Abstract
Platelet factor 4 (PF4) is one of the most abundant cationic proteins secreted from α-granules of activated platelets. Based on its structure, PF4 was assigned to the CXC family of chemokines and has been shown to have numerous effects on myeloid leukocytes. However, the receptor for PF4 remains unknown. Here, we demonstrate that PF4 induces leukocyte responses through the integrin Mac-1 (αMβ2, CD11b/CD18). Human neutrophils, monocytes, U937 monocytic and HEK293 cells expressing Mac-1 strongly adhered to immobilized PF4 in a concentration-dependent manner. The cell adhesion was partially blocked by anti-Mac-1 mAb and inhibition was enhanced when anti-Mac-1 antibodies were combined with glycosaminoglycans, suggesting that cell-surface proteoglycans act cooperatively with Mac-1. PF4 also induced Mac-1-dependent migration of human neutrophils and murine WT, but not Mac-1-deficient macrophages. Coating of Escherichia coli bacteria or latex beads with PF4 enhanced their phagocytosis by macrophages by ∼4-fold, and this process was blocked by different Mac-1 antagonists. Furthermore, PF4 potentiated phagocytosis by WT, but not Mac-1-deficient macrophages. As determined by biolayer interferometry, PF4 directly bound the αMI-domain, the major ligand-binding region of Mac-1, and this interaction was governed by a Kd of 1.3 ± 0.2 μm. Using the PF4-derived peptide library, synthetic peptides duplicating the αMI-domain recognition sequences and recombinant mutant PF4 fragments, the binding sites for αMI-domain were identified in the PF4 segments Cys12–Ser26 and Ala57–Ser70. These results identify PF4 as a ligand for the integrin Mac-1 and suggest that many immune-modulating effects previously ascribed to PF4 are mediated through its interaction with Mac-1.
Keywords: platelet, integrin, chemokine, macrophage, phagocytosis, alarmins, Mac-1, PF4
Introduction
Platelet adhesion and aggregation at sites of vessel wall injury are crucial events to prevent blood loss and initiate wound healing. During these processes, activated platelets secrete numerous molecules from their dense and α-granules that aid in thrombus formation and participate in blood coagulation. Thrombi are also known to contain leukocytes, mainly neutrophils and monocytes that invade formed thrombi and apparently contribute to the removal of platelet and fibrin deposits at later stages of thrombus remodeling (1, 2). Previous studies suggested that molecules secreted from platelet α-granules promote directed intravascular leukocyte migration to and through platelet thrombi (2, 3). Moreover, it has been shown that platelet-released products exert numerous in vitro immune-modulating effects. These mediators, which include platelet factor 4 (PF4),2 platelet basic protein and its derivatives (CTAP-III and NAP-2), epithelial-activating peptide-78 (ENA-78), thymosin-β4, MIP-1α, RANTES (regulated on activation normal T cell expressed and secreted), and others, induce leukocyte migration, activation, and degranulation, and promote phagocytosis of bacteria (4–7). Among these, PF4 and NAP-2 are the most abundant (3, 4). These molecules are known as chemokines based on their structural similarity with other members of the CXC chemokine subfamily and chemotactic activity (4, 8). However, whereas chemotactic activity of NAP-2 (CXCL7) has partially been attributed to the CXCR1/2 G protein-coupled receptors on leukocytes (9, 10), no receptor for PF4 (CXCL4) was identified.
We have recently characterized the binding properties of integrin receptor αMβ2 (Mac-1, CD11b/CD18), a major receptor on the surface of myeloid leukocytes that exhibits broad ligand recognition specificity and mediates numerous responses of these cells (11, 12). These investigations identified motifs present in many Mac-1 ligands (12). In particular, we found that the αMI-domain, a ligand-binding region of Mac-1, binds not to specific amino acid sequence(s), but rather has a preference for the sequence patterns consisting of a core of basic residues flanked by hydrophobic residues. Such αMI-domain recognition motifs have been discovered in several known Mac-1 ligands, including neutrophil elastase (13), myeloperoxidase (14), and azurocidin (15). Based on this finding, we proposed that many cationic host defense proteins/peptides stored in leukocyte granules, which are strikingly enriched in the αMI-domain recognition patterns represent a new class of Mac-1 ligands. Furthermore, many of these cationic proteins/peptides also belong to a group of the so-called alarmins, i.e. the molecules that are sequestered within cells under normal physiological conditions but would function as alarm signals for the immune system upon being exposed during tissue injury by exerting chemotactic and activating effects on leukocytes (16, 17). Indeed, by testing several cationic proteins/peptides, including the human cathelicidin peptide LL-37 and dynorphin A/B we showed that they induce a potent Mac-1-dependent chemotactic response in monocytes/macrophages, activate neutrophils, and augment phagocytosis by opsonizing bacteria (12, 18, 19).
Because PF4 is a basic protein and in its native tetrameric form displays a prominent equatorial ring of positively charged and hydrophobic amino acids, we hypothesized that it may be a candidate ligand for Mac-1. In the present study, we demonstrated that PF4 contains the sequences that represent a distinctive feature of the αMI-domain recognition specificity toward cationic proteins and provided direct evidence that PF4 binds the αMI-domain. We also demonstrated that PF4 supported various Mac-1-dependent leukocyte responses, including adhesion, migration, phagocytosis, and integrin clustering. Furthermore, we have identified two segments in PF4 as binding sites for the αMI-domain. Collectively, these data identify PF4 as a ligand of Mac-1 and suggest that similar to other cationic Mac-1 ligands, PF4's ability to induce leukocyte responses qualifies it as a platelet-derived alarmin.
Results
Screening of the PF4-derived peptide library for αMI-domain binding
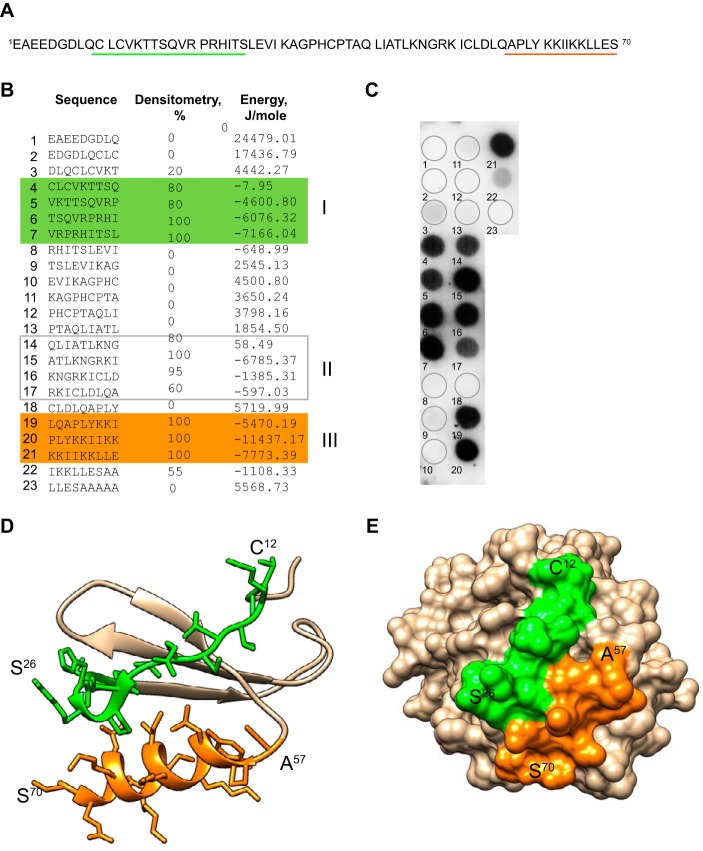
We previously developed the computer program that allows the prediction of potential Mac-1 ligands by examining the presence of putative binding sites for the αMI-domain, a ligand recognition region of Mac-1 (12). The program analyzes a peptide library made of overlapping peptides spanning the sequence of a prospective Mac-1 ligand and assigns each peptide the energy value that serves as a measure of probability that the αMI-domain binds this sequence: the lower the energy, the higher the likelihood that the sequence binds the αMI-domain. The analyses of the library spanning the sequence of PF4 (Fig. 1A) predicted the presence of several αMI-domain recognition sequences (Fig. 1B). To confirm this prediction, we synthesized a library consisting of 9-mer peptides with a 3-residue offset spanning the sequence of PF4 (Fig. 1B) and tested it for binding of 125I-labeled active αMI-domain (αM Glu123–Lys315) (Fig. 1C). Densitometry analyses of the library indicated the presence of three segments containing strong αMI-domain-binding peptides (spots 4–7, spots 14–17, and spots 19–21) (Fig. 1B). Moreover, energy scores of these peptides correlated with their αMI-domain-binding activity (Fig. 1, B and C). In particular, two overlapping peptides in region 3, 58PLYKKIIKK66 (spot 20) and 61KKIIKKLLE69 (spot 21) contain the strong αMI-domain motif 58PLYKKIIKKLL68, in which two lysine-containing clusters are surrounded by hydrophobic residues. Based on the 3D structure of PF4 (Fig. 1, D and E), only two of these segments, 12CVKTTSQVRPRHITS26 (spots 4–7) and 57APLYKKIIKKLLES70 (spots 19–21), are fully exposed on the surface, either in the monomer or tetrameric forms. In the third sequence, 43ATLKNGRKI51 (spot 15), only LKNG, a sequence that targets PF4 to α-granules (20) forms an exposed loop, whereas side chains of other residues are buried. The finding that PF4 contains sequences enriched in basic and hydrophobic residues characteristic for the αMI-domain recognition specificity toward cationic proteins suggests that PF4 has the capacity to interact with the αMI-domain.
Figure 1.
Identification of the αMI-domain recognition motifs in PF4. A, the amino acid sequence of human PF4. The underlined sequences Cys12–Ser26 and Ala57–Ser70 denote the αMI-domain-binding sites and are colored in green and orange, respectively. B, the peptide library derived from the PF4 sequence (left column). The peptide energies (right column) that serve as a measure of probability each peptide can interact with the αMI-domain were calculated as described (12). C, autoradiography analysis of binding of 125I-labeled αMI-domain to the PF4-derived peptide library. The membrane was blocked with 1% BSA and then incubated with 10 μg/ml of 125I-labeled αMI-domain in Tris-buffered saline containing 1 mm MgCl2. After washing, the membrane was dried and the αMI-domain binding was visualized by autoradiography. The αMI-domain binding observed as dark spots was analyzed by densitometry. The numbers shown in the middle column in B indicate the relative binding of the αMI-domain expressed as a percentage of the intensity of spot 6. The ribbon model of the PF4 monomer (D) and space-filling models of PF4 tetramer (E) were based on PDB code 1F9Q (45) with putative αMI-domain-binding sites identified by screening of the PF4-derived peptide library. The Cys12–Ser26 and Ala57–Ser70 sequences are shown in green and orange, respectively.
