med. 2004;350:1179–1188. doi: 10.1056/NEJMoa040419. [DOI] [PubMed] [Google Scholar]
- 27.Ungchusak K, Auewarakul P, Dowell SF, Kitphati R, Auwanit W, et al. Probable person-to-person transmission of avian influenza A (H5N1) N Engl J Med. 2005;352:333–340. doi: 10.1056/NEJMoa044021. [DOI] [PubMed] [Google Scholar]
- 28.Gao R, Cao B, Hu Y, Feng Z, Wang D, et al. Human infection with a novel avian-origin influenza A (H7N9) virus. N Engl J Med. 2013;368:1888–1897. doi: 10.1056/NEJMoa1304459. [DOI] [PubMed] [Google Scholar]
- 29.Fouchier RA, Schneeberger PM, Rozendaal FW, Broekman JM, Kemink SA, et al. Avian influenza A virus (H7N7) associated with human conjunctivitis and a fatal case of acute respiratory distress syndrome. Proc Natl Acad Sci USA. 2004;101:1356–1361. doi: 10.1073/pnas.0308352100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.WHO Cumulative number of confirmed human cases for avian influenza A(H5N1) reported to WHO, 2003-2017. 2017. www.who.int/influenza/human_animal_interface/2017_09_27_tableH5N1.pdf?ua=1 [accessed 11 November 2017]
- 31.WHO Influenza at the human-animal interface. 2017. www.who.int/influenza/human_animal_interface/Influenza_Summary_IRA_HA_interface_09_27_2017.pdf?ua=1 [accessed 11 November 2017]
- 32.Lam TT, Wang J, Shen Y, Zhou B, Duan L, et al. The genesis and source of the H7N9 influenza viruses causing human infections in China. Nature. 2013;502:241–244. doi: 10.1038/nature12515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Wu A, Su C, Wang D, Peng Y, Liu M, et al. Sequential reassortments underlie diverse influenza H7N9 genotypes in China. Cell Host Microbe. 2013;14:446–452. doi: 10.1016/j.chom.2013.09.001. [DOI] [PubMed] [Google Scholar]
- 34.Li KS, Guan Y, Wang J, Smith GJ, Xu KM, et al. Genesis of a highly pathogenic and potentially pandemic H5N1 influenza virus in eastern Asia. Nature. 2004;430:209–213. doi: 10.1038/nature02746. [DOI] [PubMed] [Google Scholar]
- 35.Guan Y, Shortridge KF, Krauss S, Webster RG. Molecular characterization of H9N2 influenza viruses: were they the donors of the "internal" genes of H5N1 viruses in Hong Kong? Proc Natl Acad Sci USA. 1999;96:9363–9367. doi: 10.1073/pnas.96.16.9363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Lin YP, Shaw M, Gregory V, Cameron K, Lim W, et al. Avian-to-human transmission of H9N2 subtype influenza A viruses: relationship between H9N2 and H5N1 human isolates. Proc Natl Acad Sci USA. 2000;97:9654–9658. doi: 10.1073/pnas.160270697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Chen H, Yuan H, Gao R, Zhang J, Wang D, et al. Clinical and epidemiological characteristics of a fatal case of avian influenza A H10N8 virus infection: a descriptive study. Lancet. 2014;383:714–721. doi: 10.1016/S0140-6736(14)60111-2. [DOI] [PubMed] [Google Scholar]
- 38.Zhang Z, Li R, Jiang L, Xiong C, Chen Y, et al. The complexity of human infected AIV H5N6 isolated from China. BMC Infect Dis. 2016;16:600. doi: 10.1186/s12879-016-1932-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Huang Y, Li X, Zhang H, Chen B, Jiang Y, et al. Human infection with an avian influenza A (H9N2) virus in the middle region of China. J Med Virol. 2015;87:1641–1648. doi: 10.1002/jmv.24231. [DOI] [PubMed] [Google Scholar]
- 40.Yuan J, Zhang L, Kan X, Jiang L, Yang J, et al. Origin and molecular characteristics of a novel 2013 avian influenza A(H6N1) virus causing human infection in Taiwan. Clin Infect Dis. 2013;57:1367–1368. doi: 10.1093/cid/cit479. [DOI] [PubMed] [Google Scholar]
- 41.Lubeck MD, Palese P, Schulman JL. Nonrandom association of parental genes in influenza A virus recombinants. Virology. 1979;95:269–274. doi: 10.1016/0042-6822(79)90430-6. [DOI] [PubMed] [Google Scholar]
- 42.Greenbaum BD, Li OT, Poon LL, Levine AJ, Rabadan R. Viral reassortment as an information exchange between viral segments. Proc Natl Acad Sci USA. 2012;109:3341–3346. doi: 10.1073/pnas.1113300109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Pauly MD, Procario MC, Lauring AS. A novel twelve class fluctuation test reveals higher than expected mutation rates for influenza A viruses. eLife. 2017;6 doi: 10.7554/eLife.26437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Wrammert J, Koutsonanos D, Li GM, Edupuganti S, Sui J, et al. Broadly cross-reactive antibodies dominate the human B cell response against 2009 pandemic H1N1 influenza virus infection. J Exp Med. 2011;208:181–193. doi: 10.1084/jem.20101352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Ekiert DC, Bhabha G, Elsliger MA, Friesen RH, Jongeneelen M, et al. Antibody recognition of a highly conserved influenza virus epitope. Science. 