Abstract
Thrips tabaci Lindeman is an important polyphagous insect pest species estimated to cause losses of more than U.S. $1 billion worldwide annually. Chemical insecticides are of limited use in the management of T. tabaci due to the thigmokinetic behavior and development of resistance to insecticides. There is an urgent need to find alternative management strategies. Small noncoding RNAs (sncRNAs) especially microRNAs (miRNAs) hold great promise as key regulators of gene expression in a wide range of organisms. MiRNAs are a group of endogenously originated sncRNA known to regulate gene expression in animals, plants, and protozoans. In this study, we explored these RNAs in T. tabaci using deep sequencing to provide a basis for future studies of their biological and physiological roles in governing gene expression. Apart from snoRNAs and piRNAs, our study identified nine novel and 130 known miRNAs from T. tabaci. Functional classification of the targets for these miRNAs predicted that majority are involved in regulating transcription, translation, signal transduction and genetic information processing. The higher expression of few miRNAs (such as tta‐miR‐281, tta‐miR‐184, tta‐miR‐3533, tta‐miR‐N1, tta‐miR‐N7, and tta‐miR‐N9) in T. tabaci pupal and adult stages reflected their possible role in larval and adult development, metamorphosis, parthenogenesis, and reproduction. This is the first exploration of the miRNAome in T. tabaci, which not only provides insights into their possible role in insect metamorphosis, growth, and development but also offer an important resource for future pest management strategies.
Keywords: deep‐sequencing, miRNAs, sRNAs, Stem‐loop RT‐PCR, Thrips tabaci, tospovirus
1. INTRODUCTION
Onion thrips, Thrips tabaci Lindemann (Figure 1), is an important polyphagous insect pest species (Lewis, 1973) belonging to the family Thripidae. Besides onions, it is known to infest around 300 plant species, including economically important crops such as tobacco, leek, cabbage, pea, melon, lettuce, potato, tomato, carnation (Diaz‐Montano, Fuchs, Nault, Fail, & Shelton, 2011; Lewis, 1997; Mandal et al., 2012). Thrips tabaci is also a vector of two viral pathogens, Iris yellow spot virus (IYSV) (Srinivasan et al., 2012) and Tomato spotted wilt virus (TSWV) (Pittman, 1927) causing significant disease around the world (German, Ullman, & Moyer, 1992). Thrips tabaci is estimated to cause more than U.S. $1 billion in crop losses annually worldwide. To date, chemical insecticides have been widely used for the management of T. tabaci, but due to its thigmokinetic behavior and frequent development of insecticide resistance, they have had little use. Therefore, the design of novel insecticides, resistance breeding strategies, an in‐depth understanding of genes and gene regulation is necessary for targeting important developmental factors/processes for effective management of this insect. MiRNA analysis is an effective tool to understand gene regulation and expression in both insect and host plant.
Figure 1.
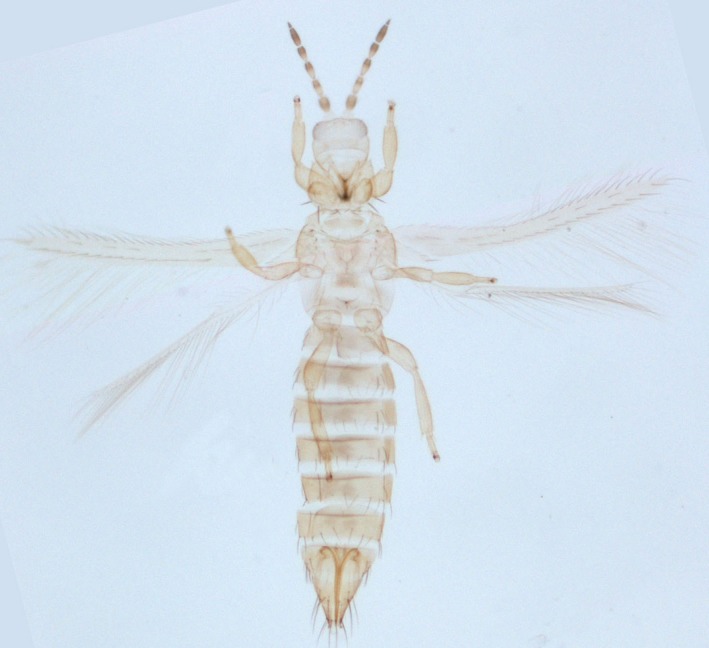

Photograph of the adult Thrips tabaci Lindeman, an important polyphagous insect pest species belonging to the family Photograph of the Thripidae. Image Credit : Dr. Ramaiyer Varatharajan (Manipur University, Imphal) and Rachana R R (ICAR‐ NBAIR, Bengaluru).
MicroRNAs (miRNAs) are a group of small, sequence‐specific, endogenously originated noncoding RNA (ncRNA) molecules containing ~18–25 nucleotides (nts), and their main function is to regulate gene expression in animals, plants, and protozoans. MiRNAs controls around 60% of protein‐coding gene activities and regulates many cellular processes (Fabian, Sonenberg, & Filipowicz, 2010; Friedman, Farh, Burge, & Bartel, 2009). The function of miRNAs appears to regulate gene expression either by translation repression or by degradation of mRNA through deadenylation (Chekulaeva & Filipowicz, 2009). MiRNA‐mediated gene regulation plays a significant role in cellular and developmental processes, for instance in cell division, cell death, disease, hormone secretion, and neural development (Ambros, 2004; Miska et al., 2007; Nohata, Hanazawa, Kinoshita, Okamoto, & Seki, 2012; Singh & Nagaraju, 2008). The first miRNA, Lin‐4 gene, was discovered by Lee, Feinbaum, and Ambros (1993) in Caenorhabditis elegans. Consequently, several miRNAs have been discovered from wide varieties of organisms including insects (Lagos‐Quintana, Rauhut, Lendeckel, & Tuschl, 2001), plants (Bartel, 2004), viruses (Cullen, 2006), and vertebrates (Lim, Glasner, Yekta, Burge, & Bartel, 2003).
Identification of miRNA includes three principle approaches, forward genetics, bioinformatics prediction (Rebijith et al., 2014; Zhang, Pan, Cannon, Cobb, & Anderson, 2006), and direct cloning and sequencing (Chen et al., 2005; Lagos‐Quintana et al., 2001; Lee & Ambros, 2001). High‐throughput next‐generation sequencing (NGS) emerged as a powerful tool to identify miRNAs from animals and plants (Calla & Geib, 2015; Guillem, Bastian, Maria‐Dolors, & Xavier, 2016; Nandety, Sharif, Kamita, Ramasamy, & Falk, 2015; Song et al., 2011; Wang et al., 2012; Wu et al., 2013). It has accelerated the pace of miRNA discovery from various animals and plants (Avesson, Reimegard, Wagner, & Söderbom, 2012; Burnside et al., 2008; Ge et al., 2013; Hu et al., 2012; Kang et al., 2012; Koh et al., 2010; Zhang et al., 2012).
So far, the miRNAome for insects is far behind nematodes, plants, and mammals (Kakumani et al., 2015). MiRNAs are reported from about 25 species of insects belonging to various orders (Stark et al., 2007; Wu et al., 2013). No information is available on T. tabaci miRNA content and function. Our study reports the detailed profile of miRNAs from T. tabaci. Further analysis identified putative target genes for these miRNAs, which will shed more light on the identification of highly specific miRNAs for thysanopteran pest management in the near future.
2. MATERIALS AND METHODS
2.1. Insect culture and RNA isolation
Thrips tabaci cultures were maintained on Phaseolus vulgaris in controlled laboratory conditions at 25°C (DeGraaf & Wood, 2009) with an 8 hr:16 hr light:dark cycle. Total RNA was isolated from whole‐body homogenates of sample mix, containing a total of 50 mg of different life stages viz. eggs, larvae, pupae, and adults of T. tabaci using TRIzol reagent (Invitrogen, Carlsbad, CA, USA).
2.2. Sample preparation and Illumina sequencing
Samples were processed according to Illumina TruSeq™ Small RNA sample preparation guide. Size fractionated small RNA populations (18–28 nts) were extracted, purified, and ligated to 3′ and 5′ adapters using T4 RNA Ligase (Life Technologies, Ambion, USA). Ligated products were reverse transcribed using SuperScript II (Life Technologies, Invitrogen, USA) followed by PCR amplification with 11 cycles and two size selection gels. High‐throughput sequencing of the small RNA libraries was performed on Illumina Hiseq2000.
2.3. Bioinformatics analysis of small RNA sequencing data
The obtained sequenced dataset was subjected to initial quality check, and the raw reads were taken for adapter trimming and filtering of low‐quality data. Thus, obtained sequencing data were queried against Rfam (http://rfam.sanger.ac.uk/) and RepBase (http://www.girinst.org/repbase/) as references to annotate the ncRNAs viz. rRNAs, tRNAs, snRNAs, snoRNAs, and repeat‐associated small RNAs and degraded fragments of expressed genes (exons and introns) in the remaining sequences. Remaining unique sequences were aligned with the miRBase (v21, http://www.miRBase.org/) entries to identify the conserved miRNAs. Novel miRNAs and their star reads were identified using the miRDeep2 (Friedlander, Mackowiak, Li, Chen, & Rajewsky, 2012) and miRCat (http://srna-workbench.cmp.uea.ac.uk/tools/mircat/). Potential secondary hairpin structures for identified novel miRNAs were predicted by employing Mfold (http://mfold.rna.albany.edu/?q_mfold/RNA-folding-form).
Homology analysis was performed with conserved miRNAs of T. tabaci with the miRNAs of other organisms from the miRBase database (Release 21.0; Griffiths‐Jones, Saini, van Dongen, & Enright, 2008). BLASTn embedded in the miRBase database was used to compare the T. tabaci miRNAs with other species, with an E‐value of .01 to find out more miRNA homologs. The naming of the miRNAs in this study has been performed according to Griffiths‐Jones, Grocock, van Dongen, Bateman, & Enright, 2006. As these miRNAs were predicted from T. tabaci, the prefix for all miRNAs was fixed as “tta.” The rest of the naming convention criteria were in accordance with miRBase (Griffiths‐Jones et al., 2006).
2.4. Phylogenetic analysis of microRNA family
All the identified miRNAs were classified into different miRNA precursor families (http://www.rfam.sanger.ac.uk), and primary sequence analyses were performed by employing Bioedit (Hall, 1999) and Weblogo (http://weblogo.berkeley.edu/logo.cgi). Few miRNA families such as miR‐8, miR‐14, miR‐276, and miR‐281 were selected for phylogenetic analysis employing RaxML.v.7.0.4 (Stamatakis, 2008).
2.5. Target prediction
Targets for identified miRNAs were predicted employing the miRanda program (Enright et al., 2004), against the expressed sequence tags (ESTs) and transcriptome (NCBI Accession: PRJNA203209) database of Frankliniella occidentalis. An alignment score (Smith & Waterman, 1981) greater than or equal to 100 and miRNA:mRNA Minimum Free Energy (MFE, ∆G) less than −20 kcal/mol were considered as putative target genes. The targets were further annotated against NCBI‐RefSeq invertebrate protein database and Gene Ontology (GO) terms were assigned (using Blast‐2‐GO) based on the annotation. The circos plot was generated using Circos (Krzywinski et al., 2009) to visualize the interaction between miRNAs and their targets.
2.6. Validation of Thrips tabaci miRNAs using Stem‐loop RT‐PCR
We were able to validate six conserved and four novel microRNAs employing Stem‐loop RT‐PCR primers designed based on previous reports (Chen et al., 2005).
2.7. Differential expression of Thrips tabaci miRNAs using Quantitative Real‐Time PCR
Differentially expressed and functionally significant ten miRNAs (six conserved and four novel) were selected for quantitative reverse transcriptase PCR (qRT‐PCR). Total RNA was isolated from different life stages viz. larvae, pupae, and adults of T. tabaci using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). Mir‐X‐miRNA qRT‐PCR SYBR Kit (Clontech Laboratories, Inc., USA) was used for the qRT‐PCR reactions. qRT‐PCR was performed on Light Cycler 480 (Roche, USA) using 1:20 diluted cDNAs and SYBR Advantage Premix (Clontech Laboratories, Mountain View, USA), according to the manufacturer's instructions. All the qRT‐PCR assays were conducted according to the MIQE guidelines (Bustin et al., 2009). qRT‐PCR assays were performed in triplicates for three independent biological replicates, and the relative gene expression data were analyzed using method (Livak & Schmittgen, 2001). U6 snRNAs was used as an internal control gene for normalization. The values of these three independent experiments were statistically analyzed using one‐way ANOVA to calculate the statistical significance.