Direct interaction of αMI-domain and PF4
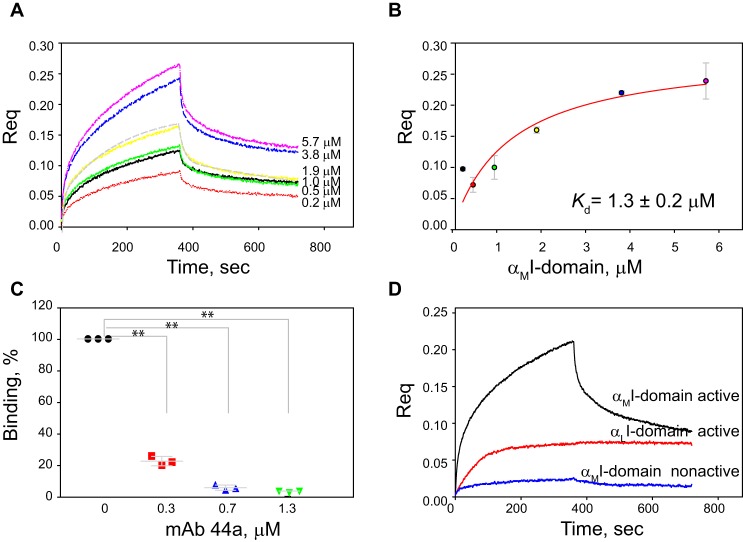
Biolayer interferometry was used to examine the interaction of PF4 coupled on the sensor chip with the mobile phase αMI-domain. Titration of the αMI-domain resulted in the binding isotherm with the interaction Kd of 1.3 ± 0.2 μm (Fig. 2, A and B). Analyses of the isotherm indicated that the binding kinetics were not consistent with a simple 1:1 interaction model as also noted for several other αMI ligands (21). Rather, the analysis using a 2:1 heterogeneous ligand fitting model showed a high-quality fit of the experimental data (Fig. 2A, shown in gray for 1.9 μm). As a specificity control, anti-αM mAb 44a inhibited PF4–αMI-domain interaction in a concentration-dependent manner (Fig. 2C). In contrast to the active form of the αMI-domain, no interaction of PF4 with nonactive αMI-domain (αM Gln119–Glu333) was detected (Fig. 2D). As expected, binding of active αLI-domain of the integrin αLβ2 (LFA-1), a receptor with a narrow ligand binding specificity (22) was low (Fig. 2D). These data demonstrate that soluble PF4 directly binds αMI-domain and this interaction requires the active form of αMI-domain.
Figure 2.
Biolayer interferometry analysis of the PF4–αMI-domain interaction. A, representative sensograms of binding of active αMI-domain (0; 0.2; 0.5; 1.0; 1.9; 3.8; 5.7 μm) to PF4 immobilized on the ForteBio sensor. The 2:1 heterogeneous ligand fitting model for 1.9 μm of the αMI-domain is shown in gray. B, the binding isotherm of the αMI-domain–PF4 interaction. Data shown are mean ± S.E. from 3 separate experiments. C, effect of anti-αM mAb 44a on the PF4–αMI domain interaction. Data shown are mean ± S.E. from 3 separate experiments. **, p ≤ 0.01. D, comparison of different I-domains for their ability to bind PF4. Representative sensograms were obtained with active αMI-domain, inactive αMI-domain, and active αLI-domain tested at 3.8 μm.
PF4 supports adhesion of Mac-1-expressing cells
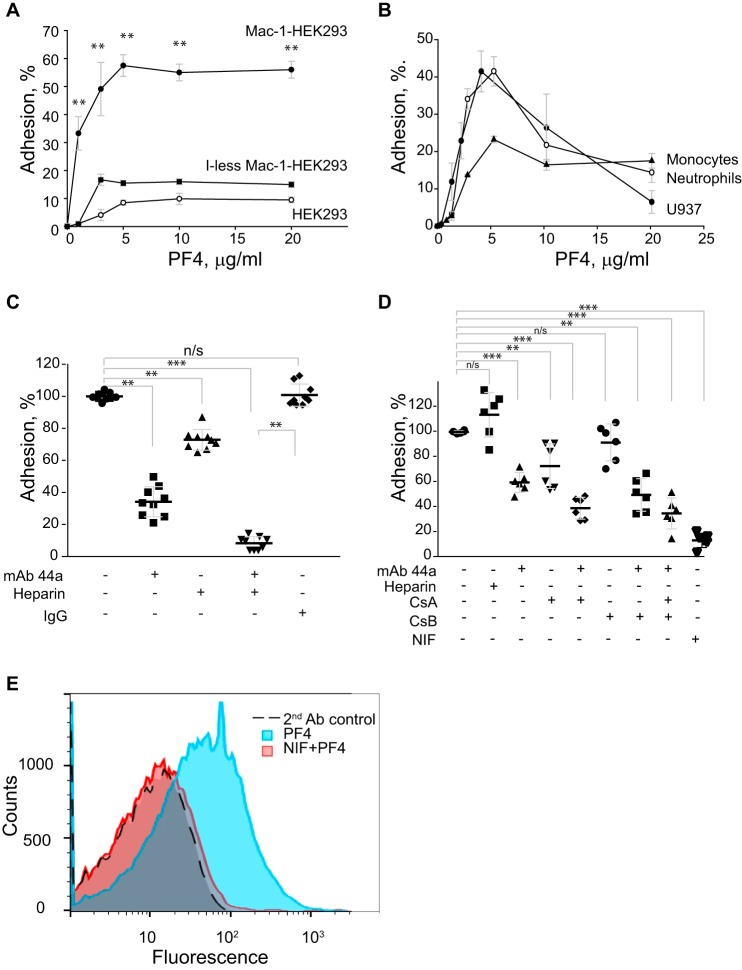
The finding that the αMI-domain directly interacts with PF4 suggests that PF4 may bind Mac-1 on the cell surface. To investigate this possibility, we performed adhesion with immobilized PF4 using various Mac-1-expressing cells. As shown in Fig. 3A, PF4 supported efficient adhesion of HEK293 cells stably transfected with Mac-1 (Mac-1-HEK293), a well-established model to test the interaction of Mac-1 with its ligands (11, 23). The adhesion was dose-dependent with saturation achieved at ∼5 μg/ml of PF4 and 55 ± 4% of added cells adhered. In contrast, WT HEK293 cells and HEK293 cells expressing the I-less form of Mac-1 adhered poorly (8 ± 1 and 16 ± 2% for WT and the I-less HEK293 cells, respectively) (Fig. 3A). Non-stimulated human neutrophils, monocytes, and monocytic U937 cells, naturally expressing Mac-1, also adhered to PF4; however, the pattern of adhesion was different. Cell adhesion was dose-dependent in the range 0–5 μg/ml of PF4 coating concentrations, after which adhesion gradually decreased (Fig. 3B). The role of Mac-1 in the interaction of Mac-1–HEK293 cells with PF4 was further determined using function-blocking mAb 44a directed to the human αM integrin subunit. This mAb inhibited adhesion of Mac-1–HEK293 cells and U937 in a dose-dependent manner (Fig. S1) and at 10 μg/ml inhibited cell adhesion by 69 ± 7 and 63 ± 2% for Mac-1–HEK293 and U937 cells, respectively. An isotype control IgG1 for mAb 44a was not active.
Figure 3.
PF4 supports adhesion of the αMβ2-expressing cells. A, aliquots (100 μl; 5 × 105/ml) of Mac-1-expressing HEK293 (Mac-1–HEK293), WT HEK293 (HEK293), and HEK293 cells expressing the I-less Mac-1 labeled with calcein were added to microtiter wells coated with different concentrations of PF4 and post-coated with 1% PVP. After 30 min at 37 °C, nonadherent cells were removed by washing and fluorescence of adherent cells was measured in a fluorescence plate reader. The number of adherent cells was determined by using the fluorescence of 100-μl aliquots with a known number of labeled cells. Data are expressed as a percentage of added cells and are mean ± S.E. from 3 separate experiments with triplicate measurements. **, p ≤ 0.01. B, adhesion of human neutrophils, monocytes, and U937 monocytic cells to microtiter wells coated with different concentrations of PF4. Data are expressed as a percentage of added cells and are mean ± S.E. from 3 separate experiments with triplicate measurements. C, Mac-1–HEK293 cells were preincubated with anti-αM mAb 44a (10 μg/ml), heparin (10 μg/ml; 2 units/ml), or their mixture (5 μg/ml of mAb44a + 5 μg/ml of heparin (1 units/ml)) and added to wells coated with 5 μg/ml of PF4. Adhesion in the absence of Mac-1 inhibitors and heparin was assigned a value of 100%. Data shown are mean ± S.E. from 3 separate experiments with triplicate measurements. **, p ≤ 0.01; ***, p ≤ 0.001 compared with control adhesion in the absence of inhibitors. D, calcein-labeled neutrophils were preincubated with 10 μg/ml of each anti-αM mAb 44a, heparin, CsA, or CsB for 15 min. Cells were also preincubated with the mixtures of mAb 44a with each glycosaminoglycan or the mixture of all three reagents. In addition, neutrophils were preincubated with 1 μg/ml of NIF. Adhesion in the absence of Mac-1 inhibitors and glycosaminoglycans was assigned a value of 100%. Data shown are mean ± S.E. from 3 separate experiments with duplicate measurements. **, p ≤ 0.01; ***, p ≤ 0.001 compared with control adhesion in the absence of inhibitors. E, binding of PF4 to human neutrophils assessed by flow cytometry. Human neutrophils were incubated with PF4 (20 μg/ml) in the presence or absence of NIF (1 μg/ml). PF4 binding was detected using polyclonal anti-PF4 antibody and Alexa 488-conjugated goat anti-rabbit antibody.
