Abstract
With the advent of new crosslinking chemistries, analytical technologies, and search algorithms, crosslinking has become an increasingly popular strategy for evaluating tertiary and quaternary structures of proteins. Collisional activated dissociation remains the primary MS/MS method for identifications of peptide crosslinks in high throughput workflows. 193 nm ultraviolet photodissociation (UVPD) has emerged as an alternative ion activation method well-suited for characterization of peptides and has been found in some cases to identify different peptides or provide distinctive sequence information than obtained by collisional activation methods. Complementary high energy collision dissociation (HCD) and UVPD were used in the present study to characterize protein crosslinking for bovine serum albumin, hemoglobin, and E. coli ribosome. Crosslinks identified by HCD and UVPD using bis(sulfosuccinimidyl)suberate (BS3), a homobifunctional amine-to-amine crosslinker, and 4-(4,6-dimethoxy-1,3,5-triazin-2-yl)-4-methylmorpholinium chloride (DMTMM), a heterofunctional amine-to-carboxylic acid crosslinker, were evaluated in the present study. While more unique BS3 crosslinks were identified by HCD, UVPD and HCD provided a complementary panel of DMTMM crosslinks which extended the degree of structural insight obtained for the proteins.
Introduction
Protein crosslinking integrated with mass spectrometry has become an indispensable strategy in the field of structural biology, with applications ranging from structural studies of single proteins and monitoring conformational changes to deciphering networks of protein-protein interactions.1–4 Extending this crosslinking methodology to even greater challenges depends on the ability to extract and interpret the most complete information possible from experiments that are both data rich and saturated with potential false positives. Nearly all crosslinking experiments use collisional activation (CID or HCD) for crosslink identification, with electron activation (like electron transfer dissociation, ETD) and hybrids like EThcD applied more recently.5,6 Ultraviolet photodissociation (UVPD) has emerged as a complementary activation technique for both bottom-up and top-down proteomics7–11 and has recently been used to detect and elucidate crosslinks via a top-down approach demonstrated for ubiquitin as well as hexameric-insulin.12 These successes have prompted the evaluation of UVPD for the identification of crosslinked peptides via a bottom-up workflow in the present study. A particular benefit of UVPD is the production of fragmentation patterns which enhance confidence in localizing modification sites of peptides or proteins,7 a feature that is ideal for confirming crosslinked sites and identifying the sequences of constituent peptides connected by a crosslink. As reported in the present study, the complementarity between HCD and UVPD reveals a more complete picture of protein crosslinks. Two crosslinkers are used here: bis(sulfosuccinimidyl)suberate (BS3), an amine-amine crosslinker, and a relatively new zero-length amine-to-carboxylic acid crosslinker, DMTMM (4-(4,6-dimethoxy-1,3,5-triazin-2-yl)-4-methylmorpholinium chloride).13,14 Three protein systems are evaluated: bovine serum albumin, hemoglobin (a tetrameric complex), and the more complex multi-protein E. coli ribosome.
Methods
Bovine serum albumin (BSA) and hemoglobin (Hb) were reconstituted into 1× phosphate buffered saline (PBS) at pH 7.2 at 10 uM and 50 uM, respectively. These solutions were reacted at a 1:10 or 1:50 protein:crosslinker molar ratio for BS3 or DMTMM, respectively, for 1 hour at room temperature, and subsequently quenched with ammonium acetate in 50× excess of the crosslinker. 