Abstract
The goal of this research was to elucidate the relationship between WHO 2016 molecular classifications of newly diagnosed, nonenhancing lower grade gliomas (LrGG), tissue sample histopathology, and magnetic resonance (MR) parameters derived from diffusion, perfusion, and 1H spectroscopic imaging from the tissue sample locations and the entire tumor. A total of 135 patients were scanned prior to initial surgery, with tumor cellularity scores obtained from 88 image-guided tissue samples. MR parameters were obtained from corresponding sample locations, and histograms of normalized MR parameters within the T2 fluid-attenuated inversion recovery lesion were analyzed in order to evaluate differences between subgroups. For tissue samples, higher tumor scores were related to increased normalized apparent diffusion coefficient (nADC), lower fractional anisotropy (nFA), lower cerebral blood volume (nCBV), higher choline (nCho), and lower N-acetylaspartate (nNAA). Within the T2 lesion, higher tumor grade was associated with higher nADC, lower nFA, and higher Cho to NAA index. Pathological analysis confirmed that diffusion and metabolic parameters increased and perfusion decreased with tumor cellularity. This information can be used to select targets for tissue sampling and to aid in making decisions about treating residual disease.
Introduction
Gliomas are the most common type of primary brain cancer in adults and are classified by the WHO 2016 classification scheme as either glioblastoma (GBM, grade IV) or lower-grade (grades II and III) diffuse gliomas. These lower-grade gliomas (LrGG) are further defined as astrocytomas (isocitrate dehydrogenase (IDH)–mutant or IDH wild-type) or as oligodendrogliomas (IDH-mutant with chromosome 1p/19q codeletion) [1]. While overall survival for LrGG has lengthened to 5 to 20 years due to advances in diagnosis and therapy, the disease remains almost inevitably fatal [2]. While the initial step in treatment is surgical resection, at the current time, there is no definitive standard of care for subsequent treatment [3]. The lack of consensus stems partly from the difficulty in predicting prognosis and defining response to the various therapeutic approaches for individual patients.
Although the WHO 2016 classification is obtained by analyzing tissue samples that are acquired at the time of initial surgery, there is still considerable variability in the course of the disease for individual patients [4]. The identification of imaging markers that can guide personalized treatment is especially important. Multiparametric MR examinations that integrate anatomic imaging with diffusion (DWI), perfusion (PWI), and spectroscopic (MRSI) imaging have been developed to provide a more objective and comprehensive evaluation of the structural, physiological, and metabolic properties of gliomas. These advanced imaging parameters are particularly important for treatment planning in patients newly diagnosed as having LrGG because the majority of them have no gadolinium contrast enhancement on T1-weighted anatomical imaging and the definition of tumor burden relies upon the evaluation of regions with hyperintensity on T2-weighted images.
The goal of this study was to elucidate the relationship between molecular classifications of LrGG, histological characteristics, and MR parameters. The population comprised 135 patients with newly diagnosed, nonenhancing LrGG, who were scanned before their initial surgery. Image-guided tissue samples were collected in a subset of patients and analyzed to relate their molecular characteristics and pathology to imaging characteristics.
Methods
UCSF Institutional Review Board approval was obtained to study patients with a suspected pathological diagnosis of a WHO grade II or III glioma without gadolinium contrast enhancement on T1-weighted MR images. Patients were recruited immediately prior to surgical resection and gave written informed consent.
MR Acquisition
MR examinations were performed on 1.5-T or 3-T scanners (GE Healthcare Technologies) using an eight-channel phased-array headcoil (MRI Devices). Standard anatomical imaging included T2-weighted fluid-attenuated inversion recovery (FLAIR), fast spin echo, and T1-weighted pre- and postgadolinium sequences. Diffusion-weighted images (DWI) were obtained in the axial plane with six gradient directions [repetition time (TR)/echo time (TE) = 1000/108 milliseconds, voxel size = 1.7 × 1.7 × 3 mm, b = 1000 s/mm]. Dynamic susceptibility contrast-enhanced perfusion-weighted images (PWI) obtained following a 5-ml/s bolus injection of 0.1 mmol/kg body weight gadolinium diethyltriamine pentaacetic acid and were acquired using a series of T2*-weighted echo-planar images [TR/TE/flip angle = 1250-1500/35-54 milliseconds/30°-35°, 128 × 128 matrix, slice thickness = 3-5 mm, 7-15 slices with 60-80 time points] before, during, and after arrival of the contrast agent. Lactate-edited 3D proton MR spectroscopic imaging data were obtained using point-resolved spectroscopic selection or volume localization and very selective saturation pulses for lipid signal suppression [excited volume = 80 × 80 × 40 mm, TR/TE = 1104/144 milliseconds, overpress factor = 1.5, field of view = 16 × 16 × 16 cm, nominal voxel size = 1 × 1 × 1 cm, flyback echo-planar readout gradient in the SI direction, 988 Hz sweep width and 712 dwell points] [5].
MR Data Processing
The anatomic, diffusion, perfusion, and spectroscopic images were aligned to the T1 postcontrast image using FMRIB's Linear Image Registration Tool [6], [7] (http://fsl.fmrib.ox.ac.uk). Each ROI was defined by a single investigator (either the first or second author), with the guidance of UCSF neuroradiologists. The T2 lesion ROI was defined using semiautomated software (BrainLAB Systems, Munich) to include all T2 FLAIR hyperintensity, relative to the surrounding normal tissue. These T2 lesion ROIs are 3D volumetric ROIs that include all T2 FLAIR hyperintensity on all slices. (Figure 1). When tissue samples were available, 5-mm–diameter spherical ROIs centered on the spatial coordinates recorded using the surgical navigation software (BrainLAB, Munich) were generated. The accuracy of the registration of the biopsy location to the T1 image within the Brainlab system has been documented previously [8], particularly by Paraskevopoulos et al. [9], who assessed this accuracy using a viscoelastic phantom to replicate tissue shift. They reported that biopsy location was accurate within 1.5 mm. A 5-mm–diameter ROI was chosen to balance the potential biopsy localization error and the need to restrict the ROI to immediate vicinity of the sampled tissue.
