Comparative transcriptomics and targeted analyses provide insights into the roles of phytohormone synthesis, signaling, and turnover during submergence-induced internode elongation in deepwater rice.
Abstract
Water submergence is an environmental factor that limits plant growth and survival. Deepwater rice (Oryza sativa) adapts to submergence by rapidly elongating its internodes and thereby maintaining its leaves above the water surface. We performed a comparative RNA sequencing transcriptome analysis of the shoot base region, including basal nodes, internodes, and shoot apices of seedlings at two developmental stages from two varieties with contrasting deepwater growth responses. A transcriptomic comparison between deepwater rice cv C9285 and nondeepwater rice cv Taichung 65 revealed both similar and differential expression patterns between the two genotypes during submergence. The expression of genes related to gibberellin biosynthesis, trehalose biosynthesis, anaerobic fermentation, cell wall modification, and transcription factors that include ethylene-responsive factors was significantly different between the varieties. Interestingly, in both varieties, the jasmonic acid content at the shoot base decreased during submergence, while exogenous jasmonic acid inhibited submergence-induced internode elongation in cv C9285, suggesting that jasmonic acid plays a role in the submergence response of rice. Furthermore, a targeted de novo transcript assembly revealed transcripts that were specific to cv C9285, including submergence-induced biotic stress-related genes. Our multifaceted transcriptome approach using the rice shoot base region illustrates a differential response to submergence between deepwater and nondeepwater rice. Jasmonic acid metabolism appears to participate in the submergence-mediated internode elongation response of deepwater rice.
Submergence stress is harmful to plants. In addition to causing oxygen- and CO2-deficient conditions by restricting environmental gas exchange, submergence reduces the light available for photosynthesis, perturbs cellular energy generation, and disrupts ionic balance (Bailey-Serres and Voesenek, 2008; Voesenek and Bailey-Serres, 2015). The mechanism of the submergence response has been well studied using rice (Oryza sativa), two diverging Rumex spp. (Rumex acetosa and Rumex palustris), and Arabidopsis (Arabidopsis thaliana), and several global analyses using transcriptomic or metabolomic approaches to examine submergence or hypoxia/anoxia stress have been reported (Lasanthi-Kudahettige et al., 2007; Magneschi and Perata, 2009; Mustroph et al., 2009, 2010; Narsai et al., 2009, 2015; Lakshmanan et al., 2013; van Veen et al., 2013; Rivera-Contreras et al., 2016).
Rice is the most important staple crop in Asia, and water availability is a crucial factor for rice cultivation. In tropical southeast Asia, rice is produced in paddy fields with water-controlling irrigation systems using rivers, lakes, ponds, and swamps. However, in some parts of south and southeast Asia, such as Bangladesh, India, Thailand, Vietnam, and Cambodia, the paddy fields are frequently submerged during the rainy season. The general cultivated rice cannot survive in these submergence-prone areas, but some cultivars, such as floating or deepwater rice, can grow and survive in such conditions even under several-months-long periods of deep flooding.
The adaptation of plants to submergence stress involves two different opposing mechanisms, the quiescence strategy (e.g. rice SUBMERGENCE1 [SUB1] varieties, Arabidopsis ecotypes, and R. acetosa) and the escape strategy (e.g. deepwater/floating rice varieties and R. palustris) (Jackson, 2008; Bailey-Serres et al., 2012; Loreti et al., 2016). Submergence-tolerant rice such as cv Flood Resistant 13A survives flooded conditions for a few weeks through the quiescence strategy, wherein the plants temporarily cease shoot elongation to conserve energy and then resume growth when the water recedes. Quantitative trait locus (QTL) mapping revealed that the SUB1 locus regulates the quiescence strategy (Fukao et al., 2006; Xu et al., 2006). The SUB1 locus on chromosome 9 contains a cluster of three group VII ethylene response factor (ERF) genes (SUB1A, SUB1B, and SUB1C), and the presence of the SUB1A-1 allele restricts underwater shoot growth and confers submergence tolerance. SUB1A-1 suppresses ethylene production, leading to the suppression of GA synthesis and GA responsiveness mediated by the negative regulators of GA signaling, SLENDER RICE1 (SLR1) and SLR-LIKE1 (SLRL1; Fukao et al., 2006; Fukao and Bailey-Serres, 2008). Moreover, SUB1A-1 activates the expression of various genes, including other ERF genes and genes encoding transcription factors, reactive oxygen species scavengers, and enzymes involved in brassinosteroid synthesis and several metabolic pathways facilitating survival during the quiescent period (Fukao et al., 2006, 2011; Jung et al., 2010; Schmitz et al., 2013; Tamang and Fukao, 2015).
Conversely, the escape strategy involves stem elongation to keep the leaves above the water surface. This typically involves metabolic activation and the mobilization of energy reserves to drive elongation growth (Fig. 1A). Utilizing this strategy, deepwater rice rapidly elongates its internodes (∼20–25 cm per day) and reaches a length of several meters in deep water (Catling, 1992; Kende et al., 1998). The deepwater rice cv C9285 from Bangladesh belongs to the japonica varietal group (Wang et al., 2013) and shows strong internode elongation in response to submergence (Hattori et al., 2009). Our previous QTL mapping showed that the submergence-induced elongation in cv C9285 is caused by three major QTLs located on chromosomes 1, 3, and 12 and two minor QTLs on chromosomes 2 and 4 (Hattori et al., 2009; Nagai et al., 2012). The major QTL on chromosome 12 contains the two ERF family genes named SNORKEL1 and SNORKEL2 (SK1/2), which are positive regulators of internode elongation. During submergence, the gaseous hormone ethylene accumulates, triggering SK1/2 gene expression in cv C9285. Although the downstream factors directly regulated by SK1/2 are still unknown, it is clearly established that internode elongation in deepwater rice requires active GA biosynthesis (Ayano et al., 2014; Nagai et al., 2014).
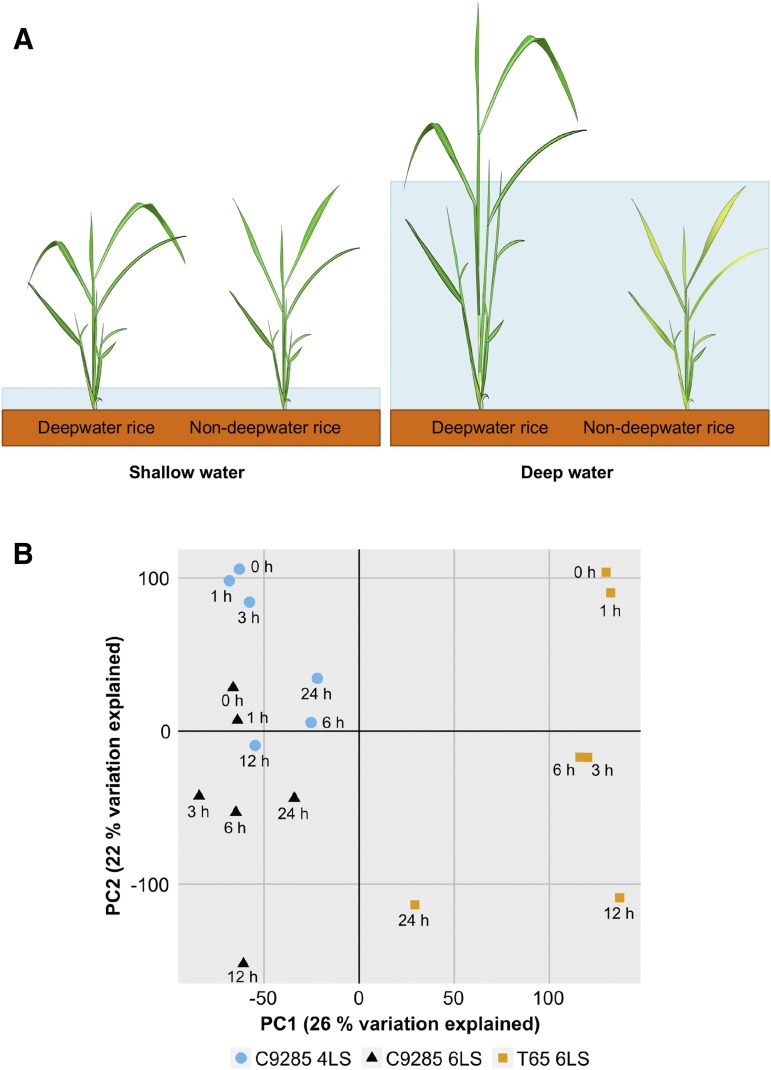
Figure 1.
Submergence responses of deepwater rice and nondeepwater rice. A, Deepwater rice can survive and escape from submerged conditions by rapid elongation of stems (internodes) and leaves, while nondeepwater rice cannot elongate internodes during submergence. B, PCA of 18 RNA-Seq samples from cv C9285 and T65 submerged at different leaf stages and time points. A plot of all transcriptome samples along the first two principal components (PC1 and PC2) is shown. The percentage of variation explained is indicated at each axis. Colors indicate samples from different genotypes and leaf stages. Each data point represents averaged data from three independent time series, and one individual plant was sampled for each data point.
Both the underwater quiescence strategy and the escape strategy involve ethylene signaling and regulation by strategy-specific group VII ERFs: SUB1A limits stem elongation underwater, whereas SK1/2 promotes the submergence-induced elongation. The molecular mechanisms of the SUB1A-mediated quiescence strategy are well studied (Bailey-Serres and Voesenek, 2010; Voesenek and Bailey-Serres, 2015), and SUB1-related gene expression profiles during submergence have been analyzed (Jung et al., 2010; Fukao et al., 2011). However, there is little information about the responses of deepwater rice under the escape strategy. In this study, we explored the transcriptional responses to submergence of the deepwater rice cv C9285. We aimed to compare the transcriptional responses associated with phytohormone signaling, regulation of gene expression, and energy metabolism following the submergence of two contrasting rice varieties (deepwater and nondeepwater rice) at different leaf developmental stages to better understand the mechanism involved in the escape strategy. Our findings expose temporal and genotype-specific responses to submergence and implicate a role for jasmonic acid (JA) catabolism in the pronounced submergence-induced internode elongation of deepwater rice.
RESULTS
Transcriptome Profiles of Deepwater Rice and Nondeepwater Rice Plants Exposed to Submergence
To investigate the gene expression dynamics of deepwater rice in response to submergence, we compared the transcriptome profiles between the deepwater rice cv C9285 and the nondeepwater rice cv Taichung 65 (T65; Supplemental Fig. S1). The internodes of cv C9285 plants can elongate in response to submergence once plants reach the six-leaf stage (6LS; Ayano et al., 2014). Hence, for the transcriptome analysis, cv C9285 and T65 plants at the 6LS were used for expression profiling. We also included submerged cv C9285 plants at the four-leaf stage (4LS), which cannot respond to submergence by elongation. The plants were completely submerged for 0, 1, 3, 6, 12, and 24 h, and we sampled the shoot base region of the stem, as this region in cv C9285 plants elongates rapidly in response to submergence. Per sample, on average, 8.04 × 106 single-end reads were generated, of which an average of 6.83 × 106 reads could be mapped to the transcripts annotated in the japonica cv Nipponbare reference genome (Kawahara et al., 2013). The average rates of mapped reads for cv C9285 and T65 samples in the cv Nipponbare reference genome were 84.8% and 86.1%, respectively (Supplemental Table S1). After quality control and filtering, we detected 27,812 expressed genes in at least one sample (Supplemental Data Set S1).
For global comparison of the transcriptomes derived from all 18 tissue samples, we performed a principal component analysis (PCA). The first two principal components (PC1 and PC2) accounted for 48% of the total variance (Fig. 1B). PC1 accounted for 26% of all variance in the data and clearly separated the cv T65 samples (T65 6LS) from the cv C9285 samples at two different stages (C9285 4LS and C9285 6LS). Among the T65 6LS samples, only the sample from the 24-h submergence treatment showed lower PC1 values closer to the cv C9285 samples. PC2 (accounting for 22% of all variance) separated the samples based on the duration of submergence treatment. That is, each genotype showed higher and lower PC2 values, reflecting the samples submerged for shorter and longer periods, respectively. The results from the PCA analysis suggest that there are different gene expression patterns between the two rice genotypes dependent on and independent of submergence treatment. Furthermore, the expression profiles become variable depending on the duration of submergence treatment but were less affected by the leaf developmental stage in cv C9285 plants. In addition, a more targeted attempt at isolating genes specifically induced by submergence at the 4LS, but not at the 6LS, in cv C9285 plants found only 36 such genes (Supplemental Results S1).
Submergence induced expression changes for a large number of genes in both genotypes (Supplemental Table S2; Supplemental Fig. S2, A and B). To group genes with similar expression profiles, we performed k-means clustering of all 27,812 expressed genes. We empirically chose k = 40 for k-means clusters and manually grouped the clusters into four major groups: (1) higher expression in cv C9285 plants (1,019 genes; Supplemental Data Set S2); (2) higher expression in cv T65 plants (1,205 genes; Supplemental Data Set S3); (3) induced expression in both cv C9285 and T65 plants (2,683 genes); and (4) repressed expression in both cv C9285 and T65 plants (4,125 genes), excluding clusters that contained genes with no discernible pattern of expression (Fig. 2A). In addition, to achieve a better separation of submergence-responsive genes, we next performed k-means clustering (k = 20) for each genotype separately (Supplemental Data Sets S4–S7). In cv C9285 samples, five (B6, B9, B11, B12, and B16) and four (B7, B8, B15, and B17) clusters contained submergence-induced and -repressed genes, respectively (Fig. 2B). A total of 2,429 genes in B6, B12, and B16 clusters showed the strongest expression around 12 h of submergence, and a total of 775 early-responsive genes were classified into B9 and B11 clusters. In cv T65 samples, five clusters (C1, C3, C5, C13, and C17) contained a total of 4,563 submergence-induced genes, and most of the genes showed relatively slow response to submergence, except for the 374 early-responsive genes in the C5 and C17 clusters (Fig. 2C). In cluster C2, the expression of 514 genes in cv T65 plants was repressed by submergence. We evaluated the biological functions of genes enriched in each group of clusters based on the MAPMAN ontology (Thimm et al., 2004; http://mapman.gabipd.org/web/guest/mapman). The hierarchical enrichment analysis of MAPMAN bins of genes with higher expression in cv C9285 or T65 plants (P < 0.05) and the data of submergence-changed genes in cv C9285 or T65 plants (P < 0.05) are shown in Supplemental Tables S3 and S4, respectively (described in Supplemental Results S1). Selected enriched MAPMAN bins were then chosen for further analysis.
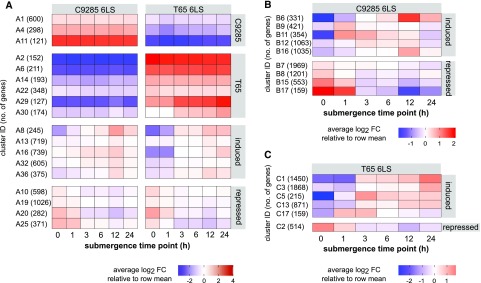
Figure 2.