100 μg of E. coli ribosome (E. coli rib) was reacted with 1 mM BS3 or 5 mM DMTMM for 30 minutes at room temperature at 1 mg/ml ribosome concentration. The structures of the crosslinkers and the types of resulting products are illustrated in Scheme S1. After the crosslinking reaction, solutions were diluted in 150 mM ammonium bicarbonate and digested with trypsin using a 1:40 ratio of protease:protein for 16 hours at 37°C. Digested samples were separated by reversed phase C18 nano-scale liquid chromatography and analyzed using a Thermo Scientific Orbitrap Fusion Lumos Tribrid mass spectrometer modified for UVPD using a 500 Hz 193 nm excimer laser as described previously.15 Sample analysis was performed at top-speed mode with a MS1 resolution of 60,000 at m/z 200 or MS2 resolution of 30,000 at m/z 200 and averaging 1 or 2 μscans, respectively. In the top-speed mode, the user selects the length of time between sequential MS1 scans, and as many precursor peptides as possible are isolated and activation during the intervening period. The number of MS2 spectra per MS1 spectrum is variable using the top-speed mode, unlike a traditional top N method where exactly N isolation/activation events (MS/MS spectra) are performed following each MS1 spectrum. 30% NCE was applied for HCD experiments, and 2 pulses at 2.25 mJ per 5 ns laser pulse were used for UVPD activation. The eluting peptides were subjected to HCD and UVPD in separate runs. Kojak-MS was used for crosslink-identification, and the reported scores are cross-correlation scores.16,17
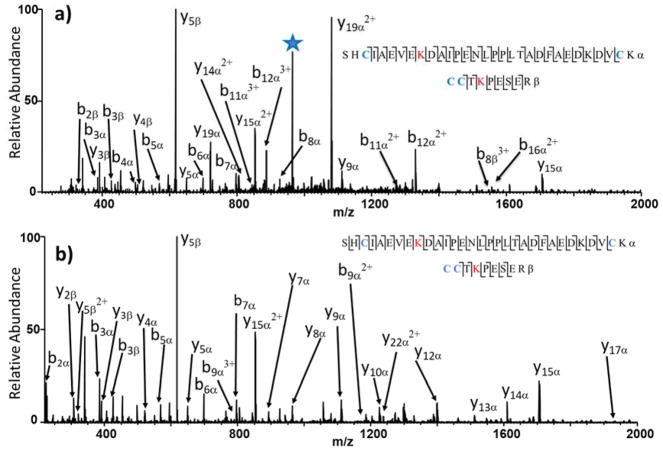
The two constituent peptides are labeled α (larger peptide) and β (smaller peptide), and cleavage along the backbone of each connected peptide yields product ions such as bnα or ynα depending on its peptide of origin and whether it contains the N- or C-terminus. If the number of residues in the designated b or y ion span the crosslinking site, then the entire other peptide is also included in the product. As an example, there is a fragment ion labelled as b12α (2+, m/z 1329.11) in Figure 1a. This large peptide contains the first 12 residues of the alpha peptide (SHCIAEVEKDAI) plus the entire beta peptide (CCTKPESER) plus the crosslinker moiety. The crosslink site for the alpha peptide in Figure 1 occurs at K9. This means that b-type fragment ions of the alpha peptide containing less than nine residues (b1, b2, b3, b4, b5, b6, b7, b8) are small peptides that do not contain the crosslink moiety nor the appended beta peptide. Percolator was used to calculate crosslinking false discovery rate (FDR).16,18 Crosslink searching was limited to two allowed missed cleavages, and the searches focused on b- and y-type ions for both HCD and UVPD with MS1 and MS2 mass tolerances of 10 ppm. While it is known that UVPD typically generates an array of ions (a,b,c,x,y,z) for larger peptides and proteins, in fact the production of b/y ions is most prominent for UVPD of the crosslinked peptides, as illustrated by the fragment ion distributions shown in Figure S1. This feature is further reflected by the sharp drop in identified crosslinks based on FDR calculations for UVPD when all fragment ion types were considered relative to inclusion of only b/y ion types (Figure S2). Based on the dominance of b/y ions in the spectra, the other fragment ion types were excluded from all searches. BS3 crosslinks were searched between all free amines (protein N-terminus and side-chains of lysines) with a mass shift of 138.0681 Da. DMTMM crosslinks were searched between carboxylic acids (C-terminus, side-chains of glutamic and aspartic acids) and free amines with a mass shift of −18.0109 Da. Crosslinks under 5% FDR that were matched in at least two of the three replicate runs were considered confidently identified.
Figure 1.
MS/MS spectra obtained for a BS3-crosslinked product (5+) from BSA obtained by (a) UVPD (2 pulses at 2.5 mJ) and (b) HCD (NCE 30%). The precursor ion is labelled with a star. Not all ions are labelled due to spectral congestion. The crosslinked sites are shown in red font. The relative locations of the two crosslinked peptides are shown in the sequence of BSA in Figure S6. Lists of fragment ions identified in the UVPD and HCD spectra are shown in Tables S5a and S5b with the respective intensity, m/z, theoretical m/z, deviation in ppm, and charge state of each observed ion.