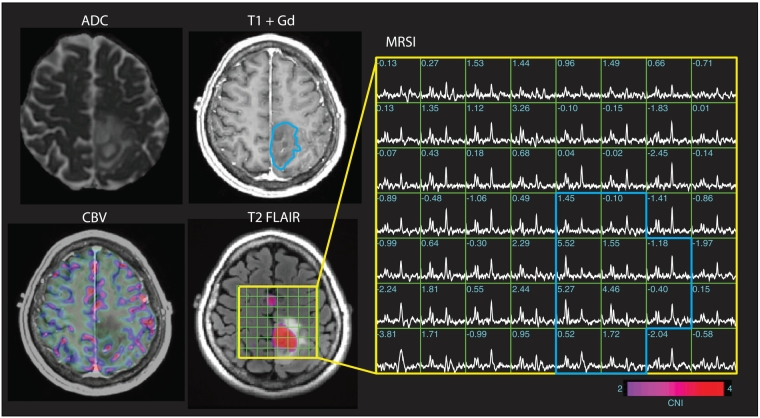
Figure 1.
Example of ADC map, T1 contrast (+Gd), CBV perfusion map, T2 FLAIR, and MRSI data from a single LrGG patient. The blue outline on the T1 + Gd image and MRSI grid indicate the T2 lesion ROI. On the CBV map, pink regions reflect higher CBV. The MRSI grid indicates the spectra and CNI values from each voxel, and the color map superimposed on the T2 FLAIR image indicates voxels with abnormally high CNI.
For DWI data, a previously published algorithm [10] was applied to estimate relevant DWI parameters and normalize between field strengths using estimates from normal-appearing brain tissue. In order to provide metrics that described regions with abnormal intensities, the voxel values for ADC and FA maps were first normalized to the mode of intensities in normal-appearing brain tissue (calculated across the entire cerebrum, excluding the T2 lesion ROI). Percentiles (10th, 50th, 90th) were then calculated from histograms of normalized intensities within T2 lesion ROIs. ADC and FA are typically interpreted as measures of tissue microarchitecture and cellularity in gliomas [11], [12], [13], [14]. By measuring normalized ADC (nADC) and normalized FA (nFA) in the voxels with the highest and lowest diffusion values across the tumor, it was possible to explore whether the presence of some regions of very high or very low diffusion had a significant relationship to molecular characteristics and pathology.
For PWI data, cerebral blood volume (CBV) and peak height (PH) were calculated for each voxel using software developed in our research group [15], [16]. The T2* signal-intensity time curves acquired during the first pass of the gadolinium bolus were converted to change in the relaxation rate (DR2*) and resampled to match the spatial resolution of the anatomic image series. PH was defined as the maximum DR2* value of the first-pass curve. CBV maps were calculated on a voxel-by-voxel basis utilizing a modified gamma-variate function that takes into account leakage of the contrast agent. PH is a simpler, nonparametric approach to characterizing the DR2* curve. Like CBV, PH is predictive of the relative contribution of simple vascular density in glioma [17]. This parameter requires minimal computational time for postprocessing the data, which makes them a common choice for use in the clinic. The CBV and PH intensities from the T2 lesion and tissue sample ROIs were normalized by values within normal brain [18], [19].
For MRSI data, the methods used for reconstruction and postprocessing are fully automated and have been described in previous publications [20], [21], [22]. In brief, the start point was the raw data file obtained from the lactate edited MRSI sequence. This comprised interleaved acquisitions (or cycles) obtained with radiofrequency pulses that modulated the phase of the lactate peak, with each cycle having eight channels of data corresponding to the multiple receiver coils. The k-space time domain data for each cycle and channel were first filtered with a 4-Hz exponential function in the time domain, zero filled to 1024 points, and Fourier transformed to produce a k-space array of spectra. The next step was to apply the k-space Fourier transforms to produce 3D spatial arrays of spectral data, followed by combination using in-house–developed software that weights the data by coil sensitivities estimated from low-resolution proton density–weighted images [22]. Additional phase corrections were applied in the SI dimension to account for the flyback echo-planar readout gradient [23]. The cycles were summed to produce an array of spectra containing choline, creatine, NAA, and lipid and subtracted to produce an array of spectra containing lactate. Spectra were baseline subtracted and phase and frequency corrected using parameters estimated from the summed array. Metabolite levels were obtained by estimating peak heights and integrated areas from the spectral arrays [20].
Peak heights and areas were determined from baseline-subtracted, frequency- and phase-corrected spectra on a voxel-by-voxel basis [24]. Metabolite peak heights were normalized by median peak heights in normal brain for Cho and Cr (nCho and nCr), and normalized Lac, Lip, and LL (nLac, nLip, and nLL) were estimated by metabolite peak heights divided by the median peak NAA intensity in normal brain. The Cho signal reflects changes in membrane synthesis and turnover associated with cell proliferation and remodeling. NAA is a marker of normal brain tissue associated with the presence of actively functioning neurons. Lac is an end product of anaerobic metabolism in normal cells and aerobic glycolysis in cancer and reflects ischemia and/or hypoxia. The Cr peak includes both creatine and phosphocreatine and is indicative of cellular bioenergetic processes. The choline-to-NAA index (CNI) is a z-score that reflects changes in the relative levels of these two metabolites compared to normal brain voxels, and the choline-to-creatine (CCRI) index is defined in a similar manner [25]. The metabolic lesion was defined as the region with CNI values greater than 2 (Figure 1). Cho, Cr, and NAA intensities were normalized by their median value in voxels that had been identified during the CNI calculation as being from normal brain (nCho, nCr, and nNAA). Levels of Lac were normalized (nLac) by the median level of NAA from voxels within the selected volume but outside of the CNI> 2 region.
Image-Guided Brain Tumor Tissue Sampling
Tissue sample targets were planned for a subset of the patients based on surgically accessible regions of both abnormally decreased ADC and abnormally increased ADC in order to sample a variety of histopathology. Intraoperative navigation guided the neurosurgeons to these locations and was used to obtain a 3D screenshot of the actual target coordinates of the excised tissue sample. The neurosurgeons acquired tissue samples as early as safely possible in the surgical procedure and only from locations that had not yet experienced significant tissue shift. To minimize any unnecessary risks to the patient, no regions of normal-appearing brain were targeted. Samples were immediately fixed, processed, and embedded as previously described [19].
