Abstract
Recent studies have investigated the epigenetic effects of environmental exposure to chemicals on human health. The associations of DNA methylation, environmental exposure and human diseases have been widely demonstrated. However, the use of gene methylation patterns as a predictive biomarker for exposure to environmental toxicants is relatively poorly understood. Here, we focused on low-molecular-weight saturated aliphatic aldehydes (LSAAs), which are important environmental risk factors in humans as major indoor air pollutants. Based on DNA methylation profiling in gene promoter regions, we analysed DNA methylation profiles following exposure of A549 cells to seven LSAAs (propanal, butanal, pentanal, hexanal, heptanal, octanal, and nonanal) to identify LSAA-characterized methylated sites and target genes, as well as to investigate whether exposure to LSAAs contributes to inducing of pulmonary toxicity. Additionally, by integrating DNA methylation and mRNA expression profile analyses, we identified core anti-correlated target genes. Gene ontology analysis of these target genes revealed several key biological processes. These findings suggest that alterations in DNA methylation by exposure to LSAAs provide novel epigenetic biomarkers for risk assessments. This DNA methylation-mRNA approach also reveals potential new mechanistic insights into the epigenetic actions of pulmonary toxicity.
Introduction
The field of epigenetics including DNA methylation, histone modification, and microRNAs is rapidly expanding, and these processes are associated with environmental exposure to pollutants. Recent studies have suggested that environmental pollutants can cause human diseases based on an epigenetic mechanism1. Epigenetics is essential for a wide range of biological processes including the regulation of gene expression, normal development, and cellular differentiation2. All epigenetic mechanisms play an important role in the pathogenesis of environmental diseases and cancers3,4.
Among the epigenetic mechanisms, there has been an increase in the number of DNA methylome studies investigating the effects of human health risk factors including environmental chemicals on disease. DNA methylation, which is the addition of methyl groups at the 5′ position on the pyrimidine ring of cytosine, is a crucial epigenetic modification of the genome1 and is important in regulating gene expression. Aberrant DNA methylation is associated with disease progression and human cancer5,6. To date, DNA methylation has been widely studied. Particularly, numerous studies demonstrating the influence of environmental toxicants on DNA methylation have been linked to human health7.
Various environmental toxicants such as air pollutants, toxic chemicals, and radiation that affect epigenetic processes result from the effects of acute or chronic exposure. Recent studies investigated whether multiple environmental exposure to polycyclic aromatic hydrocarbons, diesel exhaust particles, and particulate matter is a major risk factor for the development of environmental lung disease by epigenetic modifications3. However, the effects of exposure to major indoor air pollutants such as aldehydes on the respiratory system via DNA methylation remain unclear. Therefore, among environmental chemicals, we focused on low-molecular-weight saturated aliphatic aldehydes (LSAAs) for DNA methylome analysis. In previous studies, we investigated the genetic and epigenetic responses of LSAAs based on transcriptome and microRNA analyses8,9. Here, we performed epigenome-wide DNA methylome analysis in aldehyde-exposed A549 lung adenocarcinoma cells.
Aldehydes are categorized as volatile organic compounds, which are ubiquitous in indoor and outdoor air as common air pollutants10. They are emitted from diverse indoor sources such as carpets, floor coverings, paints, panels, and furniture11. Recognition of the health effects of indoor air pollutants has also increased. Although studies on the pulmonary effect of aldehydes have been conducted12, the data remain limited for several aldehydes such as formaldehyde and acetaldehyde. In this study, we investigated the pulmonary toxic effects of LSAAs including propanal, butanal, pentanal, hexanal, heptanal, octanal, and nonanal. To investigate the epigenetic DNA methylation biomarkers of aldehydes on the human respiratory system for health risk assessment, we performed DNA methylome analysis. Following DNA methylation profiling, we performed integrated analyses of the methylation and mRNA data in A549 cells exposed to aldehydes to identify correlations.
Taken together, these findings indicate the potential pulmonary toxic effects of aldehyde exposure. Such DNA methylation biomarkers also improve the understanding of the underlying mechanisms of epigenetic regulation associated with aldehyde exposure.
Results
Cytotoxicity of A549 cells exposed to aldehydes
To determine the optimal exposure concentrations, the cytotoxicities of seven aldehydes were evaluated in the MTT assay (Fig. S1). Based on the MTT assay data, the cell viability inhibitory concentrations (IC) of each aldehyde at 20% (IC20) were calculated and compared to those of the dimethyl sulfoxide (DMSO) vehicle control group. Each inhibitory concentration of the seven aldehydes is shown in Table 1.
Table 1.
The 20% cell viability inhibitory concentrations (IC20) of seven aldehydes compared to vehicle control (DMSO) sample.
| Aldehydes | IC20 (mM) |
|---|---|
| Propanal (C3) | 2.5 |
| Butanal (C4) | 4.6 |
| Pentanal (C5) | 1.7 |
| Hexanal (C6) | 0.8 |
| Heptanal (C7) | 0.6 |
| Octanal (C8) | 0.58 |
| Nonanal (C9) | 0.44 |
DNA methylation signatures following exposure to the seven aldehydes
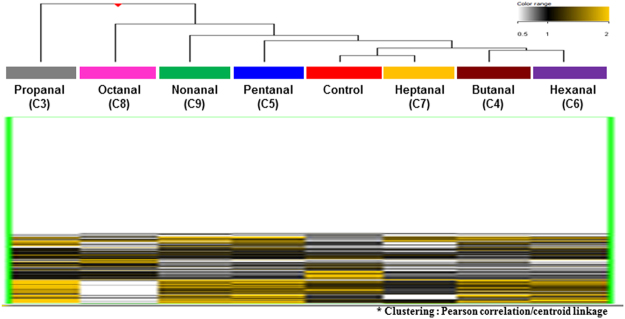
To investigate the genome-wide promoter DNA methylation expression profiles in A549 cells exposed to aldehydes, we performed a human 2 × 400 k DNA methylation microarray (Agilent Technologies, Santa Clara, CA, USA). The 414,043 methylation probes resulted in the DNA methylation of A549 cells based on unsupervised hierarchical clustering analysis. We identified the total DNA methylation expression patterns in A549 cells exposed to the aldehydes (Fig. 1), and then examined the overall DNA methylation levels to identify aldehyde-specific epigenetic DNA methylation markers.