PF4 has a high affinity for heparin (24) and negatively charged proteoglycans (25, 26). Furthermore, it is well-known that Mac-1 on monocytes and Mac-1–HEK293 cells cooperates with cell-surface heparan sulfate proteoglycans (HSPGs) during cell adhesion to the extracellular matrix protein CCN1 and to several other Mac-1 ligands (18, 19, 27, 28). To investigate whether HSPGs participate in cell adhesion to PF4, we examined the effect of heparin on adhesion of Mac-1–HEK293 cells. Heparin at 10 μg/ml partially inhibited adhesion (∼30%) (Fig. 3C). In addition, when cells were treated with both anti-αM mAb 44a (5 μg/ml) and heparin (5 μg/ml) cell adhesion was completely blocked (Fig. 3C).
We next examined the ability of Mac-1 to cooperate with glycosaminoglycans during adhesion of neutrophils to PF4. Although mAb 44a partially inhibited adhesion (39 ± 3%), heparin had no effect (Fig. 3D), consistent with previous findings that neutrophils lack detectable amounts of HSPGs (29, 30). It has been reported that chondroitin sulfates (CSs) inhibit binding of PF4 to neutrophils (29) and that CS proteoglycans (CSPGs) bind PF4 (26, 29, 30). Therefore, we tested the effect of CSs alone and in combination with mAb 44a on neutrophil adhesion to PF4. At 10 μg/ml, CSA and CSB inhibited neutrophil adhesion only by 27 ± 7 and 9 ± 6%, respectively (Fig. 3D). However, when cells were treated with the mixture of mAb 44a and each glycosaminoglycan, adhesion was reduced by 59 ± 3 and 49 ± 5% for CSA and CSB, respectively. The combination of all three reagents reduced adhesion by 63 ± 4%, suggesting that on the surface of neutrophils Mac-1 may cooperate with CSPGs in PF4 binding. Supporting the role of Mac-1 in adhesion to PF4, NIF (neutrophil inhibitory factor), a specific inhibitor of Mac-1, inhibited neutrophil adhesion by 86 ± 2% (Fig. 3D).
Because immobilization of PF4 on plastic can potentially alter its conformation leading to exposure of the Mac-1-binding sites (12) and because binding of soluble PF4 may better reproduce its physiologic presentation to cells, we also examined binding of soluble PF4 to human neutrophils using flow cytometry. As shown on Fig. 3E, PF4 bound to neutrophils and this interaction was completely inhibited by NIF, indicating that this interaction was mediated by Mac-1.
Identification of the Mac-1-binding sequences within PF4
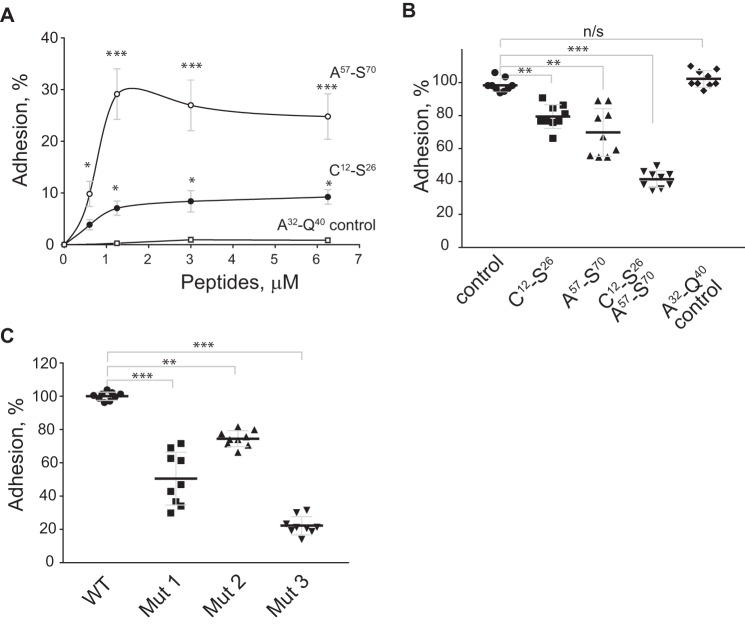
We examined the ability of the αMI-domain–binding peptides identified through screening of the PF4 library to support adhesion of Mac-1–HEK293 cells. As shown in Fig. 4A, two synthetic peptides Cys12–Ser26 and Ala57–Ser70 supported adhesion, with Ala57–Ser70 being significantly more active. A control peptide 32AGPHCPTAQ40 (ΔG = 6631 J/mol) was not active. Both soluble peptides also inhibited cell adhesion to immobilized PF4, and on a molar basis, Ala57–Ser70 was more active than Cys12–Ser26 (23 ± 3 versus 34 ± 8%, respectively), whereas the control peptide was inactive. When added together at equimolar concentrations (30 μm), the peptides blocked adhesion by 59 ± 2% (Fig. 4B). To examine whether these sequences within PF4 might participate in Mac-1 binding, we expressed recombinant mutant PF4 fragments in which Arg20 and Arg22 in Cys12–Ser26 were mutated to glycine (Mutant 1), or the C-terminal Ala57–Ser70 helix was truncated (Mutant 2) or both mutations were present (Mutant 3) (Fig. S2). Mutants 1 and 2 supported cell adhesion less efficiently than WT PF4 (52 ± 12 and 75 ± 3%), and adhesion-promoting activity of Mutant 3 was reduced by ∼5-fold compared with WT protein (Fig. 4C). These results indicate that both Cys12–Ser26 and Ala57–Ser70 are involved in the interaction with Mac-1.
Figure 4.
PF4-derived peptides Cys12–Ser26 and Ala57–Ser70 support adhesion of the Mac-1-expressing cells. A, aliquots (100 μl; 5 × 105/ml) of Mac-1–HEK293 cells were labeled with calcein and added to microtiter wells coated with different concentrations of the Cys12–Ser26 and Ala57–Ser70 peptides (0–6.5 μm). After 30 min at 37 °C, nonadherent cells were removed and adhesion was measured. The data shown are mean ± S.E. from four experiments with triplicate determinations at each point. B, calcein-labeled Mac-1–HEK293 cells were incubated with soluble Cys12–Ser26 and Ala57–Ser70 (30 μm) or their mixture for 15 min at 22 °C and added to wells coated with 5 μg/ml of PF4 and post-coated with 1% PVP. Adhesion in the absence of peptides was assigned a value of 100%. The data shown are the mean ± S.E. from three experiments each with triplicate determinations. **, p ≤ 0.01; ***, p ≤ 0.001. C, adhesion of Mac-1–HEK293 cells to WT and mutant PF4 immobilized at 10 μm. Cell adhesion to WT PF4 was assigned a value of 100%. The data shown are the mean ± S.E. from three experiments each with triplicate determinations. **, p ≤ 0.01; ***, p ≤ 0.001.
Mac-1-mediated cell migration to PF4
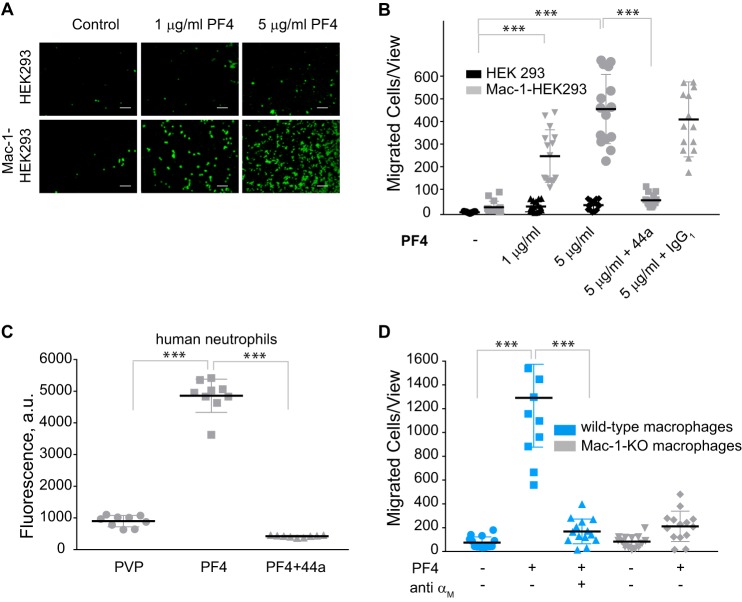
Previous studies demonstrated that PF4 and its C-terminal peptide Pro58–Ser70 were capable of inducing cell migration (8, 31). We re-examined the ability of PF4 to trigger the migratory response of various cells using a Transwell system and also determined whether migration depended on Mac-1. The Transwell membranes were coated with different concentrations of PF4 and Mac-1–HEK293 cells placed in the upper chambers and cells were allowed to migrate. As shown in Fig. 5, A and B, PF4 induced a strong concentration-dependent migratory response of Mac-1–HEK293 cells. WT HEK293 cells did not migrate to PF4 and Mac-1–HEK293 cells migrated only slightly through PVP-coated membranes (Fig. 5, A and B). Direct evidence that migration depended on Mac-1 was obtained in the experiments in which cells were preincubated with anti-Mac-1 function blocking mAb 44a. The mAb inhibited migration to the level observed with HEK293 cells. IgG1 isotype control for these antibodies had no effect (Fig. 5B).
Figure 5.
Migration of Mac-1-expressing cells to PF4 in a Transwell system. A, Transwell inserts were coated with different concentrations of PF4 (1–5 μg/ml) for 60 min at 37 °C. Mac-1-expressing or WT HEK293 cells (100 μl at 3 × 106/ml) were added to the upper wells of the Transwell chamber, and their ability to migrate was analyzed. After 16 h at 37 °C, the cells were labeled with calcein for 30 min at 37 °C. The cells from the upper chamber of the Transwells were removed by wiping with a cotton-tipped applicator, and images of the cells on the underside of the Transwell filter were taken. The figure is representative of 5 experiments. The scale bar is 100 μm. B, images from A were analyzed, and cells that migrated were counted. Data are presented as numbers of migrated cells per field ± S.E. for five random fields per well from five individual experiments. ***, p ≤ 0.001. C, calcein-labeled human neutrophils were placed in the upper chamber and allowed to migrate to PF4 (5 μg/ml) for 90 min. Data are expressed as the fluorescence of c-labeled cells migrated to the lower chamber. Results are the mean ± S.E. from three independent experiments with triplicate samples. ***, p ≤ 0.001. D, migration of macrophages isolated from WT and Mac-1-deficient mice. Macrophages (3 × 105) were placed in the upper chamber and allowed to migrate to PF4 (5 μg/ml) for 90 min. The number of migrated cells was determined as in A. Data are presented as number of migrated cells per field ± S.E. for five random fields per well from 3 experiments. ***, p ≤ 0.001.
