Abstract
Initially identified as a factor involved in tyrosine kinase receptor signaling, Grb10-interacting GYF protein 2 (GIGYF2) has later been shown to interact with the 5′ cap-binding protein 4EHP as part of a translation repression complex, and to mediate post-transcriptional repression of tethered reporter mRNAs. A current model proposes that GIGYF2 is indirectly recruited to mRNAs by specific RNA-binding proteins (RBPs) leading to translation repression through its association with 4EHP. Accordingly, we recently observed that GIGYF2 also interacts with the miRNA-induced silencing complex and probably modulates its translation repression activity. Here we have further investigated how GIGYF2 represses mRNA function. In a tethering reporter assay, we identify three independent domains of GIGYF2 with repressive activity. In this assay, GIGYF2-mediated repression is independent of 4EHP but largely dependent on the CCR4/NOT complex that GIGYF2 recruits through multiple interfaces. Importantly, we show that GIGYF2 is an RBP and identify for the first time endogenous mRNA targets that recapitulate 4EHP-independent repression. Altogether, we propose that GIGYF2 has two distinct mechanisms of repression: one depends on 4EHP binding and mainly affects translation; the other is 4EHP-independent and involves the CCR4/NOT complex and its deadenylation activity.
INTRODUCTION
Grb10-interacting GYF protein 2 (GIGYF2) was initially identified in a yeast two-hybrid screen as a protein linked to the murine insulin-like growth factor (IGF-I) receptor pathway through an interaction with growth factor receptor-bound protein (Grb)10 (1). Later studies suggested that GIGYF2 and its paralog GIGYF1 modulate IGF-I and EGF receptors signaling (1–4). GIGYF2 has a potential role in the development of Parkinson’s disease, though this is under debate due to contradictory evidence from different reports (5,6). Knockout studies in mouse support a role of GIGYF2 in the development of neurological disorders (3). Indeed, whereas homozygous animals died shortly after birth due to their inability to feed, heterozygous mice appeared to develop normally but, with age, gradually showed motor dysfunction correlated with signs of neurodegeneration and defect in IGF signaling (3). Interestingly, later studies identified GIGYF2 as part of a translation repression complex when bound to the mammalian mRNA 5′-cap binding protein 4EHP (eIF4E homologous protein) (7). In contrast to the canonical cap-binding protein eIF4E, 4EHP does not bind to eIF4G (8), a translation initiation factor that bridges the 5′-cap to the 3′ poly(A) tail of mRNAs through an interaction with the poly(A)-binding protein (PABP) (9). While the canonical eIF4E/eIF4G/PABP axis is thought to induce the formation of an mRNA closed-loop that stimulates translation (10), GIGYF2 is proposed to serve as a bridging protein between RNA-binding proteins (RBP) and 4EHP to promote translational repression through an unproductive closed-loop (7). One such RBP is tristetraprolin (TTP) that partly represses reporter target mRNAs through a direct interaction with GIGYF2 (11). We recently described a novel conditional proteomics approach that allowed us identifying GIGYF2 as associating with Argonaute (Ago)2, the core RBP involved in the miRNA-mediated silencing pathway (12). Furthermore, we showed that GIGYF2 directly but transiently interacts with the miRNA-induced silencing complex (miRISC) component GW182 (also known as TNRC6 in mammals) (12). The interaction is mediated by a conserved proline-proline-glycine-leucine (PPGL) motif within GW182 that is recognized by the GYF domain of GIGYF2. Importantly, we also showed that GIGYF2 positively regulates the activity of the miRISC at early stages of miRNA-mediated silencing (12) when translation repression is the dominant mechanism of repression (13). Further supporting a role of GIGYF2 in the post-transcriptional regulation of mRNAs, GIGYF2 was shown to both repress translation efficiency and induce decay when artificially tethered to a reporter mRNA (14). More recently, the ability of 4EHP to induce repression was shown to depend on GIGYF2 rather than on cap binding when artificially tethered to a reporter mRNA lacking a canonical poly(A) tail (15). In the same study, tethered GIGYF2 or a variant thereof that cannot bind 4EHP were both found to repress the same artificial mRNA reporter lacking a poly(A) tail. Taken together this suggests that, by contrast to 4EHP-mediated repression, which is dependent on GIGYF2, GIGYF2-mediated repression does not rely on 4EHP. However, this observation might be specific to the artificial RNA-tethering assay set up. Indeed, loss of repression in GIGYF2 knockout cells of a non-tethered reporter mRNA harboring an AU-rich element bound by TTP is only rescued by expression of wild-type GIGYF2 but not of the variant that cannot bind 4EHP (15). Moreover, in this context, the cap-binding activity of 4EHP is also necessary (15). Hence it is not clear what is the relevant physiological mechanism of GIGYF2-mediated repression. As a further complication, the studies that directly addressed the action of GIGYF2 on mRNAs were performed on artificial reporter mRNAs (14,15), sometimes lacking a canonical 3′ poly(A) tail (15). The question is thus still open if GIGYF2 represses any endogenous transcript, and if so whether it does so through its interaction with 4EHP or with other downstream factors.
We show that, when tethered to a polyadenylated reporter mRNA, GIGYF2 is a repressor of mRNA function independent of the interaction with 4EHP. We mapped three main effector domains within GIGYF2 and show that the CCR4/NOT complex and its deadenylation activity strongly contribute to GIGYF2-mediated repression. We further provide evidence that GIGYF2 interacts with the CCR4/NOT complex through multiple interfaces. Importantly, we show that GIGYF2 is a genuine RBP that can directly bind to endogenous mRNAs. Finally, we identified for the first time several endogenous targets of GIGYF2-mediated silencing that are repressed in a 4EHP-independent manner. Altogether, we propose that GIGYF2 is involved in two distinct mechanisms of mRNA repression. When GIGYF2 directly binds to its targets, it may repress them by a combination of mRNA decay and translation repression independently of 4EHP but largely relying on the CCR4/NOT complex. In the second mechanism, GIGYF2 may be indirectly recruited to mRNAs through another RBP such as TTP (11) or TNRC6 within the miRISC (12), and may then mainly promote 4EHP-dependent translational repression as previously suggested (7,12,15).
MATERIALS AND METHODS
Plasmids
Plasmids for the tethering reporter assay pRL-5boxB, pGL3-FL, pCIneo-λHA-LacZ (16), pCIneo-λHA- and HA-TNRC6C (17) and a plasmid encoding for Ago2 were a kind gift from Witold Filipowicz. The plasmid encoding eGFP-GIGYF2 (18) was a kind gift of Christian Freund (Freie Universität Berlin). The plasmids encoding CNOT7, CNOT8 and CNOT9 were a kind gift of Sebastiaan Winkler (University of Nottingham), the plasmids containing CNOT1, CNOT2 and CNOT3 cDNA were a gift from Elisa Izaurralde (Addgene 37370, 37371 and 37372) and the plasmid encoding CNOT6 was a kind gift from Ann-Bin Shyu (University of Texas). Plasmids encoding dominant negative mutants of Caf1 and CCR4a in a pBEG-3xFLAG vector were obtained from Marina Chekulaeva (Max Delbrück Center for Molecular Medicine, Berlin). The plasmid encoding PABP was a kind gift of Georg Stoecklin (Universitätsmedizin Mannheim). Plasmids encoding λHA-GIGYF2, and its deletion fragments were generated by polymerase chain reaction (PCR) amplification of the corresponding fragments, digested with MfeI and NotI, and cloning into EcoRI and NotI- digested pCIneo-λHA vectors. For the generation of pCIneo-HA-LacZ and -HA-GIGYF2, the respective λHA plasmids were digested with NheI and XhoI, the overhangs filled and the plasmid re-ligated. To generate pCIneo-λHA-mut GIGYF2, previously described mutations (7) were introduced in pCIneo-λHA-GIGYF2 by site directed mutagenesis. pMIR-FL-5boxB was generated by exchange the 3′UTR of pMIR- HMGA2 3′UTR (19) with a 5boxB PCR fragment using SacI and NaeI. The plasmids encoding CNOT subunits, GIGYF2 and its fragments, PABP or Ago2 fused with a GFP tag, were cloned by PCR amplification of the corresponding open reading frames (ORF) and ligated into peGFP vector using BamHI and XhoI. Removing GIGYF2 from peGFP-GIGYF2 using BamHI and XhoI generated the plasmid peGFP-C3, then the overhangs were filled and the plasmid was re-ligated. The cDNA from 4EHP was obtained using reverse transcriptase PCR using total RNA from HeLa 11ht cells and cloned into EcoRI/NotI-digested pCIneo vector that contains an HA tag or in an SbfI/NotI-digested pBEG-3xFLAG vector. Correctness of all plasmids was verified by Sanger sequencing.
Cell culture, generation of GIGYF2 KO cells, transfections and RNAi
Hela 11ht cells, a subclonal HeLa-CCL2 cell line, stably expressing the reverse tetracycline-controlled transcription activator rtTA-M2 and containing a locus for Flp-recombinase-mediated cassette exchange (20) were grown in Dulbecco’s modified Eagle’s medium (DMEM) medium (SIGMA) supplemented with 2 mM L-glutamine (GIBCO), 10% (v/v) tet-free fetal bovine serum (FBS) (TH. Geyer) and 200 μg/ml G418 (Sigma). Cells were regularly tested for mycoplasma contamination.
DNA transfections were performed using Polyethylenimide (PEI) (Polysciences, Inc) in a ratio 1:2 (DNA:PEI). Cells had their medium changed directly before transfection. Plasmid DNA (2-3 μg for 6 well and 10–15 μg for 10 cm), PEI and serum-free DMEM were mixed, incubated for 10 min at RT and added to the cells. In tethering experiments, cells were transfected with 2 ng pRL-5boxB, 50 ng of pGL3-FL and 10 ng HA or λHA fusion constructs per well or alternatively with 25 ng pMir-FL-5boxB and 20 ng HA or λHA fusion constructs per well of a 96-well plate. When indicated 69 ng of each plasmid encoding CNOT6 and CNOT7 catalytic mutants were co-transfected. For experiments in which the same lysate was used for RNA extraction, Western blot and tethering assay, the transfection was performed either using 200 ng pMIR-FL-5boxB and 200 ng HA-λHA fusion constructs per well of a 6-well plate or 25 ng RL-5boxB, 300 ng of pGL3-FL and 300 ng HA or λHA fusion constructs. For mRNA half-lives measurements, cells were seeded in a 12-well plate, transfected with tethering reporters 24 h after seeding, and after an additional 24 h treated with 5 μg/ml Actinomycin D (Sigma) as indicated.
siRNAs and esiRNAs were previously described (12). Transfections of siRNA/esiRNA or co-transfections with DNA were performed with the jetPRIME reagent (Polyplus Transfection). siRNAs/esiRNAs were transfected at a final concentration of 50 nM following the manufacturer’s instructions. The medium of the cells was changed 24 h after transfection and the cell were lysed after 48 h.