Discussion
After crosslinking, tryptic digestion, and LC-MS/MS analysis using both HCD and UVPD, the resulting crosslinked peptide identifications were enumerated for each of the three protein systems for each of the two crosslinkers (BS3 and DMTMM) (Table 1). The specific crosslinked peptides identified from the HCD and UVPD datasets for all protein systems are summarized in Tables S1 and S2 for BS3 and DMTMM identifications, respectively (and complete lists of hits are provided in Tables S3 and S4. Examples of mass spectra from high scoring unique and common crosslinks are shown in Figures 1, S3, S4, and S5 along with lists of identified fragment ions in Tables S5, S6, S7 and S8. In particular, the HCD and UVPD spectra obtained for one crosslink found in common by each of the two activation methods are shown in Figure 1 for a BS3 crosslinked product (5+) of BSA, and relative locations of the crosslinked peptides are highlighted on the sequence of BSA shown in Figure S6a and companion PYMOL structures in Figure S6b,c. The MS/MS spectra obtained for one DMTMM crosslinked product (4+) found in common by HCD and UVPD are shown Figure S3 for a pair of E.coli ribosomal proteins RS1 and RS17. Examples of UVPD and HCD spectra for unique BS3 crosslinked products from the E. coli ribosome and unique DMTMM crosslinked products from BSA are shown in Figures S4 (along with an expansion of sections of the UVPD spectrum in Figure S4c) and S5, respectively (with relative locations of the alpha and beta peptides in the protein sequences and companion PYMOL structures shown in Figures S7–S9). Both UVPD and HCD produced mainly b and y type ions, achieving high sequence coverage for the pairs of peptides involved in each crosslink. There was little fragmentation of the crosslinker moiety by either activation method for any of the crosslinked peptides, as confirmed by the MS/MS spectra in Figures 1, S3, S4 and S5.
Table 1.
Number of unique and common crosslinked peptides for bovine serum albumin (BSA), hemoglobin (Hb), and E. coli ribosome (E. coli rib) using the two crosslinkers BS3 and DMTMM.
| Protein/XL | HCD unique | UVPD unique | Common | Total |
|---|---|---|---|---|
| BSA/DMTMM | 13 | 21 | 29 | 63 |
| BSA/BS3 | 36 | 6 | 40 | 82 |
| Hb/DMTMM | 8 | 5 | 11 | 24 |
| Hb/BS3 | 37 | 1 | 42 | 80 |
| E. coli rib/DMTMM | 15 | 7 | 8 | 39 |
| E. coli rib/BS3 | 16 | 6 | 19 | 41 |
Across all systems a large number of peptide crosslinks were identified in common by HCD and UVPD (Table 1), in addition to a surprisingly large number of unique ones identified by only HCD or only UVPD, suggesting a degree of complementarity between the two MS/MS methods. The number of unique BS3-type crosslinks identified by UVPD was relatively low for BSA, Hb and E.coli ribosome compared to the number identified by HCD (Table 1). Conversely, the number of unique DMTMM-type crosslinks identified by UVPD and HCD were similar for all three protein systems. The numbers of specific residues that were identified as crosslinking sites are summarized in Table S1. These latter numbers are smaller than the total numbers of crosslinked products in Table 1, indicating that some of the residues are engaged in multiple crosslinks.
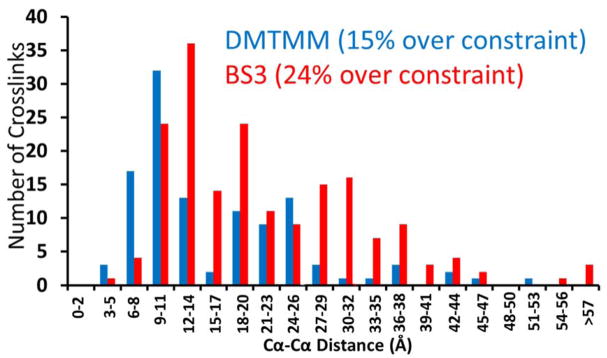
Distances between crosslinked sites were calculated from known BSA (PDB: 3V03) and Hb (PDB: 1GZX) crystal structures and are graphically displayed in Figure 2. BS3 has a maximum distance constraint of 30 Å between Cα atoms of lysines.13,19 Typical zero-length crosslinkers, such as EDC or DMTMM, have a distance constraint of approximately 15 Å between Cα atoms of crosslinked residues. However, it has been suggested that DMTMM has a distance constraint of 25 Å between a primary amine and carboxylic acid site.13 Approximately 24% of the BS3 crosslinks and 15% of the DMTMM crosslinks exceed the expected Cα-Cα distance derived from the BSA and Hb datasets (i.e. 30 Å for BS3 and 25 Å for DMTMM13). This is an acceptable level of agreement, particularly considering that oligomeric protein states which may contribute to the formation of intermolecular crosslinks were excluded from the distance Cα-Cα calculations. Additionally, the crosslinks found for the ribosome sample were congruent with previous crosslinking experiments and known protein-protein interactions.20
Figure 2.
