Abstract
Leaf-stripe smuts on grasses are a highly polyphyletic group within Ustilaginomycotina, occurring in three genera, Tilletia, Urocystis, and Ustilago. Currently more than 12 Ustilago species inciting stripe smuts are recognised. The majority belong to the Ustilago striiformis-complex, with about 30 different taxa described from 165 different plant species. This study aims to assess whether host distinct-lineages can be observed amongst the Ustilago leaf-stripe smuts using nine different loci on a representative set. Phylogenetic reconstructions supported the monophyly of the Ustilago striiformis-complex that causes leaf-stripe and the polyphyly of other leaf-stripe smuts within Ustilago. Furthermore, smut specimens from the same host genus generally clustered together in well-supported clades that often had available species names for these lineages. In addition to already-named lineages, three new lineages were observed, and described as new species on the basis of host specificity and molecular differences: namely Ustilago jagei sp. nov. on Agrostis stolonifera, U. kummeri sp. nov. on Bromus inermis, and U. neocopinata sp. nov. on Dactylis glomerata.
Keywords: DNA-based taxonomy, host specificity, molecular species discrimination, multigene phylogeny, new taxa, species complex, Ustilaginaceae
INTRODUCTION
The term “stripe smut” is commonly used to refer to Ustilaginomycotina species that cause dark brown to black linear sori of varying length in the leaves of grasses (Poaceae). Black spore masses are released after the spores have matured beneath the epidermis in the mesophyll of the host leaves (Fischer 1953, Vánky 2012). The spore release process of sori is unknown, but may be facilitated either by the withering of dead epidermal cells or by enzymatic action, as in the white blister rusts (Heller & 2009). Of the smut genera that infect grasses, Ustilago is the most prevalent (Stoll et al. 2003, 2005, Vánky 2012).
The term stripe smut does not reflect phylogenetic relatedness, as at least two other genera, Tilletia and Urocystis, contain species that manifest similar symptoms. The vast majority of leaf-stripe smuts belong to Ustilago, including U. agropyri, U. bahuichivoensis, U. bethelii, U. calamagrostidis, U. calcarea, U. davisii, U. deyeuxiicola, U. echinata, U. filiformis, U. phlei, U. scrobiculata, U. serpens s. lat., U. sporoboli-indici, U. striiformis s. lat., U. trebouxii, U. trichoneurana, and U. ulei (Vánky 2012). Of these species, U. striiformis s. lat., with the type species described on Holcus lanatus, is a complex occurring on 164 species of Poaceae representing 44 different genera (Achnatherum, Agropyron, Agrostis, Alopecurus, Ammophila, Anthoxanthum, Arctagrostis, Arrhenatherum, Avena, Beckmannia, Brachypodium, Briza, Bromus, Calamagrostis, Cleistogenes, Cynosurus, Dactylis, Danthonia, Deschampsia, Deyeuxia, Elymus, Festuca, Helictotrichon, Hierochloë, Holcus, Hordeum, Hystrix, Koeleria, Leymus, Lolium, Melica, Milium, Pennisetum, Phalaris, Phleum, Piptatherum, Poa, Polypogon, Puccinellia, Sesleria, Setaria, Sitanion, Trisetaria, and Trisetum). Based on host specificity and minor differences in spore size and surface ornamentation, approximately 30 different taxa have been described in the U. striiformis species complex on various host plants (Vánky 2012, Savchenko et al. 2014a). Ustilago serpens probably represents an overlooked species complex, occuring on five host genera: Agropyron, Brachypodium, Bromus, Elymus, and Leymus. Whether other species with large warts on their spores also belong to this complex, such as U. echinata and U. scrobiculata, is currently unclear.
Ustilago striiformis s. lat. on Alopecurus pratensis has often been the sole representative of this group in phylogenetic analyses (Stoll et al. 2005, Begerow et al. 2006, McTaggart et al. 2012a). Stoll et al. (2005) supported the recognition of U. calamagrostidis, a parasite of several species of Calamagrostis, as separate from U. striiformis. The morphological difference was mainly in spore size and ornamentation. Savchenko et al. (2014a) provided a more detailed analysis of the U. striifomis species complex using several host-fungus combinations and phylogenetic reconstructions based on the nrITS and partial LSU regions. However, while two additional species were proposed as distinct in the U. striiformis-complex, the phylogenetic resolution was too low to draw further conclusions regarding host specificity and potential species boundaries. To resolve undescribed lineages within this species complex, Savchenko et al. (2014a) suggested that several additional gene loci and host-fungus combinations should be included. However, in line with Vánky (2012), Savchenko et al. (2014a) suggested that it would be difficult to distinguish between these lineages based on morphological characters. DNA-based characteristics, such as diagnostic SNPs, along with host specificity might be a solution towards characterizing and describing previously-named and new species (Denchev et al. 2009, Piątek et al. 2013). The aim of this study was to use a multigene phylogeny to infer the phylogenetic differentiation in the leaf stripe smuts in the genus Ustilago, particularly those in the U. striiformis species complex.
MATERIAL AND METHODS
Plant and fungal material
Specimens used in the study are listed in Table 1. The names of the hosts and fungi was derived from the latest version of The International Plant Names Index (www.ipni.org), Index Fungorum (www.indexfungorum.org/) and Vánky (2012), and partly following a broad generic concept for Ustilago (Thines 2016). A majority of the samples were collected in Germany (about 76) and most collections were not older than 20 years. Samples are deposited in Herbarium Senckenbergianum Görlitz (GLM). All host identifications were confirmed by ITS sequences.
Table 1.
Smut specimens used for phylogenetic analysis.
| DNA-no. | Species | Host |
Collection details |
gene loci |
|||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Location | Date | Collector | Fungarium no. | ITS | atp2 | ssc1 | map | myosin | rpl4A | rpl3 | sdh1 | tif2 | |||
| 2354 | Sporisorium aff. inopiatum (Langdonia) | Aristida adscensionis | Zambia | 12 Apr. 2001 | C., T. & K. Vánky | M-0215944 | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929824 | KY929964 | KY930127 |
| 474 | Ustilago agrostidis-palustris | Agrostis cf. gigantea | Germany, Bavaria | 22 Jun. 2012 | J. Kruse | GLM-F105832 | KY929551 | KY930157 | KY929994 | KY929709 | KY929639 | KY929849 | KY929779 | KY929919 | KY930082 |
| 1374 | Agrostis gigantea | Switzerland | 9 Jul. 2004 | V. Kummer | GLM-F107425 | KY929582 | KY930188 | KY930025 | KY929729 | KY929659 | KY929869 | KY929799 | KY929939 | KY930102 | |
| 2395 | Agrostis sp. | Germany, Lower Saxony | 12 Jul. 2014 | J. Kruse & H. Jage | GLM-F107439 | KY929596 | KY930202 | KY930039 | KY929739 | KY929669 | KY929879 | KY929809 | KY929949 | KY930112 | |
| 2287 | Ustilago airae-caespitosae | Deschampsia caespitosa | Polen | 13 Jul. 1994 | H. Scholz | B 70 0014901 | KY929526 | KY930132 | KY929969 | KY929688 | KY929618 | KY929828 | KY929758 | KY929898 | KY930061 |
| 2401 | Deschampsia caespitosa | Austria, Upper Austria | 15 Aug. 2014 | J. Kruse | GLM-F107444 | KY929601 | KY930207 | KY930044 | KY929744 | KY929674 | KY929884 | KY929814 | KY929954 | KY930117 | |
| 2402 | Deschampsia caespitosa | Austria, Upper Austria | 15 Aug. 2014 | J. Kruse | GLM-F107445 | KY929602 | KY930208 | KY930045 | KY929745 | KY929675 | KY929885 | KY929815 | KY929955 | KY930118 | |
| 477 | Ustilago alopecurivora | Alopecurus pratensis | Germany, Hesse | 22 May 2010 | J. Kruse | GLM-F105834 | KY929553 | KY930159 | KY929996 | KY929711 | KY929641 | KY929851 | KY929781 | KY929921 | KY930084 |
| 1376 | Alopecurus pratensis | Germany, Saxony-Anhalt | 20 May 2013 | H. Jage | GLM-F107426 | KY929583 | KY930189 | KY930026 | – | – | – | – | – | – | |
| 1822 | Ustilago aff. andropogonis (Sporisorium) | Bothriochloa ischaemum | Germany, Saxony-Anhalt | 25 Jul. 2004 | H. Jage & H. John | GLM-F062665 | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929764 | KY929904 | KY930067 |
| 432 | Ustilago perennans | Arrhenatherum elatius | Germany, Schleswig-Holstein | 21 Jun. 2007 | J. Kruse | GLM-F105817 | KY929536 | KY930142 | KY929979 | KY929697 | KY929627 | KY929837 | KY929767 | KY929907 | KY930070 |
| 2398 | Ustilago brizae | Briza media | Austria, Tirol | 21 Jul. 2014 | J. Kruse | GLM-F107442 | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929812 | KY929952 | KY930115 |
| 2399 | Briza media | Germany, Bavaria | 19 Jul. 2014 | J. Kruse | GLM-F107443 | KY929600 | KY930206 | KY930043 | KY929743 | KY929673 | KY929883 | KY929813 | KY929953 | KY930116 | |
| 498 | Ustilago bromina | Bromus inermis | Germany, Saxony-Anhalt | 04 Jun. 2011 | J. Kruse | GLM-F105843 | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929785 | KY929925 | KY930088 |
| 500 | Bromus inermis | Germany, Thuringia | 15 Jun. 2013 | J. Kruse | GLM-F105844 | KY929563 | KY930169 | KY930006 | KY929716 | KY929646 | KY929856 | KY929786 | KY929926 | KY930089 | |
| 1180 | Bromus inermis | Germany, Berlin | May 1983 | H. Scholz | HUV No 498 (TUB) | KY929613 | KY930219 | KY930056 | – | – | – | – | – | – | |
| 2070 | Bromus inermis | Germany, Berlin | Aug. 1892 | P. Sydow | B 70 0014775 | KY929525 | – | – | – | – | – | – | – | – | |
| 2275 | Bromus inermis | Germany, Brandenburg | 17 Jul. 2005 | H. & I. Scholz | B 70 0014755 | KY929524 | KY930131 | KY929968 | – | – | – | – | – | – | |
| 2276 | Bromus inermis | Germany, Thuringia | 10 Sep. 1999 | I. Scholz | B 70 0021843 | KY929527 | KY930133 | KY929970 | – | – | – | – | – | – | |
| 1591 | Ustilago aff. bromivora | Bromus rigidus | Greece | 23 Apr. 2013 | C. & F. Klenke | GLM-F107429 | KY929586 | KY930192 | KY930029 | KY929731 | KY929661 | KY929871 | KY929801 | KY929941 | KY930104 |
| 3370 | Bromus sterilis | Spain, Andalusia | 2 May 2015 | J. Kruse | GLM-F107449 | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929818 | KY929958 | KY930121 | |
| 442 | Ustilago calamagrostidis | Calamagrostis epigejos | Germany, Lower Saxony | 03 Aug. 2011 | J. Kruse | GLM-F105818 | KY929537 | KY930143 | KY929980 | – | – | – | – | – | – |
| 445 | Calamagrostis epigejos | Germany, Baden-Württemberg | 20 Jul. 2013 | J. Kruse | GLM-F105819 | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929768 | KY929908 | KY930071 | |
| 1383 | Calamagrostis epigejos | Germany, Saxony-Anhalt | 29 Jun. 2013 | H. Zimmermann, U. Richter | GLM-F107427 | KY929584 | KY930190 | KY930027 | KY929730 | KY929660 | KY929870 | KY929800 | KY929940 | KY930103 | |
| 1912 | Calamagrostis epigejos | Germany, Saxony-Anhalt | 09 Aug. 1996 | H. Jage | GLM-F048100 | KY929530 | KY930136 | KY929973 | KY929691 | KY929621 | KY929831 | KY929761 | KY929901 | KY930064 | |
| 1182 | Ustilago corcontica | Calamagrostis villosa | Germany, Saxony | 22 Aug. 1987 | W. Dietrich | HUV No 794 (TUB) | KY929615 | KY930221 | KY930058 | – | – | – | – | – | – |
| 1611 | Calamagrostis villosa | Germany, Saxony-Anhalt | 26 Jul. 2003 | H. & U. Richter | GLM-F107434 | KY929591 | KY930197 | KY930034 | – | – | – | – | – | – | |
| 1825 | Ustilago cruenta (Sporisorium) | Sorghum bicolor | Greece | 11 May 2006 | H-W, Otto | GLM-F078871 | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929766 | KY929906 | KY930069 |
| 3375 | Ustilago cynodontis | Cynodon dactylon | Spain, Andalusia | 3 May 2015 | J. Kruse | GLM-F107450 | KY929607 | KY930213 | KY930050 | KY929749 | KY929679 | KY929889 | KY929819 | KY929959 | KY930122 |
| 3376 | Cynodon dactylon | Spain, Andalusia | 3 May 2015 | J. Kruse | GLM-F107451 | KY929608 | KY930214 | KY930051 | KY929750 | KY929680 | KY929890 | KY929820 | KY929960 | KY930123 | |
| 1596 | Ustilago aff. dactyloctaenii (Sporisorium) | Dactyloctenium australe | South-Africa | 22 Feb. 2000 | V. Kummer | GLM-F107430 | KY929587 | KY930193 | KY930030 | KY929732 | KY929662 | KY929872 | KY929802 | KY929942 | KY930105 |
| 478 | Ustilago denotarisii | Arrhenatherum elatius | Germany, Schleswig-Holstein | 13 May 2007 | J. Kruse | GLM-F105835 | KY929554 | KY930160 | KY929997 | – | – | – | – | – | – |
| 481 | Arrhenatherum elatius | Germany, Rhineland-Palatinate | 23 May 2010 | J. Kruse | GLM-F105836 | KY929555 | KY930161 | KY929998 | – | – | – | – | – | – | |