GIGYF2 KO cell lines were generated with a CRISPR/Cas9 approach using as a previously described sgRNA, targeting GIGYF2 (15), that was cloned into the pSpCas9(BB)-2A-GFP (PX458) vector (Addgene 48138). HeLa 11ht cells were transfected with the sgRNA-Cas9 vector using Attractene (Qiagen) and GFP positive cells were selected via FACS single cell sorting (ZMBH-Flow Cytometry &FACS Core Facility). Individual clonal cell lines were screened for efficient depletion of GIGYF2 by western blot. As further validation, the Cas9-targeted genomic locus of the knock-out clonal cell lines was amplified by PCR and subcloned in the pCR-blunt plasmid (Thermo). Sanger sequencing revealed two types of Cas9-induced mutations: a 3 bp deletion leading to the removal of the initial ATG and a 1 bp deletion leading to a frame shift in the coding sequence of GIGYF2. Similar to the previously described GIGYF1/2-knockout HEK-293 cells, a residual GIGYF2 antibody signal was observed by Western blot analysis, originating from bands running faster than full length GIGYF2. These bands were suggested to represent minor translation products of the GIGYF2 mRNA using downstream AUG start codons (15). Supporting this interpretation, the bands were not observed in GIGYF2-knockout HeLa cells additionally transfected with a pool of GIGYF2-targeting siRNAs.
Luciferase assay
Cells were lysed 24 h after transfection using passive lysis buffer (Promega). In case the same lysate was used for RNA extraction, cells were lysed using a cytoplasmic RNA lysis buffer (50 mM Tris–HCl pH 7.4, 150 mM NaCl, 1 mM ethylenediaminetetraacetic acid (EDTA) pH 8, 0.5% NP-40) for 20 min at 4°C. After centrifugation at 10 000 g for 10 min at 4°C, part of the supernatant was used for a Luciferase Assay, the equivalent of 10–15 μg protein content was kept for a Western blot analysis and the rest was used for RNA extraction. Firefly (FL) and Renilla luciferase (RL) activities were measured with a Xenius XL microplate luminometer (SAFAS, Monaco) using the Dual-Luciferase Reporter Assay System (Promega). To determine the contribution of translation repression (decrease in translation efficiency) to the overall repression observed in the luciferase assay, we first calculated translation efficiencies (TE) as the ratios of protein levels (normalized luciferase values) over mRNA levels (normalized qPCR values). The decrease in translation efficiency due to the tethering of a λHA-tagged protein was thus calculated as the TE in the presence of the HA-tagged protein (control condition, set to 100%) minus the TE in the presence of the λHA-tagged protein.
Immunofluorescence
Immunofluorescence was performed using HeLa-11ht cells seeded in a 12 well removable chamber slide (Ibidi). Cells were fixed with 3.7% formaldehyde for 10 min at RT followed by three washing steps with phosphate-buffered saline (PBS), each 3 min. Permeabilization was performed with 100% ice-cold methanol for 5 min at RT, followed by three washing steps with PBS, each 3 min. Before incubation of primary antibodies for 1 h at RT, cells were blocked in 3% FBS in PBS for 30 min at RT. Cells were washed three times with PBS for 3 min each wash and then incubated with secondary antibodies for 1h at RT followed by three washing steps in PBS, each 3 min. 4′,6-diamidino-2-phenylindole (DAPI) staining (300 nM) was done for 5 min at RT and cells were afterward washed again two times with PBS prior fixation with Fluoromount-G (Southern Biotech). Pictures were taken with the 63× objective of a Nikon Ni-E widefield microscope controlled by the Velocity software at the Heidelberg Nikon imaging center (Heidelberg). Minimum and maximum displayed values can be found in Supplementary Table S1. Cutoffs values were set against cells, which had not been treated with primary antibodies.
Antibodies
For western blot analysis, antibodies directed against the following proteins were used: GFP (SC-9996, Santa Cruz, 1:2000), α-tubulin (T6074, Sigma, 1:10000), α-tubulin (Ab18251, Abcam, 1:10000), GAPDH (60004-1lg, Proteintech, 1:20000), HA (16B12, Covance, 1:1000), GIGYF2 (NBP2-12812, Novusbio, 1:3000), FLAG-tag (F1804, Sigma, 1:500), DDX6 (A300-461A, Bethyl Laboratories, 1:1000) and GIGYF1 (A304-132A-M, Bethyl Laboratories, 1:500). Secondary antibodies were: IRDye 800CW goat anti-rabbit (LICOR, 1:10000), IRDye 680RD goat anti-rabbit (LICOR, 1:10000), IRDye 800CW goat anti-mouse (LICOR, 1:10000) and Alexa 680 goat anti-mouse (A21057, Life Technologies 1:10000). For Immunoprecipitation: HA (Covance), GIGYF2 (Novusbio) and mouse or rabbit IgG (Sigma). For Immunofluorescence: GIGYF2 (NBP2-12812, Novusbio, 1:500), HA (16B12, Covance, 1:1000), anti-mouse Alexa 488 (A28175, Invitrogen, 1:1000) and anti-rabbit Alexa 633 (A21070, Invitrogen, 1:1000).
GFP pull-down and immunoprecipitation
For GFP pull-down assays, HeLa 11ht cells grown in a 10-cm dish were transfected with 10 μg of plasmid expressing eGFP-CNOT9 or eGFP-GIGYF2, or co-transfected with 7.5 μg plasmid expressing eGFP-CNOT fusion proteins and 7.5 μg plasmid expressing HA-GIGYF2 or its deletions fragments. Twenty four hours after transfection, cells were lysed in 1.4 ml lysis buffer A (50 mM Tris–HCl pH 7.4, 150 mM NaCl, 2 mM EDTA pH 8, 0.5% NP-40, 0.5 mM DTT) supplemented with Complete protease inhibitor cocktail (Roche) for 30 min at 4°C and cleared lysates were then treated with micrococcal nuclease (NEB, 14 gel Unit/μl) and 2.5 mM CaCl2 for 25 min at RT, we have tested that this treatment eliminates RNA-dependent interactions. The lysates were then incubated with 25 μl pre-equilibrated GFP-Trap magnetic beads (Chromotek) for 2–3 h at 4°C on a rotating wheel. Beads were then washed three times with the same equilibration buffer 1 (50 mM Tris pH 7.4, 300 mM NaCl, 1 mM MgCl2, 0.1% NP-40) and GFP-fusion proteins were eluted by boiling the beads at 95°C for 5 min in Laemmli sodium dodecyl sulphate (SDS) loading buffer. Sample preparation for MS was done as previously described (12).
For immunoprecipitation assays, Hela 11ht cells were grown in a 10-cm dish were transfected with 10 μg plasmid expressing HA-4EHP, HA-GIGYF2 or variants thereof (HA-IP), or with 7.5 μg plasmid for FLAG-4EHP expression and 7.5 μg plasmid coding for HA-LacZ, HA-GIGYF2 or its mutant (FLAG-IP). Twenty four hours after transfection, cells were lysed as in the GFP pulldown assay. The lysates were then incubated with 25 μl pre-equilibrated protein G magnetic beads (NEB) previously coupled with 5 μg of the relevant antibody or control IgG overnight at 4°C on a rotating wheel. Beads were then treated as for the GFP pulldown assay, using an equilibration buffer 2 (50 mM Tris pH 7.4, 300 mM NaCl, 1 mM MgCl2, 0.5% NP-40). For FLAG immunoprecipitations, bound proteins were eluted with 150 ng/μl FLAG Peptide (BACHEM) in TBS for 1 h at 4°C.
RNA immunoprecipitation
A total of 25 μl of protein G magnetic beads (NEB) were washed with equilibration buffer 2 and coupled with 5 μg of anti-GIGYF2 or rabbit IgG overnight at 4°C on a rotating wheel in equilibration buffer 2 supplemented with 35 μg Heparin, 50 μg tRNA from Escherichia coli and 50U/ml Ribolock. On the next day, two confluent 15-cm dishes of HeLa-11ht cells were used. After washing twice with ice-cold PBS, cells were scrapped in PBS and harvested by centrifugation at 300 g for 10 min at 4°C. Cells were lysed in 1.2 ml lysis buffer A, supplemented with 50U/ml Ribolock (Thermo Scientific) for 30 min on ice and the lysates cleared by centrifugation at 10 000 g × 15 min at 4°C. The cleared lysates were then divided and incubated with the coupled anti-GIGYF2 or control beads for 3 h at 4°C on a rotating wheel. The beads were then washed three times with equilibration buffer 2 supplemented with complete protease inhibitor cocktail and 50U/ml Ribolock. For elution the beads were re-suspended in Proteinase K Buffer (300 mM NaCl, 200 mM Tris–HCl pH 7.5, 25 mM EDTA pH 8, 2% SDS) with 200 μg of Proteinase K (NEB) and incubated at 65°C for 15 min.
RNA extraction, cDNA synthesis and qPCR
For RNA extraction and purification, cells or RNA-IPed material were mixed with TRI or TRI LS Reagent (Sigma) and purified on affinity columns using the Direct-Zol RNA miniprep kit (Zymo) that includes a DNase-treatment step. For cDNA synthesis, 1 μg of purified RNA for first treated with DNaseI (NEB) and then reverse-transcribed using the Transcriptor first strand cDNA synthesis kit (Roche) with oligo(dT) or random hexamers primers. For qPCR analysis, the cDNA were typically diluted in water (1:5) and used as a template for SYBR Green based qPCR using the SYBR Green qPCR Mix (Roche) and gene specific primers (See Supplementary Table S2) in a Step One Plus Real-Time PCR System (Applied Biosystems). Each reaction was performed in technical duplicates or triplicates. Relative expression levels were calculated with the ΔCt and the data was normalized to GAPDH or GUSB. Relative expression levels were calculated with the formula 2−(ΔCt), where ΔCt is Ct(RL or FL)-Ct(GAPDH) or Ct(endogenous targets)-Ct(GUSB) and Ct is the equivalent cycle number at which the chosen threshold is crossed.