Comparison of crosslinking distance constraints from the identified DMTMM and BS3 crosslinked products for BSA and Hb. DMTMM yields a constraint of 25 Å, whereas BS3 exhibits a constraint of 30 Å.
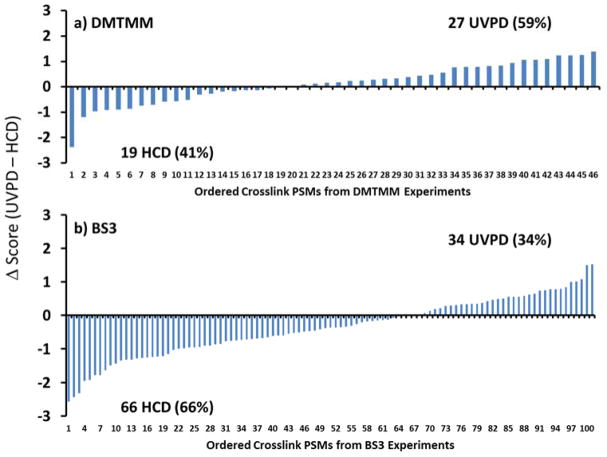
In total there were 48 DMTMM crosslinks and 101 BS3 crosslinks identified in common for HCD and UVPD after combining all the results for the three protein systems (BSA + Hb + E. coli ribosome). The corresponding Kojak-derived confidence scores with standard deviations for each of the common crosslink identifications are displayed in Figure S10. To highlight the variations in the scores returned by UVPD versus HCD, the differences (UVPD score – HCD score) were calculated and sorted (Figure 3). This comparison revealed that UVPD yielded better scores than HCD for a large portion (59%) of the commonly identified DMTMM crosslinks. In contrast, HCD provided better scores for more of the BS3 crosslinks (66%). Additionally, while a majority of the crosslinks found in common by UVPD and HCD had the same charge state, UVPD consistently found a number of crosslinks in higher charge states and HCD identified quite a few in lower charge states (Figure S11). HCD and UVPD can also be implemented in alternating scans in a single run. This requires changing the spectral acquisition mode from top speed to top N (where N = 10), a change that gives less flexibility with respect to the number of MS/MS spectra acquired. Unique peptide identifications were still found for UVPD and HCD; however, the total number of crosslinks identified decreased owing to dividing the MS/MS period between two activation modes (Figure S12).
Figure 3.
Differences in Kojak scores between UVPD and HCD (UVPD score - HCD score) for (a) DMTMM crosslinked products and (b) BS3 crosslinked products for all crosslinked peptides found in common for BSA, Hb, and the E.coli ribosomal proteins. The peptides are ordered based on the difference in HCD and UVPD scores, starting with the ones for which HCD performed best and extending to the ones for which UVPD performed best.
The features of those crosslinked products that were uniquely identified by HCD versus UVPD were examined in greater detail in order to decipher characteristics that might shed light on the variation in analytical performance for HCD versus UVPD. Two properties of particular interest were the charge states and sizes of the crosslinked species (in terms of the net mass). The results are summarized in Figure S13. For BS3, although there were many crosslinks identified in common by HCD and UVPD, there were insufficient unique BS3 crosslinks identified by UVPD to uncover any trends related to the size or charge state of the crosslinked products (Figure S13a,b). In contrast, the average crosslink masses and charge states determined for the DMTMM crosslinks (Table S2, Figure S13c,d) revealed that UVPD in general identified species in higher charge states and with greater masses. The charge state finding is echoed in Figure S11 which illustrates the number of crosslinks found in higher or lower charge states for UVPD and HCD based on comparison of all species found in common by both MS/MS methods. Another notable difference is observed in the charge states of the fragment ions created by HCD and UVPD upon analysis of the same crosslinked products. (see Figures 1, S3–5) There are far more highly charged (3+, 2+) fragment ions identified in the UVPD spectra, whereas nearly all of the fragment ions identified by HCD are singly or doubly charged. This latter finding is not surprising considering that the fast, high energy deposition of UVPD may promote fragmentation directly from excited states in a manner that does not entail significant proton migration.