Migration of human neutrophils to PF4 was also examined and found to increase by ∼4-fold compared with migration through membranes coated with PVP (Fig. 5C). Preincubation of cells with mAb 44a almost completely eliminated migration. In another set of experiments, we evaluated migration of macrophages isolated from the peritoneum of WT and Mac-1-deficient mice. mAb M1/70 against the mouse αM integrin subunit efficiently blocked migration of WT macrophages (Fig. 5D). Furthermore, whereas WT microphages migrated to PF4, migration of Mac-1-deficient cells was significantly impaired (Fig. 5D) suggesting that the engagement by Mac-1 of PF4 triggers the migratory response.
Binding of PF4 induces redistribution of Mac-1 on the cell surface
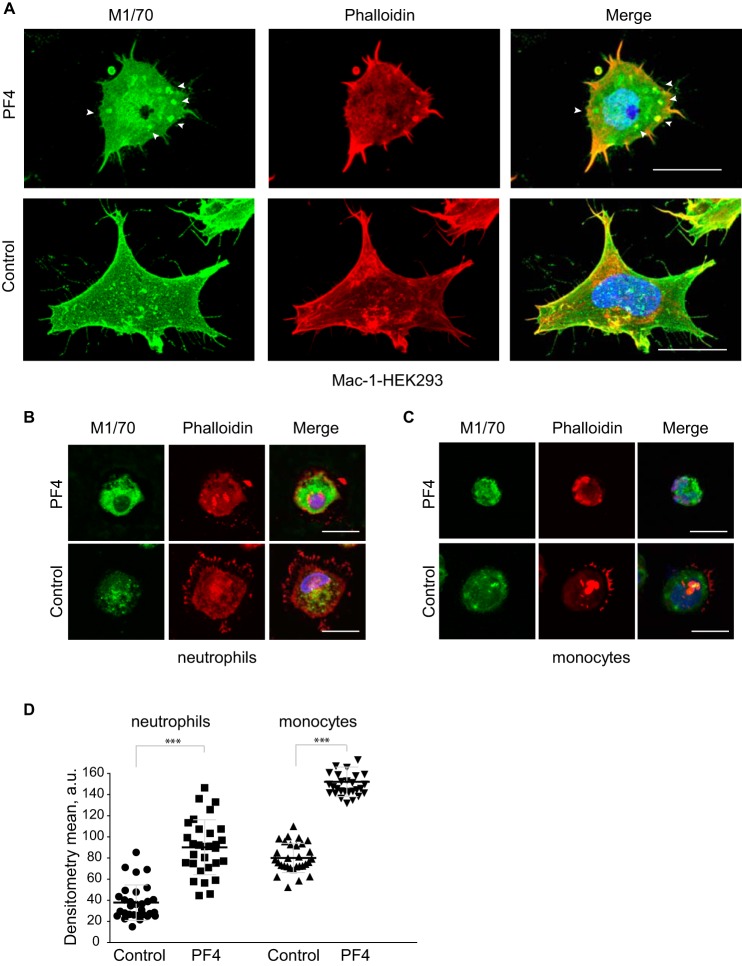
To examine whether binding of PF4 induces changes of Mac-1 distribution on the cell surface, soluble PF4 was added to adherent Mac-1–HEK293 cells. In control untreated cells, Mac-1 was concentrated at the cell periphery with some integrins evenly distributed over the cell body (Fig. 6A, lower panel, and Fig. S3B). After addition of PF4 for 30 min, 72 ± 5% of examined cells (n = 90) contained integrin clusters at the apical side of the cells (Fig. 6A, upper panel, and Fig. S3A). In addition, many of the Mac-1-containing clusters colocalized with actin. Quantitative analyses showed that PF4-treated cells were more rounded than untreated cells (Fig. S4). Moreover, the number of adherent PF4-treated cells was decreased by 33 ± 4% compared with untreated cells, suggesting that PF4 might have weakened adhesion.
Figure 6.
Effect of PF4 on the redistribution of Mac-1 on the cell surface. A–C, upper panels: adherent Mac-1–HEK293 cells (A), neutrophils (B), and monocytes (C) were treated with soluble PF4 (100 μg/ml) for 30 min, fixed, and incubated with anti-αM mAb M1/70 followed by Alexa 488-conjugated goat anti-rat secondary antibody. Bottom panels in A, B, and C, control cells were incubated with medium alone. In addition, cells were stained with Alexa Fluor 546-conjugated phalloidin and 4′,6-diamidino-2-phenylindole. The scale bars are 15 μm. D, analysis of Mac-1 expression on the surface of neutrophils and monocytes. Data are mean ± S.E. from the measurement of fluorescence of 30 cells using ImageJ software. ***, p ≤ 0.001.
To test whether the effect of PF4 could be general to other Mac-1-expressing cells, soluble PF4 was added to neutrophils and monocytes. Treatment of adherent cells with PF4 resulted in up-regulation of Mac-1 on the cell surface by ∼2.4- and 1.9-fold, respectively (Fig. 6, B–D). Moreover, similar to Mac-1–HEK293 cells, PF4 caused changes in the morphology of neutrophils and monocytes rendering them more rounded (Fig. 6, B and C, upper panels).
PF4 augments phagocytosis of bacteria or plastic beads acting via Mac-1
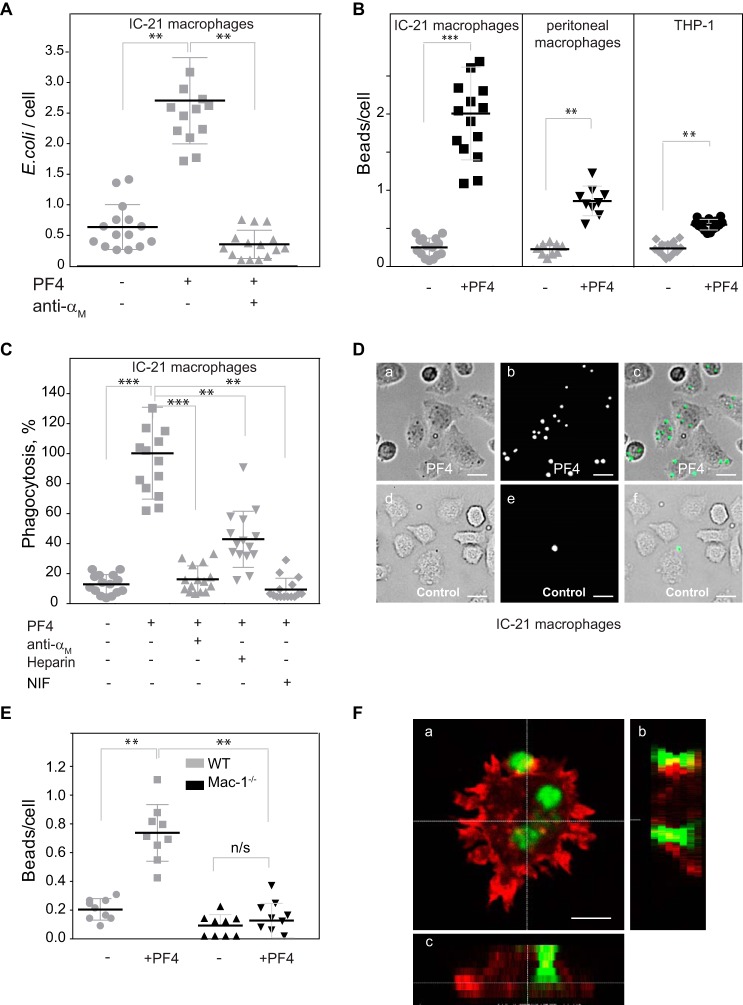
Previous studies demonstrated that PF4 was capable of binding to both Gram-positive and Gram-negative bacteria (32). We have recently shown that selected Mac-1 ligands can augment phagocytosis of bacteria and plastic beads in a Mac-1-dependent manner (18, 19). Moreover, this effect was exerted through the opsonic activity of proteins. We hypothesized that PF4 bound to the surface of bacteria may bind Mac-1 on macrophages followed by their phagocytosis. To examine this possibility, we compared control and PF4-treated Escherichia coli in a phagocytosis assay. Fluorescently-labeled bacteria were incubated with 40 μg/ml of PF4 and then unbound protein was washed by centrifugation. Adherent IC-21 murine macrophages were then incubated with either untreated or PF4-treated E. coli and their phagocytosis was determined as described (19). As shown in Fig. 7A, PF4 augmented bacteria uptake by 4.5 ± 0.2-fold. Preincubation of IC-21 cells with 20 μg/ml of M1/70 directed against murine αM integrin subunit efficiently inhibited phagocytosis of PF4-treated E. coli by IC-21 macrophages. In addition, using various leukocytes, including IC-21 macrophages, mouse peritoneal macrophages and differentiated human THP-1 macrophages, we also examined phagocytosis of plastic beads, a well-established system for phagocytosis of foreign bodies. All cells types phagocytosed fluorescence beads more efficiently when the beads were coated with PF4 (Fig. 7, B–D). Quantification of phagocytosed beads indicated that phagocytosis was increased by 10-, 4-, and 2-fold by IC-21, peritoneal, and THP-1 macrophages, respectively.
Figure 7.
