Abstract
Breast cancer is the most common cancer among women in the United States with up to 30% of those diagnosed displaying a family history of breast cancer. To date, 18% of the familial risk of breast cancer can be explained by common single nucleotide polymorphisms (SNPs). This review summarizes the discovery of risk-associated SNPs using candidate gene and genome-wide association studies (GWAS), including discovery and replication in large collaborative efforts such as COGS and OncoArray. We discuss the evolution of GWAS studies, efforts to discover additional SNPs, and methods for identifying causal variants. We summarize findings associated with overall breast cancer, pathologic subtypes, and mutation carriers (BRCA1, BRCA2, and CHEK2). In addition, we summarize the development of polygenic risk scores (PRS) using the risk-associated SNPs and show how PRS can contribute to estimation of individual risks for developing breast cancer.
Introduction
Breast cancer is the most common cancer among women in the United States, accounting for 15 percent of all new cancer cases. An estimated 15 to 30% of breast cancer cases are heritable or due to underlying genetic transmission, but the genetic alterations accounting for these breast cancers are not fully defined (1, 2). Mutations in dominant high and moderate penetrance breast cancer susceptibility genes, such as BRCA1, BRCA2, PALB2, ATM, and CHEK2 have been identified in 5% of breast cancer cases in the general population and only 30–40% of cases associated with a family history of breast cancer (3–7). Over the last decade, efforts to explain the missing heritability of breast cancer have focused on discovery of other moderate and high-risk genes as well as identification of common genetic variants. Investigations of common variants by genome-wide association studies (GWAS) have successfully identified many genetic loci that are associated with breast cancer risk, and explain up to 18% heritability, suggesting that breast cancer is a complex, polygenic disease.
Initial association studies of common genetic variants
Early investigations of common genetic risk factors for breast cancer focused on candidate gene and candidate variant studies using hundreds of breast cancer cases and unaffected controls. The majority of candidate variants associated with breast cancer in these studies have subsequently failed replication and have therefore been excluded as risk factors for breast cancer (8). Only a coding variant (D302H) in the caspase 8 gene (CASP8) has consistently shown associations with breast cancer, and ongoing studies have recently identified multiple independent risk-associated signals in this locus (9, 10).
Earliest breast cancer GWAS (2007–2013)
Beginning in 2005 with the development of large collaborative groups such as the Cancer Genetic Markers of Susceptibility (CGEMS) Breast Cancer Consortium and the Breast Cancer Association Consortium (BCAC), studies increased in size and statistical power and used genome-wide association studies (GWAS) to identify single nucleotide polymorphisms (SNPs) linked to increased breast cancer risk (11, 12). In 2007, Easton and colleagues identified five breast cancer risk loci (FGFR2—rs2981582, 8q24—rs13281615, LSP1—rs3817198, TNRC9—rs3803662, and MAP3K1—rs889312) in a two-stage GWAS study of Caucasian women followed by a third replication stage (12). The initial GWAS phase evaluated 227,876 SNPs among only 390 breast cancer cases and 364 controls from the UK, but replication in the second and third stages involved many thousands of cases and controls.
FGFR2: the first consistently replicated hit
Soon after the ground-breaking publication by Easton and colleagues, additional evidence quickly accumulated implicating the FGFR2 locus in breast cancer risk (11–15). A study by CGEMS showed an increased risk of breast cancer associated with rs1219648 (in high LD with rs2981582) and a related haplotype in FGFR2 in Caucasian post-menopausal women (11). Additionally, the FGFR2 rs2981582 SNP was more strongly associated with ER-positive than ER-negative disease, and was associated with increased risk of breast cancer among BRCA2 mutation carriers (13–15). To this day, SNPs in the FGFR2 locus are still among those with a strong influence on breast cancer risk (rs2981578 OR=1.23; rs35054928 OR=1.27; rs45631563 OR=1.19) (Table 1).
Table 1.
Risk associated with overall breast cancer for 172 SNPs
| Locus | Locus Discovery Reference* | Genes | rs ID | Alleles | MAF# | Refined SNP Reference^ | Subtype& | Overall Breast Cancer Risk (24) | |||
|---|---|---|---|---|---|---|---|---|---|---|---|
| ORa | CIb | P-valuec | Combined P-valued | ||||||||
| 2q33.1 | (99) | CASP8/ALS2CR12 | rs1830298 | T/C | 0.28 | (100) | Overall | 1.06 | 1.04–1.08 | 1.7 × 10−08 | 1.9 × 10−16 |
|
| |||||||||||
| 5q11.2 | (12) | MAP3K1 | rs62355902 | A/T | 0.16 | (12) | Overall | 1.18 | 1.15–1.21 | 8.5 × 10−42 | 6.8 × 10−98 |
| 8q24.21 | - | rs13281615 | A/G | 0.41 | (12) | Overall | 1.11 | 1.09–1.13 | 5 × 10−28 | 1.9 × 10−57 | |
| 11p15.5 | LSP1 | rs3817198 | T/C | 0.32 | (12) | Overall | 1.05 | 1.03–1.07 | 6.2 × 10−07 | 9.9 × 10−19 | |
| 16q12.1 | TOX3 | rs4784227 | C/T | 0.24 | (101) | Overall | 1.23 | 1.2–1.25 | 7 × 10−88 | 6.8 × 10−201 | |
| 10q26.13 | FGFR2 | rs2981578 | T/C | 0.47 | (102) | Overall | 1.23 | 1.21–1.25 | 1 × 10−114 | 1.3 × 10−245 | |
| 10q26.13 | FGFR2 | rs35054928 | G/GC | 0.4 | (102) | Overall | 1.27 | 1.25–1.3 | 3.5 × 10−154 | 2.3 × 10−322 | |
| 10q26.13 | FGFR2 | rs45631563 | A/T | 0.05 | (102) | Overall | 0.81 | 0.78–0.85 | 9.1 × 10−21 | 7.3 × 10−37 | |
|
| |||||||||||
| 2q35 | (46) | IGFBP5 | rs4442975 | G/T | 0.5 | (103) | ER+ | 0.89 | 0.87–0.9 | 3.6 × 10−40 | 1.1 × 10−95 |
| 2q35 | IGFBP5 | rs34005590 | C/A | 0.05 | (104) | Overall | 0.82 | 0.79–0.86 | 1.6 × 10−19 | 3.2 × 10−41 | |
|
| |||||||||||
| 5p12 | (13) | FGF10/MRPS30 | rs10941679 | A/G | 0.25 | (13) | ER+ | 1.15 | 1.13–1.18 | 6.2 × 10−43 | 5.6 × 10−73 |
|
| |||||||||||
| 3p24.1 | (16) | SLC4A7 | rs4973768 | C/T | 0.47 | (16) | Overall | 1.11 | 1.09–1.13 | 1.7 × 10−28 | 4.8 × 10−57 |
| 17q22 | - | rs2787486 | A/C | 0.3 | (105) | Overall | 0.93 | 0.91–0.94 | 1.2 × 10−14 | 5.6 × 10−29 | |
|
| |||||||||||
| 22q12.1 | (106) | CHEK2 | rs17879961 | A/G | 0.01 | (22) | Overall | 1.26 | 1.11–1.42 | 2.4 × 10−04 | 9.7 × 10−09 |
|
| |||||||||||
| 1p11.2 | (107) | EMBP1 | rs11249433 | A/G | 0.41 | (107) | Overall | 1.11 | 1.09–1.13 | 2.2 × 10−31 | 1.8 × 10−52 |
| 14q24.1 | RAD51B | rs999737 | C/T | 0.23 | (107) | Overall | 0.91 | 0.89–0.93 | 1.1 × 10−18 | 6.5 × 10−39 | |
|
| |||||||||||
| 19p13.11 | (40) | - | rs67397200 | C/G | 0.3 | (108) | BRCA1 | 1.03 | 1.01–1.05 | 4.2 × 10−03 | 1.6 × 10−08 |
|
| |||||||||||
| 9p21.3 | (20) | CDKN2A/CDKN2B | rs1011970 | G/T | 0.16 | (20) | Overall | 1.07 | 1.04–1.09 | 1.4 × 10−07 | 1 × 10−15 |
| 10p15.1 | ANKRD16 | rs2380205 | C/T | 0.44 | (20) | Overall | 0.98 | 0.96–0.99 | 1.1 × 10−02 | 1.7 × 10−04 | |
| 10q22.3 | ZMZ1 | rs704010 | C/T | 0.38 | (20) | Overall | 1.07 | 1.05–1.09 | 1.1 × 10−14 | 1.7 × 10−35 | |
| 11q13.3 | CCND1 | rs554219 | C/G | 0.13 | (97) | ER+ | 1.21 | 1.18–1.24 | 5.8 × 10−47 | 5.8 × 10−47 | |
| 11q13.3 | CCND1 | rs75915166 | C/A | 0.06 | (97) | ER+ | 1.28 | 1.24–1.33 | 4.1 × 10−42 | 4.1 × 10−95 | |
| 6q25 | ESR1 | rs3757322 | T/G | 0.32 | (93) | ER− | 1.08 | 1.06–1.1 | 1.1 × 10−16 | 3.3 × 10−41 | |
| 6q25 | ESR1 | rs9397437 | G/A | 0.07 | (93) | ER− | 1.17 | 1.14–1.21 | 6.3 × 10−21 | 4.8 × 10−54 | |
| 6q25 | ESR1 | rs2747652 | C/T | 0.48 | (93) | ER− | 0.94 | 0.92–0.960 | 1.2 × 10−11 | 1.3 × 10−26 | |
|
| |||||||||||
| 10q21.2 | (109) | ZNF365 | rs10995201 | A/G | 0.16 | (110) | Overall | 0.9 | 0.88–0.92 | 4.7 × 10−17 | 1.6 × 10−51 |
|
| |||||||||||
| 9q31.2 | (21) | - | rs10816625 | A/G | 0.06 | (111) | Overall | 1.11 | 1.07–1.15 | 2.3 × 10−08 | 5 × 10−18 |
| 9q31.2 | - | rs13294895 | C/T | 0.18 | (111) | Overall | 1.06 | 1.03–1.08 | 1.9 × 10−06 | 6.5 × 10−17 | |
| 9q31.2 | - | rs676256 | T/C | 0.38 | (111) | Overall | 0.91 | 0.9–0.93 | 1.9 × 10−21 | 3.5 × 10−53 | |
|
| |||||||||||
| 5p15.33 | (47) | TERT | rs10069690 | C/T | 0.26 | (47) | ER− | 1.06 | 1.04–1.08 | 2.5 × 10−08 | 7.8 × 10−17 |
|
| |||||||||||
| 12q24.21 | (19) | TBX3 | rs1292011 | A/G | 0.42 | (19) | Overall | 0.92 | 0.9–0.94 | 2.4 × 10−19 | 4.4 × 10−39 |
| 21q21.1 | NRIP1 | rs2823093 | G/A | 0.27 | (19) | Overall | 0.94 | 0.92–0.96 | 1.1 × 10−09 | 1.5 × 10−20 | |
|
| |||||||||||
| 6q25.1 | (112) | TAB2 | rs9485372 | G/A | 0.19 | (112) | Overall | 0.96 | 0.93–0.98 | 7.7 × 10−05 | 3.5 × 10−06 |
|
| |||||||||||
| 6q14.1 | (113) | - | rs17529111 | T/C | 0.22 | (113) | ER− | 1.02 | 1–1.04 | 4.2 × 10−02 | 1.3 × 10−09 |
| 20q11.22 | RALY | rs2284378 | C/T | 0.32 | (113) | Overall | 1 | 0.98–1.02 | 7.9 × 10−01 | 3.2 × 10−02 | |
|
| |||||||||||
| 5p15.33 | (92) | TERT | rs3215401 | A/AG | 0.31 | (92) | Overall | 0.93 | 0.91–0.95 | 7.1 × 10−13 | 1.1 × 10−20 |