2009;324:246–251. doi: 10.1126/science.1171491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Memoli MJ, Shaw PA, Han A, Czajkowski L, Reed S, et al. Evaluation of antihemagglutinin and antineuraminidase antibodies as correlates of protection in an influenza A/H1N1 virus healthy human challenge model. MBio. 2016;7:e00417-16. doi: 10.1128/mBio.00417-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Gerhard W, Yewdell J, Frankel ME, Webster R. Antigenic structure of influenza virus haemagglutinin defined by hybridoma antibodies. Nature. 1981;290:713–717. doi: 10.1038/290713a0. [DOI] [PubMed] [Google Scholar]
- 48.Plotkin JB, Dushoff J, Levin SA. Hemagglutinin sequence clusters and the antigenic evolution of influenza A virus. Proc Natl Acad Sci USA. 2002;99:6263–6268. doi: 10.1073/pnas.082110799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Smith DJ, Lapedes AS, de Jong JC, Bestebroer TM, Rimmelzwaan GF, et al. Mapping the antigenic and genetic evolution of influenza virus. Science. 2004;305:371–376. doi: 10.1126/science.1097211. [DOI] [PubMed] [Google Scholar]
- 50.Cox NJ, Brammer TL, Regnery HL. Influenza: global surveillance for epidemic and pandemic variants. Eur J Epidemiol. 1994;10:467–470. doi: 10.1007/BF01719678. [DOI] [PubMed] [Google Scholar]
- 51.Nelson MI, Viboud C, Simonsen L, Bennett RT, Griesemer SB, et al. Multiple reassortment events in the evolutionary history of H1N1 influenza A virus since 1918. PLoS Pathog. 2008;4:e1000012. doi: 10.1371/journal.ppat.1000012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Nelson MI, Holmes EC. The evolution of epidemic influenza. Nat Rev Genet. 2007;8:196–205. doi: 10.1038/nrg2053. [DOI] [PubMed] [Google Scholar]
- 53.Simonsen L, Viboud C, Grenfell BT, Dushoff J, Jennings L, et al. The genesis and spread of reassortment human influenza A/H3N2 viruses conferring adamantane resistance. Mol Biol Evol. 2007;24:1811–1820. doi: 10.1093/molbev/msm103. [DOI] [PubMed] [Google Scholar]
- 54.Nelson MI, Simonsen L, Viboud C, Miller MA, Holmes EC. The origin and global emergence of adamantane resistant A/H3N2 influenza viruses. Virology. 2009;388:270–278. doi: 10.1016/j.virol.2009.03.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Wright P, Neumann G, Kawaoka Y. Orthomyxoviruses. In: Knipe D, Howley P, editors. Fields Virology. Philadelphia, PA: Lippincott Williams & Wilkins; 2013. (editors) [Google Scholar]
- 56.Ellis JS, Alvarez-Aguero A, Gregory V, Lin YP, Hay A, et al. Influenza AH1N2 viruses, United Kingdom, 2001-02 influenza season. Emerg Infect Dis. 2003;9:304–310. doi: 10.3201/eid0903.020404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Chen MJ, La T, Zhao P, Tam JS, Rappaport R, et al. Genetic and phylogenetic analysis of multi-continent human influenza A(H1N2) reassortant viruses isolated in 2001 through 2003. Virus Res. 2006;122:200–205. doi: 10.1016/j.virusres.2006.07.010. [DOI] [PubMed] [Google Scholar]
- 58.Gregory V, Bennett M, Orkhan MH, Al Hajjar S, Varsano N, et al. Emergence of influenza A H1N2 reassortant viruses in the human population during 2001. Virology. 2002;300:1–7. doi: 10.1006/viro.2002.1513. [DOI] [PubMed] [Google Scholar]
- 59.Xu X, Smith CB, Mungall BA, Lindstrom SE, Hall HE, et al. Intercontinental circulation of human influenza A(H1N2) reassortant viruses during the 2001-2002 influenza season. J Infect Dis. 2002;186:1490–1493. doi: 10.1086/344738. [DOI] [PubMed] [Google Scholar]
- 60.Mukherjee TR, Agrawal AS, Chakrabarti S, Chawla-Sarkar M. Full genomic analysis of an influenza A (H1N2) virus identified during 2009 pandemic in Eastern India: evidence of reassortment event between co-circulating A(H1N1)pdm09 and A/Brisbane/10/2007-like H3N2 strains. Virol J. 2012;9:1–10. doi: 10.1186/1743-422X-9-233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Rith S, Chin S, Sar B, Phalla Y, Horm SV, et al. Natural co-infection of influenza A/H3N2 and A/H1N1pdm09 viruses resulting in a reassortant A/H3N2 virus. J Clin Virol. 2015;73:108–111. doi: 10.1016/j.jcv.2015.11.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Liu W, Li ZD, Tang F, Wei MT, Tong YG, et al. Mixed infections of pandemic H1N1 and seasonal H3N2 viruses in 1 outbreak. Clin Infect Dis. 2010;50:1359–1365. doi: 10.1086/652143. [DOI] [PubMed] [Google Scholar]