Tumor Cellularity, Immunohistochemistry, and Fluorescence In Situ Hybridization (FISH) Determinations
For H&E-stained slides, a tumor score was given on the basis of the contribution of tumor cellularity to total cellularity. Scores of 0 denoted neuropil without tumor; 1 indicated an infiltrating tumor margin containing detectable, but not abundant, numbers of tumor cells; 2 denoted a more cellular-infiltrated zone; 3 denoted highly cellular tumor with few non-neoplastic cells. Immunohistochemistry for the IDH1 p.R132H mutation and FISH to assess 1p/19q codeletion status were performed as previously described [26], [27].
WHO 2016 Diagnostic Classification
Tumor grade and diagnostic subtype were determined by a board-certified neuropathologist. Subtype classification was based on the 2016 WHO classification molecular and histological criteria [1]. LrGG were classified as either 1) oligodendrogliomas, IDH-mutant and 1p/19q-codeleted; 2) astrocytoma, IDH-mutant (if they were not 1p/19q co-deleted and expressed IDH1 R132H mutant protein); or 3) astrocytoma, not otherwise specified (NOS) (if they had astrocytoma histology and IDH status was unknown or if the IDH1 R132H IHC stain was negative). Note that the IDH wild-type status was not confirmed with IDH1 and IDH2 codon sequencing, and hence, the IDH1 R132H immunonegative astrocytomas were included in the astrocytoma, NOS category. Grade determinations were performed utilizing WHO 2016 criteria. For astrocytoma, elevated mitotic index was the main determinant for distinguishing the grade II from III, and microvascular proliferation and necrosis were not present by definition. For anaplastic oligodendroglioma, IDH-mutant and 1p/19q-codeleted, WHO grade III, brisk mitotic activity (>/=6 per 10 high-power fields) or conspicuous microvascular proliferation was required.
Statistical Analysis
The relationship between histological and molecular tumor characteristics (tumor score, grade, and WHO 2016 subgroup) and MRI parameters was assessed with repeated-measures analyses of variance (using JMP Pro 13.0, SAS Institute Inc.). The cutoff for defining a significant result in this exploratory study was a P value of .05. Effects that remain significant using a Benjamini-Hochberg false discovery rate [28] correction for multiple comparisons are indicated in Table 1, Table 2, Table 3. Significant main effects for variables with three or more levels were followed up with pairwise Tukey-Kramer HSD post hoc tests.
Table 1.
Imaging Results from Tissue Sample ROIs by Tissue Sample Tumor Scores
| Group | Number of Samples with DWI | nADC | nFA | Number of Samples with PWI | nCBV | Number of Samples with MRSI | nNAA | nCho | CNI |
|---|---|---|---|---|---|---|---|---|---|
| All samples | 88 | 1.88 | 0.60 | 77 | 0.72 | 57 | 0.36 | 1.42 | 4.48 |
| All samples vs tumor score (TS) | 81 | 73 | 54 | ||||||
| TS = 0 | 5 | 1.47 | 0.72 | 5 | 1.06 | 4 | 0.43 | 0.87 | 1.69 |
| TS = 1 | 15 | 1.53 | 0.70 | 11 | 0.89 | 9 | 0.52 | 1.34 | 3.72 |
| TS = 2 | 38 | 1.86 | 0.64 | 38 | 0.71 | 24 | 0.44 | 1.69 | 5.45 |
| TS = 3 | 23 | 2.17 | 0.47 | 19 | 0.53 | 17 | 0.21 | 1.33 | 4.01 |
| F value | 6.62 | 3.97 | 3.99 | 3.55 | 2.98 | ns | |||
| P value | .0005⁎ | .012⁎ | .011⁎ | .021 | .048 | ||||
| IDH-mutant astrocytoma | 46 | 39 | 30 | ||||||
| TS = 0 | 0 | 0 | 0 | ||||||
| TS = 1 | 9 | 1.58 | 0.78 | 5 | 0.97 | 6 | 0.49 | 1.47 | 4.37 |
| TS = 2 | 24 | 1.88 | 0.58 | 25 | 0.62 | 14 | 0.40 | 1.64 | 5.38 |
| TS = 3 | 13 | 2.35 | 0.43 | 9 | 0.52 | 10 | 0.25 | 1.06 | 3.35 |
| F value | 8.24 | 5.93 | 3.04 | ns | ns | ns | |||
| P value | .0009⁎ | .005⁎ | .06 | ||||||
| 1p/19q co-deleted oligodendroglioma | 25 | 22 | 12 | ||||||
| TS = 0 | 3 | 1.43 | 0.76 | 5 | 1.06 | 3 | 0.44 | 0.90 | 2.07 |
| TS = 1 | 5 | 1.45 | 0.60 | 4 | 0.95 | 2 | 0.63 | 1.28 | 3.16 |
| TS = 2 | 9 | 1.66 | 0.79 | 6 | 0.90 | 3 | 0.72 | 2.49 | 6.77 |
| TS = 3 | 8 | 1.98 | 0.53 | 7 | 0.60 | 4 | 0.21 | 1.39 | 5.70 |
| F value | 3.67 | ns | ns | ns | ns | ns | |||
| P value | .029 |
Imaging values are the group averages of the median imaging values.
Significant after Benjamini-Hochberg false discovery rate correction for multiple comparisons.
Table 2.
