Abstract
This cohort study examines the association of concomitant genetics alterations with response to treatment with epidermal growth factor receptor tyrosine kinase inhibitors among patients with EGFR-mutant advanced non–small cell lung cancer.
Treatment with epidermal growth factor receptor (EGFR) tyrosine kinase inhibitors (TKIs) demonstrates superior efficacy, compared with chemotherapy, for patients with EGFR-mutant advanced non–small cell lung cancer (NSCLC).1 However, approximately 20% to 30% of patients who receive such treatment exhibit de novo resistance to therapy with EGFR TKIs.2 A better understanding of the underlying mechanisms of this phenomenon is imperative to optimize clinical outcomes.
Methods
This retrospective cohort study (conducted from January 1, 2012, to April 30, 2014) included 58 patients with metastatic NSCLC associated with EGFR (OMIM 131550) mutation (determined in tumor tissue) treated with first-line EGFR TKIs. The institutional review board of Sun Yat-sen University Cancer Center approved the study, and all of the patients provided written informed consent. Cell-free DNA obtained before treatment was subjected to next-generation sequencing of 49 cancer-related genes with use of a kit (Ion Pi Sequencing 200 kit v2; Thermo Fisher Scientific), as reported elsewhere, at a mean depth of 15 370X.3 The association between response and concomitant mutation was determined by logistic regression analysis. Data were analyzed from April 30 through November 30, 2017. Kaplan-Meier survival curves and Cox proportional hazards regression analysis were performed, and hazard ratios (HRs) and 95% CIs were calculated to determine the survival difference. Statistical significance was defined as a 2-sided P < .05.
Results
Among the 58 patients, the median age was 58.5 years (range, 40-81 years), 28 (48%) were male, 11 (19%) were smokers, and 56 (97%) had adenocarcinoma. Median progression-free survival and overall survival were 11.53 months (95% CI, 7.90-15.17) and 41.23 months (95% CI, not computable), respectively. Median survival time and 95% CIs are provided appropriately in the Figure, B and C. Objective response rate and disease control rate were 58.6% and 89.7%, respectively.
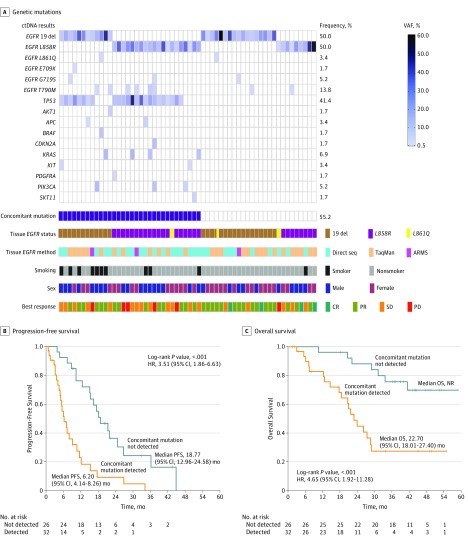
Figure. Genetic Alteration Profile of Circulating Tumor DNA (ctDNA) From Tyrosine Kinase Inhibitor (TKI)–Naive Patients With EGFR-Mutant Non–Small Cell Lung Cancer (NSCLC) and Its Association With Survival.
A, Genetic mutations identified by targeted next-generation sequencing in the plasma of 58 patients with NSCLC harboring EGFR mutation in tumor tissue. The sequencing covers 49 cancer-related genes and involves 2659 Catalogue of Somatic Mutations in Cancer (COSMIC) mutations. B, Kaplan-Meier curves of progression-free survival (PFS) in patients whose ctDNA had concomitant mutations compared with those without concomitant mutations. C, Kaplan-Meier curves of overall survival (OS) in patients whose ctDNA had concomitant mutations compared with those without concomitant mutations. ARMS indicates amplification refractory mutation system; CR, complete response; del, deletion; EGFR, epidermal growth factor receptor; HR, hazard ratio; NR, not reached; PD, progressive disease; PR, partial response; SD, stable disease; seq, sequencing; and VAF, variant allele frequency.
All the patients harbored actionable EGFR mutations in plasma, and 13 (22%) had multiple EGFR mutations, including 8 patients (14%) with co-occurring T790M mutations. Thirty-two patients (55%) harbored concomitant mutations. Patients with EGFR exon 21 mutation (69% [20 of 29] vs 41% [12 of 29]; P = .04) and those with a history of smoking (91% [10 of 11] vs 47% [22 of 47]; Fisher test, P = .02) were significantly associated with a higher incidence of concomitant mutation (Figure, A).
Concomitant mutation was significantly associated with reduced objective response rate (44% vs 77%; P = .01), shorter duration of progression-free survival (Figure, B; median survival, 6.20 months (95% CI, 4.14-8.26 months) vs 18.77 (95% CI, 12.96-24.58) months; HR, 3.51 [95% CI, 1.86-6.63]; P < .001), and shorter duration of overall survival (Figure, C; median survival, 22.70 months (95% CI, 18.01-22.30 months) vs not reached; HR, 4.65 [95% CI, 1.92-11.28]; P < .001). In multivariable analysis, concomitant mutation remained significantly associated with poorer survival after adjusting for EGFR subtype and patient characteristics (Table).