- 63.Myers CA, Kasper MR, Yasuda CY, Savuth C, Spiro DJ, et al. Dual infection of novel influenza viruses A/H1N1 and A/H3N2 in a cluster of Cambodian patients. Am J Trop Med Hyg. 2011;85:961–963. doi: 10.4269/ajtmh.2011.11-0098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Yamane N, Arikawa J, Odagiri T, Sukeno N, Ishida N. Isolation of three different influenza A viruses from an individual after probable double infection with H3N2 and H1N1 viruses. Jpn J Med Sci Biol. 1978;31:431–434. doi: 10.7883/yoken1952.31.431. [DOI] [PubMed] [Google Scholar]
- 65.Falchi A, Arena C, Andreoletti L, Jacques J, Leveque N, et al. Dual infections by influenza A/H3N2 and B viruses and by influenza A/H3N2 and A/H1N1 viruses during winter 2007, Corsica Island, France. J Clin Virol. 2008;41:148–151. doi: 10.1016/j.jcv.2007.11.003. [DOI] [PubMed] [Google Scholar]
- 66.Kendal AP, Lee DT, Parish HS, Raines D, Noble GR, et al. Laboratory-based surveillance of influenza virus in the United States during the winter of 1977-1978. II. Isolation of a mixture of A/Victoria- and A/USSR-like viruses from a single person during an epidemic in Wyoming, USA, January 1978. Am J Epidemiol. 1979;110:462–468. doi: 10.1093/oxfordjournals.aje.a112827. [DOI] [PubMed] [Google Scholar]
- 67.Frank AL, Taber LH, Wells JM. Individuals infected with two subtypes of influenza A virus in the same season. J Infect Dis. 1983;147:120–124. doi: 10.1093/infdis/147.1.120. [DOI] [PubMed] [Google Scholar]
- 68.Nishikawa F, Sugiyama T. Direct isolation of H1N2 recombinant virus from a throat swab of a patient simultaneously infected with H1N1 and H3N2 influenza A viruses. J Clin Microbiol. 1983;18:425–427. doi: 10.1128/jcm.18.2.425-427.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Phipps KL, Marshall N, Tao H, Danzy S, Onuoha N, et al. Seasonal H3N2 and 2009 pandemic H1N1 influenza A viruses reassort efficiently but produce attenuated progeny. J Virol. 2017;91:e00830-17. doi: 10.1128/JVI.00830-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Schrauwen EJ, Herfst S, Chutinimitkul S, Bestebroer TM, Rimmelzwaan GF, et al. Possible increased pathogenicity of pandemic (H1N1) 2009 influenza virus upon reassortment. Emerg Infect Dis. 2011;17:200–208. doi: 10.3201/eid1702.101268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Mena I, Nelson MI, Quezada-Monroy F, Dutta J, Cortes-Fernández R, et al. Origins of the 2009 H1N1 influenza pandemic in swine in Mexico. eLife. 2016;5 doi: 10.7554/eLife.16777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Ma W, Liu Q, Qiao C, del Real G, García-Sastre A, et al. North American triple reassortant and Eurasian H1N1 swine influenza viruses do not readily reassort to generate a 2009 pandemic H1N1-like virus. mBio. 2014;5:e00919-13. doi: 10.1128/mBio.00919-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Kaverin NV, Gambaryan AS, Bovin NV, Rudneva IA, Shilov AA, et al. Postreassortment changes in influenza A virus hemagglutinin restoring HA-NA functional match. Virology. 1998;244:315–321. doi: 10.1006/viro.1998.9119. [DOI] [PubMed] [Google Scholar]
- 74.Ilyushina NA, Rudneva IA, Shilov AA, Klenk HD, Kaverin NV. Postreassortment changes in a model system: HA-NA adjustment in an H3N2 avian-human reassortant influenza virus. Arch Virol. 2005;150:1327–1338. doi: 10.1007/s00705-005-0490-4. [DOI] [PubMed] [Google Scholar]
- 75.Neverov AD, Lezhnina KV, Kondrashov AS, Bazykin GA. Intrasubtype reassortments cause adaptive amino acid replacements in H3N2 influenza genes. PLoS Genet. 2014;10:e1004037. doi: 10.1371/journal.pgen.1004037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Steel J, Lowen AC. Influenza A virus reassortment. Curr Top Microbiol Immunol. 2014;385:377–401. doi: 10.1007/82_2014_395. [DOI] [PubMed] [Google Scholar]
- 77.Marshall N, Priyamvada L, Ende Z, Steel J, Lowen AC. Influenza virus reassortment occurs with high frequency in the absence of segment mismatch. PLoS Pathog. 2013;9:e1003421. doi: 10.1371/journal.ppat.1003421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Wittwer CT, Reed GH, Gundry CN, Vandersteen JG, Pryor RJ. High-resolution genotyping by amplicon melting analysis using LCGreen. Clin Chem. 2003;49:853–860. doi: 10.1373/49.6.853. [DOI] [PubMed] [Google Scholar]