3. RESULTS
3.1. Illumina sequencing of Thrips tabaci small RNAs
The small RNA library prepared for deep sequencing resulted in a total of 13,192,454 raw reads (Table 1). After various mapping (Table 1), the trimmed high‐quality small RNA reads were employed to identify both known and novel miRNAs. Size distributions of the trimmed high‐quality reads were varied from 18 to 26 nts with a peak at the 23 nts (Figure 2). A small portion of our library consisted of read length of around 26–28 nts, which could be putative piwi‐interacting RNAs (piRNAs) from T. tabaci as the homology search against the piRNABank database revealed that some of these were similar to previously reported piRNAs (Table 2).
Table 1.
Summary Statistics of Thrips tabaci small RNA data analysis
| Number of trimmed reads | 13,192,454 |
| Mapped to mRNA | 2,378,671 |
| Repbase mapped reads | 1,396,829 |
| Rfam mapped reads | 4,181,894 |
| Rfam unmapped reads | 9,010,560 |
| miRBase mapped reads | 47,570 |
| Total unmappable for miRNA | 5,187,490 |
| Average length | 23 |
Figure 2.
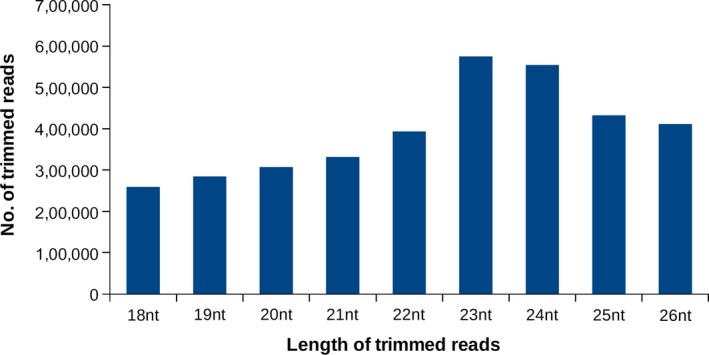
Length distribution of mappable reads (≥18 nt to ≤26 nt) obtained from Thrips tabaci deep sequencing
Table 2.
Small RNAs (Piwi RNAs) with nucleotide lengths larger than 25 nucleotides obtained from Thrips tabaci sequencing data
| smallRNA ID | Sequence (5′‐>3′ | Length (nt) | Hit in the piRNABank | E‐value |
|---|---|---|---|---|
| tta_piR1 | ATTGTGGTTCAGTGGTAGAATTCTCGCC | 28 | hsa_piR_018570 | .00065 |
| tta_piR2 | GGGTTCGATTCCCGGTCAGGGAACCA | 26 | dr_piR_0017650 | .1 |
| tta_piR3 | TTTCCGTAGTGTAGTGGTTATCACGTTC | 28 | rno_piR_005901 | 1.70E‐05 |
| tta_piR4 | CCAAAGCAUCGCGAAGGCCCACGGCG | 26 | dr_piR_0052831 | .0047 |
| tta_piR5 | ATTGGTGGTTCAGTGGTAGAATTCTCGC | 28 | hsa_piR_001312 | 1.80E‐05 |
| tta_piR6 | CCCTCGGTTCTGGCGTCAAGCGGGCCG | 27 | No Hit | NA |
| tta_piR7 | CCTGTGGTCTAGTGGTTAGGATTCGGCG | 28 | ona_piR_166322 | .00049 |
3.2. Identification of known miRNAs from Thrips tabaci
Our analyses on the trimmed high‐quality reads resulted in a total of 130 conserved miRNAs representing 55 different miRNA families (Table 3). Among the known miRNAs, miR‐276, miR‐281, miR‐8, and miR‐14 are highly expressed with an expression value of 26,418, 18,063, 16,204, and 12,453, respectively (Table 3). Analysis of the 55 miRNA families revealed that most of them were present in arthropod species (Table 4), with many homologous miRNAs from Aedes aegypti, Apis mellifera, Bombyx mori, Acyrthosiphon pisum, and Tribolium castaneum (Figure S1).
Table 3.
Expression value of known miRNAs in Thrips tabaci. The first column represents miRNA family; the second column represents the number of reads annotated on the particular miRNA family; the third column represents length of the mature miRNA sequences; the fourth column represents the name of the miRNAs in T. tabaci; the fifth column represents the miRNA sequence; the sixth column represents homologous species of organism where it has the highest similarity
| miRNA family | Expression valuesa (Reads) | Length (nt) | Name of the miRNA | Sequence (5′–3′) | Resource |
|---|---|---|---|---|---|
| mir‐281 | 468 | 19 | tta‐miR‐281a | AAGAGAGCUAUCCGUCGAC | Aedes aegypti |
| mir‐281 | 17583 | 22 | tta‐miR‐281b | AAGAGAGCUAUCCGUCGACAGU | Bombyx mori |
| mir‐281 | 4 | 22 | tta‐miR‐281c | AAGAGAGCUGUCCGUCGACAGU | Drosophila ananassae |
| mir‐281 | 5 | 21 | tta‐miR‐281d | AAGGGAGCAUCUGUCGACAGU | Lottia gigantea |
| mir‐281 | 3 | 22 | tta‐miR‐281e | UGUCAUGGAGUUGCUCUCUUUU | Branchiostoma belcheri |
| mir‐276 | 507 | 21 | tta‐miR‐276a | UAGGAACUUCAUACCGUGCUC | Aedes aegypti |
| mir‐276 | 25904 | 22 | tta‐miR‐276b | UAGGAACUUCAUACCGUGCUCU | Locusta migratoria |
| mir‐276 | 7 | 22 | tta‐miR‐276c | UAGGAACUUAAUACCGUGCUCU | Drosophila ananassae |
| mir‐306 | 173 | 21 | tta‐miR‐306a | UCAGGUACUAGGUGACUCUGA | Bombyx mori |
| mir‐306 | 3507 | 22 | tta‐miR‐306b | UCAGGUACUGAGUGACUCUGAG | Apis mellifera |
| mir‐306 | 1976 | 22 | tta‐miR‐306c | UCAGGUACUGAGUGACUCUCAG | Aedes aegypti |
| bantam | 61 | 21 | tta‐miR‐bantam‐a | UGAGAUCAUUGUGAAAGCUAU | Brugia malayi |
| bantam | 33 | 22 | tta‐miR‐bantam‐b | UGAGAUCAUUUUGAAAGCUGAU | Aedes aegypti |
| bantam | 1889 | 23 | tta‐miR‐bantam‐c | UGAGAUCAUUGUGAAAGCUGAUU | Apis mellifera |
| bantam | 5 | 23 | tta‐miR‐bantam‐d | UGAGAUCAUUGUGAAAGCUAAUU | Acyrthosiphon pisum |
| mir‐92 | 5 | 20 | tta‐miR‐92a | UAUUGCACUCGUCCCGGCCU | Brugia malayi |
| mir‐92 | 8 | 22 | tta‐miR‐92b | UAUUGCACCAGUCCCGGCCUAU | Bombyx mori |
| mir‐92 | 6 | 22 | tta‐miR‐92c | UAUUGCACCUGUCCCGGCCGAU | Ciona savignyi |
| mir‐92 | 76 | 22 | tta‐miR‐92d | UAUUGCACUCGUCCCGGCCUUG | Oikopleura dioica |
| mir‐92 | 4 | 23 | tta‐miR‐92e | UAUUGCACCAGUCCCGGCCUGAC | Tribolium castaneum |
| mir‐92 | 1861 | 22 | tta‐miR‐92f | UAUUGCACUCGUCCCGGCCUGU | Saccoglossus kowalevskii |
| mir‐92 | 629 | 22 | tta‐miR‐92 g | UAUUGCACUCGUCCCGGCCUGC | Lytechinus variegatus |
| mir‐92 | 46 | 22 | tta‐miR‐92 h | AAUUGCACCCGUCCCGGCCUGA | Apis mellifera |
| mir‐750 | 4 | 22 | tta‐miR‐750a | CAGAUCUAACUCUUCCAGCUCA | Lottia gigantea |
| mir‐750 | 1242 | 22 | tta‐miR‐750b | CCAGAUCUAACUCUUCCAGCUC | Apis mellifera |
| mir‐750 | 107 | 23 | tta‐miR‐750c | CCAGAUCUAACUCUUCCAGCUCA | Capitella teleta |
| mir‐10 | 433 | 21 | tta‐miR‐10a | ACCCUGUAGAUCCGAAUUUGU | Acyrthosiphon pisum |
| mir‐10 | 6 | 21 | tta‐miR‐10b | UACCCUGUAGAUCCGAAUUUG | Ovis aries |
| mir‐10 | 4 | 22 | tta‐miR‐10c | UACCCUGUAGAACCGAAUUUGU | Anolis carolinensis |
| mir‐10 | 6 | 22 | tta‐miR‐10d | ACCCUGUAGAUCCGAAUUUGUU | Aedes aegypti |
| mir‐10 | 9 | 22 | tta‐miR‐10e | AACCCUGUAGACCCGAAUUUGA | Gyrodactylus salaris |
| mir‐10 | 52 | 22 | tta‐miR‐10f | UACCCUGUAGAUCCGAAUUUGU | Lottia gigantea |
| mir‐10 | 3 | 23 | tta‐miR‐10 g | UACCCUGUAGAACCGAAUUUGUG | Bos taurus |
| mir‐10 | 16 | 23 | tta‐miR‐10 h | UACCCUGUAGAAUCGAAUUUGUG | Anolis carolinensis |
| mir‐10 | 3 | 23 | tta‐miR‐10i | AACCCUGUAGAUCCGAGUUAGAU | Schmidtea mediterranea |
| mir‐100 | 5 | 21 | tta‐miR‐100a | AACCCGUAGAUCCGAACUUGU | Capra hircus |
| mir‐100 | 20 | 22 | tta‐miR‐100b | AACCCGUAGAUCCGAACUUGUG | Ateles geoffroyi |
| mir‐100 | 57 | 23 | tta‐miR‐100c | AACCCGUAGAUCCGAACUUGUGU | Branchiostoma floridae |
| mir‐100 | 3 | 24 | tta‐miR‐100d | AACCCGUAGAUCCGAACUUGUGUU | Ascaris suum |
| mir‐1000 | 11 | 18 | tta‐miR‐1000a | AUAUUGUCCUGUCACAGC | Tribolium castaneum |
| mir‐1000 | 192 | 21 | tta‐miR‐1000b | AUAUUGUCCUGUCACAGCAGU | Drosophila melanogaster |
| mir‐1000 | 183 | 22 | tta‐miR‐1000c | AUAUUGUCCUGUCACAGCAGUA | Drosophila pseudoobscura |
| mir‐8 | 248 | 22 | tta‐miR‐8a | UAAUACUGUCAGGUAAAGAUGU | Culex quinquefasciatus |
| mir‐8 | 15951 | 23 | tta‐miR‐8b | UAAUACUGUCAGGUAAAGAUGUC | Capitella teleta |
| mir‐8 | 5 | 22 | tta‐miR‐8c | CAUCUUACCGGGCAGCAUUAGA | Aedes aegypti |
| mir‐9 | 8 | 18 | tta‐miR‐9a | UCUUUGGUAUCCUAGCUG | Bombyx mori |
| mir‐9 | 7 | 21 | tta‐miR‐9b | UCUUUGGUGAUCUAGUUGUAU | Tribolium castaneum |