Clustering analysis of the transcriptomes of cv C9285 and T65 plants after submergence. Heat map representations of expression data from selected clusters in both cv C9285 and T65 at the 6LS (C9286 6LS and T65 6LS; A) and from clusters only in cv C9285 6LS (B) or cv T65 6LS (C) plants are shown. The clusters in the figure derived from Supplemental Data Set S1 were arranged manually. Rows represent clusters of genes generated by k-means clustering. Columns represent samples from different time points after submergence. Colors represent the average expression profile for each cluster, with red showing high expression and blue showing low expression. Before averaging within each cluster, the expression of each gene was normalized to its average expression across all samples and transformed to log2 values. Numbers on the left side of each heat map show the assigned cluster identifiers, and the number of genes in each cluster is in parentheses. The total number of clusters was k = 40 in A and k = 20 in B and C. FC, Fold change.
Different Expression Patterns of Genes Related to Plant Hormone Biosynthesis during Submergence in Both Genotypes
Kende et al. (1998) proposed a model describing the relationship among the three phytohormones ethylene, abscisic acid (ABA), and GA in regulating rapid internode elongation of deepwater rice. During submergence, the reduced diffusion of ethylene causes its accumulation, which then triggers the ethylene signaling cascade, leading to the reduction of ABA content and the increase of GA content (Kende et al., 1998; Fukao and Bailey-Serres, 2008). To investigate the genes related to hormone biosynthesis that are regulated during submergence, we compared the expression levels of hormone biosynthetic genes between cv C9285 and T65 plants.
Ethylene is produced from its precursor, S-adenosyl-Met (SAM), derived from the Yang cycle through two steps (Yang and Hoffman, 1984; Wang et al., 2002). The first step is catalyzed by ACC synthase (ACS) converting SAM into 1-aminocyclopropane-1-carboxylic acid (ACC), while the second step involves ACC oxidase (ACO), which converts the ACC to ethylene (Rudus et al., 2013; Fig. 3A). The reaction catalyzed by ACS is the major rate-limiting step of ethylene biosynthesis, and ACO also probably becomes the rate-limiting enzyme at low-oxygen and high-ethylene conditions (Yang and Hoffman, 1984; Kende, 1993). Rice has six putative ACS genes (OsACS1–OsACS6; Rzewuski and Sauter, 2008), but only OsACS6 expression was detected in our data set, showing a similar expression pattern during submergence in both genotypes. In contrast, all seven rice ACO transcripts (OsACO1–OsACO7; Rzewuski and Sauter, 2008) were detected. In both genotypes, OsACO1, OsACO2, OsACO3, and OsACO7 showed moderate to high expression levels, while OsACO4, OsACO5, and OsACO6 were expressed only at very low levels. OsACO7 was induced markedly by submergence in both genotypes, showing slightly higher expression in cv C9285 than in cv T65. The expression profiles of genes involved in ethylene signaling were mostly similar in both cv C9285 and T65 plants (Supplemental Fig. S3A), which coincides with the similarly increased levels of ethylene during submergence in both genotypes (Hattori et al., 2009).
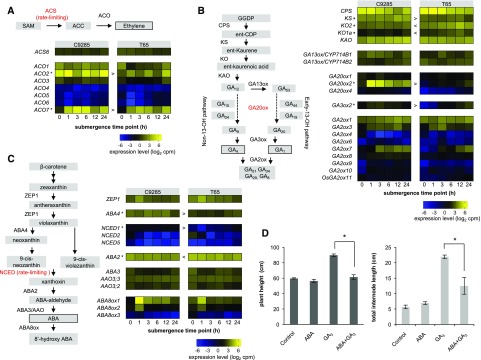
Figure 3.
Ethylene, GA, and ABA metabolism during submergence. A to C, Schematic overviews of ethylene (A), GA (B), and ABA (C) metabolism alongside heat maps showing the expression of relevant genes. Each row represents one gene, columns represent samples from different time points after submergence, and colors represent gene expression levels as log2-transformed counts per million (cpm). Lower levels of expression are represented in blue and higher expression in yellow. AAO3, ABA-aldehyde oxidase; ABA8ox, ABA-8′ oxidase; CPS, copalyl-phosphate synthase; CYP714, cytochrome P714; GAXox, GAX oxidase; GGDP, geranyl-geranyl diphosphate; KAO, ent-kaurenoic acid oxidase; KO, ent-kaurene oxidase; KS, ent-kaurene synthase; NCED, 9-cis-epoxycarotenoid dioxygenase; ZEP1, zeaxanthin epoxidase1. Asterisks indicates genes with significant differences (false discovery rate < 0.05) at more than four time points between cv C9285 and T65 samples, and < and > indicate the direction of the difference relative to cv C9285. D, Effects of ABA and GA3 on plant height and total internode length. The cv C9285 4LS plants were grown in shallow water containing 10 µm GA3 and/or 10 µm ABA for 18 d. Bars represent averages of at least seven biological replicates, and error bars show se. Asterisks show significant differences (P < 0.05) between the indicated treatments as calculated by Student’s t tests.
GA is the key hormone for submergence-induced stem elongation, and particularly, the concentrations of bioactive GA1 and GA4 increase in submerged cv C9285 plants (Hattori et al., 2009). GA biosynthesis starts from the conversion of trans-geranyl-geranyl diphosphate followed by four enzymatic steps to generate GA12 (Fig. 3B; Richards et al., 2001; Yamaguchi, 2008). In the later biosynthetic steps, the conversion of GA12 to GA53 is catalyzed by GA13 oxidase (GA13ox; Magome et al., 2013), and both GA12 and GA53 are catalyzed through two parallel pathways (the early-13-hydroxylation and non-13-hydroxylation pathways) by GA20ox. GA20ox catalyzes the conversion of GA12 and GA53 into the bioactive GA precursors GA9 and GA20, respectively. Finally, GA3ox converts GA9 and GA20 into the bioactive GA forms GA4 and GA1. The expression patterns of genes encoding the enzymes before the catalytic step of GA13ox were not prominently different between cv C9285 and T65 plants. GA20ox is the key enzyme of bioactive GA synthesis, and rice has four putative GA20ox genes (OsGA20ox1–OsGA20ox4; Sakamoto et al., 2004). The transcripts of three OsGA20ox genes were detected in our data set, and only one of the OsGA20ox genes, OsGA20ox2, was strongly expressed 1 h after submergence (greater than 15-fold increase) in cv C9285 (Fig. 3B). On the other hand, in cv T65, the expression level of OsGA20ox2 was considerably lower (1.5 cpm in cv T65 versus 72 cpm in cv C9285 1 h after submergence). OsGA20ox1 and OsGA20ox4 were expressed at low levels (less than 2.5 cpm) in both genotypes during submergence. Among the two GA3ox genes in rice, only OsGA3ox2 expression was detected in both genotypes, showing a slight increase after submergence. GA is inactivated by GA2ox and CYP714D1/EUI1 enzymes (Sakamoto et al., 2004; Zhu et al., 2006). Among the 11 putative OsGA2ox genes in rice, the expression of eight OsGA2ox genes was detected, and the expression patterns were mostly similar between the two genotypes. In our data, the expression of CYP714D1/EUI1, which inactivates GA12, GA9, and GA4, could not be detected. We also investigated the expression profiles of GA signaling-related genes during submergence and found that SLRL1 was expressed after submergence in both genotypes, although SLR1 was not induced by submergence (Supplemental Fig. S3B). However, the results remain elusive, because GA homeostasis interacts tightly with GA metabolism and GA signaling pathways, which are controlled by feedback and feed-forward regulations and various environmental factors (Sun, 2011; Hedden and Thomas, 2012).
ABA is a negative regulator of stem elongation during submergence, corresponding to the decrease of ABA content in both submerged genotypes (Hoffmann-Benning and Kende, 1992; Hattori et al., 2009). In higher plants, the ABA precursor β-carotene is synthesized from isopentenyl pyrophosphate through the plastidial methylerythritol phosphate pathway (Endo et al., 2014). β-Carotene is catalyzed by several enzymes and eventually converted into ABA (Fig. 3C). NCED is a rate-limiting enzyme in ABA biosynthesis, and in rice, NCED is putatively encoded by five genes (OsNCED1–OsNCED5; Zhu et al., 2009). OsNCED1, OsNCED2, and OsNCED5 genes were detected in our data set, and the expression of OsNCED2 was reduced in both genotypes 1 h after submergence. Additionally, in the rice genome, there are three genes that encode the ABA-inactivating enzyme, ABA 8′-hydroxylase (ABA8ox). In our data, the OsABA8ox1 and OsABA8ox2 genes in both genotypes showed transiently increased expression after 1 h of submergence. The genes involved in ABA signaling were expressed similarly in both cv C9285 and T65 plants (Supplemental Fig. S3C).
Hoffmann-Benning and Kende (1992) showed that ABA negatively regulates GA-mediated stem elongation by experiments with excised stem sections of the deepwater rice cv Habiganj Aman II. To test the effect of ABA on the GA activity in whole plants, cv C9285 plants were grown from seeds for 18 d in shallow water containing GA3, ABA, or a combination of both GA3 and ABA. Treatment with GA3 increased the total plant height and internode length compared with untreated plants (Fig. 3D). While treatment with ABA alone had no observable effect, treatment with combined ABA and GA3 reduced the GA-induced elongation of cv C9285 plants.
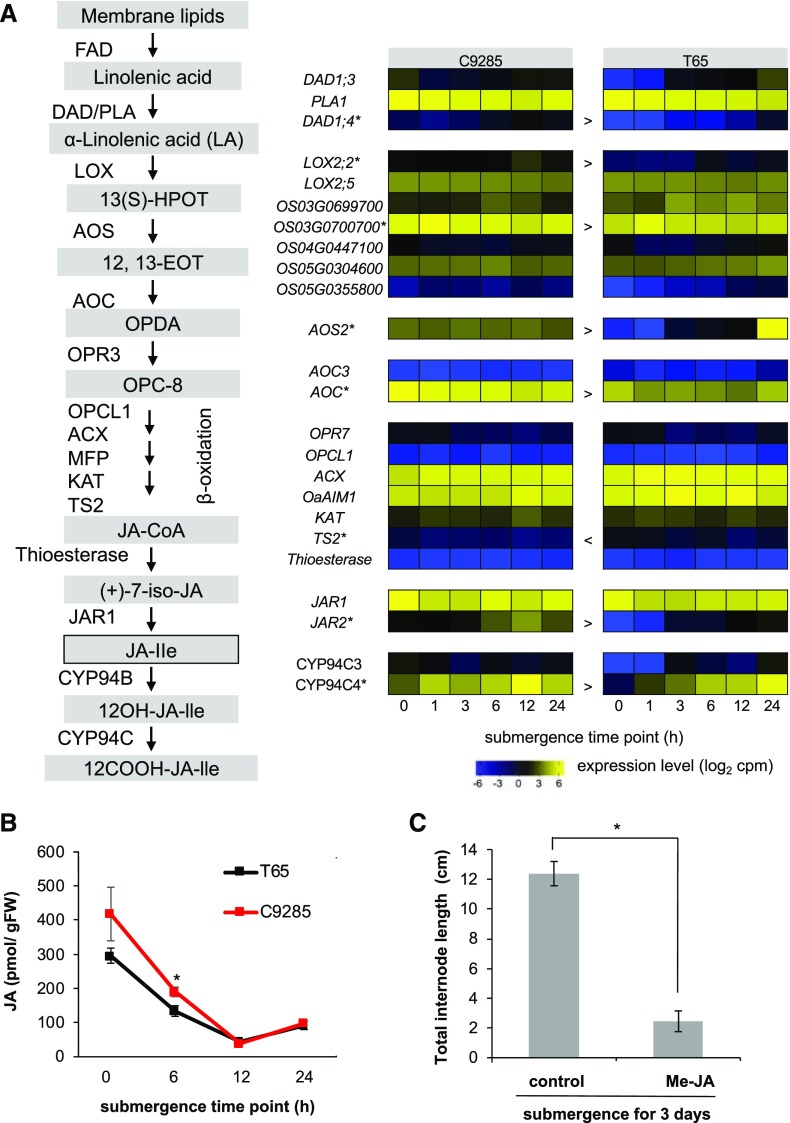
JAs control plant growth, development, and responses to abiotic and biotic stresses. JAs are derived from α-linolenic acid, which is released from galactolipids of chloroplast membranes by fatty acid desaturase and phospholipase A1, including defective in anther dehiscence1 (Wasternack and Hause, 2013; Fig. 4A). α-Linolenic acid is converted to OPC-8 through the action of several enzymes, including lipoxygenase, allene oxide synthase, allene oxide cyclase, and 12-oxo-phytodienoic acid reductase3. After the three steps of β-oxidation, OPC-8 is converted into jasmonoyl-CoA and then cleaved to (+)-7-iso-jasmonoyl by thioesterase. Finally, (+)-7-iso-jasmonoyl-Ile, which is a ligand binding a coreceptor of JA receptors, is catalyzed by jasmonate resistant1 in the cytosol. Through the activity of the members of the CYP94 family, (+)-7-iso-jasmonoyl-Ile is inactivated through hydroxylation. The putative JA-inactivating CYP94C4 gene showed higher expression during submergence compared with before submergence in both genotypes. In addition, endogenous JA levels decreased after 24 h of submergence in both genotypes (Fig. 4B). To examine the effect of JA on the submergence response, we submerged cv C9285 plants in water containing 50 µm methyl jasmonate and measured the total internode length after 3 d of treatment (Fig. 4C). Surprisingly, treatment with JA remarkably inhibited the submergence-induced internode elongation.
Figure 4.
JA metabolism during submergence. A, Schematic overview of JA metabolism alongside heat maps showing the expression of relevant genes. Each row represents one gene, columns represent samples from different time points after submergence, and colors represent gene expression levels as log2-transformed cpm. Lower levels of expression are represented in blue and higher expression in yellow. FDA, Fatty acid desaturase; DAD1, defective in anther dehiscence1; PLA, phospholipase A1; LOX, lipoxygenase; 13(S)-HPOT, 13(S)-hydroperoxyoctadecatrienoic acid; AOS, allene oxide synthase; 12,13-EOT, 12,13(S)-epoxy-octadecatrienoic acid; AOC, allene oxide cyclase; OPDA, cis-(+)-12-oxo-phytodienoic acid; OPR, 12-oxo-phytodienoic acid reductase; OPC-8, 3-oxo-2-(2′-pentenyl)-cyclopentane-1-octanoic acid; OPCL1, acyl-activating enzyme; ACX, acyl-CoA oxidase; MFP, multifunctional protein; KAT, 3-keto-acyl-CoA thiolase; TS2, tasselseed2; JA-CoA, jasmonoyl-CoA; (+)-7-iso-JA, (+)-7-iso-jasmonoyl; JAR1, jasmonate resistant1; JA-lle, (+)-7-iso-jasmonoly-l-Ile; CYP94, cytochrome P94. Asterisks indicate genes with significant differences (false discovery rate < 0.05) at more than four time points between cv C9285 and T65 samples, and < and > indicate the direction of the difference relative to cv C9285. B, Endogenous JA levels in the shoot base regions of cv C9285 and T65 plants during submergence for 24 h. Symbols represent averages of at least five biological replicates, and error bars represent se. The asterisk shows a significant difference (P < 0.05) between cv C9285 and T65 samples as calculated by Student’s t tests. FW, Fresh weight. C, JA inhibits internode elongation during submergence. The cv C9285 6LS plants were submerged under water containing 50 µm methyl jasmonate (Me-JA) for 3 d. Bars represent averages of at least 12 biological replicates, and error bars show se. The asterisk shows a significant difference (P < 0.05) between the indicated treatments as calculated by Student’s t tests.