Effect of PF4 on phagocytosis of bacteria and latex beads by various macrophages. A, fluorescently labeled E. coli were incubated with PF4 (40 μg/ml) for 30 min at 37 °C. Soluble peptide was removed by centrifugation and PF4-coated bacteria were subsequently incubated with adherent IC-21 mouse macrophages for 60 min at 37 °C. Part of IC-21 macrophages were preincubated with anti-αM mAb M1/70 (20 μg/ml) for 15 min before adhesion. Phagocytosis was determined as described under “Experimental Procedures.” Data are expressed as mean ratios of bacteria per macrophage ± S.E. **, p ≤ 0.01. B, fluorescent latex beads (2.5 × 107/ml) were preincubated with PF4 (40 μg/ml) for 30 min at 37 °C. Soluble PF4 was removed from beads by high-speed centrifugation. PF4-coated latex beads were incubated with adherent IC-21 mouse macrophages, mouse peritoneal macrophages, or differentiated THP-1 human macrophages for 60 min at 37 °C. C, adherent IC-21 macrophages (106/ml) were preincubated with anti-αM mAb M1/70 (20 μg/ml), heparin (20 μg/ml), or NIF (2 μg/ml) for 20 min at 22 °C. PF4-coated latex beads were incubated with cells for 60 min at 37 °C, and nonphagocytosed beads were removed and phagocytosis was measured. Phagocytosis of PF4-coated latex beads in the absence of Mac-1 inhibitors and heparin was assigned a value of 100%. Data shown are mean ± S.E. of five random fields determined for each condition and are representative of 3 separate experiments. **, p ≤ 0.01; ***, p ≤ 0.001. D, a representative experiment showing fluorescence of IC-21 macrophages exposed to PF4-coated latex beads. Shown are bright field (a and d), fluorescence (b and e), and merge (c and f) images of IC-21 macrophages incubated with PF4-coated (a–c) or uncoated (d–f) control beads. The scale bar is 15 μm. E, PF4-coated beads were incubated with adherent mouse peritoneal macrophages isolated from WT and Mac-1−/− mice for 30 min at 37 °C. Nonphagocytosed beads were removed, and the ratio of beads per macrophage was quantified from three fields of fluorescent images. Data shown are mean ± S.E. of triplicate measurements from three experiments. **, p ≤ 0.01 compared with untreated control beads. F, representative confocal image showing phagocytosed beads inside a macrophage. Horizontal (b) and vertical (c) cross-sections of the macrophage were taken at the positions shown by white lines. The scale bar is 5 μm.
To investigate whether integrin Mac-1 and HSPGs on macrophages are involved in promoting phagocytosis of PF4-coated beads, we examined the effects of anti-Mac-1 reagents and heparin on phagocytosis of beads. Both mAb M1/70 and NIF, which binds directly to the αMI-domain of Mac-1 inhibited PF4-mediated phagocytosis by 91 ± 7 and 75 ± 6%, respectively (Fig. 7C). Heparin also reduced the number of phagocytized beads; however, in contrast to anti-Mac-1 reagents its potency was less (56 ± 14%).
Further evidence supporting the role of Mac-1 in PF4-induced phagocytosis was obtained using macrophages isolated from the peritoneum of WT and Mac-1−/− mice. Control and PF4-coated fluorescent beads were added to adherent WT and Mac-1-deficient macrophages and their rates of phagocytosis were determined. Pretreatment of beads with PF4 enhanced their uptake by WT macrophages by ∼4-fold (Fig. 7, E and F). In agreement with previous data that phagocytosis of Mac-1-deficient neutrophils is impaired (33), phagocytosis of control beads by Mac-1-deficient macrophages was strongly reduced (Fig. 7E). Phagocytosis of PF4-coated beads by Mac-1-deficient macrophages was also increased; however, PF4 failed to enhance phagocytosis to the level observed with WT macrophages. These data suggest that on the surface of macrophages, Mac-1 is the major receptor responsible for the opsonic function of PF4.
Discussion
PF4 is an abundant 7.8-kDa basic heparin-binding protein released from platelet α-granules upon their activation. Although PF4 has been assigned to the CXC chemokine subfamily based on its structure, its chemotactic activity for neutrophils and monocytes has remained controversial and it was concluded that PF4 does not behave as a classic chemokine (4, 6). Moreover, to date the receptor on leukocytes mediating PF4 responses has not been identified.
In the present study we identified leukocyte integrin Mac-1 as a receptor for PF4. In support of this finding, we showed that PF4 supports a potent migratory response in human neutrophils, mouse macrophages, and Mac-1-expressing HEK293 cells, which are entirely dependent on Mac-1. Consistent with the cell migration results, PF4 supported efficient adhesion of Mac-1-expressing cells. In addition, PF4 enhanced phagocytosis of E. coli and latex beads by macrophages in a Mac-1-dependent manner. When added to adherent Mac-1-expressing cells, soluble PF4 induced clustering of Mac-1 in Mac-1-expressing HEK293 cells and up-regulation of Mac-1 on the surface of neutrophils and monocytes. The latter data are consistent with a previous report showing that soluble PF4 added to suspended neutrophils induced expression of Mac-1 (34). Finally, two αMI-domain–binding sites, whose sequences conform to recognition specificity of Mac-1 toward cationic proteins (12) have been identified in PF4. These observations establish Mac-1 as a receptor for PF4 and suggest a role for the Mac-1–PF4 interactions in the induction of inflammatory responses during activation of platelets.
Although PF4 was the first chemokine discovered, its ability to induce the chemotactic response was not strictly established. While several earlier studies showed that platelet-derived PF4 induced migration of neutrophils and monocytes (8, 35, 36), some reports did not observe the chemotactic activity (3, 37–39). It has been debated (38) that the chemotactic effect of PF4 might have potentially been due to contamination of preparations with NAP-2. However, the fact that the synthetic peptide duplicating the C-terminal part of PF4 (residues 57–70) had chemotactic activity for monocytes (31) argued in favor of the idea that PF4 itself may mediate cell migration. Our data clearly demonstrate that purified recombinant PF4 depleted of endotoxin promotes leukocyte migration and does so by binding to integrin Mac-1, whose ability to support cell migration to various ligands has been reported in numerous studies (18, 19, 40–42). Interestingly, one of the two αMI-domain binding sites for PF4 encompasses the C-terminal segment 57APLYKKIIKKLLES70 (Figs. 1 and 4). It appears likely that the lack of chemotactic activity observed in some previous studies could have been due to the low concentrations of PF4 used in migration assays because PF4 was expected to act in the nanomolar range, typical of classical chemokines. In line with previous investigations (8, 35, 36) the migratory activity of PF4 in our experiments has been observed at concentrations of 1–5 μg/ml (0.13–0.65 μm) (Fig. 5). The latter values seem to be in good accordance with the Kd determined for the interaction of PF4 with the αMI-domain (∼1.3 μm) and also agree with the concentrations of PF4 released at sites of injury (≥2 μm). Although PF4 triggers leukocyte migration in vitro, the biological meaning of this capacity is unclear. Activated platelets release NAP-2, a classic chemokine that exerts its chemotactic response in the expected nanomolar range in vitro and has been shown to be important in migration of leukocytes at sites of vascular injury (43). Nevertheless, blockade of CXCR1/2 and NAP-2 deficiency in the mouse models of endothelial injury that generated platelet-rich thrombi was associated with a partial decrease in leukocyte migration to thrombi (43), suggesting the role of other attractants. Interestingly, Mac-1 deficiency also strongly impaired leukocyte migration through thrombi, implicating a Mac-1-dependent response, which potentially might have been mediated by PF4.
Previous attempts to identify the PF4 receptor showed that a significant portion of PF4's binding to neutrophils involved its interaction with cell-surface glycosaminoglycans, in particular CSPGs (29, 30). Our data demonstrating that the inhibitory effect of anti-Mac-1 function blocking mAb on adhesion of neutrophils to immobilized PF4 was potentiated by CSA and CSB, but not by heparin, suggest that CSPGs cooperate with Mac-1 in PF4 binding (Fig. 3D). Nevertheless, heparin, which has a high affinity for PF4 (30, 44) partially blocked adhesion of Mac-1-expressing HEK293 cells to PF4 and the inhibitory effect of anti-Mac-1 function blocking mAb was enhanced by heparin (Fig. 3C). Similar effects of heparin and anti-Mac-1 function blocking reagents on adhesion of Mac-1–HEK293 cells as well as monocytes have been documented for other Mac-1 ligands (18, 19, 27), including a multifunctional cationic glycosaminoglycan-binding cytokine and growth factor pleiotrophin (28). These data suggest that on the surface of cells that express HS glycosmanoglycans, HSPGs may serve as co-receptors with Mac-1 to mediate PF4 binding. Because PF4 appears to exist at equilibrium among monomers, dimers and tetramers (45) and each monomeric protein contains two αMI-domain-binding sites (Figs. 1 and 4), PF4 can potentially bridge Mac-1 and proteoglycans into large multimolecular clusters.
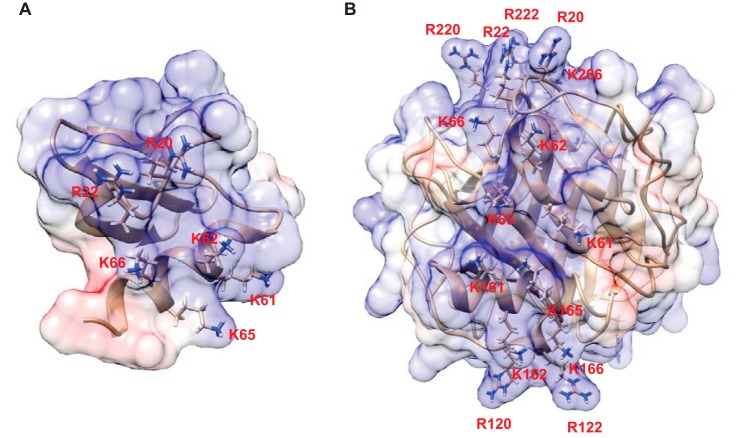
The αMI-domain binding sequences in PF4, 12CVKTTSQVRPRHITS26 and 57APLYKKIIKKLLES70, identified through screening of peptide libraries (Fig. 1) and mutational analyses (Fig. 4) conform to the expected recognition specificity of Mac-1. In particular, we recently showed that within its numerous ligands Mac-1 has a preference for the short sequences enriched in positively charged residues flanked by hydrophobic residues, in which basic residues are main contributors to interaction with the αMI-domain (12, 21). As shown in Fig. 8, the very same segments in PF4 that bind αMI-domain contain the residues that are important in heparin binding, namely arginines 20 and 22 in Cys12–Ser26 and four lysines in the C-terminal sequence Ala57–Ser70 (residues 61, 62, 65, and 66) (46, 47). As shown in Fig. 1D both αMI-domain binding sequences are situated in close proximity in the PF4 monomer forming a basic surface. In the asymmetric PF4 tetramer, these sequences in the AC and BD dimers form even larger basic surfaces and are displayed on the opposite sides of the complex, providing potential sites for the interactions with both αMI-domain and GAGs (Fig. 8B). At present, the complex relationships between PF4, Mac-1, and cell-surface proteoglycans are poorly understood and would require further analyses, but the property of PF4 to form dimers and tetramers with a unique surface chemistry may play a critical role.