|
| |||||||||||
| 1q32.1 | (48) | LGR6 | rs6678914 | G/A | 0.41 | (48) | Overall | 1 | 0.99–1.02 | 7.3 × 10−01 | 3 × 10−01 |
| 1q32.1 | MDM4 | rs4245739 | A/C | 0.26 | (48) | Overall | 1.02 | 1–1.04 | 2.5 × 10−02 | 1.3 × 10−04 | |
| 2p24.1 | - | rs12710696 | C/T | 0.37 | (48) | Overall | 1.03 | 1.01–1.04 | 6.6 × 10−03 | 1.3 × 10−08 | |
| 16q12.2 | FTO | rs11075995 | T/A | 0.24 | (48) | Overall | 1.03 | 1.01–1.06 | 1.3 × 10−03 | 8.7 × 10−09 | |
|
| |||||||||||
| 6p24.3 | (114) | TFAP2A | rs9348512 | C/A | 0.33 | (114) | BRCA2 | 1 | 0.99–1.02 | 6.4 × 10−01 | 8 × 10−01 |
|
| |||||||||||
| 22q13.1 | (60) | APOBEC3A/APOBEC3B | chr22:39359355 | I/D7 | 0.1 | (60) | Overall | 1.1 | 1.07–1.14 | 6.1 × 10−09 | 4.9 × 10−12 |
|
| |||||||||||
| 1p36.22 | (22) | PEX14 | rs616488 | A/G | 0.33 | (22) | Overall | 0.94 | 0.93–0.96 | 1.8 × 10−09 | 5 × 10−20 |
| 1p13.2 | DCLRE1B | rs11552449 | C/T | 0.17 | (22) | Overall | 1.04 | 1.01–1.06 | 2.3 × 10−03 | 4.6 × 10−11 | |
| 2q14.1 | - | rs4849887 | C/T | 0.1 | (22) | Overall | 0.91 | 0.88–0.94 | 1.2 × 10−10 | 6.9 × 10−20 | |
| 2q31.1 | DLX2-AS1 | rs2016394 | G/A | 0.47 | (22) | Overall | 0.95 | 0.94–0.97 | 3.1 × 10−07 | 6.2 × 10−12 | |
| 2q31.1 | CDCA7 | rs1550623 | A/G | 0.15 | (22) | Overall | 0.95 | 0.93–0.98 | 8.8 × 10−05 | 5.4 × 10−10 | |
| 2q35 | DIRC3 | rs16857609 | C/T | 0.26 | (22) | Overall | 1.06 | 1.04–1.09 | 1.4 × 10−09 | 1.8 × 10−25 | |
| 3p26.1 | EGPT/ITPR1 | rs6762644 | A/G | 0.38 | (22) | Overall | 1.05 | 1.03–1.07 | 1.6 × 10−08 | 4 × 10−18 | |
| 3p24.1 | TGFBR2 | rs12493607 | G/C | 0.34 | (22) | Overall | 1.05 | 1.03–1.07 | 4.9 × 10−07 | 6.9 × 10−14 | |
| 4q24 | TET2 | rs9790517 | C/T | 0.23 | (22) | Overall | 1.04 | 1.01–1.06 | 1.1 × 10−03 | 5 × 10−11 | |
| 4q34.1 | ADAM29 | rs6828523 | C/A | 0.12 | (22) | Overall | 0.91 | 0.88–0.93 | 1.2 × 10−11 | 1.8 × 10−25 | |
| 5q11.2 | RAB3C | rs10472076 | T/C | 0.38 | (22) | Overall | 1.03 | 1.01–1.04 | 6.6 × 10−03 | 9.6 × 10−09 | |
| 5q11.2 | PDE4D | rs1353747 | T/G | 0.09 | (22) | Overall | 0.96 | 0.93–0.99 | 7.6 × 10−03 | 4.1 × 10−09 | |
| 5q33.3 | EBF1 | rs1432679 | T/C | 0.43 | (22) | Overall | 1.08 | 1.06–1.1 | 2.9 × 10−17 | 6.6 × 10−31 | |
| 6p25.3 | FOXQ1 | rs11242675 | T/C | 0.37 | (22) | Overall | 1 | 0.98–1.02 | 9.7 × 10−01 | 1 × 10−04 | |
| 6p23 | RANBP9 | rs204247 | A/G | 0.44 | (22) | Overall | 1.04 | 1.02–1.06 | 6.4 × 10−05 | 7.9 × 10−13 | |
| 7q35 | NOBOX/ARHGEF6 | rs720475 | G/A | 0.25 | (22) | Overall | 0.96 | 0.94–0.98 | 3 × 10−04 | 1.2 × 10−11 | |
| 8p12 | - | rs9693444 | C/A | 0.32 | (22) | Overall | 1.06 | 1.04–1.08 | 1.7 × 10−10 | 1.6 × 10−21 | |
| 8q21.11 | - | rs6472903 | T/G | 0.17 | (22) | Overall | 0.94 | 0.92–0.96 | 4.4 × 10−21 | 4.4 × 10−21 | |
| 8q21.11 | HNF4G | rs2943559 | A/G | 0.08 | (22) | Overall | 1.1 | 1.07–1.14 | 4.2 × 10−09 | 4 × 10−24 | |
| 8q24.21 | MYC | rs11780156 | C/T | 0.17 | (22) | Overall | 1.05 | 1.03–1.08 | 2.5 × 10−05 | 1.1 × 10−13 | |
| 9q31.2 | - | rs10759243 | C/A | 0.29 | (22) | Overall | 1.06 | 1.04–1.08 | 4.2 × 10−10 | 2.2 × 10−18 | |
| 10p12.31 | DNAJC1 | rs7072776 | G/A | 0.29 | (22) | Overall | 1.05 | 1.03–1.07 | 2.7 × 10−07 | 1.8 × 10−19 | |
| 10p12.31 | DNAJC1 | rs11814448 | A/C | 0.02 | (22) | Overall | 1.12 | 1.06–1.19 | 1.4 × 10−06 | 6.1 × 10−18 | |
| 10q25.2 | TCFL2 | rs7904519 | A/G | 0.46 | (22) | Overall | 1.03 | 1.01–1.05 | 8.6 × 10−04 | 1.5 × 10−13 | |
| 10q26.12 | (22) | - | rs11199914 | C/T | 0.32 | (22) | Overall | 0.96 | 0.94–0.98 | 2.1 × 10−05 | 6.5 × 10−12 |
| 11q13.1 | - | rs3903072 | G/T | 0.47 | (22) | Overall | 0.97 | 0.95–0.97 | 9.1 × 10−04 | 2.3 × 10−12 | |
| 11q24.3 | - | rs11820646 | C/T | 0.4 | (22) | Overall | 0.96 | 0.94–0.98 | 2.5 × 10−05 | 2.1 × 10−14 | |
| 12p13.1 | - | rs12422552 | G/C | 0.26 | (22) | Overall | 1.06 | 1.04–1.08 | 3.2 × 10−08 | 3.6 × 10−15 | |
| 12q22 | NTN4 | rs17356907 | A/G | 0.3 | (22) | Overall | 0.91 | 0.9–0.93 | 6.6 × 10−20 | 1 × 10−39 | |
| 13q13.1 | BRCA2 | rs11571833 | A/T | 0.01 | (22) | Overall | 1.35 | 1.23–1.48 | 4 × 10−10 | 3.1 × 10−15 | |
| 14q13.3 | PAX9 | rs2236007 | G/A | 0.21 | (22) | Overall | 0.93 | 0.91–0.95 | 5.8 × 10−10 | 4.2 × 10−21 | |
| 14q24.1 | RAD51B | rs2588809 | C/T | 0.17 | (22) | Overall | 1.06 | 1.03–1.08 | 2.6 × 10−06 | 6.3 × 10−14 | |
| 14q32.11 | CCDC88C | rs941764 | A/G | 0.35 | (22) | Overall | 1.03 | 1.02–1.05 | 3.6 × 10−04 | 8.2 × 10−13 | |
| 16q12.2 | FTO | rs17817449 | T/G | 0.41 | (22) | Overall | 0.95 | 0.93–0.96 | 4.9 × 10−09 | 2.5 × 10−21 | |
| 16q23.2 | CDYL2 | rs13329835 | A/G | 0.23 | (22) | Overall | 1.07 | 1.05–1.09 | 8.3 × 10−11 | 8.8 × 10−27 | |
| 18q11.2 | - | rs527616 | G/C | 0.38 | (22) | Overall | 0.97 | 0.95–0.98 | 2.8 × 10−04 | 6.7 × 10−15 | |
| 18q11.2 | CHST9 | rs1436904 | T/G | 0.4 | (22) | Overall | 0.95 | 0.94–0.97 | 1.1 × 10−07 | 9.9 × 10−15 | |
| 19p13.11 | ELL | rs4808801 | A/G | 0.34 | (22) | Overall | 0.93 | 0.91–0.95 | 2 × 10−12 | 4.7 × 10−28 | |
| 19q13.31 | KCCN4/LYPD5 | rs3760982 | G/A | 0.46 | (22) | Overall | 1.05 | 1.03–1.07 | 2.1 × 10−08 | 1.4 × 10−16 | |
| 22q12.2 | EM1D1 | rs132390 | T/C | 0.04 | (22) | Overall | 1.04 | 0.99–1.09 | 8.1 × 10−02 | 1.2 × 10−08 | |
| 22q13.1 | MKL1 | rs6001930 | T/C | 0.1 | (22) | Overall | 1.12 | 1.09–1.16 | 5.7 × 10−16 | 4.4 × 10−34 | |
|
| |||||||||||
| 1q32.1 | (72) | ZC3H11A | rs4951011 | A/G | 0.16 | (72) | Overall | 1.04 | 1.02–1.07 | 5.4 × 10−04 | 1.3 × 10−05 |
| 5q14.3 | ARRDC3 | rs10474352 | C/T | 0.16 | (72) | Overall | 0.94 | 0.92–0.97 | 4.5 × 10−11 | 4.5 × 10−11 | |
| 15q26.1 | PRC1 | rs2290203 | G/A | 0.21 | (72) | Overall | 0.94 | 0.92–0.96 | 1.8 × 10−07 | 8.07 × 10−10 | |
|
| |||||||||||
| 3p14.1 | (115) | ATNX7 | rs1053338 | A/G | 0.14 | (115) | Overall | 1.05 | 1.02–1.07 | 5 × 10−04 | 5.3 × 10−11 |
| 7q21.2 | AKAP9 | rs6964587 | G/T | 0.39 | (115) | Overall | 1.03 | 1.02–1.05 | 2.2 × 10−04 | 9 × 10−11 | |
|
| |||||||||||
| 7q34 | (116) | - | rs11977670 | G/A | 0.43 | (116) | Lobular | 1.06 | 1.04–1.08 | 1.9 × 10−11 | 1 × 10−16 |
|
| |||||||||||
| 1q21.1 | (23) | RNF115 | rs12405132 | C/T | 0.37 | (23) | Overall | 0.97 | 0.95–0.99 | 1 × 10−03 | 6.3 × 10−10 |
| 1q21.2 | OTUD7B | rs12048493 | A/C | 0.38 | (23) | Overall | 1.04 | 1.02–1.06 | 9.4 × 10−06 | 8.6 × 10−14 | |
| 1q43 | EXO1 | rs72755295 | A/G | 0.03 | (23) | Overall | 1.15 | 1.09–1.2 | 8 × 10−08 | 1.7 × 10−14 | |
| 3p21.31 | - | rs6796502 | G/A | 0.1 | (23) | Overall | 0.92 | 0.89–0.95 | 2.5 × 10−08 | 5.5 × 10−15 | |
| 5p15.1 | - | rs13162653 | G/T | 0.45 | (23) | Overall | 0.99 | 0.97–1.010 | 1.8 × 10−01 | 5.4 × 10−07 | |
| 5p13.3 | - | rs2012709 | C/T | 0.48 | (23) | Overall | 1.02 | 1–1.04 | 2.1 × 10−02 | 1.2 × 10−08 | |
| 5q14.2 | ATG10 | rs7707921 | A/T | 0.25 | (23) | Overall | 0.96 | 0.94–0.98 | 1.2 × 10−04 | 1.7 × 10−12 | |
| 6p22.1 | - | rs9257408 | G/C | 0.41 | (23) | Overall | 1.02 | 1–1.040 | 5.5 × 10−02 | 6.9 × 10−08 | |
| 7q32.3 | FLJ43663 | rs4593472 | C/T | 0.35 | (23) | Overall | 0.97 | 0.95–0.99 | 7.9 × 10−04 | 1.8 × 10−11 | |
| 8p11.23 | - | rs13365225 | A/G | 0.18 | (23) | Overall | 0.91 | 0.89–0.93 | 1.2 × 10−15 | 1.4 × 10−20 | |
| 8q23.3 | LINC00536 | rs13267382 | G/A | 0.36 | (23) | Overall | 1.03 | 1.01–1.05 | 8.7 × 10−04 | 1.6 × 10−11 | |
| 14q32.12 | (23) | RIN3 | rs11627032 | T/C | 0.25 | (23) | Overall | 0.96 | 0.94–0.98 | 1.6 × 10−04 | 4.1 × 10−11 |
| 17q11.2 | ATAD5 | rs146699004 | GGT/G | 0.27 | (23) | Overall | 0.97 | 0.95–0.99 | 1.3 × 10−03 | 2 × 10−09 | |
| 17q25.3 | - | rs745570 | G/A | 0.5 | (23) | Overall | 1.03 | 1.01–1.05 | 2.1 × 10−03 | 3.9 × 10−10 | |
| 18q12.3 | SETBP1 | rs6507583 | A/G | 0.07 | (23) | Overall | 0.92 | 0.89–0.96 | 9.5 × 10−06 | 2.2 × 10−12 | |
|
| |||||||||||
| 2p23.2 | (10) | WDR43 | rs4577244 | C/T | 0.23 | (10) | ER-/BRCA1 | 1.01 | 0.99–1.03 | 2.4 × 10−01 | 4.3 × 10−01 |
| 12p11.22 | - | rs7297051 | C/T | 0.24 | (10) | ER-/BRCA1 | 0.89 | 0.87–0.91 | 2.9 × 10−27 | 3 × 10−60 | |
| 13q22.1 | - | rs6562760 | G/A | 0.24 | (10) | ER-/BRCA1 | 0.95 | 0.93–0.97 | 8.6 × 10−06 | 1.5 × 10−09 | |
|
| |||||||||||
| 1p36.13 | (24) | KLHDC7A | rs2992756 | C/T | 0.49 | (24) | Overall | 1.06 | 1.04–1.08 | 1.3 × 10−11 | 1.6 × 10−15 |
| 1p34.2 | - | rs4233486 | T/C | 0.36 | (24) | Overall | 0.97 | 0.95–0.98 | 2.3 × 10−04 | 9.1 × 10−09 | |
| 1p34.2 | HIVEP3 | rs79724016 | T/G | 0.03 | (24) | Overall | 0.93 | 0.88–0.97 | 3.3 × 10−03 | 3.5 × 10−08 | |
| 1p34.1 | PIK3R3, LOC101929626 | rs1707302 | G/A | 0.34 | (24) | Overall | 0.96 | 0.95–0.98 | 1.4 × 10−04 | 3.0 × 10−08 | |
| 1p32.3 | - | rs140850326 | I/D9 | 0.49 | (24) | Overall | 0.97 | 0.95–0.99 | 3.4 × 10−04 | 3.9 × 10−08 | |
| 1p22.3 | - | rs17426269 | G/A | 0.15 | (24) | Overall | 1.05 | 1.02–1.07 | 1.7 × 10−04 | 1.7 × 10−08 | |
| 1p12 | - | rs7529522 | T/C | 0.23 | (24) | Overall | 1.06 | 1.04–1.08 | 1.6 × 10−08 | 1.7 × 10−10 | |
| 1q22 | TRIM46 | rs4971059 | G/A | 0.35 | (24) | Overall | 1.05 | 1.03–1.07 | 3.9 × 10−08 | 4.8 × 10−11 | |