- 79.Fonville JM, Marshall N, Tao H, Steel J, Lowen AC. Influenza virus reassortment is enhanced by semi-infectious particles but can be suppressed by defective interfering particles. PLoS Pathog. 2015;11:e1005204. doi: 10.1371/journal.ppat.1005204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Huang IC, Li W, Sui J, Marasco W, Choe H, et al. Influenza A virus neuraminidase limits viral superinfection. J Virol. 2008;82:4834–4843. doi: 10.1128/JVI.00079-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Tao H, Steel J, Lowen AC. Intrahost dynamics of influenza virus reassortment. J Virol. 2014;88:7485–7492. doi: 10.1128/JVI.00715-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Tao H, Li L, White MC, Steel J, Lowen AC. Influenza A virus coinfection through transmission can support high levels of reassortment. J Virol. 2015;89:8453–8461. doi: 10.1128/JVI.01162-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Wille M, Tolf C, Avril A, Latorre-Margalef N, Wallerström S, et al. Frequency and patterns of reassortment in natural influenza A virus infection in a reservoir host. Virology. 2013;443:150–160. doi: 10.1016/j.virol.2013.05.004. [DOI] [PubMed] [Google Scholar]
- 84.Deng G, Tan D, Shi J, Cui P, Jiang Y, et al. Complex reassortment of multiple subtypes of avian influenza viruses in domestic ducks at the Dongting Lake Region of China. J Virol. 2013;87:9452–9462. doi: 10.1128/JVI.00776-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Abolnik C, Gerdes GH, Sinclair M, Ganzevoort BW, Kitching JP, et al. Phylogenetic analysis of influenza A viruses (H6N8, H1N8, H4N2, H9N2, H10N7) isolated from wild birds, ducks, and ostriches in South Africa from 2007 to 2009. Avian Dis. 2010;54:313–322. doi: 10.1637/8781-040109-Reg.1. [DOI] [PubMed] [Google Scholar]
- 86.Dugan VG, Chen R, Spiro DJ, Sengamalay N, Zaborsky J, et al. The evolutionary genetics and emergence of avian influenza viruses in wild birds. PLoS Pathog. 2008;4:e1000076. doi: 10.1371/journal.ppat.1000076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Nelson MI, Detmer SE, Wentworth DE, Tan Y, Schwartzbard A, et al. Genomic reassortment of influenza A virus in North American swine, 1998-2011. J Gen Virol. 2012;93:2584–2589. doi: 10.1099/vir.0.045930-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Vijaykrishna D, Poon LL, Zhu HC, Ma SK, Li OT, et al. Reassortment of pandemic H1N1/2009 influenza A virus in swine. Science. 2010;328:1529. doi: 10.1126/science.1189132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Rabadan R, Levine AJ, Krasnitz M. Non-random reassortment in human influenza A viruses. Influenza Other Respir Viruses. 2008;2:9–22. doi: 10.1111/j.1750-2659.2007.00030.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Maines TR, Chen LM, Matsuoka Y, Chen H, Rowe T, et al. Lack of transmission of H5N1 avian-human reassortant influenza viruses in a ferret model. Proc Natl Acad Sci USA. 2006;103:12121–12126. doi: 10.1073/pnas.0605134103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Jackson S, van Hoeven N, Chen LM, Maines TR, Cox NJ, et al. Reassortment between avian H5N1 and human H3N2 influenza viruses in ferrets: a public health risk assessment. J Virol. 2009;83:8131–8140. doi: 10.1128/JVI.00534-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Chen LM, Davis CT, Zhou H, Cox NJ, Donis RO. Genetic compatibility and virulence of reassortants derived from contemporary avian H5N1 and human H3N2 influenza A viruses. PLoS Pathog. 2008;4:e1000072. doi: 10.1371/journal.ppat.1000072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Schrauwen EJ, Bestebroer TM, Rimmelzwaan GF, Osterhaus AD, Fouchier RA, et al. Reassortment between Avian H5N1 and human influenza viruses is mainly restricted to the matrix and neuraminidase gene segments. PLoS One. 2013;8:e59889. doi: 10.1371/journal.pone.0059889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Kimble JB, Sorrell E, Shao H, Martin PL, Perez DR. Compatibility of H9N2 avian influenza surface genes and 2009 pandemic H1N1 internal genes for transmission in the ferret model. Proc Natl Acad Sci USA. 2011;108:12084–12088. doi: 10.1073/pnas.1108058108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Dlugolenski D, Jones L, Howerth E, Wentworth D, Tompkins SM, et al. Swine influenza virus PA and neuraminidase gene reassortment into human H1N1 influenza virus is associated with an altered pathogenic phenotype linked to increased MIP-2 expression. J Virol. 2015;89:5651–5667. doi: 10.1128/JVI.00087-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Ma J, Shen H, Liu Q, Bawa B, Qi W, et al. Pathogenicity and transmissibility of novel reassortant H3N2 influenza viruses with 2009 pandemic H1N1 genes in pigs. J Virol. 2015;89:2831–2841. doi: 10.1128/JVI.03355-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Enami M, Sharma G, Benham C, Palese P. An influenza virus containing nine different RNA segments. Virology. 1991;185:291–298. doi: 10.1016/0042-6822(91)90776-8. [DOI] [PubMed] [Google Scholar]