Figure 8.
Basic residues in PF4 important for the αMI-domain and heparin binding. A and B, electrostatic potential surface representation (positive, blue; negative, red) of the PF4 monomer (A) and tetramer (B) based on PDB code 1F9Q (45). Basic residues Arg20, Arg22 in the 12Cys–Ser26 segment and Lys61, Lys62, Lys65, and Lys66 in the 57Ala–Ser70 segment in the PF4 monomer (A) and PF4 tetramer (B) that have been identified as critical for αMI-domain binding are shown. The same residues have been demonstrated to be important for heparin binding (45).
We now show that PF4 significantly augments phagocytosis of bacteria and latex beads by human and mouse macrophages acting via Mac-1, a well-known phagocytic receptor. Previous studies documented the ability of PF4 to bind to bacteria (32, 48). Therefore, we tested the idea that PF4 may serve as an opsonin in Mac-1-mediated phagocytosis. Indeed, pretreatment of E. coli and latex beads with PF4 enhanced phagocytosis by macrophages by severalfold (Fig. 7). Because the process was completely abrogated by anti-Mac-1 function blocking reagents, it appears that Mac-1 serves as a receptor for PF4 on the bacterial surface even without cooperation with HSPGs. The ability of PF4 to promote phagocytosis, in conjunction with its ability to induce generation of oxygen radicals in phagocytes (39) lends further support to the idea that PF4 acts as a host defense protein involved in the first line of defense against bacterial infection at the sites of vascular injury. Although PF4 may promote phagocytosis either by activating macrophages (39) or directly opsonizing bacteria (Fig. 7) the fact that it is released from platelets in extraordinarily high amounts seems to support its role as an opsonin.
The numerous immune-modulating effects induced by PF4 are reminiscent of those of several cationic host defense peptides/proteins released from neutrophils and other cells. This group includes the human cathelicidin LL-37, defensins, eosinophil-derived neurotoxin, and others. These molecules, which in addition to their primary anti-microbial function also induce numerous immune-modulating responses, have been collectively termed alarmins, referring to their ability to serve as alarm/danger signals for the immune system (16). Such molecules are sequestered within neutrophil granules and released during the immune-inflammatory response. We recently showed that the established alarmin LL-37 induces leukocyte responses, including a potent augmentation of phagocytosis via Mac-1 (18). Moreover, many other cationic host defense peptides/proteins are Mac-1 ligands (19). Because the PF4 functional spectrum shares with leukocyte cationic host defense proteins/peptides common characteristics and similar to leukocyte proteins exerts its effects on myeloid cells through Mac-1, it would qualify as a host defense protein against microbial invaders and the first platelet alarmin, thus further supporting the idea of platelets as immune cells (49, 50).
Experimental procedures
Reagents
The mouse mAb 44a directed against the human αM-integrin subunit and rat mAb M1/70, which recognizes both mouse and human αM-integrin subunits were purified from conditioned media of hybridoma cells obtained from the American Tissue Culture Collection (Manassas, VA) using protein A-agarose. Alexa Fluor 546 phalloidin was from Molecular Probes (Eugene, OR). Mouse mAb IgG1 isotype control for mAb 44a was from Cell Signaling (Danvers, MA) and rat IgG2b isotype control for mAb M1/70 was from Bio-Rad. Polyclonal antibody against human PF4 was from Santa Cruz Biotechnology (Dallas, TX). Synthetic peptides corresponding to the 12CVKTTSQVRPRHITS26, 32AGPHCPTAQ40, and 57APLYKKIIKKLLES70 sequences of human PF4 were obtained from Peptide 2.0 (Chantilly, VA). NIF was a gift from Corvas International. Polyvinylpyrrolidone (PVP), heparin (sodium salt; from porcine intestinal mucosa), and chondroitin sulfate A and B were purchased from Sigma. Calcein-AM and fluorescent latex beads (FluoroSpheres, 1 μm) were from Thermo.
Cells
Human embryonic kidney cells (HEK293) and HEK293 cells stably expressing integrin Mac-1 were previously described (11, 23). The U937 monocytic cells and IC-21 murine macrophages were grown in RPMI containing 10% FBS and antibiotics. The THP-1 cells were cultured in RPMI containing 10% FBS, antibiotics, and 0.05 mm 2-mercaptoethanol. The THP-1 were differentiated by adding 10 ng/ml of PMA into the medium for 48 h. Human neutrophils were isolated under sterile conditions from peripheral blood obtained from healthy volunteers as described (51). Human monocytes were isolated from peripheral blood obtained from healthy volunteers using the EasySep Human monocyte isolation kit (StemCell Technologies, Cambridge, MA) according to the manufacturer's protocol. The studies were approved by the Institutional Review Board of Arizona State University and performed in accordance with the Declaration of Helsinki. E. coli MG-1655 (ATCC 700926) were from ATCC (Manassas, VA). Thioglycollate-elecited peritoneal macrophages were obtained from 8-week-old WT C57BL/6 and Mac-1-deficient mice (The Jackson Laboratories, Bar Harbor, ME) by lavage using cold PBS containing 5 mm EDTA as described (52).
Mice
C57BL/6J and Mac-1−/− (B6.129S4-Itgamtm1Myd/J) mice were obtained from The Jackson Laboratory. All procedures were performed in accordance with the animal protocols approved by the Institutional Animal Care and Use Committee at the Arizona State University. Mac-1−/− mice were housed in irradiated cages and all animals were maintained under constant temperature (22 °C) and humidity, on a 12-h light/dark cycle in the Animal Facility of Arizona State University. Animal experiments were performed using both genders and appropriate age-matching controls.
Expression of recombinant proteins
Full-length PF4 cDNA (residues 1–70) was amplified from plasmid PF4 (RC211322) (Origen) and was cloned in the expression vector pGEX-4T-1 (GE Healthcare). Recombinant PF4 mutants were prepared by site-directed mutagenesis. The full-length and mutant PF4 proteins were expressed as fusion proteins with glutathione S-transferase and purified from soluble fractions of E. coli lysates by affinity chromatography. Glutathione S-transferase-tagged PF4 bound to the column was cleaved with thrombin and eluted. All recombinant proteins were monomeric as determined by size-exclusion chromatography. PF4 obtained from ERL (South Bend, IN) was used as a control. To obtain endotoxin-free proteins High Capacity Endotoxin Removed Spin columns (Pierce) were used.
The active conformer of the αMI-domain (residues αM Glu123–Lys315) was prepared as described previously (21) and labeled with 125iodine using IODO-GEN (Pierce, Rockford, IL). Nonactive αMI-domain (residues αM Gln119–Glu333) and active αLI-domain (Gly127–Tyr307) were isolated as described (53).
Synthesis of cellulose-bound peptide libraries
The PF4-derived peptide library assembled on a single cellulose membrane support was prepared by parallel spot synthesis and the membrane-bound peptides were tested for their ability to bind the αMI-domain according to a previously described procedure (21, 54).
Detection measurement of direct protein interaction by biolayer interferometry
Biolayer interferometry experiments were performed using an Octet K2 instrument (fortéBIO, Pall Corporation). Purified PF4 was immobilized on the Amine Reactive Second-generation (AR2G) biosensor using the amine coupling kit according to the manufacturer's protocol. Different concentrations of αM- and αLI-domains were applied in the mobile phase and the association between the immobilized and flowing proteins was detected. Experiments were performed in 20 mm HEPES, 150 mm NaCl, 0.05% (v/v) Tween 20, pH 7.5 containing 1 mm MgCl2 or 5 mm EDTA. The PF4-coated surface was regenerated with 25 mm NaOH. The analysis of the binding kinetics was performed using ForteBio Data Analysis 9.0 software. The dissociation rate constant (Kd) was obtained by curve fitting of the association and dissociation phases of sensograms using heterogeneous ligand model.
Cell adhesion assays
Adhesion assays were performed essentially as described previously (11, 41). Briefly, the wells of 96-well microtiter plates (Immulon 4HBX, Thermo Labsystems, Franklin, MA) were coated with various concentrations of WT or mutant PF4, or PF4-derived peptides for 3 h at 37 °C and post-coated with 1.0% PVP for 1 h at 37 °C. Cells were labeled with 5 μm calcein for 30 min at 37 °C and washed twice with Hanks' balanced salt solution containing 0.1% BSA. Aliquots (100 μl) of labeled cells (5 × 105/ml) were added to each well and allowed to adhere for 30 min at 37 °C. The nonadherent cells were removed by two washes with PBS and fluorescence was measured in a fluorescence plate reader. The number of adherent cells was determined by using the fluorescence of 100-μl aliquots with a known number of labeled cells.
Flow cytometry
Freshly isolated human neutrophils were incubated with or without NIF (1 μg/ml) for 15 min at 22 °C. Following this, PF4 (20 μg/ml) was added and cells were incubated for an additional 1 h. After the removal of non-bound PF4 by washing, cells were fixed using 2% paraformaldehyde. Fixed cells were incubated with polyclonal rabbit anti-human PF4 antibody followed by goat anti-rabbit Alexa 488-conjugated secondary antibody and analyzed using a FACSCelesta instrument (BD Biosciences, San Jose, CA).
Transwell migration assays
Transwell migration assays with WT HEK293, Mac-1-expressing HEK293 cells, human neutrophils, and purified murine macrophages using Transwell inserts (8 μm pore size for HEK293 cells, 3 μm for neutrophils, and 5 μm for macrophages) were performed as previously described (19, 40).