Figure 1.
Total genome-wide profiling of DNA methylation of A549 cells exposed to the seven aldehydes compared to vehicle control group (DMSO). The heatmap shows the DNA methylation profiles of aldehydes exposed A549 cells based on hierarchical clustering (Yellow: hypermethylation; Black: hypomethylation).
Identification of characteristic aldehydes methylation markers
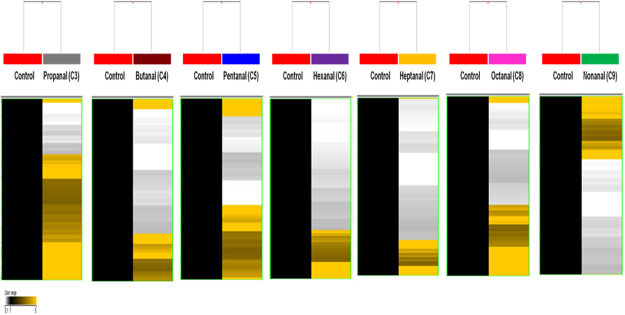
To determine whether exposure to aldehydes alters DNA methylation, we conducted DNA methylome analysis of seven aldehydes (C3–C9) using an in vitro model. First, we isolated methylated DNA using MeDIP from genomic DNA. Using this methylated DNA, a DNA methylation profile was acquired and analysed in the group exposed to the seven aldehydes and matched vehicle control group (Fig. 2, Table 2). In the propanal (C3) exposure group, we identified 4,345 genes that were differentially methylated (hyper-: 3,115 and hypo-: 1,230). In the butanal (C4) exposure group, 4,316 differentially methylated genes (hyper-: 1,584 and hypo-: 2,732) were identified. In the pentanal (C5) exposure group, 4,266 methylated genes (hyper-: 2,304 and hypo-: 1,962) were identified and 4,712 genes (hyper-: 1,572 and hypo-: 3,140) were methylated by hexanal (C6) exposure, and 3,598 target genes (hyper-: 897 and hypo-: 2,701) were identified following heptanal (C7) exposure. In the octanal (C8) exposure group, 3,822 methylated genes (hyper-: 1,611 and hypo-: 2,211) were identified, and 4,468 genes (hyper-: 1,898 and hypo-: 2,570) were methylated by nonanal (C9) exposure.
Figure 2.
DNA methylation signatures of differentiated seven aldehydes compared to vehicle control group (DMSO) (Yellow: hypermethylation; Black: hypomethylation).
Table 2.
Number of differentially methylated genes under exposure to seven aldehydes with 3.0-fold change and p-value < 0.05.
| Aldehydes | |||||||
|---|---|---|---|---|---|---|---|
| Propanal (C3) | Butanal (C4) | Pentanal (C5) | Hexanal (C6) | Heptanal (C7) | Octanal (C8) | Nonanal (C9) | |
| Methylated site | 9,741 | 10,405 | 9,931 | 12,164 | 9,023 | 8,891 | 11,316 |
| Promoter region | 4,896 | 4,957 | 4,859 | 5,608 | 4,170 | 4,263 | 5,268 |
| Regulated gene | 4,345 | 4,316 | 4,266 | 4,712 | 3,598 | 3,822 | 4,468 |
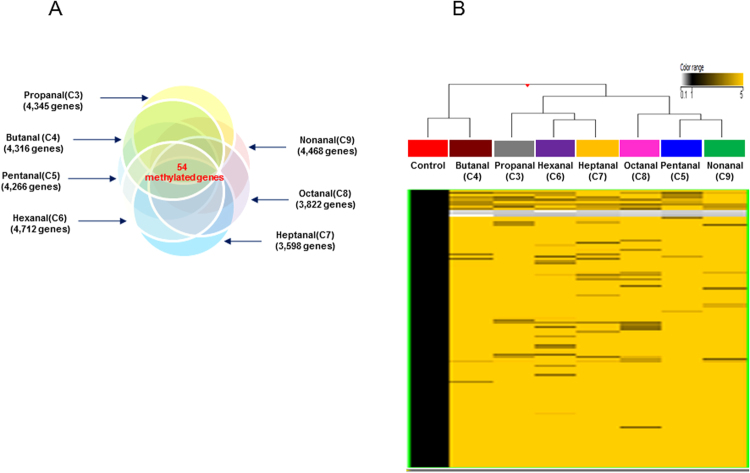
Among them, 54 genes showed common methylated expression patterns in the seven aldehydes exposure group (Fig. 3, Table 3). All methylated genes showed significant changes of 3.0-fold with p-values < 0.05.
Figure 3.
(A) Venn diagram shows the differentially methylated genes exposed under each aldehyde exposure. The intersection regions are the number of common differentially methylated genes following exposure to the seven aldehydes. (B) Hierarchical clustering of methylated genes that commonly altered DNA methylation in A549 cells exposed to the seven aldehydes with a fold-change ≥3.0-fold and p-value < 0.05 compared to the vehicle control group (DMSO) (Yellow: hypermethylation; Black: hypomethylation).
Table 3.
Commonly methylated genes of seven aldehydes exposed-A549 cell compared to vehicle control with 3.0-fold change and p-value < 0.05.