Imaging Results from Tissue Sample ROIs between and within 1p/19q Co-Deleted Oligodendroglioma and IDH-Mutant Astrocytoma Subgroups
| Group | Number of Samples with DWI | nADC | nFA | Number of Samples with PWI | nCBV | Number of Samples with MRSI | nNAA | nCho | CNI |
|---|---|---|---|---|---|---|---|---|---|
| Grade II | 52 | 1.73 | .63 | 44 | .82 | 37 | .37 | 1.43 | 4.40 |
| Grade III | 36 | 2.11 | .54 | 33 | .59 | 20 | .34 | 1.39 | 4.64 |
| F value | 10.73 | ns | 4.97 | ns | ns | ns | |||
| P value | .002⁎ | .031 | |||||||
| IDH-mutant astrocytoma | |||||||||
| Grade II | 22 | 1.75 | 0.64 | 17 | 0.68 | 17 | 0.32 | 1.30 | 4.19 |
| Grade III | 28 | 2.15 | 0.50 | 24 | 0.59 | 15 | 0.35 | 1.47 | 4.71 |
| F value | 8.80 | 4.34 | ns | ns | ns | ns | |||
| P value | .005⁎ | .043 | |||||||
| IDH-mutant astrocytoma | 50 | 1.98 | 0.56 | 41 | 0.63 | 32 | 0.33 | 1.38 | 4.44 |
| 1p/19q co-deleted oligodendroglioma | 28 | 1.70 | 0.64 | 23 | 0.89 | 13 | 0.48 | 1.60 | 4.83 |
| F value | 5.9 | ns | 4.08 | ns | ns | ns | |||
| P value | .018⁎ | .022 |
Imaging result values are the group averages of the median imaging values.
Significant after Benjamini-Hochberg false discovery rate correction for multiple comparisons.
Table 3.
Imaging Results from T2 Lesion ROIs between and within 1p/19q Co-Deleted Oligodendroglioma and IDH-Mutant Astrocytoma Subgroups
| Group | Comparison | Diffusion Parameters | MRSI Parameters |
|---|---|---|---|
| All patients | Grade II vs grade III | 71 vs 39 patients nADC: 10% (P = .039) nFA: 90% (P = .023) |
60 vs 33 patients CNI (P = .015⁎) |
| IDH-mutant astrocytoma | Grade II vs grade III | 20 vs 36 patients ns |
23 vs 23 patients nCHO (P = .041) |
| IDH-mutant astrocytoma vs 1p/19q co-deleted oligodendroglioma |
All patients | 56 vs 29 patients nADC: 10%, median, 90% (P = .0006⁎, .0001⁎, .0001⁎) nFA: 10%, median (P = .006⁎, .015⁎) |
46 vs 28 patients CNI, nCHO, CCRI (P = .01, .032, .012) nLAC (P = .015) |
| Grade II only | 30 vs 29 patients nADC: 10%, median, 90% (P = .045, .0001⁎, .0001⁎) nFA: 10%, median (P = .029, .046) |
23 vs 28 patients ns |
Significant after Benjamini-Hochberg false discovery rate correction for multiple comparisons.
Results
Patient Characteristics
Of the 135 patients with lower-grade glioma enrolled in this study, 89 were characterized as grade II and 46 as grade III. The median age of the population was 41 (interquartile range = 31-49 years), and 64 (47%) were female. Evaluation of postoperative imaging indicated that 44 (33%) patients received gross total resections, 88 (65%) had subtotal resections, and 3 (2%) had biopsies only. There were 63 patients with IDH-mutant astrocytoma, 39 patients with IDH-mutant, 1p/19q co-deleted oligodendroglioma, 26 astrocytomas that were IDH negative by immunohistochemistry, and 7 astrocytomas for whom the molecular status was unknown. All patients with 1p/19q co-deleted oligodendroglioma were classified as being WHO grade II. Confirmed locations for image-guided tissue samples on presurgical MRIs were acquired for 88 samples from 45 patients. Table 1 indicates the number of samples from patients in each category of the WHO 2016 classification.
Imaging Values in Tissue Sample ROIs Versus Tumor Score
There were 81 samples with tumor scores and diffusion parameter values, 73 with tumor scores and perfusion parameter values, and 54 with tumor scores and spectroscopic parameter values (see Table 1). The overall median nADC, nCho, and CNI were higher (1.88, 1.42, and 4.48) than normal brain, with the nFA, nCBV, and nFAA being lower (0.60, 0.72, and 0.36) than normal brain. In all cases, the majority of samples had tumor scores greater than zero (76/81 = 94%, 68/73 = 93%, and 49/54 = 93%, respectively).
When all the relevant samples were evaluated based upon their tumor score, there were consistent, significant increases in nADC (median values from 1.53 to 2.17 for tumor scores of 1 to 3) and decreases in nFA (median from 0.70 to 0.47 for tumor scores of 1 to 3) (see Table 1). The same patterns in median nADC and nFA were observed when the analysis was restricted to the subset of samples with IDH-mutant astrocytoma (see Figure 2A, Table 1). For the subset of samples from 1p19q co-deleted oligodendroglioma, the median nADC values also increased with tumor score but with lower values (1.45 to 1.98 for tumor scores of 1 to 3). In this case, there was no clear relationship between the median nFA and tumor score (see Figure 2A, Table 1).
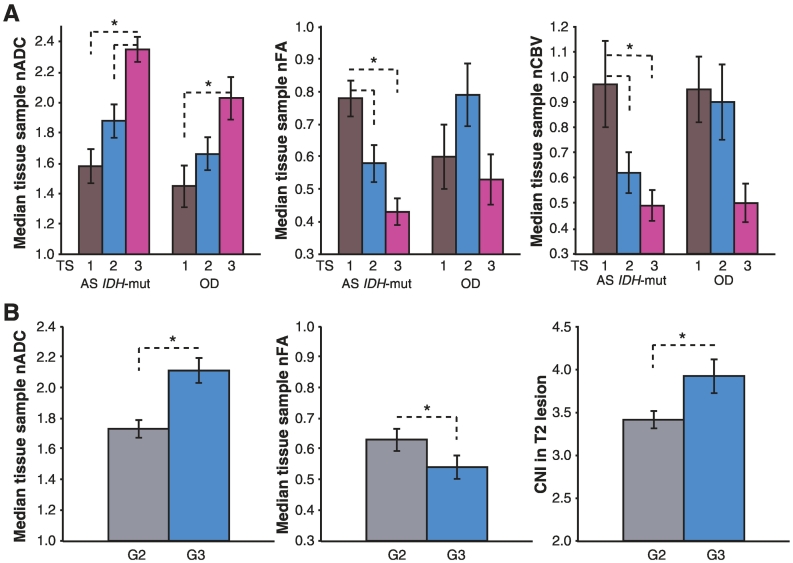
Figure 2.