Deepwater Treatment Has Genotype-Specific Effects on Trehalose Metabolism and Fermentation-Related Pathways
We further investigated the expression of genes in other relevant pathways to understand the metabolic adaptation of deepwater rice under submergence. Comparative analysis of metabolic pathways using MAPMAN software showed that genes related to the metabolism of trehalose (Supplemental Fig. S4A) and fermentation (Supplemental Fig. S4B) were induced in response to 1 h of submergence in cv C9285 plants. The expression of genes in the NO3-generating pathway (Supplemental Fig. S4C) also was up-regulated after submergence, but their absolute expression was at very low levels.
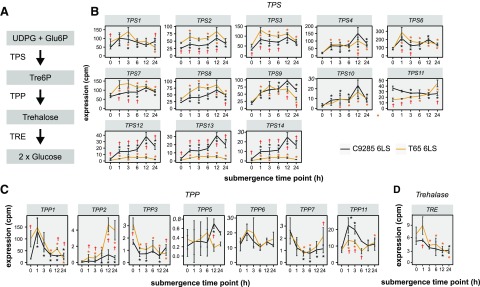
Trehalose is a disaccharide that functions in carbon metabolism and tolerance to biotic and abiotic stresses (Lunn et al., 2014). In plants, trehalose is synthesized in a two-step process from UDP-Glc and Glc-6-P by the subsequent actions of trehalose-6-phosphate synthases (TPS) and trehalose-6-phosphate phosphatases (TPP), and the degradation of trehalose is catalyzed by trehalase (TRE; Fig. 5A; Ponnu et al., 2011). In our data set, we detected the expression of trehalose metabolism-related genes, including 13 TPS (out of 14), seven TPP (out of 13), and one TRE (Ge et al., 2008) genes. Some TPS genes clearly responded to submergence in cv C9285 plants. Notably, three TPS genes (OsTPS12, OsTPS13, and OsTPS14) were strongly induced in response to submergence in cv C9285 plants but not in cv T65 plants (Fig. 5B). OsTPP1 and OsTPP11 showed elevated expression in response to submergence in cv C9285 plants, while OsTPP2 expression was induced at a relatively low level in cv T65 after submergence (Fig. 5C). In the case of TRE, the expression pattern was similar in both genotypes (Fig. 5D).
Figure 5.
Expression of trehalose metabolism-related genes during submergence. A, Schematic overview of the trehalose metabolic pathway. Glu-6P, Glc-6-P; UDPG, UDP-Glc. B to D, Expression of genes involved in trehalose metabolism during submergence in cv C9285 6LS (black) and cv T65 6LS (orange) plants at the indicated time points after submergence. Data points show expression levels as average cpm in three biological replicates, and error bars show sd. Black and orange asterisks indicate significant differences (P < 0.05) from 0 h at each time point for cv C9285 and T65 samples, respectively. Daggers indicate significant differences (false discovery rate < 0.05) between cv C9285 and T65 samples. P values were calculated using likelihood ratio tests and corrected for multiple testing by the Bonferroni-Holm method.
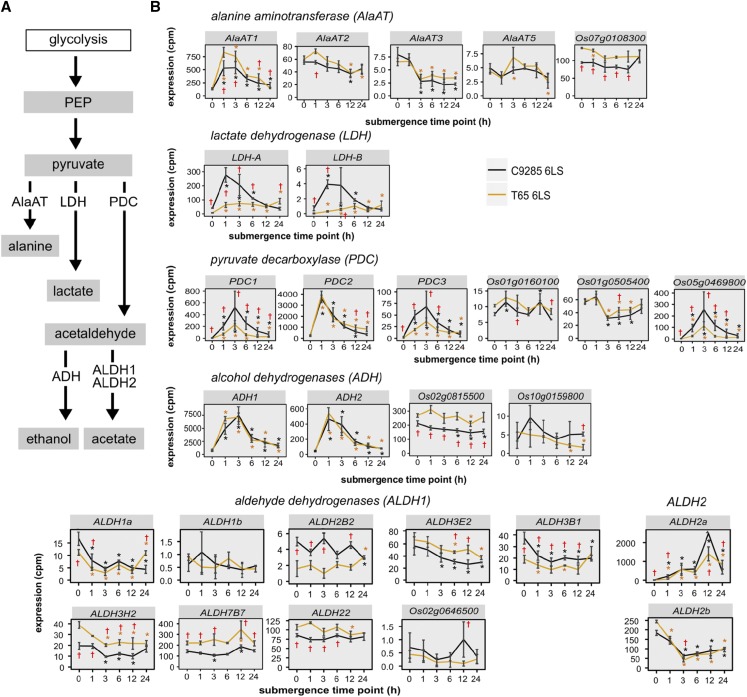
Under anaerobic conditions, oxygen is limited for aerobic respiration and plant cells implement anaerobic respiration (glycolysis and fermentation) for energy production instead (Perata et al., 1998; Fukao and Bailey-Serres, 2004; Bailey-Serres and Voesenek, 2008; Miro and Ismail, 2013). The glycolysis pathway generates ATP, and the following fermentation steps produce NAD+ for the maintenance of glycolysis and some metabolites. Fermentation converts the glycolysis-derived pyruvate into other metabolites such as Ala, lactate, ethanol, and acetate (Fig. 6A). Ala is synthesized from pyruvate and Glu by Ala aminotransferase (AlaAT), while lactate is formed by the reduction of pyruvate via lactate dehydrogenase (LDH). Pyruvate decarboxylase (PDC) converts pyruvate to acetaldehyde. To detoxify acetaldehyde, alcohol dehydrogenase (ADH) or acetaldehyde dehydrogenase (ALDH) subsequently catalyzes the conversion of acetaldehyde to ethanol or acetate, respectively. Interestingly, we found that the two LDH genes (LDH-A and LDH-B) were strongly and specifically induced 1 h after submergence in cv C9285 but not in cv T65 plants (Fig. 6B). Among other fermentation-related genes, OsAlaAT1, OsPDC1, OsPDC2, and OsPDC3, Os05g0469800, OsADH1 and OsADH2, and OsALDH2a appeared to increase after submergence in both genotypes, whereas genes in the ALDH1 family seemed to show variable expression patterns during submergence.
Figure 6.
Expression of fermentation-related genes during submergence. A, Schematic overview of three fermentation pathways. B, Expression of genes related to fermentation during submergence in cv C9285 6LS (black) and cv T65 6LS (orange) plants at the indicated time points after submergence. Data points show expression levels as average cpm in three biological replicates, and error bars show sd. Graphs are arranged to reflect the different possibilities of anaerobic energy production. PEP, Phosphoenolpyruvate. Black and orange asterisks indicate significant differences (P < 0.05) from 0 h at each time point for cv C9285 and T65 samples, respectively. Daggers indicate significant differences (P < 0.05) between cv C9285 and T65 samples. P values were calculated using likelihood ratio tests and corrected for multiple testing by the Bonferroni-Holm method.
Submergence Induces Differential Expression Profiles of Genes Related to Cell Wall Formation between cv C9285 and T65 Plants
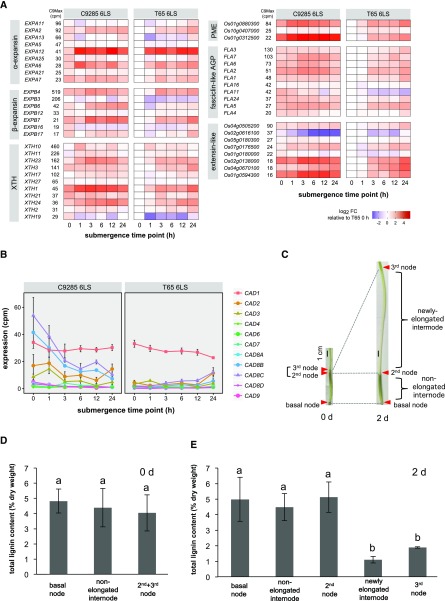
Under the nonsubmerged condition, genes related to cell wall formation and secondary metabolic pathways showed higher expression in cv C9285 plants compared with cv T65 plants (Supplemental Fig. S5, A–D). The plant cell wall provides mechanical strength, regulates growth, serves as a barrier to diffusion, and protects the plant against pathogens. The cell wall-related transcripts with higher expression in cv C9285 plants encoded cell wall modification proteins α- and β-expansins (EXPAs and EXPBs), xyloglucan endotransglycosylases/hydrolases (XTHs), pectin esterases (PMEs), and structural cell wall proteins (fasciclin-like arabinogalactans [FLAs] and extensins; Fig. 7A). In cv T65 plants, almost all genes responded to submergence, and the expression level finally reached that of cv C9285 within 24 h of submergence. In cv C9285 plants, several genes were strongly induced by submergence, especially the genes EXPB6, EXPB7, OsXTH23, and OsXTH1, and two PMEs (Os01g0880300 and Os01g0312500), three FLAs (FLA7, FLA6, and FLA2), and two extensins (Os02g0138000 and Os01g0594300) remarkably responded to submergence in cv C9285 plants.
Figure 7.
Effects of submergence on the expression of cell wall formation-related genes. A, Expression pattern of cell wall-related genes during 24 h of submergence. The gene expression levels in cv C9285 6LS and cv T65 6LS samples are indicated by log2 fold change (FC) at each time point (0, 1, 3, 12, and 24 h) relative to that of cv T65 samples before submergence. Genes with a maximal cpm value of more than 15 are shown. AGP, Arabinogalactan protein. B, Changes in expression of 11 CAD/SAD genes during submergence for 24 h. Data points show expression levels as average cpm in three biological replicates, and error bars show sd. C, Sampled regions of internodes for lignin quantification. The cv C9285 plants at the seven-leaf stage (7LS) that have already formed internodes were submerged for 2 d. Images of stems before (0 d) and after (2 d) submergence treatment of cv C9285 plants at the 7LS are shown. Red arrowheads indicate stem nodes. D and E, Lignin content in the nodes and internodes before (D) and after (E) 2 d of submergence of cv C9285 plants at 7LS. Bars represent averages of at least six biological replicates, and error bars represent se. Different letters indicate significant (P < 0.05) differences according to the Tukey-Kramer test.
Under nonsubmerged conditions, genes involved in flavonoid, anthocyanin, and phenylpropanoid metabolism, as well as in the biosynthesis of lignin and simple phenols (e.g. laccases involved in lignin polymerization), were expressed higher in cv C9285 compared with cv T65 (Supplemental Fig. S5C). Lignin is a complex polymer of monolignols and strengthens the cell wall as a component of secondary cell walls. Thus, cell wall lignification probably functions in the suppression of rapid growth in deepwater rice, resulting in a slow rate of elongation of its stems in nonsubmerged conditions (Sauter and Kende 1992). We investigated the transcriptional responses of lignin biosynthesis-related genes to submergence and focused on genes encoding coniferyl/sinapyl alcohol dehydrogenase (CAD/SAD), which catalyzes the final step of monolignol biosynthesis (Supplemental Fig. S6A). In cv C9285 plants, among the 11 CAD/SAD genes (out of a total of 12 genes in the genome), OsCAD1, OsCAD2, OsCAD8B, and OsCAD8C were highly expressed in nonsubmerged conditions and, except for OsCAD1, showed decreased expression in response to submergence (Fig. 7B). OsCAD1 showed a similar expression level in both genotypes, while the other CAD genes in cv T65 maintained lower expression levels during submergence. To determine whether the decrease in the expression of CAD genes affects lignin biosynthesis in elongating internodes of cv C9285 plants, we measured the lignin content of the cv C9285 stem, including (1) nodes from different positions, (2) nonelongated internodes (an internode that already stopped elongating), and (3) newly elongated internodes (a newly elongating internode during submergence; Fig. 7C). Under the nonsubmerged condition, the basal node, nonelongated internode, and second and third nodes showed similar lignin contents (Fig. 7D). The total lignin content in the basal and second nodes did not change after 2 d of submergence (Fig. 7, D and E). On the other hand, the third node on the internode newly elongated after 2 d of submergence exhibited a lower lignin content (Fig. 7E). The total lignin content in the nonelongating internode was not affected by 2 d of submergence, while the content of the newly elongated internodes was lower than that of the nonelongated internode. These results suggest that the suppression of lignin biosynthesis through the reduction of CAD transcripts contributes to the elongation of internodes in cv C9285 plants. In agreement with this, the lignin content in the basal nodes of cv T65 plants did not change during submergence (Supplemental Fig. S6B).
Expression of ERF Transcription Factor Genes in Response to Submergence
The MAPMAN overrepresentation analysis revealed that transcription factor genes from the AP2/EREBP family were enriched significantly among genes induced by submergence in both genotypes (Supplemental Table S4, A and B). To investigate the responses of all transcription factor genes to submergence, the genes in our data set were categorized according to the Plant Transcription Factor Database version 3.0 (http://planttfdb.cbi.pku.edu.cn/; Jin et al., 2014; Supplemental Data Set S8; Supplemental Fig. S7; Supplemental Results S1). Interestingly, those clusters contained a total of 10 ERF genes (Supplemental Fig. S7). The ERFs belong to the AP2/EREBP superfamily, which is a plant-specific transcription factor family with three subfamilies: (1) AP2 family proteins containing two repeated AP2 DNA-binding domains; (2) ERF family proteins containing a single AP2 domain; and (3) RAV family proteins containing a B3 DNA-binding domain and a single AP2 domain (Riechmann and Meyerowitz, 1998; Sakuma et al., 2002; Nakano et al., 2006). Nakano et al. (2006) divided the rice ERF family (a total of 139 genes) into 15 subfamilies including 28 subgroups. In our data set, we categorized the ERF genes into these subgroups. Among all transcription factors in our data set, 115 were found to be ERF genes, including 41, 10, and 64 genes with increased, decreased, and unchanged expression during submergence, respectively (Supplemental Table S5). The ERF genes from subgroups IIa, IIIc, VIIa, VIIb, IXa, Xa, Xb, Xc, and XI predominantly showed increased expression in response to submergence in cv C9285 plants, while those of subgroups IIIe and Va showed decreased expression under submergence treatment. Supplemental Figure S8 shows expression profiles of the submergence-induced ERF genes. The SUB1 locus encodes three ERF transcription factor genes (SUB1A, SUB1B, and SUB1C; Xu et al., 2006); however, the genome of cv C9285 contains only SUB1B and SUB1C but does not have SUB1A (Hattori et al., 2009). In cv C9285 plants, SUB1B (OsERF#063) and SUB1C (OsERF#073) increased 3.2- and 2.7-fold 1 h after submergence, respectively (Supplemental Fig. S8A). The expression of SK1 and SK2 genes, which are positive regulators of internode elongation in cv C9285 plants and absent in cv T65 plants, also was increased remarkably 5.3- and 3.2-fold by submergence, respectively (Supplemental Fig. S8B).