| Gene Symbol | Gene name | Methylation degree (Fold-change) | ||||||
|---|---|---|---|---|---|---|---|---|
| C3 | C4 | C5 | C6 | C7 | C8 | C9 | ||
| ZFHX2 | zinc finger homeobox protein 2 | 12.98 | 3.32 | 10.00 | 8.15 | 3.33 | 11.36 | 13.54 |
| TTTY5 | testis-specific transcript, Y-linked 5 | 55.06 | 59.56 | 334.00 | 95.77 | 17.98 | 10.80 | 80.09 |
| CCDC21;CEP85 | centrosomal protein 85 | 66.14 | 49.39 | 52.05 | 164.01 | 134.69 | 129.86 | 86.00 |
| GJA9-MYCBP | GJA9-MYCBP readthrough | 8.75 | 3.55 | 6.45 | 6.10 | 3.69 | 8.61 | 12.25 |
| PRKACB | protein kinase cAMP-activated catalytic subunit beta | 823.47 | 78.87 | 1169.04 | 362.87 | 560.59 | 1057.51 | 1139.15 |
| LPPR5 | lipid phosphate phosphatase-related protein type 5 isoform 2 | 7.93 | 5.74 | 6.59 | 7.30 | 8.36 | 7.68 | 7.68 |
| TDRKH | tudor and KH domain containing | 6.99 | 10.40 | 7.49 | 7.88 | 6.52 | 9.72 | 13.72 |
| OR10Z1 | olfactory receptor family 10 subfamily Z member 1 | 3.17 | 3.88 | 3.62 | 4.89 | 3.68 | 5.29 | 6.37 |
| NUF2 | NUF2, NDC80 kinetochore complex component | 8.99 | 27.81 | 37.15 | 15.14 | 4.05 | 5.39 | 6.90 |
| KCNF1 | potassium voltage-gated channel modifier subfamily F member 1 | 10.08 | 6.90 | 9.53 | 12.95 | 11.10 | 5.69 | 7.97 |
| C2orf61; STPG4 | sperm-tail PG-rich repeat containing 4 | 9.47 | 5.94 | 8.98 | 6.75 | 4.32 | 5.60 | 5.89 |
| BMP10 | bone morphogenetic protein 10 | 6.25 | 5.38 | 12.21 | 5.50 | 4.86 | 3.78 | 18.05 |
| REG1A | regenerating family member 1 alpha | 3.97 | 3.70 | 3.82 | 4.09 | 3.79 | 3.34 | 4.05 |
| MIR128-1 | microRNA 128-1 | 6.73 | 6.26 | 6.47 | 6.92 | 6.42 | 5.66 | 6.27 |
| UBXN4 | UBX domain protein 4 | 5.16 | 3.73 | 4.29 | 4.75 | 5.44 | 5.00 | 5.00 |
| WDR75 | WD repeat domain 75 | 6.02 | 5.01 | 6.58 | 6.02 | 6.12 | 14.02 | 7.04 |
| THUMPD3 | THUMP domain containing 3 | 5.45 | 9.95 | 5.95 | 5.45 | 5.54 | 5.78 | 3.27 |
| COX17 | COX17,cytochrome c oxidase copper chaperone | 22.46 | 18.67 | 24.53 | 22.46 | 22.82 | 15.61 | 28.73 |
| MFN1 | mitofusin 1 | 12.89 | 13.22 | 10.36 | 3.21 | 4.97 | 3.23 | 27.85 |
| NFXL1 | nuclear transcription factor, X-box binding like 1 | 16.26 | 4.96 | 8.71 | 7.16 | 11.07 | 7.78 | 5.77 |
| UGT2B10 | UDP glucuronosyltransferase family 2 member B10 | 9.04 | 8.45 | 13.47 | 3.98 | 6.16 | 11.61 | 12.51 |
| IGJ;JCHAIN | joining chain of multimeric IgA and IgM | 10.61 | 9.91 | 15.07 | 4.68 | 7.22 | 13.63 | 14.68 |
| HSD17B11 | hydroxysteroid 17-beta dehydrogenase 11 | 13.20 | 8.61 | 10.78 | 5.08 | 4.48 | 6.00 | 7.62 |
| SNCA | synuclein alpha | 12.42 | 11.60 | 11.19 | 5.47 | 8.45 | 15.95 | 17.18 |
| MIR583 | microRNA 583 | 15.37 | 18.05 | 12.78 | 14.15 | 16.21 | 5.69 | 14.89 |
| PCDHA9 | protocadherin alpha 9 | 300.99 | 40.15 | 5.39 | 76.53 | 254.52 | 56.58 | 64.89 |
| PCDHA10 | protocadherin alpha 10 | 105.50 | 98.53 | 149.77 | 46.49 | 229.99 | 135.49 | 145.95 |
| PHACTR1 | phosphatase and actin regulator 1 | 21.89 | 16.73 | 9.00 | 6.72 | 5.74 | 15.07 | 8.65 |
| HIST1H2BE | histone cluster 1 H2B family member e | 42.08 | 21.83 | 91.80 | 60.34 | 13.59 | 3.23 | 41.49 |
| TRIM27 | tripartite motif containing 27 | 3.50 | 13.33 | 47.39 | 7.21 | 9.71 | 6.19 | 12.52 |
| MRPS18A | mitochondrial ribosomal protein S18A | 292.96 | 127.71 | 246.55 | 239.77 | 219.35 | 91.89 | 370.17 |
| NMBR | neuromedin B receptor | 15.41 | 14.39 | 21.87 | 6.79 | 10.49 | 17.90 | 21.32 |
| LHFPL3 | lipoma HMGIC fusion partner-like 3 | 27.40 | 51.13 | 17.34 | 61.21 | 16.23 | 25.98 | 19.18 |
| PEX2 | peroxisomal biogenesis factor 2 | 101.62 | 124.05 | 115.75 | 156.58 | 117.68 | 169.28 | 109.65 |
| SPINK4 | serine peptidase inhibitor, Kazal type 4 | 13.95 | 12.61 | 6.96 | 10.09 | 7.85 | 5.19 | 8.88 |
| SLK | STE20 like kinase | 3.89 | 6.52 | 25.12 | 4.00 | 3.71 | 3.27 | 35.02 |
| PIK3C2A | phosphatidylinositol-4-phosphate 3-kinase catalytic subunit type 2 alpha | 47.86 | 23.32 | 31.00 | 13.92 | 7.86 | 39.95 | 54.35 |
| CD163L1 | CD163 molecule like 1 | 8.33 | 16.73 | 25.48 | 3.28 | 7.12 | 9.91 | 14.92 |