RNA binding assay
The RNA binding assay was adapted from a previous report (21). HeLa 11ht cells grown in a 10 cm dish were transfected with 10 μg plasmid expressing eGFP, eGFP-Ago2, eGFP-PABP or eGFP-GIGYF2 or its truncation variants. Twenty four hours after transfection, cells were washed once with PBS and 4 ml of PBS was added to the dishes. Cells were then placed on ice and immediately irradiated with 0.15 J/cm2 UV light at 254 nm (UV Stratalinker 1800, Stratagene). Cells were scrapped, collected by centrifugation and re-suspended in 300 μl lysis buffer B (100 mM KCl, 5 mM MgCl2, 10 mM Tris pH 7.4, 1% NP-40, 1 mM DTT, 100 units/ml Ribolock, 1× protease inhibitor cocktail and 200 μM ribonucleoside vanvadyl complex (NEB)). Lysates were incubated on ice for 10 min, snap-frozen, and thawed to achieve complete lysis. Lysates were centrifuged for 10 min at 10 000 rpm at 4°C and the supernatant was dispatched into three tubes. Samples were mixed with 400 μl of dilution buffer (500 mM NaCl, 1 mM Mg2Cl, 0.05% SDS, 0.05% NP-40, 50 mM Tris–HCl pH 7.4, 100 units/ml Ribolock and 1× protease inhibitor cocktail) and 10 μl of pre-equilibrated GFP-Trap A beads (Chromotek). GFP beads were pre-equilibrated as in GFP pulldown experiments (see above). GFP-tagged proteins were immunoprecipitated for 2 h at 4° on a rotating wheel. Beads were then washed with 500 μl medium salt buffer (250 mM NaCl, 1 mM Mg2Cl, 0.025% SDS, 0.05% NP-40, 20 mM Tris–HCl pH 7.4, 50 units/ml Ribolock, 1× protease inhibitor cocktail) and incubated for 15 min at 4°C on a rotating wheel with 250 μl of blocking solution (200 mM LiCl, 20 mM Tris–HCl pH 7.5, 1 mM EDTA pH 8, 0.01% NP-40, 100 μg/ml E. Coli tRNA, 100 μg/ml bovine serum albumin, 50 units/ml Ribolock, 1× protease inhibitor cocktail). Beads were then incubated for 1 h at 4°C on a rotating wheel with 250 μl of hybridization buffer (500 mM LiCl, 20 mM Tris–HCl pH 7.4, 0.05% LiDS, 1 mM EDTA pH 8, 5 mM DTT, 0.01% NP-40, 8nM oligo(DT)25 WellRED (Sigma), 100 units/ml Ribolock and 1× protease inhibitor cocktail) and the excess of fluorescent was removed by washing once with 500 μl of washing buffer A (500 mM LiCl, 20 mM Tris–HCl 7.4, 0.01% LiDS, 0.01% NP-40, 1 mM EDTA pH 8, 5 mM DTT, 50 units/ml Ribolock, 1× protease inhibitor cocktail) and twice with 500 μl of washing buffer B (200 mM LiCl, 20 mM Tris–HCl 7.4, 0.01% LiDS, 0.01% NP-40, 1 mM EDTA pH 8, 5 mM DTT, 50 units/ml Ribolock, 1× protease inhibitor cocktail). Beads were re-suspended in 100 μl of washing buffer B and transferred to a 96-well optical plate (UV Plates, 4titude).
Fluorescence measurements
Fluorescence signals from the RNA binding assay were measured using a Spectra Max M5 plate reader (Molecular Devices), using the following parameters: eGFP: excitation 475 nm, emission 509 nm; WellRED: excitation 650 nm, emission 670 nm. The measurements were done from the bottom of the plate.
Microarray analysis
RNA extracted from control, GIGYF2-KO and GIGYF2 overexpressing HeLa cells (all from three biological replicates) was sent to the GeneCore Facility (EMBL, Heidelberg) for Affymetrix GeneChip array analysis. When comparing to GIGYF2-KO cells, control cells were HeLa-11ht cells that had been transfected with the pSpCas9(BB)-2A-GFP plasmid with no single guide RNA and then underwent the same clonal selection as the KO cells. When comparing to GIGYF2 overexpressing cells, control cells were HeLa-11ht cells that had been transfected with the parental plasmid. RNA samples (500 ng) were processed and labeled for array hybridization using the Ambion WT Expression kit (Life Technologies, catalogue number 4411974). Labeled, fragmented cDNA (Affymetrix GeneChip® WT Terminal Labeling and Controls Kit, catalog number 901524) was hybridized to Affymetrix GeneChip® Human Gene 2.0 ST arrays (catalogue number 902113) for 16 h at 45°C (at 60 rpm) (Affymetrix GeneChip® Hybridization, Wash, and Stain Kit, catalog number 900720). Arrays were washed and stained using the Affymetrix Fluidics Station 450, and scanned using the Hewlett-Packard GeneArray Scanner 3000 7G. Data analysis was performed with the Transcriptome Analysis Console (TAC) 4.0 Software (Affymetrix).
RESULTS
GIGYF2 represses mRNAs by a combination of mRNA decay and translation repression
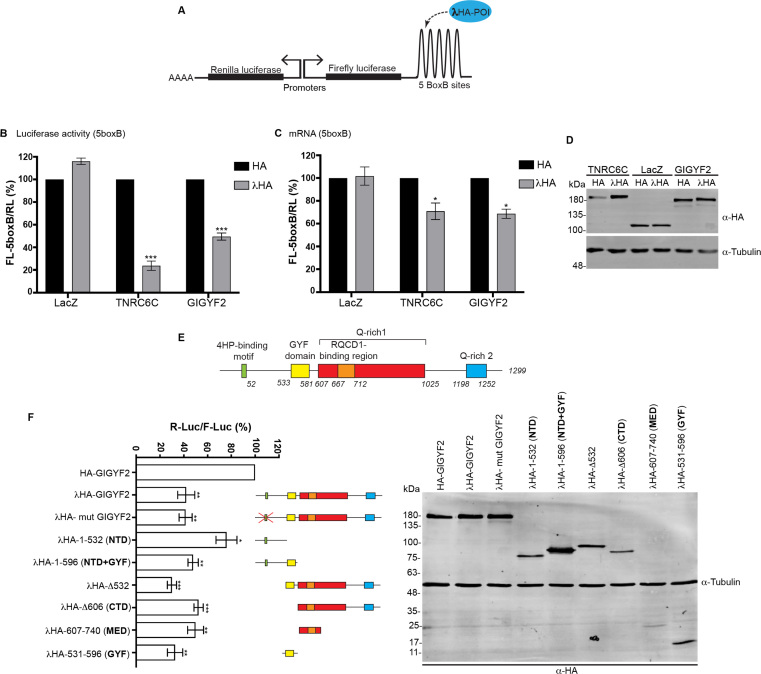
Previous data performed in HEK-293 cells showed that GIGYF2 represses mRNA function when tethered to reporter mRNAs with a canonical poly(A) tail (14) or with an internal poly(A) stretch followed by the non-polyadenylated 3′end of the MALAT non-coding RNA (15). In the former study, repression was shown to be the result of a combination of translational repression and mRNA decay (14). As we wanted to map the domains of GIGYF2 that are responsible for this activity, we first verified if GIGYF2 also represses mRNA reporters in HeLa cells, our model cell line. To this end, we used an RNA-protein tethering assay (Figure 1A). GIGYF2 fused to a λN peptide and an HA-tag (λHA-GIGYF2) was co-expressed with a thymidine kinase (TK) promoter-driven FL luciferase mRNA reporter (termed FL-5boxB) that contains in its 3′UTR five B-box hairpins, which specifically bind the λN peptide (22). As a normalization control, an RL reporter without B-box was co-expressed with the FL-5boxB reporter. As protein controls, we used a λN peptide fused to HA-tagged β-galactosidase (λHA-lacZ), which is not expected to repress FL-5boxB, and a λHA fusion to the GW182 protein TNRC6C (λHA-TNRC6C), an efficient repressor of boxB-tagged mRNA reporters in tethering assays (17). HA-tagged GIGYF2, LacZ and TNRC6C, all without λN peptide, were used as additional controls to ensure that any observation was due to the tethering of the proteins to the FL-5boxB reporter mRNA and not merely to pleiotropic overexpression effects. To calculate the repression induced by the tethered proteins, the normalized expression of the FL-5boxB reporter in the presence of the tethered λHA-tagged proteins was divided by the normalized expression of the FL-5boxB reporter in the presence of the non-tethered HA-tagged proteins that was set to 100%. As expected, λHA-TNRC6C very efficiently repressed FL-5boxB activity to 24% of control conditions (Figure 1B). Similarly, λHA-GIGYF2 also significantly repressed FL-5boxB albeit to a lesser extent than λHA-TNRC6C (to 50% of control). By contrast, tethering of λHA-LacZ did not result in repression of FL-5boxB. Importantly, similar to the TK promoter-driven FL-5boxB reporter, a cytomegalovirus promoter-driven RL-5boxB mRNA reporter (with five B-box hairpins in its 3′UTR) was also repressed by λHA-GIGYF2 but not by HA-GIGYF2 (Supplementary Figure S1). Hence, λHA-GIGYF2-mediated inhibition is not dependent on a specific reporter or promoter.
Figure 1.