| 483 | Arrhenatherum elatius | Germany, Lower Saxony | 31 Jul. 2011 | J. Kruse | GLM-F105837 | KY929556 | KY930162 | KY929999 | – | – | – | – | – | – | |
| 486 | Arrhenatherum elatius | Germany, Thuringia | 04 Jun. 2012 | J. Kruse | GLM-F105838 | KY929557 | KY930163 | KY930000 | – | – | – | – | – | – | |
| 488 | Arrhenatherum elatius | Germany, Bavaria | 16 May 2013 | J. Kruse | GLM-F105839 | KY929558 | KY930164 | KY930001 | – | – | – | – | – | – | |
| 447 | Ustilago echinata | Phalaris arundinacea | Germany, Lower Saxony | 01 Jul. 2010 | J. Kruse | GLM-F105820 | KY929539 | KY930145 | KY929982 | KY929699 | KY929629 | KY929839 | KY929769 | KY929909 | KY930072 |
| 449 | Phalaris arundinacea | Germany, Lower Saxony | 29 Aug. 2011 | J. Kruse | GLM-F105821 | KY929540 | KY930146 | KY929983 | KY929700 | KY929630 | KY929840 | KY929770 | KY929910 | KY930073 | |
| 1914 | Phalaris arundinacea | Switzerland, St. Gallen | 26 Jul. 2000 | H. Jage | GLM-F048338 | KY929531 | KY930137 | KY929974 | KY929692 | KY929622 | KY929832 | KY929762 | KY929902 | KY930065 | |
| 451 | Ustilago aff. filiformis | Glyceria fluitans | Germany, Lower Saxony | 17 May 2007 | J. Kruse | GLM-F105822 | KY929541 | KY930147 | KY929984 | KY929701 | KY929631 | KY929841 | KY929771 | KY929911 | KY930074 |
| 454 | Glyceria fluitans | Germany, Bavaria | 24 Jun. 2012 | J. Kruse | GLM-F105823 | KY929542 | KY930148 | KY929985 | KY929702 | KY929632 | KY929842 | KY929772 | KY929912 | KY930075 | |
| 455 | Glyceria fluitans | Germany, Bavaria | 10 May 2013 | J. Kruse | GLM-F105824 | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929773 | KY929913 | KY930076 | |
| 456 | Ustilago filiformis | Glyceria maxima | Germany, Lower Saxony | 01 Jul. 2010 | J. Kruse | GLM-F105825 | KY929544 | KY930150 | KY929987 | KY929704 | KY929634 | KY929844 | KY929774 | KY929914 | KY930077 |
| 472 | Ustilago jagei sp. nov. | Agrostis rupestris | Switzerland, Grisons | 02 Aug. 2009 | J. Kruse | GLM-F105830 | KY929549 | KY930155 | KY929992 | – | – | – | – | – | – |
| 473 | Agrostis stolonifera | Germany, Bavaria | 20 May 2012 | J. Kruse | GLM-F105831 | KY929550 | KY930156 | KY929993 | – | – | – | – | – | – | |
| 476 | Agrostis stolonifera | Germany, Hesse | 22 May 2010 | J. Kruse | GLM-F105833 | KY929552 | KY930158 | KY929995 | KY929710 | KY929640 | KY929850 | KY929780 | KY929920 | KY930083 | |
| 551 | Agrostis sp. | Germany, Lower Saxony | 11 Jun. 2010 | J. Kruse | GLM-F107423 | KY929580 | KY930186 | KY930023 | KY929727 | KY929657 | KY929867 | KY929797 | KY929937 | KY930100 | |
| 2396 | Agrostis stolonifera | Germany, Bavaria | 20 Jul. 2014 | J. Kruse | GLM-F107440 | KY929597 | KY930203 | KY930040 | KY929740 | KY929670 | KY929880 | KY929810 | KY929950 | KY930113 | |
| 2397 | Agrostis stolonifera | Germany, Hesse | 27 Jun. 2014 | J. Kruse | GLM-F107441 | KY929598 | KY930204 | KY930041 | KY929741 | KY929671 | KY929881 | KY929811 | KY929951 | KY930114 | |
| 494 | Agrostis sp. | Germany, Bavaria | 04 Jul. 2013 | J. Kruse | GLM-F105841 | KY929560 | KY930166 | KY930003 | KY929713 | KY929643 | KY929853 | KY929783 | KY929923 | KY930086 | |
| 1375 | Agrostis stolonifera | Germany, Saxony-Anhalt | 16 Sep. 2001 | H. Jage | GLM-F047379 | KY929528 | KY930134 | KY929971 | KY929689 | KY929619 | KY929829 | KY929759 | KY929899 | KY930062 | |
| 1612 | Ustilago kummeri sp. nov. | Bromus inermis | Germany, Brandenburg | 19 Jun. 2010 | V. Kummer | GLM-F107435 | KY929592 | KY930198 | KY930035 | KY929736 | KY929666 | KY929876 | KY929806 | KY929946 | KY930109 |
| 1948 | Bromus inermis | Germany, Saxony-Anhalt | 17 Jul. 2001 | H. Jage, W. Lehman | GLM-F047380 | KY929529 | KY930135 | KY929972 | KY929690 | KY929620 | KY929830 | KY929760 | KY929900 | KY930063 | |
| 501 | Ustilago loliicola | Lolium perenne | Germany, Bavaria | 14 May 2013 | J. Kruse | GLM-F105845 | KY929564 | KY930170 | KY930007 | – | – | – | – | – | – |
| 2288A | Festuca pratensis | Germany, Hesse | 25 May 2014 | J. Kruse | GLM-F107437 | KY929594 | KY930200 | KY930037 | – | – | – | – | – | – | |
| 3386 | Festuca arundinacea | Germany, Hesse | 02 Nov. 2014 | J. Kruse | GLM-F107454 | KY929611 | KY930217 | KY930054 | KY929753 | KY929683 | KY929893 | KY929823 | KY929963 | KY930126 | |
| 2815A | Ustilago maydis | Zea mays | Germany, Saxony-Anhalt | 10 Jul. 2007 | H. Jage | GLM-F107446 | KY929603 | KY930209 | KY930046 | KY929746 | KY929676 | KY929886 | KY929816 | KY929956 | KY930119 |
| 1404 | Ustilago milii | Milium effusum | Germany, Saxony-Anhalt | 02 Jun. 2002 | H. Jage | GLM-F107428 | KY929585 | KY930191 | KY930028 | – | – | – | – | – | – |
| 2303 | Milium effusum | Germany, Saxony | 03 Jun. 2012 | W. Dietrich | GLM-F107438 | KY929595 | KY930201 | KY930038 | KY929738 | KY929668 | KY929878 | KY929808 | KY929948 | KY930111 | |
| 3385 | Milium effusum | Germany, Hesse | 11 Jun. 2015 | J. Kruse | GLM-F107453 | KY929610 | KY930216 | KY930053 | KY929752 | KY929682 | KY929892 | KY929822 | KY929962 | KY930125 | |
| 503 | Ustilago neocopinata sp. nov. | Dactylis glomerata | Germany, Lower Saxony | 01 Jul. 2010 | J. Kruse | GLM-F105846 | KY929565 | KY930171 | KY930008 | – | – | – | – | – | – |
| 505 | Dactylis glomerata | Germany, Bavaria | 20 Jun. 2010 | J. Kruse | GLM-F105847 | KY929566 | KY930172 | KY930009 | – | – | – | – | – | – | |
| 506 | Dactylis glomerata | Germany, Lower Saxony | 19 May 2011 | J. Kruse | GLM-F105848 | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929787 | KY929927 | KY930090 | |
| 508 | Dactylis glomerata | Germany, Bavaria | 19 Jul. 2011 | J. Kruse | GLM-F105849 | KY929568 | KY930174 | KY930011 | KY929718 | KY929648 | KY929858 | KY929788 | KY929928 | KY930091 | |
| 510 | Dactylis glomerata | Germany, Bavaria | 24 May 2012 | J. Kruse | GLM-F105850 | KY929569 | KY930175 | KY930012 | KY929719 | KY929649 | KY929859 | KY929789 | KY929929 | KY930092 | |
| 512 | Dactylis glomerata | Germany, Bavaria | 15 Jun. 2012 | J. Kruse | GLM-F107413 | KY929570 | KY930176 | KY930013 | – | – | – | – | – | – | |
| 521 | Dactylis glomerata | Germany, Thuringia | 15 Jun. 2013 | J. Kruse | GLM-F107414 | KY929571 | KY930177 | KY930014 | – | – | – | – | – | – | |
| 463 | Ustilago nuda | Hordeum vulgare | Germany, Bavaria | 12 May 2012 | J. Kruse | GLM-F105826 | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929775 | KY929915 | KY930078 |
| 884 | Sporisorium aff. occidentale | Andropogon gerardii | USA | 30 Jul. 1989 | not known | HUV No 758 (TUB) | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929825 | KY929965 | KY930128 |
| 471 | Ustilago salweyi | Holcus mollis | Germany, Bavaria | 11 Jun. 2012 | J. Kruse | GLM-F105829 | KY929548 | KY930154 | KY929991 | KY929708 | KY929638 | KY929848 | KY929778 | KY929918 | KY930081 |
| 489 | Holcus mollis | Germany, Bavaria | 16 May 2013 | J. Kruse | GLM-F105840 | KY929559 | KY930165 | KY930002 | KY929712 | KY929642 | KY929852 | KY929782 | KY929922 | KY930085 | |
| 523 | Holcus lanatus | Germany, Lower Saxony | 24 May 2009 | J. Kruse | GLM-F107415 | KY929572 | KY930178 | KY930015 | KY929720 | KY929650 | KY929860 | KY929790 | KY929930 | KY930093 | |
| 524 | Holcus lanatus | Germany, Lower Saxony | 22 May 2010 | J. Kruse | GLM-F107416 | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929791 | KY929931 | KY930094 | |
| 525 | Holcus lanatus | Germany, Lower Saxony | 27 May 2010 | J. Kruse | GLM-F107417 | KY929574 | KY930180 | KY930017 | KY929722 | KY929652 | KY929862 | KY929792 | KY929932 | KY930095 | |
| 531 | Holcus lanatus | Germany, Bavaria | 17 May 2012 | J. Kruse | GLM-F107418 | KY929575 | KY930181 | KY930018 | – | – | – | – | – | – | |
| 541 | Holcus mollis | Germany, Saxony | 03 Jun. 2011 | J. Kruse | GLM-F107419 | KY929576 | KY930182 | KY930019 | KY929723 | KY929653 | KY929863 | KY929793 | KY929933 | KY930096 | |
| 543 | Holcus mollis | Germany, Saxony-Anhalt | 05 Jun. 2011 | J. Kruse | GLM-F107420 | KY929577 | KY930183 | KY930020 | KY929724 | KY929654 | KY929864 | KY929794 | KY929934 | KY930097 | |
| 544 | Holcus mollis | Germany, Saxony-Anhalt | 05 Jun. 2011 | J. Kruse | GLM-F107421 | KY929578 | KY930184 | KY930021 | KY929725 | KY929655 | KY929865 | KY929795 | KY929935 | KY930098 | |
| 545 | Holcus mollis | Germany, Lower Saxony | 17 Aug. 2011 | J. Kruse | GLM-F107422 | KY929579 | KY930185 | KY930022 | KY929726 | KY929656 | KY929866 | KY929796 | KY929936 | KY930099 | |
| 497 | Ustilago scaura | Helictotrichon pubescens | Germany, Rhineland-Palatinate | 23 May 2010 | J. Kruse | GLM-F105842 | KY929561 | KY930167 | KY930004 | KY929714 | KY929644 | KY929854 | KY929784 | KY929924 | KY930087 |
| 3384 | Helictotrichon pubescens | Germany, Hesse | 10 Jun. 2015 | J. Kruse | GLM-F107452 | KY929609 | KY930215 | KY930052 | KY929751 | KY929681 | KY929891 | KY929821 | KY929961 | KY930124 | |
| 1359 | Ustilago aff. schroeteriana (Sporisorium) | Paspalum virgatum | Costa Rica | 15 Mar. 1991 | T. & K. Vánky | HUV No 888 (TUB) | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929826 | KY929966 | KY930129 |
| 1608 | Ustilago scrobiculata | Calamagrostis epigejos | Germany, Brandenburg | 17 Aug. 2011 | V. Kummer & C. Buhr | GLM-F107431 | KY929588 | KY930194 | KY930031 | KY929733 | KY929663 | KY929873 | KY929803 | KY929943 | KY930106 |
| 1609 | Calamagrostis epigejos | Germany, Thuringia | 27 May 2010 | V. Kummer | GLM-F107432 | KY929589 | KY930195 | KY930032 | KY929734 | KY929664 | KY929874 | KY929804 | KY929944 | KY930107 | |
| 1610 | Calamagrostis epigejos | Germany, Brandenburg | 24 Jun. 2007 | V. Kummer | GLM-F107433 | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929805 | KY929945 | KY930108 | |
| 467 | Ustilago serpens | Elymus repens | Germany, Schleswig-Holstein | 31 Jul. 2012 | J. Kruse | GLM-F105827 | KY929546 | KY930152 | KY929989 | KY929706 | KY929636 | KY929846 | KY929776 | KY929916 | KY930079 |
| 469 | Elymus repens | Germany, Thuringia | 15 Jun. 2013 | J. Kruse | GLM-F105828 | KY929547 | KY930153 | KY929990 | KY929707 | KY929637 | KY929847 | KY929777 | KY929917 | KY930080 | |
| 3110 | Elymus repens | Germany, Brandenburg | 29 Jun. 2014 | V. Kummer | GLM-F107447 | KY929604 | KY930210 | KY930047 | – | – | – | – | – | – | |
| 1305 | Ustilago aff. sorghi (Sporisorium) | Sorghum plumosum | Australia | 20 Feb. 1996 | A. A. Mitchell, C. & K. Vánky | HUV No 970 (TUB) | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929827 | KY929967 | KY930130 |
| 1951 | Ustilago aff. syntherismae | Digitaria sanguinalis | Germany, Saxony-Anhalt | 01 Oct . 2004 | H. Jage | GLM-F064759 | KY929534 | KY930140 | KY929977 | KY929695 | KY929625 | KY929835 | KY929765 | KY929905 | KY930068 |
| 1617 | Digitaria sanguinalis | Germany, Brandenburg | 11 Aug. 2001 | V. Kummer | GLM-F107436 | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929807 | KY929947 | KY930110 | |
| 553 | Ustilago trichophora | Echinochloa crus-galli | Germany, North Rhine-Westphalia | 04 Oct . 2010 | J. Kruse | GLM-F107424 | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | Kruse et al. 2017b | KY929798 | KY929938 | KY930101 |
| 1957 | Echinochloa crus-galli | Germany, Saxony-Anhalt | 01 Oct . 2003 | H. Jage | GLM-F062638 | KY929532 | KY930138 | KY929975 | KY929693 | KY929623 | KY929833 | KY929763 | KY929903 | KY930066 | |
| 3347 | Ustilago aff. vanderystii (Sporisorium) | Hyparrhenia hirta | Spain, Andalusia | 22 Apr. 2015 | J. Kruse | GLM-F107448 | KY929605 | KY930211 | KY930048 | KY929747 | KY929677 | KY929887 | KY929817 | KY929957 | KY930120 |
Type specimens are printed in bold face.
DNA extraction and PCR
About 2–20 mg of infected plant tissue was taken from fungarium samples, placed in 2 mL plastic reaction tubes and homogenized in a mixer mill (MM2, Retsch) using a combination of three to five 1 mm and two 3 mm metal beads at 25 Hz for 5–10 min. Genomic DNA was extracted using the BioSprint 96 DNA Plant Kit (Qiagen, Hilden) loaded to a KingFisher Flex robot (Thermo Scientific, Dreieich).
The complete nrITS of all DNA extracts were amplified using PCR following the procedure of White et al. (1990). The primer pairs M-ITS1 (Stoll et al. 2003) / ITS4 (White et al. 1990) or M-ITS1 / smITS-R1 (Kruse et al. 2017a) were used as the reverse and forward primers, respectively. For DNA samples from historic specimens, including type specimens, the Ustilaginaceae-optimised reverse primer ITS-US3R (5’TATCAAAACCCGGCAGGGAAG3’), located at the ITS2 region, was used.