- 98.Donald HB, Isaacs A. Counts of influenza virus particles. J Gen Microbiol. 1954;10:457–464. doi: 10.1099/00221287-10-3-457. [DOI] [PubMed] [Google Scholar]
- 99.Mclain L, Armstrong SJ, Dimmock NJ. One defective interfering particle per cell prevents influenza virus-mediated cytopathology: an efficient assay system. J Gen Virol. 1988;69:1415–1419. doi: 10.1099/0022-1317-69-6-1415. [DOI] [PubMed] [Google Scholar]
- 100.Isaacs A, Donald HB. Particle counts of haemagglutinating viruses. J Gen Microbiol. 1955;12:241–247. doi: 10.1099/00221287-12-2-241. [DOI] [PubMed] [Google Scholar]
- 101.Bancroft CT, Parslow TG. Evidence for segment-nonspecific packaging of the influenza a virus genome. J Virol. 2002;76:7133–7139. doi: 10.1128/JVI.76.14.7133-7139.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Gerber M, Isel C, Moules V, Marquet R. Selective packaging of the influenza A genome and consequences for genetic reassortment. Trends Microbiol. 2014;22:446–455. doi: 10.1016/j.tim.2014.04.001. [DOI] [PubMed] [Google Scholar]
- 103.Mcgeoch D, Fellner P, Newton C. Influenza virus genome consists of eight distinct RNA species. Proc Natl Acad Sci USA. 1976;73:3045–3049. doi: 10.1073/pnas.73.9.3045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Nakajima K, Sugiura A. Three-factor cross of influenza virus. Virology. 1977;81:486–489. doi: 10.1016/0042-6822(77)90165-9. [DOI] [PubMed] [Google Scholar]
- 105.Bergmann M, Muster T. The relative amount of an influenza A virus segment present in the viral particle is not affected by a reduction in replication of that segment. J Gen Virol. 1995;76:3211–3215. doi: 10.1099/0022-1317-76-12-3211. [DOI] [PubMed] [Google Scholar]
- 106.Smith GL, Hay AJ. Replication of the influenza virus genome. Virology. 1982;118:96–108. doi: 10.1016/0042-6822(82)90323-3. [DOI] [PubMed] [Google Scholar]
- 107.Chou YY, Vafabakhsh R, Doğanay S, Gao Q, Ha T, et al. One influenza virus particle packages eight unique viral RNAs as shown by FISH analysis. Proc Natl Acad Sci USA. 2012;109:9101–9106. doi: 10.1073/pnas.1206069109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Inagaki A, Goto H, Kakugawa S, Ozawa M, Kawaoka Y. Competitive incorporation of homologous gene segments of influenza A virus into virions. J Virol. 2012;86:10200–10202. doi: 10.1128/JVI.01204-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Duhaut SD, Dimmock NJ. Defective segment 1 RNAs that interfere with production of infectious influenza A virus require at least 150 nucleotides of 5' sequence: evidence from a plasmid-driven system. J Gen Virol. 2002;83:403–411. doi: 10.1099/0022-1317-83-2-403. [DOI] [PubMed] [Google Scholar]
- 110.Duhaut SD, Mccauley JW. Defective RNAs inhibit the assembly of influenza virus genome segments in a segment-specific manner. Virology. 1996;216:326–337. doi: 10.1006/viro.1996.0068. [DOI] [PubMed] [Google Scholar]
- 111.Odagiri T, Tashiro M. Segment-specific noncoding sequences of the influenza virus genome RNA are involved in the specific competition between defective interfering RNA and its progenitor RNA segment at the virion assembly step. J Virol. 1997;71:2138–2145. doi: 10.1128/jvi.71.3.2138-2145.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Odagiri T, Tominaga K, Tobita K, Ohta S. An amino acid change in the non-structural NS2 protein of an influenza A virus mutant is responsible for the generation of defective interfering (DI) particles by amplifying DI RNAs and suppressing complementary RNA synthesis. J Gen Virol. 1994;75:43–53. doi: 10.1099/0022-1317-75-1-43. [DOI] [PubMed] [Google Scholar]
- 113.Fujii Y, Goto H, Watanabe T, Yoshida T, Kawaoka Y. Selective incorporation of influenza virus RNA segments into virions. Proc Natl Acad Sci USA. 2003;100:2002–2007. doi: 10.1073/pnas.0437772100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Liang Y, Hong Y, Parslow TG. cis-Acting packaging signals in the influenza virus PB1, PB2, and PA genomic RNA segments. J Virol. 2005;79:10348–10355. doi: 10.1128/JVI.79.16.10348-10355.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Watanabe T, Watanabe S, Noda T, Fujii Y, Kawaoka Y. Exploitation of nucleic acid packaging signals to generate a novel influenza virus-based vector stably expressing two foreign genes. J Virol. 2003;77:10575–10583. doi: 10.1128/JVI.77.19.10575-10583.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Ozawa M, Fujii K, Muramoto Y, Yamada S, Yamayoshi S, et al. Contributions of two nuclear localization signals of influenza A virus nucleoprotein to viral replication. J Virol. 