While there are insufficient crosslink identifications to identify the differentiating features contributing to the complementarity of the HCD and UVPD mass spectra, some speculation is possible based on the factors that modulate fragmentation pathways promoted by HCD versus UVPD. First, the duty cycles for UVPD and HCD are comparable. However, the S/N ratio of UVPD spectra is typically lower than that of HCD spectra because the photon flux is restricted to limit secondary dissociation. For BS3-crosslinked peptides two very basic sites (lysine side-chains) are converted into non-basic amides, thus potentially increasing the proton mobility, a factor that is expected to enhance non-specific fragmentation upon HCD.21 Non-specific fragmentation denotes production of diagnostic b/y sequence ions throughout the peptide backbone as opposed to specific preferential cleavages adjacent to Pro for peptides with mobile protons and adjacent to Glu and Asp for peptides with no mobile protons.22 For DMTMM-crosslinked peptides, one basic lysine side-chain is converted to an amide (a less basic site) and one acidic site (e.g., side-chain of Glu or Asp) of the second peptide is converted to an amide, a factor that seems to suppress preferential Glu/Asp cleavages which are prominent upon collisional activation of peptides in low charge states (lacking mobile protons).21,22 These conversions of acidic and basic residues to ones that alter proton mobility have little impact on UVPD, a process that is largely independent of charge state and does not rely on mobile protons. In contrast, HCD is a process mediated by mobile protons and prone to enhanced preferential cleavages adjacent to Asp, Glu and Pro residues depending on the charge state.21,22
In summary 193 nm UVPD and HCD serve as complementary MS/MS techniques for identifying both amine-to-amine (BS3) and amine-to-carboxylic acid (DMTMM) crosslinks. Both UVPD and HCD provided extensive sequence coverage of the connected peptides and allow unambiguous localization of the sites of crosslinks. Integrating the findings from UVPD and HCD for identifying DMTMM type crosslinks should be particularly beneficial for expanding the degree of structural information obtained from a protein crosslinking experiment. The impact of combining UVPD and HCD is less significant for identification of BS3 crosslinks owing to the low number of crosslinks uniquely identified by UVPD. UVPD consistently returned higher confidence scores for DMTMM crosslinks, whereas HCD yielded better scores for BS3 crosslinks. Differences in the types of unique DMTMM crosslinks identified by UVPD versus HCD were found based on the sizes of the crosslinked products and charge states, with UVPD favoring the identification of larger crosslinked species in higher charge states. The performance of UVPD was enhanced for crosslinks involving acidic residues, an interesting outcome that merits further exploration.
Supplementary Material
Scheme S1. Crosslinking reagents and their reactions for (a) BS3 and (b) DMTMM.
Table S1. Number of specific crosslinked amino acids identified among all pairs of identified crosslinked peptides in Table 1 (for bovine serum albumin (BSA), hemoglobin (Hb), and E. coli ribosome (E. coli rib).
Table S2. Mass and charge state weighted averages between those crosslinks uniquely identified by UVPD and HCD for crosslinker DMTMM.
Tables S3 and S4: Excel spreadsheets listing all crosslinked products for BS3 (Table S3) and DMTMM (Table S4) for BSA, Hb, and the ribosomal protein data sets.
Tables S5-S8. Lists of identified fragment ions, experimental and theoretical masses, mass error, type of ion, and charge state for the companion UVPD and HCD spectra.
Figure S1. Representative relative ion abundances from BS3 and DMTMM crosslinks identified by UVPD from spectra seen in Figure 1a and Figure S2a.
Figure S2. FDR plot of BS3 crosslinked BSA fragmented with UVPD. The red line indicates the search was performed with only b/y type ions while the blue line indicates the search was performed with abc/xyz type ions.
Figure S3. MS/MS spectra obtained for a DMTMM-crosslinked product (4+) between E. coli ribosomal proteins RS1 and RL17 obtained by (a) UVPD (2 pulses at 2.5 mJ) and (b) HCD (NCE 30).
Figure S4. Fragmentation of uniquely identified BS3 crosslinks: (a) E. coli ribosomal protein L5 (4+) based on UVPD, and (b) Hb crosslink (6+) based on HCD and expanded sections of UVPD spectrum in (c).
Figure S5. Fragmentation of uniquely identified DMTMM crosslinks from BSA based on (a) UVPD (6+) and (b) HCD (6+).