GCC-Box-Like Promoter Elements Are Enriched in the Promoters of Genes Expressed Specifically in cv C9285 Plants
Using k-means clustering, we isolated 189 genes that are expressed specifically in cv C9285 plants (Supplemental Data Set S9). Their putative promoter regions (1 kb upstream from the transcriptional start site) were constructed by aligning reads from genomic DNA of cv C9285 plants to the japonica cv Nipponbare reference genome. The constructed promoter sequences were then analyzed for enriched DNA sequence motifs using the MEME suite. We found two GCC-box like elements (GCGGCGGCGG and CGCCGCCGCC) enriched in sequence among the analyzed promoter sequences (Supplemental Fig. S9). The consensus DNA-binding motif (GCCGCC) is known to be recognized by the ERF proteins (Hao et al., 1998; Fujimoto et al., 2000).
Identification and Analysis of Differentially Expressed Genes Unique to cv C9285
Although about 85% of cv C9285 RNA sequencing (RNA-Seq) reads could be mapped to the cv Nipponbare reference genome (Supplemental Table S1), there is a possibility that cv C9285 has additional genes missing in the reference genome, such as SK1 and SK2. Hence, we performed a targeted de novo assembly of transcripts to detect cv C9285-specific transcripts (Supplemental Fig. S10; Supplemental Methods S1). We used 47,978,704 paired-end reads from genomic DNA of cv C9285 (approximately 20× coverage) to assemble C9285-specific genomic regions and quantified transcripts detected in those regions using RNA-Seq reads. We then restricted our analysis to those transcripts that are expressed at high levels, are strongly induced by submergence, or have an interesting annotation. In total, 86 transcribed loci on 33 genomic contigs were found to be either putatively unique to cv C9285 or sufficiently different from the cv Nipponbare reference so that mapping with standard parameters failed to produce alignments (Supplemental Data Sets S10 and S12–S14).
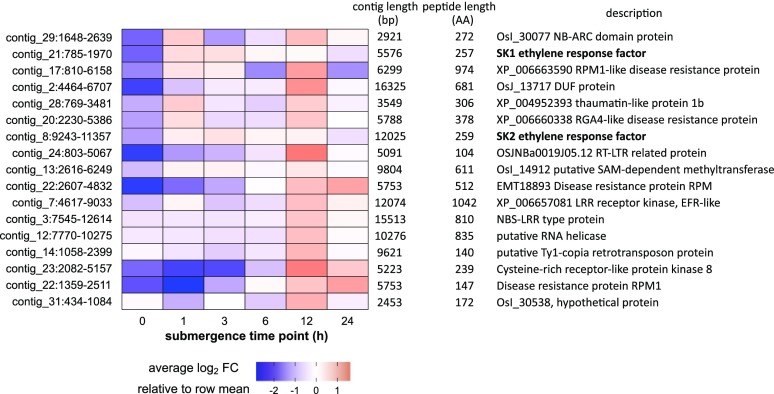
Among those 86 loci, two encoded the full-length SK1 and SK2 transcripts. The genomic contigs that contained the SK1 (contig_21; 5,776 bp) and SK2 (contig_8; 12,025 bp) loci aligned almost perfectly with parts of the bacterial artificial chromosome clone that was used for map-based cloning of SK1 and SK2 (Hattori et al., 2009). When we quantified the expression of SK1 and SK2 genes from those contigs, we found the typical early increase of transcriptional activity 1 h after submergence with a subsequent decline (Supplemental Fig. S8B). This shows that our targeted assembly strategy was able to correctly identify unique cv C9285 transcripts, including their genomic sequence and their expression patterns. We found 11 novel transcripts that had at least 2-fold higher expression levels after 1 h of submergence (Fig. 8). Except for SK1 and SK2, most of those transcripts did not encode well-characterized proteins. Instead, a sequence comparison using BLASTP reported at least six pathogen- or disease-related proteins. The remaining three proteins were annotated as domain of unknown function, retrotransposon related, and SAM-dependent methyl transferase.
Figure 8.
Submergence-induced transcripts from cv C9285-unique loci. Expression from selected loci that are presumably absent in the IRGSP-1.0 reference genome is shown. Each row represents one locus, columns represent samples from different time points after submergence, and colors represent gene expression levels as log2-transformed cpm normalized to the average expression in all samples of different time points. Genes that have at least a 2-fold increase in expression after submergence in cv C9285 6LS plants are shown. Complete expression data from cv C9285-unique transcripts can be found in Supplemental Data Set S10, and sequence data are provided in Supplemental Data Sets S12 to S14. AA, Amino acids; FC, fold change.
DISCUSSION
The overall goal of this study was to understand the transcriptional responses associated with the escape strategy of the deepwater rice cv C9285 during submergence. Cataloging the whole transcriptome through an RNA-Seq approach revealed differences in gene expression between deepwater rice (cv C9285) and nondeepwater rice (cv T65) plants as well as unique and common responses to submergence in the two genotypes. The results of our comprehensive transcriptome analysis contribute to the finding of key genes regulating the metabolic pathways during submergence in rice.
Transcripts Associated with Ethylene, GA, and ABA Metabolism Are Differentially Expressed in Response to Submergence
Previous studies have demonstrated that the hormones ethylene, GA, and ABA modulate the internode elongation response of submerged deepwater rice (Kende et al., 1998). Among the ethylene biosynthesis genes, it is reported that OsACS1, OsACS5, and OsACO1 were increased by submergence in deepwater rice (Mekhedov and Kende, 1996; Zarembinski and Theologis, 1997; Van Der Straeten et al., 2001). On the other hand, in our data, OsACO7 transcript and protein accumulated during submergence in both genotypes (Fig. 3A; Supplemental Fig. S11A). Ethylene accumulation in submerged deepwater rice is attributed to the decrease in ethylene diffusion and the enhancement of ethylene biosynthesis (Métraux and Kende, 1983). Our previous study showed that ethylene accumulation is a common occurrence in both cv C9285 and T65 plants (Hattori et al., 2009). Our transcript analysis supports that OsACO7 is involved in ethylene accumulation in both genotypes during submergence. OsACO7 has not been reported to participate in ethylene biosynthesis during submergence, although most previous studies used nondeepwater rice varieties.
Under the submergence condition, the accumulated ethylene enhances GA accumulation for the promotion of internode elongation in deepwater rice (Métraux and Kende, 1983, 1984; Raskin and Kende, 1984; Kende et al., 1998). Interestingly, unlike ethylene biosynthesis, the activation of GA biosynthesis occurs only in cv C9285 plants (Hattori et al., 2009). In our data, OsGA20ox2 (also known as SD1) expression levels were strongly and specifically induced by submergence in cv C9285 but not in cv T65 (Fig. 3B). GA20ox2 is a key locus for GA production; thus, its increased expression is most likely responsible for the higher GA1 and GA4 accumulation in submerged cv C9285 plants (Supplemental Fig. S12F; Hattori et al., 2009; Ayano et al., 2014). In the case of the submergence-tolerant rice with SUB1A, SUB1A suppresses GA signaling through the accumulation of GA signaling repressors, SLR1 and SLRL1, leading to the restricted elongation of stems and leaves under submergence (Fukao and Bailey-Serres, 2008; Colebrook et al., 2014). Thus, the absence of SUB1A in the deepwater rice cv C9285 may explain the accumulation of bioactive GA levels promoting the elongation of stems and leaves during submergence. It has been reported that the GA-induced gene Growth Regulating Factor1 (OsGRF1) and a GA stimulated transcript (GAST)-like gene, OsGSR1, enhance stem elongation in rice (van der Knaap et al., 2000; Ben-Nissan et al., 2004; Choi et al., 2004; Wang et al., 2009). Interestingly, two GAST-like genes (Os05g0376800 and OsGSR1) were expressed higher in cv C9285 plants during submergence, although OsGRF1 was not induced during submergence in our data set (Supplemental Fig. S11, B and C), suggesting that the GAST-like genes may be involved in GA-induced stem elongation during submergence in cv C9285 plants (Furukawa et al., 2006).
The content of ABA, which has the antagonistic effect to GA on internode elongation, decreased under submergence and ethylene treatments (Hoffmann-Benning and Kende, 1992; Azuma et al.,1995; Benschop et al., 2006; Yang and Choi, 2006; Saika et al., 2007; Weiss and Ori, 2007; Fukao and Bailey-Serres, 2008; Hattori et al., 2009; Chen et al., 2010). The decrease in ABA content is a common response to submergence in both cv T65 and C9285 plants (Hattori et al., 2009). Our data revealed that transcript levels of OsNCED2, encoding one of the rate-limiting enzymes in ABA biosynthesis, decreased, and OsABA8ox1 and OsABA8ox2 encoding ABA inactivation enzymes were up-regulated in both genotypes during submergence (Fig. 3C), indicating that the decrease in ABA content may be explained by changes in the expression levels of genes related to ABA metabolism in both genotypes.
JA Is a Novel Regulatory Factor Associated with Stem Elongation in Deepwater Rice
Kende et al. (1998) reported that ethylene, ABA, and GA are key regulators of internode elongation in deepwater rice. In this study, we demonstrated that submergence regulated the endogenous JA content in node samples of rice and that JA inhibited the submergence-induced internode elongation in cv C9285 plants (Fig. 4). CYP94C2b encodes a cytochrome P450 enzyme and was involved in the inactivation of JA-Ile (Kurotani et al., 2015). Our data showed that CYP94C4, which is a major expressed CYP94C gene, showed increased expression levels during submergence in both genotypes. Thus, the increase in CYP94C4 might regulate the decrease in endogenous JA content. The decrease in JA is a common response to submergence in both genotypes, like the changes in ethylene and ABA contents during submergence. The nondeepwater rice cv T65 at 6LS has no ability to elongate its internodes even in the case of GA treatment (Nagai et al., 2014), suggesting that the changes in JA contents in cv T65 plants do not affect stem elongation.
Studies of cross talk between JA and GA signaling pathways have shown that GA antagonistically functions in JA signaling pathways involved in plant growth and development processes in Arabidopsis (Hou et al., 2013; Wasternack and Hause, 2013). In addition, Yang et al. (2012) showed that JA-mediated growth inhibition in rice was caused by changes in the levels of DELLA repressors and interference with GA signaling. Our data suggest that GA-mediated internode elongation in deepwater rice requires the suppression of JA function through the reduction of JA content in submergence.
The cv C9285 and T65 Plants Differentially Express Trehalose and Fermentation Metabolism-Related Genes
Trehalose, which accumulates in response to various stresses, acts as a carbon source and an osmoprotectant by stabilizing proteins and membranes against the stresses (Krasensky and Jonak, 2012). On the other hand, trehalose-6-phosphate (Tre6P), a precursor of trehalose, functions as a signaling metabolite that coordinates carbon assimilation, starch synthesis, nitrogen metabolism, growth, and development (Schluepmann et al., 2003; Martins et al., 2013; Lunn et al., 2014; Yadav et al., 2014; Figueroa et al., 2016; Figueroa and Lunn, 2016). The change in the Tre6P-Suc ratio is an important homeostatic mechanism of plants under stress conditions; thus, Tre6P also acts as a signal for Suc status through negative feedback regulation of Suc levels (Yadav et al., 2014; Figueroa and Lunn, 2016). In rice, overexpression of OsTPS1 or OsTPP1 enhanced the tolerance to abiotic stresses, leading to the expression of stress-related genes (Ge et al., 2008; Li et al., 2011a). OsTPP7 (OsTPP11 in this article) also enhances the anaerobic germination tolerance in young rice by regulating the trehalose content and Tre6P/Suc homeostasis in sugar signaling, but not the Tre6P content (Kretzschmar et al., 2015). In our data, although several OsTPS genes, including OsTPS1, OsTPP1, and OsTPP11, were expressed by submergence in both genotypes, the expression of OsTPS12, OsTPS13, and OsTPS14 was induced dramatically by submergence in cv C9285 but maintained at a lower level in cv T65 (Fig. 5B). This result implies that trehalose metabolism during submergence is differentially regulated between cv C9285 and T65 and that trehalose and Tre6P may accumulate more in submerged cv C9285 plants, contributing to the ability of deepwater rice to withstand and adapt to submergence stress. In submerged SUB1A-1-containing M202(SUB1) rice seedlings and anoxic coleoptiles, the trehalose biosynthesis pathway is activated (Jung et al., 2010), suggesting that such a pathway also is utilized by the deepwater rice as a part of metabolic adaptation to submergence.
Under low-oxygen conditions, glycolysis is channeled predominantly to fermentation pathways (instead of aerobic respiration), which is necessary for cell survival to produce energy and recycle carbon for other pathways (Gibbs and Greenway, 2003; Voesenek et al., 2006). Our data exhibited the increased expression of fermentation-related genes, PDCs, ADH1 and ADH2, and ALDH2a, during submergence in both genotypes (Fig. 6B). A functional SUB1 locus regulates the gene expression and enzyme activities of PDC and ADH during submergence (Fukao et al., 2006). Thus, a SUB1A-1-containing rice, cv Flood Resistant 13A, accumulates less aldehydes because high expression of ADH enhances the detoxification of acetaldehyde into the neutral and diffusible ethanol (Singh et al., 2001; Xu et al., 2006). Submergence also induces the expression of ALDH2a, but not ALDH1, in coleoptiles of rice (Nakazono et al., 2000). Our results imply that the fermentation pathway is active in both genotypes and that the enzymes ADH and ALDH2a are likely involved in removing toxic alcohol and acetaldehyde in response to low oxygen during submergence (Fig. 6A).
The increased expression of OsAlaAT1 (Fig. 6B) further supports that the active fermentation pathway in cv C9285 and T65 also generates Ala. These results are consistent with the increased activities of AlaAT, PDC, and ADH in the flooded coleoptile of rice cv Nipponbare (Kato-Noguchi, 2006); however, in their study, the activity of LDH and lactate content were similar both in the presence and absence of oxygen. In our data set, the transcripts of two LDH genes in cv C9285 plants were strongly up-regulated after submergence compared with T65 plants (Fig. 6B). The formation of lactate leads to a decrease in cytosolic pH, and the initial acidification of the cytoplasm helps to achieve optimum activity of PDC, which, in turn, promotes the switch from lactate to ethanol fermentation (Roberts et al., 1984; Kennedy et al., 1992; Magneschi and Perata, 2009). Hence, it is possible that the abundance of LDH transcripts results in higher accumulation of lactate in the cytoplasm and a stronger lactate-dependent reduction of cytosolic pH favoring ethanol fermentation more in cv C9285 than in cv T65 plants. Recently, Lee et al. (2015) reported that lactate binds to NDRG3, an oxygen-regulated protein, and activates Raf-ERK signaling to promote cell growth and angiogenesis during hypoxia in human cells, suggesting that lactate in plant cells might function as a signaling molecule in a similar signaling pathway in response to hypoxia.