| mir‐9 | 6 | 21 | tta‐miR‐9c | UCUUUGGUACUUUAGCUGUAG | Acyrthosiphon pisum |
| mir‐9 | 13 | 23 | tta‐miR‐9d | UCUUUGGUUAUCUAGCUGUAUGA | Capitella teleta |
| mir‐9 | 4 | 24 | tta‐miR‐9e | UCUUUGGUUUUCUAGCUGUAUGAU | Schmidtea mediterranea |
| mir‐2 | 21 | 23 | tta‐miR‐2a | UAUCACAGCCAGCUUUGAUGAGC | Apis mellifera |
| mir‐2 | 27 | 23 | tta‐miR‐2b | UAUCACAGCCAGCUUUGAUGAGU | Lottia gigantea |
| mir‐2 | 27 | 24 | tta‐miR‐2c | UAUCACAGCCAGCUUUGAUGAGCU | Aedes aegypti |
| mir‐184 | 7145 | 21 | tta‐miR‐184a | UGGACGGAGAACUGAUAAGGG | Anopheles gambiae |
| mir‐184 | 142 | 22 | tta‐miR‐184b | UGGACGGAGAACUGAUAAGGGU | Anolis carolinensis |
| mir‐184 | 118 | 22 | tta‐miR‐184c | UGGACGGAGAACUGAUAAGGGC | Ixodes scapularis |
| mir‐279 | 20 | 21 | tta‐miR‐279a | UGACUAGAUCCACACUCAUCC | Acyrthosiphon pisum |
| mir‐279 | 58 | 22 | tta‐miR‐279b | UGACUAGAUCCACACUCAUCCA | Lottia gigantea |
| mir‐279 | 14 | 22 | tta‐miR‐279c | UGACUAGAUCCACACUCAUUAA | Anopheles gambiae |
| mir‐279 | 4 | 22 | tta‐miR‐279d | UGACUAGAUCUACACUCAUUGA | Bombyx mori |
| mir‐279 | 105 | 22 | tta‐miR‐279e | UGACUAGAGUCACACUCGUCCA | Apis mellifera |
| mir‐279 | 635 | 22 | tta‐miR‐279f | UGACUAGAUCCAUACUCGUCUG | Bombyx mori |
| mir‐279 | 26 | 24 | tta‐miR‐279 g | UGACUAGAUCGAAAUACUCGUCCC | Apis mellifera |
| mir‐279 | 103 | 25 | tta‐miR‐279 h | UGACUAGAUCCAUACUCGUCUAUAG | Tribolium castaneum |
| mir‐2796 | 65 | 21 | tta‐miR‐2796a | AGGCCGGCGGAAACUACUUGC | Nasonia vitripennis |
| mir‐2796 | 5 | 22 | tta‐miR‐2796b | GUAGGCCGGCGGAAACUACUAG | Acyrthosiphon pisum |
| mir‐2796 | 168 | 23 | tta‐miR‐2796c | GUAGGCCGGCGGAAACUACUUGC | Apis mellifera |
| mir‐14 | 26 | 21 | tta‐miR‐14a | UCAGUCUUUUUCUCUCUCCUA | Anopheles gambiae |
| mir‐14 | 12427 | 22 | tta‐miR‐14b | UCAGUCUUUUUCUCUCUCCUAU | Acyrthosiphon pisum |
| mir‐993 | 3 | 20 | tta‐miR‐993a | UACCCUGUAGAUCCGGGCUU | Tribolium castaneum |
| mir‐993 | 110 | 23 | tta‐miR‐993b | GAAGCUCGUCUCUACAGGUAUCU | Acyrthosiphon pisum |
| mir‐993 | 10 | 23 | tta‐miR‐993c | UACCCUGUAGAUCCGGGCUUUUG | Manduca sexta |
| mir‐993 | 3 | 23 | tta‐miR‐993d | UACCCUGUAGUUCCGGGCUUUUG | Drosophila melanogaster |
| mir‐1175 | 106 | 23 | tta‐miR‐1175a | AAGUGGAGCAGUGGUCUCUUCAC | Tribolium castaneum |
| mir‐1175 | 17 | 22 | tta‐miR‐1175b | AAGUGGAGUAGUGGUCUCAUCG | Aedes aegypti |
| mir‐1175 | 4 | 23 | tta‐miR‐1175c | UGAGAUUCACUCCUCCAACUUAC | Apis mellifera |
| mir‐1175 | 56 | 24 | tta‐miR‐1175d | UGAGAUUCAACUCCUCCAACUUAA | Bombyx mori |
| mir‐124 | 106 | 21 | tta‐miR‐124a | UAAGGCACGCGGUGAAUGCCA | Schmidtea mediterranea |
| mir‐124 | 84 | 21 | tta‐miR‐124b | UAAGGCACGCGGUGAAUGCUA | Anolis carolinensis |
| mir‐263 | 4 | 21 | tta‐miR‐263a | AAUGGCACUGGAAGAAUUCAC | Bombyx mori |
| mir‐263 | 18 | 23 | tta‐miR‐263b | AAUGGCACUGGAAGAAUUCACGG | Aedes aegypti |
| mir‐263 | 20 | 24 | tta‐miR‐263c | AAUGGCACUGGAAGAAUUCACGGG | Drosophila melanogaster |
| mir‐2944 | 18 | 22 | tta‐miR‐2944a | UAUCACAGCAGUAGUUACCUGA | Aedes aegypti |
| mir‐2944 | 13 | 23 | tta‐miR‐2944b | UAUCACAGCAGUAGUUACCUGGU | Apis mellifera |
| mir‐13 | 399 | 22 | tta‐miR‐13a | UAUCACAGCCACUUUGAUGAGC | Tribolium castaneum |
| mir‐13 | 17 | 23 | tta‐miR‐13b | UAUCACAGCCAUUUUUGACGAGU | Bombyx mori |
| mir‐34 | 15 | 22 | tta‐miR‐34a | UGGCAGUGUGGUUAGCUGGUUG | Aedes aegypti |
| mir‐34 | 5 | 23 | tta‐miR‐34b | UGGCAGUGUGGUUAGCUGGUUGU | Ascaris suum |
| mir‐34 | 3 | 23 | tta‐miR‐34c | UGGCAGUGUGGUUAGCUGGUAGU | Lottia gigantea |
| mir‐133 | 15 | 22 | tta‐miR‐133a | UUGGUCCCCGUCAACCAGCUGU | Schmidtea mediterranea |
| mir‐133 | 14 | 22 | tta‐miR‐133b | UUGGUCCCCUUCAACCAGCUGU | Drosophila persimilis |
| mir‐317 | 5 | 21 | tta‐miR‐317a | UGAACACAGCUGGUGGUAUCU | Acyrthosiphon pisum |
| mir‐317 | 13 | 24 | tta‐miR‐317b | UGAACACAGCUGGUGGUAUCUUCU | Lottia gigantea |
| mir‐317 | 13 | 25 | tta‐miR‐317c | UGAACACAGCUGGUGGUAUCUCAGU | Apis mellifera |
| mir‐317 | 4 | 25 | tta‐miR‐317d | UGAACACAGCUGGUGGUAUCUCUUU | Capitella teleta |
| mir‐12 | 13 | 21 | tta‐miR‐12a | UGAGUAUUACAUCAGGUACUG | Tribolium castaneum |
| mir‐12 | 3 | 23 | tta‐miR‐12b | UGAGUAUUACAUCAGGUACUGGU | Daphnia pulex |
| mir‐252 | 4 | 22 | tta‐miR‐252a | CUAAGUACUAGUGCCGCAGGAG | Drosophila melanogaster |
| mir‐252 | 5 | 23 | tta‐miR‐252b | CUAAGUACUAGUGCCGCAGGAGU | Saccoglossus kowalevskii |
| mir‐277 | 11 | 22 | tta‐miR‐277a | UAAAUGCACUAUCUGGUACGAC | Aedes aegypti |
| mir‐277 | 5 | 23 | tta‐miR‐277b | UAAAUGCACUAUCUGGUACGACA | Acyrthosiphon pisum |
| mir‐31 | 3 | 21 | tta‐miR‐31a | AGGCAAGAUGUCGGCAUAGCU | Tribolium castaneum |
| mir‐31 | 7 | 22 | tta‐miR‐31b | GGCAAGAUGUCGGCAUAGCUGA | Apis mellifera |
| mir‐3477 | 69 | 23 | tta‐miR‐3477a | UAAUCUCAUGCGGUAACUGUGAG | Apis mellifera |
| mir‐3477 | 121 | 22 | tta‐miR‐3477b | UAAUCUCAUGUGGUAACUGUGA | Apis mellifera |
| mir‐2779 | 5 | 20 | tta‐miR‐2779 | AUAUCCGGCUCGAAGGACCA | Bombyx mori |
| mir‐929 | 4 | 22 | tta‐miR‐929 | AAAUUGACUCUAGUAGGGAGUC | Drosophila melanogaster |
| mir‐71 | 172 | 22 | tta‐miR‐71 | UCUCACUACCUUGUCUUUCAUG | Tribolium castaneum |
| mir‐375 | 4 | 22 | tta‐miR‐375 | UUUGUUCGUUCGGCUCGAGUUA | Apis mellifera |
| mir‐190 | 3 | 24 | tta‐miR‐190 | AGAUAUGUUUGAUAUUCUUGGUUG | Acyrthosiphon pisum |
| mir‐7550 | 3 | 18 | tta‐miR‐7550 | AUCCGGCUCGAAGGACCA | Ictalurus punctatus |
| mir‐482 | 3 | 22 | tta‐miR‐482 | GGAAUGGGCUGAUUGGGAAGCA | Phaseolus vulgaris |
| mir‐2478 | 3 | 20 | tta‐miR‐2478 | GUAUCCCACUUCUGACACCA | Bos taurus |
| mir‐316 | 3 | 21 | tta‐miR‐316 | UGUCUUUUUCCGCUUUGCUGC | Heliconius melpomene |
| mir‐3049 | 98 | 23 | tta‐miR‐3049 | UCGGGAAGGUAGUUGCGGCGGAU | Apis mellifera |
| mir‐996 | 57 | 21 | tta‐miR‐996 | UGACUAGAUACAUACUCGUCU | Apis mellifera |
| mir‐275 | 40 | 23 | tta‐miR‐275 | UCAGGUACCUGAAGUAGCGCGCG | Anopheles gambiae |
| mir‐965 | 31 | 22 | tta‐miR‐965 | UAAGCGUAUAGCUUUUCCCCUU | Tribolium castaneum |
| mir‐67 | 25 | 24 | tta‐miR‐67 | UCACAACCUCCUUGAGUGAGUUGA | Ascaris suum |
| mir‐315 | 21 | 23 | tta‐miR‐315 | UUUUGAUUGUUGCUCAGAAAGCC | Acyrthosiphon pisum |
| mir‐305 | 14 | 23 | tta‐miR‐305 | UUUGUACUUCAUCAGGUGCUCUG | Tetranychus urticae |
| mir‐894 | 11 | 20 | tta‐miR‐894 | CGUUUCACGUCGGGUUCACC | Physcomitrella patens |
| mir‐3533 | 9 | 20 | tta‐miR‐3533 | AUGAAGUGUGACGUGGACAU | Bos taurus |
| mir‐307 | 9 | 20 | tta‐miR‐307 | UCACAACCUCCUUGAGUGAG | Daphnia pulex |
| mir‐2765 | 664 | 22 | tta‐miR‐2765 | UGGUAACUCCACCACCGUUGGC | Bombyx mori |
| mir‐210 | 22 | 21 | tta‐miR‐210 | CUUGUGCGUGUGACAGCGGCU | Drosophila melanogaster |
| mir‐1 | 650 | 22 | tta‐miR‐1 | UGGAAUGUAAAGAAGUAUGGAG | Drosophila melanogaster |
| mir‐87 | 18 | 21 | tta‐miR‐87 | GUGAGCAAAGUUUCAGGUGUG | Ixodes scapularis |
| let‐7 | 279 | 21 | tta‐let‐7 | TGAGGTAGTAGGTTGTATAGT | Drosophila melanogaster |
| mir‐3791 | 15 | 21 | tta‐miR‐3791 | UCACCGGGUAGGAUUCAUCCA | Apis mellifera |
| Plant‐specific miRNA | |||||
| mir‐9774 | 6 | 22 | – | CAAGATATTGGGTATTTCTGTC | Triticum aestivum |
Expression value is equivalent to number of miRNA reads from the library.
Table 4.