GIGYF2 is a repressor of mRNA function. (A) Schematic representation of the RNA tethering assay, POI: protein of interest. (B) Normalized expression levels of the FL-5boxB reporter measured with a dual luciferase assay (protein) or (C) by RT-qPCR (mRNA). In both cases FL-5boxB readings were normalized to RL values. The normalized FL values obtained in the presence of the proteins without λ-peptide (HA-tagged (non-tethered) protein) were set to 100%. Mean values are shown with s.e.m. from six independent experiments. (D) Expression of the fusion proteins analyzed by western blotting. (E) Schematic representation of human GIGYF2. Positions of binding motif for 4EHP, GYF, Q-rich1, Q-rich2 and RQCD1-binding domains are indicated. (F) Left: Normalized expression of RL-5boxB measured with a dual luciferase assay. HeLa cells were co-transfected with plasmids expressing the indicated constructs, RL-5BoxB and FL transfection control reporters. RL was normalized to FL and values of normalized RL produced in the presence of HA-GIGYF2 were set to 100%. Mean values are shown with s.e.m. from four to six independent experiments. Right: Protein expression was analyzed by western blotting. P-values were calculated from the average of the ratios with a two-tailed Student’s t-test. *P-value < 0.05, **P < 0.01, *P-value < 0.05, ***P < 0.001.
Total RNA was extracted from the tethering assay samples to evaluate if the observed decrease of luciferase activities correlates to the expression levels of the mRNA reporters. As previously described (17), FL-5boxB mRNA levels only partially correlate with decreased luciferase activity mediated by the λHA-TNRC6C positive control (Figure 1C), confirming its well-known mode of repression mediated by a combination of mRNA decay and translational repression. Similarly, the significant decrease in mRNA levels (to 69% of control non-tethered conditions) did not match luciferase activities upon expression of λHA-GIGYF2, indicating that GIGYF2-mediated repression is also due to a combination of mRNA destabilization and translational repression. Importantly, western blot analysis revealed that all the fusion proteins were expressed at comparable levels (Figure 1D). As similar data were obtained in a study performed in HEK-293 cells (14), this suggests that the mechanism of GIGYF2-mediated repression in conserved in different human cell lines. Of note, in HeLa cells, mRNA decay and translation repression contribute to a similar extent to GIGYF2-mediated repression of the reporter mRNAs with a 31% decrease of mRNA levels and a 28% decrease in translation efficiency (see methods). This is in good agreement with the contributions observed in HEK-293 cells in which repression is due to a decrease of 40% at the transcript levels and a decrease of 42% at the translation level (estimated from Figure 4 in (14)).
Figure 4.
GIGYF2 is an RBP. (A) Schematic representation of the RNA binding assay. (B) HeLa cells transfected with the indicated constructs were irradiated with 254 nm UV light. The GFP fusions were then affinity purified from cells lysates using GFP Trap Agarose beads. Co-immunoprecipitated mRNAs were detected by hybridization with WellRED-labeled oligo(dT)25. Shown are western blot analysis of the expression of each fusion protein (left) and the corresponding red (WellRED) to green (GFP) fluorescence ratios (right), the ratio obtained for the GFP condition was set to 1. The graph was generated from three biological replicates each done in three technical replicates. (C) same as in (B) with the indicated fusion proteins. *P-value < 0.05, **P < 0.01.
GIGYF2 represses tethered mRNAs independent of 4EHP through three main distinct domains
Using λHA-tagged fusions of GIGYF2 truncations, we mapped which parts of the protein are responsible for its repressive activity (Figure 1E). Analysis of these variants in the tethering assay revealed that an N-terminal domain (aa 1 to 532, NTD) of GIGYF2, which comprises the 4EHP-binding motif, shows modest but significant repressive activity, indicating that it contributes to GIGYF2-mediated repression (Figure 1F, NTD). This domain is however dispensable for repression as a truncation of GIGYF2 lacking the first 532 amino acids shows comparable activity to the full-length protein (Figure 1F, Δ532). A recent study reported that GIGYF2 represses a tethered reporter mRNA lacking a canonical poly(A) tail independently of 4EHP binding (15). We thus verified whether a variant of GIGYF2 with a disrupted 4EHP-binding site (mut GIGYF2) also represses our mRNA reporter that harbors a canonical poly(A) tail. In agreement with previous data (7), the introduced mutations efficiently abolished binding of HA-tagged mut GIGYF2 to FLAG-tagged 4EHP in co-IP experiments (Supplementary Figure S2A). When tested in the tethering assay, λHA-tagged mut GIGYF2 was as effective as the WT protein (Figure 1F, mut GIGYF2), demonstrating that GIGYF2 repressing activity on a tethered mRNA with a canonical poly(A) tail is also independent of 4EHP. When the NTD was extended to add the GYF domain (fragment 1–596, termed NTD+GYF), repressive activity was significantly enhanced and comparable to the full-length protein (Figure 1F, NTD+GYF). A truncation lacking both NTD and GYF domains (fragment Δ606, termed CTD for C-terminal domain) also showed repressive activity comparable to the full-length protein (Figure 1F, CTD). These observations let us hypothesize that GIGYF2 possesses three repressive regions: the NTD (1–532), and to a larger extent the GYF domain and the CTD (Δ606).
To delineate more precisely the latter region, we also tested a fragment corresponding to amino acids 607–740 that comprises a previously described binding site for the protein RQCD1 (see below). This fragment displayed repressive activity comparable to the full-length protein (Figure 1F, MED) and was thus termed MED for mid effector domain. We have also tested the GYF domain (Figure 1F, GYF) in isolation and it showed repression comparable to the full-length protein. Importantly, we verified that all λHA-tagged protein fragments were expressed at comparable levels, which was the case for all except the MED, which although weakly expressed still exhibited silencing activity (Figure 1F).
To test whether some of the HA-tagged GIGYF2 fragments may have interacted with endogenous GIGYF2 possibly leading to false interpretation of the repression activity, co-immunoprecipitation experiments from HeLa cell lysates were performed. In contrast to the positive control HA-4EHP, no interaction with endogenous GIGYF2 could be detected for any of the tested GIGYF2 fragments (Supplementary Figure S2B). Altogether, the data suggest that three distinct domains of GIGYF2 support mRNA repression and that GIGYF2-mediated repression of mRNAs with a canonical poly(A) tail is independent of 4EHP.
GIGYF2-mediated repression involves the CCR4/NOT complex and its deadenylation activity
To identify potential downstream effectors of GIGYF2 that may be involved in mRNA repression, we expressed GFP-tagged GIGYF2, or GFP as a negative control, in HeLa cells and performed affinity purification on GFP-trap beads with nuclease-treated lysates to exclude RNA-mediated interactions. Co-purified proteins were then analyzed by mass spectrometry. We considered as specific interacting partners of GIGYF2 proteins that were identified by at least three unique peptides in the GFP-GIGYF2 pulldown and that were not detected at all in the control GFP pulldown. With this filter, 65 GIGYF2-interacting proteins were found (Supplementary Table S3). In addition to the known GIGYF2 interacting proteins 4EHP (gene name EIF4E2) and ZNF598 (7), numerous splicing factors specifically associated with GIGYF2. Many splicing factors harbor a PPGΦ sequence that is recognized by the GYF domain of GIGYF2, and accordingly many of them were found to associate with a recombinantly prepared GYF domain in pulldown experiments from HeLa cell lysates (18). Interestingly, endogenous DDX6 (also known as Rck/p54) a known translation inhibitor and activator of decapping (23), and CNOT1, the scaffold subunit of the CCR4/NOT complex, also specifically interacted with GFP-tagged GIGYF2. While we could map the interaction site of DDX6 to the NTD of GIGYF2 (Supplementary Figure S3), GIGYF2-mediated repression was not sensitive to DDX6 depletion in the RNA tethering assay (not shown) and thus we did not further characterize this interaction. We thus decided to focus on the CCR4/NOT complex, which is known to stimulate deadenylation and decay when recruited to mRNAs (24). In this complex, made of up at least eight CNOT subunits (25), the largest subunit CNOT1 acts as a scaffold necessary for the assembly of the whole complex whereas CNOT6 (also known as CCR4A) or its paralog CNOT6L (also known as CCR4B) and CNOT7 (also known as CAF1A) or its paralog CNOT8 (also known as CAF1B) are the two subunits with deadenylation activity (24). To assess the functional relevance of the CCR4/NOT complex for GIGYF2-mediated repression, RNA tethering reporter assays were performed with λHA-GIGYF2 under conditions in which CNOT subunits were depleted by specific siRNAs (Figure 2A). To ensure we efficiently impaired the function of the CCR4/NOT complex, the same experiments were also performed on λHA-TNRC6C, which is known to rely on an intact CCR4/NOT complex to exert mRNA repression (13,26). Previously, we have shown that a triple knock down of CNOT1, CNOT7 and CNOT8 efficiently disrupts CCR4/NOT-mediated deadenylation of mRNA reporters of miRNA targets (13). Using the same conditions, GIGYF2-mediated repression was completely alleviated in the tethering assay (Figure 2A). As expected, TNRC6C-mediated repression was also alleviated upon knock-down of CNOT1, 7 and 8 (Figure 2A). A previous study showed that CNOT7 and CNOT8 can also lead to translational repression independent of their deadenylation activity (27). To pinpoint which effect is responsible for GIGYF2-mediated silencing, we used catalytic inactive variants of the two deadenylation subunits CNOT6 (28) and CNOT7 (29). When expressed in cells, they act dominant negatively and specifically disrupt CCR4/NOT-mediated deadenylation. Under these conditions, GIGYF2-mediated repression was largely, but not completely, alleviated (Figure 2B), indicating that it relies on mRNA deadenylation to a large extent. Of note, the residual repression observed in the latter conditions (to about 80% of control, Figure 2B) might reflect a CCR4/NOT complex-mediated translation repression activity that contributes to GIGYF2-mediated repression. By contrast, TNRC6C was hardly affected by the dominant negative deadenylases (Figure 2B), reflecting its mechanism of action that involves a strong translation inhibition component in the context of tethering assays (30,31). Interestingly, GIGYF2-mediated repression was not affected by sole depletion of the CNOT1 subunit (Figure 2A). This is unlikely due to inefficient CNOT1 depletion, as repression mediated by the CNOT1-interacting protein TNRC6C was relieved under the same conditions (Figure 2A). A possible explanation for this observation may be that GIGYF2 can directly interact with the deadenylase subunits of the CCR4/NOT complex, similar to BTG/TOB proteins that can directly bind to CNOT7 or CNOT8, and were shown to be sensitive to CNOT7/8 depletion but insensitive to CNOT1 depletion in an RNA-tethering assay (32). Finally, since GIGYF2-mediated repression involves the deadenylation activity of the CCR4/NOT complex, it is expected that GIGYF2 should stimulate the decay rates of bound mRNAs. We determined boxB-tagged reporter mRNA half-lives using RT-qPCR analysis following an actinomycin D-mediated transcription arrest. As predicted, co-expression of λHA-GIGYF2 led to faster decay kinetics of the boxB-tagged reporter (Figure 2C), but not of a control mRNA (Figure 2D), as compared to conditions in which (untethered) HA-GIGYF2 was co-expressed. Altogether this suggests that GIGYF2 mainly relies on the CCR4/NOT complex and its deadenylation activity to mediate repression.