The NL1 and NL4 primer pair (O’Donnell 1993) were used to amplify the Large Subunit (LSU) of the nrDNA with an annealing temperature of 53 °C. For other loci, the following regions were amplified with their respective primer pairs and annealing temperatures in brackets: myosin R0.5/F3 (55 °C), map R6/F2 (56 °C), rpl3 R1/F1 (53 °C), tif2 R3/F3 (53 °C), ssc1 R1/F2 (53 °C), sdh1 R3/F2 (53 °C), rpl4A R1/F4 (53 °C), and atp2 R4/F6 (53 °C) (Kruse et al. 2017b).
The plant ITS was amplified using the primer pair ITS1P and ITS4 (Ridgway et al. 2003) at 53 °C annealing temperature. The cycling reaction was performed in a thermocycler (Eppendorf Mastercycler 96 vapo protect; Eppendorf, Hamburg) with an initial denaturation at 95 °C for 4 min, 36 PCR cycles of denaturation at 95 °C for 40 s, annealing between 53–56 °C (depending on the specific primer pair) for 40 s and elongation at 72 °C for 60 s, followed by a final elongation at 72 °C for 4 min. For DNA samples older than 50 years, PCR cycles were increased to 46 cycles and a larger amount of DNA (1.5 μL of extracted DNA in a reaction volume of 11 μL) was used. The resulting amplicons were sequenced at the Biodiversity and Climate Research Centre (BiK-F) laboratory using the abovementioned PCR primers. However, amplicons from M-ITS1/smITS-R1 were sequenced using the ITS4 reverse primer. The resulting sequences were deposited in GenBank (https://www.ncbi.nlm.nih.gov/genbank/, Table 1).
Alignments and phylogenetic tree reconstruction
We used 93 samples (including 62 of the Ustilago striiformis-complex) for the phylogenetic analysis; 93 had sequences from nrITS, atp2 (ATP synthase subunit 2) and ssc1 (member of the heat shock proteins), and 70 had additional sequences from myosin (myosin group I), map (methionine aminopeptidase), rpl3 (ribosomal protein L3), tif2 (initial translation factor of elF4A), sdh1 (succinate dehydrogenase ubiquinone flavoprotein), and rpl4A (ribosomal protein L4-A) (Table 1). About two thirds of the samples (62) belonged to the U. striiformis species complex. Sporisorium inopinatum (syn. Langdonia inopinata) was chosen as outgroup, according to the findings of McTaggart et al. (2012a).
Alignments were made on individual loci using mafft v. 7 (Katoh & Standley 2013) using the G-INS-i algorithm. Both leading and trailing gaps of the alignments were removed manually. Two different sets of concatenated alignments for the phylogenetic constructions were generated. The first multigene-alignment includes three loci (ITS, atp2, and ssc1) from 93 smut samples. The resulting total alignment was 1502 bp (ITS: 643 bp, atp2: 595 bp, ssc1: 264 bp). The second multigene-alignment included nine genes with a final alignment of 3156 bp (ITS: 643 bp, atp2: 595 bp, ssc1 264 bp, map: 251 bp, myosin: 257 bp, rpl4A: 415 bp, rpl3: 218 bp, sdh1: 269 bp, tif2: 244 bp).
The diagnostic bases for the U. striiformis species complex for all gene markers were determined using the above mentioned alignments. One further ITS alignment was created (443 bp), with the sequence of the type specimen of U. bromina (Table 1), the U. bromina sequences from GenBank (KF381006-8) and sequences from the same host-fungus-combination from this study, to check if all specimens were sequence-identical with the type collection of U. bromina on Bromus inermis (data not shown).
For phylogenetic tree constructions, Minimum Evolution (ME) analysis was done using Mega 6.06 (Tamura et al. 2013) with the Tamura-Nei substitution model and assuming complete deletion at 80 % cut-off with 1000 bootstrap replicates. All other parameters were set to default values. Maximum Likelihood (ML) analysis was done using RAxML (Stamatakis 2014) with parameters set to default values and Bayesian analysis was done using MrBayes 3.2 (Ronquist & Huelsenbeck 2003) running five times with model 6 (GTR) using four incrementally heated chains for 10 million generations, sampling every 1000th tree discarding the first 30 % of the obtained trees, all other parameters were set to default on the TrEase webserver (http://www.thines-lab.senckenberg.de/trease).
To account for potentially deviating evolutionary properties, the analysis in ME was done also on a partitioned concatenated dataset. As no supported differences within the topology of the trees were observed in comparison with the un-partitioned dataset, the other analyses were carried out without partitioning.
Morphological examination
For light microscopy, fungarium specimens (GLM-F107417, GLM-F105836, GLM-F107435, GLM-F107413, GLM-F047379, GLM-F105827) were transferred to 60 % lactic acid on a slide. Morphological examination was carried out using a Zeiss Imager M2 AX10 microscope (Carl Zeiss, Göttingen). Measurements of the spores were performed at x400. The measurements are reported as maxima and minima in parentheses, and the mean plus and minus the standard deviation of a number of measurements is given in parenthesis. The means are placed in italics.
RESULTS
Phylogenetic inference
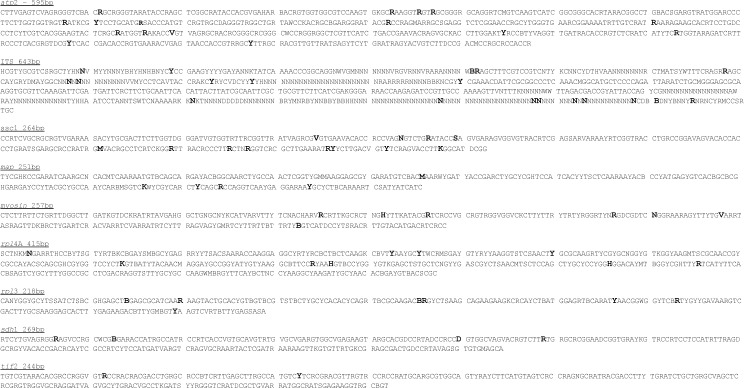
The LSU sequence data were excluded from further analysis since sequences were identical for all members of the Ustilago striiformis species complex (data not shown). All other loci showed SNPs within the U. striiformis cluster. The diagnostic bases (SNPs) with their specific positions are given in Fig. 6.
Fig. 6.
Alignment consensus sequences for the alignments used in this study with positions of diagnostic bases highlighted in bold face.
There were no supported conflicts in the topology of the trees of the single loci and the concatenated trees. Thus, the datasets were combined and used as concatenated for further analysis. The multigene tree based on nine different loci (Fig. 1) showed strong to maximum support for a monophyly of the U. striiformis species complex. If multiple specimens from one host species were included, these grouped together with strong to maximum support, except for the clades corresponding to U. scaura s. lat. (ME 64, ML 63, BA 0.99), U. brizae (ME 63, ML 68, BA 0.99), and U. agrostidis-palustris (ME 71, ML 68, BA 0.99), which received weak to strong support (Fig. 1).
Fig. 1.
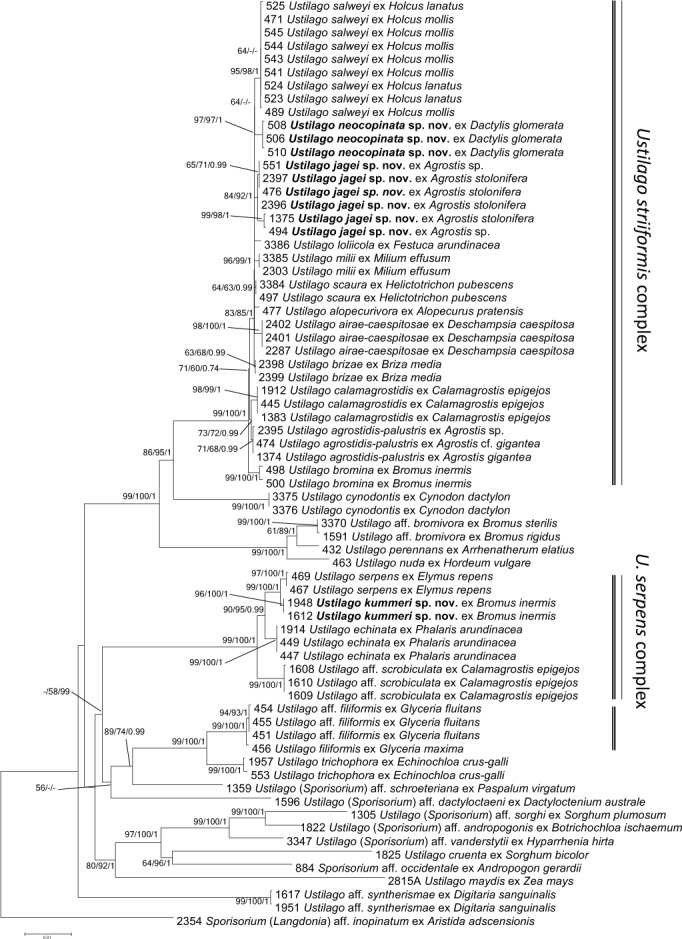
Phylogenetic tree based on Minimum Evolution analysis of nine loci (ITS, myosin, map, rpl3, tif2, ssc1, sdh1, rpl4A, atp2). Numbers on branches denote support in Minimum Evolution, Maximum Likelihood and Bayesian Analyses, in the respective order. Values below 55 % are denoted by ‘-‘. The bar indicates the number of substitutions per site.
A phylogenetic reconstruction (Fig. 2) with an additional 21 specimens but based on only half of the characters per specimen (ITS, atp2, and ssc1) revealed the same groups as the double-sized alignment, but expectedly with weaker statistical support. For example, the three weak to strongly supported lineages shown in Fig. 1 still grouped together, but with no or weak support (U. brizae – ME 64, ML -, BA 0.79; U. scaura s. lat. – ME -, ML -, BA 0.79; U. agrostidis-palustris – no support), highlighting the importance of gene selection.
Fig. 2.
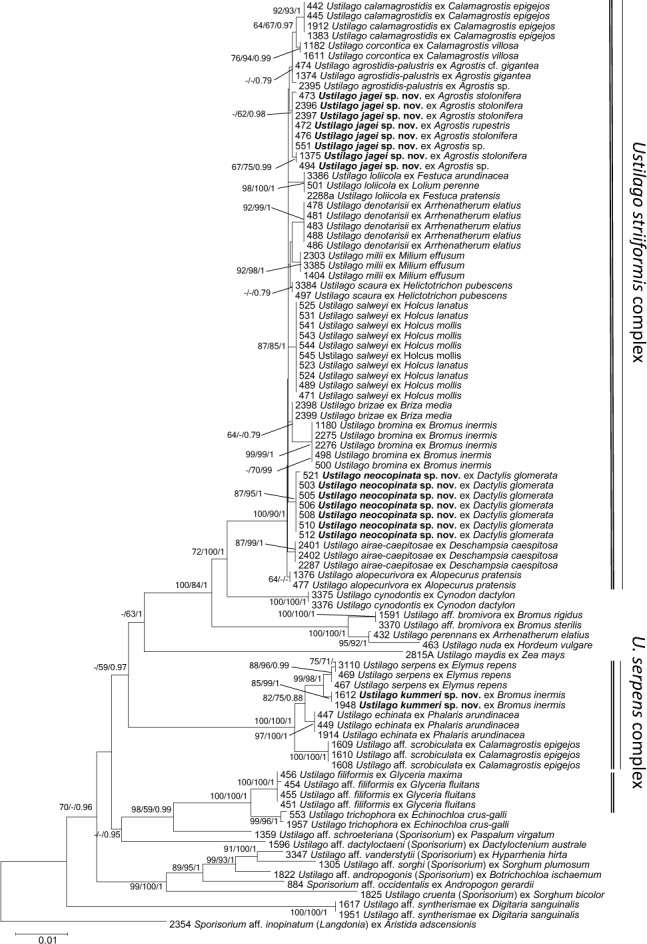

Phylogenetic tree based on Minimum Evolution analysis of three loci (ITS, ssc1, atp2). Numbers on branches denote support in Minimum Evolution, Maximum Likelihood and Bayesian Analyses, in the respective order. Values below 55 % are denoted by ‘-‘.. The bar indicates the number of substitutions per site.
In the phylogenetic reconstruction based on fewer genes (Fig. 2) additional specimens were included, which further supported the high degree of genetic differentiation in conjunction with the host species infected. Specimens from Festuca and Lolium grouped together with strong support, while the monophyly of the clade containing samples from Alopecurus species was unsupported to weakly supported (ME 64, ML -, BA -). Two monophyletic groups were absent from the tree with more loci (Fig. 1): one on Calamagrostis villosa and another on Arrhenatherum elatius. Both of these groups were highly supported (Calamagrostis: ME 76, ML 94, BA 0.99; Arrhenatherum: ME 92, ML 99, BA 1) in the tree based on fewer loci (Fig. 2).
In both phylogenetic trees (Figs 1,2), U. cynodontis was inferred as the sister species to the whole U. striiformis species complex. To illustrate the relationships within this species complex further, two additional phylogenetic trees with a reduced sampling and U. cynodontis as outgroup are shown in Figs 3 (9 loci) and 4 (3 loci). The support values and the topology were comparable to the phylogenetic reconstructions in Figs 1–2. In both phylogenetic trees, U. serpens on Elymus repens and on Bromus inermis grouped together with high to maximum support. This group clustered with two further lineages with larger echinulate spores compared to the U. striiformis species complex, which is considered a synapomorphy of this lineage.
Fig. 3.
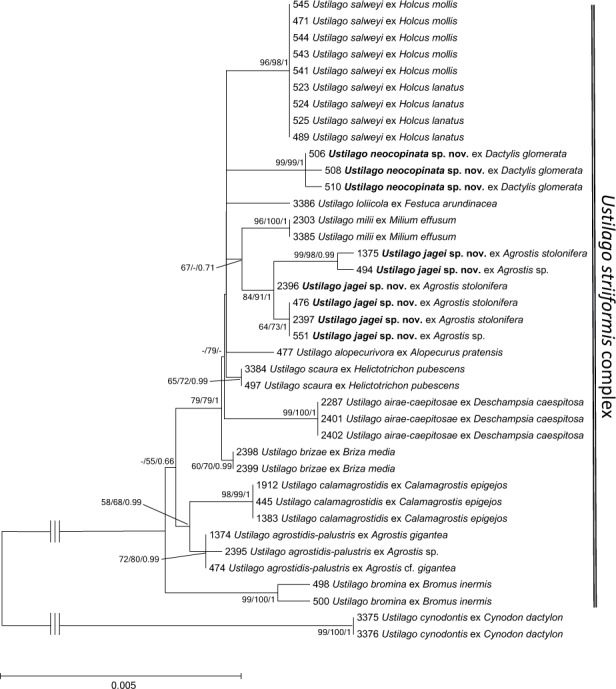
Phylogenetic tree based on Minimum Evolution analysis of nine loci (ITS, myosin, map, rpl3, tif2, ssc1, sdh1, rpl4A, atp2) detailed showing the Ustilago striiformis-complex with the outgroup U. cynodontis. Numbers on branches denote support in Minimum Evolution, Maximum Likelihood and Bayesian Analyses, in the respective order. Values below 55 % are denoted by ‘-‘. The bar indicates the number of substitutions per site.
Fig. 4.