Homology analysis of Thrips tabaci miRNA homologs
| tta‐miR | Insects | Other Arthropods | Other Invertbrates | Vertebrates | Note |
|---|---|---|---|---|---|
| tta‐bantam | √ | √ | √ | — | Invertebrate specific |
| tta‐let‐7 | √ | √ | √ | √ | Highly conserved |
| tta‐miR‐1 | √ | — | — | — | Insect specific |
| tta‐miR‐10 | √ | √ | √ | √ | Highly conserved |
| tta‐miR‐100 | √ | √ | √ | √ | Highly conserved |
| tta‐miR‐1000 | √ | — | — | — | Insect specific |
| tta‐miR‐1175 | √ | — | √ | — | Invertebrate specific |
| tta‐miR‐12 | √ | — | — | — | Insect specific |
| tta‐miR‐124 | √ | √ | √ | √ | Highly conserved |
| tta‐miR‐13 | √ | — | — | — | Insect specific |
| tta‐miR‐133 | √ | √ | √ | √ | Highly conserved |
| tta‐miR‐14 | √ | — | — | — | Insect specific |
| tta‐miR‐184 | √ | √ | √ | √ | Highly conserved |
| tta‐miR‐190 | √ | — | √ | √ | Highly conserved |
| tta‐miR‐2 | √ | √ | √ | — | Invertebrate specific |
| tta‐miR‐210 | √ | — | √ | √ | Highly conserved |
| tta‐miR‐2478 | — | — | — | √ | Vertebrate specific |
| tta‐miR‐252 | √ | — | √ | — | Invertebrate specific |
| tta‐miR‐263 | √ | √ | √ | — | Invertebrate specific |
| tta‐miR‐275 | √ | √ | — | — | Arthropod specific |
| tta‐miR‐276 | √ | √ | — | — | Arthropod specific |
| tta‐miR‐2765 | √ | — | — | — | Insect specific |
| tta‐miR‐277 | √ | — | — | — | Insect specific |
| tta‐miR‐2779 | √ | — | — | — | Insect specific |
| tta‐miR‐279 | √ | √ | √ | — | Invertebrate specific |
| tta‐miR‐2796 | √ | — | — | — | Insect specific |
| tta‐miR‐281 | √ | √ | √ | √ | Highly conserved |
| tta‐miR‐2944 | √ | — | — | — | Insect specific |
| tta‐miR‐3049 | √ | — | — | — | Insect specific |
| tta‐miR‐305 | √ | √ | — | — | Arthropod specific |
| tta‐miR‐306 | √ | √ | — | — | Arthropod specific |
| tta‐miR‐307 | √ | √ | √ | — | Invertebrate specific |
| tta‐miR‐31 | √ | — | — | — | Insect specific |
| tta‐miR‐315 | √ | √ | √ | √ | Highly conserved |
| tta‐miR‐316 | √ | √ | — | — | Arthropod specific |
| tta‐miR‐317 | √ | √ | √ | — | Invertebrate specific |
| tta‐miR‐34 | √ | √ | √ | — | Invertebrate specific |
| tta‐miR‐3477 | √ | — | — | — | Insect specific |
| tta‐miR‐3533 | — | — | — | √ | Vertebrate specific |
| tta‐miR‐375 | √ | √ | √ | — | Invertebrate specific |
| tta‐miR‐3791 | √ | — | — | — | Insect specific |
| tta‐miR‐482 | — | — | √ | — | Invertebrate specific |
| tta‐miR‐67 | — | — | √ | — | Invertebrate specific |
| tta‐miR‐71 | — | — | √ | — | Invertebrate specific |
| tta‐miR‐750 | √ | — | — | — | Insect specific |
| tta‐miR‐7550 | — | — | — | √ | Vertebrate specific |
| tta‐miR‐8 | √ | √ | √ | — | Invertebrate specific |
| tta‐miR‐87 | √ | — | — | — | Insect specific |
| tta‐miR‐894 | — | — | — | √ | Vertebrate specific |
| tta‐miR‐9 | √ | √ | √ | √ | Highly conserved |
| tta‐miR‐92 | √ | √ | √ | √ | Highly conserved |
| tta‐miR‐929 | √ | √ | √ | √ | Highly conserved |
| tta‐miR‐965 | √ | √ | — | — | Arthropod specific |
| tta‐miR‐993 | √ | √ | √ | — | Invertebrate specific |
| tta‐miR‐996 | √ | — | — | — | Insect specific |
3.3. Identification of novel miRNAs from Thrips tabaci
Miranalyzer pipeline identified a total of nine novel miRNAs from T. tabaci for the first time (Table 5), with their predicted precursor secondary structures (Figure 3). The complete details of the mature miRNAs and their corresponding pre‐miRNAs have been given in Table 5. The length of the novel miRNAs ranged from 21 to 22 nucleotides with a preference of Uracil (66.7%) followed by Adenine (22.2%) at the 5′ end. The length of the pre‐miRNAs was in the range of 63–76 nucleotides with an average Minimum Free Energy (MFE) of −35.97 kcal/mol, indicating pre‐miRNAs are readily folded into their secondary structures. Among these nine miRNAs, three were located in the 5′ arm while the other six arose from 3′ arm (Table 5, Figure 3). tta‐miR‐N4 (3414 copies) and tta‐miR‐N7 (1978 copies) were having the highest abundance compared to the remaining novel miRNAs (Table 5).
Table 5.
Details of Thrips tabaci novel miRNAs and its star strands obtained from this study. Information regarding mature, star and precursor sequences, start and end position, orientation, expression values, MFE value and (A+U) content, etc. have been given
| MicroRNA Family | Name of the Novel miRNA | miRNA sequence | miRNA* sequence | Hairpin sequence | Start | End | Orientation | miRNA Reads | miRNA* Reads | MFE |
|---|---|---|---|---|---|---|---|---|---|---|
| Novel miRNA‐1 | tta‐miR‐N1 | AGGUAACUAACUUGCAGGCCA | NO | GUGGGAUGGCCAGUAAGUUAGAGCCCUCUUGUUUAGAUGAAGUUGGAGGUAACUAACUUGCAGGCCAAGCCAC | 3630 | 3650 | ‐ [Negative strand] | 28 | Nil | −32.9 |
| Novel miRNA‐2 | tta‐miR‐N2 | UUCGUUGUGCGGAAAAAUGGAU | NO | CGCCAGGACCACAUUUUUCUGCACGGAUGGACUGAGAUUGAUACGGUUCGUUGUGCGGAAAAAUGGAUUCUUGGCG | 1439 | 1460 | + [Positive strand] | 6 | Nil | −40 |
| Novel miRNA‐3 | tta‐miR‐N3 | AUCAGCGAGUUCUGGCACUAC | NO | GGGAGGGAUCAGCGAGUUCUGGCACUACGUGCAGAUUUGAGUGCGUGUGUCAGAACUAAUUGAACCCGCCC | 11956 | 11976 | + [Positive strand] | 15 | Nil | −39.6 |
| Novel miRNA‐4 | tta‐miR‐N4 | UGACUAGACUCUCACUCGUCU | NO | UCCCUCGGCGAGUGAGUUUCUGGCUCAUGUUGUCAGUUCAUGACUAGACUCUCACUCGUCUAGGGA | 8647 | 8667 | + [Positive strand] | 3414 | Nil | −37.2 |
| Novel miRNA‐5 | tta‐miR‐N5 | UGGUAACUAACUUGCGGGCCA | NO | GGUGGCUCGUAAGUUAAGUUCCCGCUGUGAUUUAAACUAGUGGUAACUAACUUGCGGGCCACU | 13477 | 13497 | ‐ [Negative strand] | 305 | Nil | −35.1 |
| Novel miRNA‐6 | tta‐miR‐N6 | UUUGUUCGCUCGGCUCGAUGUA | CCUCGAGCCUGGCGGACAGGU(5) CCUCGAGCCUGGCGGACAGGUU(9) | CCUAAUGCCUCGAGCCUGGCGGACAGGUUGUCCUGUUCGAGUAAUUUGUUCGCUCGGCUCGAUGUAUGAGG | 1925 | 1946 | + [Positive strand] | 528 | 14 | −37.9 |
| Novel miRNA‐7 | tta‐miR‐N7 | UCAGGUACCAGAAGUAGCGCG | GUGCUGCAUCCGGUGCUAGUG(1) | CUUGCCGGUGCUGCAUCCGGUGCUAGUGGCUGUGAUUUUAAACCAGUCAGGUACCAGAAGUAGCGCGCGGGGAG | 12973 | 12993 | ‐ [Negative strand] | 1978 | 1 | −36 |
| Novel miRNA‐8 | tta‐miR‐N8 | UCUUUGGUGAUUUGGCGGUAUG | AUAAAGCUAGAUUACCAAAGC(6) AUAAAGCUAGAUUACCAAAGCA(6) | UGUUGCUUCUUUGGUGAUUUGGCGGUAUGUAAUAAUUGAAAGGCCAUAAAGCUAGAUUACCAAAGCAGGGACA | 14289 | 14310 | + [Positive strand] | 22 | 12 | −29.3 |
| Novel miRNA‐9 | tta‐miR‐N9 | CGCGUCGGUGUGCGCAGAAGG | CCUGCCUGGAGCCGCCGACGG(3) | GUAGGGUCGCGUCGGUGUGCGCAGAAGGGUCUAUGUGUGGGCCUGCCUGGAGCCGCCGACGGCUGUGC | 274 | 294 | + [Positive strand] | 12 | 3 | −34.8 |
Figure 3.
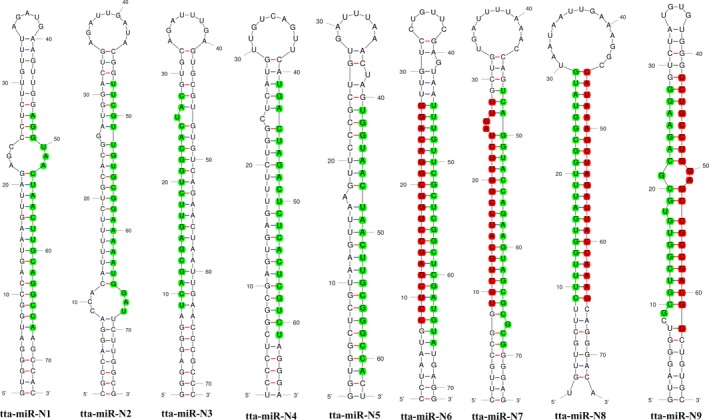
Stem‐loop structures of nine novel Thrips tabaci miRNAs indicating mature miRNA sequence (green color) and miRNA star strand sequences (red color)
3.4. The presence of miRNA star strands
It is very difficult to identify the star strand (miRNA*) sequences from the library, as it will be degraded soon after being exported to the cytosol. However, our results revealed that ten T. tabaci miRNA* families (mir‐14, mir‐184, mir‐8, mir‐276, mir‐210, mir‐1, mir‐3477, mir‐71, mir‐13, and let‐7) were identified within the known miRNA category (Table 6). The expression values (number of reads) of all miRNA*s were lower than that of their corresponding miRNAs (Table 6). Among the miRNA* family, mir‐8 and mir‐276 families were having the highest abundance with 308 and 258 copies, respectively. Our results also indicated the presence of miRNA* sequences in four of our novel miRNAs such as tta‐miR‐N6, tta‐miR‐N7, tta‐miR‐N8, and tta‐miR‐N9, although the abundance was low (Table 5). The complete characteristic features of these miRNA* sequences and their corresponding pre‐miRNA*s have been given in Tables 5 and 6.
Table 6.