| PRH1-PRR4 | PRH1-PRR4 readthrough | 8.49 | 9.50 | 24.44 | 3.64 | 8.42 | 6.82 | 16.24 |
| CCNA1 | cyclin A1 | 15.92 | 9.51 | 13.60 | 5.87 | 3.17 | 12.83 | 15.04 |
| CCNB1IP1 | cyclin B1 interacting protein 1 | 3.45 | 16.63 | 16.41 | 3.20 | 9.03 | 9.13 | 11.86 |
| C14orf104; DNAAF2 | dynein axonemal assembly factor 2 | 12.81 | 8.80 | 7.24 | 4.70 | 3.62 | 3.95 | 5.63 |
| C14orf166B; LRRC74A | leucine rich repeat containing 74 A | 4.87 | 3.74 | 6.05 | 4.08 | 4.38 | 3.21 | 4.33 |
| CSNK1A1P1 | casein kinase 1 alpha 1 pseudogene 1 | 11.74 | 5.67 | 8.68 | 5.57 | 7.31 | 14.50 | 6.04 |
| KIF7 | kinesin family member 7 | 11.73 | 14.32 | 3.39 | 18.08 | 6.31 | 10.21 | 12.66 |
| CHD3 | chromodomain helicase DNA binding protein 3 | 28.65 | 23.21 | 35.98 | 15.25 | 14.61 | 15.85 | 10.57 |
| PDE4A | phosphodiesterase 4A | 7.78 | 11.77 | 7.15 | 5.46 | 9.41 | 8.80 | 8.16 |
| ZNF525 | zinc finger protein 525 | 11.18 | 7.40 | 24.49 | 4.92 | 7.61 | 17.13 | 15.78 |
| C20orf191; NCOR1P1 | nuclear receptor corepressor 1 pseudogene 1 | 8.70 | 10.62 | 9.91 | 13.40 | 10.07 | 14.49 | 16.16 |
| MYT1 | myelin transcription factor 1 | 16.18 | 7.61 | 13.72 | 5.89 | 3.82 | 3.87 | 6.82 |
| AIRE | autoimmune regulator | 4.90 | 4.56 | 19.72 | 5.04 | 4.68 | 4.12 | 4.57 |
| ACOT9 | acyl-CoA thioesterase 9 | 15.50 | 12.40 | 20.91 | 10.68 | 28.30 | 24.78 | 18.05 |
| RPA4 | replication protein A4 | 245.44 | 74.49 | 183.14 | 399.31 | 242.59 | 133.71 | 106.36 |
| C9orf57 | chromosome 9 open reading frame 57 | 0.29 | 0.23 | 0.28 | 0.08 | 0.26 | 0.23 | 0.23 |
Integrated analysis of methylated DNA and mRNA expression profiles
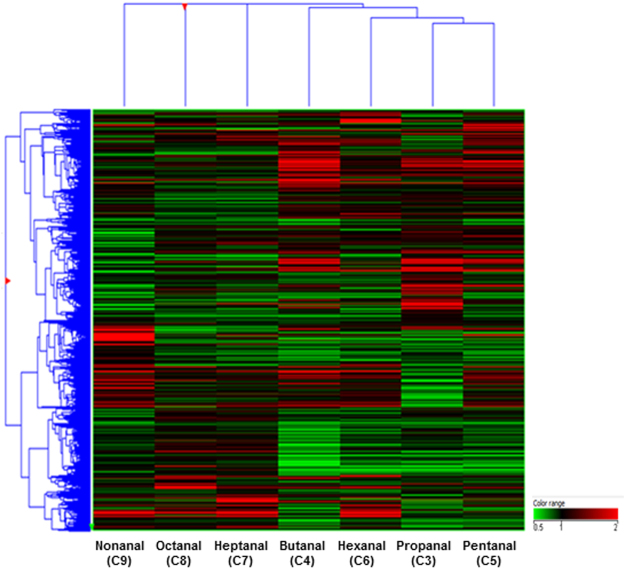
We also conducted integrated analysis of DNA methylation and mRNA expression. First, we conducted gene expression profiling of cells exposed to the seven aldehydes to identify differentially expressed genes (Fig. 4). The differentially expressed genes in the A549 cells exposed to aldehydes were detected using the Human Whole Genome Microarray 44 K array (GSE56005). Next, we performed an integrative analysis of DNA methylation and mRNA expression to investigate whether DNA methylation has a regulatory impact on gene expression following aldehyde exposure. We identified hyper-methylated and down-regulated genes and hypo-methylated and up-regulated ones in each of aldehyde exposure groups (Table 4). These genes were considered potential epigenetic biomarkers for determining the exposure of the LSAAs.
Figure 4.
Heat map of differentially expressed genes in A549 cells exposed to the seven aldehydes using unsupervised hierarchical clustering. Colour intensity shows the differences in expression (Red: up-regulation; Green: down-regulation).
Table 4.
Anti-correlated matching number of DNA methylation and mRNA expression by aldehydes exposure.
| Hyper-methylated vs. down-regulated | Hypo-methylated vs. up-regulated | |
|---|---|---|
| Propanal (C3) | 252 | 166 |
| Butanal (C4) | 248 | 483 |
| Pentanal (C5) | 157 | 242 |
| Hexanal (C6) | 123 | 338 |
| Heptanal (C7) | 72 | 269 |
| Octanal (C8) | 68 | 132 |
| Nonanal (C9) | 146 | 299 |
GO and KEGG pathway analyses of candidate DNA methylation biomarkers of aldehydes
To investigate the related biological process and pathway of aldehyde-specific methylated genes, we performed GO and KEGG pathway analyses using DAVID bioinformatics resources. We analysed each aldehyde-matched DNA methylated genes. The common key biological processes were mainly involved in cell surface receptor linked signal transductions (GO:0007166) such as the G-protein coupled receptor (GPCR) signalling pathway (GO:0007186) and neurological system process (GO:0050877). KEGG pathway analysis revealed that aldehyde exposure was associated with neuroactive ligand-receptor interaction, calcium signalling pathway, and pathways in cancer. Although we identified overlapping similar biological processes and pathways for seven aldehydes, each aldehyde has distinct biological functions (Tables 5 and 6).
Table 5.