Cell Wall Synthesis and Modification Genes Are Differentially Expressed in cv C9285 Plants in Response to Submergence
Genes with a differential response to submergence between cv C9285 and T65 plants were enriched in the cell wall-related category (Supplemental Fig. S2C). Additionally, genes encoding cell wall-related proteins, such as EXPs, XTHs, PMEs, and structural cell wall proteins (FLAs and extensins), were categorized as submergence-induced genes in both genotypes (Supplemental Table S4, A and B) and showed higher expression in cv C9285 plants under the nonsubmerged condition (Fig. 7A). Elongating cells require the relaxation of cell walls, involving cell wall-loosening factors such as EXPs. EXP activity increases under acidic conditions, which are controlled by plasma membrane H+-ATPase activity via auxin-induced SMALL AUXIN UP-RNA proteins (Perrot-Rechenmann, 2010; Spartz et al., 2014). In deepwater rice, a positive correlation was observed between the expression of EXPs and acid-induced cell wall extensibility, and they may function in internode elongation (Cho and Kende, 1997a, 1997b; Lee and Kende, 2001). The EXPs together with XTHs control fiber orientation and viscoelastic properties of the matrix facilitating cell wall expansions (Cosgrove, 2000; Van Sandt et al., 2007). XTH activity possibly enhances EXP activity during the shade-avoidance elongation response in Stellaria longipes (Sasidharan et al., 2008). In rice, XTHs could function in the cell wall formation of the vascular bundles in elongating stems (Hara et al., 2014).
PMEs catalyze the demethylesterification of pectin, a GalUA-rich cell wall polymer, resulting in the formation of a gel-like structure of polymers and thereby increasing wall porosity and extension of cells (Jolie et al., 2010). Jeong et al. (2015) showed that high PME activity was detected in germinating shoots that actively undergo cell elongation. The cell wall also is regulated by structural cell wall proteins, such as the arabinogalactan proteins, which are involved in elongation and growth of Arabidopsis and cucumber (Cucumis sativus; van Hengel and Roberts, 2002; Park et al., 2003). We found that that the expression of several genes related to cell wall loosening was induced. These genes might contribute to cell wall reconstruction and increased cell wall extensibility, leading to further enhancement of rapid internode elongation of cv C9285 plants during submergence.
The accumulation of lignin is part of the cell wall modifications that lead to an increase in mechanical strength but also inhibit cell elongation. In cv C9285 plants, the expression of several CAD genes, encoding key enzymes that catalyze the final step in monolignol synthesis, decreased during submergence (Fig. 7B), leading to a reduced lignin content in newly elongated internodes during submergence (Fig. 7E).
Peroxidase genes showed differential expression patterns between both genotypes under submergence (Supplemental Fig. S2C; Supplemental Results S1) and were classified as submergence-induced genes (Supplemental Table S4, A and B). Interestingly, the expression levels of the peroxidase genes were higher in cv C9285 plants than in cv T65 plants during the early submergence condition (Supplemental Fig. S6C). Peroxidases are involved in lignification, stress defense, and the regulation of reactive oxygen species production (Shigeto and Tsutsumi, 2016). In addition, hydroxyl radicals produced by peroxidases also may function in cell wall loosening and cell elongation, as in maize (Zea mays) roots (Liszkay et al., 2004). Thus, submergence-responsive peroxidases probably function in cell wall loosening and the lignification of elongating internodes in cv C9285 plants.
ERF Genes Are Differentially Regulated in Submerged cv C9285 and T65 Plants
ERFs are major downstream components of the ethylene signaling pathway. In our analysis, among the 115 detected ERF family transcripts, 39 genes in the Ib, IIa, IIIc, VII, VIIIa, IXa, X, and XI subgroups were up-regulated and 10 genes in the IIIe and Va subgroups were down-regulated by submergence in cv C9285 plants (Supplemental Table S5), indicating that ethylene signaling is an important regulatory pathway for the response of deepwater rice to submergence.
Interestingly, among all 15 members of the group VII ERF family, 12 genes, including SUB1B (OsERF#063) and SUB1C (OsERF#073), were induced by submergence in cv C9285 plants (Supplemental Table S5; Supplemental Fig. S8A). Members of the group VII ERF family act as oxygen sensors under hypoxic conditions by the N-end rule pathway for protein degradation in Arabidopsis (Gibbs et al., 2011; Licausi et al., 2011; Gasch et al., 2016) and function in multiple stress tolerances, such as drought, cold, pathogen attack, salinity, osmotic stress, and submergence, in various plants (Licausi et al., 2010; Mizoi et al., 2013; Gibbs et al., 2015; Papdi et al., 2015). We found that the other three genes (OsERF#70, OsERF#71, and OsERF#72) encoding a CMVII-4 motif, which is predicted to be a phosphorylation site of mitogen-activated protein kinases (MPKs; Nakano et al., 2006), showed nonresponsive expression to submergence in cv C9285 plants (Supplemental Fig. S13). MPKs phosphorylate a large number of transcription factors, including ERFs (Popescu et al., 2009), and the MPK-induced phosphorylations in rice ERFs enhance their transcriptional activities and environmental stress tolerances (Cheong et al., 2003; Schmidt et al., 2013). Recently, it was reported that OsMPK3 phosphorylated SUB1A-1 to control shoot elongation in a SUB1A-1-dependent manner, although the phosphorylation site was not located in the CMVII-4 motif (Singh and Sinha, 2016). In cv C9285 plants, all members of the group VII ERFs that encode proteins without the phosphorylation sites were induced by submergence, and the submergence-induced expression levels of some group VII ERFs (OsERF#59, OsERF#61, OsERF#63, OsERF#64, OsERF#66, and OsERF#67) were higher during submergence compared with cv T65 plants (Supplemental Fig. S13). These results suggest that phosphorylation-independent group VII ERFs may be important for the submergence-induced stress response in deepwater rice, although there is still the possibility that submergence induces phosphorylation of the ERFs by MPKs.
Other submergence-induced ERFs in subgroups IIa and VIIIa encode proteins that contain an ERF-associated amphiphilic repression (EAR) motif (DLNxxP), which acts as a transcriptional repressor domain (Fujimoto et al., 2000; Ohta et al., 2001). As EAR-containing transcription factors regulate various stress responses (Kazan, 2006; Dong and Liu, 2010; Licausi et al., 2013), the submergence-induced ERFs might be involved in negatively regulating the expression of dispensable genes for the submergence response.
In our data, group IIIc, IXa, and X ERFs also showed up-regulated expression in response to submergence (Supplemental Table S5). In Arabidopsis, the group IIIc, IXa, and X ERFs function in tolerance to environmental stresses such as freezing, salt, and dehydration (e.g. cold-binding factor/dehydration-responsive element binding1 protein; Dubouzet et al., 2003), the ethylene-JA signaling pathway (Lorenzo et al., 2003; Champion et al., 2009), and the response to ABA and various stresses (Pandey et al., 2005; Zhu et al., 2010), respectively. In the case of the group XI ERFs, this family exists in rice but not in Arabidopsis, indicating that these genes have rice-specific functions.
A group IIIe ERF, TINY, functions in the reduction of cell expansion and differentiation, leading to a dwarf phenotype in Arabidopsis (Wilson et al., 1996). In cv C9285 plants, two group IIIe ERF family genes were down-regulated by submergence, suggesting that the group IIIe ERFs possibly affect cell expansion during submergence. Arabidopsis ERF11 in group VIIIa represses ethylene biosynthesis genes but promotes cell elongation by promoting GA accumulation and inhibiting DELLA function (Li et al., 2011b; Zhou et al., 2016). In rice, OsEATB (OsERF#102) in group Xb and OsERF3 (OsERF#075) in group VIIIa negatively regulate internode elongation (Qi et al., 2011; Zhang et al., 2013). In cv C9285 plants, OsEATB was not expressed and OsERF3 was not differentially expressed during submergence (Supplemental Fig. S11D), suggesting that the two negative regulators do not participate in GA-mediated internode elongation during submergence in cv C9285 plants.
SUB1A up-regulates the transcriptional expression level of 12 ERF family genes (Jung et al., 2010). Although the SUB1A gene is not present in the cv C9285 genome (Hattori et al., 2009), the six SUB1A-dependent expressed genes (i.e. OsERF#025, OsERF#066, OsERF#067, OsERF#068, OsERF#076, and OsERF#077) also were induced by submergence in cv C9285 plants (Supplemental Fig. S8A). All SUB1A downstream ERF genes belong to the IIIc, VIIa, and VIIIa subfamilies. This indicates that these genes also are regulated by other factors in cv C9285 plants and may indicate that their expression is not associated directly with internode elongation in deepwater rice.
The ERF transcription factors bind to the promoter regions of their target genes, such as pathogen-related genes, through the recognition of a GCC-box cis-regulatory sequence (GCCGCC [core motif]; Ohme-Takagi and Shinshi, 1995; Hao et al., 1998; Solano et al., 1998). Our in silico analysis showed that the two GCC-box-like motifs (G/CCGGCGGCGG and CGCCGCCGCC) were enriched in the promoter regions of genes with higher expression in cv C9285 plants (Supplemental Fig. S9), suggesting that these genes with GCC-box-like motifs are regulated preferentially by ERFs. Some submergence-induced ERF genes showed higher expression in cv C9285 than those of cv T65 plants before and after submergence (Supplemental Fig. S8A). This result suggests that the ERF genes in cv C9285 are associated with the constitutively higher expression of their putative downstream target genes. Recently, Gasch et al. (2016) identified a novel hypoxia-responsive promoter element that is a binding site of ERF VIIs, RELATED TO AP2 2 (RAP2.2) and RAP2.12, the key regulators of the N-end rule pathway in Arabidopsis. However, we did not identify the hypoxia-responsive promoter element motif in our promoter analysis of cv C9285-enriched genes. The oxygen levels in underwater nodes could be much higher than expected for two reasons: (1) diffusion from the water layer into the node, and (2) underwater photosynthesis. Under submergence with light illumination, the oxygen concentration of deepwater rice initially dropped but recovered within 90 min by underwater photosynthesis (Stünzi and Kende, 1989). Thus, our data may not be suitable to study hypoxia, since we collected all samples under light conditions.
A Model for the Transcriptional Response of Deepwater Rice to Submergence
We propose a model for submergence responses during the escape strategy in the deepwater rice cv C9285 (Supplemental Fig. S14). Submergence subjects plant cells to multiple stresses, such as limitation of gas diffusion, lower light intensity, high risk of pathogen infection, and decreased oxygen uptake. To avoid critical damage from submergence, cv C9285 plants have evolved to rapidly elongate their internodes and leaves. We revealed that changes in the contents of the plant hormones ethylene, ABA, GA, and JA during submergence were transcriptionally regulated. Ethylene signaling leads to biotic and abiotic stress tolerances by changing the expression of many genes and plant hormone levels (Müller and Munné-Bosch, 2015), and our data largely exhibited changes in the putative downstream components of ethylene signaling during submergence. We also proposed that JA is a novel regulator for submergence-induced internode elongation in cv C9285 plants. In deepwater rice, GA has an important role in the elongation response, because GA treatment induces the expression of cell wall-related genes such as EXPs and changes the activity of CAD (Sauter and Kende, 1992; Cho and Kende 1997c). We found that cv C9285-specific accumulation of active GAs was probably controlled by the expression of GA20ox2 during submergence (Fig. 3B), and the cell wall-related genes showed higher expression in cv C9285 plants (Fig. 7A). Furthermore, the reduced lignin content in cv C9285 plants also may facilitate the elongation of internodes under the submerged condition. Overall modifications in cell wall metabolism also may contribute to the rapid internode elongation of deepwater rice. Among the transcription factor genes, the ERF family genes were expressed especially at high levels during submergence in cv C9285 plants. Interestingly, genes expressed specifically in cv C9285 plants contain the GCC-box-like motifs that are recognized by ERFs. This indicates that ethylene signaling and the transcriptional response pathway via ERFs could be key factors for the submergence response in cv C9285 plants. In fact, rice ETHYLENE INSENSITIVE3-LIKE1, which is the master regulator of ethylene signaling, binds to the promoter region of the ethylene/submergence-induced ERF transcription factors SK1/2 that positively, but not exclusively, regulate internode elongation in cv C9285 plants (Hattori et al., 2009). Since we could not find any cv C9285-specific novel submergence-induced transcription factors in the cv C9285 genome by our de novo assembly analysis and the presence of SK1/2 alone does not lead to a full stem elongation response, additional transcription factors in coordination with SK1/2 are most likely involved.
In R. palustris, genes associated with photomorphogenesis and shade avoidance seem to regulate the underwater elongation response (van Veen et al., 2013). In our data, the expression pattern of light signaling-regulated genes in cv C9285 plants did not show any obvious genotype or submergence-specific pattern (Supplemental Fig. S15). The genes involved in fermentation and trehalose metabolic pathways responded to submergence in both cv C9285 and T65 plants, and we found that TPSs and LDHs were expressed preferentially during submergence in cv C9285 plants (Figs. 5 and 6), suggesting that these metabolic activities increase more strongly in cv C9285 plants during submergence compared with normal rice such as cv T65 plants, resulting in adaptation to long-term submergence.
Our experiments showed that many pathogenesis related proteins were expressed preferentially in each rice variety (Supplemental Table S3), and several genes encoding disease resistance proteins were identified as cv C9285-unique transcripts (Fig. 8). Waterlogged conditions increase the risk of pathogen infection in plants (Tamang and Fukao, 2015). Accordingly, rice in paddy fields have protection mechanisms that prevent damage rendering them vulnerable to pathogens (Hsu et al., 2013). Therefore, these cv C9285-specific genes might function as a pathogen defense pathway during partial long-term submergence in cv C9285 plants.
Our studies identified genotype-specific responsive genes to submergence in cv C9285 plants and assumed that the putative key genes would be associated with submergence-induced physiological responses in cv C9285 plants. Furthermore, we showed that ethylene is one of the main drivers of submergence responses and that JA is a new negative regulator of submergence-induced internode elongation in cv C9285 plants. The network involving submergence-induced changes and genetic factors complexly regulates underwater elongation and adaptation in deepwater rice.
MATERIALS AND METHODS
Plant Material and Cultivation
A deepwater rice (Oryza sativa) cultivar (cv C9285) and a nondeepwater rice cultivar (cv T65; japonica) were used. Rice seeds were incubated at 60°C for 10 min followed by pregermination at 29°C in water for 3 to 4 d. Afterward, germinated seeds were transferred to plastic pots containing soil mixture (Mikawa Baido; AICHI Mederu) and grown in a greenhouse in Nagoya, Japan, in June with a natural light cycle of approximately 14 h of light/10 h of dark. For deepwater treatment, rice seedlings that reached the indicated leaf stages (4LS and 6LS) were completely submerged for 1, 3, 6, 12, and 24 h (Supplemental Fig. S1A). To avoid differences in gene expression between samples due to circadian rhythms, submergence treatments were initiated at different times of the day and all samples were collected in the afternoon, although varying durations of underwater and abovewater photosynthesis cause changes in the availability of photosynthates. After submergence treatment, 5 mm of the shoot base region containing internodes, nodes, the shoot apex, and basal regions of leaves was sampled, rapidly frozen in liquid nitrogen, and stored at −80°C until RNA extraction (Supplemental Fig. S1B).