2007;81:30–41. doi: 10.1128/JVI.01434-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Ozawa M, Maeda J, Iwatsuki-Horimoto K, Watanabe S, Goto H, et al. Nucleotide sequence requirements at the 5' end of the influenza A virus M RNA segment for efficient virus replication. J Virol. 2009;83:3384–3388. doi: 10.1128/JVI.02513-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Fujii K, Fujii Y, Noda T, Muramoto Y, Watanabe T, et al. Importance of both the coding and the segment-specific noncoding regions of the influenza A virus NS segment for its efficient incorporation into virions. J Virol. 2005;79:3766–3774. doi: 10.1128/JVI.79.6.3766-3774.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Cobbin JC, Ong C, Verity E, Gilbertson BP, Rockman SP, et al. Influenza virus PB1 and neuraminidase gene segments can cosegregate during vaccine reassortment driven by interactions in the PB1 coding region. J Virol. 2014;88:8971–8980. doi: 10.1128/JVI.01022-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Gilbertson B, Zheng T, Gerber M, Printz-Schweigert A, Ong C, et al. Influenza NA and PB1 gene segments interact during the formation of viral progeny: localization of the binding region within the PB1 Gene. Viruses. 2016;8:238. doi: 10.3390/v8080238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Gavazzi C, Yver M, Isel C, Smyth RP, Rosa-Calatrava M, et al. A functional sequence-specific interaction between influenza A virus genomic RNA segments. Proc Natl Acad Sci USA. 2013;110:16604–16609. doi: 10.1073/pnas.1314419110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Moreira ÉA, Weber A, Bolte H, Kolesnikova L, Giese S, et al. A conserved influenza A virus nucleoprotein code controls specific viral genome packaging. Nat Commun. 2016;7:12861. doi: 10.1038/ncomms12861. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Noda T, Sagara H, Yen A, Takada A, Kida H, et al. Architecture of ribonucleoprotein complexes in influenza A virus particles. Nature. 2006;439:490–492. doi: 10.1038/nature04378. [DOI] [PubMed] [Google Scholar]
- 124.Nakatsu S, Sagara H, Sakai-Tagawa Y, Sugaya N, Noda T, et al. Complete and Incomplete genome packaging of influenza A and B viruses. mBio. 2016;7:e01248-16. doi: 10.1128/mBio.01248-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Noda T, Sugita Y, Aoyama K, Hirase A, Kawakami E, et al. Three-dimensional analysis of ribonucleoprotein complexes in influenza A virus. Nat Commun. 2012;3:639. doi: 10.1038/ncomms1647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Fournier E, Moules V, Essere B, Paillart JC, Sirbat JD, et al. Interaction network linking the human H3N2 influenza A virus genomic RNA segments. Vaccine. 2012;30:7359–7367. doi: 10.1016/j.vaccine.2012.09.079. [DOI] [PubMed] [Google Scholar]
- 127.White MC, Steel J, Lowen AC. Heterologous packaging signals on segment 4, but not segment 6 or segment 8, limit influenza A virus reassortment. J Virol. 2017;91:e00195-17. doi: 10.1128/JVI.00195-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Baker SF, Nogales A, Finch C, Tuffy KM, Domm W, et al. Influenza A and B virus intertypic reassortment through compatible viral packaging signals. J Virol. 2014;88:10778–10791. doi: 10.1128/JVI.01440-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Essere B, Yver M, Gavazzi C, Terrier O, Isel C, et al. Critical role of segment-specific packaging signals in genetic reassortment of influenza A viruses. Proc Natl Acad Sci USA. 2013;110:E3840. doi: 10.1073/pnas.1308649110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Gao Q, Palese P. Rewiring the RNAs of influenza virus to prevent reassortment. Proc Natl Acad Sci USA. 2009;106:15891–15896. doi: 10.1073/pnas.0908897106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Li C, Hatta M, Watanabe S, Neumann G, Kawaoka Y. Compatibility among polymerase subunit proteins is a restricting factor in reassortment between equine H7N7 and human H3N2 influenza viruses. J Virol. 2008;82:11880–11888. doi: 10.1128/JVI.01445-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Hatta M, Halfmann P, Wells K, Kawaoka Y. Human influenza a viral genes responsible for the restriction of its replication in duck intestine. Virology. 2002;295:250–255. doi: 10.1006/viro.2002.1358. [DOI] [PubMed] [Google Scholar]