| 1q32.1 | PHLDA3 | rs35383942 | C/T | 0.06 | (24) | Overall | 1.12 | 1.08–1.17 | 12 × 10−09 | 3.8 × 10−13 | |
| 1q41 | ESRRG | rs11117758 | G/A | 0.21 | (24) | Overall | 0.95 | 0.93–0.97 | 7.7 × 10−07 | 3.9 × 10−09 | |
| 2p25.1 | GRHL1 | rs113577745 | C/G | 0.1 | (24) | Overall | 1.08 | 1.05–1.11 | 3.7 × 10−07 | 3.9 × 10−10 | |
| 2p23.3 | ADCY3 | rs6725517 | A/G | 0.41 | (24) | Overall | 0.96 | 0.94–0.98 | 7.5 × 10−06 | 2.9 × 10−12 | |
| 2q13 | BCL2L11 | rs71801447 | CTTATGTT/C | 0.06 | (24) | Overall | 1.09 | 1.05–1.13 | 7.7 × 10−06 | 3.7 × 10−08 | |
| 2q36.3 | - | rs12479355 | A/G | 0.21 | (24) | Overall | 0.96 | 0.94–0.98 | 4.7 × 10−04 | 2.4 × 10−08 | |
| 3p13 | FOXP1 | rs6805189 | T/C | 0.48 | (24) | Overall | 0.97 | 0.95–0.99 | 3.3 × 10−04 | 4.6 × 10−08 | |
| 3p12.1 | VGLL3 | rs13066793 | A/G | 0.09 | (24) | Overall | 0.94 | 0.91–0.97 | 1.5 × 10−04 | 1.0 × 10−09 | |
| 3p12.1 | CMSS1, FILIP1L | rs9833888 | G/T | 0.22 | (24) | Overall | 1.06 | 1.04–1.08 | 2.6 × 10−07 | 5.2 × 10−10 | |
| 3q23 | ZBTB38 | rs34207738 | CTT/C | 0.41 | (24) | Overall | 1.06 | 1.04–1.08 | 1.4 × 10−09 | 3.2 × 10−15 | |
| 3q26.31 | - | rs58058861 | G/A | 0.21 | (24) | Overall | 1.06 | 1.04–1.09 | 1.6 × 10−08 | 1.9 × 10−10 | |
| 4p14 | - | rs6815814 | A/C | 0.26 | (24) | Overall | 1.06 | 1.04–1.08 | 6.1 × 10−08 | 6.1 × 10−13 | |
| 4q21.23 | HELQ | 4:84370124 | TA/TAA | 0.47 | (24) | Overall | 1.04 | 1.02–1.05 | 1.7 × 10−04 | 2.2 × 10−09 | |
| 4q22.1 | LOC105369192 | rs10022462 | C/T | 0.44 | (24) | Overall | 1.04 | 1.02–1.06 | 9.4 × 10−06 | 1.6 × 10−09 | |
| 4q28.1 | - | rs77528541 | G/T | 0.13 | (24) | Overall | 0.95 | 0.92–0.97 | 4.8 × 10−05 | 1.4 × 10−09 | |
| 5p15.33 | AHRR | rs116095464 | T/C | 0.05 | (24) | Overall | 1.06 | 1.02–1.1 | 2.6 × 10−03 | 3.8 × 10−09 | |
| 5q11.1 | - | rs72749841 | T/C | 0.16 | (24) | Overall | 0.93 | 0.91–0.96 | 8.5 × 10−06 | 7.2 × 10−10 | |
| 5q11.1 | - | rs35951924 | A/AT | 0.32 | (24) | Overall | 0.95 | 0.93–0.97 | 4.0 × 10−07 | 1.3 × 10−11 | |
| 5q22.1 | NREP | rs6882649 | T/G | 0.34 | (24) | Overall | 0.97 | 0.95–0.99 | 2.7 × 10−03 | 3.7 × 10−09 | |
| 5q31.1 | (24) | HSPA4 | rs6596100 | C/T | 0.25 | (24) | Overall | 0.94 | 0.92–0.96 | 5.2 × 10−08 | 7.7 × 10−09 |
| 5q35.1 | - | rs4562056 | G/T | 0.33 | (24) | Overall | 1.05 | 1.03–1.07 | 4.1 × 10−07 | 4.7 × 10−10 | |
| 6p22.3 | ATXN1 | rs3819405 | C/T | 0.33 | (24) | Overall | 0.96 | 0.94–0.97 | 2.2 × 10−06 | 1.7 × 10−08 | |
| 6p22.3 | CDKAL1 | rs2223621 | C/T | 0.38 | (24) | Overall | 1.04 | 1.02–1.06 | 1.0 × 10−04 | 3.0 × 10−10 | |
| 6p22.2 | - | rs71557345 | G/A | 0.07 | (24) | Overall | 0.92 | 0.88–0.96 | 8.4 × 10−05 | 3.9 × 10−10 | |
| 6q14.1 | - | rs12207986 | A/G | 0.47 | (24) | Overall | 0.97 | 0.95–0.98 | 2.0 × 10−04 | 1.5 × 10−09 | |
| 6q23.1 | L3MBTL3 | rs6569648 | T/C | 0.24 | (24) | Overall | 0.94 | 0.92–0.96 | 4.8 × 10−08 | 3.0 × 10−12 | |
| 7p15.3 | DNAH11, CDCA7L | rs7971 | A/G | 0.35 | (24) | Overall | 0.96 | 0.94–0.98 | 1.4 × 10−05 | 1.9 × 10−08 | |
| 7p15.1 | CREB5 | rs17156577 | T/C | 0.11 | (24) | Overall | 1.05 | 1.02–1.08 | 3.8 × 10−04 | 4.3 × 10−09 | |
| 7q21.3 | - | rs17268829 | T/C | 0.28 | (24) | Overall | 1.05 | 1.03–1.07 | 1.3 × 10−06 | 4.5 × 10−13 | |
| 7q22.1 | CUX1 | rs71559437 | G/A | 0.12 | (24) | Overall | 0.93 | 0.91–0.96 | 9.1 × 10−07 | 5.1 × 10−12 | |
| 8q22.3 | - | rs514192 | T/A | 0.32 | (24) | Overall | 1.05 | 1.03–1.07 | 3.7 × 10−06 | 5.6 × 10−09 | |
| 8q23.1 | ZFPM3 | rs12546444 | A/T | 0.1 | (24) | Overall | 0.93 | 0.91–0.96 | 5.8 × 10−06 | 7.5 × 10−11 | |
| 8q24.13 | - | rs58847541 | G/A | 0.15 | (24) | Overall | 1.08 | 1.05–1.1 | 7.3 × 10−09 | 5.5 × 10−13 | |
| 9q33.1 | ASTN2 | rs1895062 | A/G | 0.41 | (24) | Overall | 0.94 | 0.92–0.95 | 6.9 × 10−13 | 1.1 × 10−14 | |
| 9q33.3 | LMX1B | rs10760444 | A/G | 0.43 | (24) | Overall | 1.03 | 1.02–1.05 | 2.8 × 10−04 | 9.1 × 10−09 | |
| 9q34.2 | ABO | rs8176636 | I/D10 | 0.2 | (24) | Overall | 1.03 | 1.01–1.06 | 3.2 × 10−03 | 1.4 × 10−08 | |
| 10p14 | - | rs67958007 | TG/T | 0.12 | (24) | Overall | 1.09 | 1.06–1.12 | 1.8 × 10−09 | 1.7 × 10−10 | |
| 10q23.33 | - | rs140936696 | C/CAA | 0.18 | (24) | Overall | 1.04 | 1.02–1.07 | 7.4 × 10−04 | 4.2 × 10−08 | |
| 11p15 | PIDD1 | rs6597981 | G/A | 0.48 | (24) | Overall | 0.96 | 0.94–0.97 | 5.7 × 10−07 | 1.4 × 10−12 | |
| 12q21.31 | - | rs202049448 | T/C | 0.34 | (24) | Overall | 0.95 | 0.93–0.97 | 2.5 × 10−07 | 2.7 × 10−08 | |
| 12q24.31 | - | rs206966 | C/T | 0.16 | (24) | Overall | 1.05 | 1.02–1.07 | 2.7 × 10−04 | 3.8 × 10−08 | |
| 14q32.33 | ADSSL1 | rs10623258 | C/CTT | 0.45 | (24) | Overall | 1.04 | 1.02–1.06 | 2.7 × 10−05 | 2.3 × 10−08 | |
| 16q12.2 | - | rs28539243 | G/A | 0.49 | (24) | Overall | 1.05 | 1.03–1.07 | 3.6 × 10−08 | 9.1 × 10−15 | |
| 16q13 | AMFR | rs2432539 | G/A | 0.4 | (24) | Overall | 1.03 | 1.02–1.05 | 3.1 × 10−04 | 4.0 × 10−08 | |
| 16q24.2 | - | rs4496150 | C/A | 0.25 | (24) | Overall | 0.96 | 0.94–0.98 | 3.4 × 10−05 | 8.1 × 10−09 | |
| 17q21.2 | CNTNAP1 | rs72826962 | C/T | 0.01 | (24) | Overall | 1.2 | 1.11–1.3 | 5.1 × 10−06 | 4.6 × 10−09 | |
| 17q21.31 | KANSL1 | rs2532263 | G/A | 0.19 | (24) | Overall | 0.95 | 0.93–0.97 | 4.7 × 10−06 | 6.9 × 10−13 | |
| 18q12.1 | GAREM1 | rs117618124 | T/C | 0.05 | (24) | Overall | 0.89 | 0.85–0.92 | 4.5 × 10−08 | 5.5 × 10−12 | |
| 19p13.13 | NFIX1 | rs78269692 | T/C | 0.05 | (24) | Overall | 1.09 | 1.04–1.13 | 3.9 × 10−05 | 1.9 × 10−09 | |
| 19p13.12 | - | rs2594714 | G/A | 0.23 | (24) | Overall | 0.97 | 0.95–0.99 | 6.7 × 10−03 | 1.1 × 10−08 | |
| 19p13.11 | GATAD2A, MIR640 | rs2965183 | G/A | 0.35 | (24) | Overall | 1.04 | 1.02–1.06 | 9.6 × 10−06 | 6.3 × 10−12 | |
| 19q13.22 | GIPR | rs71338792 | A/AT | 0.23 | (24) | Overall | 1.05 | 1.03–1.07 | 8.1 × 10−06 | 3.5 × 10−09 | |
| 20p12.3 | MCM8 | rs16991615 | G/A | 0.06 | (24) | Overall | 1.1 | 1.06–1.14 | 1.4 × 10−07 | 1.9 × 10−09 | |
| 20q13.13 | (24) | - | rs6122906 | A/G | 0.18 | (24) | Overall | 1.05 | 1.03–1.07 | 2.9 × 10−05 | 2.5 × 10−10 |
| 22q13.1 | PLA2G6 | rs738321 | C/G | 0.38 | (24) | Overall | 0.95 | 0.93–0.97 | 2.7 × 10−08 | 1.0 × 10−13 | |
| 22q13.2 | XRCC6 | rs73161324 | C/T | 0.06 | (24) | Overall | 1.06 | 1.02–1.09 | 3.8 × 10−03 | 2.0 × 10−09 | |
| 22q13.31 | - | rs28512361 | G/A | 0.11 | (24) | Overall | 1.05 | 1.02–1.08 | 5.7 × 10−04 | 2.3 × 10−08 | |
Risk locus was originally discovered in the associated reference
MAF reported in OncoArray controls
Reference for the refined SNP of interest
Breast cancer subtype in which the refined SNP was evaluated
Odds ratio (OR) reported for overall breast cancer OncoArray analysis
95% Confidence Interval (CI) reported for overall breast cancer OncoArray analysis
p-value reported for overall breast cancer OncoArray analysis
Combined meta-analysis p-value reported for overall breast cancer
Early strategies to identify additional susceptibility loci
Though most early GWAS studies had small sample sizes by today’s standards, studies improved power by enriching for family history, performing meta-analyses, and evaluating marginally significant GWAS SNPs in larger populations. For example, 925 SNPs displaying near genome-wide significant association with breast cancer in the initial study by Easton et al were further evaluated by BCAC using 37,012 cases and 40,069 controls resulting in two new susceptibility loci at 3p24 near SLC4A7 and NEK10 and at 17q23.2 near COX11 (12, 16). Additional efforts by the Shanghai Breast Cancer study in a three-stage study also identified risk-associated SNPs in the ESR1 locus; these SNPs were replicated in an independent study of European women from BCAC (1591 cases and 1466 controls) (17). Thus, the initial phase of GWAS using limited numbers of cases and controls in one, two, or three stage GWAS designs and the examination of marginally significant loci from these initial GWAS in larger sample sizes resulted in identification of the first 14 loci containing SNPs displaying genome-wide associations (p<1×10−6) with breast cancer and conferring low to moderate risk of disease (odds ratio (OR)=1.07 to 1.29) in European populations (18). Furthermore, 72 potential breast cancer SNPs that did not reach genome-wide significance in pooled data from two GWAS (UK2 and British Breast Cancer Study (BBCS)) were evaluated using cases and controls from 41 studies in BCAC and 9 individual GWAS. One SNP on 12p11 near PTHLH (rs10771399) was associated with overall breast cancer risk, and two others (rs1292011on 12q24; rs2823093 on 21q21 near NRIP1) associated with ER-positive breast cancer risk (19). Subsequently, five additional breast cancer susceptibility loci were identified in a multi-stage GWAS that included 3,659 cases with a family history of breast cancer and 4,897 controls (20). Similarly, Fletcher and colleagues identified a new susceptibility locus at 9q31.2 in a GWAS including 1,694 cases with a personal and/or family history of breast cancer compared to 2,365 controls (21).