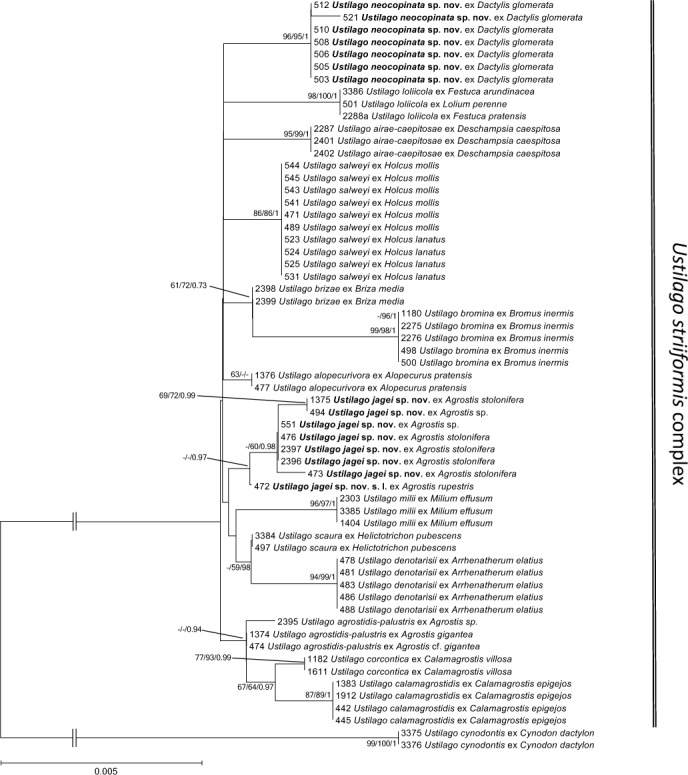
Phylogenetic tree based on Minimum Evolution analysis of three loci (ITS, ssc1, atp2) detailed showing the Ustilago striiformis-complex with the outgroup U. cynodontis. Numbers on branches denote support in Minimum Evolution, Maximum Likelihood and Bayesian Analyses, in the respective order. Values below 55 % are denoted by ‘-‘. The bar indicates the number of substitutions per site.
The resolution on the backbone was rather low, as highlighted also by the ambiguous placement of U. maydis, which was resolved as a sister group to the pathogens on the majority of panicoid hosts in the tree based on 9 loci (Fig. 1) with moderate to maximum support, while being inferred as a sister to the clade containing the U. species complex as well as the U. nuda species group with lacking to maximum support in the tree based on three loci (Fig. 2).
Morphology
The degree of overlap in morphological characteristics was too high in both species complexes to provide easily accessible characteristics for species delimitation (Fig. 5). The individual measurements are included in the species descriptions below and summarized in Table 3.
Fig. 5.
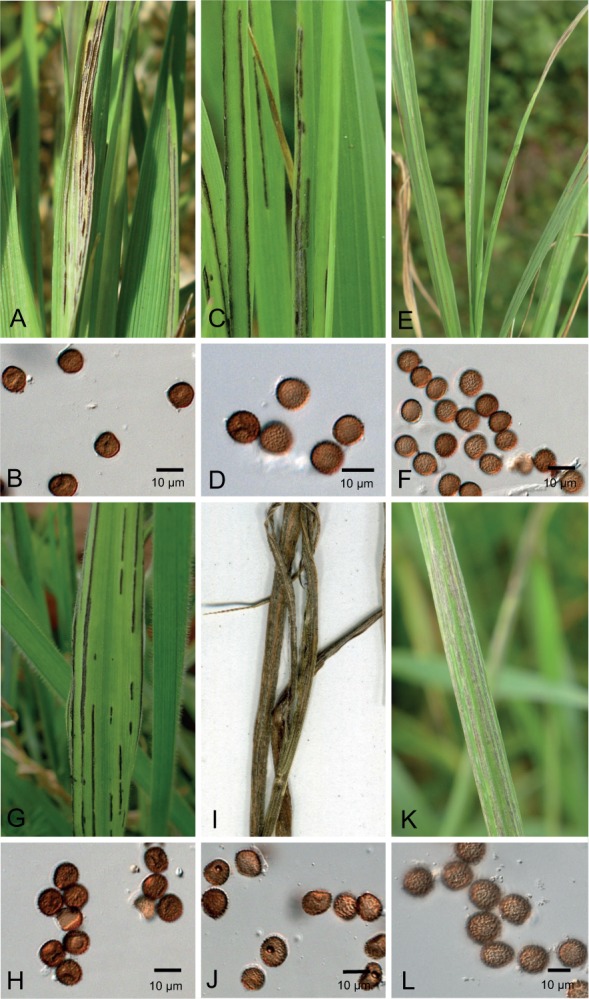

Sori and spores of Ustilago jagei (A–B), U. denotarisii (C–D), U. neocopinata (E–F), U. salweyi (G–H), U. kummeri (I–J), and U. serpens s. str. (K–L). A. Sori of U. jagei on Agrostis stolonifera (GLM-F047379); B. Teliospores seen by LM; C. Sori of U. denotarisii on Arrhenatherum elatius (GLM-F105836); D. Teliospores seen by LM; E. Sori of U. neocopinata on Dactylis glomerata (GLM-F107413); F. Teliospores seen by LM; G. Sori of U. salweyi on Holcus lanatus (GLM-F107417); H. Teliospores seen by LM; I. Sori of U. kummeri on Bromus inermis (GLM-F107435); J. Teliospores seen by LM; K. Sori of U. serpens s. str. on Elymus repens (GLM-F105827); and L. Teliospores seen by LM.
Table 3.
Measurements from 100 teliospores for four different species of the Ustilago striiformis-complex on Agrostis stolonifera, Dactylis glomerata, Arrhenatherum elatius, and Holcus lanatus, as well as two species of the Ustilago serpens-complex on Elymus repens and Bromus inermis.
| Ustilago striiformis-complex | Ustilago serpens-complex | |||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| U. jagei sp. nov. | U. denotarisii | U. neocopinata sp. nov. | U. salveii | U. serpens | U. kummeri sp. nov. | |||||||||||||
| on Agrostis stolonifera | on Arrhenatherum elatius | on Dactylis glomerata | on Holcus lanatus | on Elymus repens | on Bromus inermis | |||||||||||||
| spores | spores | spores | spores | spores | spores | |||||||||||||
| No. | length | width | l/b | length | width | l/b | length | width | l/b | length | width | l/b | length | width | l/b | length | width | l/b |
| 1 | 10.5 | 9.5 | 1.11 | 10 | 9 | 1.11 | 11 | 10.5 | 1.05 | 11.5 | 10.5 | 1.1 | 11 | 10.5 | 1.05 | 14 | 13 | 1.08 |
| 2 | 10 | 7.5 | 1.33 | 11.5 | 9 | 1.28 | 9.5 | 9.5 | 1 | 11.5 | 10 | 1.15 | 12 | 10 | 1.2 | 13.5 | 11 | 1.23 |
| 3 | 10.5 | 8.5 | 1.24 | 9 | 8 | 1.13 | 11 | 10 | 1.1 | 10 | 9.5 | 1.05 | 12.5 | 10 | 1.25 | 14.5 | 12 | 1.21 |
| 4 | 13.5 | 9.5 | 1.42 | 10 | 8 | 1.25 | 10.5 | 9.5 | 1.11 | 11 | 9.5 | 1.16 | 13 | 12 | 1.08 | 14 | 12.5 | 1.12 |
| 5 | 11 | 9 | 1.22 | 10.5 | 8.5 | 1.24 | 10.5 | 10 | 1.05 | 12 | 9.5 | 1.26 | 12.5 | 10.5 | 1.19 | 14 | 12 | 1.17 |
| 6 | 11 | 10 | 1.1 | 11.5 | 9 | 1.28 | 11 | 9.5 | 1.16 | 12 | 9 | 1.33 | 13 | 12.5 | 1.04 | 11.5 | 11.5 | 1 |
| 7 | 9.5 | 8 | 1.19 | 10.5 | 9.5 | 1.11 | 10 | 8.5 | 1.18 | 11 | 9 | 1.22 | 12.5 | 11.5 | 1.09 | 14 | 12 | 1.17 |
| 8 | 11 | 8 | 1.38 | 10.5 | 9.5 | 1.11 | 10.5 | 10 | 1.05 | 10.5 | 9 | 1.17 | 12.5 | 9.5 | 1.32 | 14 | 13.5 | 1.04 |
| 9 | 10.5 | 10 | 1.05 | 11.5 | 10 | 1.15 | 10.5 | 10 | 1.05 | 10 | 10 | 1 | 13.5 | 11 | 1.23 | 13 | 12.5 | 1.04 |
| 10 | 11.5 | 9 | 1.28 | 11.5 | 8.5 | 1.35 | 10.5 | 10.5 | 1 | 10.5 | 9.5 | 1.11 | 13 | 11 | 1.18 | 13.5 | 13.5 | 1 |
| 11 | 11.5 | 10 | 1.15 | 11 | 8 | 1.38 | 11 | 10 | 1.1 | 10.5 | 9.5 | 1.11 | 14.5 | 13.5 | 1.07 | 13.5 | 11.5 | 1.17 |
| 12 | 11.5 | 8 | 1.44 | 11 | 10 | 1.1 | 11 | 11 | 1 | 10.5 | 10 | 1.05 | 14.5 | 12 | 1.21 | 12.5 | 11 | 1.14 |
| 13 | 12 | 8 | 1.5 | 10.5 | 9 | 1.17 | 12 | 10.5 | 1.14 | 12.5 | 9 | 1.39 | 15.5 | 11 | 1.41 | 13.5 | 12.5 | 1.08 |
| 14 | 12 | 10.5 | 1.14 | 12 | 9 | 1.33 | 10 | 10 | 1 | 10 | 8 | 1.25 | 13 | 12.5 | 1.04 | 12 | 12 | 1 |
| 15 | 10 | 8.5 | 1.18 | 10.5 | 9 | 1.17 | 10.5 | 10.5 | 1 | 11 | 10 | 1.1 | 12.5 | 12 | 1.04 | 13.5 | 12.5 | 1.08 |
| 16 | 12 | 11.5 | 1.04 | 10.5 | 9.5 | 1.11 | 10.5 | 9.5 | 1.11 | 10.5 | 9.5 | 1.11 | 13 | 12.5 | 1.04 | 12 | 11.5 | 1.04 |
| 17 | 11 | 8 | 1.38 | 12 | 9 | 1.33 | 10 | 9 | 1.11 | 10 | 9 | 1.11 | 12 | 11.5 | 1.04 | 13.5 | 13 | 1.04 |
| 18 | 11 | 9.5 | 1.16 | 12.5 | 10.5 | 1.19 | 10 | 9.5 | 1.05 | 11.5 | 9.5 | 1.21 | 13 | 10.5 | 1.24 | 14.5 | 13 | 1.12 |
| 19 | 11 | 9 | 1.22 | 10 | 9 | 1.11 | 10.5 | 9.5 | 1.11 | 10 | 9 | 1.11 | 13 | 11.5 | 1.13 | 13 | 11.5 | 1.13 |
| 20 | 12 | 9.5 | 1.26 | 12.5 | 11 | 1.14 | 11 | 10.5 | 1.05 | 10.5 | 9.5 | 1.11 | 13 | 12 | 1.08 | 13.5 | 13 | 1.04 |
| 21 | 11 | 9.5 | 1.16 | 12.5 | 11.5 | 1.09 | 10.5 | 10 | 1.05 | 10.5 | 9 | 1.17 | 12.5 | 11 | 1.14 | 14.5 | 12 | 1.21 |
| 22 | 13 | 9.5 | 1.37 | 13.5 | 12 | 1.13 | 11.5 | 10.5 | 1.1 | 10 | 8.5 | 1.18 | 12 | 11.5 | 1.04 | 13.5 | 12.5 | 1.08 |
| 23 | 12.5 | 10 | 1.25 | 13.5 | 10 | 1.35 | 11 | 11 | 1 | 11.5 | 9 | 1.28 | 13 | 11.5 | 1.13 | 13 | 12.5 | 1.04 |
| 24 | 11.5 | 10 | 1.15 | 11.5 | 10.5 | 1.1 | 10 | 9.5 | 1.05 | 10 | 9.5 | 1.05 | 13.5 | 12 | 1.13 | 12.5 | 12 | 1.04 |
| 25 | 10.5 | 8.5 | 1.24 | 11.5 | 9.5 | 1.21 | 11 | 10.5 | 1.05 | 11 | 9.5 | 1.16 | 13 | 10.5 | 1.24 | 15 | 13.5 | 1.11 |
| 26 | 10.5 | 10 | 1.05 | 12.5 | 11 | 1.14 | 11 | 10.5 | 1.05 | 9.5 | 9 | 1.06 | 12 | 10.5 | 1.14 | 13 | 11.5 | 1.13 |
| 27 | 11 | 9 | 1.22 | 12.5 | 11.5 | 1.09 | 11 | 10 | 1.1 | 10.5 | 9 | 1.17 | 12.5 | 12 | 1.04 | 13.5 | 13.5 | 1 |
| 28 | 10.5 | 10.5 | 1 | 11 | 10.5 | 1.05 | 10.5 | 9 | 1.17 | 10 | 9.5 | 1.05 | 13 | 12 | 1.08 | 13.5 | 11.5 | 1.17 |
| 29 | 11 | 9.5 | 1.16 | 11 | 11 | 1 | 11 | 11 | 1 | 10 | 9 | 1.11 | 14 | 12.5 | 1.12 | 13.5 | 11.5 | 1.17 |
| 30 | 10.5 | 7.5 | 1.4 | 11 | 9.5 | 1.16 | 10 | 10 | 1 | 10 | 9 | 1.11 | 12 | 11 | 1.09 | 13 | 12.5 | 1.04 |
| 31 | 10.5 | 9 | 1.17 | 11 | 9 | 1.22 | 10 | 9.5 | 1.05 | 10 | 9 | 1.11 | 12.5 | 11.5 | 1.09 | 13 | 11 | 1.18 |
| 32 | 10 | 8.5 | 1.18 | 11.5 | 10.5 | 1.1 | 10 | 9.5 | 1.05 | 11 | 10 | 1.1 | 14.5 | 12.5 | 1.16 | 14 | 13 | 1.08 |
| 33 | 10.5 | 9.5 | 1.11 | 11 | 8.5 | 1.29 | 10.5 | 10.5 | 1 | 10 | 9 | 1.11 | 13 | 11.5 | 1.13 | 14 | 13 | 1.08 |
| 34 | 10.5 | 9.5 | 1.11 | 11.5 | 9 | 1.28 | 10.5 | 10.5 | 1 | 11 | 8.5 | 1.29 | 14 | 12.5 | 1.12 | 14 | 13 | 1.08 |
| 35 | 11.5 | 10 | 1.15 | 12.5 | 9.5 | 1.32 | 11 | 10 | 1.1 | 11 | 10 | 1.1 | 12 | 11.5 | 1.04 | 12.5 | 12 | 1.04 |
| 36 | 12 | 9 | 1.33 | 10.5 | 8.5 | 1.24 | 10.5 | 9.5 | 1.11 | 10 | 9 | 1.11 | 14.5 | 11.5 | 1.26 | 13 | 11 | 1.18 |
| 37 | 11 | 9.5 | 1.16 | 12.5 | 10.5 | 1.19 | 11 | 10.5 | 1.05 | 10 | 8.5 | 1.18 | 12.5 | 11.5 | 1.09 | 15 | 13.5 | 1.11 |
| 38 | 10.5 | 9 | 1.17 | 11 | 9.5 | 1.16 | 10.5 | 10 | 1.05 | 10.5 | 9.5 | 1.11 | 14 | 12 | 1.17 | 14 | 13 | 1.08 |
| 39 | 9.5 | 9 | 1.06 | 10.5 | 10 | 1.05 | 11 | 10 | 1.1 | 10.5 | 10.5 | 1 | 13 | 10 | 1.3 | 14 | 13.5 | 1.04 |
| 40 | 10 | 8.5 | 1.18 | 12.5 | 10.5 | 1.19 | 10.5 | 9.5 | 1.11 | 10 | 9 | 1.11 | 11.5 | 11 | 1.05 | 13 | 13 | 1 |