RNA Extraction, Sequencing, and Read Mapping
For each RNA extraction, frozen tissues from one individual plant were homogenized and up to 100 mg was used. Total RNA of each sample was isolated using the RNeasy Plant Mini Kit (Qiagen) with the RNase-Free DNase Set (Qiagen). RNA purity was checked using a NanoDrop spectrophotometer (Thermo-Fisher), and RNA was quantified using the QuantiFluor RNA system (Promega) and the EnSpire Multimode Plate Reader (PerkinElmer). For HiSeq library construction, 2 µg of total RNA was used with the TruSeq RNA Sample Preparation Kit version 2 (Illumina) according to the manufacturer’s instructions. Agencourt AMPure XP beads (Beckman Coulter) were used to remove small DNA fragments. The clustering of index-coded samples was performed on the cBot Cluster Generation System using the TruSeq SR Cluster Kit v2-cBot-GA (Illumia) and the TruSeq SBS Kit v5-GA (Illumia). After cluster generation, the library preparations were sequenced on the Illumina Genome Analyzer IIx, and 36-bp single-ended reads were generated. Reads were mapped to the IRGSP-1.0 reference transcripts of japonica cv Nipponbare (Goff et al., 2002) using bowtie (version 0.12.7; Langmead et al., 2009) with the -all -best -strata options (Supplemental Table S1), and read counts were quantified using PostgreSQL and a custom PHP script.
Data Analysis
Statistical data analysis was performed using R. The edgeR package was used to normalize raw count data and generate cpm values (Robinson et al., 2010). Transcripts with less than 10 averaged counts in at least one condition were removed from the data set, and only the strongest expressed isoform per transcript was used for analysis. To determine differentially expressed genes, a negative binominal generalized log-linear model was used (function glmFit) with each genotype and time point after submergence, defined as one group. Then, likelihood ratio tests (function glmLRT) were performed to compare between groups or to test for interactions between genotype and submergence treatment. The Bonferroni-Holm method was used to correct for multiple testing (function topTags). PCA was performed using the prcomp function with scaled and centered cpm data as the input. The k-means clustering was performed using the MBClusterSeq package with a negative binomial model and the EM algorithm (Si et al., 2014). The number of clusters was varied between 10 and 100 in intervals of 10, and empirically k = 40 (for all samples) or k = 20 (for samples from one genotype only) was chosen as the most appropriate number of clusters. Hierarchical clustering of transcription factor genes was performed using the dist and hclust functions. The distance matrix was calculated using Euclidian distance, and clustering was performed using the average algorithm.
Gene set enrichment analysis was based on the MAPMAN ontology, and mappings (Rice Annotation Project Database locus identifier to MAPMAN bin) were obtained from http://www.mapman.gabipd.org (Ramsak et al., 2014). Significant enrichment of bins was determined using Fisher’s exact test, and corrections for multiple testing were done by the Bonferroni-Holm method using the R functions fisher.test and p.adjust, respectively. Separate analyses were performed for the first three hierarchy levels of the MAPMAN ontology.
Metabolic maps were based on a combination of Rice Annotation Project Database Kyoto Encyclopedia of Genes and Genomes (Kanehisa et al., 2016), MAPMAN, and in-house annotations. Data visualization was performed using ggplot2 (Wickham, 2009).
ABA and GA3 Treatments of Whole Plants
Germinated seeds of cv C9285 plants were sown in plastic pots filled with soil and then grown in a phytotron at 25°C with a 14-h photoperiod. When the plants were grown at the 4LS, they were transferred to new plastic containers for hormone treatment. For hormone treatment, plants were watered with 10 µm ABA (Sigma-Aldrich), 10 µm GA3 (Wako), or a combination of both into the containers. The water level in each container was controlled to keep the hormone concentration constant during the treatment. Plant height and internode length were measured after 18 d of treatment. Internode length was calculated using the total length of all internodes.
JA Treatment of Whole Plants during Submergence
Germinated seeds of cv C9285 plants were sown in plastic pots filled with soil and then grown in a controlled-environment chamber at 25°C with a 14-h photoperiod. When the plants reached the 6LS, they were submerged in water with or without 50 µm methyl jasmonate (Wako) for 3 d. Total internode length was calculated as the sum of the length of all internodes.
Measurement of Plant Hormone Contents
The concentrations of endogenous hormones were measured using ultra-performance liquid chromatography-tandem mass spectrometry (UPLC-Xevo TQ-S; Waters) as described by Kojima et al. (2009).
Measurement of Lignin Content
Nodes and internodes were sampled from cv C9285 and T65 plants at the 6LS or 7LS before and after submergence treatment for 2 d. Lignin content was determined by the thioglycolic acid method according to Suzuki et al. (2009).
Promoter Motif Enrichment Analysis
Transcripts were clustered into six groups based on the expression in cv C9285 and T65 plants (6LS) at 0, 1, 3, 6, 12, and 24 h after submergence using k-means clustering as described above. For tentative construction of a cv C9285 genome, all reads obtained from whole-genome sequencing of cv C9285 were mapped against the IRGSP-1.0 pseudomolecules using bwa-mem with the -M option (Li and Durbin, 2009). Mapped reads were realigned using RealignerTargetCreator and indelRealigner from the GATK software suite (DePristo et al., 2011). To identify single-nucleotide polymorphisms and insertions/deletions, UnifiedGenotyper of GATK was used with the -glm BOTH option. The cv C9285 genome sequence was tentatively constructed by modifying the IRGSP-1.0 genome with the identified variants of cv C9285 by custom Perl scripts. DNA sequences 1 kb upstream of the translational start sites from the cv C9285 genome were regarded as promoter regions and used for motif enrichment analysis. CENSOR was used to mask low-complexity regions in the promoter sequences (Kohany et al., 2006). The MEME suite was used to find enriched DNA sequence motifs using the following parameters: -dna -nmotifs 10 -maxw 10 (Bailey and Elkan, 1994).
Accession Numbers
All reads used in this study can be found in the DNA Database of Japan under bioproject PRJDB5294 (cv C9285 RNA-Seq reads) and bioproject PRJDB5300 (cv C9285 genomic reads).
Supplemental Data
The following supplemental materials are available.
Supplemental Figure S1. Schematic overview of the experimental setup and transcriptome sequencing.
Supplemental Figure S2. Significance analysis using a generalized log-linear model.
Supplemental Figure S3. Expression of genes related to ethylene, GA, and ABA signaling during submergence.
Supplemental Figure S4. Overview of MAPMAN visualization of differentially expressed genes in cv C9285 plants after 1 h of submergence.
Supplemental Figure S5. Overview of MAPMAN visualization of differentially expressed genes in cv C9285 and T65 plants under the nonsubmerged condition.
Supplemental Figure S6. Expression of lignin biosynthesis genes during submergence.
Supplemental Figure S7. Clustering of transcription factor genes according to expression in cv C9285 relative to cv T65 plants.
Supplemental Figure S8. Submergence-induced AP2/EREBP family genes in cv C9285 plants.
Supplemental Figure S9. Enriched sequence motifs in the promoters of genes that are differentially expressed in cv C9285 plants.
Supplemental Figure S10. Analysis pipeline to detect novel cv C9285 transcripts.
Supplemental Figure S11. Submergence responses related to plant hormones.
Supplemental Figure S12. Isolation of genes specifically induced by submergence in cv C9285 4LS plants.
Supplemental Figure S13. Expression of group VII ERF genes during submergence.
Supplemental Figure S14. Molecular model for the submergence response in the deepwater rice cv C9285.
Supplemental Figure S15. Expression of light signaling genes during submergence.
Supplemental Table S1. Summary of read mapping statistics.
Supplemental Table S2. Number of genes with significantly different expression in pairwise comparisons.
Supplemental Table S3. Enriched MAPMAN bins among genes preferentially expressed in cv C9285 or T65 plants.
Supplemental Table S4. Enriched MAPMAN bins among genes induced or repressed by submergence in either genotype.
Supplemental Table S5. Response to submergence of rice ERF subfamily genes in cv C9285 plants.
Acknowledgments
We thank Dr. Ko Hirano for assisting with lignin quantification and Dr. M. Fujita for providing cv C9285 genomic sequencing data. Seeds of cv C9285 used in this study were distributed from the National Institute of Genetics supported by the National Bioresource Project, AMED, Japan.
Footnotes
This work was supported by JST Core Research for Evolutional Science and Technology, a MEXT Grant-in-Aid for Scientific Research on Innovative Areas (22119007 and 17H06473) and JICA-JST SATREPS, by JSPS Grand-in Aid for Young Scientists (B) Grant Number 17K15136 to A.M., and by a JST ERATO Grant (JPMJER1004) to T.H.
References
- Ayano M, Kani T, Kojima M, Sakakibara H, Kitaoka T, Kuroha T, Angeles-Shim RB, Kitano H, Nagai K, Ashikari M (2014) Gibberellin biosynthesis and signal transduction is essential for internode elongation in deepwater rice. Plant Cell Environ 37: 2313–2324 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Azuma T, Hirano T, Deki Y, Uchida N, Yasua T, Yamaguchi T (1995) Involvement of the decrease in levels of abscisic acid in the internodal elongation of submerged floating rice. J Plant Physiol 146: 323–328 [Google Scholar]
- Bailey TL, Elkan C (1994) Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc Int Conf Intell Syst Mol Biol 2: 28–36 [PubMed] [Google Scholar]
- Bailey-Serres J, Voesenek LA (2008) Flooding stress: acclimations and genetic diversity. Annu Rev Plant Biol 59: 313–339 [DOI] [PubMed] [Google Scholar]
- Bailey-Serres J, Voesenek LA (2010) Life in the balance: a signaling network controlling survival of flooding. Curr Opin Plant Biol 13: 489–494 [DOI] [PubMed] [Google Scholar]
- Bailey-Serres J, Lee SC, Brinton E. (2012) Waterproofing Crops: Effective Flooding Survival Strategies. Plant Physiol 2: 1698–1709 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ben-Nissan G, Lee JY, Borohov A, Weiss D (2004) GIP, a Petunia hybrida GA-induced cysteine-rich protein: a possible role in shoot elongation and transition to flowering. Plant J 37: 229–238 [DOI] [PubMed] [Google Scholar]
- Benschop JJ, Bou J, Peeters AJM, Wagemaker N, Gühl K, Ward D, Hedden P, Moritz T, Voesenek LA (2006) Long-term submergence-induced elongation in Rumex palustris requires abscisic acid-dependent biosynthesis of gibberellin1. Plant Physiol 141: 1644–1652 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Catling D. (1992) Rice in Deep Water. Macmillan Press, London [Google Scholar]
- Champion A, Hebrard E, Parra B, Bournaud C, Marmey P, Tranchant C, Nicole M (2009) Molecular diversity and gene expression of cotton ERF transcription factors reveal that group IXa members are responsive to jasmonate, ethylene and Xanthomonas. Mol Plant Pathol 10: 471–485 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen X, Pierik R, Peeters AJM, Poorter H, Visser EJ, Huber H, de Kroon H, Voesenek LA (2010) Endogenous abscisic acid as a key switch for natural variation in flooding-induced shoot elongation. Plant Physiol 154: 969–977 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheong YH, Moon BC, Kim JK, Kim CY, Kim MC, Kim IH, Park CY, Kim JC, Park BO, Koo SC, et al. (2003) BWMK1, a rice mitogen-activated protein kinase, locates in the nucleus and mediates pathogenesis-related gene expression by activation of a transcription factor. Plant Physiol 132: 1961–1972 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cho HT, Kende H (1997a) Expansins in deepwater rice internodes. Plant Physiol 113: 1137–1143 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cho HT, Kende H (1997b) Expansins and internodal growth of deepwater rice. Plant Physiol 113: 1145–1151 [DOI] [PMC free article] [PubMed] [Google Scholar]; Cho HT, Kende H (1997c) Expression of expansin genes is correlated with growth in deepwater rice. Plant Cell 9: 1661–1671 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi D, Kim JH, Kende H (2004) Whole genome analysis of the OsGRF gene family encoding plant-specific putative transcription activators in rice (Oryza sativa L.). Plant Cell Physiol 45: 897–904 [DOI] [PubMed] [Google Scholar]
- Colebrook EH, Thomas SG, Phillips AL, Hedden P (2014) The role of gibberellin signalling in plant responses to abiotic stress. J Exp Biol 217: 67–75 [DOI] [PubMed] [Google Scholar]
- Cosgrove DJ. (2000) Loosening of plant cell walls by expansins. Nature 407: 321–326 [DOI] [PubMed] [Google Scholar]
- DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, Philippakis AA, del Angel G, Rivas MA, Hanna M, et al. (2011) A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet 43: 491–498 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dong CJ, Liu JY (2010) The Arabidopsis EAR-motif-containing protein RAP2.1 functions as an active transcriptional repressor to keep stress responses under tight control. BMC Plant Biol 10: 47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dubouzet JG, Sakuma Y, Ito Y, Kasuga M, Dubouzet EG, Miura S, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold-responsive gene expression. Plant J 33: 751–763 [DOI] [PubMed] [Google Scholar]
- Endo A, Okamoto M, Koshiba T (2014) ABA biosynthetic and catabolic pathways. In Zhang DP, ed, Abscisic Acid: Metabolism, Transport and Signaling. Springer, Dordrecht, The Netherlands, pp 21–45 [Google Scholar]
- Figueroa CM, Feil R, Ishihara H, Watanabe M, Kölling K, Krause U, Höhne M, Encke B, Plaxton WC, Zeeman SC, et al. (2016) Trehalose 6-phosphate coordinates organic and amino acid metabolism with carbon availability. Plant J 85: 410–423 [DOI] [PubMed] [Google Scholar]
- Figueroa CM, Lunn JE (2016) A tale of two sugars: trehalose 6-phosphate and sucrose. Plant Physiol 172: 7–27 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujimoto SY, Ohta M, Usui A, Shinshi H, Ohme-Takagi M (2000) Arabidopsis ethylene-responsive element binding factors act as transcriptional activators or repressors of GCC box-mediated gene expression. Plant Cell 12: 393–404 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukao T, Bailey-Serres J (2004) Plant responses to hypoxia: is survival a balancing act? Trends Plant Sci 9: 449–456 [DOI] [PubMed] [Google Scholar]