(A) Median nADC, nFA, and nCBV values within tissue sample ROIs with tumor scores of 1, 2, and 3 (indicating the level of tumor cellularity within the sample) for IDH mutant astrocytoma (AS IDH+) and 1p/19q co-deleted oligodendroglioma (OD) LrGG patients. (B) Median nADC, nFA within tissue sample ROIs, and CNI values within T2/CNI >2 lesion ROIs for grade II and grade III LrGG patients.
For the samples with positive tumor score that had perfusion values, the median nCBV significantly decreased with increasing score (0.89 to 0.53 for tumor scores of 1 to 3) (see Table 1). The same range of values and trends was observed when the analysis was restricted within each molecular subgroup. Within samples with IDH-mutant astrocytoma, nCBV showed a trend towards lower values with higher tumor scores (P = .06), with significantly lower nCBV values for tumor scores of 2 and 3 than 1 (see Figure 2A). Within samples with 1p19q co-deleted oligodendroglioma, the relationship between tumor score and nCBV did not reach significance.
For the spectroscopic data, there was a significant decrease in nNAA with tumor score (0.52 to 0.21 for tumor scores of 1 to 3) (see Table 1). The nCho and CNI were both elevated in samples with positive tumor score, but the highest nCho and CNI values were in samples with tumor scores of 2. A similar pattern was observed in subgroups of samples based upon the WHO 2016 criteria, but the relationships to tumor score were not significant.
Imaging Values in Tissue Sample ROIs Versus Grade and WHO 2016 Subgroup
For lesions characterized as grade II, there were 52 samples with diffusion, 44 samples with perfusion, and 37 samples with spectroscopic values (see Table 2). The corresponding numbers of samples from grade III lesions were 36, 33, and 20, respectively. When values from all relevant samples were considered, the only parameters that were significantly different between grades were the median nADC (1.73 for grade II and 2.11 for grade III, P = .002) and median nCBV (0.82 for grade II and 0.59 for grade III, P = .031). When the analysis was restricted to the IDH-mutant astrocytomas, only the median nADC (1.75 vs 2.15, P = .005) and the median nFA (0.64 vs 0.50, P = .043) were significantly different between samples from grade II and III lesions (see Figure 2B). When all samples from IDH-mutant astrocytoma were compared with those from 1p19q co-deleted oligodendroglioma, the only parameters that were significantly different were the median nADC (1.98 vs 1.70, P = .018, see Figure 3A) and the nCBV (0.63 vs 0.89, P = .022).
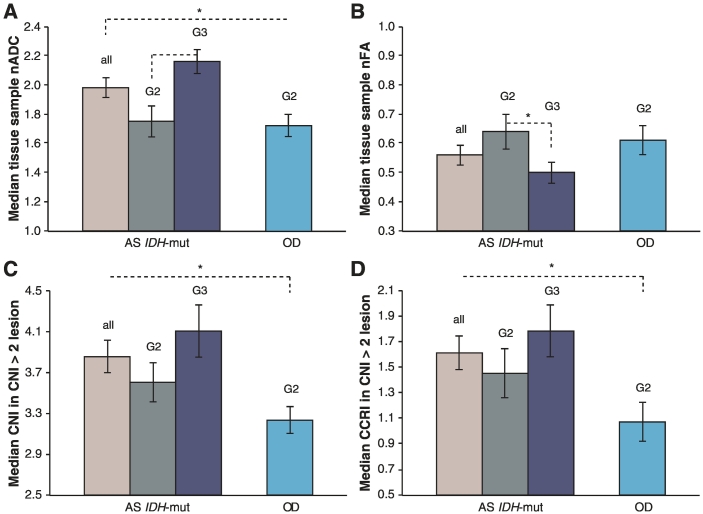
Figure 3.
(A) Median nADC within tissue sample ROIs, (B) nFA within tissue sample ROIs, (C) median CNI within T2/CNI >2 lesion ROIs, and (D) median CCRI values within T2/CNI >2 lesion ROIs for IDH mutant astrocytoma (AS IDH+) and 1p/19q co-deleted oligodendroglioma (OD) LrGG patients.
Imaging Values in Lesion Level ROIs Versus Grade and WHO 2016 Subgroup
While none of the perfusion parameters within the T2 lesion ROIs showed significant differences between grades and WHO 2016 subgroups, there were several findings for diffusion and spectroscopic parameters (see Table 3). When comparing values for grade II lesions versus grade III lesions, there were significant differences detected in the 10th percentile nADC (P = .039), the 90th percentile nFA (P = .023), and the median CNI (3.41 vs 3.92, P = .015, see Figure 2B). Within the IDH-mutant astrocytomas, there was a significant difference in the median nCHO between grade II and grade III lesions but not in summary metrics from histograms of nADC and nFA.
When comparing between WHO 2016 subgroups, there were significant differences in the 10th percentile, median, and 90th percentile nADC and the 10th percentile and median nFA, as well as in the median CNI (see Figure 3C), nCho, CCrI (see Figure 3D), and nLac between the IDH-mutant astrocytomas and the 1p19q co-deleted oligodendroglioma. If the analysis was restricted to the subset of IDH-mutant astrocytomas that were characterized as grade II, the nADC and nFA metrics showed similar results, but there were no significant findings for the spectroscopic parameters.
Discussion
The goal of this study was to provide information that could be used in interpreting diffusion, perfusion, and spectroscopic imaging data from patients with newly diagnosed, nonenhancing LrGG. Determining how parameters derived from these advanced imaging modalities vary between and within the WHO 2016 molecular subgroups is critical for evaluating prognosis and resolving ambiguities associated with conventional anatomic images. By obtaining imaging data at the presurgical examination and acquiring image-guided tissue samples from a subset of the patients, we were able to make direct comparisons of in vivo parameters with molecular and histological characteristics. Our findings have highlighted the complementary nature of these advanced imaging modalities and detected significant differences in parameter values between molecular subgroups of LrGG at both the tissue sample and whole lesion level.