| 41 | 10.5 | 9.5 | 1.11 | 11 | 9.5 | 1.16 | 10 | 9.5 | 1.05 | 10 | 9 | 1.11 | 13.5 | 10.5 | 1.29 | 14.5 | 12.5 | 1.16 |
| 42 | 11.5 | 10.5 | 1.1 | 11.5 | 11 | 1.05 | 10 | 10 | 1 | 10.5 | 10 | 1.05 | 12.5 | 9.5 | 1.32 | 13 | 12 | 1.08 |
| 43 | 11 | 10.5 | 1.05 | 10 | 10 | 1 | 10.5 | 9.5 | 1.11 | 10.5 | 9.5 | 1.11 | 13.5 | 11 | 1.23 | 13.5 | 11.5 | 1.17 |
| 44 | 10 | 9 | 1.11 | 11 | 9.5 | 1.16 | 10.5 | 10 | 1.05 | 10 | 10 | 1 | 14 | 12 | 1.17 | 14 | 11.5 | 1.22 |
| 45 | 10.5 | 8.5 | 1.24 | 11.5 | 10 | 1.15 | 10.5 | 9.5 | 1.11 | 12 | 10 | 1.2 | 13.5 | 10.5 | 1.29 | 13 | 11.5 | 1.13 |
| 46 | 10.5 | 8 | 1.31 | 11.5 | 11.5 | 1 | 10.5 | 9.5 | 1.11 | 10.5 | 10 | 1.05 | 14 | 12 | 1.17 | 13.5 | 12 | 1.13 |
| 47 | 12.5 | 10.5 | 1.19 | 11 | 10.5 | 1.05 | 9.5 | 8.5 | 1.12 | 9.5 | 9.5 | 1 | 12 | 11.5 | 1.04 | 12.5 | 11.5 | 1.09 |
| 48 | 11 | 9.5 | 1.16 | 10.5 | 10 | 1.05 | 10 | 10 | 1 | 11.5 | 10 | 1.15 | 13.5 | 12 | 1.13 | 13.5 | 11.5 | 1.17 |
| 49 | 11 | 9.5 | 1.16 | 11.5 | 9.5 | 1.21 | 10.5 | 9.5 | 1.11 | 10.5 | 10 | 1.05 | 13 | 11.5 | 1.13 | 13 | 12 | 1.08 |
| 50 | 10 | 9.5 | 1.05 | 11.5 | 9.5 | 1.21 | 11 | 10.5 | 1.05 | 10.5 | 8 | 1.31 | 13.5 | 12 | 1.13 | 13 | 12.5 | 1.04 |
| 51 | 10 | 9.5 | 1.05 | 10 | 8.5 | 1.18 | 10 | 9 | 1.11 | 11 | 9 | 1.22 | 14 | 11.5 | 1.22 | 12.5 | 10.5 | 1.19 |
| 52 | 10.5 | 8.5 | 1.24 | 10.5 | 8 | 1.31 | 10 | 11 | 0.91 | 10.5 | 10.5 | 1 | 13.5 | 10.5 | 1.29 | 14.5 | 12 | 1.21 |
| 53 | 12 | 9.5 | 1.26 | 10 | 8.5 | 1.18 | 9 | 9 | 1 | 11.5 | 10 | 1.15 | 14 | 11.5 | 1.22 | 14 | 12 | 1.17 |
| 54 | 10.5 | 10.5 | 1 | 11 | 8.5 | 1.29 | 10 | 9.5 | 1.05 | 10.5 | 9 | 1.17 | 13 | 12 | 1.08 | 13.5 | 12 | 1.13 |
| 55 | 10.5 | 10.5 | 1 | 12 | 9.5 | 1.26 | 10 | 9.5 | 1.05 | 10 | 10 | 1 | 12 | 11 | 1.09 | 14 | 12 | 1.17 |
| 56 | 11 | 10 | 1.1 | 9.5 | 9 | 1.06 | 9.5 | 9.5 | 1 | 10.5 | 10.5 | 1 | 12 | 12 | 1 | 13 | 13 | 1 |
| 57 | 10.5 | 9 | 1.17 | 10 | 8.5 | 1.18 | 11.5 | 10.5 | 1.1 | 11 | 9.5 | 1.16 | 13 | 10.5 | 1.24 | 14 | 13 | 1.08 |
| 58 | 10 | 10 | 1 | 11.5 | 9.5 | 1.21 | 10 | 9 | 1.11 | 10.5 | 10 | 1.05 | 14.5 | 10.5 | 1.38 | 13 | 12.5 | 1.04 |
| 59 | 11 | 10 | 1.1 | 11 | 10 | 1.1 | 10 | 7.5 | 1.33 | 10.5 | 9.5 | 1.11 | 13 | 11.5 | 1.13 | 13.5 | 12 | 1.13 |
| 60 | 10.5 | 10.5 | 1 | 12 | 9.5 | 1.26 | 10 | 10 | 1 | 10 | 9.5 | 1.05 | 13 | 12 | 1.08 | 14.5 | 12 | 1.21 |
| 61 | 10.5 | 8.5 | 1.24 | 11 | 10 | 1.1 | 10.5 | 10 | 1.05 | 10.5 | 9.5 | 1.11 | 13.5 | 10 | 1.35 | 14.5 | 13 | 1.12 |
| 62 | 11.5 | 9 | 1.28 | 10.5 | 10 | 1.05 | 11 | 9 | 1.22 | 10.5 | 10 | 1.05 | 13 | 11.5 | 1.13 | 14.5 | 13 | 1.12 |
| 63 | 10.5 | 8.5 | 1.24 | 10.5 | 9.5 | 1.11 | 10.5 | 9.5 | 1.11 | 9.5 | 9.5 | 1 | 12.5 | 12 | 1.04 | 13.5 | 12 | 1.13 |
| 64 | 10.5 | 9.5 | 1.11 | 10.5 | 10 | 1.05 | 11 | 10.5 | 1.05 | 10 | 9.5 | 1.05 | 12.5 | 12 | 1.04 | 14 | 12.5 | 1.12 |
| 65 | 10 | 10 | 1 | 10 | 8.5 | 1.18 | 10.5 | 10.5 | 1 | 11 | 9 | 1.22 | 14.5 | 10.5 | 1.38 | 13 | 12.5 | 1.04 |
| 66 | 10.5 | 8.5 | 1.24 | 11.5 | 11 | 1.05 | 11 | 11 | 1 | 10.5 | 8.5 | 1.24 | 11.5 | 10.5 | 1.1 | 13 | 12.5 | 1.04 |
| 67 | 11 | 10.5 | 1.05 | 11 | 9.5 | 1.16 | 11.5 | 10 | 1.15 | 11 | 9.5 | 1.16 | 15 | 12 | 1.25 | 13.5 | 12.5 | 1.08 |
| 68 | 10.5 | 8.5 | 1.24 | 11 | 9.5 | 1.16 | 11 | 11 | 1 | 10.5 | 9.5 | 1.11 | 12.5 | 11 | 1.14 | 14 | 13 | 1.08 |
| 69 | 10 | 10 | 1 | 11.5 | 10.5 | 1.1 | 11.5 | 11 | 1.05 | 10 | 9 | 1.11 | 14 | 11 | 1.27 | 14.5 | 14 | 1.04 |
| 70 | 10 | 10 | 1 | 11.5 | 11 | 1.05 | 10 | 9.5 | 1.05 | 11 | 10 | 1.1 | 12 | 11 | 1.09 | 13.5 | 12.5 | 1.08 |
| 71 | 11 | 9 | 1.22 | 11 | 10 | 1.1 | 9 | 9 | 1 | 10.5 | 9 | 1.17 | 13 | 10.5 | 1.24 | 13 | 12.5 | 1.04 |
| 72 | 10 | 10 | 1 | 10.5 | 10 | 1.05 | 10 | 9.5 | 1.05 | 11 | 10.5 | 1.05 | 13 | 12 | 1.08 | 13.5 | 12.5 | 1.08 |
| 73 | 10 | 10 | 1 | 13 | 10 | 1.3 | 11 | 10 | 1.1 | 9.5 | 9 | 1.06 | 11.5 | 11.5 | 1 | 13.5 | 13 | 1.04 |
| 74 | 10.5 | 8 | 1.31 | 11 | 9 | 1.22 | 10.5 | 9.5 | 1.11 | 11 | 9.5 | 1.16 | 14 | 11 | 1.27 | 13.5 | 13 | 1.04 |
| 75 | 10 | 9.5 | 1.05 | 11.5 | 10 | 1.15 | 10.5 | 9.5 | 1.11 | 10.5 | 10.5 | 1 | 12 | 10.5 | 1.14 | 15.5 | 13.5 | 1.15 |
| 76 | 11.5 | 9 | 1.28 | 12 | 10.5 | 1.14 | 11 | 9.5 | 1.16 | 10.5 | 10 | 1.05 | 11.5 | 9 | 1.28 | 13 | 12.5 | 1.04 |
| 77 | 11 | 10 | 1.1 | 10.5 | 10 | 1.05 | 9 | 9 | 1 | 11.5 | 9.5 | 1.21 | 12.5 | 11 | 1.14 | 14 | 12.5 | 1.12 |
| 78 | 11.5 | 9.5 | 1.21 | 10.5 | 10 | 1.05 | 9.5 | 9.5 | 1 | 9.5 | 8.5 | 1.12 | 11 | 10.5 | 1.05 | 14.5 | 12 | 1.21 |
| 79 | 11 | 9 | 1.22 | 11.5 | 8.5 | 1.35 | 11 | 10 | 1.1 | 10 | 9 | 1.11 | 13 | 11 | 1.18 | 13.5 | 12 | 1.13 |
| 80 | 11.5 | 9.5 | 1.21 | 10.5 | 9.5 | 1.11 | 10 | 8.5 | 1.18 | 11 | 8.5 | 1.29 | 11.5 | 11 | 1.05 | 13.5 | 12.5 | 1.08 |
| 81 | 9.5 | 9.5 | 1 | 11 | 9.5 | 1.16 | 10.5 | 10 | 1.05 | 11.5 | 9 | 1.28 | 11.5 | 11 | 1.05 | 13.5 | 10.5 | 1.29 |
| 82 | 11 | 9.5 | 1.16 | 10.5 | 10 | 1.05 | 11 | 9.5 | 1.16 | 10.5 | 10 | 1.05 | 12.5 | 11 | 1.14 | 13 | 13 | 1 |
| 83 | 10.5 | 10.5 | 1 | 11.5 | 9 | 1.28 | 11.5 | 9.5 | 1.21 | 11 | 9.5 | 1.16 | 12.5 | 12 | 1.04 | 14.5 | 13 | 1.12 |
| 84 | 11.5 | 10 | 1.15 | 11 | 9 | 1.22 | 10 | 9.5 | 1.05 | 10 | 9.5 | 1.05 | 12 | 9.5 | 1.26 | 13.5 | 12.5 | 1.08 |
| 85 | 11 | 10 | 1.1 | 10 | 10 | 1 | 11 | 9.5 | 1.16 | 10 | 8 | 1.25 | 13.5 | 11 | 1.23 | 14 | 13 | 1.08 |
| 86 | 11 | 9 | 1.22 | 10.5 | 9.5 | 1.11 | 11.5 | 11 | 1.05 | 9.5 | 7.5 | 1.27 | 14 | 9.5 | 1.47 | 14 | 12 | 1.17 |
| 87 | 11 | 10.5 | 1.05 | 10.5 | 9 | 1.17 | 11.5 | 10.5 | 1.1 | 11 | 8.5 | 1.29 | 12.5 | 12 | 1.04 | 15 | 13.5 | 1.11 |
| 88 | 11 | 9 | 1.22 | 11 | 9.5 | 1.16 | 10 | 9 | 1.11 | 10 | 9 | 1.11 | 12 | 12 | 1 | 12.5 | 12.5 | 1 |
| 89 | 10 | 7.5 | 1.33 | 11.5 | 8.5 | 1.35 | 9.5 | 9 | 1.06 | 10 | 9.5 | 1.05 | 15 | 12.5 | 1.2 | 13.5 | 11.5 | 1.17 |
| 90 | 11 | 9.5 | 1.16 | 10 | 9.5 | 1.05 | 11 | 9.5 | 1.16 | 11 | 10 | 1.1 | 14.5 | 12 | 1.21 | 12 | 12 | 1 |
| 91 | 10.5 | 9 | 1.17 | 13.5 | 11 | 1.23 | 11 | 9.5 | 1.16 | 11 | 10.5 | 1.05 | 12 | 11.5 | 1.04 | 13.5 | 13 | 1.04 |
| 92 | 10.5 | 9 | 1.17 | 13.5 | 10.5 | 1.29 | 9.5 | 9.5 | 1 | 11.5 | 10 | 1.15 | 14 | 11 | 1.27 | 13 | 12 | 1.08 |
| 93 | 10 | 8.5 | 1.18 | 13 | 10.5 | 1.24 | 9.5 | 7.5 | 1.27 | 11.5 | 9.5 | 1.21 | 13.5 | 11 | 1.23 | 14 | 12.5 | 1.12 |
| 94 | 11 | 9.5 | 1.16 | 11.5 | 10.5 | 1.1 | 11 | 11 | 1 | 10.5 | 9.5 | 1.11 | 13 | 11 | 1.18 | 13.5 | 11.5 | 1.17 |
| 95 | 10.5 | 9 | 1.17 | 11 | 11 | 1 | 11 | 10 | 1.1 | 10.5 | 10 | 1.05 | 14.5 | 12 | 1.21 | 14.5 | 12 | 1.21 |
| 96 | 10.5 | 7.5 | 1.4 | 12 | 10.5 | 1.14 | 10.5 | 8.5 | 1.24 | 10.5 | 9.5 | 1.11 | 12.5 | 10.5 | 1.19 | 14.5 | 13 | 1.12 |
| 97 | 11.5 | 9.5 | 1.21 | 11 | 11 | 1 | 10.5 | 10 | 1.05 | 10.5 | 9.5 | 1.11 | 12.5 | 11.5 | 1.09 | 13 | 11 | 1.18 |
| 98 | 11.5 | 9.5 | 1.21 | 11 | 9 | 1.22 | 13 | 11 | 1.18 | 11 | 10 | 1.1 | 11.5 | 11 | 1.05 | 13.5 | 11.5 | 1.17 |
| 99 | 13.5 | 11.5 | 1.17 | 10.5 | 10 | 1.05 | 10 | 9 | 1.11 | 10.5 | 9.5 | 1.11 | 12.5 | 11.5 | 1.09 | 14.5 | 13.5 | 1.07 |
| 100 | 10.5 | 9.5 | 1.11 | 11 | 10 | 1.1 | 10 | 10 | 1 | 11 | 9.5 | 1.16 | 13.5 | 12.5 | 1.08 | 13.5 | 11.5 | 1.17 |
TAXONOMY
Based on our phylogenetic analyses, the following nomenclature and taxonomic changes are proposed for leaf stripe smuts caused by species of Ustilago. The positions given for the diagnostic bases refer to specific positions in the alignments as highlighted in the alignment consensus sequences in Fig. 4. Only selected synonyms are given here. For a complete synonymy reference should be made to Vánky (2012) and references therein.