SNP Discovery: COGS era (2012–2015)
The Collaborative Oncologic Gene-environment Study (COGS) was a multi-consortium initiative to study prostate, ovarian, and breast cancer using a customized iSelect SNP genotyping array from Illumina (iCOGS) in which more than 200,000 SNPs were tested in each sample. In 2013, Michailidou and colleagues reported on the evaluation of 199,961 SNPs across 52,675 breast cancer cases and 49,436 controls from BCAC (22). At that time, 27 breast cancer risk loci had already been identified. Twenty-three of the 27 loci were replicated in COGS, 3 were more weakly associated with breast cancer risk, and 1 SNP was not included on the iCOGS genotyping array. In addition, 41 new breast cancer susceptibility loci were identified, bringing the total number of breast cancer susceptibility loci to 68. Thirteen of the newly identified SNPs were more strongly associated with ER-positive breast cancer, while one SNP was associated with ER-negative breast cancer. The 68 susceptibility SNPs were estimated to account for ~14% of familial relative risk (risk of breast cancer in first degree relatives of women with breast cancer).
By 2015, 79 breast cancer susceptibility loci had been published, and 71 of these were confirmed in a 2015 meta-analysis including the COGS data from BCAC and 11 additional GWAS (23). The meta-analysis included 62,533 breast cancer cases and 60,976 controls. Using the 1000 Genomes Project, which catalogues a haplotype map for 38 million SNPs, genotypes were imputed for ~11.6 million SNPs, with imputation r2>0.3, that had not been included on the original arrays. After excluding SNPs within 500kb of previously identified SNPs, 15 new breast cancer susceptibility loci were identified. This increased the total number of independent signals in known susceptibility loci to 94. Variation in these loci was estimated to account for ~16% of familial breast cancer risk.
SNP Discovery: OncoArray era (2015-present)
The most recent and largest breast cancer GWAS used the Illumina OncoArray BeadChip, which included approximately 570,000 SNPs and was imputed to ~11.8 million SNPs (imputation r2>0.3) (24, 25). From BCAC, 61,282 breast cancer cases and 45,494 controls of European ancestry were genotyped on the OncoArray platform and results were used in a meta-analysis including data from iCOGS and 11 GWAS studies (24). In total, 122,977 breast cancer cases and 105,974 controls of European ancestry were analyzed along with 14,068 breast cancer cases and 13,104 controls of East Asian ancestry. In this study, SNPs from 49 previously identified breast cancer loci were replicated with genome-wide significance (p<5×10−8), and SNPs from another 50 known risk loci did not reach but were associated with breast cancer (p<0.05) (Table 1). In addition, 65 new breast cancer risk loci (also genome-wide significant) were identified for women of European ancestry (Table 1). Of these, 19 SNPs were associated more strongly with ER-positive disease, and 2 SNPs were associated more strongly with ER-negative disease (Supplemental Table 1). The total 172 risk-associated SNPs account for an estimated ~18% of familial relative risk (Table 1).
Divide and conquer: SNPs associated with breast cancer subtypes and mutation carriers
Studies of more homogeneous populations have been conducted to increase the statistical power of analyses, including BRCA1 and BRCA2 mutation carriers and breast cancer subtypes, such as ER-negative disease and triple negative breast cancer (TNBC).
BRCA1 and BRCA2 mutation carriers
In 2007, the CIMBA consortium was formed with the goal of investigating genetic modifiers of cancer risk in BRCA1 and BRCA2 mutation carriers (26). Using up to 30,000 BRCA1 and BRCA2 mutation carriers with breast or ovarian cancer or unaffected by these diseases, CIMBA evaluated SNPs discovered in previous candidate SNP studies and GWAS including FGFR2, TNRC9, RAD51, TP53, ERCC4, CASP8, ZRCC1, IRS1, and a locus on chromosome 6p22 (15, 27–39). SNP associations were not always consistent among BRCA1 and BRCA2 mutation carriers. For example, the FGFR2 locus was found to be associated with breast cancer in BRCA2 mutation carriers only, whereas TNRC9 was associated with both BRCA1 and BRCA2 breast cancer (15).
The first GWAS focusing on BRCA1 and BRCA2 mutation carriers was performed in 2010 and modeled genotype data from women diagnosed with breast cancer under the age of 40 and unaffected women over the age of 35 using a retrospective likelihood approach (40). Multiple genome-wide significant SNPs associated with two independent signals at 19p13 were identified among BRCA1 but not BRCA2 mutation carriers. These SNPs were also associated with TNBC in a subsequent analysis (41). These results were later replicated by Couch and colleagues in a study investigating associations between SNPs at 19p13.1 and ZNF365 and both breast and ovarian cancer (42). SNPs at 19p13.1 were associated with ER-negative breast cancer for BRCA1 and BRCA2 mutation carriers and increased risk of ovarian cancer. Additionally, the SNP of interest in ZNF365 was associated with ER-positive breast cancer for both BRCA1 and BRCA2 mutation carriers.
As genetic susceptibility to breast cancer in BRCA1 and BRCA2 mutation carriers unfolded, it became evident that breast cancer risk for BRCA1 and BRCA2 mutation carriers may be influenced by different loci. SNPs associated with overall breast cancer risk in the general population were also associated with breast cancer risk in BRCA2 mutation carriers, whereas SNPs associated with ER-negative breast cancer in the general population were associated with breast cancer risk in BRCA1 mutation carriers. These findings are consistent with the fact that BRCA2 mutation carriers are substantially more likely to have ER-positive disease than BRCA1 mutation carriers (BRCA2 ER-positive OR=11.4; 95% CI: 9.8–13.2) (43).
Similar to the general population, CIMBA identified stronger associations with ER-positive breast cancer for eleven out of the first twelve known susceptibility loci (the exception was rs2046210, near ESR1) in 11,421 BRCA1 and 7,080 BRCA2 mutation carriers (44). In BRCA1 mutation carriers, there were significantly stronger associations with ER-positive cancer risk than ER-negative cancer risk for SNPs in FGFR2 and SLC4A7/NEK10. In addition, a SNP in TOX3/TNRC9 was associated with ER-positive disease, but not ER-negative disease, and a SNP in LSP1 was associated with ER-negative but not ER-positive disease. For BRCA2 carriers, one SNP near ESR1 was significantly different by ER-status. In 2014, Kuchenbaecker and colleagues evaluated 74 known breast cancer susceptibility loci in 15,252 BRCA1 mutation carriers and 8,211 BRCA2 mutation carriers using the iCOGS array (45). This study confirmed genome-wide significant associations for SNPs in three loci previously known to modify BRCA1 breast cancer risk: 19p13, 6q25.1, and 12p11. In addition, four novel SNPs of interest were identified, with a SNP at 1q32 in MDM4 reaching genome-wide significance. Furthermore, 15 and 8 breast cancer susceptibility loci were associated (p<0.05) with risk of ER-negative disease in BRCA1 and BRCA2 mutation carriers, respectively. With respect to ER-positive disease, 10 SNPs were associated (p<0.05) with risk in BRCA1 carriers and 14 SNPs were associated with risk in BRCA2 carriers. More recently, associations with BRCA1 breast cancer and ER-negative disease in the general population from iCOGS and the Oncoarray study have been combined in meta-analyses to increase statistical power to identify SNPs and loci associated with ER-negative disease (10, 25).
Pathological Subtypes
Following identification of breast cancer risk loci in the early GWAS, subsequent studies investigated associations between these risk loci and breast tumor histopathology. In 2008, Garcia-Closas and colleagues found that of the five SNPs previously identified by Easton and colleagues, two risk loci were associated with ER-positive breast cancer, and three were associated with ER-negative breast cancer (12, 14). Both SNPs associated with ER-positive breast cancer were also associated with lower tumor grade, but none of the SNPs were associated with lymph node status following corrections for multiple comparisons (14).
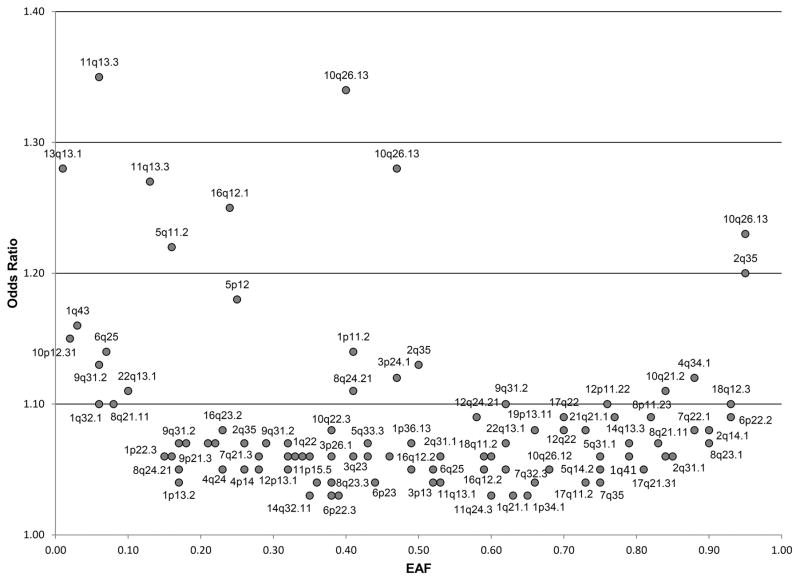
Because the overwhelming majority of breast cancer cases are ER-positive, it is not surprising that most early GWAS studies identified subtype-specific associations predominantly with ER-positive disease. For example, Stacey and colleagues evaluated Icelandic breast cancer cases (n=1,600) and controls (n=11,563) for 311,524 SNPs on the Illumina Infinium HumanHap 300 microarray, and then replicated the top ten hits in an independent dataset (46). Three new loci associated with ER-positive breast cancer were identified (13, 46). To date, many of the breast cancer SNPs associated with overall breast cancer are also associated with ER-positive breast cancer (Supplemental Table 1; Figure 1).
Figure 1.
Allelic architecture of SNPs associated with ER-positive breast cancer
Odds ratio and effect allele frequency (EAF) of the risk-associated SNP for all SNPs associated with ER-positive breast cancer in the OncoArray meta-analysis (p<5 × 10−8) (24)
As a result, several studies specifically focused on ER-negative breast cancer and TNBC were conducted. In 2011, Haiman and colleagues evaluated SNPs in a study including women of both European and African ancestry, and identified the TERT-CLPTM1L locus as a risk factor for both ER-negative breast cancer and TNBC (47). Additionally, four novel SNPs associated with ER-negative breast cancer were identified in a meta-analysis of 3 GWAS of 4,193 ER-negative breast cancer cases and 35,194 controls, with confirmation in the iCOGS dataset using 6,514 ER-negative cases and 41,455 controls (48).
Focusing specifically on risk of triple negative disease, the Triple Negative Breast Cancer Consortium (TNBCC) evaluated 21 variants previously associated with overall breast cancer risk in 2,980 TNBC cases and 4,978 controls in 2011 and found that 4 were strongly associated with TNBC (49). Subsequently, in 2012 SNPs in the 19p13.1 locus were associated with ER-negative and triple negative disease using 48,869 breast cancer cases and 49,787 controls from BCAC (41). In 2014, 25 of 74 known breast cancer susceptibility SNPs were associated with triple negative breast cancer (p<0.05) based on data from 3,677 TNBC cases and 4,708 controls from TNBCC and iCOGS (50). Additionally, Purrington and colleagues identified two SNPs associated with TNBC in ESR1, that were independent of other breast cancer risk associations in this locus (50).
In 2016, Couch and colleagues identified 4 novel loci (two at 13q22) displaying genome wide significant associations with ER-negative breast cancer by pooling data from 11 GWAS and the iCOGS BCAC study (including 12,472 total ER-negative breast cancer cases and 56,820 controls) (10). Two of the novel SNPs (rs67073037 and rs115635831) were not significantly associated with ER-positive breast cancer, whereas rs17181761 and rs6562760 were weakly associated with ER-positive disease. In addition, 15 of 94 previously reported general breast cancer risk SNPs were associated with ER-negative breast cancer at genome-wide significance in this study. Novel SNPs associated with ER-negative breast cancer were also identified in 3 previously discovered breast cancer risk loci. This study also included 7,797 BRCA1 mutation carriers from CIMBA that had been diagnosed with breast cancer and 7,455 that had not yet had cancer. Out of 19 loci that were known to be associated with ER-negative breast cancer in the general population, 15 SNPs showed some association with risk in BRCA1 mutation carriers (p<0.05).