Details of Thrips tabaci miRNA*s obtained from this study. Information regarding mature, star and precursor sequences, start and end position, orientation, expression values, MFE value and (A+U) content, etc. have been given
| MicroRNA Family | Name of the miRNA* | miRNA | miRNA* | Hairpin sequence | Start | End | miRNA* Abundance | Minimum Free Energy(MFE) | Hairpin G/C% |
|---|---|---|---|---|---|---|---|---|---|
| mir‐14 | tta‐tta‐miR*‐14 | UCAGUCUUUUUCUCUCUCCUAU | GGGGAGAGAUAAGGGCUUUGGCU(5) | GGGGAGAGAUAAGGGCUUUGGCUCGAUUUUAAAGUCAGUCAGUCUUUUUCUCUCUCC | 345 | 366 | 5 | −26.7 | 45.614033 |
| mir‐184 | tta‐miR*‐184 | UGGACGGAGAACUGAUAAGGG | CCUUGUCAUUCUCGUGUCCGGU(21) | CGCCUCCUUGUCAUUCUCGUGUCCGGUUGUGCAUUCAACUUACUGGACGGAGAACUGAUAAGGGCGCG | 592 | 612 | 21 | −33.5 | 54.411762 |
| mir‐8 | tta‐miR*‐8 | UAAUACUGUCAGGUAAAGAUGUC | CAUCUUACCGGGCAGCAUUAGAC(308) | UCUGUUCACAUCUUACCGGGCAGCAUUAGACUUGGAUUGAUAGCCUCUAAUACUGUCAGGUAAAGAUGUCGUCAGA | 3,662 | 3,684 | 308 | −27.7 | 43.421055 |
| mir‐276 | tta‐miR*‐276 | UAGGAACUUCAUACCGUGCUCU | UAGCGAGGUAUAGAGUUCCUACG(196) AGCGAGGUAUAGAGUUCCUACG(62) | UCCAGUAGCGAGGUAUAGAGUUCCUACGUGGUGUUGGGUACAGUAGGAACUUCAUACCGUGCUCUUGGA | 4,052 | 4,073 | 258 | −33.9 | 49.275364 |
| mir‐210 | tta‐miR*‐210 | CUUGUGCGUGUGACAGCGGCU | AGCUGCUGGACACUGCACAAG(1) AGCUGCUGGACACUGCACAAGA(7) | AGCUGCUGGACACUGCACAAGAUUAGACUUUGGAAAACUCUUGUGCGUGUGACAGCGGCU | 10,847 | 1,0867 | 8 | −28.09 | 50 |
| mir‐1 | tta‐miR*‐1 | UGGAAUGUAAAGAAGUAUGGAG | CCAUACUUCCUUGCUUCCCAU(7) CCAUACUUCCUUGCUUCCCAUA(4) | GUUCCAUACUUCCUUGCUUCCCAUAUUGCCAUUUGAAACUUAUGGAAUGUAAAGAAGUAUGGAGC | 581 | 602 | 11 | −24.86 | 38.46154 |
| mir‐3477 | tta‐miR*‐3477 | UAAUCUCAUGUGGUAACUGUGA | UCAGGGUUCCGCGUGAGGUUG(1) | GUAAUCUCAUGUGGUAACUGUGAGUUGUACUUGUACCUCAGGGUUCCGCGUGAGGUUGC | 5,735 | 5,756 | 1 | −31.3 | 49.152542 |
| mir‐71 | tta‐miR*‐71 | UCUCACUACCUUGUCUUUCAUG | UGAAAGACAUGGGUAGUGAGAU(19) UGAAAGACAUGGGUAGUGAGAUG(20) | GGGUGACGUGAAAGACAUGGGUAGUGAGAUGUUUGCUGCUGUACAUCUCACUACCUUGUCUUUCAUGUUGCUC | 1,176 | 1,197 | 39 | −47.1 | 46.575344 |
| mir‐13 | tta‐miR*‐13 | UAUCACAGCCACUUUGAUGAGC | GCCAUCAAUACGGCUGUGAGAGC(64) CCAUCAAUACGGCUGUGAGAGC(17) | GAGGCUGGAGCCAUCAAUACGGCUGUGAGAGCGUGAAUUUGAUACCGUAUCACAGCCACUUUGAUGAGCUCUGGCUUC | 1,427 | 1,449 | 81 | −41 | 51.282055 |
| let‐7 | tta‐miR*‐let‐7 | UGAGGUAGUAGGUUGUAUAGU | CUGUACAACUUGCUAACUUUC(2) CUGUACAACUUGCUAACUUUCC(4) | GCCGGGUUGAGGUAGUAGGUUGUAUAGUAAUGAACUACAACACUUGGGAGUACUGUACAACUUGCUAACUUUCCCUCGC | 1,826 | 1,846 | 6 | −31.9 | 45.56962 |
3.5. Identification of plant miRNA family in Thrips tabaci sRNA library
Interestingly, this study has identified mir‐9774 (Expression value 6), a plant microRNA family in our T. tabaci sRNA library (Table 3).
3.6. Phylogenetic analysis of Thrips tabaci miRNAs
Phylogenetic analyses revealed that most of the known miRNAs are highly conserved (Table 4, Figure 4a1–d1 and Figure 4a3–d3) among various species within the Kingdom and the phylogenetic trees for miR‐8, miR‐14, miR‐276, and miR‐281 revealed that T. tabaci miRNAs grouped with the closely related species of insects (Figure 4a2–d2). Figure 4 also revealed that T. tabaci miRNAs are well conserved, particularly in the seed region compared to the homologous miRNAs from other species.
Figure 4.
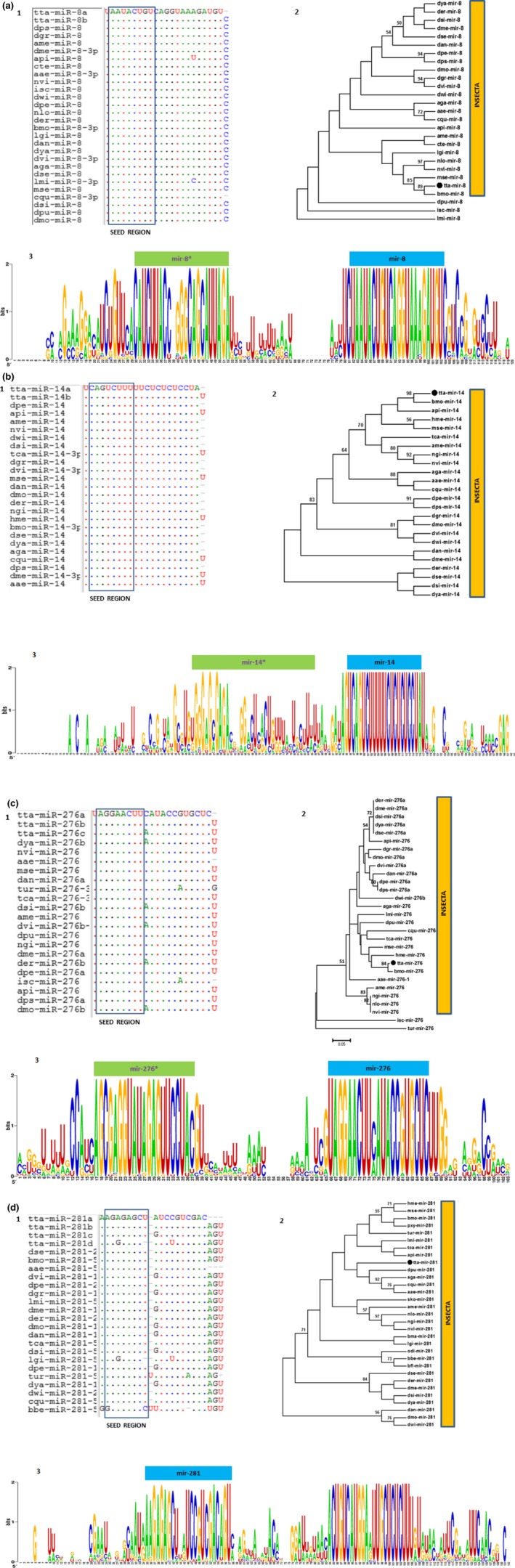
(a–d): 1. Homology in the seed region of the Thrips tabaci miRNAs (a–d are for mir‐8, mir‐14, mir‐276, and mir‐281, respectively) with respect to its counterpart from other insect species. The first three letters of each miRNAs indicating the name of the species (e.g.,: dya‐ Drosophila yakuba). (a–d): 2. Maximum Likelihood tree (RaxML.v.7.0.4) indicating the phylogenetic relationship of precursor miRNA sequences from various members of the animal kingdom. (a–d): 3. Thrips tabaci pre‐miRNAs weblogo indicating both mature (blue bar) and the star (green bars) sequences. Each logo consists of stacks of symbols, one for each nucleotide position in the sequence. The height indicates the sequence conservation at that nucleotide position and the height of symbols within the stack indicates the relative frequency of each nucleotide at that position
3.7. Identification of targets for Thrips tabaci miRNAs
Targets were predicted for known and novel miRNAs of T. tabaci employing miRanda with a scale of 0–7 to indicate the stringency of miRNA‐target pairing with the smaller numbers representing higher stringency. ESTs and transcriptome of F. occidentalis were used as a reference for target searches with a cut‐off score 140.
3.7.1. Targets for known miRNAs from Thrips tabaci
One hundred and thirty known miRNAs were searched for targets against ESTs and transcriptome sequences of F. occidentalis. A total of 218 and 1,025 targets were obtained from ESTs and transcriptome, respectively (Tables S1 and S2). The Blast‐2‐GO enrichment analysis was performed employing gene ontology (GO) terms for genes targeted by these miRNAs (Figure 5a,b). For those targets in the ESTs, three motifs were over‐represented in GO–BP (biological process) category viz. “metabolic process,” “transport,” and “catabolic process.” The GO–MF (molecular function) category was over‐represented by the motif “oxidoreductase activity” and “catalytic activity” (Figure 5a). On the other hand, GO terms enrichment analysis of miRNA targets in the transcriptome yielded motifs for “transport,” “signal transduction,” and “metabolic process” in GO‐BP category; while, GO‐MF category was over‐represented with motifs for “ATP binding,” “transferase activity,” and “binding” (Figure 5b). Complete details of the Blast‐2‐GO analysis were provided in Tables S3 and S4.
Figure 5.
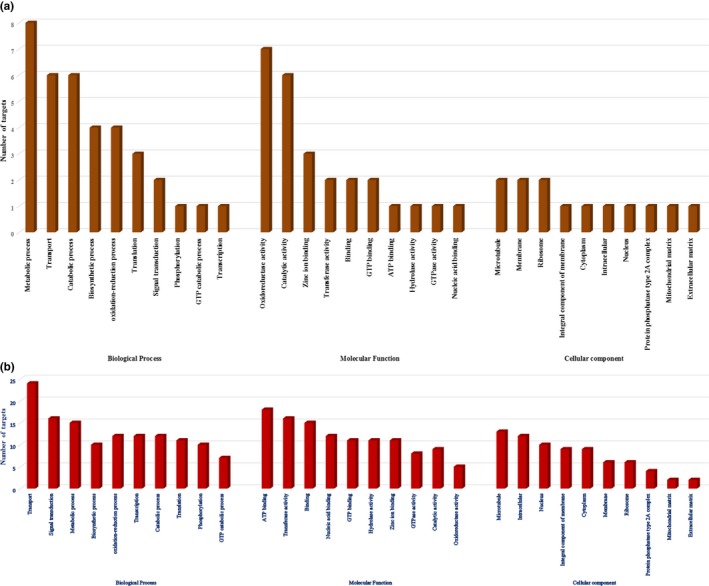
Gene Ontology (GO) classification of the putative target genes for the conserved T. tabaci miRNAs against ESTs (a) and transcriptome (b) sequences of F. occidentalis. GO terms was assigned to each target gene based on the annotation and were summarized into three main GO categories viz. (1) biological process (BP) (2) molecular function (MF), and (3) cellular component (CC). Only top ten subcategories are presented here
3.7.2. Targets for novel miRNAs from Thrips tabaci
Novel miRNAs were searched for their targets in the F. occidentalis transcriptome. A total of 65 miRNA‐target pairs were obtained (Table S5), and further Blast‐2‐GO analysis indicated the over‐representation of “Transport” and “ATP binding” as GO‐BP and GO‐MF category, respectively (Figure 6 and Table S6).
Figure 6.