- Fukao T, Bailey-Serres J (2008) Submergence tolerance conferred by Sub1A is mediated by SLR1 and SLRL1 restriction of gibberellin responses in rice. Proc Natl Acad Sci USA 105: 16814–16819 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukao T, Xu K, Ronald PC, Bailey-Serres J (2006) A variable cluster of ethylene response factor-like genes regulates metabolic and developmental acclimation responses to submergence in rice. Plant Cell 18: 2021–2034 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukao T, Yeung E, Bailey-Serres J (2011) The submergence tolerance regulator SUB1A mediates crosstalk between submergence and drought tolerance in rice. Plant Cell 23: 412–427 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Furukawa T, Sakaguchi N, Shimada H (2006) Two OsGASR genes, rice GAST homologue genes that are abundant in proliferating tissues, show different expression patterns in developing panicles. Genes Genet Syst 81: 171–180 [DOI] [PubMed] [Google Scholar]
- Gasch P, Fundinger M, Müller JT, Lee T, Bailey-Serres J, Mustroph A (2016) Redundant ERF-VII transcription factors bind to an evolutionarily conserved cis-motif to regulate hypoxia-responsive gene expression in Arabidopsis. Plant Cell 28: 160–180 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ge LF, Chao DY, Shi M, Zhu MZ, Gao JP, Lin HX (2008) Overexpression of the trehalose-6-phosphate phosphatase gene OsTPP1 confers stress tolerance in rice and results in the activation of stress responsive genes. Planta 228: 191–201 [DOI] [PubMed] [Google Scholar]
- Gibbs DJ, Conde JV, Berckhan S, Prasad G, Mendiondo GM, Holdsworth MJ (2015) Group VII ethylene response factors coordinate oxygen and nitric oxide signal transduction and stress responses in plants. Plant Physiol 169: 23–31 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gibbs DJ, Lee SC, Isa NM, Gramuglia S, Fukao T, Bassel GW, Correia CS, Corbineau F, Theodoulou FL, Bailey-Serres J, et al. (2011) Homeostatic response to hypoxia is regulated by the N-end rule pathway in plants. Nature 479: 415–418 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gibbs J, Greenway H (2003) Mechanisms of anoxia tolerance in plants. I. Growth, survival and anaerobic catabolism. Funct Plant Biol 30: 353. [DOI] [PubMed] [Google Scholar]
- Goff SA, Ricke D, Lan TH, Presting G, Wang R, Dunn M, Glazebrook J, Sessions A, Oeller P, Varma H, et al. (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. japonica). Science 296: 92–100 [DOI] [PubMed] [Google Scholar]
- Hao D, Ohme-Takagi M, Sarai A (1998) Unique mode of GCC box recognition by the DNA-binding domain of ethylene-responsive element-binding factor (ERF domain) in plant. J Biol Chem 273: 26857–26861 [DOI] [PubMed] [Google Scholar]
- Hara Y, Yokoyama R, Osakabe K, Toki S, Nishitani K (2014) Function of xyloglucan endotransglucosylase/hydrolases in rice. Ann Bot 114: 1309–1318 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hattori Y, Nagai K, Furukawa S, Song XJ, Kawano R, Sakakibara H, Wu J, Matsumoto T, Yoshimura A, Kitano H, et al. (2009) The ethylene response factors SNORKEL1 and SNORKEL2 allow rice to adapt to deep water. Nature 460: 1026–1030 [DOI] [PubMed] [Google Scholar]
- Hedden P, Thomas SG (2012) Gibberellin biosynthesis and its regulation. Biochem J 444: 11–25 [DOI] [PubMed] [Google Scholar]
- Hoffmann-Benning S, Kende H (1992) On the role of abscisic acid and gibberellin in the regulation of growth in rice. Plant Physiol 99: 1156–1161 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsu FC, Chou MY, Chou SJ, Li YR, Peng HP, Shih MC (2013) Submergence confers immunity mediated by the WRKY22 transcription factor in Arabidopsis. Plant Cell 25: 2699–2713 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jackson MB. (2008) Ethylene-promoted elongation: an adaptation to submergence stress. Ann Bot 101: 229–248 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jeong HY, Nguyen HP, Lee C (2015) Genome-wide identification and expression analysis of rice pectin methylesterases: implication of functional roles of pectin modification in rice physiology. J Plant Physiol 183: 23–29 [DOI] [PubMed] [Google Scholar]
- Jin J, Zhang H, Kong L, Gao G, Luo J (2014) PlantTFDB 3.0: a portal for the functional and evolutionary study of plant transcription factors. Nucleic Acids Res 42: D1182–D1187 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jolie RP, Duvetter T, Van Loey AM, Hendrickx ME (2010) Pectin methylesterase and its proteinaceous inhibitor: a review. Carbohydr Res 345: 2583–2595 [DOI] [PubMed] [Google Scholar]
- Jung KH, Seo YS, Walia H, Cao P, Fukao T, Canlas PE, Amonpant F, Bailey-Serres J, Ronald PC (2010) The submergence tolerance regulator Sub1A mediates stress-responsive expression of AP2/ERF transcription factors. Plant Physiol 152: 1674–1692 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M (2016) KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 44: D457–D462 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kato-Noguchi H. (2006) Pyruvate metabolism in rice coleoptiles under anaerobiosis. Plant Growth Regul 50: 41–46 [Google Scholar]
- Kawahara Y, de la Bastide M, Hamilton JP, Kanamori H, McCombie WR, Ouyang S, Schwartz DC, Tanaka T, Wu J, Zhou S, et al. (2013) Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice (N Y) 6: 4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kazan K. (2006) Negative regulation of defence and stress genes by EAR-motif-containing repressors. Trends Plant Sci 11: 109–112 [DOI] [PubMed] [Google Scholar]
- Kende H. (1993) Ethylene biosynthesis. Annu Rev Plant Physiol Plant Mol Biol 44: 283–307 [Google Scholar]
- Kende H, van der Knaap E, Cho HT (1998) Deepwater rice: a model plant to study stem elongation. Plant Physiol 118: 1105–1110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kennedy RA, Rumpho ME, Fox TC (1992) Anaerobic metabolism in plants. Plant Physiol 100: 1–6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohany O, Gentles AJ, Hankus L, Jurka J (2006) Annotation, submission and screening of repetitive elements in Repbase: RepbaseSubmitter and Censor. BMC Bioinformatics 7: 474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kojima M, Kamada-Nobusada T, Komatsu H, Takei K, Kuroha T, Mizutani M, Ashikari M, Ueguchi-Tanaka M, Matsuoka M, Suzuki K, et al. (2009) Highly sensitive and high-throughput analysis of plant hormones using MS-probe modification and liquid chromatography-tandem mass spectrometry: an application for hormone profiling in Oryza sativa. Plant Cell Physiol 50: 1201–1214 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krasensky J, Jonak C (2012) Drought, salt, and temperature stress-induced metabolic rearrangements and regulatory networks. J Exp Bot 63: 1593–1608 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kretzschmar T, Pelayo MAF, Trijatmiko KR, Gabunada LFM, Alam R, Jimenez R, Mendioro MS, Slamet-Loedin IH, Sreenivasulu N, Bailey-Serres J, et al. (2015) A trehalose-6-phosphate phosphatase enhances anaerobic germination tolerance in rice. Nat Plants 1: 15124. [DOI] [PubMed] [Google Scholar]
- Kurotani K, Hayashi K, Hatanaka S, Toda Y, Ogawa D, Ichikawa H, Ishimaru Y, Tashita R, Suzuki T, Ueda M, et al. (2015) Elevated levels of CYP94 family gene expression alleviate the jasmonate response and enhance salt tolerance in rice. Plant Cell Physiol 56: 779–789 [DOI] [PubMed] [Google Scholar]
- Lakshmanan M, Zhang Z, Mohanty B, Kwon JY, Choi HY, Nam HJ, Kim DI, Lee DY (2013) Elucidating rice cell metabolism under flooding and drought stresses using flux-based modeling and analysis. Plant Physiol 162: 2140–2150 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10: R25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lasanthi-Kudahettige R, Magneschi L, Loreti E, Gonzali S, Licausi F, Novi G, Beretta O, Vitulli F, Alpi A, Perata P (2007) Transcript profiling of the anoxic rice coleoptile. Plant Physiol 144: 218–231 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee DC, Sohn HA, Park ZY, Oh S, Kang YK, Lee KM, Kang M, Jang YJ, Yang SJ, Hong YK, et al. (2015) A lactate-induced response to hypoxia. Cell 161: 595–609 [DOI] [PubMed] [Google Scholar]
- Lee Y, Kende H (2001) Expression of beta-expansins is correlated with internodal elongation in deepwater rice. Plant Physiol 127: 645–654 [PMC free article] [PubMed] [Google Scholar]
- Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25: 1754–1760 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li HW, Zang BS, Deng XW, Wang XP (2011a) Overexpression of the trehalose-6-phosphate synthase gene OsTPS1 enhances abiotic stress tolerance in rice. Planta 234: 1007–1018 [DOI] [PubMed] [Google Scholar]
- Li J, Jiang J, Qian Q, Xu Y, Zhang C, Xiao J, Du C, Luo W, Zou G, Chen M, et al. (2011b) Mutation of rice BC12/GDD1, which encodes a kinesin-like protein that binds to a GA biosynthesis gene promoter, leads to dwarfism with impaired cell elongation. Plant Cell 23: 628–640 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Licausi F, Kosmacz M, Weits DA, Giuntoli B, Giorgi FM, Voesenek LA, Perata P, van Dongen JT (2011) Oxygen sensing in plants is mediated by an N-end rule pathway for protein destabilization. Nature 479: 419–422 [DOI] [PubMed] [Google Scholar]
- Licausi F, Ohme-Takagi M, Perata P (2013) APETALA2/Ethylene Responsive Factor (AP2/ERF) transcription factors: mediators of stress responses and developmental programs. New Phytol 199: 639–649 [DOI] [PubMed] [Google Scholar]
- Licausi F, van Dongen JT, Giuntoli B, Novi G, Santaniello A, Geigenberger P, Perata P (2010) HRE1 and HRE2, two hypoxia-inducible ethylene response factors, affect anaerobic responses in Arabidopsis thaliana. Plant J 62: 302–315 [DOI] [PubMed] [Google Scholar]
- Liszkay A, van der Zalm E, Schopfer P (2004) Production of reactive oxygen intermediates (O2.−, H2O2, and .OH) by maize roots and their role in wall loosening and elongation growth. Plant Physiol 136: 3114–3123 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lorenzo O, Piqueras R, Sánchez-Serrano JJ, Solano R (2003) ETHYLENE RESPONSE FACTOR1 integrates signals from ethylene and jasmonate pathways in plant defense. Plant Cell 15: 165–178 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loreti E, van Veen H, Perata P (2016) Plant responses to flooding stress. Curr Opin Plant Biol 33: 64–71 [DOI] [PubMed] [Google Scholar]
- Lunn JE, Delorge I, Figueroa CM, Van Dijck P, Stitt M (2014) Trehalose metabolism in plants. Plant J 79: 544–567 [DOI] [PubMed] [Google Scholar]
- Magneschi L, Perata P (2009) Rice germination and seedling growth in the absence of oxygen. Ann Bot 103: 181–196 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Magome H, Nomura T, Hanada A, Takeda-Kamiya N, Ohnishi T, Shinma Y, Katsumata T, Kawaide H, Kamiya Y, Yamaguchi S (2013) CYP714B1 and CYP714B2 encode gibberellin 13-oxidases that reduce gibberellin activity in rice. Proc Natl Acad Sci USA 110: 1947–1952 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martins MCM, Hejazi M, Fettke J, Steup M, Feil R, Krause U, Arrivault S, Vosloh D, Figueroa CM, Ivakov A, et al. (2013) Feedback inhibition of starch degradation in Arabidopsis leaves mediated by trehalose 6-phosphate. Plant Physiol 163: 1142–1163 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mekhedov SI, Kende H (1996) Submergence enhances expression of a gene encoding 1-aminocyclopropane-1-carboxylate oxidase in deepwater rice. Plant Cell Physiol 37: 531–537 [DOI] [PubMed] [Google Scholar]
- Métraux JP, Kende H (1983) The role of ethylene in the growth response of submerged deep water rice. Plant Physiol 72: 441–446 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Métraux JP, Kende H (1984) The cellular basis of the elongation response in submerged deep-water rice. Planta 160: 73–77 [DOI] [PubMed] [Google Scholar]
- Miro B, Ismail AM (2013) Tolerance of anaerobic conditions caused by flooding during germination and early growth in rice (Oryza sativa L.). Front Plant Sci 4: 269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mizoi J, Ohori T, Moriwaki T, Kidokoro S, Todaka D, Maruyama K, Kusakabe K, Osakabe Y, Shinozaki K, Yamaguchi-Shinozaki K (2013) GmDREB2A;2, a canonical DEHYDRATION-RESPONSIVE ELEMENT-BINDING PROTEIN2-type transcription factor in soybean, is posttranslationally regulated and mediates dehydration-responsive element-dependent gene expression. Plant Physiol 161: 346–361 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Müller M, Munné-Bosch S (2015) Ethylene response factors: a key regulatory hub in hormone and stress signaling. Plant Physiol 169: 32–41 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mustroph A, Lee SC, Oosumi T, Zanetti ME, Yang H, Ma K, Yaghoubi-Masihi A, Fukao T, Bailey-Serres J (2010) Cross-kingdom comparison of transcriptomic adjustments to low-oxygen stress highlights conserved and plant-specific responses. Plant Physiol 152: 1484–1500 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mustroph A, Zanetti ME, Jang CJH, Holtan HE, Repetti PP, Galbraith DW, Girke T, Bailey-Serres J (2009) Profiling translatomes of discrete cell populations resolves altered cellular priorities during hypoxia in Arabidopsis. Proc Natl Acad Sci USA 106: 18843–18848 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagai K, Kondo Y, Kitaoka T, Noda T, Kuroha T, Angeles-Shim RB, Yasui H, Yoshimura A, Ashikari M (2014) QTL analysis of internode elongation in response to gibberellin in deepwater rice. AoB Plants 6: plu028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagai K, Kuroha T, Ayano M, Kurokawa Y, Angeles-Shim RB, Shim JH, Yasui H, Yoshimura A, Ashikari M (2012) Two novel QTLs regulate internode elongation in deepwater rice during the early vegetative stage. Breed Sci 62: 178–185 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakano T, Suzuki K, Fujimura T, Shinshi H (2006) Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol 140: 411–432 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakazono M, Tsuji H, Li Y, Saisho D, Arimura S, Tsutsumi N, Hirai A (2000) Expression of a gene encoding mitochondrial aldehyde dehydrogenase in rice increases under submerged conditions. Plant Physiol 124: 587–598 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Narsai R, Edwards JM, Roberts TH, Whelan J, Joss GH, Atwell BJ (2015) Mechanisms of growth and patterns of gene expression in oxygen-deprived rice coleoptiles. Plant J 82: 25–40 [DOI] [PubMed] [Google Scholar]