Biological process of significant anti-correlated methylated genes by aldehydes exposure.
| GO Annotation (Biological process; BP) | Gene count | p-value (<0.05) | |
|---|---|---|---|
| Propanal (C3) | G-protein coupled receptor protein signalling pathway | 34 | 0.0123 |
| Response to hormone stimulus | 18 | 9.64E-04 | |
| Mitotic cell cycle | 14 | 0.031 | |
| Regulation of secretion | 12 | 0.002 | |
| Regulation of protein amino acid phosphorylation | 10 | 0.007 | |
| Butanal (C4) | Cell surface receptor-linked signal transduction | 84 | 0.006 |
| Neurological system process | 56 | 0.017 | |
| Cell motion | 27 | 0.012 | |
| Regulation of phosphorylation | 24 | 0.049 | |
| Response to hormone stimulus | 23 | 0.008 | |
| Protein kinase cascade | 21 | 0.029 | |
| Pentanal (C5) | Cell surface receptor-linked signal transduction | 46 | 0.017 |
| Regulation of cell proliferation | 22 | 0.040 | |
| Induction of apoptosis | 13 | 0.012 | |
| Reproductive developmental process | 11 | 0.019 | |
| Sensory organ development | 10 | 0.021 | |
| Hexanal (C6) | Cell surface receptor-linked signal transduction | 65 | 3.12E-05 |
| G-protein coupled receptor protein signaling pathway | 44 | 8.25E-05 | |
| Neurological system process | 36 | 0.032 | |
| Cell adhesion | 29 | 8.21E-04 | |
| Acute inflammatory response | 7 | 0.017 | |
| Heptanal (C7) | Cell surface receptor-linked signal transduction | 49 | 3.27E-04 |
| G-protein coupled receptor protein signaling pathway | 34 | 3.80E-04 | |
| Neurological system process | 34 | 0.001 | |
| Sensory perception | 24 | 0.004 | |
| Neuron differentiation | 14 | 0.022 | |
| Cell morphogenesis involved in differentiation | 10 | 0.016 | |
| Octanal (C8) | Cell surface receptor-linked signal transduction | 32 | 0.004 |
| Homeostatic process | 14 | 0.047 | |
| Cell motion | 12 | 0.010 | |
| Cell morphogenesis | 9 | 0.032 | |
| T cell activation | 5 | 0.042 | |
| Sensory perception of pain | 3 | 0.043 | |
| Nonanal (C9) | Cell surface receptor-linked signal transduction | 60 | 3.49E-04 |
| Neurological system process | 40 | 0.003 | |
| G-protein coupled receptor protein signaling pathway | 36 | 0.009 | |
| Sensory perception | 26 | 0.028 | |
| Response to hormone stimulus | 14 | 0.040 | |
| Regulation of cell migration | 9 | 0.024 |
Table 6.
KEGG pathway analysis of selected aldehydes specific anti-correlated methylated genes.
| KEGG pathway | Gene count | p-value (<0.05) | |
|---|---|---|---|
| Propanal (C3) | Olfactory transduction | 14 | 0.03 |
| cAMP signalling pathway | 10 | 0.009 | |
| Calcium signalling pathway | 8 | 0.042 | |
| Mineral absorption | 4 | 0.049 | |
| Butanal (C4) | Pathways in cancer | 22 | 0.030 |
| PI3K-Akt signalling pathway | 21 | 0.016 | |
| MAPK signalling pathway | 18 | 0.007 | |
| Focal adhesion | 17 | 0.002 | |
| Calcium signaling pathway | 16 | 0.001 | |
| Neurotrophin signaling pathway | 12 | 0.003 | |
| Pentanal (C5) | Pathways in cancer | 19 | 1.48E-04 |
| MAPK signalling pathway | 13 | 0.002 | |
| Calcium signalling pathway | 12 | 2.79E-04 | |
| Ras signalling pathway | 11 | 0.006 | |
| cAMP signalling pathway | 10 | 0.008 | |
| Hexanal (C6) | Neuroactive ligand-receptor interaction | 19 | 4.08E-06 |
| Pathways in cancer | 15 | 0.016 | |
| Ras signalling pathway | 10 | 0.027 | |
| Calcium signalling pathway | 9 | 0.020 | |
| Rap1 signalling pathway | 9 | 0.046 | |
| Heptanal (C7) | Serotonergic synapse | 7 | 0.007 |
| Vascular smooth muscle contraction | 6 | 0.034 | |
| Insulin secretion | 5 | 0.040 | |
| Octanal (C8) | Neuroactive ligand-receptor interaction | 9 | 0.008 |
| Calcium signalling pathway | 6 | 0.038 | |
| Nonanal (C9) | Pathways in cancer | 17 | 0.002 |
| Neuroactive ligand-receptor interaction | 13 | 0.004 | |
| cAMP signalling pathway | 12 | 9.25E-04 | |
| Calcium signalling pathway | 11 | 0.002 | |
| MAPK signalling pathway | 11 | 0.018 | |
| Serotonergic synapse | 8 | 0.004 |
Discussion
Saturated aliphatic aldehydes are widely used in perfumes, essential oils, and flavoring13. They are also considered as indoor and outdoor air pollutants14. Human exposure to aldehydes occurs mainly by inhalation. Therefore, several studies have monitored the levels of aldehydes in the environment and reported the potential adverse health effects on humans15. Additionally, previous studies demonstrated that aldehydes may contribute to the development of various disease including pulmonary inflammatory diseases such as tracheitis, bronchitis, and bronchiolitis16. Among the aldehydes, formaldehyde and acetaldehyde are the most widely studied in compaison with other aldehydes. Little information is available regarding the toxicity of the other aldehydes, although numerous studies have conducted aldehyde measurement in indoor environments. Therefore, we investigated the relationship between exposure to other aldehydes and potential adverse health effects associated with pulmonary toxicity.
Epigenetic studies are important for examining the association between environmental exposure and health outcomes and have been utilized for exposure assessment. However, few studies have investigated the association between environmental factors including aldehydes and genome-wide epigenome in humans. Therefore, we focused on epigenetic alterations of saturated aliphatic aldehydes and aldehyde-specific DNA methylation changes as potential biomarkers.