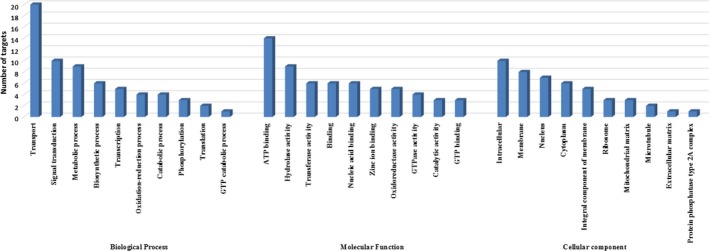
Gene Ontology (GO) classification of the putative target genes for the T. tabaci miRNAs against transcriptome sequences of F. occidentalis. GO terms was assigned to each target gene based on the annotation and were summarized into three main GO categories viz. (1) biological process (BP) (2) molecular function (MF), and (3) cellular component (CC). Only top ten subcategories are presented here
3.7.3. Synteny analysis using Circos
The synteny analysis of the T. tabaci miRNAs and their targets were performed by employing circos (Krzywinski et al., 2009). In brief, the Blast analysis was performed using T. tabaci miRNA sequences (known and novel) against F. occidentalis scaffolds (Approx. largest 200). The positions of miRNAs were identified and their targets are represented in the Circos plot (Figure 7).
Figure 7.
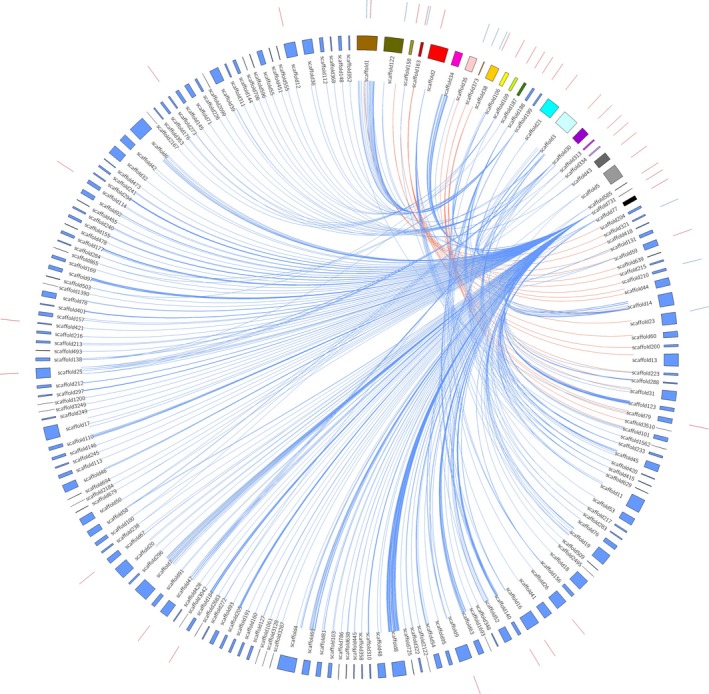
Map of the Western Flower Thrips, F. occidentalis scaffolds linking T. tabaci miRNAs and their putative targets prepared using Circos (Krzywinski et al., 2009). The outer circle represents the highlights of nine novel miRNA represented in blue and 34 known miRNA represented in red color. The inner circle marks each scaffold in a different color. The blue lines in the center of the figure connect a known miRNAs with its target that are represented across 173 scaffolds of F. occidentalis genome. Whereas, the orange lines in the center represent the interaction of novel miRNA with its target positions
3.8. Validation of Thrips tabaci microRNAs
This study revealed 130 known and nine novel miRNAs from T. tabaci. However, further validation of these miRNAs was performed by (1) stem‐loop endpoint reverse transcriptase PCR (RT‐PCR) and (2) real‐time quantitative reverse transcriptase PCR (RT‐qPCR). Using stem‐loop endpoint RT‐PCR, we have validated six conserved viz. tta‐miR‐281, tta‐miR‐276, tta‐miR‐10, tta‐miR‐100, tta‐miR‐184, and tta‐miR‐3533 and four novel miRNAs viz. tta‐miR‐N1, tta‐miR‐N4, tta‐miR‐N7, tta‐miR‐N9 from T. tabaci using the primer sets as described (Table S7). All of these miRNAs were amplified with an approximate product size of 75 bp (Figure 8a).
Figure 8.
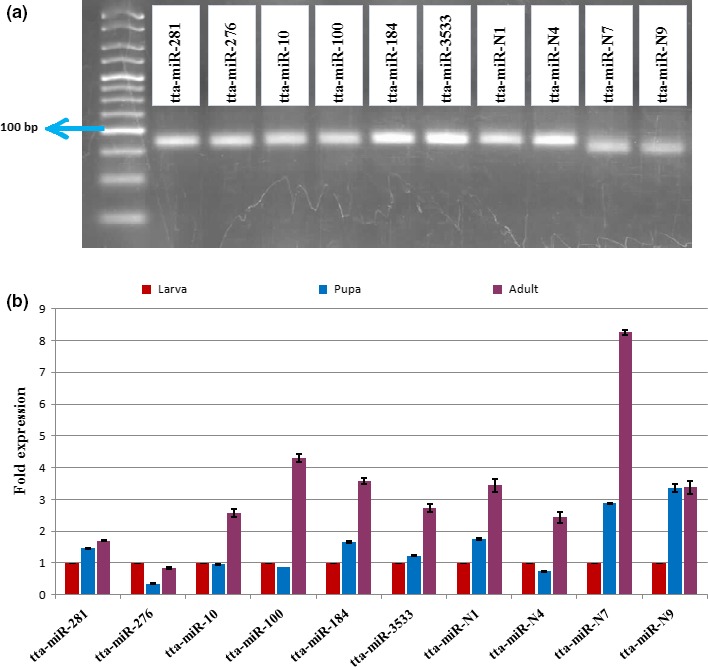
(a) Stem‐loop RT‐PCR analyses of six conserved and four novel miRNAs from Thrips tabaci. The products were resolved on 3% agarose gel in 1X TBE stained with ethidium bromide and HyperLadder™ 25 bp (Bioline, USA) used as marker. (b) Stem‐loop RT‐qPCR analysis of spatiotemporally expressed T. tabaci miRNAs in larva, pupa and adults. “*” and “**” means a statistically significant difference at level p < .05 and p < .001, respectively, for these miRNAs in the larva, pupae, and adult T. tabaci. The error bars indicate standard deviation for three biological replications
Our study also quantified the expression level of the above‐mentioned ten miRNAs from T. tabaci larva, pupa, and adult using RT‐qPCR (Table S8, Figure 8b). Results suggested that the miRNA expression was higher in pupal and adult stages compared to larval stages in six microRNAs such as tta‐miR‐281, tta‐miR‐184, tta‐miR‐3533, tta‐miR‐N1, tta‐miR‐N7, and tta‐miR‐N9 (Figure 8b).
4. DISCUSSION
The onion thrips, Thrips tabaci, is an important pest species and a tospovirus vector causing significant negative impacts on yield and quality of various economically important crops (German et al., 1992). Although microRNAs are key gene regulators and are involved in many biological processes, including growth and development, no previous study has been conducted on the identification and validation of miRNAs in T. tabaci. MicroRNAs are known from more than 25 insect species, (Stark et al., 2007). Several miRNAs have been reported from various orders of insects such as Diptera, Hymenoptera, Coleoptera, Orthoptera, Lepidoptera, Hemiptera, Homoptera (Wu et al., 2013), and Thysanoptera (Rebijith, Asokan, Hande, & Krishna Kumar, 2016). This study reports the complete miRNA profile from onion thrips, Thrips tabaci. A small RNA library was prepared from the pooled samples of different developmental stages of T. tabaci and the high‐throughput Illumina deep‐sequencing technology (Avesson et al., 2012; Burnside et al., 2008; Ge et al., 2013; Koh et al., 2010) was used to identify miRNAs from the prepared library.
We used the F. occidentalis genome sequence as a reference for T. tabaci, as the complete genome T. tabaci is still not available in the database. The higher percentage of mapping (91%) was possible only because both these insects belong to the same family, Thripidae. Employing this approach, our study revealed 130 conserved and nine novel miRNAs from T. tabaci. The size distributions of the high‐quality reads were varied from 18 to 28 nts in our library and the peak was at the 25 nt, which was on par with previous studies (Ge et al., 2013; Liang, Feng, Zhou, & Gao, 2013; Sattar et al., 2012). Our study indicated the unique read distributes of 26–28 nts with a relative lower abundance, which is common in many small RNA libraries (Chang et al., 2016; Jagadeeswaran et al., 2010; Surridge et al., 2011; Zhang et al., 2013), indicating the presence of piRNAs. Piwi RNAs (piRNAs) are the class of small RNAs mediating chromatin modifications (Ross, Weiner, & Lin, 2014) which are derived mainly from retrotransposons and other repetitive elements with high sequence diversity (Ross et al., 2014; Siomi, Sato, Pezic, & Aravin, 2011; Zhang et al., 2013). Thus, our results indicated that T. tabaci genome not only harbors miRNAs but also other small RNAs such as piRNAs that might be involved in the transgenerational epigenetic inheritance (Weick & Miska, 2014).
MiRNAs are evolutionarily conserved regulators of gene expression (Rebijith et al., 2014; Zhang et al., 2009), and few can even act as markers in defining the evolutionary relationship in a wide range of insect species (Kakumani et al., 2015). Our homology and phylogeny analysis revealed that insect miRNAs are well‐conserved, despite considerable diversity in the genome (Figure 4a–d). MiRNA*s are not easily detectable as it degrades soon after being exported to the cytosol (Wu et al., 2013). However, our results indicated the presence of several miRNA*s (Tables 5 and 6) that matched to the same precursor sequences with their mismatched complementary mature miRNAs.
We identified the presence of a plant‐specific miRNA family, mir‐9774 in the T. tabaci sRNA library, and the same has been recently reported from Triticum aestivum L. and Brachypodium distachyon (L.) Beauv (Wei et al., 2009). Previous miRNA studies on cotton/melon aphid, A. gossypii also reported six plant miRNA family (Sattar et al., 2012). They also showed that such microRNAs were transformed into the aphid tissues (especially in gut contents) during the phloem sap ingestion. However, none of those six have been identified in our sRNA library.
Our results showed that the highest expression is for tta‐miR‐276 with an expression value of 26,418. Very recent studies showed that miR‐276 expressed in the ovaries of female locusts mediates progeny egg‐hatching synchrony by upregulating its target brahma (brm), a transcription coactivator gene (He et al., 2016). Thus, it is plausible that miR‐276 enhances brm expression to promote developmental synchrony and provide insight into the regulation of developmental homeostasis in T. tabaci. The second highest expression is for miR‐281 with an expression value of 18,063 and might be involved in the development and metamorphosis of T. tabaci as recent studies showed that miR‐281 regulates the expression of ecdysone receptor (EcR) isoform B, in Bombyx mori (Jiang et al., 2013). Another interesting microRNA obtained in the current study was miR‐8, and it can target the Wingless signalling pathway to regulate secretion of yolk protein precursors by the female mosquito fat body and accumulation into the developing ovaries (Lucas et al., 2015, http://www.smartscitech.com/index.php/RD/article/view/815). Therefore, it is quite possible that miR‐8 may play a key role in the reproductive processes of T. tabaci. An insect‐specific miR‐14 was identified in T. tabaci with an expression value of 12,453 and studies on lepidopteran insects showed the antiapoptotic role of miR‐14 (Kumarswamy & Chandna, 2010). The rest of the species‐specific miRNAs identified in T. tabaci might play important role in insect‐specific features, such as metamorphosis, parthenogenesis, and biogenesis of pheromones (Zhang et al., 2007). Whereas, the other invertebrate‐ and vertebrate‐specific miRNAs (Table 3) identified from T. tabaci required special attention, as their nonexistence in other species of insects could be due to the absence of complete genomic information for most of those insects (Ge et al., 2013).
The expression profile of miRNA varies spatiotemporally among different developmental stages (Li, Cassidy, Reinke, Fischboeck, & Carthew, 2009; Xu, Zhou, Wang, Auersperg, & Peng, 2006), and the developmental expression profiles (larval, pupal and adult stage) of ten microRNAs were studied by RT‐ qPCR (Figure 8b). The higher expression of tta‐miR‐281, tta‐miR‐184, tta‐miR‐3533, tta‐miR‐N1, tta‐miR‐N7, and tta‐miR‐N9 in T. tabaci pupal and adult stages reflected their possible role in parthenogenesis, adult development, and sexual reproduction. The high levels of miR‐276 in the larval stage indicated their possible involvement in insect‐specific features such as metamorphosis.
miRNAs regulate the gene expression through targeting transcripts that can bring about mRNA cleavage, mRNA decay or translational repression of target mRNAs by binding to 3′ UTRs, 5′ UTRs, and even to coding regions (Lytle, Yario, & Steitz, 2007). Thus, it is important to identify the gene targets and thereby we can understand the biological role of a particular miRNA. As miRNA targets have been identified using the (1) expressed sequence tags (ESTs) and (2) transcriptomic sequences of F. occidentalis. The GO annotations for the predicted targets were classified as potential biological process, cellular component, and molecular function. The putative targeted genes included signal transduction pathways, transcription factors, reproduction, embryo development, insect molting, immune response, and even metabolism. Overall, the results from our study indicated that these conserved and novel miRNAs identified from T. tabaci might play crucial regulatory role in the regulation of thrips growth and development.