The parameters that showed the largest number of significant findings and emphasized the potential for using DWI in evaluating LrGG were the nADC and nFA. Of particular interest in this case was that higher nADC and lower nFA were associated with higher tumor scores and grade III rather than grade II histology. This is in contrast to previous reports, which indicated that lower ADC was associated with worse clinical outcomes for patients with GBM and could be interpreted as a surrogate marker of tumor cellularity [11], [13]. Our results suggest that in these newly diagnosed, nonenhancing LrGG, the values of nADC and nFA are dominated by tumor infiltration causing disruption of neuronal, axonal, and glia microstructure rather than by changes due to abnormal cellularity [29]. The association of higher nADC with lesions that may have a worse prognosis is further supported by the differences between the WHO 2016 subgroups, for which IDH-mutant astrocytomas have higher nADC than the 1p19q co-deleted oligodendrogliomas, both at the tissue sample and at the whole lesion level. The values observed in this analysis were consistent with our previously published data [30], which were not specific to molecular subgroups but did separate results based upon histological assessments of astrocytoma versus oligodendroglioma. Of note is that we previously observed lower nADC in regions of contrast enhancement in newly diagnosed grade III glioma relative to nonenhancing regions [31], and in a study of recurrent grade II glioma, where a substantial number of the lesions had regions of enhancement, we found that lower nADC was associated with malignant progression [32]. Taken together, this suggests that the nADC values evolve as a result of treatment and tumor progression so that cellularity may ultimately become a more dominant factor.
The analysis of the PWI data from our study indicated that the nCBV values in grade II and III glioma were less than in normal-appearing white matter and decreased in samples with higher tumor score and grade. As was the case for parameters derived from the DWI data, these results are different from those obtained in prior studies of patients with GBM, which indicate that lesions with higher nCBV have a worse prognosis [16]. It seems likely that these newly diagnosed, nonenhancing LrGG tumors have not yet caused a significant change in angiogenesis and that the values observed are therefore more reflective of the disruption of normal microvasculature. The observation that the nCBV is significantly lower in tissue samples from IDH-mutant astrocytomas than in 1p19q co-deleted oligodendroglioma is consistent with our prior study that compared values in grade II lesions based upon histological subtypes [30].
In contrast to results obtained from the analysis of parameters from DWI and PWI data, the pattern of relationships between MRSI parameters, molecular characteristics, and pathology for this study was consistent with those reported for patients with GBM [16], [33], [34]. Tissue samples with positive tumor scores had higher nCho and lower nNAA than normal brain, and the CNI values in the intersection of the T2 and metabolic lesions were significantly higher for grade III versus grade II. Of particular interest in evaluating the WHO 2016 subgroups is that the median CNI, CCRI, nCho, and nLac were all higher in lesion ROIs for patients with IDH-mutant astrocytomas compared with those with 1p19q co-deleted oligodendroglioma. Given that there were no significant differences in metabolite levels when the analysis was restricted to comparing the results from grade II lesions and given the results from our previous analysis of grade II lesions that was based upon histological subtypes [30], it appears that this may be accounted for by the differences in metabolite levels between grades.
Because treatment decisions can be based on the histopathologic analysis of surgical biopsies, the selection of intratumoral targets to biopsy can profoundly impact patient care. Our results indicate that regions with increased nCHO and decreased nNAA have high tumor cellularity, whether they are in enhancing or nonenhancing regions and across all grades of glioma [16], [33], [34]. Color overlays of the CNI are particularly helpful in highlighting regions with abnormal levels of these metabolites and, because they represent a combined z-score, are easier to interpret than maps of relative metabolite levels [25]. For nonenhancing lesions that are more likely to be LrGG, regions with higher nADC and lower nFA may provide samples with a higher fraction of tumor cells. The use of ADC color-coded histogram maps [29], [33] may be helpful for targeting tissue samples from regions with both high and low values in order to get a complete picture of the tumor and potentially target tumor regions with different molecular characteristics.
Other applications that are particularly critical for the ongoing management of patients with glioma are assessing the spatial extent of residual tumor and monitoring changes in tumor properties that are associated with response to therapy and tumor progression. The results of this and our prior studies of LrGG [29,39,33,34] indicate that there are differences in the values of nADC, nCBV, and metabolic parameters between WHO 2016 subgroups, which should be considered in interpreting findings from serial examinations of individual patients. Whether changes in these in vivo imaging parameters provide additional prognostic information is still under investigation, as this requires a long-term study of patient outcomes.
It is important to note limitations of the current study. One is that the size of the tissue sample ROIs (5 mm) is considerably larger than the size of the actual tissue samples, and despite efforts by the surgeons to collect tissue samples early in the resection, the location accuracy can be further impacted by brain shift during surgery. Another is that, because the sample sizes for some of the molecular subgroups of LrGG was small, the comparative analysis was restricted to the two most common types (IDH-mutant astrocytomas and 1p19q co-deleted oligodendroglioma). Future studies will be strengthened by the routine acquisition of molecular characteristics for WHO 2016 classification by using sequencing to confirm IDH wild-type status and through the evaluation of additional molecular characteristics, such as ATRX loss and telomerase reverse transcriptase promoter mutations, which are emerging as important biomarkers for LrGG [35], [36].
In conclusion, this study achieved its goal by highlighting differences in advanced imaging parameters between and within molecular subgroups of patients with newly diagnosed, nonenhancing LrGG. Pathological analysis confirmed the interpretation of spectroscopy, diffusion, and perfusion parameters relative to tumor score and histological characteristics. While the spectroscopy results were consistent with the metabolic pattern of malignant features observed in GBM, the nADC was higher and nCBV was lower in regions with higher tumor score and histologic grade, which are the reverse of patterns observed in GBM. The findings of the study may be important for directing tissue sampling to more malignant regions of the T2 lesion, for making decisions about treating residual disease, and for interpreting the results from serial MR examinations.
Footnotes
Funding Sources: This work was supported by the National Institutes of Health (NIH R01 CA159869 and NIH P50 CA097257).