Figure 2.
GIGYF2-mediated silencing relies on the CCR4/NOT complex and its deadenylation activity. (A) Dual luciferase assay of HeLa cell lysates co-transfected with RL-5boxB, FL and λHA (dark gray bars)- or HA (black bars)-TNRC6C plasmids, or λHA (light gray bars)- or HA (white bars)-GIGYF2 plasmids. The cells were additionally co-transfected with siRNAs against CNOT1, CNOT1/7/8 or a control siRNA as indicated. (B) Same as in (A) except that in addition to the reporters and fusion proteins plasmids, the cells were co-transfected with a plasmids coding for dominant negative variants of CNOT6 and CNOT7 (CNOT6/7 mut) or with a control plasmid (mock) as indicated. In both (A) and (B), RL readings were normalized to FL and values of normalized RL produced in the presence of the HA-GIGYF2 were set to 100%. Mean values are shown with s.e.m. from four to six independent experiments. *P-value < 0.05; **P < 0.005; ***P < 0.001, n.s.: non-significant. (C). Expression levels of the RL-5BoxB mRNA reporter over time following Actinomycin D-mediated transcription arrest in the presence of λHA—or HA-GIGYF2. (D) Same as (C) assessing the p21 mRNA as a control for effectiveness of the ActD treatment. RL-5boxB or p21 levels (normalized to GAPDH) were analyzed at the indicated time points by RT-qPCR in five independent experiments and values obtained at time 0 were set to 100%. Error bars are s.e.m.
GIGYF2 interacts with the CCR4/NOT complex
In the context of mammary carcinogenesis, an interaction between GIGYF2 and the CCR4-NOT complex subunit RQCD1 (also known as CNOT9) has been described and suggested to enhance the association of GIGYF2 with EGFR via the adaptor protein Grb10 (2). The region of GIGYF2 involved in RQCD1/CNOT9 binding has been mapped to amino acids 667–712 (2) and is thus comprised within the MED (Figure 1E). To test whether this interaction is based on protein-protein interactions, GFP-tagged CNOT9 was expressed in HeLa cells and captured on GFP-trap beads following nuclease treatment. Western blot analysis revealed that endogenous GIGYF2 specifically co-purified with GFP-CNOT9 but not with the GFP control (Supplementary Figure S4A). Since we detected interaction of GFP-tagged GIGYF2 with endogenous CNOT1 by mass spectrometry, we asked if GIGYF2 interacts with the whole CCR4–NOT complex. To this end, GFP-tagged CNOT subunits were co-expressed with HA-tagged GIGYF2 in HeLa cells and, following nuclease treatment, captured on GFP-trap beads (Supplementary Figure S4B and C). HA-GIGYF2 associated with all the tested CNOT subunits, suggesting an RNA-independent protein–protein interaction with the whole complex.
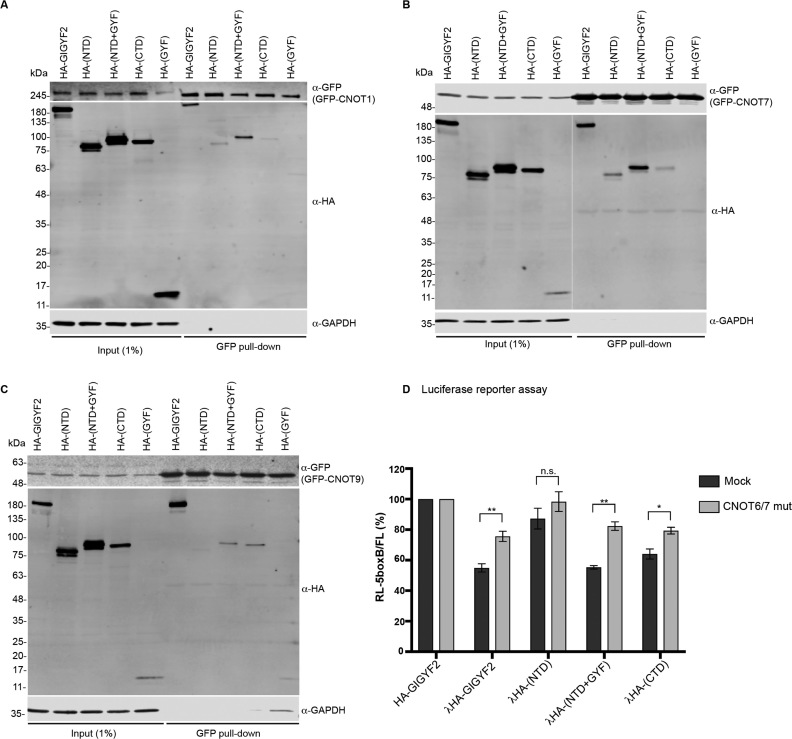
We next wondered which parts of GIGYF2 mediate an interaction with the CCR4/NOT complex. To this end, we tested four truncations, namely 1–532 (the NTD), 533–596 (corresponding to the GYF domain), 1–596 (NTD+GYF) and Δ606 (the CTD that comprises the minimal MED). The CTD was used instead of the MED itself because the latter showed much weaker expression levels. The four variants were co-expressed with GFP-CNOT1 (the scaffold subunit), -CNOT7 (a deadenylase subunit) or -CNOT9 (previously shown to bind to GIGYF2) that were affinity isolated on GFP-trap beads following cell lysis and nuclease treatment. After affinity isolation, co-isolation of NTD, NTD+GYF and CTD was detected with CNOT1 and CNOT7 (Figure 3A and B) while NTD+GYF and CTD co-purified with CNOT9 (Figure 3C). Unexpectedly, no interaction between the isolated GYF domain and the GFP-tagged CNOT subunits could be detected, although adding the GYF domain to the NTD increased binding to CNOT1 and CNOT7 (compare NTD and NTD+GYF in Figure 3A–C), and although CNOT1 harbors a PPGL motif, which is the typical signature recognized by the Smy2-type GYF domain found in GIGYF2. Previously, a recombinant isolated GYF domain of GIGYF2 was shown to associate with nuclear splicing factors in pulldown experiments from whole cell lysates (18). This was attributed to the large number of spliceosomal proteins that harbor a PPGΦ motif recognized by this domain and that may mask interactions of the GYF domain with cytoplasmic proteins (18). In light of these observations, we analyzed the subcellular localization of the GIGYF2 fragments we used in the pulldown approach by indirect immuno-fluorescence (Supplementary Figure S5). Strikingly, and by contrast to the full-length protein and all the other GIGYF2 truncations that were detected in the cytosol, the GYF domain mainly localized to the nucleus at steady-state. Hence, similar to the previous pulldown experiments performed with the recombinant protein, the numerous PPGΦ motif-containing splicing factors may preferentially interact with the isolated GYF domain in the nucleus and mask potential interactions it might have in the cytoplasm in the context of the full-length protein. As the nuclear localization of the GYF domain may complicate the interpretation of our data, its contribution to the activity of full-length GIGYF2 was assessed by comparing the NTD to the NTD+GYF construct in all further experiments.
Figure 3.
Characterization of the interaction between GIGYF2 and the CCR4/NOT complex. Blots of GFP-pull-down experiments. GFP-fusions of CNOT1 (A), CNOT7 (B) and CNOT9 (C) were co-expressed with the indicated constructs in HeLa cells. The GFP fusions were affinity purified from cells lysates using GFP-trap magnetic beads. (D) Dual luciferase assay of cells transfected with RL-5boxB, FL and the indicated fusion proteins in the presence (CNOT6/7 mut) or absence (mock) of dominant negative variants of CNOT6 and CNOT7. Mean values are shown with s.e.m. for four to six independent experiments. *P-value < 0.05; **P < 0.01; n.s., not significant.
While NTD, NTD+GYF and CTD all interacted with CNOT proteins, comparing the pulldown efficiencies for each of the tested GFP-tagged CNOT subunit suggests that CNOT1 and CNOT7 preferentially bind to the GYF domain (when appended to the NTD) while CNOT9 preferentially bind to CTD and NTD+GYF (Figure 3A–C). More quantitative statements on how the different domains recruit the CCR4/NOT complex are however not possible as the differences might reflect direct and indirect interactions between different CNOT proteins and GIGYF2 domains, and are influenced by the unknown amount of each GFP-tagged CNOT protein that assembled within the endogenous CCR4/NOT complex. Indeed, as we have mapped a binding site for DDX6 within the NTD of GIGYF2 (Supplementary Figure S3) and DDX6 is known to directly bind to CNOT1 (33,34), NTD and CCR4/NOT complex might interact through an indirect interaction mediated by DDX6. By contrast, a direct binding site of GIGYF2 for CNOT9 has been previously mapped to a region within the CTD (2). Yet, as a fraction of overexpressed GFP-CNOT9 is expected to assemble into the endogenous CCR4/NOT complex, other subunits of the complex may mediate an indirect interaction of NTD and NTD+GYF with affinity purified GFP-CNOT9.
To complement the pulldown studies, functional tethering assays were performed with the GIGYF2 fragments upon expression of the dominant negative CNOT6 and CNOT7 deadenylase variants. Similar to the full-length protein, NTD+GYF and CTD showed significantly decreased repression when CCR4/NOT complex-mediated deadenylation was disrupted (Figure 3D). Repression mediated by the NTD was also reproducibly weaker under the same conditions in every experiment we conducted, however since silencing induced by this fragment is modest, the differences were small and did not lead to statistical significance with the number of repeats we performed (Figure 3D). Altogether, the data suggest a physical and functional interaction of GIGYF2 with the CCR4/NOT complex mediated by multiple interfaces involving several domains of GIGYF2. Such a complex network of interactions is reminiscent to how Drosophila melanogaster GW182 interacts with the CCR4/NOT complex (35) and might contribute to stabilizing multiple transient interactions between individual CNOT proteins and GIGYF2.