We analysed the DNA methylation patterns of the seven saturated aliphatic aldehydes from propanal (C3) to nonanal (C9) in A549 cells. To identify the critical DNA methylation-based biomarkers of aldehydes, we investigated the correlation between DNA methylation profiles and mRNA expression using microarrays. Microarray technology is a powerful tool for evaluating environmental chemicals on human health, providing valuable genomic information for identifying biomarkers related to occupational exposure and disease prognosis17–19.
Using a DNA methylation array, we identified methylated genes specific for each aldehyde as well as commonly methylated genes among the seven aldehydes at the IC20, which shows low cytotoxicity and is useful for identifying genes. A total of 54 genes were commonly methylated compared to in control cells, with a greater than 3.0-fold change cut-off and p-value < 0.05. These methylation alterations were considered as important potential epigenetic-based biomarkers, which can be used to assess cases of aldehyde exposure and predict pulmonary toxicity induced by aldehyde exposure.
Aberrant DNA methylation has been associated with numerous human diseases, cancer, and environmental exposure20–22. Furthermore, hyper/hypo-methylated genes can modify the function of genes and regulate gene expression. Generally, DNA hypermethylation leads to down-regulation of genes and hypomethylation allows for up-regulation of genes. The inverse correlation between DNA methylation and mRNA expression plays critical roles and can serve as potential epigenetic biomarkers. Therefore, we also investigated the correlation between DNA methylation and mRNA expression to identify critical epigenetic biomarkers of aldehydes. In each aldehyde exposure group, we identified numerous anti-correlated methylated genes (Table 4). To further investigate the biological relevance of these genes, we performed functional analyses such as GO and KEGG pathway analyses using the DAVID bioinformatics tool (https://david.ncifcrf.gov/). Furthermore, GO terms and KEGG pathways are important indicators for elucidating the biological functions of genes23.
First, GO analysis showed that aldehyde-associated epigenetic biomarkers were associated with cell surface receptor-linked signal transduction, cell adhesion, inflammatory response, neurological system process, apoptosis, and the GPCR signalling pathway. Among the biological processes, cell surface receptor-linked signal transduction and the GPCR signalling pathway were mainly involved in biological processes related to aldehyde exposure. Further studies are needed to determine the molecular mechanisms involved in the effects of aldehyde exposure on the GPCR signalling pathway. Next, we conducted KEGG pathway analysis to identify detailed information on the key biological pathways relevant to aldehyde exposure. The calcium signalling pathway, MAPK signalling pathway, and cancer pathways were linked to the biological significance of aldehydes (Table 5). Among the related biological pathways, we previously demonstrated that exposure to butanal and octanal induced MAPK cascades in A549 cells24,25. The MAPK cascades are central signalling pathways in cellular processes that mediate a variety of physiological and pathological functions26. Therefore, further studies are required to determine whether aldehydes induce cellular metabolism via MAPK cascades.
Taken together, these findings demonstrate that DNA methylation-based epigenetic biomarkers of aldehydes in the human respiratory system can be used for exposure assessment and predicting pulmonary toxicity. Our results also improve the understanding of the underlying mechanisms of aldehyde exposure, which may be useful for determining the developmental toxicological mechanisms of aldehyde-induced environmental lung disease.
Materials and Methods
Chemicals and reagents
Aldehydes (propanal, butanal, pentanal, hexanal, heptanal, octanal, nonanal), dimethyl sulfoxide (DMSO), and 3-(4,5-dimethylthaizol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) were purchased from Sigma–Aldrich (St. Louis, MO, USA). The following cell culture media and supplemented buffer solutions were purchased from GIBCO™ (Grand Island, NY, USA): Roswell Park Memorial Institute (RPMI) 1640, Dulbecco’s phosphate buffered saline, foetal bovine serum, and antibiotics (penicillin and streptomycin). All chemicals used in this study were analytical grade or the highest grade available.
Cell culture
The human lung adenocarcinoma epithelial cell line (A549) was obtained from the Korea Cell Line Bank (Seoul, Korea) and grown in RPMI1640(Gibco) supplemented with 10% foetal bovine serum, sodium bicarbonate, HEPES, and penicillin under a humidified atmosphere of 5% CO2 and 95% air at 37 °C. The culture medium was replaced every 2 or 3 days.
Chemical treatment
A549 cells were treated with each chemical compound at 1.0% of final solvent (DMSO) concentration in multi-well plates. Because of the volatility of the compounds, the plates were sealed with sealing films after chemical treatment. A549 cells were seeded into a 24-well plate at a density of 7.0 × 104 cells/mL per well in 500 μL of media for the cytotoxicity assay and seeded into a 6-well plate at a density of 25.0 × 104 cells/mL for RNA and genomic DNA extraction. After incubation for 24 h, the cells were treated with the seven aldehydes for 48 h.
Determination of cell viability
To determine the cell viability and cytotoxic effects of the aldehydes, an MTT cell proliferation assay was performed as described previously by Mosmann27.
Preparation of genomic DNA
Genomic DNA was extracted from A549 cells exposed to the seven aldehydes using the QIAamp DNA Mini Kit (Qiagen, Hilden, Germany) according to manufacturer’s instructions. The amount of each genomic DNA was measured using a NanoDrop ND 1000 spectrophotometer (NanoDrop Technologies, Inc., Wilmington, DE, USA). Only clear samples without a substantial smear were considered suitable for use and their quality was checked by electrophoresis in a 1.5% agarose gel in 1X TAE buffer (4.8 g of Tris, 1.14 mL of acetic acid, 2 mL of 0.5 M EDTA at pH 8.0, and ethidium bromide) at a constant 100 V for 15 min.
Fragmentation of genomic DNA
Genomic DNA was randomly fragmented using a Sonic Dismembrator 550 (Fisher Scientific, USA) as described previously by Song et al.28. Briefly, 5 μg of DNA in 0.2 mL was placed into 1.5-mL tubes immersed in ice and sonicated for various times (10, 20, and 40 s) with a model Sonic Dismembrator 550 (Fischer Scientific, USA) set at 30 W, 1/8-inch horn, and 5% maximum output placed in the centre of the DNA solution at a depth of 5 mm. The size range of the fragmented DNA was evaluated by agarose gel electrophoresis and ethidium bromide staining using DNA size markers 500–10,000 base pairs in size.