References
- 1.Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol. 2016;131:803–820. doi: 10.1007/s00401-016-1545-1. [DOI] [PubMed] [Google Scholar]
- 2.Laack NN, Sarkaria JN, Buckner JC. Radiation Therapy Oncology Group 9802: Controversy or Consensus in the Treatment of Newly Diagnosed Low-Grade Glioma? Semin Radiat Oncol. 2015;25:197–202. doi: 10.1016/j.semradonc.2015.02.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Field KM, Rosenthal MA, Khasraw M, Sawkins K, Nowak AK. Evolving management of low grade glioma: No consensus amongst treating clinicians. J Clin Neurosci. 2016;23:81–87. doi: 10.1016/j.jocn.2015.05.038. [DOI] [PubMed] [Google Scholar]
- 4.Le Rhun E, Taillibert S, Chamberlain MC. Current Management of Adult Diffuse Infiltrative Low Grade Gliomas. Curr Neurol Neurosci Rep. 2016;16:15. doi: 10.1007/s11910-015-0615-4. [DOI] [PubMed] [Google Scholar]
- 5.Park I, Chen AP, Zierhut ML, Ozturk-Isik E, Vigneron DB, Nelson SJ. Implementation of 3T lactate-edited 3D 1H MR spectroscopic imaging with flyback echo-planar readout for gliomas patients. Ann Biomed Eng. 2011;39:193–204. doi: 10.1007/s10439-010-0128-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Jenkinson M, Bannuster P, Brady M, Smith S. Improved optimization for the robust and accurate linear registration and motion correction of brain images. NeuroImage. 2002;17:825–841. doi: 10.1016/s1053-8119(02)91132-8. [DOI] [PubMed] [Google Scholar]
- 7.Duarte-Carvajalino J, Sapiro G, Harel N, Lenglet C. A framework for linear and non-linear registration of diffusion-weighted MRIs using angular interpolation. Front Neurosci. 2013;7:1–15. doi: 10.3389/fnins.2013.00041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Keles GE, Lamborn KR, Berger MS. Coregistration accuracy and detec- tion of brain shift using intraoperative sononavigation during resection of hemispheric tumors. Neurosurgery. 2003;53:556–562. doi: 10.1227/01.neu.0000080949.44837.4c. [DOI] [PubMed] [Google Scholar]
- 9.Paraskevopoulos D, Unterberg A, Metzner R, Dreyhaupt J, Eggers G, Wirtz CR. Comparative study of application accuracy of two frameless neuronavigation systems: experimental error assessment quantifying registration methods and clinically influencing factors. Neurosurg Rev. 2010;34:217–228. doi: 10.1007/s10143-010-0302-5. [DOI] [PubMed] [Google Scholar]
- 10.Basser PJ, Pierpaoli C. Microstructural and physiological features of tissues elucidated by quantitative-diffusion-tensor MRI. J Magn Reson. 1996;111:209–219. doi: 10.1006/jmrb.1996.0086. [DOI] [PubMed] [Google Scholar]
- 11.Wen Q, Jalilian L, Lupo JM, Molinaro AM, Chang SM, Clarke J, Prados M, Nelson SJ. Comparison of ADC metrics and their association with outcome for patients with newly diagnosed glioblastoma being treated with radiation therapy, temozolomide, erlotinib and bevacizumab. J Neuro-Oncol. 2015;121:331–339. doi: 10.1007/s11060-014-1636-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hilario A, Sepulveda JM, Perez-Nuñez A, Salvador E, Millan JM, Hernandez-Lain A, Rodriguez-Gonzalez V, Lagares A, Ramos A. A Prognostic Model Based on Preoperative MRI Predicts Overall Survival in Patients with Diffuse Gliomas. Am J Neuroradiol. 2014;35:1096–1102. doi: 10.3174/ajnr.A3837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Romano A, Calabria LF, Tavanti F, Minniti G, Rossi-Espagnet MC, Coppola V, Pugliese S, Guida D, Francione G, Colonnese C. Apparent diffusion coefficient obtained by magnetic resonance imaging as a prognostic marker in glioblastomas: correlation with MGMT promoter methylation status. Eur Radiol. 2013;23:513–520. doi: 10.1007/s00330-012-2601-4. [DOI] [PubMed] [Google Scholar]
- 14.Zulfiqar M, Yousem D, Lai H. ADC Values and Prognosis of Malignant Astrocytomas: Does Lower ADC Predict Worse Prognosis Independent of Grade of Tumor? – A Meta-Analysis. Am J Roentgenol. 2013;200:624–629. doi: 10.2214/AJR.12.8679. [DOI] [PubMed] [Google Scholar]
- 15.Lee MC, Cha S, Chang SM, Nelson SJ. Dynamic susceptibility contrast perfusion imaging of radiation effects in normal-appearing brain tissue: changes in the first-pass and recirculation phases. J Magn Reson Imaging. 2005;21:683–693. doi: 10.1002/jmri.20298. [DOI] [PubMed] [Google Scholar]
- 16.Li Y, Lupo JM, Polley MY, Crane JC, Bian W, Cha S, Chang S, Nelson SJ. Serial analysis of imaging parameters in patients with newly diagnosed glioblastoma multiforme. Neuro-Oncology. 2011;13:546–557. doi: 10.1093/neuonc/noq194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Essock-Burns E, Phillips JJ, Molinaro AM, Lupo JM, Cha S, Chang SM, Nelson SJ. Comparison of DSC-MRI post-processing techniques in predicting microvascular histopathology in patients newly diagnosed with GBM. J Magn Reson Imaging. 2013;38:388–400. doi: 10.1002/jmri.23982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Essock-Burns E, Lupo JM, Cha S, Polley M-Y, Butowski NA, Chang SM, Nelson SJ. Assessment of perfusion MRI-derived parameters in evaluating and predicting response to antiangiogenic therapy in patients with newly diagnosed glioblastoma. Neuro-Oncology. 