Figure S6. Relative locations of alpha (red) and beta (blue) peptides corresponding to the crosslink illustrated in Figure 1.
Figure S7. Relative locations of alpha (red) and beta (blue) peptides corresponding to the crosslink illustrated in Figure S4b for hemoglobin Hb.
Figure S8. Relative locations of alpha (red) and beta (blue) peptides corresponding to the crosslink illustrated in Figure S5a.
Figure S9. Relative locations of alpha (red) and beta (blue) peptides corresponding to the crosslink illustrated in Figure S5b.
Figure S10. Common crosslinks identified between HCD and UVPD for (a) DMTMM and (b) BS3, sorted by highest to lowest Kojak score based on the UVPD data.
Figure S11. Charge difference between those crosslinks found in common by UVPD and HCD for (a) DMTMM and (b) BS3 crosslinks.
Figure S12. Comparison of (a,c) charge states and (b,d) masses of the identified BS3 (a,b) and DMTMM (c,d) crosslinks.
Figure S13. Comparing crosslink identifications for BS3-crosslinked BSA obtained by standard top N HCD-only or UVPD-only runs and runs in which HCD and UVPD spectra were acquired in alternating scans.
Acknowledgments
We acknowledge the following funding sources: NSF (Grant CHE1402753 and CHE1559838), the Welch Foundation (Grant F-1155), and NIH 1K12GM102745 (fellowship to MBC). Funding from the UT System for support of the UT System Proteomics Core Facility Network is gratefully acknowledged.
References
- 1.Holding AN. Methods. 2015;89:54–63. doi: 10.1016/j.ymeth.2015.06.010. [DOI] [PubMed] [Google Scholar]
- 2.Kahraman A, Herzog F, Leitner A, Rosenberger G, Aebersold R, Malmström L. PLOS ONE. 2013;8:e73411. doi: 10.1371/journal.pone.0073411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Pham ND, Parker RB, Kohler JJ. Curr Opin Chem Biol. 2013;17:90–101. doi: 10.1016/j.cbpa.2012.10.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Sinz A. Expert Rev Proteomics. 2014;11:733–743. doi: 10.1586/14789450.2014.960852. [DOI] [PubMed] [Google Scholar]
- 5.Liu F, Rijkers DTS, Post H, Heck AJR. Nat Methods. 2015;12(12):1179–1184. doi: 10.1038/nmeth.3603. [DOI] [PubMed] [Google Scholar]
- 6.Kolbowski L, Mendes ML, Rappsilber J. Anal Chem. 2017;89:5311–5318. doi: 10.1021/acs.analchem.6b04935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Robinson MR, Taliaferro JM, Dalby KN, Brodbelt JS. J Proteome Res. 2016;15:2739–2748. doi: 10.1021/acs.jproteome.6b00289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Madsen JA, Xu H, Robinson MR, Horton AP, Shaw JB, Giles DK, Kaoud TS, Dalby KN, Trent MS, Brodbelt JS. Mol Cell Proteomics. 2013;12:2604–2614. doi: 10.1074/mcp.O113.028258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Shaw JB, Li W, Holden DD, Zhang Y, Griep-Raming J, Fellers RT, Early BP, Thomas PM, Kelleher NL, Brodbelt JS. J Am Chem Soc. 2013;135:12646–12651. doi: 10.1021/ja4029654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cleland TP, DeHart CJ, Fellers RT, VanNispen AJ, Greer JB, LeDuc RD, Parker WR, Thomas PM, Kelleher NL, Brodbelt JS. J Proteome Res. 2017;16:2072–2079. doi: 10.1021/acs.jproteome.7b00043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Greer SM, Parker WR, Brodbelt JS. J Proteome Res. 2015;14:2626–2632. doi: 10.1021/acs.jproteome.5b00165. [DOI] [PubMed] [Google Scholar]
- 12.Cammarata MB, Brodbelt JS. ChemistrySelect. 2016;1:590–593. [Google Scholar]