Immunoprecipitation of methylated DNA (MeDIP)
MeDIP was performed using the MethylMiner Methylated DNA Enrichment Kit (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s instructions. One microgram of fragmented DNA and 3 μg of untreated control DNA (Input) were used for downstream quality procedures and labelling. First, 7 μL of MBD-Biotin Protein was coupled to 10 μL of Dynabeads M-280 Streptavidin according to the manufacturer’s instructions.
The MBD-magnetic bead conjugates were washed three times and resuspended in 1 volume of 1X bind/ wash buffer. The capture reaction was conducted by adding of 1 μg sonicated DNA to the MBD magnetic beads on a rotating mixer for 1 h at room temperature (RT). All capture reactions were performed in duplicate. Next, the beads were washed three times with 1X bind/wash buffer. The methylated DNA was eluted as a single fraction with a high-salt elution buffer (2,000 mM NaCl). Subsequently, each fraction was concentrated by ethanol precipitation using 1 μL glycogen (20 μg/μL), 1/10th volume of 3 M sodium acetate (pH 5.2), and two volumes of 100% ethanol, and then resuspended in 60 μL of DNase-free water. The eluted MeDIP pellet was stored at −20 °C for long-term storage.
Whole genome amplification (WGA)
After purification and quantification, MeDIP and Input DNA were amplified using the GenomePlex® Complete Whole Genome Amplification (WGA2) Kit (Sigma-Aldrich) according to the modified manufacturer’s instructions. Two points were modified from the supplier’s recommendations: first, the initial DNA fragmentation step was omitted because sonicated DNA was used. Second, 20 ng of IP and Input DNA were used rather than 10 ng. The reactions were cleaned by purification with a column-based technique (QIAquick PCR cleanup columns, Qiagen). Each amplified DNA was quantified using the NanoDrop ND 1000 spectrophotometer. The quality and success of WGA were assessed by agarose gel electrophoresis to evaluate the conservation of size distribution after WGA.
Analysis of promoter DNA methylation and gene expression profile
DNA methylation analysis was conducted on the WGA samples using the Human promoter 2 × 400 K methylation array (Agilent Technologies, Santa Clara, CA, USA). Briefly, 1 μg of amplified DNA and methylated IP samples were annealed with 5 μL of random primer for 3 min at 95 °C and then placed on ice for 5 min. WGA samples were labelled with Cy5 (red) for fully methylated DNA or Cy3 (green) for fragmented DNA using a randomly primed Klenow polymerase reaction using the Agilent Genomic DNA Enzymatic Labeling Kit (Agilent Technologies) according to the manufacturer’s instructions. After purification, labelled samples were resuspended with blocking reagent and hybridization buffer, followed by boiling for 3 min at 95 °C and incubated for 30 min at 37 °C, and then hybridized to arrays for 40 h at 67 °C while rotating at 800 RPM in a hybridization oven for 3 min. After washing, the slide was dried and hybridization images on the slides were scanned by the Agilent C scanner and analysed using Agilent Feature Extraction software (v10.7.3.1). All data normalization and selection of fold-changed probes were performed using GeneSpringGX 7.3 (Agilent Technologies). Probes showing a greater than 3.0-fold difference in the ratio between the test and control samples were selected and considered as differentially methylated probes.
Gene expression profiling of A549 cells exposed to aldehydes was conducted using the 44 k Whole Human Genome Microarray (Agilent Technologies) as described previously by Song et al.25. The data were normalized by dividing the average normalized treated signal intensity by the average normalized control intensity.
Comparative analysis of DNA methylation and mRNA
Comparative analysis of methylation and mRNA data was performed to identify the anti-correlation associated with exposure to aldehydes using GeneSpring GX.
Gene Ontology (GO) category analysis
To determine the functional categories of methylated target genes, the DAVID Gene Functional Classification Tool (https://david.ncifcrf.gov/) was used. Using the DAVID tool, pathway analyses were also conducted based on the Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathways, which was linked to the KEGG pathway map.
Statistical analysis
The differences between control and exposure subjects were evaluated using the unpaired t-test. The p-value criterion was set at p-value < 0.05 as the level of statistical significance.
Data availability
The datasets generated during the current study are available from the corresponding author upon reasonable request.
Electronic supplementary material
Acknowledgements
This research was supported by the Korea Research Foundation grants from Korea Ministry of Environment as “The Ecoinnovation Project (412-111-010)”, and KIST Program to Ryu, J.C. of the Republic of Korea.
Author Contributions
Y.C. performed the experiments, analysed the data, and wrote the manuscript; M.K.S. designed the research and performed the experiments; T.S.K. was involved in provided feedback and interpretation of the results; J.C.R. conceived and designed the research and supervised the work.
Competing Interests
The authors declare no competing interests.
Footnotes
Electronic supplementary material
Supplementary information accompanies this paper at 10.1038/s41598-018-28813-z.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Hou L, Zhang X, Wang D, Baccarelli A. Environmental chemical exposures and human epigenetics. Int. J. Epidemiol. 2012;41:79–105. doi: 10.1093/ije/dyr154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Jaenisch R, Bird A. Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat. Genet. 2003;33:245–254. doi: 10.1038/ng1089. [DOI] [PubMed] [Google Scholar]
- 3.Ho SM, et al. Environmental epigenetics and its implication on disease risk and health outcomes. ILAR. J. 2012;53(3–4):289–305. doi: 10.1093/ilar.53.3-4.289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Pogribny, I.P. et al. Alterations in DNA methylation resulting from exposure to chemical carcinogens. In: eLS. (John Wiley & Sons, Ltd., 2014).