5. CONCLUSIONS
In summary, the result from our study add to the pool of miRNA databases and is the first report of small RNAs from T. tabaci, a nonmodel insect lacking genome information. One hundred and thirty conserved and nine novel miRNAs were identified with high confidence and sufficient evidence is the major contribution of our study. Sequence analyses revealed that most of the T. tabaci miRNAs are highly conserved in various species, making miRNAs, a hallmark of evolutionarily conserved regulators of gene expression. To harmonize the data and to provide more useful biological insights, we have also carried out in silico analysis of identifying potential targets for these miRNAs. Our results indicated that the list of putative mRNA targets was very extensive and most of the putative target genes for T. tabaci miRNAs were associated with several KEGG pathways such as metabolic process, transport, translation, signal pathways, and oxidative phosphorylation. However, further experiments are required for the validation of these targets. Expression levels of T. tabaci miRNAs were validated by RT‐qPCR, and the results indicated few of these miRNAs have been predicted in the adult development process, which can be further utilized in gene functional studies through RNAi‐based approach or in developing miRNA mimics both for feeding and in planta expression (Agrawal, Sachdev, Rodrigues, Sowjanya Sree, & Bhatnagar, 2013; Jayachandran, Hussain, & Asgari, 2013; Nandety et al., 2015) as novel pest management strategies based on gene silencing and insect transgenesis.
DATA AVAILABILITY
All relevant data are within the paper and its Supporting Information files. The small RNA Sequence data has been submitted to NCBI under the BioSample project ‘PRJNA350618’; BioSample Accession: ‘SAMN05943039’.
CONFLICT OF INTEREST
The authors have declared that no competing interests exist.
AUTHORS’ CONTRIBUTIONS
Conceptualization: KBR HRH, Experiments: KBR HRH, Reagents/materials: KBR RA SG, Writing—original draft: KBR, Writing—review and editing: KBR HRH RA SG NKK.
Supporting information
ACKNOWLEDGMENTS
We thank Sonal Dsouza, Communication and Programme Assistant; Bioversity International for language editing that greatly improved the manuscript. Our sincere thanks are due to The Director, IIHR, Bangalore for providing necessary facilities.
Balan RK, Ramasamy A, Hande RH, Gawande SJ, Krishna Kumar NK. Genome‐wide identification, expression profiling, and target gene analysis of microRNAs in the Onion thrips, Thrips tabaci Lindeman (Thysanoptera: Thripidae), vectors of tospoviruses (Bunyaviridae). Ecol Evol. 2018;8:6399–6419. 10.1002/ece3.3762
Contributor Information
Rebijith K. Balan, Email: rebijith@gmail.com.
Asokan Ramasamy, Email: asokaniihr@gmail.com.
REFERENCES
- Agrawal, N. , Sachdev, B. , Rodrigues, J. , Sowjanya Sree, K. , & Bhatnagar, R. K. (2013). Development associated profiling of chitinase and microRNA of Helicoverpa armigera identified chitinase repressive microRNA. Scientific Reports, 3, 2292 10.1038/srep02292 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ambros, V. (2004). The functions of animal microRNAs. Nature, 431, 350–355. 10.1038/nature02871 [DOI] [PubMed] [Google Scholar]
- Avesson, L. , Reimegard, J. , Wagner, E. G. , & Söderbom, F. (2012). MicroRNAs in Amoebozoa: Deep sequencing of the small RNA population in the social amoeba, Dictyostelium discoideum reveals developmentally regulated microRNAs. RNA, 18, 1771–1782. 10.1261/rna.033175.112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartel, D. P. (2004). MicroRNAs: Genomics biogenesis mechanism and function. Cell, 116, 281–297. 10.1016/S0092-8674(04)00045-5 [DOI] [PubMed] [Google Scholar]
- Burnside, J. , Ouyang, M. , Anderson, A. , Bernberg, E. , Lu, C. , Meyers, B. C. , … Morgan, R. W. (2008). Deep sequencing of chicken microRNAs. BMC Genomics, 9, 185. 10.1186/1471-2164-9-185 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bustin, S. A. , Benes, V. , Garson, J. A. , Hellemans, J. , Huggett, J. , & Kubista, M. (2009). The MIQE guidelines‐minimum information for publication of quantitative real‐time PCR experiments. Clinical Chemistry, 55, 611–622. 10.1373/clinchem.2008.112797 [DOI] [PubMed] [Google Scholar]
- Calla, B. , & Geib, S. M. (2015). MicroRNAs in the oriental fruit fly, Bactrocera dorsalis: Extending Drosophilid miRNA conservation to the Tephritidae. BMC Genomics, 16, 740 10.1186/s12864-015-1835-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang, Z. X. , Tang, N. , Wang, L. , Zhang, L. Q. , Akinyemi, I. A. , & Wu, Q. F. (2016). Identification and characterization of microRNAs in the white‐backed planthopper, Sogatella furcifera . Insect Science, 23, 452–468. 10.1111/1744-7917.12343 [DOI] [PubMed] [Google Scholar]
- Chekulaeva, M. , & Filipowicz, W. (2009). Mechanisms of miRNA‐mediated post‐transcriptional regulation in animal cells. Current Opinion in Cell Biology, 21, 452–460. 10.1016/j.ceb.2009.04.009 [DOI] [PubMed] [Google Scholar]
- Chen, C. , Ridzon, D. A. , Broomer, A. J. , Zhou, Z. , Lee, D. H. , Nguyen, J. T. , … Lao, K. Q. (2005). Real‐time quantification of microRNAs by stem–loop RT‐PCR. Nucleic Acids Research, 33, e179 10.1093/nar/gni178 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cullen, B. R. (2006). Viruses and microRNAs. Nature Genetics, 38(Suppl.), S25–S30. 10.1038/ng1793 [DOI] [PubMed] [Google Scholar]
- DeGraaf, H. E. , & Wood, G. M. (2009). An improved method for rearing western flower thrips Frankliniella occidentalis . Florida Entomologist, 92(4), 664–666. 10.1653/024.092.0424 [DOI] [Google Scholar]
- Diaz‐Montano, J. , Fuchs, M. , Nault, B. A. , Fail, J. , & Shelton, A. M. (2011). Onion thrips (Thysanoptera: Thripidae): A global past of increasing concern in onion. Journal of Economic Entomology, 104, 1–13. 10.1603/EC10269 [DOI] [PubMed] [Google Scholar]
- Enright, A. J. , John, B. , Gaul, U. , Tuschl, T. , Sander, C. , & Marks, D. S. (2004). MicroRNA targets in Drosophila . Genome Biology, 5(1), R1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fabian, M. R. , Sonenberg, N. , & Filipowicz, W. (2010). Regulation of mRNA translation and stability by microRNAs. Annual Review of Biochemistry, 79, 351–379. 10.1146/annurev-biochem-060308-103103 [DOI] [PubMed] [Google Scholar]
- Friedlander, M. R. , Mackowiak, S. D. , Li, N. , Chen, W. , & Rajewsky, N. (2012). miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Research, 40, 37–52. 10.1093/nar/gkr688 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Friedman, R. C. , Farh, K. K. , Burge, C. B. , & Bartel, D. P. (2009). Most mammalian mRNAs are conserved targets of microRNAs. Genome Research, 19, 92–105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ge, X. , Zhang, Y. , Jiang, J. , Zhong, Y. , Yang, X. , Li, Z. , … Tan, A. (2013). Identification of MicroRNAs in Helicoverpa armigera and Spodoptera litura based on Deep sequencing and homology analysis. International Journal of Biological Sciences, 9(1), 1–15. 10.7150/ijbs.5249 [DOI] [PMC free article] [PubMed] [Google Scholar]
- German, T. L. , Ullman, D. E. , & Moyer, J. W. (1992). Tospoviruses: Diagnosis, molecular biology, phylogeny, and vector relationships. Annual Review of Phytopathology, 30, 315–348. 10.1146/annurev.py.30.090192.001531 [DOI] [PubMed] [Google Scholar]
- Griffiths‐Jones, S. , Grocock, R. J. , van Dongen, S. , Bateman, A. , & Enright, A. J. (2006). miRBase: MicroRNA sequences, targets and gene nomenclature. Nucleic Acids Research, 34(Database Issue), D140–D144. 10.1093/nar/gkj112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Griffiths‐Jones, S. , Saini, H. K. , van Dongen, S. , & Enright, A. J. (2008). miRBase: Tools for microRNA genomics. Nucleic Acids Research, 36(Database issue), D154–D158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guillem, Y. , Bastian, F. , Maria‐Dolors, P. , & Belles, X. (2016). The microRNA toolkit of insects. Scientific Reports, 6, 37736 10.1038/srep37736 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hall, T. A. (1999). BioEdit: A user‐friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series, 41, 95–98. [Google Scholar]
- He, J. , Chen, Q. , Wei, Y. , Jiang, F. , Yang, M. , Hao, S. , … Kang, L. (2016). MicroRNA‐276 promotes egg‐hatching synchrony by up‐regulating brmin locusts. Proceedings of National Academy of Sciences of the United States of America, 113(3), 584–589. 10.1073/pnas.1521098113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu, H. Y. , He, L. , Fominykh, K. , Yan, Z. , Guo, S. , Zhang, X. , … Wang, W. (2012). Evolution of the human‐specific microRNA miR‐941. Nature Communications, 3, 1145 10.1038/ncomms2146 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jagadeeswaran, G. , Zheng, Y. , Sumathipala, N. , Jiang, H. B. , Arrese, E. L. , Soulages, J. L. , … Sunkar, R. (2010). Deep sequencing of small RNA libraries reveals dynamic regulation of conserved and novel microRNAs and microRNA‐stars during silkworm development. BMC Genomics, 11, 52 10.1186/1471-2164-11-52 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jayachandran, B. , Hussain, M. , & Asgari, S. (2013). An insect trypsin‐like serine protease as a target of miRNA: Utilization of miRNA mimics and inhibitors by oral feeding. Insect Biochemistry and Molecular Biology, 43(4), 398–406. 10.1016/j.ibmb.2012.10.004 [DOI] [PubMed] [Google Scholar]
- Jiang, J. , Ge, X. , Li, Z. , Wang, Y. , Song, Q. , Stanley, D. W. , … Huang, Y. (2013). MicroRNA‐281 regulates the expression of ecdysone receptor (EcR) isoform B in the silkworm, Bombyx mori . Insect Biochemistry and Molecular Biology, 43, 692–700. 10.1016/j.ibmb.2013.05.002 [DOI] [PubMed] [Google Scholar]
- Kakumani, P. K. , Chinnappan, M. , Singh, A. K. , Malhotra, P. , Mukherjee, S. K. , & Bhatnagar, R. K. (2015). Identification and characteristics of microRNAs from army worm, Spodoptera frugiperda cell line Sf21. PLoS One, 10(2), e0116988 10.1371/journal.pone.0116988 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kang, Y. , Gao, J. , Yang, T. , Ma, Q. , Qiu, X. , Fan, Q. , & Ma, B. (2012). MicroRNA‐34a inhibits the proliferation and metastasis of osteosarcoma cells both in vitro and in vivo. PLoS One, 7(3), e33778 10.1371/journal.pone.0033778 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koh, W. , Sheng, C. T. , Tan, B. , Lee, Q. Y. , Kuznetsov, V. , Kiang, L. S. , & Tanavde, V. (2010). Analysis of deep‐sequencing microRNA expression profile from human embryonic stem cells derived mesenchymal stem cells reveals possible role of let‐7 microRNA family in downstream targeting of hepatic nuclear factor 4 alpha. BMC Genomics, 11(Suppl. 1), S6 10.1186/1471-2164-11-S5-S6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krzywinski, M. , Schein, J. , Birol, İ. , Connors, J. , Gascoyne, R. , Horsman, D. , … Marra, M. A. (2009). Circos: An information aesthetic for comparative genomics. Genome Research, 19(9), 1639–1645. 10.1101/gr.092759.109 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kumarswamy, R. , & Chandna, S. (2010). Inhibition of microRNA‐14 contributes to actinomycin‐D induced apoptosis in the Sf9 insect cell line. Cell Biology International, 34, 851–857 (Printed in Great Britain). 10.1042/CBI20100035 [DOI] [PubMed] [Google Scholar]