GIGYF2 is an RNA-binding protein
GIGYF2 was identified as a candidate RBP in mRNA-interactome capture experiments performed in HEK-293 (36), Huh7 (37) and in mES (38) cell lines. However, GIGYF2 was absent from the RBP dataset obtained from HeLa cells, our model cell line, using the same approach (39). While mRNA-interactome datasets provide lists of high confidence potential RBPs, not all of these are bona fide RBPs (39,40). It is thus necessary and important to validate RBP candidates from such datasets. To test whether GIGYF2 directly binds to RNA, we used a quantitative fluorescent-based assay that was previously designed to validate direct binding to polyadenylated RNA of specific proteins in living cells (21,39). To this end, GFP-tagged GIGYF2 was expressed in HeLa cells, which were then UV-irradiated to induce crosslinking of RBPs to their target RNAs in the native cellular environment (Figure 4A). After cell lysis and affinity purification of tagged GIGYF2 on GFP-trap beads, co-purifying polyadenylated RNA was detected through hybridization with WellRED-labeled oligo-dT probes. Ratios of red over green fluorescence were then compared to GFP as a negative control and to two well-established GFP-tagged RBPs: Ago2 and PABP. Ago2 and PABP have been previously characterized through cross-linking immunoprecipitation experiments (41–44), which similar to the fluorescence-based assay also involves cross-linking of a protein of interest to endogenous RNA in living cells. As shown in Figure 4B, a clear and significant higher ratio of red over green fluorescent signal was observed for both Ago2 and GIGYF2 when compared to the control GFP sample, indicating that GIGYF2 is a bona fide RBP in HeLa cells.
We next tried to delineate which part of GIGYF2 confers RNA-binding ability to the full-length protein. The fluorescent-based RNA-binding assay was thus applied to GFP-tagged NTD, NTD+GYF and CTD and compared to GFP as a negative control as well as to PABP as a positive control. NTD and CTD both showed a significant increase of their red to green ratio, revealing an RNA-binding activity. As NTD and NTD+GYF showed similar red to green ratios (Figure 4C), the GYF domain seems not to significantly contribute to the RNA-binding activity of GIGYF2. Altogether, the data suggest the N- and C-terminal domains of GIGYF2, flanking its GYF domain, participate in direct binding to endogenous mRNAs.
Identification of endogenous targets of GIGYF2
We have shown that, when artificially brought to a reporter mRNA, GIGYF2 stimulates mRNA decay and represses translation independently of 4EHP. However, these observations may be specific to the tethering assay set up, as contradictory conclusions on to the importance of 4EHP have been previously reported when comparing tethered to non-tethered reporter mRNAs (15). More critical, whether GIGYF2 regulates the expression of endogenous transcripts is still unknown. As we found that GIGYF2 is an RBP, we next sought to identify endogenous RNA targets. We generated a GIGYF2-knockout HeLa cell line using the CRISPR/Cas9 system programmed with a single guide RNA that had been previously used to deplete GIGYF2 from HEK-293 cells (15). With this approach, we obtained cell lines in which GIGYF2 levels were considerably reduced (see ‘Materials and Methods’ section), whereas the expression levels of the GIGYF1 paralog remained unaffected (Supplementary Figure S6). To identify genes that are affected by GIGYF2 depletion, high-resolution microarrays were used to compare expression profiles from control and GIGYF2-knockout HeLa cells in three biological replicates (Figure 5A). Whereas the tethering assay suggested that recruitment of GIGYF2 to mRNA induces their destabilization, GIGYF2 depletion led to a similar number of upregulated (124) and downregulated (142) annotated transcripts by at least 1.5-fold (P < 0.05). However, the array data do not discriminate between direct and indirect targets. Hence, the downregulation of many transcripts might be an indirect consequence of the upregulation of GIGYF2 direct targets, as was previously observed upon disruption of the miRNA pathway (45). While we cannot exclude that GIGYF2 may also directly stabilize a pool of endogenous transcripts, we focused on the GIGYF2-mediated mRNA destabilization mechanism suggested by the tethering assay. To refine the set of potential targets we compared expression profiles from control and GIGYF2-overexpressing HeLa cells (Figure 5B). Indeed, it is expected that whereas direct targets of GIGYF2 should be stabilized in GIGYF2-depleted cells, they should show downregulation upon overexpression of the protein. GIGYF2 overexpression led to an up regulation of 362 and a down regulation of 224 annotated transcripts by at least 1.5-fold (P < 0.05). When comparing both microarray datasets, we found that 23 transcripts were significantly upregulated upon GIGYF2 depletion and concomitantly downregulated upon GIGYF2 overexpression (Figure 5C). Out of these 23 transcripts we picked seven for further analysis. To discriminate direct RNA targets of GIGYF2 from transcripts misregulated by secondary effects triggered by non-physiological expression levels of GIGYF2, we performed RNA-IP experiments using HeLa cells and pulling on endogenous GIGYF2, and subjected the isolated RNA to RT-qPCR analysis (Figure 5D). From the seven selected microarray hits, five transcripts (SVOPL, COL8A1, NPR3, CPM and ECEL1) showed enrichment in the GIGYF2 IP when compared to two control housekeeping mRNAs that were not identified as potential GIGYF2 targets in the array datasets (GUSB that was arbitrarily used for normalization, and PPIA). Altogether, these five mRNAs are very likely genuine direct targets of GIGYF2 as they associate with GIGYF2, are downregulated upon overexpression of the protein and upregulated upon depletion of the protein.
Figure 5.
Identification of endogenous targets of GIGYF2. (A) Volcano plot showing mRNA expression level changes upon depletion of GIGYF2 from HeLa cells analyzed with microarrays. (B) Same as in (A) but showing changes in mRNA expression levels upon overexpression of GIGYF2. In both (A) and (B) the logarithmic ratios of mRNA levels were plotted against negative logarithmic P values of a two-sided two samples t-test (n = 3 biological replicates). Red and green points are significantly (P < 0.05) upregulated and downregulated transcripts respectively. Highlighted in purple are the transcripts that are both upregulated upon GIGYF2 depletion and downregulated upon GIGYF2 overexpression. (C) Venn graph showing the transcripts that are upregulated upon GIGYF2 depletion and downregulated upon GIGYF2 overexpression, and the corresponding overlap between the two datasets. (D) Top: enrichment of selected transcripts in GIGYF2 immunoprecipitates analyzed by RT-qPCR. RNA samples were purified from input cell lysates and immuno-precipitation samples performed with an antibody against GIGYF2, or a negative control antibody (Control IgG). Transcripts levels in the IP were normalized to the input RNA fraction and enrichment in the GIGYF2 IP over the control IP was calculated. As a negative control, enrichment obtained for GUSB mRNA was set to 1 (indicated by the dotted line). Mean values are shown with s.e.m. Bottom: efficiency and specificity of the GIGYF2 immunoprecipitation was analyzed by western blotting.
Endogenous targets of GIGYF2 are repressed independently of 4EHP
Previously, a model was proposed in which GIGYF2 would act as a bridging protein between the cap binding protein 4EHP and RBPs to promote repression of specific transcripts (7). In contrast, data from the tethering assays suggest an alternative model in which GIGYF2 recruits the CCR4/NOT complex to mRNAs and promotes their repression independently of 4EHP. To test which mechanism holds true for the newly identified endogenous targets, GIGYF2-knockout cells were rescued either with wild-type GIGYF2 or with the variant of GIGYF2 (mut GIGYF2) unable to bind to 4EHP (Figure 6A) and the outcome on expression levels of the five identified target mRNAs were analyzed by RT-qPCR (Figure 6B). In both rescue conditions, we found that all five transcripts were downregulated, demonstrating that 4EHP binding to GIGYF2 is not essential for the repression of these five endogenous mRNAs.
Figure 6.
4EHP is not essential to GIGYF2-mediated repression of endogenous targets. (A) Western blot analysis of the expression levels of the indicated proteins in GIGYF2 knockout cells that were rescued with a control plasmid (mock) or plasmids encoding GIGYF2 or a variant thereof that cannot bind to 4EHP (mut GIGYF2). (B) Expression levels, normalized to GUSB mRNA, of the five validated endogenous target transcripts and a control mRNA (PPIA) analyzed by RT-qPCR from the rescued cells analyzed in (A). Log2 values obtained from mock-transfected cells were set to 0. Mean values are shown with s.e.m. from two to seven independent experiments. *P-value < 0.01; **P < 0.05; n.s., not significant. (C) A speculative model for two modes of GIGYF2-mediated repression depending on direct or indirect binding to target mRNAs.
DISCUSSION
Murine GIGYF2 was initially identified as a binding partner for the adaptor protein Grb10 that binds to phosphorylated insulin/IGF receptors and modulates their signaling (1). GIGYF2 was later shown to be part of a translation repression complex when associated with the alternative cap-binding protein 4EHP (7). Recent studies showed that GIGYF2 represses reporter mRNAs in RNA tethering assays (14,15). Yet, with contradictory data obtained from tethered or non-tethered artificial reporters it was unclear whether 4EHP is necessary for GIGYF2-mediated repression (15). More importantly, whether GIGYF2 regulates endogenous transcripts was unknown. Here we have mapped three regions of GIGYF2 that cooperate to repress reporter mRNAs and recruit the CCR4/NOT complex. Furthermore, we showed that the deadenylation activity of the CCR4/NOT complex is involved in GIGYF2-mediated repression. Finally, we found that GIGYF2 is a direct RBP and identified endogenous targets that are repressed independently of 4EHP.
GIGYF2 is an RBP
GIGYF2 was identified as a candidate RBP in mRNA interactome datasets obtained from HEK-293 (36), mES (38), HepG2 (37) but not from HeLa cells (39). As in all large-scale datasets, hits from mRNA interactomes are high-confidence candidates but not always bona fide RBPs (39,40). By using a fluorescence-based ratiometric assay (21) in HeLa cells, we identified GIGYF2 as an RBP that binds to RNA through two independent domains: NTD and CTD. Interestingly, GIGYF2 does not display any known or predicted RNA binding domain and is thus one of the so-called ‘enigmRBP’ with no clear function assigned to the RNA binding activity (37). We noted that GIGYF1, the closest homolog of GIGYF2 in mammals with ca. 50% identity, was not found in any of the mRNA interactome datasets and thus might not bind RNA. Interestingly, compared to GIGYF1, GIGYF2 contains in both RNA-binding regions many tripeptide repeats, motifs that are often enriched in RBPs (37). This may guide further studies required to more precisely delineate and confirm the RNA binding motifs of GIGYF2.