- 5.Jelinek J, et al. Aberrant DNA methylation is associated with disease progression, resistance to imatinib and shortened survival in chronic myelogenous leukemia. PLoS One. 2011;6(7):e22110. doi: 10.1371/journal.pone.0022110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zhou R, Man Y. Integrated analysis of DNA methylation profiles and gene expression profiles to identify genes associated with pilocytic astrocytomas. Mol. Med. Rep. 2016;13(4):3491–3497. doi: 10.3892/mmr.2016.4943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Marsit CJ. Influence of environmental exposure on human epigenetic regulation. J. Exp. Biol. 2015;218(Pt 1):71–79. doi: 10.1242/jeb.106971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Song MK, Choi HS, Lee HS, Ryu JC. Transcriptome profile analysis of saturated aliphatic aldehydes reveals carbon number-specific molecules involved in pulmonary toxicity. Chem. Res. Toxicol. 2014;27:1362–1370. doi: 10.1021/tx500171r. [DOI] [PubMed] [Google Scholar]
- 9.Song MK, Lee HS, Ryu JC. Integrated analysis of microRNA and mRNA expression profiles highlights aldehyde-induced inflammatory responses in cells relevant for lung toxicity. Toxicology. 2015;334:111–121. doi: 10.1016/j.tox.2015.06.007. [DOI] [PubMed] [Google Scholar]
- 10.Villanueva, F. et al. Aldehyde measurements in indoor and outdoor environments in Central-Southern Spain. Current Air Quality Issues10.5772/60016 (2015).
- 11.Wang S, Ang HM, Tade MO. Volatile organic compounds in indoor environment and photocatalytic oxidation: State of the art. Environ. Int. 2007;33(5):694–705. doi: 10.1016/j.envint.2007.02.011. [DOI] [PubMed] [Google Scholar]
- 12.Lecureur V, et al. MAPK- and PKC/CREB-dependent induction of interleukin-11 by the environmental contaminant formaldehyde in human bronchial epithelial cells. Toxicology. 2012;292:13–22. doi: 10.1016/j.tox.2011.11.011. [DOI] [PubMed] [Google Scholar]
- 13.Lyndon, B. Spacecraft maximum allowable concentrations for selected airborne contaminants. (Johnson Space Center, 1999).
- 14.Marchand C, Bulliot B, Le CalvéS, Mirabel P. Aldehyde measurements in indoor environments in Strasbourg (France) Atmospheric. Environ. 2006;40:1336–1345. doi: 10.1016/j.atmosenv.2005.10.027. [DOI] [Google Scholar]
- 15.Kim J, et al. Indoor aldehydes concentration and emission rate of formaldehyde in libraries and private reading rooms. Atmospheric. Environ. 2013;71:1–6. doi: 10.1016/j.atmosenv.2013.01.059. [DOI] [Google Scholar]
- 16.Villanueva, F., et al Aldehyde measurements in indoor and outdoor environments in Central-Southern Spain. Current Air Quality Issues Farhad Nejadkoorki, IntechOpen, Available from: https://www.intechopen.com/books/current-air-quality-issues/aldehyde-measurements-in-indoor-and-outdoor-environments-in-central-southern-spain, 10.5772/60016 (2015)
- 17.Jung J, Hah K, Lee W, Jang W. Meta-analysis of microarray datasets for the risk assessment of coplanar polychlorinated biphenyl 77 (PCB77) on human health. Toxicol. Environ. Health. Sci. 2017;9(2):161–168. doi: 10.1007/s13530-017-0317-1. [DOI] [Google Scholar]
- 18.Gwinn MR, Weston A. Application of oligonucleotide microarray technology to toxic occupational exposures. J. Toxicol. Environ. Health A. 2008;71(5):315–324. doi: 10.1080/15287390701738509. [DOI] [PubMed] [Google Scholar]
- 19.Kim SY, et al. Identification of potential biomarkers for xylene exposure by microarray analyses of gene expression and methylation. Mol. Cell. Toxicol. 2016;12:15–20. doi: 10.1007/s13273-016-0003-4. [DOI] [Google Scholar]
- 20.Robertson KD. DNA methylation and human disease. Nat. Rev. Genet. 2005;6:597–610. doi: 10.1038/nrg1655. [DOI] [PubMed] [Google Scholar]
- 21.Olsson M, Beck S, Kogner P, Martinsson T, Carén H. Genome-wide methylation profiling identifies novel methylated genes in neuroblastoma tumors. Epigenetics. 2016;11(1):74–84. doi: 10.1080/15592294.2016.1138195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lambert MP, et al. Aberrant DNA methylation of imprinted loci in hepatocellular carcinoma and after in vitro exposure to common risk factors. Clin. Epigenetics. 2015;7:15. doi: 10.1186/s13148-015-0053-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Yao QY, Huang H, Tong Y, Xia EH, Gao LZ. Transcriptome analysis identifies candidate genes related to triacylglycerol and pigment biosynthesis and photoperiodic flowering in the ornamental and oil-producing plant, Camellia reticulata (Theaceae) Front. Plant. Sci. 2016;7:163. doi: 10.3389/fpls.2016.00163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Lee HS, et al. Analysis of mRNA expression profiles highlights alterations in modulation of the DNA damage-related genes under butanal exposure in A549 human alveolar epithelial cells. Mol. Cell. Toxicol. 2013;9(1):85–94. doi: 10.1007/s13273-013-0012-5. [DOI] [Google Scholar]
- 25.Song MK, et al. Analysis of microRNA and mRNA expression profiles highlights alterations in modulation of the MAPK pathway under octanal exposure. Environ. Toxicol. Pharmacol. 2014;37(1):84–94. doi: 10.1016/j.etap.2013.11.005. [DOI] [PubMed] [Google Scholar]
- 26.Plotnikov A, Zehorai E, Procaccia S, Seger R. The MAPK cascades: signaling components, nuclear roles and mechanisms of nuclear translocation. Biochim. Biophys. Acta. 2011;1813(9):1619–1633. doi: 10.1016/j.bbamcr.2010.12.012. [DOI] [PubMed] [Google Scholar]
- 27.Mosmann T. Rapid colorimetric assay for cellular growth and survival: Application to proliferation and cytotoxicity assays. J. Immunol. Methods. 1983;65:55–63. doi: 10.1016/0022-1759(83)90303-4. [DOI] [PubMed] [Google Scholar]
- 28.Song MK, et al. The optimal conditions for the estimation of DNA methylation levels using high throughput microarray derived DNA immunoprecipitation(MeDIP)-enrichment in human bloods. Toxicol. Environ. Health. Sci. 2011;3(3):185–192. doi: 10.1007/s13530-011-0097-y. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets generated during the current study are available from the corresponding author upon reasonable request.