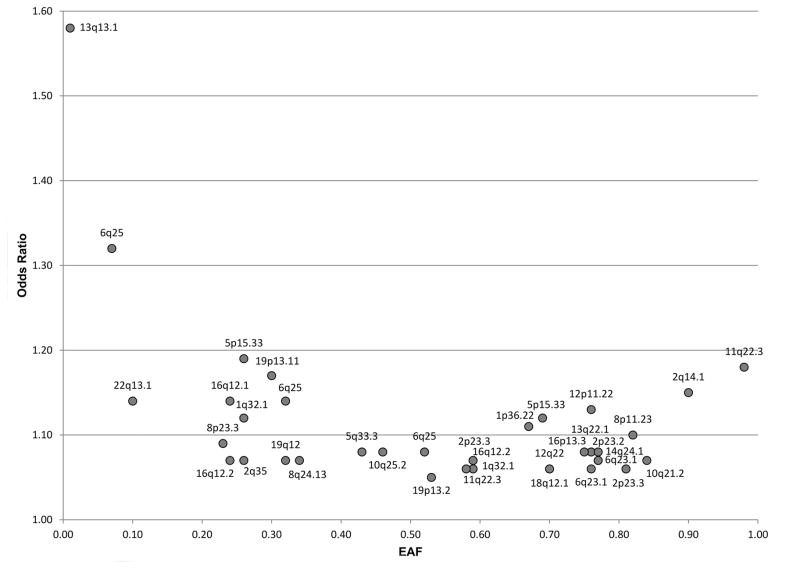
Milne and colleagues recently evaluated ~11.5 million SNPs using imputed OncoArray data from 21,468 ER-negative breast cancer cases and 100,594 controls (25). Of these, 9,655 cases and 45,494 served as an independent replication for 10 of the 11 SNPs previously identified in ER-negative or BRCA1 studies (Supplemental Table 1; Figure 2). In addition, 10 new genome wide significant ER-negative susceptibility SNPs were identified (Supplemental Table 1; Figure 2). Five of these 20 SNPs associated with ER-negative disease were also associated with TNBC; this may be due to decreased statistical power due to a smaller sample of patients with TNBC, or it may be that HER2-driven tumors have unique genetics that have not been well-studied to date. Furthermore, 105 additional SNPs that were previously associated with overall breast cancer risk yielded associations with ER-negative breast cancer (5×10−8<p<0.05), bringing the total number of ER-negative variants to 125 (Supplemental Table 1). These SNPs account for ~14% of familial relative risk for ER-negative breast cancer. Of the SNPs associated with overall breast cancer risk, 24 were also associated with breast cancer risk for BRCA1 mutation carriers (25).
Figure 2.
Allelic architecture of SNPs associated with ER-negative breast cancer
Odds ratio and effect allele frequency (EAF) of the risk-associated SNP for all SNPs associated with ER-negative breast cancer in the OncoArray meta-analysis (p<5 × 10−8) (24, 25)
Race/Ethnicity
LD structures in populations
The majority of SNPs associated with breast cancer risk have been discovered in populations of European descent, presenting an etiological and clinical challenge for non-European populations. There are known differences in LD structures between Europeans and other populations, which may influence risks associated with breast cancer for selected SNPs (51). Additionally, women of African descent have an increased incidence of ER-negative breast cancer, which may be partially explained by genetic risk factors (52).
LD structures influencing GWAS results in breast cancer were first reported by Stacey and colleagues in 2008 when utilizing the Multiethnic Cohort to replicate variants associated with breast cancer risk (13). Although power was limited by small sample sizes, certain SNPs that were associated with breast cancer risk in Europeans and Latinas (rs13387042 and rs3803662) were not significantly associated with increased breast cancer risk in Hawaiians, Japanese, or African Americans. In fact, one risk SNP in Europeans (rs3803662) displayed a protective effect in African Americans, suggesting that it was in LD with a different causative SNP (13).
African American
Discovering risk loci underlying breast cancer etiology in African American women may allow investigators to further understand and address health disparities in African American women with breast cancer (53). Several studies investigating associations between SNPS in known breast cancer risk loci, candidate genes and pathways with breast cancer risk in the African American population have been conducted (54–66). For example, in 2013, Long and colleagues evaluated 67 known breast cancer susceptibility SNPs in 1231 cases and 2069 controls, and found that 7 SNPs were significantly associated (p<0.05) with breast cancer risk in African American women, 3 were borderline significant (p<0.10), and 57 did not show association (60).
There have also been several GWAS for breast cancer in African American women. In 2011, a GWAS by Haiman and colleagues included women of African American ancestry and women of European ancestry resulting in the identification of the TERT-CLPTM1L loci (47). To increase sample size and power, three consortia have been formed to study breast cancer in women of African ancestry: African American Breast Cancer (AABC; n=3,153 cases), GWAS in Breast Cancer in the African Diaspora (ROOT; n=1,657 cases), and African American Breast Cancer Epidemiology and Risk (AMBER; n~6,000 cases) (63, 66, 67). Two epidemiological studies, Multi-Ethnic Cohort (MEC) and Women’s Circle of Health Study (WCHS) contributed samples to both AABC and AMBER.
GWAS have been performed by both AABC and ROOT. However, no SNPs yielded genome-wide significant associations with breast cancer (66, 68). The AMBER consortium performed a two-stage meta-analysis of breast cancer GWAS in women of African descent using discovery phase data from 4,673 breast cancer cases and 4,774 controls from the ROOT and AABC consortia followed by replication in the AMBER consortium. Three SNPs, one of which was novel (rs13074711 in 3q26.21) were associated with overall breast cancer risk (69). In subsequent analyses, 2 of the SNPs were associated with ER-negative disease and one with ER-positive disease (69). A separate study of AMBER followed by replication in an independent case-control study identified two novel loci associated with ER-positive cancer in women of African ancestry (14p16.1 and 17q25.1) (70). The study also identified two loci (10q26 and 11q13) previously associated with breast cancer risk that have an excess of African ancestry. Subsequent fine-mapping studies suggested that 11q13 is associated with ER-positive breast cancer in women of African ancestry (70).
Asian populations
The breast cancer GWAS studies from COGS and OncoArray have included women of European and Asian ancestry (22–24). There have also been several independent GWAS studies in Asian women. In 2009, a GWAS of 1,505 cases and 1,522 controls from the Shanghai Breast Cancer Study was performed (17). After two further stages of replication of 29 and 4 SNPs, respectively, a SNP upstream of ESR1 was associated with breast cancer risk. The association was stronger in post-menopausal women and women with ER-negative breast cancer. These findings were subsequently confirmed in a population of European descent and provided the first glimpses into ESR1 as a complex locus (17). Studies of 23,637 East Asian breast cancer cases and 25,579 East Asian controls using the iCOGs array found that 31 of the 67 breast cancer loci at that time also conferred an increased risk of breast cancer in East Asian women (71). Similar studies of 22,780 breast cancer cases and 24,181 controls from the East Asian population identified three novel breast cancer loci (1q32.1, 5q14.3, and 15q26.1) at genome-wide significance that replicated in a European-ancestry population (72).
Putting it all together—Polygenic Risk Scores
Individually, SNPs have quite small effect sizes and may not be informative for evaluating risk of developing breast cancer. However, the recent OncoArray study suggested that all of the breast cancer susceptibility markers to date explain up to 18 percent of familial relative risk (24). Thus, combining these common variants together may provide some insight into individual risks of breast cancer. While the first 7 breast cancer susceptibility SNPs were shown to have little influence on risks predicted by the National Cancer Institute Breast Cancer Risk Assessment Tool (BCRAT or Gail model), subsequent studies using polygenic risk scores (PRS) based on larger numbers of SNPs have successfully demonstrated an ability to stratify or individualize breast cancer risk in a number of populations (73).
In 2015, BCAC investigators generated a PRS using 77 known breast cancer susceptibility SNPs that improved risk estimates for women in the general population (74). Specifically, lifetime risk for a woman without a family history of breast cancer in the highest quintile of the PRS was 16.6%. However, women with a first-degree family history of breast cancer and in the highest quintile had a lifetime risk of 24.4%. Separately, Vachon and colleagues showed that a 76 SNP PRS was independent from breast density as a risk factor and enhanced the breast cancer risk prediction by the Breast Cancer Surveillance Consortium breast cancer risk model (75). With the recent discovery of an additional 75 breast cancer susceptibility loci, it is anticipated that an updated PRS including all 182 breast cancer SNPs will likely stratify risk even further (24, 25). However, understanding how the PRS influences other risk factors in these models is needed. Recent analyses examining the combined effects of a 77-SNP PRS and risk factors on breast cancer risk in up to 23,000 cases showed the combined effects of the 77-SNP PRS and environmental risk factors for breast cancer are generally well described by a multiplicative model. However, there was some evidence for departure from the multiplicative model for alcohol, current menopausal hormone therapy and height with ER-positive disease that warrants further study (76).
PRS have also been designed to modify the estimated breast cancer risk of women with mutations in cancer predisposition genes, such as BRCA1, BRCA2, and CHEK2 c.1100delC. CHEK2 c.1100delC is found in up to 1.5 percent of European individuals and is known to confer a moderate risk of breast cancer (odds ratio (OR)=2.0–3.0) (7, 77, 78). Muranen and colleagues applied the PRS based on 77 breast cancer risk SNPs in the BCAC population with consideration for the effect of the PRS on CHEK2*1100delC carriers (79). For the highest quintile of the PRS, the OR for carriers was 2.03 (0.86–4.78), while the OR for the lowest quintile was much lower and more comparable to lifetime risk expected in the general population (OR=0.52, 95% CI: 0.16–1.74). In the future, PRS may be developed to refine breast cancer risk estimates for mutation carriers in other high and moderate breast cancer risk genes such as PALB2 and ATM.
In 2017, Kuchenbaecker and colleagues evaluated 94 breast cancer risk SNPs in a PRS to stratify risk for BRCA1 and BRCA2 mutation carriers (80). PRS were generated for overall breast cancer risk as well as ER-negative and ER-positive breast cancer risk. All three PRS were associated with breast cancer risk for BRCA1 and BRCA2 carriers. For BRCA1 carriers, the ER-negative PRS showed the strongest association (HR=1.27; 95% CI=1.23–1.31). Conversely, stronger associations were observed for the overall and ER-positive PRS in BRCA2 mutation carriers (HR=1.22; 95% CI=1.17–1.28). Stratification or personalization of breast cancer risk is particularly important for women with BRCA1 and BRCA2 mutations because of high age-related and lifetime risks of disease. Identification of mutation carriers at particularly high risk or with reduced risk relative to the average mutation carrier may lead to changes in risk management including decisions about uptake and timing of prophylactic mastectomy and prophylactic oophorectomy, and decisions about child-bearing.
In 2016, Dite and colleagues evaluated models combining a PRS based on 77 breast cancer susceptibility SNPs and clinical breast cancer risk prediction models including BOADICEA, BRCAPRO, BCRAT, and IBIS (81). This study suggested that including the PRS improved breast cancer prediction in women younger than 50 years by more than 20 percent. Beginning in August 2017, Myriad Genetics, a commercial genetic testing laboratory, incorporated 82 SNPs on the MyRisk multi-gene panel. The resulting Breast Cancer riskScore that includes results from the 82 SNPs and the Tyrer-Cuzick model predicts a woman’s 5-year and lifetime risk of developing breast cancer (82).
PRS may also impact clinical care by informing screening guidelines, particularly in women ages 40–49, for whom professional societies often have discordant recommendations. The Women Informed to Screen Depending On Measure of risk (WISDOM) population-based study is investigating breast cancer screening modalities in the precision medicine era (83). This study will evaluate risk thresholds and screening recommendations by integrating traditional clinical models, mutations in high and intermediate penetrance genes, and PRS. In addition, WISDOM plans to investigate prediction of breast cancer subtype-specific risk, which may result in more frequent screening for women at a higher risk of more aggressive breast cancers (e.g. ER-negative disease) (83).
Breast Cancer Gene-Environment interactions/associations with other risk factors
There are many well established factors that influence risk for breast cancer including breast density, parity, breastfeeding, alcohol use, body mass index (BMI), exercise, hormone replacement therapy (HRT), and menstrual history. For hereditary traits that influence breast cancer risk, such as breast density and menstrual history, there may be overlap between their predisposition loci and breast cancer risk loci. In fact, using GWAS of breast density and breast cancer, both traits show evidence of a shared genetic basis that is mediated through a large number of common variants (84). Further, 18 percent of known breast cancer susceptibility variants have been associated with mammographic density (85). Lindstrom and colleagues performed a two-stage GWAS investigating SNPs associated with mammographic density and found that four previously identified breast cancer susceptibility SNPs are also associated with mammographic density (rs12665607 in ESR1, rs10995190 in SNF365, rs3817198 in LSP1, and rs17001868 in SGSM2/MKL1) (86). Four additional SNPs (rs186749 in PRDM6, rs7816345 in 8p11.23, rs703556 in IGF1, and rs7289126 in TMEM184B) found to be associated with mammographic density were later shown to be associated with breast cancer risk (86). For non-hereditary traits, gene-environment interactions may be useful in assessing risk associated with breast cancer risk SNPs. However, sufficient power is frequently an obstacle in identifying gene-environment interactions (87). Following correction for multiple testing, five SNPs have been shown to have an interaction with breast cancer risk and a risk factor: two SNPs have been associated with parity (rs3817198 in LSP1 and rs11249422 in 1p11.2), one SNP with alcohol use (rs1045485 in CASP8), and two SNPs in strong LD with each other were associated with post-menopausal BMI (rs10483028 and rs2242714 in 21q22.12) (88–90). Additional gene-environment interactions may be identified in the OncoArray study due to larger sample sizes.