Ustilago agrostidis-palustris W. H. Davis ex Ciferri, Ann. Mycol. 29: 54 (1931).
Type: USA: Wisconsin: Madison, on cultivated ‘redtop’ (i.e. Agrostis “palustris Huds.”, now Agrostis gigantea), 8 July 1921, W. H. & J. J. Davis (BPI 166994 – lectotype designated here, MBT 380628).
Confirmed host: Agrostis gigantea.
Confirmed distribution: Germany and USA.
Notes: Ustilago agrostidis-palustris can be distinguished from other leaf stripe smuts of the U. striiformis species complex based on its host specific occurrence on Agrostis gigantea s. lat. Furthermore, it differs in one diagnostic base from all other species of the U. striiformis-complex included in this study – in the sdh1 gene there is a C instead of a T at position 138 (Table 2, Fig. 6).
Table 2.
Diagnostic bases within the Ustilago striiformis and the Ustilago serpens complexes.
|
Gen Loci |
||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| atp2 | map | ssc1 | myosin | rpl4A | rpl3 | sdh1 | tif2 | ITS | ||||||||||
| U. striiformis-complex | Pos. | Base | Pos. | Base | Pos. | Base | Pos. | Base | Pos. | Base | Pos. | Base | Pos. | Base | Pos. | Base | Pos. | Base |
| on Agrostis gigantea | x | x | x | x | x | x | x | x | x | x | x | x | 138 | C / T | x | x | x | x |
| on Agrostis stolonifera and A. rupestris | 466 | A / G | x | x | x | x | x | x | x | x | 92 | A / G | x | x | x | x | x | x |
| on Alopecurus pratensis | 358 | A / G | 192 | G / T | x | x | 83 | T / C | x | x | x | x | x | x | x | x | x | x |
| on Arrhenatherum elatius | 346 | A / G | x | x | 182 | A / C | x | x | x | x | x | x | x | x | x | x | x | x |
| on Bromus inermis | 191, 244 | G / A | x | x | 232 | C / T | x | x | 228, 311 292 |
A / G |
x | x | x | x | 23 | A / G | 621 | C / T |
| on Calamagrostis epigejos | 91 | A / G | x | x | x | x | x | x | x | x | x | x | x | x | 65 | T / C | 102 | T / C |
| on Calamagrostis villosa | 535 | T / C | x | x | x | x | x | x | x | x | x | x | x | x | x | x | x | x |
| on Dactylis glomerata | x | x | x | x | 69, 198 | A / G | x | x | 120 | T / C | 40 | A / G | x | x | x | x | 617 | A / G |
| on Deschampsia caespitosa | 22, 94 | A / G | 227 | T / C | x | x | 133 | A / G | x | x | 199 | T / C | x | x | x | x | 576 | A / G |
| on Festuca spp. and Lolium spp. | x | x | x | x | 210, 214, 231 243 |
A / G |
x | x | x | x | x | x | x | x | x | x | x | x |
| on Holcus spp. | x | x | x | x | x | x | x | x | 85 | T / C | 133 | T / C | x | x | x | x | 103 | A / G |
| on Milium effusum | 301 | A / G | x | x | x | x | x | x | x | x | x | x | x | x | x | x | 206 | T / C |
| U. serpens-complex | Pos. | Base | Pos. | Base | Pos. | Base | Pos. | Base | Pos. | Base | Pos. | Base | Pos. | Base | Pos. | Base | Pos. | Base |
| on Bromus inermis | x | x | X | x | x | x | x | x | x | x | x | x | x | x | x | x | 260 629 |
C / T G / A |
| on Calamagrostis epigejos | 175 | G / A | 92 | A / C | 88, 99 | C / G | 70 | A / G | 7 | C / G | 28 | G / T | 13 | G / A | x | x | 215 | C / T |
| 181, 429, 496 | T / C | 203 | C / T | 93 | G / A | 225 | T / C | 91 | T / C | 25 | G / T | 522 | T / - | |||||
| 352 | A / G | 255 | G / T | 187 | G / T | 100 | A / G | 597 | A / T | |||||||||
| 606 | A / G | |||||||||||||||||
| on Elymus repens | x | x | x | x | x | x | 93 | A / G | 232 | T / C | x | x | x | x | x | x | 157 170 |
A / G C / T |
| on Phalaris arundinacea | 85 | G / A | 208 | A / G | x | x | 141 | C / A | x | x | 91 | T / C | 58, 256 | A / G | x | x | 19 | C / A |
| 156 | T / C | 146 | A / G | 38 | C / T | |||||||||||||
| 186, 596 | A / - | |||||||||||||||||
| 188, 604 | G / A | |||||||||||||||||
Slash (/) = instead of, x = no diagnostic bases.
Ustilago airae-caespitosae (Lindr.) Liro, Ann. Acad. Sci. Fenn., ser. A 17 (1): 71 (1924).
Basionym: Tilletia airae-caespitosae Lindr., Acta Soc. Fauna Flora Fenn. 26:15 (1904).
Type: Finland: Nyland: Helsingfors, Hagasund, on Aira caespitosa (i.e. Deschampsia caespitosa), 10 Aug. 1902, J. I. Lindroth [Vestergren, Micr. Rar. Sel. no. 806; Sydow, Ustil. no. 316] (M-0236198 – lectotype designated here, MBT 380628; from one of the several duplicate collections treated as “lectotype” by Lindeberg, Symb. Bot. Upsal. 16 (2): 135, 1959).
Confirmed host: Deschampsia caespitosa.
Confirmed distribution: Austria and Finland.
Notes: Within the Ustilago striiformis species complex, U. airae-caespitosae can be distinguished from other species based on the host-specific occurrence on Deschampsia caespitosa. Furthermore, it differs in six diagnostic bases from all other species within the U. striiformis species complex included in this study – in the atp2 gene it has an A instead of a G at position 22 and 94, in the map gene there is a T instead of a C at position 227, in the myosin gene there is an A instead of a G at position 133, in the rpl3 gene a T instead of a C at position 199, and an A instead of a G at position 576 in the ITS region (Table 2, Fig. 6).
Ustilago alopecurivora (Ule) Liro, Ann. Acad. Sci. fenn., ser. A 17 (1): 72 (1924).
Basionym: Tilletia alopecurivora Ule, Hedwigia 25: 113 (1886).
Synonyms: Uredo longissima var. megalospora Riess, in Rabenhorst, Herb. Viv. Myc. no. 1897 (1854).
Ustilago megalospora (Riess) Cif., Nuovo Giorn. Bot. Ital. 40: 261 (1933).
Type: Germany: Bavaria: Coburg, Hofgarten, on Alopecurus pratensis, June 1879, E. Ule (B – holotype lost); Berlin: Charlottenburg-Nord, Kolonie Königsdamm, slope of ditch, 9 Aug. 1988, H. Scholz (B 70 0014985 – neotype designated here, MBT 380629).
Confirmed host: Alopecurus pratensis.
Confirmed distribution: Germany.
Notes: Within the U. striiformis species complex, U. alopecurivora can be distinguished from other species based on the host-specific occurrence on Alopecurus pratensis. Furthermore, U. alopecurivora differs in three diagnostic bases from all other species within the U. striiformis species complex included in this study – in the atp2 gene, there is an A instead of a G at position 358, in the map gene there is a G instead of a T at position 192, and in the myosin gene there is a T instead of a C at position 83 (Table 2, Fig. 6).
Ustilago brizae (Ule) Liro, Ann. Acad. Sci. Fenn., Ser. A 17 (1): 74 (1924).
Basionym: Tilletia brizae Ule, Verh. Bot. Ver. Prov. Brandenb. 25: 214 (1884).
Type: Germany: Bavaria: Coburg, Rögener Berg, on Briza media, July 1879, E. Ule [Rabenhorst, Fungi Eur. no. 3604] (M-0147750 – lectotype designated here, MBT 380630; from one of the several duplicate collections treated as “lectotype” by Lindeberg, Symb. Bot. Upsal. 16(2): 135, 1959).
Confirmed hosts: Briza media.
Confirmed distribution: Austria and Germany.
Notes: Within the U. striiformis species complex, U. brizae can be distinguished from other species based on the host-specific occurrence on Briza media. Furthermore, U. brizae differs in one diagnostic base from all other species within the U. striiformis species complex included in this study, except U. bromina on Bromus inermis, in having a C instead of a T at position 621 in the ITS region, and differs from U. bromina by having an A instead of a G at position 223 in the ITS region (Table 2, Fig. 6).
Ustilago corcontica (Bubák) Liro, Ann. Acad. Sci. Fenn., Ser. A 17 (1): 383 (1924).
Basionym: Tilletia corcontica Bubák, Houby Ceské, Hemibasidii 2: 47 (1912).
Type: Czech Republic: on the crest of Riesengebirge Mts, on Calamagrostis halleriana (i.e. C. villosa), 20 July 1872, J. Gerhardt (BPI 172761 – lectotype designated here, MBT 380631; one of the “isolectotypes” of Lindeberg, Symb. Bot. Upsal. 16(2): 114, 1959).
Confirmed host: Calamagrostis villosa.
Confirmed distribution: Czech Republic and Germany.
Notes: Within the U. striiformis species complex, U. corcontica can be distinguished from other species based on the host-specific occurrence on Calamagrostis villosa. Furthermore, U. corcontica differs in one diagnostic bases from all other species within the U. striiformis species complex included in this study – in the atp2 gene there is an T instead of a C at position 535 (Table 2, Fig. 6).
Ustilago denotarisii A. A. Fischer v. Waldheim, Aperҁu Syst. Ustil.: 22 (1877); as “de Notarisii”.
Type: Italy: on Arrhenatherum spp. (not located but could also not be confirmed as lost; a neotype may need to be designated for this species in the future).
Confirmed hosts: Arrhenatherum species.
Confirmed distribution: Germany and Italy.
Notes: Spores globose to ovoid, standard range (9.0–)10.5– (av. 11.2)–12.0 (–13.5) × (8.0–) 9.0–(av. 9.7)–10.5(–12.0) μm, length/breadth ratio of 1.10- (av. 1.20) -1.38, olive-brown, and finely echinulate. Within the U. striiformis species complex, U. denotarisii can be distinguished from other species based on the host-specific occurrence on Arrhenatherum species. Furthermore, U. denotarisii differs in two diagnostic bases from all other species within the U. striiformis species complex included in this study – in the atp2 gene there is an A instead of a G at position 346, and in the gene ssc1 there is an A instead of a C at position 182 (Table 2, Fig. 6).
Ustilago echinata J. Schröt., Abh. Schles. Ges. Vaterl. Kult., Abth. Naturwiss.: 48: 4 (1870 [“1869”].
Type: Poland: Silesia: ‘Schwarzwasserbruch’, near Legnica, on Phalaris arundinacea, June 1869, W. G. Schneider [Rabenhorst, Fungi Eur. no. 1497] (FR – lectotype designated here, MBT 380632; one of the several duplicate collections previously treated as “lectotype” in Rabenhorst, Fungi Eur. No. 1497).
Reported hosts: Glyceria grandis, Phalaris arundinacea, and Scolochloa festucacea.
Confirmed host: Phalaris arundinacea.
Known distribution: Asia, North America, and Europe.
Notes: This species shares one sequence motif (AACCCAAC) at positions 20–27 in the ITS region with other coarsely ornamented stripe smuts (U. serpens clade in Fig. 1), and many SNPs which distinguish U. echinata from species of the U. striiformis-complex. Within the U. serpens-complex, U. echinata can be distinguished from other species based on its host-specific occurrence on Phalaris arundinacea (type host). Whether the other hosts of a similar ecotype are infected by the same species could not be clarified in the current study, but the high degree of host specificity observed in Ustilago renders it possible that specimens from other host genera will have to be described as new species. Furthermore, U. echinata differs in eight diagnostic bases from all other species within the U. serpens species complex included in this study – in the atp2 gene there is a G instead of an A at position 85, in the map gene there is an A instead of a G at position 208, in the myosin gene there is a C instead of an A at position 141 and a T instead of a C at position 156, in the rpl3 gene there is a T instead of a C at position 91 and an A instead of a G at position 146, in the sdh1 gene there is an A instead of a G and at positions 58 and 256, and in the ITS locus there is a C instead of an A at position 19, a C instead of a T at position 38, an A instead of a gap at position 186 and 596 and a G instead of an A at positions 188 and 604 (Tab. 2, Fig. 6).
Due to the generally narrow host specificity of smut fungi, it is conceivable that U. echinata will be revealed to be a species group.
Ustilago jagei J. Kruse & Thines, sp. nov.
MycoBank MB819627
Etymology: Named after mycologist Horst Jage from Kemberg (Germany), who has made significant contributions to the knowledge of phytopathogenic fungi and has enabled well-sampled phylogenetic investigations in various plant pathogens by his outstanding collections.
Diagnosis: Within the U. striiformis species complex, U. jagei can be distinguished from other species based on its host-specific occurrence on Agrostis stolonifera s. lat. Furthermore, U. jagei differs in two diagnostic bases from all other species within the U. striiformis species complex included in this study – in the atp2 gene there is an A instead of a G at position 466 and in the gene rpl3 there is an A instead of a G at position 92 (Table 2, Fig. 6).
Type: Germany: Saxony-Anhalt: Dessau, Kühnauer Sea, southern shore east-southeast of Großkuhnau, wayside, on Agrostis stolonifera, 16 Sept. 2001, H. Jage (GLM-F047379 – holotype).
Description: Sori as long narrow streaks parallel to vascular bundles, mostly in the leaves, rarely ascending into the inflorescence, initially covered by the epidermis of the plants, which soon frays. Spore mass dark brown to almost black, powdery. Infection systemic, infected plants usually sterile. Spores globose to ovoid, (9.5–) 10.0–(av. 10.9) –11.5(–13.5) x (7.5–) 8.5–(av. 9.3)–10.0(–11.5) μm, length/breadth ratio 1.04-(av. 1.24)-1.5, olive-brown, finely echinulate (Table 3, Figs 3–4).
Confirmed hosts: Agrostis rupestris and A. stolonifera.
Confirmed distribution: Germany and Switzerland.
Notes: It seems possible that U. jagei on Agrostis stolonifera s. lat. represents a species complex, and further investigations with more specimens and additional gene loci are needed to clarify this situation.
Ustilago kummeri J. Kruse & Thines, sp. nov.