- Narsai R, Howell KA, Carroll A, Ivanova A, Millar AH, Whelan J (2009) Defining core metabolic and transcriptomic responses to oxygen availability in rice embryos and young seedlings. Plant Physiol 151: 306–322 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohme-Takagi M, Shinshi H (1995) Ethylene-inducible DNA binding proteins that interact with an ethylene-responsive element. Plant Cell 7: 173–182 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohta M, Matsui K, Hiratsu K, Shinshi H, Ohme-Takagi M (2001) Repression domains of class II ERF transcriptional repressors share an essential motif for active repression. Plant Cell 13: 1959–1968 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pandey GK, Grant JJ, Cheong YH, Kim BG, Li L, Luan S (2005) ABR1, an APETALA2-domain transcription factor that functions as a repressor of ABA response in Arabidopsis. Plant Physiol 139: 1185–1193 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Papdi C, Pérez-Salamó I, Joseph MP, Giuntoli B, Bögre L, Koncz C, Szabados L (2015) The low oxygen, oxidative and osmotic stress responses synergistically act through the ethylene response factor VII genes RAP2.12, RAP2.2 and RAP2.3. Plant J 82: 772–784 [DOI] [PubMed] [Google Scholar]
- Park MH, Suzuki Y, Chono M, Knox JP, Yamaguchi I (2003) CsAGP1, a gibberellin-responsive gene from cucumber hypocotyls, encodes a classical arabinogalactan protein and is involved in stem elongation. Plant Physiol 131: 1450–1459 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perata P, Loreti E, Guglielminetti L, Alpi A (1998) Carbohydrate metabolism and anoxia tolerance in cereal grains. Acta Bot Neerl 47: 269–283 [Google Scholar]
- Perrot-Rechenmann C. (2010) Cellular responses to auxin: division versus expansion. Cold Spring Harb Perspect Biol 2: a001446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ponnu J, Wahl V, Schmid M (2011) Trehalose-6-phosphate: connecting plant metabolism and development. Front Plant Sci 2: 70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Popescu SC, Popescu GV, Bachan S, Zhang Z, Gerstein M, Snyder M, Dinesh-Kumar SP (2009) MAPK target networks in Arabidopsis thaliana revealed using functional protein microarrays. Genes Dev 23: 80–92 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qi W, Sun F, Wang Q, Chen M, Huang Y, Feng YQ, Luo X, Yang J (2011) Rice ethylene-response AP2/ERF factor OsEATB restricts internode elongation by down-regulating a gibberellin biosynthetic gene. Plant Physiol 157: 216–228 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramsak Ž, Baebler Š, Rotter A, Korbar M, Mozetic I, Usadel B, Gruden K (2014) GoMapMan: integration, consolidation and visualization of plant gene annotations within the MapMan ontology. Nucleic Acids Res 42: D1167–D1175 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raskin I, Kende H (1984) Role of gibberellin in the growth response of submerged deep water rice. Plant Physiol 76: 947–950 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richards DE, King KE, Ait-Ali T, Harberd NP (2001) How gibberellin regulates plant growth and development: a molecular genetic analysis of gibberellin signaling. Annu Rev Plant Physiol Plant Mol Biol 52: 67–88 [DOI] [PubMed] [Google Scholar]
- Riechmann JL, Meyerowitz EM (1998) The AP2/EREBP family of plant transcription factors. Biol Chem 379: 633–646 [DOI] [PubMed] [Google Scholar]
- Rivera-Contreras IK, Zamora-Hernández T, Huerta-Heredia AA, Capataz-Tafur J, Barrera-Figueroa BE, Juntawong P, Peña-Castro JM (2016) Transcriptomic analysis of submergence-tolerant and sensitive Brachypodium distachyon ecotypes reveals oxidative stress as a major tolerance factor. Sci Rep 6: 27686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roberts JKM, Callis J, Jardetzky O, Walbot V, Freeling M (1984) Cytoplasmic acidosis as a determinant of flooding intolerance in plants. Proc Natl Acad Sci USA 81: 6029–6033 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26: 139–140 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rudus I, Sasiak M, Kepczynski J (2013) Regulation of ethylene biosynthesis at the level of 1-aminocyclopropane-1-carboxylate oxidase (ACO) gene. Acta Physiol Plant 35: 295–307 [Google Scholar]
- Rzewuski G, Sauter M (2008) Ethylene biosynthesis and signaling in rice. Plant Sci 175: 32–42 [Google Scholar]
- Saika H, Okamoto M, Miyoshi K, Kushiro T, Shinoda S, Jikumaru Y, Fujimoto M, Arikawa T, Takahashi H, Ando M, et al. (2007) Ethylene promotes submergence-induced expression of OsABA8ox1, a gene that encodes ABA 8′-hydroxylase in rice. Plant Cell Physiol 48: 287–298 [DOI] [PubMed] [Google Scholar]
- Sakamoto T, Miura K, Itoh H, Tatsumi T, Ueguchi-Tanaka M, Ishiyama K, Kobayashi M, Agrawal GK, Takeda S, Abe K, et al. (2004) An overview of gibberellin metabolism enzyme genes and their related mutants in rice. Plant Physiol 134: 1642–1653 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakuma Y, Liu Q, Dubouzet JG, Abe H, Shinozaki K, Yamaguchi-Shinozaki K (2002) DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression. Biochem Biophys Res Commun 290: 998–1009 [DOI] [PubMed] [Google Scholar]
- Sasidharan R, Chinnappa CC, Voesenek LA, Pierik R (2008) The regulation of cell wall extensibility during shade avoidance: a study using two contrasting ecotypes of Stellaria longipes. Plant Physiol 148: 1557–1569 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sauter M, Kende H (1992) Levels of β-glucan and lignin in elongating internodes of deepwater rice. Plant Cell Physiol 33: 1089–1097 [Google Scholar]
- Schluepmann H, Pellny T, van Dijken A, Smeekens S, Paul M (2003) Trehalose 6-phosphate is indispensable for carbohydrate utilization and growth in Arabidopsis thaliana. Proc Natl Acad Sci USA 100: 6849–6854 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidt R, Mieulet D, Hubberten HM, Obata T, Hoefgen R, Fernie AR, Fisahn J, San Segundo B, Guiderdoni E, Schippers JHM, et al. (2013) Salt-responsive ERF1 regulates reactive oxygen species-dependent signaling during the initial response to salt stress in rice. Plant Cell 25: 2115–2131 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmitz AJ, Folsom JJ, Jikamaru Y, Ronald P, Walia H (2013) SUB1A-mediated submergence tolerance response in rice involves differential regulation of the brassinosteroid pathway. New Phytol 198: 1060–1070 [DOI] [PubMed] [Google Scholar]
- Shigeto J, Tsutsumi Y (2016) Diverse functions and reactions of class III peroxidases. New Phytol 209: 1395–1402 [DOI] [PubMed] [Google Scholar]
- Si Y, Liu P, Li P, Brutnell TP (2014) Model-based clustering for RNA-seq data. Bioinformatics 30: 197–205 [DOI] [PubMed] [Google Scholar]
- Singh HP, Singh BB, Ram PC (2001) Submergence tolerance of rainfed lowland rice: search for physiological marker traits. J Plant Physiol 158: 883–889 [Google Scholar]
- Singh P, Sinha AK (2016) A positive feedback loop governed by SUB1A1 interaction with MITOGEN-ACTIVATED PROTEIN KINASE3 imparts submergence tolerance in rice. Plant Cell 28: 1127–1143 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Solano R, Stepanova A, Chao Q, Ecker JR (1998) Nuclear events in ethylene signaling: a transcriptional cascade mediated by ETHYLENE-INSENSITIVE3 and ETHYLENE-RESPONSE-FACTOR1. Genes Dev 12: 3703–3714 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spartz AK, Ren H, Park MY, Grandt KN, Lee SH, Murphy AS, Sussman MR, Overvoorde PJ, Gray WM (2014) SAUR inhibition of PP2C-D phosphatases activates plasma membrane H+-ATPases to promote cell expansion in Arabidopsis. Plant Cell 26: 2129–2142 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stünzi JT, Kende H (1989) Gas composition in the internal air spaces of deepwater rice in relation to growth induced by submergence. Plant Cell Physiol 30: 49–56 [Google Scholar]
- Sun TP. (2011) The molecular mechanism and evolution of the GA-GID1-DELLA signaling module in plants. Curr Biol 21: R338–R345 [DOI] [PubMed] [Google Scholar]
- Suzuki S, Suzuki Y, Yamamoto N, Hattori T, Sakamoto M, Umezawa T (2009) High-throughput determination of thioglycolic acid lignin from rice. Plant Biotechnol 26: 337–340 [Google Scholar]
- Tamang BG, Fukao T (2015) Plant adaptation to multiple stresses during submergence and following desubmergence. Int J Mol Sci 16: 30164–30180 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thimm O, Bläsing O, Gibon Y, Nagel A, Meyer S, Krüger P, Selbig J, Müller LA, Rhee SY, Stitt M (2004) MAPMAN: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J 37: 914–939 [DOI] [PubMed] [Google Scholar]
- van der Knaap E, Kim JH, Kende H (2000) A novel gibberellin-induced gene from rice and its potential regulatory role in stem growth. Plant Physiol 122: 695–704 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Der Straeten D, Zhou Z, Prinsen E, Van Onckelen HA, Van Montagu MC (2001) A comparative molecular-physiological study of submergence response in lowland and deepwater rice. Plant Physiol 125: 955–968 [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Hengel AJ, Roberts K (2002) Fucosylated arabinogalactan-proteins are required for full root cell elongation in Arabidopsis. Plant J 32: 105–113 [DOI] [PubMed] [Google Scholar]
- Van Sandt VST, Suslov D, Verbelen JP, Vissenberg K (2007) Xyloglucan endotransglucosylase activity loosens a plant cell wall. Ann Bot 100: 1467–1473 [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Veen H, Mustroph A, Barding GA, Vergeer-van Eijk M, Welschen-Evertman RA, Pedersen O, Visser EJ, Larive CK, Pierik R, Bailey-Serres J, et al. (2013) Two Rumex species from contrasting hydrological niches regulate flooding tolerance through distinct mechanisms. Plant Cell 25: 4691–4707 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Voesenek LA, Bailey-Serres J (2015) Flood adaptive traits and processes: an overview. New Phytol 206: 57–73 [DOI] [PubMed] [Google Scholar]
- Voesenek LA, Colmer TD, Pierik R, Millenaar FF, Peeters AJ (2006) How plants cope with complete submergence. New Phytol 170: 213–226 [DOI] [PubMed] [Google Scholar]
- Wang KLC, Li H, Ecker JR (2002) Ethylene biosynthesis and signaling networks. Plant Cell (Suppl) 14: S131–S151 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang L, Wang Z, Xu Y, Joo S-H, Kim S-K, Xue Z, Xu Z, Wang Z, Chong K (2009) OsGSR1 is involved in crosstalk between gibberellins and brassinosteroids in rice. Plant J 57: 498–510 [DOI] [PubMed] [Google Scholar]
- Wang M, Zhu Z, Tan L, Liu F, Fu Y, Sun C, Cai H (2013) Complexity of indica-japonica varietal differentiation in Bangladesh rice landraces revealed by microsatellite markers. Breed Sci 63: 227–232 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wasternack C, Hause B (2013) Jasmonates: biosynthesis, perception, signal transduction and action in plant stress response, growth and development. An update to the 2007 review in Annals of Botany. Ann Bot 111: 1021–1058 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss D, Ori N (2007) Mechanisms of cross talk between gibberellin and other hormones. Plant Physiol 144: 1240–1246 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wickham H. (2009) ggplot2: Elegant Graphics for Data Analysis. Springer, New York [Google Scholar]
- Wilson K, Long D, Swinburne J, Coupland G (1996) A Dissociation insertion causes a semidominant mutation that increases expression of TINY, an Arabidopsis gene related to APETALA2. Plant Cell 8: 659–671 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu K, Xu X, Fukao T, Canlas P, Maghirang-Rodriguez R, Heuer S, Ismail AM, Bailey-Serres J, Ronald PC, Mackill DJ (2006) Sub1A is an ethylene-response-factor-like gene that confers submergence tolerance to rice. Nature 442: 705–708 [DOI] [PubMed] [Google Scholar]
- Yadav UP, Ivakov A, Feil R, Duan GY, Walther D, Giavalisco P, Piques M, Carillo P, Hubberten HM, Stitt M, et al. (2014) The sucrose-trehalose 6-phosphate (Tre6P) nexus: specificity and mechanisms of sucrose signalling by Tre6P. J Exp Bot 65: 1051–1068 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamaguchi S. (2008) Gibberellin metabolism and its regulation. Annu Rev Plant Biol 59: 225–251 [DOI] [PubMed] [Google Scholar]
- Yang DL, Yao J, Mei CS, Tong XH, Zeng LJ, Li Q, Xiao LT, Sun TP, Li J, Deng XW, et al. (2012) Plant hormone jasmonate prioritizes defense over growth by interfering with gibberellin signaling cascade. Proc Natl Acad Sci USA 109: E1192–E1200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang SF, Hoffman NE (1984) Ethylene biosynthesis and its regulation in higher plants. Annu Rev Plant Physiol 35: 155–189 [Google Scholar]
- Yang SH, Choi D (2006) Characterization of genes encoding ABA 8′-hydroxylase in ethylene-induced stem growth of deepwater rice (Oryza sativa L.). Biochem Biophys Res Commun 350: 685–690 [DOI] [PubMed] [Google Scholar]
- Zarembinski TI, Theologis A (1997) Expression characteristics of OS-ACS1 and OS-ACS2, two members of the 1-aminocyclopropane-1-carboxylate synthase gene family in rice (Oryza sativa L. cv. Habiganj Aman II) during partial submergence. Plant Mol Biol 33: 71–77 [DOI] [PubMed] [Google Scholar]
- Zhang H, Zhang J, Quan R, Pan X, Wan L, Huang R (2013) EAR motif mutation of rice OsERF3 alters the regulation of ethylene biosynthesis and drought tolerance. Planta 237: 1443–1451 [DOI] [PubMed] [Google Scholar]
- Zhou X, Zhang ZL, Park J, Tyler L, Yusuke J, Qiu K, Nam EA, Lumba S, Desveaux D, McCourt P, et al. (2016) The ERF11 transcription factor promotes internode elongation by activating gibberellin biosynthesis and signaling. Plant Physiol 171: 2760–2770 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu G, Ye N, Zhang J (2009) Glucose-induced delay of seed germination in rice is mediated by the suppression of ABA catabolism rather than an enhancement of ABA biosynthesis. Plant Cell Physiol 50: 644–651 [DOI] [PubMed] [Google Scholar]
- Zhu Q, Zhang J, Gao X, Tong J, Xiao L, Li W, Zhang H (2010) The Arabidopsis AP2/ERF transcription factor RAP2.6 participates in ABA, salt and osmotic stress responses. Gene 457: 1–12 [DOI] [PubMed] [Google Scholar]
- Zhu Y, Nomura T, Xu Y, Zhang Y, Peng Y, Mao B, Hanada A, Zhou H, Wang R, Li P, et al. (2006) ELONGATED UPPERMOST INTERNODE encodes a cytochrome P450 monooxygenase that epoxidizes gibberellins in a novel deactivation reaction in rice. Plant Cell 18: 442–456 [DOI] [PMC free article] [PubMed] [Google Scholar]