Understanding the biology
Most of the breast cancer susceptibility SNPs reside in intergenic or intronic regions of the genome, and counter to early expectations, do not generally impact protein-coding regions. Many follow-up studies have been conducted to identify causal variants, defined by Edwards and colleagues as variants that influence a molecular or cellular process to affect a human phenotype (91). There are many approaches to evaluation of GWAS findings including fine mapping by genotyping of additional SNPs in a locus, in silico annotations, SNP function, target gene identification, and target gene function.
Fine mapping studies often require more power than the original GWAS in order to identify small differences in risk that are associated with different SNPs. Because fine mapping studies require information on each SNP in the locus of interest, the advent of imputation techniques has alleviated the burdens of genotyping every single SNP. Several fine-mapping studies have been performed to identify causal breast cancer SNPs, revealing complex loci that contain more than one independent variant contributing to risk. One example of this is an investigation of the TERT locus that identified three independent SNP associations (92). ESR1 is another example of a complex risk locus; there have been five independent SNPs identified in ESR1 that have different associations with breast cancer according to subtype and grade (93).
One way to identify SNP function and a potential target gene is to evaluate expression quantitative trait loci (eQTLs) to determine if a SNP is impacting gene expression. After utilizing fine mapping to identify multiple independent signals in the 5p12 locus, Ghoussaini and colleagues performed an eQTL analysis that revealed a SNP associated with increased expression of two candidate genes for breast cancer pathogenesis (94). However, eQTL studies can be limited by the normal tissue that is available. While data for many normal tissues are now available from the GTEx portal, the numbers of samples analyzed limit the statistical power of eQTL analyses based on these data (95). More recently, transcriptome-wide association studies (TWAS) have been used to analyze both SNPs and gene expression to identify novel breast cancer susceptibility genes and loci. By pairing genotyping data with gene expression data, SNPs that can predict gene expression can be selected. Subsequent imputation of gene expression data can be performed using these selected SNPs in another dataset. Finally, the imputed gene expression data can be evaluated for associations with breast cancer risk. Importantly, this method reduces the correction needed for multiple comparisons, which may increase power. Additionally, findings from these studies may have clearer functional implications than traditional GWAS, and combining SNPs to predict gene expression may help to enhance signals that were too weak to detect previously. Using TWAS, Hoffman and colleagues identified five genes and twenty-three unique SNPs that may be associated with breast cancer risk (96).
Causal SNPs in regulatory regions may impact breast cancer risk in complex ways. One example is the 11q13 breast cancer risk locus, which has been associated with ER-positive disease. Fine mapping studies identified three independent signals in 11q13 (97). Subsequent functional assays including electrophorectic mobility shift assay (EMSA), promoter and enhancer luciferase reporter assays, and chromatin immunoprecipitation studies implicated these SNPs in distal, long-distance regulation of the CCND1 promoter (97). This distal enhancer on 11q13 has also been implicated in regulation of two estrogen-modulated long noncoding RNAs, suggesting that the underlying risk-driving SNPs in this locus may cause preferential chromatin looping further impacting promoter activity and transcription (98). Importantly, CRISPR/cas9 based allele-specific deletion and/or replacement is rapidly becoming the preferred method for evaluation of effects of single SNPs or combinations of SNPs on the biology of risk associated loci.
Conclusions
To date, 182 breast cancer susceptibility SNPs have been identified for breast cancer, increasing understanding of the heritability of breast cancer. The combined effect of these SNPs in PRS models may pave the way to tailored screening and intervention strategies in the general population as well as in carriers of deleterious mutations in predisposition genes (e.g., BRCA1 and BRCA2). Further functional studies of risk loci will elucidate biological mechanisms of breast cancer etiology, with potential implications for breast cancer prevention, prognosis, and treatment. New methodologies, such as TWAS, are emerging to inform functional investigations of known breast cancer SNPs as well as identify novel susceptibility breast cancer SNPs and genes. Additionally, with the growing size of consortia designed to study the genetic component of breast cancer, we may see even larger GWAS in the future that will have power to detect smaller effect sizes.
Supplementary Material
Acknowledgments
K.J. Ruddy was supported by a training grant under the CTSA Grant Program Numbers UL1 TR000135 and KL2TR000136-09 from the National Center for Advancing Translational Sciences (NCATS) of the NIH. J. Lilyquist, C.M. Vachon, and F.J. Couch were supported by a NIH training grant R25CA92049 (Mayo Cancer Genetic Epidemiology Training program). F.J. Couch was supported by Breast Cancer Research Foundation, NIH grants R01 CA192393, CA116167, CA176785, the Mayo Clinic Specialized Program of Research Excellence (SPORE) in Breast Cancer (CA116201), and the Mayo Clinic Breast Cancer Registry. K.J. Ruddy, C.M. Vachon, and F.J. Couch were also supported by the Mayo Clinic Cancer Center. The contents of this paper are solely the responsibility of the authors and do not necessarily represent the official view of NIH.
Footnotes
Conflict of Interest: Authors have no conflicts to disclose.
References
- 1.Peto J, Mack TM. High constant incidence in twins and other relatives of women with breast cancer. Nat Genet. 2000;26(4):411–4. doi: 10.1038/82533. [DOI] [PubMed] [Google Scholar]
- 2.Colditz GA, Willett WC, Hunter DJ, Stampfer MJ, Manson JE, Hennekens CH, et al. Family history, age, and risk of breast cancer. Prospective data from the Nurses’ Health Study. JAMA. 1993;270(3):338–43. [PubMed] [Google Scholar]
- 3.Miki Y, Swensen J, Shattuck-Eidens D, Futreal PA, Harshman K, Tavtigian S, et al. A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science. 1994;266(5182):66–71. doi: 10.1126/science.7545954. [DOI] [PubMed] [Google Scholar]
- 4.Wooster R, Bignell G, Lancaster J, Swift S, Seal S, Mangion J, et al. Identification of the breast cancer susceptibility gene BRCA2. Nature. 1995;378(6559):789–92. doi: 10.1038/378789a0. [DOI] [PubMed] [Google Scholar]
- 5.Rahman N, Seal S, Thompson D, Kelly P, Renwick A, Elliott A, et al. PALB2, which encodes a BRCA2-interacting protein, is a breast cancer susceptibility gene. Nat Genet. 2007;39(2):165–7. doi: 10.1038/ng1959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Renwick A, Thompson D, Seal S, Kelly P, Chagtai T, Ahmed M, et al. ATM mutations that cause ataxia-telangiectasia are breast cancer susceptibility alleles. Nat Genet. 2006;38(8):873–5. doi: 10.1038/ng1837. [DOI] [PubMed] [Google Scholar]
- 7.Meijers-Heijboer H, van den Ouweland A, Klijn J, Wasielewski M, de Snoo A, Oldenburg R, et al. Low-penetrance susceptibility to breast cancer due to CHEK2(*)1100delC in noncarriers of BRCA1 or BRCA2 mutations. Nat Genet. 2002;31(1):55–9. doi: 10.1038/ng879. [DOI] [PubMed] [Google Scholar]
- 8.Breast Cancer Association C. Commonly studied single-nucleotide polymorphisms and breast cancer: results from the Breast Cancer Association Consortium. J Natl Cancer Inst. 2006;98(19):1382–96. doi: 10.1093/jnci/djj374. [DOI] [PubMed] [Google Scholar]
- 9.Cox A, Dunning AM, Garcia-Closas M, Balasubramanian S, Reed MW, Pooley KA, et al. A common coding variant in CASP8 is associated with breast cancer risk. Nat Genet. 2007;39(3):352–8. doi: 10.1038/ng1981. [DOI] [PubMed] [Google Scholar]
- 10.Couch FJ, Kuchenbaecker KB, Michailidou K, Mendoza-Fandino GA, Nord S, Lilyquist J, et al. Identification of four novel susceptibility loci for oestrogen receptor negative breast cancer. Nat Commun. 2016;7:11375. doi: 10.1038/ncomms11375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hunter DJ, Kraft P, Jacobs KB, Cox DG, Yeager M, Hankinson SE, et al. A genome-wide association study identifies alleles in FGFR2 associated with risk of sporadic postmenopausal breast cancer. Nat Genet. 2007;39(7):870–4. doi: 10.1038/ng2075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Easton DF, Pooley KA, Dunning AM, Pharoah PD, Thompson D, Ballinger DG, et al. Genome-wide association study identifies novel breast cancer susceptibility loci. Nature. 2007;447(7148):1087–93. doi: 10.1038/nature05887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Stacey SN, Manolescu A, Sulem P, Thorlacius S, Gudjonsson SA, Jonsson GF, et al. Common variants on chromosome 5p12 confer susceptibility to estrogen receptor-positive breast cancer. Nat Genet. 2008;40(6):703–6. doi: 10.1038/ng.131. [DOI] [PubMed] [Google Scholar]
- 14.Garcia-Closas M, Hall P, Nevanlinna H, Pooley K, Morrison J, Richesson DA, et al. Heterogeneity of breast cancer associations with five susceptibility loci by clinical and pathological characteristics. PLoS Genet. 2008;4(4):e1000054. doi: 10.1371/journal.pgen.1000054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Antoniou AC, Spurdle AB, Sinilnikova OM, Healey S, Pooley KA, Schmutzler RK, et al. Common breast cancer-predisposition alleles are associated with breast cancer risk in BRCA1 and BRCA2 mutation carriers. Am J Hum Genet. 2008;82(4):937–48. doi: 10.1016/j.ajhg.2008.02.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ahmed S, Thomas G, Ghoussaini M, Healey CS, Humphreys MK, Platte R, et al. Newly discovered breast cancer susceptibility loci on 3p24 and 17q23.2. Nat Genet. 2009;41(5):585–90. doi: 10.1038/ng.354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zheng W, Long J, Gao YT, Li C, Zheng Y, Xiang YB, et al. Genome-wide association study identifies a new breast cancer susceptibility locus at 6q25.1. Nat Genet. 2009;41(3):324–8. doi: 10.1038/ng.318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Mavaddat N, Antoniou AC, Easton DF, Garcia-Closas M. Genetic susceptibility to breast cancer. Mol Oncol. 2010;4(3):174–91. doi: 10.1016/j.molonc.2010.04.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ghoussaini M, Fletcher O, Michailidou K, Turnbull C, Schmidt MK, Dicks E, et al. Genome-wide association analysis identifies three new breast cancer susceptibility loci. Nat Genet. 2012;44(3):312–8. doi: 10.1038/ng.1049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Turnbull C, Ahmed S, Morrison J, Pernet D, Renwick A, Maranian M, et al. Genome-wide association study identifies five new breast cancer susceptibility loci. Nat Genet. 2010;42(6):504–7. doi: 10.1038/ng.586. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Fletcher O, Johnson N, Orr N, Hosking FJ, Gibson LJ, Walker K, et al. Novel breast cancer susceptibility locus at 9q31.2: results of a genome-wide association study. J Natl Cancer Inst. 2011;103(5):425–35. doi: 10.1093/jnci/djq563. [DOI] [PubMed] [Google Scholar]