MycoBank MB819628
Etymology: Named after the mycologist Volker Kummer from Potsdam (Germany), who has made significant contributions to the knowledge of phytopathogenic fungi and has enabled well-sampled phylogenetic investigations in various plant pathogens by his outstanding ability to recognise easily overlooked plant pathogens.
Diagnosis: Differs from species of the U. striiformis species complex in the larger spores and taller warts. Furthermore, U. kummeri shares one sequence motif at positions 20-27 (AACCCAAC) with other coarsely ornamented stripe smuts, and many SNPs distinguishing it from species of the U. striiformis species complex. Within the U. serpens-complex, U. kummeri can be distinguished from other species based on the host-specific occurrence on Bromus inermis. Furthermore, U. kummeri differs in two diagnostic bases from U. serpens on Elymus repens – in the ITS region there is an C instead of a G at position 260 and G instead of an A at position 629 (Table 2, Fig. 6).
Type: Germany: Brandenburg: Middlemark, Uetz: Hinterer Werder, southwest corner between Sacrow-Paretzer-Channel und Havel-Channel, on Bromus inermis, 19 June 2010, V. Kummer (GLM-F107435 – holotype; VK 2577/17 – isotype).
Description: Sori as long, narrow streaks parallel to vascular bundles, mostly in the leaves, rarely ascending to the inflorescence, initially covered by the epidermis of the plants, which soon frays. Spore mass dark brown, powdery. Infection systemic, infected plants mostly sterile. Spores ovoid to globose, (11.0-) 12.0- (av. 13.0) -14.0 (-15.5) × (9.0-) 10.5- (av. 11.5) -12.0 (-13.5), length/breadth ratio 1.04- (av. 1.15) -1.41, olive-brown, coarsely verrucose to echinulate (Table 3, Figs 5–6).
Confirmed host: Bromus inermis.
Confirmed distribution: Germany.
Notes: It seems likely that additional species will be discovered in the U. serpens clade once more stripe-smuts with coarse spore ornamentation will be scrutinised.
Ustilago loliicola Ciferri, Fl. Ital. Crypt., Par. I. Fungi, Fasc. 17: 345 (1938).
Type: Germany: Berlin: Berlin-Weissensee, on Lolium perenne, Sept. 1877, E. Ule [Rabenhorst, Fungi Eur. no. 2491] (FR – lectotype designated here, MBT 380633; from one of the several duplicate collections treated as “lectotype” by Lindeberg, Symb. Bot. Upsal. 16 (2): 136, 1959).
Confirmed hosts: Festuca arundinacea s. lat. and Lolium perenne.
Confirmed distribution: Germany.
Notes: Within the U. striiformis species complex, U. loliicola can be distinguished from other species based on the specific occurrence on the closely related hosts Festuca arundinacea s. lat. and Lolium perenne. Furthermore, U. loliicola differs in four diagnostic bases from all other species within the U. striiformis species complex included in this study – in the ssc1 locus there is an A instead of a G at positions 210, 214 and 231, and a T instead of a C at position 243 (Table 2, Fig. 6).
Ustilago milii (Fuckel) Liro, Ann. Acad. Sci. Fenn., ser. A 17 (1): 78 (1924).
Basionym: Tilletia milii Fuckel, Jb. nassau. Ver. Naturk. 23–24: 40 (1870).
Type: Germany: Hesse: Rabenkopf Mt., near Oestrich, on Milium effusum, L. Fuckel [Fungi Rhenani no. 2410] (FR – lectotype designated here, MBT 380634, from one of the several duplicate collections treated as “lectotype” in Fuckel, Fungi Rhenani no. 2410).
Confirmed host: Milium effusum.
Confirmed distribution: Germany.
Notes: Within the U. striiformis species complex, U. milii can be distinguished from other species based on the host-specific occurrence on Milium effusum. Furthermore, U. milii differs in two diagnostic bases from all other species within the U. striiformis species complex included in this study – in the atp2 gene there is an A instead of a G at position 301, and in the ITS there is a T instead of a C at position 206 (Table 2, Fig. 6).
Ustilago neocopinata J. Kruse & Thines, sp. nov.
MycoBank MB819630
Etymology: Highlights the unexpected finding that there are several distinct and host-specific species within the U. striiformis species complex.
Diagnosis: Within the U. striiformis species complex, U. neocopinata can be distinguished from other species based on the host-specific occurrence on Dactylis glomerata. Furthermore, U. neocopinata differs in five diagnostic bases from all other species within the U. striiformis species complex included in this study – in the ssc1 gene there is an A instead of a G at positions 69 and 198, in the rpl4A gene there is a T instead of a C at position 120, in the rpl3 gene there is an A instead of a G at position 40, and in the ITS region there is an A instead of a G at position 617 (Table 2, Figs 5–6).
Type: Germany: Bavaria: Upper Franconia, Kronach county, Wallenfels, in the direction of the sewage treatment plant downstream of Stumpfenschneidmühle, on Dactylis glomerata, 15 July 2012, J. Kruse (GLM-F107413 – holotype).
Description: Sori as long small streaks parallel to vascular bundles, mostly in the leaves, very rarely ascending to the inflorescence, initially covered by the epidermis of the plants, which soon frays. Spore mass dark brown to almost black, powdery. Infection systemic, infected plants mostly sterile. Spores mostly globose, rarely ovoid, (9.0–) 10.0– (av. 10.5)–11.0 (–13.0) × (7.5–) 9.0– (av. 9.8) –10.5 (–11) μm, length/breadth ratio 1.00- (av. 1.07) -1.18, olive-brown, finely echinulate (Table 3, Figs 5–6).
Notes: As the host is widespread throughout the Holarctic region, it is conceivable that the species will prove to have a much wider distribution range than currently known.
Ustilago salweyi Berk. & Broome, Ann. Mag. Nat. Hist. 5: 463 (1850).
Type: UK: Channel Islands: Guernsey, St Martin’s, on Holcus lanatus [originally misidentified as Dactylis glomerata fide Hubbard, in Stevenson, PIant Dis. Rep. 30: 57, 1946], 1847, T. Salwey (K-M – holotype; K-M00022071 – isotype).
Synonyms: Uredo striiformis Westend., Bull. Acad. R. Sci. Belg., cl. sci. 18: 406 (1852); as “striaeformis”.
Uredo salveii (Berk. & Broome) Oudem., Prodromus Florae Bataviae, 2nd edn,4: 180 (1866).
Tilletia debaryana A.A. Fisch. Waldh., in Rabenhorst, Fungi eur. no. 1097 (1867).
Tilletia striiformis (Westend.) Magnus, Malpighia 1: 8 (1875).
Ustilago striiformis (Westend.) Niessl, Hedwigia 15: 1 (1876).
Tilletia salveii (Berk. & Broome) P. Karst., Bidrag. Kännedom. Finlands Naurt. Folk. 6: 102 (1884).
Confirmed hosts: Holcus lanatus and H. mollis.
Confirmed distribution: Belgium, Germany, and UK.
Notes: Spores globose to ovoid, standard range (9.5–)10.0–(av. 10.6) –11.0 (–12.5) × (7.5–) 9.0–(av. 9.4)–10.0(–10.5) μm, finely echinulate, length/breadth ratio 1.00–(av. 1.15)–1.39. Within the U. striiformis species complex, U. salweyi can be distinguished from other species based on the host-specific occurrence on Holcus lanatus and H. mollis. Furthermore, U. salweyi differs in three diagnostic bases from all other species within the striiformis species complex included in this study – in the rpl4A gene there is a T instead of a C at position 85, in rpl3 there is a T instead of a C at position 133, and in the ITS region there is an A instead of a G at positions 103 (Table 2, Fig. 6).
The original host was misidentified as Dactylis glomerata, but this was found to be incorrect and actually Holcus lanatus by the leading grass specialist C.E. Hubbard (in Stevenson 1946). David Hawksworth also studied the type materials in K-M and concurs. Hosts in their vegetative stage can be misidentified, as some characteristics, such as leaf shape, ligula, and general habit can be modified as a consequence of infection.
Ustilago scaura Liro s. lat. , Ann. Acad. Sci. Fenn., ser. A, 17(1): 73 (1924).
Replaced name: Tilletia avenae Ule, Verh. Bot. Vereins Prov. Brandenburg 25: 214 (1884).
Type: Germany: Bavaria: Coburg, Fortress, on Avena pratensis (i.e. Helictotrichon pratense), June 1879, E. Ule (s. n. – lost); Hesse: county Tann/Rhön, at Galgenmount, on Avena pubescens [now, Helictotrichon pubescens], 16 Sept. 1990, H. Scholz (B 70 0014830 – neotype designated here, MBT 380637).
Non Ustilago avenae (Pers.) Rostrup, Overs. K. danske Vidensk. Selsk. Forh. Medlemmers Arbeider: 13 (1890).
Confirmed host: Helictotrichon pubescens, H. pratense?
Confirmed distribution: Germany.
Notes: Within the U. striiformis species complex, U. scaura s. lat. can be distinguished from other species based on the host-specific occurrence on Helictotrichon pratense and H. pubescens. Furthermore, U. scaura s. lat. differs in one diagnostic base from all other species within the U. salweyi species complex included in this study, except U. denotarisii on Arrhenatherum spp., in having a T instead of a C at position 628 in the ITS region, and from U. denotarisii on Arrhenatherum elatius in having a 13 nucleotide deletion at positions 222-241 in the ITS alignment (Table 2, Fig. 6).
Since the type has been lost, we designate a neotype for Ustilago scaura with material on the closely related H. pubescens.
Ustilago scrobiculata Liro, Ann. Acad. Sci. Fenn., ser. A 17(1): 68 (1924).
Type: Finland: Nyland: Pornainen, Kirveskoski, on Calamagrostis arundinacea, 9 Aug. 1916, T. Putkonen & J. I. Liro (H – lectotype, designated by Lindeberg, Symb. Bot. Upsal. 16 (2): 130 (1959).
Synonym: ? Ustilago deyeuxiae L. Guo, Mycosystema 6: 51 (1993).
Reported hosts: Calamagrostis spp. (see Vánky 2012: 1265).
Reported distribution: Asia and Europe.
Notes: This species shares one sequence motif with other coarsely ornamented stripe smuts (AACCCAAC at positions 20–27), which distinguishes it from species of the Ustilago striiformis species complex, and many additional single SNPs. Within the U. serpens species complex, U. scrobiculata differs in 21 diagnostic bases from other species (Table 2, Fig. 6). It seems possible that U. deyeuxiae has not been sampled on Calamagrostis arundinacea, as the host of U. deyeuxiae is given as “Deyeuxia arundinacea” by Guo (1993), which is often seen as a synonym of D. pyramidalis in Asian literature (e.g. Shenglian et al. 2006). Thus, it seems possible that the species needs to be reconsidered as independent from U. scrobiculata once sequence data from the type specimen become available.
Ustilago serpens (P. Karst.) B. Lindeb., Symb. Bot. Upsal. 16(2): 133 (1959).
Basionym: Tilletia serpens P. Karst., Fungi Fenn. Exs., fasc. 6 : no. 599 (1866).
Type: Finland: Merimasku, on “Dactylis glomerata” [re-determined as Elymus repens by Lindeberg, Symb. Bot. Upsal. 16(2): 133, 1959], July 1862, P. Karsten [Fungi Fenn. Exs no. 599] (HUV 10432 – lectotype designated here; MBT 380638 from one of the several duplicate collections treated as “lectotype” by Lindeberg, Symb. Bot. Upsal. 16(2): 133, 1959).
Confirmed host: Elymus repens.
Confirmed distribution: Finland and Germany.
Notes: The spores are small to medium sized, (11.5–) 13.0– (av. 13.5) –14.5 (–15.5) × (10.5–) 11.5 (av. 12.5) –13.0 (–14.0) μm, with a length/breadth ratio of 1.00–(av. 1.09)–1.23 and with coarsely verrucose ornamentation. This species shares one sequence motif with other coarsely ornamented stripe smuts (AACCCAAC at position 20–27), which distinguishes it from species of the U. striiformis species complex and many additional SNPs. Within the U. serpens-complex, U. serpens can be distinguished from other species based on four diagnostic bases: in the myosin gene there is an A instead of G at position 93, in rpl4A gene there is a T instead of a C at position 232 and in the ITS locus there is a C instead of a T at position 260, and a G instead of an A at position 629 (Table 2, Fig. 6).
Vánky (2012) lists several additional hosts for U. serpens. Due to the narrow specialization of stripe-smut revealed in this study, however, it seems likely that these harbour several distinct species. Until sequence data become available for these host-pathogen combinations, Ustilago on these other hosts is probably best referred to as U. serpens s. lat.
DISCUSSION
In this study, the closely related species of the Ustilago striiformis-complex and some other leaf stripe Ustilago smuts were investigated using multigene phylogenetic reconstructions to clarify their relationships. In total, 62 specimens of the U. striiformis species complex (incl. U. calamagrostidis) and four other leaf stripe smuts (U. echinata, U. filiformis, U. scrobiculata, and U. serpens s. lat.) were studied.
Phylogenetic analyses provided strong support for the polyphyly of the leaf-stripe smuts within Ustilago. However, the multilocus-based phylogenetic trees support the monophyly of the U. striiformis species complex, in contrast to the analysis by Savchenko et al. (2014a), where it was concluded that the U. striiformis group was polyphyletic and the segregation of two species was necessary to render it monophyletic. That interpretation was mainly based on a combined LSU-ITS tree of U. striiformis species, where U. bromina and U. nunavutica were located outside the U. striiformis s. lat. clade. Because of this conflicting result, the ITS region of the type specimen of U. bromina was sequenced (Table 1) and compared with the deposited GenBank sequences of Savchenko et al. (2014a). The type specimen of U. bromina on Bromus inermis had an ITS sequence nearly identical (except for a base exchange in a poly A/T region) with the other specimens identified as this species in the current study. It differed in nine bases compared to the three sequences labelled as U. bromina in Savchenko et al. (2014a). It is conceivable that these specimens belong to another undescribed smut species (the three sequences were obtained from material from Israel and USA, while the type collection was from Germany), or the quality of the sequences was not optimal; almost all differences in the sequences from Savchenko et al. (2014a) in comparison to the sequences from this study were located behind a poly A/T site, which necessitated re-sequencing for several of the specimens used in this study. Furthermore, misidentification of the host plant seems also possible, as no records were found for the occurrence of Bromus inermis in the floras of Israel (http://flora.org.il/en/plants/) or Palestine (Feinbrun-Dothan 1986).
Ustilago nunavutica was the second species that led Savchenko et al. (2014a) to assume that the U. striiformis species complex was polyphyletic. Comparing the ITS and LSU sequences of U. nunavutica with sequences from the current study, the LSU sequence used by Savchenko et al. (2014a) showed several SNPs (data not shown), while all other U. striiformis samples investigated in this study were identical in the LSU region. In contrast, the ITS sequence of U. nunavutica has only few SNPs in comparison to other members of the U. striiformis species complex and is identical with U. neocopinata. It seems possible that the LSU sequence of U. nunavutica either was of bad quality or shows the amplification of a contaminant smut fungus. However, as the genera Puccinellia and Dactylis are not closely related (Schneider et al. 2009) and very high host specificity has been revealed for the closely related species of the U. striiformis species complex in this study, it is unlikely that U. neocopinata and U. nunavutica are conspecific.