- 13.Leitner A, Joachimiak LA, Unverdorben P, Walzthoeni T, Frydman J, Förster F, Aebersold R. Proc Natl Acad Sci U S A. 2014;111:9455–9460. doi: 10.1073/pnas.1320298111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Leitner A. Chem Sci. 2016;7:4792–4803. doi: 10.1039/c5sc04196a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Klein DR, Holden DD, Brodbelt JS. Anal Chem. 2016;88:1044–1051. doi: 10.1021/acs.analchem.5b04218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Hoopmann MR, Mendoza L, Deutsch EW, Shteynberg D, Moritz RL. J Am Soc Mass Spectrom. 2016;27:1728–1734. doi: 10.1007/s13361-016-1435-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hoopmann MR, Zelter A, Johnson RS, Riffle M, MacCoss MJ, Davis TN, Moritz RL. J Proteome Res. 2015;14:2190–2198. doi: 10.1021/pr501321h. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Klammer AA, Park CY, Noble WS. J Proteome Res. 2009;8:2106–2113. doi: 10.1021/pr8011107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Merkley ED, Rysavy S, Kahraman A, Hafen RP, Daggett V, Adkins JN. Protein Sci. 2014;23:747–759. doi: 10.1002/pro.2458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Lauber MA, Rappsilber J, Reilly JP. Mol Cell Proteomics MCP. 2012;11:1965–1976. doi: 10.1074/mcp.M112.019562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Wysocki VH, Tsaprailis G, Smith LL, Breci LA. J Mass Spectrom. 2000;35:1399–1406. doi: 10.1002/1096-9888(200012)35:12<1399::AID-JMS86>3.0.CO;2-R. [DOI] [PubMed] [Google Scholar]
- 22.Paizs B, Suhai S. Mass Spectrom Rev. 2005;24:508–548. doi: 10.1002/mas.20024. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Scheme S1. Crosslinking reagents and their reactions for (a) BS3 and (b) DMTMM.
Table S1. Number of specific crosslinked amino acids identified among all pairs of identified crosslinked peptides in Table 1 (for bovine serum albumin (BSA), hemoglobin (Hb), and E. coli ribosome (E. coli rib).
Table S2. Mass and charge state weighted averages between those crosslinks uniquely identified by UVPD and HCD for crosslinker DMTMM.
Tables S3 and S4: Excel spreadsheets listing all crosslinked products for BS3 (Table S3) and DMTMM (Table S4) for BSA, Hb, and the ribosomal protein data sets.
Tables S5-S8. Lists of identified fragment ions, experimental and theoretical masses, mass error, type of ion, and charge state for the companion UVPD and HCD spectra.
Figure S1. Representative relative ion abundances from BS3 and DMTMM crosslinks identified by UVPD from spectra seen in Figure 1a and Figure S2a.
Figure S2. FDR plot of BS3 crosslinked BSA fragmented with UVPD. The red line indicates the search was performed with only b/y type ions while the blue line indicates the search was performed with abc/xyz type ions.
Figure S3. MS/MS spectra obtained for a DMTMM-crosslinked product (4+) between E. coli ribosomal proteins RS1 and RL17 obtained by (a) UVPD (2 pulses at 2.5 mJ) and (b) HCD (NCE 30).
Figure S4. Fragmentation of uniquely identified BS3 crosslinks: (a) E. coli ribosomal protein L5 (4+) based on UVPD, and (b) Hb crosslink (6+) based on HCD and expanded sections of UVPD spectrum in (c).
Figure S5. Fragmentation of uniquely identified DMTMM crosslinks from BSA based on (a) UVPD (6+) and (b) HCD (6+).
Figure S6. Relative locations of alpha (red) and beta (blue) peptides corresponding to the crosslink illustrated in Figure 1.
Figure S7. Relative locations of alpha (red) and beta (blue) peptides corresponding to the crosslink illustrated in Figure S4b for hemoglobin Hb.
Figure S8. Relative locations of alpha (red) and beta (blue) peptides corresponding to the crosslink illustrated in Figure S5a.
Figure S9. Relative locations of alpha (red) and beta (blue) peptides corresponding to the crosslink illustrated in Figure S5b.
Figure S10. Common crosslinks identified between HCD and UVPD for (a) DMTMM and (b) BS3, sorted by highest to lowest Kojak score based on the UVPD data.
Figure S11. Charge difference between those crosslinks found in common by UVPD and HCD for (a) DMTMM and (b) BS3 crosslinks.
Figure S12. Comparison of (a,c) charge states and (b,d) masses of the identified BS3 (a,b) and DMTMM (c,d) crosslinks.
Figure S13. Comparing crosslink identifications for BS3-crosslinked BSA obtained by standard top N HCD-only or UVPD-only runs and runs in which HCD and UVPD spectra were acquired in alternating scans.