- 22.Michailidou K, Hall P, Gonzalez-Neira A, Ghoussaini M, Dennis J, Milne RL, et al. Large-scale genotyping identifies 41 new loci associated with breast cancer risk. Nat Genet. 2013;45(4):353–61. 61e1–2. doi: 10.1038/ng.2563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Michailidou K, Beesley J, Lindstrom S, Canisius S, Dennis J, Lush MJ, et al. Genome-wide association analysis of more than 120,000 individuals identifies 15 new susceptibility loci for breast cancer. Nat Genet. 2015;47(4):373–80. doi: 10.1038/ng.3242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Michailidou K, Lindstrom S, Dennis J, Beesley J, Hui S, Kar S, et al. Association analysis identifies 65 new breast cancer risk loci. Nature. 2017 doi: 10.1038/nature24284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Milne RL, Kuchenbaecker KB, Michailidou K, Beesley J, Kar S, Lindstrom S, et al. Identification of ten variants associated with risk of estrogen-receptor-negative breast cancer. Nat Genet. 2017 doi: 10.1038/ng.3785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Chenevix-Trench G, Milne RL, Antoniou AC, Couch FJ, Easton DF, Goldgar DE, et al. An international initiative to identify genetic modifiers of cancer risk in BRCA1 and BRCA2 mutation carriers: the Consortium of Investigators of Modifiers of BRCA1 and BRCA2 (CIMBA) Breast Cancer Res. 2007;9(2):104. doi: 10.1186/bcr1670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Antoniou AC, Sinilnikova OM, Simard J, Leone M, Dumont M, Neuhausen SL, et al. RAD51 135G-->C modifies breast cancer risk among BRCA2 mutation carriers: results from a combined analysis of 19 studies. Am J Hum Genet. 2007;81(6):1186–200. doi: 10.1086/522611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Antoniou AC, Sinilnikova OM, McGuffog L, Healey S, Nevanlinna H, Heikkinen T, et al. Common variants in LSP1, 2q35 and 8q24 and breast cancer risk for BRCA1 and BRCA2 mutation carriers. Hum Mol Genet. 2009;18(22):4442–56. doi: 10.1093/hmg/ddp372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Antoniou AC, Kartsonaki C, Sinilnikova OM, Soucy P, McGuffog L, Healey S, et al. Common alleles at 6q25.1 and 1p11.2 are associated with breast cancer risk for BRCA1 and BRCA2 mutation carriers. Hum Mol Genet. 2011;20(16):3304–21. doi: 10.1093/hmg/ddr226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Antoniou AC, Kuchenbaecker KB, Soucy P, Beesley J, Chen X, McGuffog L, et al. Common variants at 12p11, 12q24, 9p21, 9q31.2 and in ZNF365 are associated with breast cancer risk for BRCA1 and/or BRCA2 mutation carriers. Breast Cancer Res. 2012;14(1):R33. doi: 10.1186/bcr3121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Osorio A, Pollan M, Pita G, Schmutzler RK, Versmold B, Engel C, et al. An evaluation of the polymorphisms Ins16bp and Arg72Pro in p53 as breast cancer risk modifiers in BRCA1 and BRCA2 mutation carriers. Br J Cancer. 2008;99(6):974–7. doi: 10.1038/sj.bjc.6604624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sinilnikova OM, Antoniou AC, Simard J, Healey S, Leone M, Sinnett D, et al. The TP53 Arg72Pro and MDM2 309G>T polymorphisms are not associated with breast cancer risk in BRCA1 and BRCA2 mutation carriers. Br J Cancer. 2009;101(8):1456–60. doi: 10.1038/sj.bjc.6605279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Osorio A, Milne RL, Pita G, Peterlongo P, Heikkinen T, Simard J, et al. Evaluation of a candidate breast cancer associated SNP in ERCC4 as a risk modifier in BRCA1 and BRCA2 mutation carriers. Results from the Consortium of Investigators of Modifiers of BRCA1/BRCA2 (CIMBA) Br J Cancer. 2009;101(12):2048–54. doi: 10.1038/sj.bjc.6605416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Engel C, Versmold B, Wappenschmidt B, Simard J, Easton DF, Peock S, et al. Association of the variants CASP8 D302H and CASP10 V410I with breast and ovarian cancer risk in BRCA1 and BRCA2 mutation carriers. Cancer Epidemiol Biomarkers Prev. 2010;19(11):2859–68. doi: 10.1158/1055-9965.EPI-10-0517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Osorio A, Milne RL, Alonso R, Pita G, Peterlongo P, Teule A, et al. Evaluation of the XRCC1 gene as a phenotypic modifier in BRCA1/2 mutation carriers. Results from the consortium of investigators of modifiers of BRCA1/BRCA2. Br J Cancer. 2011;104(8):1356–61. doi: 10.1038/bjc.2011.91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ding YC, McGuffog L, Healey S, Friedman E, Laitman Y, Paluch-Shimon S, et al. A nonsynonymous polymorphism in IRS1 modifies risk of developing breast and ovarian cancers in BRCA1 and ovarian cancer in BRCA2 mutation carriers. Cancer Epidemiol Biomarkers Prev. 2012;21(8):1362–70. doi: 10.1158/1055-9965.EPI-12-0229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wang X, Pankratz VS, Fredericksen Z, Tarrell R, Karaus M, McGuffog L, et al. Common variants associated with breast cancer in genome-wide association studies are modifiers of breast cancer risk in BRCA1 and BRCA2 mutation carriers. Hum Mol Genet. 2010;19(14):2886–97. doi: 10.1093/hmg/ddq174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Stevens KN, Wang X, Fredericksen Z, Pankratz VS, Greene MH, Andrulis IL, et al. Evaluation of chromosome 6p22 as a breast cancer risk modifier locus in a follow-up study of BRCA2 mutation carriers. Breast Cancer Res Treat. 2012;136(1):295–302. doi: 10.1007/s10549-012-2255-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Osorio A, Milne RL, Kuchenbaecker K, Vaclova T, Pita G, Alonso R, et al. DNA glycosylases involved in base excision repair may be associated with cancer risk in BRCA1 and BRCA2 mutation carriers. PLoS Genet. 2014;10(4):e1004256. doi: 10.1371/journal.pgen.1004256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Antoniou AC, Wang X, Fredericksen ZS, McGuffog L, Tarrell R, Sinilnikova OM, et al. A locus on 19p13 modifies risk of breast cancer in BRCA1 mutation carriers and is associated with hormone receptor-negative breast cancer in the general population. Nat Genet. 2010;42(10):885–92. doi: 10.1038/ng.669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Stevens KN, Fredericksen Z, Vachon CM, Wang X, Margolin S, Lindblom A, et al. 19p13.1 is a triple-negative-specific breast cancer susceptibility locus. Cancer Res. 2012;72(7):1795–803. doi: 10.1158/0008-5472.CAN-11-3364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Couch FJ, Wang X, McGuffog L, Lee A, Olswold C, Kuchenbaecker KB, et al. Genome-wide association study in BRCA1 mutation carriers identifies novel loci associated with breast and ovarian cancer risk. PLoS Genet. 2013;9(3):e1003212. doi: 10.1371/journal.pgen.1003212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Mavaddat N, Barrowdale D, Andrulis IL, Domchek SM, Eccles D, Nevanlinna H, et al. Pathology of breast and ovarian cancers among BRCA1 and BRCA2 mutation carriers: results from the Consortium of Investigators of Modifiers of BRCA1/2 (CIMBA) Cancer Epidemiol Biomarkers Prev. 2012;21(1):134–47. doi: 10.1158/1055-9965.EPI-11-0775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Mulligan AM, Couch FJ, Barrowdale D, Domchek SM, Eccles D, Nevanlinna H, et al. Common breast cancer susceptibility alleles are associated with tumour subtypes in BRCA1 and BRCA2 mutation carriers: results from the Consortium of Investigators of Modifiers of BRCA1/2. Breast Cancer Res. 2011;13(6):R110. doi: 10.1186/bcr3052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Kuchenbaecker KB, Neuhausen SL, Robson M, Barrowdale D, McGuffog L, Mulligan AM, et al. Associations of common breast cancer susceptibility alleles with risk of breast cancer subtypes in BRCA1 and BRCA2 mutation carriers. Breast Cancer Res. 2014;16(6):3416. doi: 10.1186/s13058-014-0492-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Stacey SN, Manolescu A, Sulem P, Rafnar T, Gudmundsson J, Gudjonsson SA, et al. Common variants on chromosomes 2q35 and 16q12 confer susceptibility to estrogen receptor-positive breast cancer. Nat Genet. 2007;39(7):865–9. doi: 10.1038/ng2064. [DOI] [PubMed] [Google Scholar]
- 47.Haiman CA, Chen GK, Vachon CM, Canzian F, Dunning A, Millikan RC, et al. A common variant at the TERT-CLPTM1L locus is associated with estrogen receptor-negative breast cancer. Nat Genet. 2011;43(12):1210–4. doi: 10.1038/ng.985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Garcia-Closas M, Couch FJ, Lindstrom S, Michailidou K, Schmidt MK, Brook MN, et al. Genome-wide association studies identify four ER negative-specific breast cancer risk loci. Nat Genet. 2013;45(4):392–8. 8e1–2. doi: 10.1038/ng.2561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Stevens KN, Vachon CM, Lee AM, Slager S, Lesnick T, Olswold C, et al. Common breast cancer susceptibility loci are associated with triple-negative breast cancer. Cancer Res. 2011;71(19):6240–9. doi: 10.1158/0008-5472.CAN-11-1266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Purrington KS, Slager S, Eccles D, Yannoukakos D, Fasching PA, Miron P, et al. Genome-wide association study identifies 25 known breast cancer susceptibility loci as risk factors for triple-negative breast cancer. Carcinogenesis. 2014;35(5):1012–9. doi: 10.1093/carcin/bgt404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Pfaff CL, Parra EJ, Bonilla C, Hiester K, McKeigue PM, Kamboh MI, et al. Population structure in admixed populations: effect of admixture dynamics on the pattern of linkage disequilibrium. Am J Hum Genet. 2001;68(1):198–207. doi: 10.1086/316935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Chu KC, Anderson WF. Rates for breast cancer characteristics by estrogen and progesterone receptor status in the major racial/ethnic groups. Breast Cancer Res Treat. 2002;74(3):199–211. doi: 10.1023/a:1016361932220. [DOI] [PubMed] [Google Scholar]
- 53.Newman LA, Kaljee LM. Health Disparities and Triple-Negative Breast Cancer in African American Women: A Review. JAMA Surg. 2017;152(5):485–93. doi: 10.1001/jamasurg.2017.0005. [DOI] [PubMed] [Google Scholar]
- 54.Murphy ME, Liu S, Yao S, Huo D, Liu Q, Dolfi SC, et al. A functionally significant SNP in TP53 and breast cancer risk in African-American women. NPJ Breast Cancer. 2017;3:5. doi: 10.1038/s41523-017-0007-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Ruiz-Narvaez EA, Rosenberg L, Yao S, Rotimi CN, Cupples AL, Bandera EV, et al. Fine-mapping of the 6q25 locus identifies a novel SNP associated with breast cancer risk in African-American women. Carcinogenesis. 2013;34(2):287–91. doi: 10.1093/carcin/bgs334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Ruiz-Narvaez EA, Lunetta KL, Hong CC, Haddad S, Yao S, Cheng TD, et al. Genetic variation in the insulin, insulin-like growth factor, growth hormone, and leptin pathways in relation to breast cancer in African-American women: the AMBER consortium. NPJ Breast Cancer. 2016:2. doi: 10.1038/npjbcancer.2016.34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Ruiz-Narvaez EA, Haddad SA, Lunetta KL, Yao S, Bensen JT, Sucheston-Campbell LE, et al. Gene-based analysis of the fibroblast growth factor receptor signaling pathway in relation to breast cancer in African American women: the AMBER consortium. Breast Cancer Res Treat. 2016;155(2):355–63. doi: 10.1007/s10549-015-3672-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Yao S, Haddad SA, Hu Q, Liu S, Lunetta KL, Ruiz-Narvaez EA, et al. Genetic variations in vitamin D-related pathways and breast cancer risk in African American women in the AMBER consortium. Int J Cancer. 2016;138(9):2118–26. doi: 10.1002/ijc.29954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Feng Y, Stram DO, Rhie SK, Millikan RC, Ambrosone CB, John EM, et al. A comprehensive examination of breast cancer risk loci in African American women. Hum Mol Genet. 2014;23(20):5518–26. doi: 10.1093/hmg/ddu252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Long J, Zhang B, Signorello LB, Cai Q, Deming-Halverson S, Shrubsole MJ, et al. Evaluating genome-wide association study-identified breast cancer risk variants in African-American women. PLoS One. 2013;8(4):e58350. doi: 10.1371/journal.pone.0058350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Song C, Chen GK, Millikan RC, Ambrosone CB, John EM, Bernstein L, et al. A genome-wide scan for breast cancer risk haplotypes among African American women. PLoS One. 2013;8(2):e57298. doi: 10.1371/journal.pone.0057298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Palmer JR, Ruiz-Narvaez EA, Rotimi CN, Cupples LA, Cozier YC, Adams-Campbell LL, et al. Genetic susceptibility loci for subtypes of breast cancer in an African American population. Cancer Epidemiol Biomarkers Prev. 2013;22(1):127–34. doi: 10.1158/1055-9965.EPI-12-0769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Chen F, Chen GK, Millikan RC, John EM, Ambrosone CB, Bernstein L, et al. Fine-mapping of breast cancer susceptibility loci characterizes genetic risk in African Americans. Hum Mol Genet. 2011;20(22):4491–503. doi: 10.1093/hmg/ddr367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Hutter CM, Young AM, Ochs-Balcom HM, Carty CL, Wang T, Chen CT, et al. Replication of breast cancer GWAS susceptibility loci in the Women’s Health Initiative African American SHARe Study. Cancer Epidemiol Biomarkers Prev. 2011;20(9):1950–9. doi: 10.1158/1055-9965.EPI-11-0524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Zhang J, Yao S, Hu Q, Zhu Q, Liu S, Lunetta KL, et al. Genetic variations in the Hippo signaling pathway and breast cancer risk in African American women in the AMBER Consortium. Carcinogenesis. 2016;37(10):951–6. doi: 10.1093/carcin/bgw077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Huo D, Zheng Y, Ogundiran TO, Adebamowo C, Nathanson KL, Domchek SM, et al. Evaluation of 19 susceptibility loci of breast cancer in women of African ancestry. Carcinogenesis. 2012;33(4):835–40. doi: 10.1093/carcin/bgs093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Palmer JR, Ambrosone CB, Olshan AF. A collaborative study of the etiology of breast cancer subtypes in African American women: the AMBER consortium. Cancer Causes Control. 2014;25(3):309–19. doi: 10.1007/s10552-013-0332-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Chen F, Chen GK, Stram DO, Millikan RC, Ambrosone CB, John EM, et al. A genome-wide association study of breast cancer in women of African ancestry. Hum Genet. 2013;132(1):39–48. doi: 10.1007/s00439-012-1214-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Huo D, Feng Y, Haddad S, Zheng Y, Yao S, Han YJ, et al. Genome-wide association studies in women of African ancestry identified 3q26.21 as a novel susceptibility locus for oestrogen receptor negative breast cancer. Hum Mol Genet. 2016;25(21):4835–46. doi: 10.1093/hmg/ddw305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Ruiz-Narvaez EA, Sucheston-Campbell L, Bensen JT, Yao S, Haddad S, Haiman CA, et al. Admixture Mapping of African-American Women in the AMBER Consortium Identifies New Loci for Breast Cancer and Estrogen-Receptor Subtypes. Front Genet. 