In agreement with Stoll et al. (2005) and Spooner & Legon (2006), we found that U. calamagrostidis and U. corcontica belonged to the U. striiformis species complex. However, further resolution within the U. striiformis species complex was only achieved when the protein-coding loci introduced by Kruse et al. (2017b) were employed. The trees revealed a host genus or host species specific occurrence for almost all lineages within the U. striiformis species complex, thus they should be treated as distinct species, supported by the observations of Liro (1924). All specimens from a single host species formed a clade according to the host species (or the host genus, in case of Holcus), with the exception of the rather closely related species Lolium perenne and Festuca arundinacea (Malik & Thomas 1966, Catalán et al. 2004, Hand et al. 2010). As most of these clades received high to maximum support, they should be considered to represent distinct species, which can be distinguished based on the host and diagnostic SNPs (Fig. 6). For most of the 14 lineages of the U. striiformis species complex validly published names are available, necessitating the description of only two new species in this complex, U. neocopinata on Dactylis glomerata and U. jagei on Agrostis stolonifera s. lat. Vánky (2012) and Savchenko et al. (2014a) mentioned that different species on different hosts within this complex vary remarkably in spore shape, size, and ornamentation. However, morphological variation was observed to be high even within the same host species in the current study and also by Vánky (2012). Thus it is difficult to distinguish these closely related species based on morphology, necessitating the consideration of hosts and SNPs for diagnosis. The host range of at least two species of Ustilago parasitic to Agrostis could not be inferred with certainty, as both ITS and chloroplast loci did not resolve closely related species in the A. stolonifera and A. gigantea clusters (Amundsen & Warnke 2012).
While investigating synonymies of the U. striiformis species complex, it was found that the name U. salweyi is the correct name for the stripe smut on Holcus lanatus. Stevenson (1946) flagged U. salweyi as a “nomen ambiguum”, although no action was taken to formally reject the name. Following the ICN (McNeill et al. 2012), the name U. salweyi has priority over Uredo striiformis as it was published two years earlier (Berkeley & Broome 1850: 463). Although the group generally referred to as the U. striiformis-group does not contain a species with that as the correct name, as it is still included as a synonym we feel that it is best to continue to use “U. striiformis-group” or “species complex” for these fungi as it is so well established and recalls the symptoms all species of the complex exhibit, although this feature is shared by some leaf-stripe smuts not belonging to this complex.
The species within the U. striiformis species complex have sometimes been recognised as special forms based on infection trials (Liro 1924, Davis 1930, 1935, Fischer 1940). However, it has been shown for various biotrophic pathogens that the special form concept, in which there is a population continuum with somewhat specialised forms, cannot be upheld (Göker et al. 2004, Lutz et al. 2005, Kemler et al. 2009, Thines et al. 2009, Ploch et al. 2011, Savchenko et al. 2014b, Choi & Thines 2015).
Similar to the situation in the U. striiformis species complex, Ustilago serpens s. lat. on different hosts clustered in phylogenetically distinct subgroups. As the type host for U. serpens is Elymus repens, the collections from Bromus inermis warrants recognition as a new species. Ustilago serpens is another example illustrating the narrow host specialization among smut fungi. As for both the coarsely ornamented stripe-smuts (U. serpens clade) and the finely ornamented stripe smuts (U. striiformis clade) only a subset of the known hosts could be included in the current study. It is therefore conceivable that some older names published for specific host-pathogen combinations in these groups warrant recognition and several new species await discovery.
With respect to the global phylogeny of Ustilago it is noteworthy that even based on nine loci the backbone of the phylogenetic tree was only poorly resolved. Conflicting supported topologies were inferred with respect to the phylogenetic position of U. maydis in the reconstructions based on three (sister to a clade comprising, among others, the U. nuda and the U. salweyi clade) and nine loci (sister to a clade comprising the majority of smuts on panicoid grasses).
This highlights the high degree of uncertainty that there still is with respect to the global phylogeny of Ustilago s. lat. (Thines 2016). Considering the diversity of anatomical characteristics and disease syndromes caused, many of which have arisen several times independently (such as the stripe-smut habit; McTaggart et al. 2012a, b, c), any splitting of Ustilago s. lat. into smaller genera as suggested by McTaggart et al. (2012a, 2016) is probably premature and might become obsolete due to the high degree of parallel evolution and associated homoplasy.
Acknowledgments
This study was funded by the LOEWE research funding programme in the framework of the Cluster for Integrative Fungal Research (IPF). We are grateful to the curators of B (Berlin), GLM (Görlitz), M (Munich), and TUB (Tübingen) for allowing the investigation of specimens in their keeping, and David Hawksworth for important information on Ustilago salweyi types in K-M (Kew fungarium) and nomenclatural corrections. We furthermore want to thank the curators of B (Berlin), HBG (Hamburg) and L (Leiden) for searching for the type collection of Ustilago scaura. We thank the private collectors Horst Jage and Volker Kummer for allowing us to include some of their collections in this study. We want to thank Bagdevi Mishra for the opportunity to use TrEase (thines-lab@senckenberg.de/trease) for phylogenetic analysis. Furthermore, we are grateful to Y.-J. Choi for help with the primer design for an internal ITS primer and to Reuel Bennett for proof-reading an initial draft of the manuscript. MT and JK conceived the study; JK made collection trips; FK, HJ, HR, HZ, UR, WD, and VK provided material; JK performed laboratory experiments and microscopy, and also analysed the data; JK and MT interpreted the data and prepared the manuscript with contributions from the other authors.
REFERENCES
- Amundsen K, Warnke S. (2012) Agrostis species relationships based on trnL-trnF and atpI-atpH Intergenic Spacer Regions. Hortscience 47: 18–24. [Google Scholar]
- Begerow D, Stoll M, Bauer R. (2006) A phylogenetic hypothesis of Ustilaginomycotina based on multiple gene analyses and morphological data. Mycologia 98: 906–916. [DOI] [PubMed] [Google Scholar]
- Berkeley MJ, Broome CE. (1850) Notices of British fungi. Annals and Magazine of Natural History 5: 455–467. [Google Scholar]
- Catalán P, Torrecilla P, Ángel J, Rodríguez L, Olmstead RG. (2004) Phylogeny of the festucoid grasses of subtribe Loliinae and allies (Poeae, Pooideae) inferred from ITS and trnL–F sequences. Molecular Phylogenetics and Evolution 31: 517–541. [DOI] [PubMed] [Google Scholar]
- Choi Y-J, Thines M. (2015) Host jumps and radiation, not co-divergence drives diversification of obligate pathogens: a case study in downy mildews and Asteraceae. PLoS ONE 10(7): e0133655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis WH. (1930) Two physiologic forms of Ustilago striaeformis (Westd.) Niessl. Phytopathology 20: 65–74. [Google Scholar]
- Davis WH. (1935) Summary of investigations with Ustilago striaeformis parasitizing some common grasses. Phytopathology 25: 810–817. [Google Scholar]
- Denchev C, Giraud T, Hood ME. (2009) Three new species of anthericolous smut fungi on Caryophyllaceae. Mycologia Balcanica 6: 79–84. [Google Scholar]
- Feinbrun-Dothan N. (1986) Flora Palaestina. Vol. 4. Jerusalem: Israel Academy of Sciences and Humanities, Section of Sciences. [Google Scholar]
- Fischer GW. (1940) Fundamental studies on the stripe smut of grasses (Ustilago striaeformis) in the Pacific Northwest. Phytopathology 30: 93–118. [Google Scholar]
- Fischer GW. (1953) Manual of the North American Smut Fungi. New York: Ronald Press. [Google Scholar]
- Göker M, Riethmüller A, Voglmayr H, Weiss M, Oberwinkler F. (2004) Phylogeny of Hyaloperonospora based on nuclear ribosomal internal transcribed spacer sequences. Mycological Progress 3: 83–94. [Google Scholar]
- Guo L. (1993) Ustilago deyeuxiae sp. nov. and three smut species new to China. Mycosystema 6: 51–55. [Google Scholar]
- Hand ML, Cogan NOC, Stewart AV, Forster JW. (2010) Evolutionary history of tall fescue morphotypes inferred from molecular phylogenetics of the Lolium-Festuca species complex. BMC Evolutionary Biology 10: 303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heller A, Thines M. (2009) Evidence for the importance of enzymatic digestion of epidermal walls during subepidermal sporulation and pustule opening in white blister rusts (Albuginaceae). Mycological Research 113: 657–667. [DOI] [PubMed] [Google Scholar]
- Katoh K, Standley DM. (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Molecular Biology and Evolution 30: 772–780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kemler M, Lutz M, Göker M, Oberwinkler F, Begerow D. (2009) Hidden diversity in the non-caryophyllaceous plant-parasitic members of Microbotryum (Pucciniomycotina: Microbotryales). Systematics and Biodiversity 7: 297–306. [Google Scholar]
- Kruse J, Choi Y-J, Thines M. (2017a) New smut-specific primers for the ITS barcoding of Ustilaginomycotina. Mycological Progress 16: 213–221. [Google Scholar]
- Kruse J, Mishra B, Choi Y-J, Sharma R, Thines M. (2017b) New smut-specific primers for multilocus genotyping and phylogenetics of Ustilaginaceae. Mycological Progress 16: 1–9. [Google Scholar]
- Liro JI. (1924) Die Ustilagineen Finnlands. Annales Academiae Scientiarum Fennicae, series A 17(1): 1–636. [Google Scholar]
- Lutz M, Göker M, Piatek M, Kemler M, Begerow D, Oberwinkler F. (2005) Anther smuts of Caryophyllaceae: molecular characters indicate host-dependent species delimitation. Mycological Progress 4: 225–238. [Google Scholar]
- Malik CP, Thomas PT. (1966) Karyotypic studies in some Lolium and Festuca species. Caryologia 19: 167–196. [Google Scholar]
- McNeill J, Barrie FR, Buck WR, Demoulin V, Greuter W, et al (2012) International Code of Nomenclature for algae, fungi, and plants (Melbourne Code) adopted by the Eighteenth International Botanical Congress Melbourne, Australia, July 2011. [Regnum Vegetabile No. 154.] Königstein: Koeltz Scientific Books. [Google Scholar]
- McTaggart AR, Shivas RG, Geering AD, Callaghan B, Vánky K, Scharaschkin T. (2012a) Soral synapomorphies are significant for the systematics of the Ustilago-Sporisorium–Macalpinomyces complex (Ustilaginaceae). Persoonia 29: 63–77. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McTaggart AR, Shivas RG, Geering ADW, Vánky K, Scharaschkin T. (2012b) A review of the Ustilago–Sporisorium–Macalpinomyces complex. Persoonia 29: 55–62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McTaggart AR, Shivas RG, Geering ADW, Vánky K, Scharaschkin T. (2012c) Taxonomic revision of Ustilago, Sporisorium and Macalpinomyces. Persoonia 29: 116–132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McTaggart AR, Shivas RG, Boekhout T, Oberwinkler F, Vánky K, et al (2016) Mycosarcoma (Ustilaginaceae), a resurrected generic name for corn smut (Ustilago maydis) and its close relatives with hypertrophied, tubular sori. IMA Fungus 7: 309–315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O’Donnell K. (1993) Fusarium and its near relatives. In: The Fungal Holomorph: mitotic, meiotic and pleomorphic speciation in fungal systematics (Reynolds DR, Taylor JW, eds.): 225–233. Wallingford: CAB International. [Google Scholar]
- Piątek M, Lutz M, Chater AO. (2013) Cryptic diversity in the Antherospora vaillantii complex on Muscari species. IMA Fungus 4: 5–19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ploch S, Telle S, Choi Y-J, Cunnington J, Priest M, et al (2011) The molecular phylogeny of the white blister rust genus Pustula reveals a case of underestimated biodiversity with several undescribed species on ornamentals and crop plants. Fungal Biology 115: 214–219. [DOI] [PubMed] [Google Scholar]
- Ridgway KP, Duck JM, Young JPW. (2003) Identification of roots from grass swards using PCR-RFLP and FFLP of the plastid trnL (UAA) intron. BMC Ecology 3(8): 1–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ronquist F, Huelsenbeck JP. (2003) MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19: 1572–1574. [DOI] [PubMed] [Google Scholar]
- Savchenko KG, Carris LM, Castlebury LA, Heluta VP, Wasser SP, Nevo E. (2014a) Stripe smuts of grasses: one lineage or high levels of polyphyly? Persoonia 33: 169–181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Savchenko KG, Carris LM, Castlebury LA, Heluta VP, Wasser SP, Nevo E. (2014b) Revision of Entyloma (Entylomatales, Exobasidiomycetes) on Eryngium. Mycologia 106: 797–810. [DOI] [PubMed] [Google Scholar]
- Schneider J, Döring E, Hilu KW, Röser M. (2009) Phylogenetic structure of the grass subfamily Pooideae based on comparison of plastid matK gene–3’trnK exon and nuclear ITS sequences. Taxon 58: 405–424. [Google Scholar]
- Shenglian L, Wenli C, Phillips SM. (2006) 87. Deyeuxia Clarion ex P. Beauvois, Ess. Agrostogr. 43. 1812. In: Flora of China (Zhengyi W, Raven PH, Deyuan H, eds) 22: 348–359. Beijing: Science Press. [Google Scholar]
- Spooner BM, Legon NW. (2006) Additions and amendments to the list of British smut fungi. Mycologist 20: 90–96. [Google Scholar]
- Stamatakis A. (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30: 1312–1313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stevenson JA. (1946) A nomenclatorial discussion of Ustilago striiformis. PIant Disease Reporter 30: 53–59. [Google Scholar]
- Stoll M, Begerow D, Oberwinkler F. (2005) Molecular phylogeny of Ustilago, Sporisorium, and related taxa based on combined analyses of rDNA sequences. Mycological Research 109: 342–356. [DOI] [PubMed] [Google Scholar]
- Stoll M, Piepenbring M, Begerow D, Oberwinkler F. (2003) Molecular phylogeny of Ustilago and Sporisorium species (Basidiomycota, Ustilaginales) based on internal transcribed spacer (ITS) sequences. Canadian Journal of Botany 81: 976–984. [Google Scholar]
- Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. (2013) MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Molecular Biology and Evolution 30: 2725–2729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thines M. (2016) Proposal to conserve the name Ustilago (Basidiomycota) with a conserved type. Taxon 65: 1170–1171. [Google Scholar]
- Thines M, Choi Y-J, Kemen E, Ploch S, Holub EB, Shin H-D, Jones JDG. (2009) A new species of Albugo parasitic to Arabidopsis thaliana reveals new evolutionary patterns in white blister rusts (Albuginaceae). Persoonia 22: 123–128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vánky K. (2012) Smut Fungi of the World. St Paul, MN: American Phytopathological Society Press. [Google Scholar]
- Vánky K. (2007) Taxonomic studies on Ustilaginomycetes – 27. Mycotaxon 99: 1–70. [Google Scholar]
- White TJ, Bruns T, Lee S, Taylor J. (1990) Amplification and direct sequencing of fungal ribosomal RNA sequences for phylogenetics. In: PCR Protocols: a guide to methods and applications (Innis N, Gelfand D, Sninsky J, White T, eds): 315–322. San Diego: Academic Press. [Google Scholar]