Phagocytosis assays
Phagocytosis assays were performed as described (18). Briefly, FITC-labeled E. coli (100 μl, 3 × 108/ml) were incubated with PF4 (40 μg/ml) for 30 min at 37 °C and washed with Dulbecco's modified Eagle's medium (DMEM) by centrifugation at 1800 × g for 5 min to remove unbound protein. The pellet was re-suspended in DMEM + 10% FBS at the concentration of 107 bacterial particles/ml. For control experiments, media was substituted for PF4, but all other aspects of the procedure were the same. IC-21 macrophages and THP-1 cells were re-suspended in DMEM + 10% FBS and cultured in Costar 48-well plates (2.5 × 105/well) for 3–5 h at 37 °C. After media was aspirated, adherent cells were washed and incubated with 0.5 ml of FITC-labeled E. coli suspensions, for 1 h at 37 °C. Cells were washed with 3 × 1 ml of PBS and phagocytosed bacteria were counted in the presence of trypan blue to quench the fluorescence of any remaining bacteria outside of macrophages. The ratio of bacterial particles per macrophage was quantified taking photographs of five fields for each well using a Leica DM4000 B microscope (Leica Microsystems, Buffalo Grove, IL) with a ×20 objective. For selected experiments, fluorescent 1.0-μm latex beads were incubated with PF4, washed, and applied at 2.5 × 106 to wells containing adherent macrophages.
In studies with macrophages isolated from the peritoneum of WT and Mac-1−/− mice, the cells were allowed to adhere to glass cover slides for 2 h at 37 °C. After removing non-adherent cells, fluorescent latex beads treated with PF4 were added to the cells and incubated for 30 min at 37 °C. Cells were washed with PBS, fixed with 2% paraformaldehyde, and beads were counted. Animal studies were carried out in strict accordance with the recommendations in the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health.
Immunofluorescence
Mac-1 HEK293 cells, neutrophils, and monocytes were allowed to adhere on cover glass for 1 h. After adhesion cells were incubated with PF4 (100 μg/ml) for 30 min at 37 °C. Cells were fixed with 2% paraformaldehyde, permeabilized with 0.1% Triton X-100, and incubated with mAb M1/70 (10 μg/ml). Cells were washed and incubated with Alexa 488-conjugated secondary antibody. In addition cells were stained with Alexa Fluor 546 phalloidin and 4′,6-diamidino-2-phenylindole. Confocal images were obtained using a Leica TCS SP5 AOBS Spectral Confocal System with ×63/1.4 objective and Leica SP8 White Light Laser Confocal/Lightsheet inverted microscope with ×20/0.75 objective and ×40/1.3 objective (Exton, PA).
Statistical analysis
All data are presented as the mean ± S.E. The statistical differences between two groups were determined using a Student's t test and between three or more groups using one-way analysis of variance with SigmaPlot 11.0 software (Systat Software, San Jose, CA) and Prism software (GraphPad, La Jolla, Ca). Differences were considered significant at p < 0.05.
Author contributions
V. K. L., V. P. Y., T. P. U., and N. P. P. formal analysis; V. K. L., V. P. Y., and N. P. P. investigation; V. K. L. methodology; V. K. L., V. P. Y., T. P. U., and N. P. P. writing-review and editing; V. P. Y. resources; V. P. Y. and T. P. U. funding acquisition; T. P. U. and N. P. P. writing-original draft; N. P. P. conceptualization; N. P. P. data curation; N. P. P. visualization.
Supplementary Material
Acknowledgments
We thank Dr. Xu Wang and Di Shen from the Department of Chemistry and Biochemistry, Arizona State University, for help with analyses of proteins structures.
This work was supported by National Institutes of Health Grants HL63199 (to T. P. U.) and DK102020 (to V. P. Y.). The authors declare that they have no conflicts of interest with the contents of this article. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
This article contains Figs. S1–S4.
- PF4
- platelet factor 4
- HSPG
- heparan sulfate proteoglycan
- CS
- chondroitin sulfate
- CSPG
- CS proteoglycan
- NIF
- neutrophil inhibitory factor
- PVP
- polyvinylpyrrolidone
- DMEM
- Dulbecco's modified Eagle's medium.
References
- 1. Afshar-Kharghan V., and Thiagarajan P. (2006) Leukocyte adhesion and thrombosis. Curr. Opin. Hematol. 13, 34–39 10.1097/01.moh.0000190107.54790.de [DOI] [PubMed] [Google Scholar]
- 2. Hagberg I. A., Roald H. E., and Lyberg T. (1998) Adhesion of leukocytes to growing arterial thrombi. Thromb. Haemost. 80, 852–858 [PubMed] [Google Scholar]
- 3. Walz A., Dewald B., von Tscharner V., and Baggiolini M. (1989) Effects of the neutrophil-activating peptide NAP-2, platelet basic protein, connective tissue-activating peptide III and platelet factor 4 on human neutrophils. J. Exp. Med. 170, 1745–1750 10.1084/jem.170.5.1745 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Brandt E., Petersen F., Ludwig A., Ehlert J. E., Bock L., and Flad H. D. (2000) The β-thromboglobulins and platelet factor 4: blood platelet-derived CXC chemokines with divergent roles in early neutrophil regulation. J. Leukoc. Biol. 67, 471–478 10.1002/jlb.67.4.471 [DOI] [PubMed] [Google Scholar]
- 5. Blair P., and Flaumenhaft R. (2009) Platelet α-granules: basic biology and clinical correlates. Blood Rev. 23, 177–189 10.1016/j.blre.2009.04.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Kowalska M. A., Rauova L., and Poncz M. (2010) Role of the platelet chemokine platelet factor 4 (PF4) in hemostasis and thrombosis. Thromb. Res. 125, 292–296 10.1016/j.thromres.2009.11.023 [DOI] [PubMed] [Google Scholar]
- 7. Nurden A. T. (2011) Platelets, inflammation and tissue regeneration. Thromb. Haemost. 105, S13–S33 10.1160/THS10-11-0720 [DOI] [PubMed] [Google Scholar]
- 8. Deuel T. F., Senior R. M., Chang D., Griffin G. L., Heinrikson R. L., and Kaiser E. T. (1981) Platelet factor 4 is chemotactic for neutrophils and monocytes. Proc. Natl. Acad. Sci. U.S.A. 78, 4584–4587 10.1073/pnas.78.7.4584 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Ludwig A., Petersen F., Zahn S., Götze O., Schröder J. M., Flad H. D., and Brandt E. (1997) The CXC-chemokine neutrophil-activating peptide-2 induces two distinct optima of neutrophil chemotaxis by differential interaction with interleukin-8 receptors CXCR-1 and CXCR-2. Blood 90, 4588–4597 [PubMed] [Google Scholar]
- 10. Petersen F., Flad H. D., and Brandt E. (1994) Neutrophil-activating peptides NAP-2 and IL-8 bind to the same sites on neutrophils but interact in different ways: discrepancies in binding affinities, receptor densities, and biologic effects. J. Immunol. 152, 2467–2478 [PubMed] [Google Scholar]
- 11. Yakubenko V. P., Lishko V. K., Lam S. C., and Ugarova T. P. (2002) A molecular basis for integrin αMβ2 ligand binding promiscuity. J. Biol. Chem. 277, 48635–48642 10.1074/jbc.M208877200 [DOI] [PubMed] [Google Scholar]
- 12. Podolnikova N. P., Podolnikov A. V., Haas T. A., Lishko V. K., and Ugarova T. P. (2015) Ligand recognition specificity of leukocyte integrin αMβ2 (Mac-1, CD11b/CD18) and its functional consequences. Biochemistry 54, 1408–1420 10.1021/bi5013782 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Cai T.-Q., and Wright S. D. (1996) Human leukocyte elastase is an endogenous ligand for the integrin CRR3 (CD11b/CD18, Mac-1, αMβ2) and modulates polymorphonuclear leukocyte adhesion. J. Exp. Med. 184, 1213–1223 10.1084/jem.184.4.1213 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Johansson M. W., Patarroyo M., Oberg F., Siegbahn A., and Nilsson K. (1997) Myeloperoxidase mediates cell adhesion via the αMβ2 integrin (Mac-1, CD11b/CD18). J. Cell Sci. 110, 1133–1139 [DOI] [PubMed] [Google Scholar]
- 15. Soehnlein O., Xie X., Ulbrich H., Kenne E., Rotzius P., Flodgaard H., Eriksson E. E., and Lindbom L. (2005) Neutrophil-derived heparin-binding protein (HBP/CAP37) deposited on endothelium enhances monocyte arrest under flow conditions. J. Immunol. 174, 6399–6405 10.4049/jimmunol.174.10.6399 [DOI] [PubMed] [Google Scholar]
- 16. Oppenheim J. J., and Yang D. (2005) Alarmins: chemotactic activators of immune responses. Curr. Opin. Immunol. 17, 359–365 10.1016/j.coi.2005.06.002 [DOI] [PubMed] [Google Scholar]
- 17. Yang D., Postnikov Y. V., Li Y., Tewary P., de la Rosa G., Wei F., Klinman D., Gioannini T., Weiss J. P., Furusawa T., Bustin M., and Oppenheim J. J. (2012) High-mobility group nucleosome-binding protein 1 acts as an alarmin and is critical for lipopolysaccharide-induced immune responses. J. Exp. Med. 209, 157–171 10.1084/jem.20101354 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Lishko V. K., Moreno B., Podolnikova N. P., and Ugarova T. P. (2016) Identification of human cathelicidin peptide LL-37 as a ligand for macrophage integrin αMβ2 (Mac-1, CD11b/CD18) that promotes phagocytosis by opsonizing bacteria. Res. Rep. Biochem. 2016, 39–55 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Podolnikova N. P., Brothwell J. A., and Ugarova T. P. (2015) The opioid peptide dynorphin A induces leukocyte responses via integrin Mac-1 (αMβ2, CD11b/CD18). Mol. Pain. 11, 33 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. El Golli N., Issertial O., Rosa J. P., and Briquet-Laugier V. (2005) Evidence for a granule targeting sequence within platelet factor 4. J. Biol. Chem. 280, 30329–30335 10.1074/jbc.M503847200 [DOI] [PubMed] [Google Scholar]
- 21. Lishko V. K., Podolnikova N. P., Yakubenko V. P., Yakovlev S., Medved L., Yadav S. P., and Ugarova T. P. (2004) Multiple binding sites in fibrinogen for integrin αMβ2 Mac-1). J. Biol. Chem. 279, 44897–44906 10.1074/jbc.M408012200 [DOI] [PubMed] [Google Scholar]