GIGYF2 is a repressor of mRNAs through the recruitment of the CCR4/NOT complex
In accordance with a previous study (14), we have shown that in tethering assays GIGYF2 represses bound mRNAs with a canonical poly(A) tail through a combination of translation repression and mRNA decay mechanisms. We mapped three distinct domains of GIGYF2 that are responsible for the repression activity: the NTD, and to a larger extent the GYF and MED domains. We deciphered the mode of action of GIGYF2-mediated repression by showing that it is independent of 4EHP binding but strongly dependent on the CCR4/NOT complex that GIGYF2 recruits through its three silencing domains. Previously we have shown that GIGYF2 interacts through its GYF domain with the miRISC component TNRC6 (12). As TNRC6 directly binds to CNOT1 and CNOT9 (33,34), TNRC6 possibly could act as a bridging protein between GIGYF2 and the CCR4/NOT complex. However, this hypothesis is not supported by our data, as GIGYF2-mediated silencing was not sensitive to CNOT1 depletion in tethering assays (Figure 2), which is in contrast to CNOT1-dependent TNRC6-induced repression. Moreover, the CTD of GIGYF2 is similarly potent in repression as the full-length GIGYF2, but unable to interact with TNRC6 as it lacks the GYF domain, again arguing against a requirement of TNRC6 in our assays.
mRNA targets of GIGYF2
Previous studies that investigated a role of GIGYF2 in repressing mRNAs focused on the characterization of reporter mRNAs to which GIGYF2 was tethered (14,15) and hence, whether GIGYF2 regulates endogenous transcripts was still unknown. Here, we identified endogenous transcripts that are associated with and regulated by GIGYF2. Our microarray profiling data allowed identifying 23 endogenous RNAs that are specifically repressed by GIGYF2, including 21 protein-coding mRNAs. Interestingly, the latter transcripts showed an overrepresentation (15 out of 21 transcripts) for mRNAs coding for transmembrane, GPI-anchored or secreted proteins and hence that are translated at the endoplasmic reticulum (ER). As GIGYF2 localizes at the ER in steady-state (18), GIGYF2-mediated repression might be spatially restricted to this organelle. In RNA-IP experiments, five candidate transcripts associated with GIGYF2, and can thus be considered as endogenous targets of the protein. It was not clear from previous data whether 4EHP is required for physiological GIGYF2-mediated repression (15). Testing the 4EHP requirement in the context of the five identified target mRNAs revealed that GIGYF2-mediated repression was independent of 4EHP, which is in agreement with our RNA-tethering assay.
The targets we identified may represent a fraction of a larger set of regulated transcripts. Indeed, with the aim to identify GIGYF2-specific targets, we have used GIGYF2-depleted cells in which the GIGYF1 paralog was expressed at wild-type levels. GIGYF1 and GIGYF2 share more than 50% amino acid identity, with most of the difference resulting from additional sequences that are only present in GIGYF2. Whereas both murine GIGYF1 and GIGYF2 are ubiquitously expressed, GIGYF2 homozygous knockout mice die shortly after birth (3), suggesting that the two proteins are not functionally redundant. Yet, both paralogs bind to 4EHP (7) and CNOT9 (2), and in addition share an almost identical GYF domain. Hence, even though it was not identified as a candidate RBP, the question whether GIGYF1 may also bind and regulate mRNAs is still open and must be addressed in the future as a potential partial functional overlap of both paralogs might have masked a larger population of GIGYF2-regulated transcripts. Moreover, in line with the RNA tethering assay that showed significant GIGYF2-stimulated mRNA decay, we have focused on mRNAs that are significantly destabilized by GIGYF2. By doing so, we may have missed potential targets that are essentially repressed at the translation level.
Multiple modes of repression for GIGYF2
GIGYF2 has been described as part of a translation inhibition complex acting through binding to the cap binding protein 4EHP (7). In this context, it was proposed that GIGYF2 serves as a bridging protein between RBPs and 4EHP resulting in a translation-incompetent cap-binding complex. Such a mechanism appears to apply to TTP, which binds 4EHP through GIGYF2 to enhance repression of its targets (11), and might also be used by the miRISC (12). By contrast, our data suggest an alternative mechanism of mRNA repression in which GIGYF2 may directly bind to its target mRNAs independently of 4EHP. In this mode of repression, GIGYF2 promotes the destabilization and translation repression of its targets through the recruitment of the CCR4/NOT complex. Importantly, we identified endogenous mRNA targets that are repressed by GIGYF2 independently of 4EHP. Whether GIGYF2 can use its two modes of repression, sequentially or simultaneously, on the same transcripts is currently unknown. However, as we have found five endogenous transcripts that are repressed by GIGYF2 independently of 4EHP, we propose that GIGYF2 may have a set of targets that is repressed through the recruitment of the CCR4/NOT complex and another set that is repressed through the GIGYF2/4EHP axis possibly mainly at the translation level (Figure 6C). This would be analogous to TRIM71 in Caenorhabditis elegans (46), that either leads to transcript degradation or translational repression of its targets depending on whether it binds within a 5′ or a 3′UTR. For GIGYF2 using one mechanism or the other might depend on how it is recruited to its target mRNAs: direct binding might lead to repression through the recruitment of the CCR4/NOT complex, whereas indirect binding mediated by an additional RBP might lead to repression through 4EHP. Identification of the RNA features recognized by GIGYF2 as well as transcriptome-wide identification of its targets coupled with ribosome profiling will help shedding light on the different repression mechanisms of GIGYF2.
DATA AVAILABILITY
The microarray data have been deposited with the GEO Data bank under accession number GSE101098.
Supplementary Material
ACKNOWLEDGEMENTS
The authors thank Dr Kai Schönig (ZI, Mannheim, Germany) for the kind gift of the HeLa-11ht cell line. Dr Marina Chekulaeva (MDC, Berlin, Germany), Dr Christian Freund (FU Berlin, Germany), Dr Elisa Izzauralde (MPI for evolution biology, Tübingen, Germany), Dr Witold Filipowicz (FMI, Basel, Switzerland), Dr Ann-Bin Shyu (University of Texas at Houston, USA), Dr Sebastiaan Winkler (University of Nottingham, UK), Dr Georg Stoecklin (Universitätsmedizin Mannheim) for kindly sharing plasmids. We also thank Anne-Marie Alleaume and Dr Matthias Hentze (EMBL, Heidelberg, Germany) for their help in establishing the RBP assay as well as Jelena Pistolic and Dr Vladimir Benes at the EMBL Genecore facility for the microarray analysis, Dr Ashish Goyal (DKFZ, Heidelberg) for his help in establishing the mRNA stability assay and for sharing primers, Dr Monika Langlotz at the Flow Cytometry & FACS Core Facility (ZMBH, Heidelberg) for performing single cell sorting, Jérôme Oswald for characterising the KO cell lines and Dr Mandy Jeske for critical comments on the manuscript.
SUPPLEMENTARY DATA
Supplementary Data are available at NAR Online.
FUNDING
Deutsche Forschungsgemeinschaft, cluster of excellence CellNetworks [DFG-EXC81 to J.B.]. Funding for open access charge: Heidelberg University Open Access Publishing Fund; Deutsche Forschungsgemeinschaft.
Conflict of interest statement. None declared.