2011;13:119–131. doi: 10.1093/neuonc/noq143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Barajas RF, Phillips JJ, Parvataneni R, Molinaro A, Essock-Burns E, Bourne G, Parsa AT, Aghi MK, McDermott MW, Berger MS. Regional variation in histopathologic features of tumor specimens from treatment-naïve glioblastoma correlates with anatomic and physiologic MR imaging. Neuro-Oncology. 2012;14:942–954. doi: 10.1093/neuonc/nos128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Nelson SJ, Kadambi AK, Park I, Li Y, Crane J, Olson M, Molinaro A, Roy R, Butowski N, Cha S. Association of early changes in 1H MRSI parameters with survival for patients with newly diagnosed glioblastoma receiving a multimodality treatment regimen. Neuro-Oncology. 2017:430–439. doi: 10.1093/neuonc/now159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Li Y, Osorio JA, Ozturk-Isik E, Chen AP, Xu D, Crane JC, Cha S, Chang S, Berger MS, Vigneron DB. Considerations in applying 3D PRESS H-1 brain MRSI with an eight-channel phased-array coil at 3 T. Magn Reson Imaging. 2006;24:1295–1302. doi: 10.1016/j.mri.2006.07.012. [DOI] [PubMed] [Google Scholar]
- 22.Crane JC, Olson MP, Nelson SJ. SIVIC: Open-Source, Standards-Based Software for DICOM MR Spectroscopy Workflows. Int J Biomed Imaging. 2013;2013:169526. doi: 10.1155/2013/169526. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Cunningham CH, Vigneron DB, Chen AP, Xu D, Nelson SJ, Hurd RE, Kelley DA, Pauly JM. Design of flyback echo-planar readout gradients for magnetic resonance spectroscopic imaging. Magn Reson Med. 2005;54:1286–1289. doi: 10.1002/mrm.20663. [DOI] [PubMed] [Google Scholar]
- 24.Li Y, Lupo JM, Paravataneni R, Lamborn KR, Cha S, Chang SM, Nelson SJ. Suvival analysis in patients with newly diagnosed glioblastoma using pre- and postradiotherapy MR spectroscopic imaging. Neuro-Oncology. 2013;15:607–617. doi: 10.1093/neuonc/nos334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.McKnight TR, Noworolski SM, Vigneron DB, Nelson SJ. An automated technique for the quantitative assessment of 3D-MRSI data from patients with glioma. J Magn Reson Imaging. 2001;13:167–177. doi: 10.1002/1522-2586(200102)13:2<167::aid-jmri1026>3.0.co;2-k. [DOI] [PubMed] [Google Scholar]
- 26.Elkhaled A, Jalbert LE, Phillips JJ, Yoshihara HAI, Parvataneni R, Srinivasan R, Bourne G, Berger MS, Chang SM, Cha S. Magnetic resonance of 2-hydroxyglutarate in IDH1-mutated low-grade gliomas. Sci Transl Med. 2012;4 doi: 10.1126/scitranslmed.3002796. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Pekmezci M, Rice T, Molinaro AM, Walsh KM, Decker PA, Hansen H, Sicotte H, Kollmeyer TM, McCoy LS, Sarkar G. Adult Infiltrating Gliomas with WHO 2016 Integrated Diagnosis: Additional Prognostic Roles of ATRX and TERT. Acta Neuropathol. 2017;133:1001–1016. doi: 10.1007/s00401-017-1690-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc. 1995;57:289–300. [Google Scholar]
- 29.Khayal IS, Vandenberg SR, Smith KJ, Cloyd CP, Chang SM, Cha S, Nelson SJ. McKnight TR. MRI apparent diffusion coefficient reflects histopathologic subtype, axonal disruption, and tumor fraction in diffuse-type grade II gliomas. Neuro-Oncology. 2011;13:1192–1201. doi: 10.1093/neuonc/nor122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Bian W, Khayal IS, Lupo JM, McGue C, Vandenberg S, Lamborn KR, Chang SM, Cha S, Nelson SJ. Multiparametric characterization of grade 2 glioma subtypes using magnetic resonance spectroscopic, perfusion, and diffusion imaging. Transl Oncol. 2009;2:271–280. doi: 10.1593/tlo.09178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Ozturk-Isik E, Pirzkall A, Lamborn KR, Cha S, Chang SM, Nelson SJ. Spatial characteristics of newly diagnosed grade 3 glioma assessed by magnetic resonance metabolic and diffusion tensor imaging. Transl Oncol. 2012;5:10–18. doi: 10.1593/tlo.11208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Jalbert LE, Neill E, Phillips JJ, Lupo JM, Olson MP, Molinaro AM, Berger MS, Chang SM, Nelson SJ. Magnetic resonance analysis of malignant transformation in recurrent glioma. Neuro-Oncology. 2016;18:1169–1179. doi: 10.1093/neuonc/now008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Neill E, Luks T, Dayal M, Phillips JJ, Perry A, Jalbert LE, Cha S, Molinaro A, Chang SM, Nelson SJ. Quantitative Multi-modal MR Imaging as a Non-Invasive Prognostic Tool for Patients with Recurrent Low-Grade Glioma. J Neuro-Oncol. 2017;132:171–179. doi: 10.1007/s11060-016-2355-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.McKnight TR, Lamborn KR, Love TD, Berger MS, Chang S, Dillon WP, Bollen A, Nelson SJ. Correlation of magnetic resonance spectroscopic and growth characteristics within Grades II and III gliomas. J Neurosurg. 2007;106:660–666. doi: 10.3171/jns.2007.106.4.660. [DOI] [PubMed] [Google Scholar]
- 35.Cancer Genome Atlas Research Network Comprehensive, Integrative Genomic Analysis of Diffuse Lower-Grade Gliomas. N Engl J Med. 2015;372:2481–2498. doi: 10.1056/NEJMoa1402121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Eckel-Passow JE, Lachance DH, Molinaro AM, Walsh KM, Decker PA, Sicotte H, Pekmezci M, Rice T, Kosel ML, Smirnov IV. Glioma Groups Based on 1p/19q, IDH, and TERT Promoter Mutations in Tumors. N Engl J Med. 2015;372:2499–2508. doi: 10.1056/NEJMoa1407279. [DOI] [PMC free article] [PubMed] [Google Scholar]