- 22. Diamond M. S., Staunton D. E., Marlin S. D., and Springer T. A. (1991) Binding of the integrin Mac-1 (CD11b/CD18) to the third immunoglobulin-like domain of ICAM-1 (CD54) and its regulation by glycosylation. Cell. 65, 961–971 10.1016/0092-8674(91)90548-D [DOI] [PubMed] [Google Scholar]
- 23. Zhang L., and Plow E. F. (1997) Identification and reconstruction of the binding pocket within αMβ2 for a specific and high affinity ligand, NIF. J. Biol. Chem. 272, 17558–17564 10.1074/jbc.272.28.17558 [DOI] [PubMed] [Google Scholar]
- 24. Rucinski B., Niewiarowski S., James P., Walz D. A., and Budzynski A. Z. (1979) Antiheparin proteins secreted by human platelets. purification, characterization, and radioimmunoassay. Blood 53, 47–62 [PubMed] [Google Scholar]
- 25. Rauova L., Poncz M., McKenzie S. E., Reilly M. P., Arepally G., Weisel J. W., Nagaswami C., Cines D. B., and Sachais B. S. (2005) Ultralarge complexes of PF4 and heparin are central to the pathogenesis of heparin-induced thrombocytopenia. Blood 105, 131–138 10.1182/blood-2004-04-1544 [DOI] [PubMed] [Google Scholar]
- 26. Lord M. S., Cheng B., Farrugia B. L., McCarthy S., and Whitelock J. M. (2017) Platelet Factor 4 binds to vascular proteoglycans and controls both growth factor activities and platelet activation. J. Biol. Chem. 292, 4054–4063 10.1074/jbc.M116.760660 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Schober J. M., Chen N., Grzeszkiewicz T. M., Jovanovic I., Emeson E. E., Ugarova T. P., Ye R. D., Lau L. F., and Lam S. C. (2002) Identification of integrin αMβ2 as an adhesion receptor on peripheral blood monocytes for Cyr61 (CCN1) and connective tissue growth factor (CCN2): immediate-early gene products expressed in atherosclerotic lesions. Blood 99, 4457–4465 10.1182/blood.V99.12.4457 [DOI] [PubMed] [Google Scholar]
- 28. Shen D., Podolnikova N. P., Yakubenko V. P., Ardell C. L., Balabiyev A., Ugarova T. P., and Wang X. (2017) Pleiotrophin, a multifunctional cytokine and growth factor, induces leukocyte responses through the integrin Mac-1. J. Biol. Chem. 292, 18848–18861 10.1074/jbc.M116.773713 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Petersen F., Bock L., Flad H. D., and Brandt E. (1998) A chondroitin sulfate proteoglycan on human neutrophils specifically binds platelet factor 4 and is involved in cell activation. J. Immunol. 161, 4347–4355 [PubMed] [Google Scholar]
- 30. Petersen F., Brandt E., Lindahl U., and Spillmann D. (1999) Characterization of a neutrophil cell surface glycosaminoglycan that mediates binding of platelet factor 4. J. Biol. Chem. 274, 12376–12382 10.1074/jbc.274.18.12376 [DOI] [PubMed] [Google Scholar]
- 31. Osterman D. G., Griffin G. L., Senior R. M., Kaiser E. T., and Deuel T. F. (1982) The carboxyl-terminal tridecapeptide of platelet factor 4 is a potent chemotactic agent for monocytes. Biochem. Biophys. Res. Commun. 107, 130–135 10.1016/0006-291X(82)91679-5 [DOI] [PubMed] [Google Scholar]
- 32. Krauel K., Pötschke C., Weber C., Kessler W., Fürll B., Ittermann T., Maier S., Hammerschmidt S., Bröker B. M., and Greinacher A. (2011) Platelet factor 4 binds to bacteria, inducing antibodies cross-reacting with the major antigen in heparin-induced thrombocytopenia. Blood 117, 1370–1378 10.1182/blood-2010-08-301424 [DOI] [PubMed] [Google Scholar]
- 33. Coxon A., Rieu P., Barkalow F. J., Askari S., Sharpe A. H., von Andrian U. H., Arnaout M. A., and Mayadas T. N. (1996) A novel role for the β2 integrin CD11b/CD18 in neutrophil apoptosis: a homeostatic mechanism in inflammation. Immunity 5, 653–666 10.1016/S1074-7613(00)80278-2 [DOI] [PubMed] [Google Scholar]
- 34. Xiao Z., Visentin G. P., Dayananda K. M., and Neelamegham S. (2008) Immune complexes formed following the binding of anti-platelet factor 4 (CXCL4) antibodies to CXCL4 stimulate human neutrophil activation and cell adhesion. Blood 112, 1091–1100 10.1182/blood-2008-04-153288 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Bebawy S. T., Gorka J., Hyers T. M., and Webster R. O. (1986) In vitro effects of platelet factor 4 on normal human neutrophil functions. J. Leukoc. Biol. 39, 423–434 10.1002/jlb.39.4.423 [DOI] [PubMed] [Google Scholar]
- 36. Park K. S., Rifat S., Eck H., Adachi K., Surrey S., and Poncz M. (1990) Biologic and biochemic properties of recombinant platelet factor 4 demonstrate identity with the native protein. Blood 75, 1290–1295 [PubMed] [Google Scholar]
- 37. Clark-Lewis I., Dewald B., Geiser T., Moser B., and Baggiolini M. (1993) Platelet factor 4 binds to interleukin 8 receptors and activates neutrophils when its N terminus is modified with Glu-Leu-Arg. Proc. Natl. Acad. Sci. U.S.A. 90, 3574–3577 10.1073/pnas.90.8.3574 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Petersen F., Ludwig A., Flad H. D., and Brandt E. (1996) TNF-α renders human neutrophils responsive to platelet factor 4; comparison of PF-4 and IL-8 reveals different activity profiles of the two chemokines. J. Immunol. 156, 1954–1962 [PubMed] [Google Scholar]
- 39. Pervushina O., Scheuerer B., Reiling N., Behnke L., Schröder J. M., Kasper B., Brandt E., Bulfone-Paus S., and Petersen F. (2004) Platelet factor 4/CXCL4 induces phagocytosis and the generation of reactive oxygen metabolites in mononuclear phagocytes independently of Gi protein activation or intracellular calcium transients. J. Immunol. 173, 2060–2067 10.4049/jimmunol.173.3.2060 [DOI] [PubMed] [Google Scholar]
- 40. Forsyth C. B., Solovjov D. A., Ugarova T. P., and Plow E. F. (2001) Integrin αMβ2-mediated cell migration to fibrinogen and its recognition peptides. J. Exp. Med. 193, 1123–1133 10.1084/jem.193.10.1123 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Lishko V. K., Yakubenko V. P., and Ugarova T. P. (2003) The interplay between integrins αMβ2 and α5β1 during cell migration to fibronectin. Exp. Cell Res. 283, 116–126, 10.1016/S0014-4827(02)00024-1 [DOI] [PubMed] [Google Scholar]
- 42. Cao C., Lawrence D. A., Strickland D. K., and Zhang L. (2005) A specific role of integrin Mac-1 in accelerated efflux to the lymphatics. Blood 106, 3234–3241 10.1182/blood-2005-03-1288 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Ghasemzadeh M., Kaplan Z. S., Alwis I., Schoenwaelder S. M., Ashworth K. J., Westein E., Hosseini E., Salem H. H., Slattery R., McColl S. R., Hickey M. J., Ruggeri Z. M., Yuan Y., and Jackson S. P. (2013) The CXCR1/2 ligand NAP-2 promotes directed intravascular leukocyte migration through platelet thrombi. Blood 121, 4555–4566 10.1182/blood-2012-09-459636 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Handin R. I., and Cohen H. J. (1976) Purification and binding properties of human platelet factor 4. J. Biol. Chem. 251, 4273–4282 [PubMed] [Google Scholar]
- 45. Zhang X., Chen L., Bancroft D. P., Lai C. K., and Maione T. E. (1994) Crystal structure of recombinant human platelet factor 4. Biochemistry 33, 8361–8366 10.1021/bi00193a025 [DOI] [PubMed] [Google Scholar]
- 46. Mayo K. H., Ilyina E., Roongta V., Dundas M., Joseph J., Lai C. K., Maione T., and Daly T. J. (1995) Heparin binding to platelet factor-4; an NMR and site-directed mutagenesis study: arginine residues are crucial for binding. Biochem. J. 312, 357–365 10.1042/bj3120357 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Cai Z., Yarovoi S. V., Zhu Z., Rauova L., Hayes V., Lebedeva T., Liu Q., Poncz M., Arepally G., Cines D. B., and Greene M. I. (2015) Atomic description of the immune complex involved in heparin-induced thrombocytopenia. Nat. Commun. 6, 8277 10.1038/ncomms9277 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Krauel K., Weber C., Brandt S., Zahringer U., Mamat U., Greinacher A., and Hammerschmidt S. (2012) Platelet factor 4 binding to lipid A of Gram-negative bacteria exposes PF4/heparin-like epitopes. Blood 120, 3345–3352 10.1182/blood-2012-06-434985 [DOI] [PubMed] [Google Scholar]
- 49. von Hundelshausen P., and Weber C. (2007) Platelets as immune cells: bridging inflammation and cardiovascular disease. Circ. Res. 100, 27–40 10.1161/01.RES.0000252802.25497.b7 [DOI] [PubMed] [Google Scholar]
- 50. Yeaman M. R. (2010) Platelets in defense against bacterial pathogens. Cell Mol. Life Sci. 67, 525–544 10.1007/s00018-009-0210-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Lishko V. K., Yermolenko I. S., and Ugarova T. P. (2010) Plasminogen on the surface of fibrin clot prevents adhesion of leukocytes and platelets. J. Thromb. Haemost. 8, 799–807 10.1111/j.1538-7836.2010.03778.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Podolnikova N. P., Kushchayeva Y. S., Wu Y., Faust J., and Ugarova T. P. (2016) The Role of integrins αMβ2 (Mac-1, CD11b/CD18) and αDβ2 (CD11d/CD18) in macrophage fusion. Am. J. Pathol. 186, 2105–2116 10.1016/j.ajpath.2016.04.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Yakubenko V. P., Solovjov D. A., Zhang L., Yee V. C., Plow E. F., and Ugarova T. P. (2001) Identification of the binding site for fibrinogen recognition peptide γ383–395 within the αM I-domain of integrin αMβ2. J. Biol. Chem. 276, 13995–14003 10.1074/jbc.M010174200 [DOI] [PubMed] [Google Scholar]
- 54. Kramer A., and Schneider-Mergener J. (1998) Synthesis and screening of peptide libraries on continuous cellulose membrane supports. Methods Mol. Biol. 87, 25–39 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.