- Lagos‐Quintana, M. , Rauhut, R. , Lendeckel, W. , & Tuschl, T. (2001). Identification of novel genes coding for small expressed RNAs. Science, 294, 853–858. 10.1126/science.1064921 [DOI] [PubMed] [Google Scholar]
- Lee, R. C. , & Ambros, V. (2001). An extensive class of small RNAs in Caenorhabditis elegans . Science, 294, 862–864. 10.1126/science.1065329 [DOI] [PubMed] [Google Scholar]
- Lee, R. C. , Feinbaum, R. L. , & Ambros, V. (1993). The C. elegans heterochronic gene lin‐4 encodes small RNAs with antisense complementarity to lin‐14. Cell, 75, 843–854. 10.1016/0092-8674(93)90529-Y [DOI] [PubMed] [Google Scholar]
- Lewis, T. (1973). Thrips: Their biology, ecology and economic importance (p. 349). London, UK: Academic Press. [Google Scholar]
- Lewis, T. (1997). Pest thrips in perspective In Lewis T. (Ed.), Thrips as crop pests (pp. 1–13). New York, NY: CAB International. [Google Scholar]
- Li, X. , Cassidy, J. J. , Reinke, C. A. , Fischboeck, S. , & Carthew, R. W. (2009). A microRNA imparts robustness against environmental fluctuation during development. Cell, 137, 273–282. 10.1016/j.cell.2009.01.058 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liang, P. , Feng, B. , Zhou, X. , & Gao, X. (2013). Identification and developmental profiling of microRNAs in diamondback moth, Plutella xylostella (L.). PLoS One, 8(11), e78787 10.1371/journal.pone.0078787 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lim, L. P. , Glasner, M. E. , Yekta, S. , Burge, C. B. , & Bartel, D. P. (2003). Vertebrate microRNA genes. Science, 299(5612), 1540 10.1126/science.1080372 [DOI] [PubMed] [Google Scholar]
- Livak, K. J. , & Schmittgen, T. D. (2001). Analysis of relative gene expression data using real‐time quantitative PCR and the 2−ΔΔCT method. Methods, 25(4), 402–408. 10.1006/meth.2001.1262 [DOI] [PubMed] [Google Scholar]
- Lucas, K. J. , Roy, S. , Ha, J. , Gervaise, A. L. , Kokoza, V. A. , & Raikhel, A. S. (2015). MicroRNA‐8 targets the Wingless signaling pathway in the female mosquito fat body to regulate reproductive processes. Proceedings of National Academy of Sciences of the United States of America, 12, 1440–1445. 10.1073/pnas.1424408112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lytle, J. R. , Yario, T. A. , & Steitz, J. A. (2007). Target mRNAs are repressed as efficiently by microRNA‐binding sites in the 5′ UTR as in the 3′ UTR. Proceedings of National Academy of Sciences of the United States of America, 104, 9667–9672. 10.1073/pnas.0703820104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mandal, B. , Jain, R. K. , Krishnareddy, M. , Krishna Kumar, N. K. , Ravi, K. S. , & Pappu, H. R. (2012). Emerging problems of tospoviruses (Bunyaviridae) and their management in the Indian subcontinent. Plant Disease, 96, 468–479. 10.1094/PDIS-06-11-0520 [DOI] [PubMed] [Google Scholar]
- Miska, E. A. , Alvarez‐Saavedra, E. , Abbott, A. L. , Lau, N. C. , Hellman, A. B. , McGonagle, S. M. , … Horvitz, H. R. (2007). Most Caenorhabditis elegans microRNAs are individually not essential for development or viability. PLoS Genetics, 3(12), e215 10.1371/journal.pgen.0030215 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nandety, R. S. , Sharif, A. , Kamita, S. G. , Ramasamy, A. , & Falk, B. W. (2015). Identification of novel and conserved microRNAs in Homalodisca vitripennis, the glassy‐winged sharpshooter by expression profiling. PLoS One, 10(10), e0139771 10.1371/journal.pone.0139771 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nohata, N. , Hanazawa, T. , Kinoshita, T. , Okamoto, Y. , & Seki, N. (2012). MicroRNAs function as tumor suppressors or oncogenes: Aberrant expression of microRNAs in head and neck squamous cell carcinoma. Auris, Nasus, Larynx, 40(2), 143–149. [DOI] [PubMed] [Google Scholar]
- Pittman, H. A. (1927). Spotted wilt of tomatoes. Journal of Australia Council for Scientific and Industrial Research, 1, 74–77. [Google Scholar]
- Rebijith, K. B. , Asokan, R. , Hande, H. R. , & Krishna Kumar, N. K. (2016). The first report of miRNAs from a thysanopteran insect, Thrips palmi karny using high‐throughput sequencing. PLoS One, 11(9), e0163635 10.1371/journal.pone.0163635 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rebijith, K. B. , Asokan, R. , Krishna, V. , Ranjitha, H. H. , Kumar, N. K. , & Ramamurthy, V. V. (2014). In silico prediction and characterization of MicroRNAs from Aphis gossypii (Hemiptera: Aphididae). Annals of the Entomological Society of America, 107(2), 521–531. 10.1603/AN12158 [DOI] [Google Scholar]
- Ross, R. J. , Weiner, M. M. , & Lin, H. (2014). PIWI proteins and PIWI‐interacting RNAs in the soma. Nature, 505, 353–359. 10.1038/nature12987 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sattar, S. , Addo‐Quaye, C. , Song, Y. , Anstead, J. A. , Sunkar, R. , & Thompson, G. A. (2012). Expression of small RNA in Aphis gossypii and its potential role in the resistance interaction with melon. PLoS One, 7, e48579 10.1371/journal.pone.0048579 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Singh, J. , & Nagaraju, J. (2008). In silico prediction and characterization of microRNAs from red flour beetle (Tribolium castaneum). Insect Molecular Biology, 17, 427–436. 10.1111/j.1365-2583.2008.00816.x [DOI] [PubMed] [Google Scholar]
- Siomi, M. C. , Sato, K. , Pezic, D. , & Aravin, A. A. (2011). PIWI‐interacting small RNAs: The vanguard of genome defence. Nature Reviews Molecular Cell Biology, 12, 246–258. 10.1038/nrm3089 [DOI] [PubMed] [Google Scholar]
- Smith, T. F. , & Waterman, M. S. (1981). Identification of common molecular subsequences. Journal of Molecular Biology, 147, 195–197. 10.1016/0022-2836(81)90087-5 [DOI] [PubMed] [Google Scholar]
- Song, Q. X. , Liu, Y. F. , Hu, X. Y. , Zhang, W. K. , Ma, B. , Chen, S. Y. , & Zhang, J.‐S. (2011). Identification of miRNAs and their target genes in developing soybean seeds by deep sequencing. BMC Plant Biology, 11, 5 10.1186/1471-2229-11-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Srinivasan, R. , Sundaraj, S. , Pappu, H. R. , Diffie, S. , Riley, D. G. , & Gitaitis, R. D. (2012). Transmission of Iris yellow spot virus by Frankliniella fusca and Thrips tabaci (Thysanoptera: Thripidae). Journal of Economic Entomology, 105(1), 40–47. 10.1603/EC11094 [DOI] [PubMed] [Google Scholar]
- Stamatakis, A. (2008). The RAxML 704 manual. Retrieved from http://icwwwepflch/~stamatak/
- Stark, A. , Kheradpour, P. , Parts, L. , Brennecke, J. , Hodges, E. , Hannon, G. J. , & Kellis, M. (2007). Systematic discovery and characterization of fly microRNAs using 12 Drosophila genomes. Genome Research, 17(12), 1865–1879. 10.1101/gr.6593807 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Surridge, A. K. , Lopez‐Gomollon, S. , Moxon, S. , Maroja, L. S. , Rathjen, T. , Nadeau, N. J. , … Jiggins, C. D. (2011). Characterisation and expression of microRNAs in developing wings of the neotropical butterfly Heliconius melpomene . BMC Genomics, 12, 62 10.1186/1471-2164-12-62 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang, F. , Li, L. , Liu, L. , Li, H. , Zhang, Y. , Yao, Y. , … Gao, J. (2012). High‐throughput sequencing discovery of conserved and novel microRNAs in Chinese cabbage (Brassica rapa L. ssp. pekinensis). Molecular Genetics and Genomics, 287, 555–563. 10.1007/s00438-012-0699-3 [DOI] [PubMed] [Google Scholar]
- Wei, B. , Cai, T. , Zhang, R. , Li, A. , Huo, N. , Li, S. , … Mao, L. (2009). Novel microRNAs uncovered by deep sequencing of small RNA transcriptomes in bread wheat (Triticum aestivum L.) and Brachypodium distachyon (L.) Beauv. Functional and Integrative Genomics, 9, 499–511. 10.1007/s10142-009-0128-9 [DOI] [PubMed] [Google Scholar]
- Weick, E. M. , & Miska, E. A. (2014). piRNAs: From biogenesis to function. Development, 141, 3458–3471. 10.1242/dev.094037 [DOI] [PubMed] [Google Scholar]
- Wu, W. , Ren, Q. , Li, C. , Wang, Y. , Sang, M. , Zhang, Y. , & Li, B. (2013). Characterization and comparative profiling of MicroRNAs in a sexual dimorphism insect, Eupolyphaga sinensis walker. PLoS One, 8(4), e59016 10.1371/journal.pone.0059016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu, G. , Zhou, H. , Wang, Q. , Auersperg, N. , & Peng, C. (2006). Activin receptor‐like kinase 7 induces apoptosis through up‐regulation of Bax and down‐regulation of Xiap in normal and malignant ovarian epithelial cell lines. Molecular Cancer Research, 4, 235–246. 10.1158/1541-7786.MCR-05-0174 [DOI] [PubMed] [Google Scholar]
- Zhang, L. , Ding, L. , Cheung, T. H. , Dong, M. Q. , Chen, J. , Sewell, A. K. , … Han, M. (2007). Systematic identification of C. elegans miRISC proteins, miRNAs, and mRNA targets by their interactions with GW182 proteins AIN‐1 and AIN‐2. Molecular Cell, 28, 598–613. 10.1016/j.molcel.2007.09.014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang, B. H. , Pan, X. P. , Cannon, C. H. , Cobb, G. P. , & Anderson, T. A. (2006). Conservation and divergence of plant microRNA genes. Plant Journal, 46, 243–259. 10.1111/j.1365-313X.2006.02697.x [DOI] [PubMed] [Google Scholar]
- Zhang, Y. , Zhang, X. O. , Chen, T. , Xiang, J. F. , Yin, Q. F. , Xing, Y. H. , … Chen, L. L. (2013). Circular intronic long noncoding RNAs. Molecular Cell, 51(6), 792–806. 10.1016/j.molcel.2013.08.017 [DOI] [PubMed] [Google Scholar]
- Zhang, X. , Zheng, Y. , Jagadeeswaran, G. , Ren, R. , Sunkar, R. , & Jiang, H. (2012). Identification and developmental profiling of conserved and novel microRNAs in Manduca sexta . Insect Biochemistry and Molecular Biology, 42, 381–395. 10.1016/j.ibmb.2012.01.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang, Y. , Zhou, X. , Ge, X. , Jiang, J. , Li, M. , Jia, S. , … Li, F. (2009). Insect‐Specific microRNA involved in the development of the silkworm Bombyx mori . PLoS One, 4, e4677 10.1371/journal.pone.0004677 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All relevant data are within the paper and its Supporting Information files. The small RNA Sequence data has been submitted to NCBI under the BioSample project ‘PRJNA350618’; BioSample Accession: ‘SAMN05943039’.