- 133.Song MS, Pascua PN, Lee JH, Baek YH, Park KJ, et al. Virulence and genetic compatibility of polymerase reassortant viruses derived from the pandemic (H1N1) 2009 influenza virus and circulating influenza A viruses. J Virol. 2011;85:6275–6286. doi: 10.1128/JVI.02125-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Octaviani CP, Goto H, Kawaoka Y. Reassortment between seasonal H1N1 and pandemic (H1N1) 2009 influenza viruses is restricted by limited compatibility among polymerase subunits. J Virol. 2011;85:8449–8452. doi: 10.1128/JVI.05054-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Hara K, Nakazono Y, Kashiwagi T, Hamada N, Watanabe H. Co-incorporation of the PB2 and PA polymerase subunits from human H3N2 influenza virus is a critical determinant of the replication of reassortant ribonucleoprotein complexes. J Gen Virol. 2013;94:2406–2416. doi: 10.1099/vir.0.053959-0. [DOI] [PubMed] [Google Scholar]
- 136.Wagner R, Matrosovich M, Klenk HD. Functional balance between haemagglutinin and neuraminidase in influenza virus infections. Rev Med Virol. 2002;12:159–166. doi: 10.1002/rmv.352. [DOI] [PubMed] [Google Scholar]
- 137.Rudneva IA, Kovaleva VP, Varich NL, Farashyan VR, Gubareva LV, et al. Influenza A virus reassortants with surface glycoprotein genes of the avian parent viruses: effects of HA and NA gene combinations on virus aggregation. Arch Virol. 1993;133:437–450. doi: 10.1007/BF01313781. [DOI] [PubMed] [Google Scholar]
- 138.Rudneva IA, Sklyanskaya EI, Barulina OS, Yamnikova SS, Kovaleva VP, et al. Phenotypic expression of HA-NA combinations in human-avian influenza A virus reassortants. Arch Virol. 1996;141:1091–1099. doi: 10.1007/BF01718612. [DOI] [PubMed] [Google Scholar]
- 139.Castrucci MR, Kawaoka Y. Biologic importance of neuraminidase stalk length in influenza A virus. J Virol. 1993;67:759–764. doi: 10.1128/jvi.67.2.759-764.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Mitnaul LJ, Matrosovich MN, Castrucci MR, Tuzikov AB, Bovin NV, et al. Balanced hemagglutinin and neuraminidase activities are critical for efficient replication of influenza A virus. J Virol. 2000;74:6015–6020. doi: 10.1128/JVI.74.13.6015-6020.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.García-Sastre A, Egorov A, Matassov D, Brandt S, Levy DE, et al. Influenza A virus lacking the NS1 gene replicates in interferon-deficient systems. Virology. 1998;252:324–330. doi: 10.1006/viro.1998.9508. [DOI] [PubMed] [Google Scholar]
- 142.O'Neill RE, Talon J, Palese P. The influenza virus NEP (NS2 protein) mediates the nuclear export of viral ribonucleoproteins. Embo J. 1998;17:288–296. doi: 10.1093/emboj/17.1.288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Robb NC, Smith M, Vreede FT, Fodor E. NS2/NEP protein regulates transcription and replication of the influenza virus RNA genome. J Gen Virol. 2009;90:1398–1407. doi: 10.1099/vir.0.009639-0. [DOI] [PubMed] [Google Scholar]
- 144.Marión RM, Zürcher T, de La Luna S, Ortín J. Influenza virus NS1 protein interacts with viral transcription-replication complexes in vivo. J Gen Virol. 1997;78:2447–2451. doi: 10.1099/0022-1317-78-10-2447. [DOI] [PubMed] [Google Scholar]
- 145.Robb NC, Chase G, Bier K, Vreede FT, Shaw PC, et al. The influenza A virus NS1 protein interacts with the nucleoprotein of viral ribonucleoprotein complexes. J Virol. 2011;85:5228–5231. doi: 10.1128/JVI.02562-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Min JY, Li S, Sen GC, Krug RM. A site on the influenza A virus NS1 protein mediates both inhibition of PKR activation and temporal regulation of viral RNA synthesis. Virology. 2007;363:236–243. doi: 10.1016/j.virol.2007.01.038. [DOI] [PubMed] [Google Scholar]
- 147.de La Luna S, Fortes P, Beloso A, Ortín J. Influenza virus NS1 protein enhances the rate of translation initiation of viral mRNAs. J Virol. 1995;69:2427–2433. doi: 10.1128/jvi.69.4.2427-2433.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Enami K, Sato TA, Nakada S, Enami M. Influenza virus NS1 protein stimulates translation of the M1 protein. J Virol. 1994;68:1432–1437. doi: 10.1128/jvi.68.3.1432-1437.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Falcón AM, Marión RM, Zürcher T, Gómez P, Portela A, et al. Defective RNA replication and late gene expression in temperature-sensitive influenza viruses expressing deleted forms of the NS1 protein. J Virol. 2004;78:3880–3888. doi: 10.1128/JVI.78.8.3880-3888.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Wang Z, Robb NC, Lenz E, Wolff T, Fodor E, et al. NS reassortment of an H7-type highly pathogenic avian influenza virus affects its propagation by altering the regulation of viral RNA production and antiviral host response. J Virol. 2010;84:11323–11335. doi: 10.1128/JVI.01034-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151.Nemeroff ME, Barabino SM, Li Y, Keller W, Krug RM. Influenza virus NS1 protein interacts with the cellular 30 kDa subunit of CPSF and inhibits 3'end formation of cellular pre-mRNAs. Mol Cell. 1998;1:991–1000. doi: 10.1016/S1097-2765(00)80099-4. [DOI] [PubMed] [Google Scholar]