REFERENCES
- 1. Giovannone B., Lee E., Laviola L., Giorgino F., Cleveland K.A., Smith R.J.. Two novel proteins that are linked to insulin-like growth factor (IGF-I) receptors by the Grb10 adapter and modulate IGF-I signaling. J. Biol. Chem. 2003; 278:31564–31573. [DOI] [PubMed] [Google Scholar]
- 2. Ajiro M., Nishidate T., Katagiri T., Nakamura Y.. Critical involvement of RQCD1 in the EGFR-Akt pathway in mammary carcinogenesis. Int. J. Oncol. 2010; 37:1085–1093. [DOI] [PubMed] [Google Scholar]
- 3. Giovannone B., Tsiaras W.G., de la Monte S., Klysik J., Lautier C., Karashchuk G., Goldwurm S., Smith R.J.. GIGYF2 gene disruption in mice results in neurodegeneration and altered insulin-like growth factor signaling. Hum. Mol. Genet. 2009; 18:4629–4639. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Higashi S., Iseki E., Minegishi M., Togo T., Kabuta T., Wada K.. GIGYF2 is present in endosomal compartments in the mammalian brains and enhances IGF-1-induced ERK1/2 activation. J. Neurochem. 2010; 115:423–437. [DOI] [PubMed] [Google Scholar]
- 5. Tan E.K., Schapira A.H.. Summary of GIGYF2 studies in Parkinson's disease: the burden of proof. Eur. J. Neurol. 2010; 17:175–176. [DOI] [PubMed] [Google Scholar]
- 6. Zhang Y., Sun Q.Y., Yu R.H., Guo J.F., Tang B.S., Yan X.X.. The contribution of GIGYF2 to Parkinson's disease: a meta-analysis. Neurol. Sci. 2015; 36:2073–2079. [DOI] [PubMed] [Google Scholar]
- 7. Morita M., Ler L.W., Fabian M.R., Siddiqui N., Mullin M., Henderson V.C., Alain T., Fonseca B.D., Karashchuk G., Bennett C.F. et al. A novel 4EHP-GIGYF2 translational repressor complex is essential for mammalian development. Mol. Cell. Biol. 2012; 32:3585–3593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Joshi B., Cameron A., Jagus R.. Characterization of mammalian eIF4E-family members. Eur. J. Biochem. 2004; 271:2189–2203. [DOI] [PubMed] [Google Scholar]
- 9. Imataka H., Gradi A., Sonenberg N.. A newly identified N-terminal amino acid sequence of human eIF4G binds poly(A)-binding protein and functions in poly(A)-dependent translation. EMBO J. 1998; 17:7480–7489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Kahvejian A., Roy G., Sonenberg N.. The mRNA closed-loop model: the function of PABP and PABP-interacting proteins in mRNA translation. Cold Spring Harb. Symp. Quant. Biol. 2001; 66:293–300. [DOI] [PubMed] [Google Scholar]
- 11. Fu R., Olsen M.T., Webb K., Bennett E.J., Lykke-Andersen J.. Recruitment of the 4EHP-GYF2 cap-binding complex to tetraproline motifs of tristetraprolin promotes repression and degradation of mRNAs with AU-rich elements. RNA. 2016; 22:373–382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Schopp I.M., Amaya Ramirez C.C., Debeljak J., Kreibich E., Skribbe M., Wild K., Béthune J.. Split-BioID a conditional proteomics approach to monitor the composition of spatiotemporally defined protein complexes. Nat. Commun. 2017; 8:15690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Béthune J., Artus-Revel C.G., Filipowicz W.. Kinetic analysis reveals successive steps leading to miRNA-mediated silencing in mammalian cells. EMBO Rep. 2012; 13:716–723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Kryszke M.H., Adjeriou B., Liang F., Chen H., Dautry F.. Post-transcriptional gene silencing activity of human GIGYF2. Biochem. Biophys. Res. Commun. 2016; 475:289–294. [DOI] [PubMed] [Google Scholar]
- 15. Peter D., Weber R., Sandmeir F., Wohlbold L., Helms S., Bawankar P., Valkov E., Igreja C., Izaurralde E.. GIGYF1/2 proteins use auxiliary sequences to selectively bind to 4EHP and repress target mRNA expression. Genes Dev. 2017; 31:1147–1161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Pillai R.S., Artus C.G., Filipowicz W.. Tethering of human Ago proteins to mRNA mimics the miRNA-mediated repression of protein synthesis. RNA. 2004; 10:1518–1525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Zipprich J.T., Bhattacharyya S., Mathys H., Filipowicz W.. Importance of the C-terminal domain of the human GW182 protein TNRC6C for translational repression. RNA. 2009; 15:781–793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Ash M.R., Faelber K., Kosslick D., Albert G.I., Roske Y., Kofler M., Schuemann M., Krause E., Freund C.. Conserved beta-hairpin recognition by the GYF domains of Smy2 and GIGYF2 in mRNA surveillance and vesicular transport complexes. Structure. 2010; 18:944–954. [DOI] [PubMed] [Google Scholar]
- 19. Hock J., Weinmann L., Ender C., Rudel S., Kremmer E., Raabe M., Urlaub H., Meister G.. Proteomic and functional analysis of Argonaute-containing mRNA-protein complexes in human cells. EMBO Rep. 2007; 8:1052–1060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Weidenfeld I., Gossen M., Low R., Kentner D., Berger S., Gorlich D., Bartsch D., Bujard H., Schonig K.. Inducible expression of coding and inhibitory RNAs from retargetable genomic loci. Nucleic Acids Res. 2009; 37:e50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Strein C., Alleaume A.M., Rothbauer U., Hentze M.W., Castello A.. A versatile assay for RNA-binding proteins in living cells. RNA. 2014; 20:721–731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Gehring N.H., Neu-Yilik G., Schell T., Hentze M.W., Kulozik A.E.. Y14 and hUpf3b form an NMD-activating complex. Mol. Cell. 2003; 11:939–949. [DOI] [PubMed] [Google Scholar]
- 23. Parker R., Sheth U.. P bodies and the control of mRNA translation and degradation. Mol. Cell. 2007; 25:635–646. [DOI] [PubMed] [Google Scholar]
- 24. Wahle E., Winkler G.S.. RNA decay machines: deadenylation by the Ccr4-not and Pan2-Pan3 complexes. Biochim. Biophys. Acta. 2013; 1829:561–570. [DOI] [PubMed] [Google Scholar]
- 25. Winkler G.S., Balacco D.L.. Heterogeneity and complexity within the nuclease module of the Ccr4-Not complex. Front. Genet. 2013; 4:296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Chen C.Y., Zheng D., Xia Z., Shyu A.B.. Ago-TNRC6 triggers microRNA-mediated decay by promoting two deadenylation steps. Nat. Struct. Mol. Biol. 2009; 16:1160–1166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Cooke A., Prigge A., Wickens M.. Translational repression by deadenylases. J. Biol. Chem. 2010; 285:28506–28513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Yamashita A., Chang T.C., Yamashita Y., Zhu W., Zhong Z., Chen C.Y., Shyu A.B.. Concerted action of poly(A) nucleases and decapping enzyme in mammalian mRNA turnover. Nat. Struct. Mol. Biol. 2005; 12:1054–1063. [DOI] [PubMed] [Google Scholar]
- 29. Viswanathan P., Ohn T., Chiang Y.C., Chen J., Denis C.L.. Mouse CAF1 can function as a processive deadenylase/3′-5′-exonuclease in vitro but in yeast the deadenylase function of CAF1 is not required for mRNA poly(A) removal. J. Biol. Chem. 2004; 279:23988–23995. [DOI] [PubMed] [Google Scholar]
- 30. Braun J.E., Huntzinger E., Fauser M., Izaurralde E.. GW182 proteins directly recruit cytoplasmic deadenylase complexes to miRNA targets. Mol. Cell. 2011; 44:120–133. [DOI] [PubMed] [Google Scholar]
- 31. Chekulaeva M., Mathys H., Zipprich J.T., Attig J., Colic M., Parker R., Filipowicz W.. miRNA repression involves GW182-mediated recruitment of CCR4-NOT through conserved W-containing motifs. Nat. Struct. Mol. Biol. 2011; 18:1218–1226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Doidge R., Mittal S., Aslam A., Winkler G.S.. The anti-proliferative activity of BTG/TOB proteins is mediated via the Caf1a (CNOT7) and Caf1b (CNOT8) deadenylase subunits of the Ccr4-not complex. PLoS One. 2012; 7:e51331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Chen Y., Boland A., Kuzuoglu-Ozturk D., Bawankar P., Loh B., Chang C.T., Weichenrieder O., Izaurralde E.. A DDX6-CNOT1 complex and W-binding pockets in CNOT9 reveal direct links between miRNA target recognition and silencing. Mol. Cell. 2014; 54:737–750. [DOI] [PubMed] [Google Scholar]
- 34. Mathys H., Basquin J., Ozgur S., Czarnocki-Cieciura M., Bonneau F., Aartse A., Dziembowski A., Nowotny M., Conti E., Filipowicz W.. Structural and biochemical insights to the role of the CCR4-NOT complex and DDX6 ATPase in microRNA repression. Mol. Cell. 2014; 54:751–765. [DOI] [PubMed] [Google Scholar]
- 35. Huntzinger E., Kuzuoglu-Ozturk D., Braun J.E., Eulalio A., Wohlbold L., Izaurralde E.. The interactions of GW182 proteins with PABP and deadenylases are required for both translational repression and degradation of miRNA targets. Nucleic Acids Res. 2013; 41:978–994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Baltz A.G., Munschauer M., Schwanhausser B., Vasile A., Murakawa Y., Schueler M., Youngs N., Penfold-Brown D., Drew K., Milek M. et al. The mRNA-bound proteome and its global occupancy profile on protein-coding transcripts. Mol. Cell. 2012; 46:674–690. [DOI] [PubMed] [Google Scholar]
- 37. Beckmann B.M., Horos R., Fischer B., Castello A., Eichelbaum K., Alleaume A.M., Schwarzl T., Curk T., Foehr S., Huber W. et al. The RNA-binding proteomes from yeast to man harbour conserved enigmRBPs. Nat. Commun. 2015; 6:10127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Kwon S.C., Yi H., Eichelbaum K., Fohr S., Fischer B., You K.T., Castello A., Krijgsveld J., Hentze M.W., Kim V.N.. The RNA-binding protein repertoire of embryonic stem cells. Nat. Struct. Mol. Biol. 2013; 20:1122–1130. [DOI] [PubMed] [Google Scholar]
- 39. Castello A., Fischer B., Eichelbaum K., Horos R., Beckmann B.M., Strein C., Davey N.E., Humphreys D.T., Preiss T., Steinmetz L.M. et al. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell. 2012; 149:1393–1406. [DOI] [PubMed] [Google Scholar]
- 40. Jagannathan S., Hsu J.C., Reid D.W., Chen Q., Thompson W.J., Moseley A.M., Nicchitta C.V.. Multifunctional roles for the protein translocation machinery in RNA anchoring to the endoplasmic reticulum. J. Biol. Chem. 2014; 289:25907–25924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Chi S.W., Zang J.B., Mele A., Darnell R.B.. Argonaute HITS-CLIP decodes microRNA-mRNA interaction maps. Nature. 2009; 460:479–486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Kini H.K., Silverman I.M., Ji X., Gregory B.D., Liebhaber S.A.. Cytoplasmic poly(A) binding protein-1 binds to genomically encoded sequences within mammalian mRNAs. RNA. 2016; 22:61–74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Leung A.K., Young A.G., Bhutkar A., Zheng G.X., Bosson A.D., Nielsen C.B., Sharp P.A.. Genome-wide identification of Ago2 binding sites from mouse embryonic stem cells with and without mature microRNAs. Nat. Struct. Mol. Biol. 2011; 18:237–244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Spengler R.M., Zhang X., Cheng C., McLendon J.M., Skeie J.M., Johnson F.L., Davidson B.L., Boudreau R.L.. Elucidation of transcriptome-wide microRNA binding sites in human cardiac tissues by Ago2 HITS-CLIP. Nucleic Acids Res. 2016; 44:7120–7131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Sinkkonen L., Hugenschmidt T., Berninger P., Gaidatzis D., Mohn F., Artus-Revel C.G., Zavolan M., Svoboda P., Filipowicz W.. MicroRNAs control de novo DNA methylation through regulation of transcriptional repressors in mouse embryonic stem cells. Nat. Struct. Mol. Biol. 2008; 15:259–267. [DOI] [PubMed] [Google Scholar]
- 46. Aeschimann F., Kumari P., Bartake H., Gaidatzis D., Xu L., Ciosk R., Grosshans H.. LIN41 post-transcriptionally silences mRNAs by two distinct and position-dependent mechanisms. Mol. Cell. 2017; 65:476–489. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The microarray data have been deposited with the GEO Data bank under accession number GSE101098.