Table. Treatment Outcomes for Patients With EGFR-Mutant Advanced Non–Small Cell Lung Cancer Treated With EGFR TKIs.
| Variable | No. (%) | Objective Response | Progression-Free Survival | Overall Survival | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| No. (%) | P Value | Univariate | Multivariablea | Univariate | Multivariablea | ||||||
| HR (95% CI) | P Value | HR (95% CI) | P Value | HR (95% CI) | P Value | HR (95% CI) | P Value | ||||
| Type of TKI | |||||||||||
| Gefitinib | 50 (86.2) | 28 (56.0) | .32 | 1 [Reference] | .41 | 1 [Reference] | .75 | 1 [Reference] | .95 | 1 [Reference] | .33 |
| Erlotinib hydrochloride | 8 (13.8) | 6 (75.0) | 1.46 (0.59-3.60) | 0.85 (0.31-2.34) | 1.04 (0.36-3.02) | 0.54 (0.16-1.85) | |||||
| EGFR statusb | |||||||||||
| Exon 19 deletion | 29 (50.0) | 21 (72.4) | .04 | 1 [Reference] | .03 | 1 [Reference] | .13 | 1 [Reference] | .04 | 1 [Reference] | .29 |
| Exon 21 mutation | 29 (50.0) | 13 (44.8) | 1.95 (1.05-3.60) | 1.65 (0.87-3.11) | 2.29 (1.04-5.05) | 1.60 (0.67-3.81) | |||||
| EGFR T790M mutationb | |||||||||||
| Positive | 8 (13.8) | 2 (25.0) | .055 | 1 [Reference] | <.001 | 1 [Reference] | <.001 | 1 [Reference] | .28 | 1 [Reference] | .35 |
| Negative | 50 (86.2) | 32 (64.0) | 6.39 (2.58-15.87) | 5.80 (2.20-15.32) | 1.81 (0.62-5.29) | 1.72 (0.55-5.37) | |||||
| Concomitant mutationsb | |||||||||||
| Yes | 32 (55.2) | 14 (43.8) | .01 | 1 [Reference] | <.001 | 1 [Reference] | .001 | 1 [Reference] | <.001 | 1 [Reference] | .002 |
| No | 26 (44.8) | 20 (76.9) | 3.51 (1.86-6.63) | 3.49 (1.71-7.14) | 4.65 (1.92-11.28) | 4.82 (1.78-13.07) | |||||
Abbreviations: EGFR, epidermal growth factor receptor; HR, hazard ratio; TKI, tyrosine kinase inhibitor.
Age (stratified by 65 years old), sex, history of smoking, type of TKI, EGFR status, EGFR T790M mutation, and concomitant mutations were entered into the multivariable Cox proportional hazards regression model.
Based on circulating tumor DNA sequencing results.
Discussion
Similar to a recent study, our results might challenge the current view that EGFR-mutant NSCLC is a single-oncogene–driven disease.4 This study also reveals the widespread existence of concomitant mutations in patients with EGFR-mutant advanced NSCLC. In addition, genetic coalterations negatively affect the response and survival of patients who receive first-line treatment with EGFR TKIs, plausibly because of the bypass activation of survival signaling pathways or tumor heterogeneity.
Patients with EGFR exon 19 deletion were found to have longer survival than those with exon 21 mutation, although the underlying mechanisms remain unknown.5 Of interest, we found that patients with exon 21 mutation had a significantly higher incidence of concomitant mutation. In multivariable analysis, the EGFR subtype was no longer significantly associated with survival, which suggests that the inferior treatment outcome associated with exon 21 mutation could be attributed to or confounded by the higher incidence of concomitant mutation. The presence of EGFR T790M mutation at baseline also was associated with poorer efficacy of first-generation EGFR TKIs, which provides a rationale for the superiority of osimeritinib therapy over therapy with earlier-generation TKIs in untreated EGFR-mutant NSCLC.6
Major limitations of this study were the sample size, the single-centered design, and the lack of serial biopsies. In addition, the data from our hot spot next-generation sequencing (not including other mutations, copy number alterations, or chromosomal abnormalities) might represent an underestimation of concomitant mutation and prevent an in-depth analysis of signaling pathway or clonality.
These limitations aside, our results suggest that the common presence of co-occurring genetic alterations may be associated with resistance to EGFR TKIs in patients with EGFR-mutant NSCLC. Our study highlights the importance of deploying multiplex molecular profiling and conducting research on the use of polytherapy or sequential therapy to address the coalterations that drive drug resistance.
References
- 1.Lee JK, Hahn S, Kim DW, et al. Epidermal growth factor receptor tyrosine kinase inhibitors vs conventional chemotherapy in non–small cell lung cancer harboring wild-type epidermal growth factor receptor: a meta-analysis. JAMA. 2014;311(14):1430-1437. [DOI] [PubMed] [Google Scholar]
- 2.Maemondo M, Inoue A, Kobayashi K, et al. ; North-East Japan Study Group . Gefitinib or chemotherapy for non–small-cell lung cancer with mutated EGFR. N Engl J Med. 2010;362(25):2380-2388. [DOI] [PubMed] [Google Scholar]
- 3.Xu S, Lou F, Wu Y, et al. Circulating tumor DNA identified by targeted sequencing in advanced-stage non–small cell lung cancer patients. Cancer Lett. 2016;370(2):324-331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Blakely CM, Watkins TBK, Wu W, et al. Evolution and clinical impact of co-occurring genetic alterations in advanced-stage EGFR-mutant lung cancers. Nat Genet. 2017;49(12):1693-1704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Lee CK, Wu YL, Ding PN, et al. Impact of specific epidermal growth factor receptor (EGFR) mutations and clinical characteristics on outcomes after treatment with EGFR tyrosine kinase inhibitors versus chemotherapy in EGFR-mutant lung cancer: a meta-analysis. J Clin Oncol. 2015;33(17):1958-1965. [DOI] [PubMed] [Google Scholar]
- 6.Soria JC, Ohe Y, Vansteenkiste J, et al. Osimertinib in untreated EGFR-mutated advanced non–small-cell lung cancer. N Engl J Med. 2018;378(2):113-125. [DOI] [PubMed] [Google Scholar]