- 152.Hale BG, Steel J, Medina RA, Manicassamy B, Ye J, et al. Inefficient control of host gene expression by the 2009 pandemic H1N1 influenza A virus NS1 protein. J Virol. 2010;84:6909–6922. doi: 10.1128/JVI.00081-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153.Ayllon J, Domingues P, Rajsbaum R, Miorin L, Schmolke M, et al. A single amino acid substitution in the novel H7N9 influenza A virus NS1 protein increases CPSF30 binding and virulence. J Virol. 2014;88:12146–12151. doi: 10.1128/JVI.01567-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154.Kuo RL, Krug RM. Influenza a virus polymerase is an integral component of the CPSF30-NS1A protein complex in infected cells. J Virol. 2009;83:1611–1616. doi: 10.1128/JVI.01491-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Twu KY, Kuo RL, Marklund J, Krug RM. The H5N1 influenza virus NS genes selected after 1998 enhance virus replication in mammalian cells. J Virol. 2007;81:8112–8121. doi: 10.1128/JVI.00006-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Shelton H, Smith M, Hartgroves L, Stilwell P, Roberts K, et al. An influenza reassortant with polymerase of pH1N1 and NS gene of H3N2 influenza A virus is attenuated in vivo. J Gen Virol. 2012;93:998–1006. doi: 10.1099/vir.0.039701-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157.Ince WL, Gueye-Mbaye A, Bennink JR, Yewdell JW. Reassortment complements spontaneous mutation in influenza A virus NP and M1 genes to accelerate adaptation to a new host. J Virol. 2013;87:4330–4338. doi: 10.1128/JVI.02749-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Ma EJ, Hill NJ, Zabilansky J, Yuan K, Runstadler JA. Reticulate evolution is favored in influenza niche switching. Proc Natl Acad Sci USA. 2016;113:5335–5339. doi: 10.1073/pnas.1522921113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Li C, Hatta M, Nidom CA, Muramoto Y, Watanabe S, et al. Reassortment between avian H5N1 and human H3N2 influenza viruses creates hybrid viruses with substantial virulence. Proc Natl Acad Sci USA. 2010;107:4687–4692. doi: 10.1073/pnas.0912807107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160.Octaviani CP, Ozawa M, Yamada S, Goto H, Kawaoka Y. High level of genetic compatibility between swine-origin H1N1 and highly pathogenic avian H5N1 influenza viruses. J Virol. 2010;84:10918–10922. doi: 10.1128/JVI.01140-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Stincarelli M, Arvia R, de Marco MA, Clausi V, Corcioli F, et al. Reassortment ability of the 2009 pandemic H1N1 influenza virus with circulating human and avian influenza viruses: public health risk implications. Virus Res. 2013;175:151–154. doi: 10.1016/j.virusres.2013.04.012. [DOI] [PubMed] [Google Scholar]
- 162.Octaviani CP, Li C, Noda T, Kawaoka Y. Reassortment between seasonal and swine-origin H1N1 influenza viruses generates viruses with enhanced growth capability in cell culture. Virus Res. 2011;156:147–150. doi: 10.1016/j.virusres.2010.12.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Fabrizio TP, Sun Y, Yoon SW, Jeevan T, Dlugolenski D, et al. Virologic differences do not fully explain the diversification of swine influenza viruses in the United States. J Virol. 2016;90:10074–10082. doi: 10.1128/JVI.01218-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Hao X, Hu J, Wang J, Xu J, Cheng H, et al. Reassortant H5N1 avian influenza viruses containing PA or NP gene from an H9N2 virus significantly increase the pathogenicity in mice. Vet Microbiol. 2016;192:95–101. doi: 10.1016/j.vetmic.2016.07.002. [DOI] [PubMed] [Google Scholar]
- 165.Ma W, Brenner D, Wang Z, Dauber B, Ehrhardt C, et al. The NS segment of an H5N1 highly pathogenic avian influenza virus (HPAIV) is sufficient to alter replication efficiency, cell tropism, and host range of an H7N1 HPAIV. J Virol. 2010;84:2122–2133. doi: 10.1128/JVI.01668-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166.Cline TD, Karlsson EA, Freiden P, Seufzer BJ, Rehg JE, et al. Increased pathogenicity of a reassortant 2009 pandemic H1N1 influenza virus containing an H5N1 hemagglutinin. J Virol. 2011;85:12262–12270. doi: 10.1128/JVI.05582-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167.Zhang Y, Zhang Q, Kong H, Jiang Y, Gao Y, et al. H5N1 hybrid viruses bearing 2009/H1N1 virus genes transmit in guinea pigs by respiratory droplet. Science. 2013;340:1459–1463. doi: 10.1126/science.1229455. [DOI] [PubMed] [Google Scholar]