2016;7:170. doi: 10.3389/fgene.2016.00170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Zheng W, Zhang B, Cai Q, Sung H, Michailidou K, Shi J, et al. Common genetic determinants of breast-cancer risk in East Asian women: a collaborative study of 23 637 breast cancer cases and 25 579 controls. Hum Mol Genet. 2013;22(12):2539–50. doi: 10.1093/hmg/ddt089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Cai Q, Zhang B, Sung H, Low SK, Kweon SS, Lu W, et al. Genome-wide association analysis in East Asians identifies breast cancer susceptibility loci at 1q32.1, 5q14.3 and 15q26.1. Nat Genet. 2014;46(8):886–90. doi: 10.1038/ng.3041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Gail MH. Discriminatory accuracy from single-nucleotide polymorphisms in models to predict breast cancer risk. J Natl Cancer Inst. 2008;100(14):1037–41. doi: 10.1093/jnci/djn180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Mavaddat N, Pharoah PD, Michailidou K, Tyrer J, Brook MN, Bolla MK, et al. Prediction of breast cancer risk based on profiling with common genetic variants. J Natl Cancer Inst. 2015;107(5) doi: 10.1093/jnci/djv036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Vachon CM, Pankratz VS, Scott CG, Haeberle L, Ziv E, Jensen MR, et al. The contributions of breast density and common genetic variation to breast cancer risk. J Natl Cancer Inst. 2015;107(5) doi: 10.1093/jnci/dju397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Rudolph A, Song M, Brook MN, Milne RL, Mavaddat N, Michailidou K, et al. Joint associations of a polygenic risk score and environmental risk factors for breast cancer in the Breast Cancer Association Consortium. Int J Epidemiol. doi: 10.1093/ije/dyx242. In Press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Consortium CBCC-C. CHEK2*1100delC and susceptibility to breast cancer: a collaborative analysis involving 10,860 breast cancer cases and 9,065 controls from 10 studies. Am J Hum Genet. 2004;74(6):1175–82. doi: 10.1086/421251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Weischer M, Bojesen SE, Ellervik C, Tybjaerg-Hansen A, Nordestgaard BG. CHEK2*1100delC genotyping for clinical assessment of breast cancer risk: meta-analyses of 26,000 patient cases and 27,000 controls. J Clin Oncol. 2008;26(4):542–8. doi: 10.1200/JCO.2007.12.5922. [DOI] [PubMed] [Google Scholar]
- 79.Muranen TA, Greco D, Blomqvist C, Aittomaki K, Khan S, Hogervorst F, et al. Genetic modifiers of CHEK2*1100delC-associated breast cancer risk. Genet Med. 2017;19(5):599–603. doi: 10.1038/gim.2016.147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Kuchenbaecker KB, McGuffog L, Barrowdale D, Lee A, Soucy P, Dennis J, et al. Evaluation of Polygenic Risk Scores for Breast and Ovarian Cancer Risk Prediction in BRCA1 and BRCA2 Mutation Carriers. J Natl Cancer Inst. 2017;109(7) doi: 10.1093/jnci/djw302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Dite GS, MacInnis RJ, Bickerstaffe A, Dowty JG, Allman R, Apicella C, et al. Breast Cancer Risk Prediction Using Clinical Models and 77 Independent Risk-Associated SNPs for Women Aged Under 50 Years: Australian Breast Cancer Family Registry. Cancer Epidemiol Biomarkers Prev. 2016;25(2):359–65. doi: 10.1158/1055-9965.EPI-15-0838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Hughes E, Judkins T, Wagner S, Wenstrup R, Lanchbury JS, Gutlin A. Development and Validation of a Residual Risk Score to Predict Breast Cancer Risk in Unaffected Women Negative for Mutations on a Multi-Gene Hereditary Cancer Panel. ASCO Annual Meeting; 2017. [Google Scholar]
- 83.Shieh Y, Eklund M, Madlensky L, Sawyer SD, Thompson CK, Stover Fiscalini A, et al. Breast Cancer Screening in the Precision Medicine Era: Risk-Based Screening in a Population-Based Trial. J Natl Cancer Inst. 2017;109(5) doi: 10.1093/jnci/djw290. [DOI] [PubMed] [Google Scholar]
- 84.Varghese JS, Thompson DJ, Michailidou K, Lindstrom S, Turnbull C, Brown J, et al. Mammographic breast density and breast cancer: evidence of a shared genetic basis. Cancer Res. 2012;72(6):1478–84. doi: 10.1158/0008-5472.CAN-11-3295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Stone J, Thompson DJ, Dos Santos Silva I, Scott C, Tamimi RM, Lindstrom S, et al. Novel Associations between Common Breast Cancer Susceptibility Variants and Risk-Predicting Mammographic Density Measures. Cancer Res. 2015;75(12):2457–67. doi: 10.1158/0008-5472.CAN-14-2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Lindstrom S, Thompson DJ, Paterson AD, Li J, Gierach GL, Scott C, et al. Genome-wide association study identifies multiple loci associated with both mammographic density and breast cancer risk. Nat Commun. 2014;5:5303. doi: 10.1038/ncomms6303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Rudolph A, Chang-Claude J, Schmidt MK. Gene-environment interaction and risk of breast cancer. Br J Cancer. 2016;114(2):125–33. doi: 10.1038/bjc.2015.439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Travis RC, Reeves GK, Green J, Bull D, Tipper SJ, Baker K, et al. Gene-environment interactions in 7610 women with breast cancer: prospective evidence from the Million Women Study. Lancet. 2010;375(9732):2143–51. doi: 10.1016/S0140-6736(10)60636-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Nickels S, Truong T, Hein R, Stevens K, Buck K, Behrens S, et al. Evidence of gene-environment interactions between common breast cancer susceptibility loci and established environmental risk factors. PLoS Genet. 2013;9(3):e1003284. doi: 10.1371/journal.pgen.1003284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Schoeps A, Rudolph A, Seibold P, Dunning AM, Milne RL, Bojesen SE, et al. Identification of new genetic susceptibility loci for breast cancer through consideration of gene-environment interactions. Genet Epidemiol. 2014;38(1):84–93. doi: 10.1002/gepi.21771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Edwards SL, Beesley J, French JD, Dunning AM. Beyond GWASs: illuminating the dark road from association to function. Am J Hum Genet. 2013;93(5):779–97. doi: 10.1016/j.ajhg.2013.10.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Bojesen SE, Pooley KA, Johnatty SE, Beesley J, Michailidou K, Tyrer JP, et al. Multiple independent variants at the TERT locus are associated with telomere length and risks of breast and ovarian cancer. Nat Genet. 2013;45(4):371–84. 84e1–2. doi: 10.1038/ng.2566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Dunning AM, Michailidou K, Kuchenbaecker KB, Thompson D, French JD, Beesley J, et al. Breast cancer risk variants at 6q25 display different phenotype associations and regulate ESR1, RMND1 and CCDC170. Nat Genet. 2016;48(4):374–86. doi: 10.1038/ng.3521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Ghoussaini M, French JD, Michailidou K, Nord S, Beesley J, Canisus S, et al. Evidence that the 5p12 Variant rs10941679 Confers Susceptibility to Estrogen-Receptor-Positive Breast Cancer through FGF10 and MRPS30 Regulation. Am J Hum Genet. 2016;99(4):903–11. doi: 10.1016/j.ajhg.2016.07.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Lonsdale J, Thomas J, Salvatore M, Phillips R, Lo E, Shad S, et al. The Genotype-Tissue Expression (GTEx) project. Nature Genetics. 2013;45:580. doi: 10.1038/ng.2653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Hoffman JD, Graff RE, Emami NC, Tai CG, Passarelli MN, Hu D, et al. Cis-eQTL-based trans-ethnic meta-analysis reveals novel genes associated with breast cancer risk. PLoS Genet. 2017;13(3):e1006690. doi: 10.1371/journal.pgen.1006690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.French JD, Ghoussaini M, Edwards SL, Meyer KB, Michailidou K, Ahmed S, et al. Functional variants at the 11q13 risk locus for breast cancer regulate cyclin D1 expression through long-range enhancers. Am J Hum Genet. 2013;92(4):489–503. doi: 10.1016/j.ajhg.2013.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Betts JA, Moradi Marjaneh M, Al-Ejeh F, Lim YC, Shi W, Sivakumaran H, et al. Long Noncoding RNAs CUPID1 and CUPID2 Mediate Breast Cancer Risk at 11q13 by Modulating the Response to DNA Damage. Am J Hum Genet. 2017;101(2):255–66. doi: 10.1016/j.ajhg.2017.07.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.MacPherson G, Healey CS, Teare MD, Balasubramanian SP, Reed MW, Pharoah PD, et al. Association of a common variant of the CASP8 gene with reduced risk of breast cancer. J Natl Cancer Inst. 2004;96(24):1866–9. doi: 10.1093/jnci/dji001. [DOI] [PubMed] [Google Scholar]
- 100.Lin WY, Camp NJ, Ghoussaini M, Beesley J, Michailidou K, Hopper JL, et al. Identification and characterization of novel associations in the CASP8/ALS2CR12 region on chromosome 2 with breast cancer risk. Hum Mol Genet. 2015;24(1):285–98. doi: 10.1093/hmg/ddu431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Udler MS, Ahmed S, Healey CS, Meyer K, Struewing J, Maranian M, et al. Fine scale mapping of the breast cancer 16q12 locus. Hum Mol Genet. 2010;19(12):2507–15. doi: 10.1093/hmg/ddq122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Meyer KB, O’Reilly M, Michailidou K, Carlebur S, Edwards SL, French JD, et al. Fine-scale mapping of the FGFR2 breast cancer risk locus: putative functional variants differentially bind FOXA1 and E2F1. Am J Hum Genet. 2013;93(6):1046–60. doi: 10.1016/j.ajhg.2013.10.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Ghoussaini M, Edwards SL, Michailidou K, Nord S, Cowper-Sal Lari R, Desai K, et al. Evidence that breast cancer risk at the 2q35 locus is mediated through IGFBP5 regulation. Nat Commun. 2014;4:4999. doi: 10.1038/ncomms5999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Wyszynski A, Hong CC, Lam K, Michailidou K, Lytle C, Yao S, et al. An intergenic risk locus containing an enhancer deletion in 2q35 modulates breast cancer risk by deregulating IGFBP5 expression. Hum Mol Genet. 2016;25(17):3863–76. doi: 10.1093/hmg/ddw223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Darabi H, Beesley J, Droit A, Kar S, Nord S, Moradi Marjaneh M, et al. Fine scale mapping of the 17q22 breast cancer locus using dense SNPs, genotyped within the Collaborative Oncological Gene-Environment Study (COGs) Sci Rep. 2016;6:32512. doi: 10.1038/srep32512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Hall J, Marcel V, Bolin C, Fernet M, Tartier L, Vaslin L, et al. The associations of sequence variants in DNA-repair and cell-cycle genes with cancer risk: genotype-phenotype correlations. Biochem Soc Trans. 2009;37(Pt 3):527–33. doi: 10.1042/BST0370527. [DOI] [PubMed] [Google Scholar]
- 107.Thomas G, Jacobs KB, Kraft P, Yeager M, Wacholder S, Cox DG, et al. A multistage genome-wide association study in breast cancer identifies two new risk alleles at 1p11.2 and 14q24.1 (RAD51L1) Nat Genet. 2009;41(5):579–84. doi: 10.1038/ng.353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Lawrenson K, Kar S, McCue K, Kuchenbaeker K, Michailidou K, Tyrer J, et al. Functional mechanisms underlying pleiotropic risk alleles at the 19p13.1 breast-ovarian cancer susceptibility locus. Nat Commun. 2016;7:12675. doi: 10.1038/ncomms12675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Cai Q, Long J, Lu W, Qu S, Wen W, Kang D, et al. Genome-wide association study identifies breast cancer risk variant at 10q21.2: results from the Asia Breast Cancer Consortium. Hum Mol Genet. 2011;20(24):4991–9. doi: 10.1093/hmg/ddr405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Darabi H, McCue K, Beesley J, Michailidou K, Nord S, Kar S, et al. Polymorphisms in a Putative Enhancer at the 10q21.2 Breast Cancer Risk Locus Regulate NRBF2 Expression. Am J Hum Genet. 2015;97(1):22–34. doi: 10.1016/j.ajhg.2015.05.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Orr N, Dudbridge F, Dryden N, Maguire S, Novo D, Perrakis E, et al. Fine-mapping identifies two additional breast cancer susceptibility loci at 9q31.2. Hum Mol Genet. 2015;24(10):2966–84. doi: 10.1093/hmg/ddv035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Long J, Cai Q, Sung H, Shi J, Zhang B, Choi JY, et al. Genome-wide association study in east Asians identifies novel susceptibility loci for breast cancer. PLoS Genet. 2012;8(2):e1002532. doi: 10.1371/journal.pgen.1002532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Siddiq A, Couch FJ, Chen GK, Lindstrom S, Eccles D, Millikan RC, et al. A meta-analysis of genome-wide association studies of breast cancer identifies two novel susceptibility loci at 6q14 and 20q11. Hum Mol Genet. 2012;21(24):5373–84. doi: 10.1093/hmg/dds381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Gaudet MM, Kuchenbaecker KB, Vijai J, Klein RJ, Kirchhoff T, McGuffog L, et al. Identification of a BRCA2-specific modifier locus at 6p24 related to breast cancer risk. PLoS Genet. 2013;9(3):e1003173. doi: 10.1371/journal.pgen.1003173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Milne RL, Burwinkel B, Michailidou K, Arias-Perez JI, Zamora MP, Menendez-Rodriguez P, et al. Common non-synonymous SNPs associated with breast cancer susceptibility: findings from the Breast Cancer Association Consortium. Hum Mol Genet. 2014;23(22):6096–111. doi: 10.1093/hmg/ddu311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Sawyer E, Roylance R, Petridis C, Brook MN, Nowinski S, Papouli E, et al. Genetic predisposition to in situ and invasive lobular carcinoma of the breast. PLoS Genet. 2014;10(4):e1004285. doi: 10.1371/journal.pgen.1004285. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.