Abstract
This study examines the impact of cefepime and APP-β (antipseudomonal penicillin/ β-lactamase inhibitor combinations) on Gram-negative bacterial colonization and resistance in two Australian ICUs. While resistance did not cumulatively increase, cefepime (but not APP-β treatment) was associated with acquisition of antibiotic resistant Enterobacteriaceae, consistent with an ecological effect. Analysis of the resident gut E. coli population in a subset of patients showed an increase in markers of horizontal gene transfer after cefepime exposure that helps explain the increase in APP-β resistance and reminds us that unmeasured impacts on the microbiome are key outcome determinants that need to be fully explored.
Electronic supplementary material
The online version of this article (10.1186/s40635-018-0185-2) contains supplementary material, which is available to authorized users.
To the Editor,
Effects of late-generation cephalosporins such as cefepime (FEP) on resistance acquisition and the gut microflora are uncertain [1–4]. In a previous study in two Australian ICUs, nearly 70% of all prescriptions were allocated in respective cycles to either cefepime or an antipseudomonal penicillin/β-lactamase inhibitor (APP-β) like piperacillin/tazobactam [5]. Under this strong sustained selection, cefepime exposure resulted in more methicillin-resistant Staphylococcus aureus (MRSA) and Pseudomonas aeruginosa colonization and infection than APP-β despite equivalent in vitro susceptibility [5]. In order to determine whether clinically important Enterobacteriaceae were similarly affected, perineal samples from patients within this cohort who had been admitted directly to the ICU (n = 206) were cultured at admission (< 48-h ICU stay) and again after 3 days of cycle-specified antibiotic (FEP or APP-β) [5]. Resistance to gentamicin and APP-β were chosen as key phenotypes not associated with cefepime resistance. Resistant Enterobacteriaceae were cultured from a modest proportion at admission (to gentamicin, 14%; to APP-β, 26%) but with no cumulative increase over time, as for MRSA and Pseudomonas [5]. Colonization by APP-β-resistant Enterobacteriaceae increased significantly overall after ICU admission (p = 0.015) but was almost 2.5 times more likely after cefepime than APP-β exposure (p < 0.05) (Fig. 1).
Fig. 1.
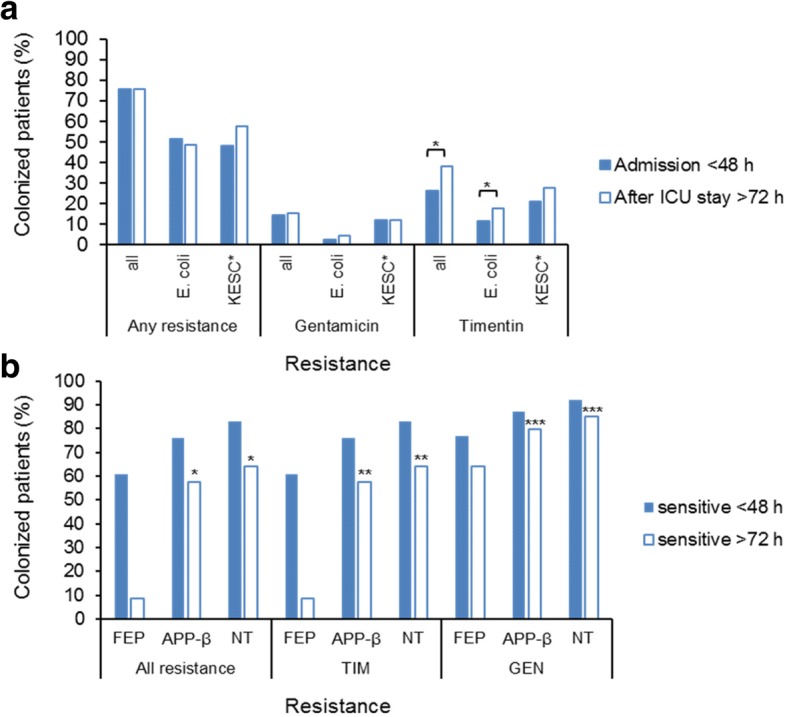
Variation in colonization patterns before and after ICU stay. a Rates of colonization with antibiotic-resistant species at admission and after ICU stay. b Change in susceptibility rates after ICU stay in different treatment cycles. Proportion of the entire study group of patients (n = 206) with positive cultures. “Any resistance” indicates any bacterial growth on antibiotic supplemented media. “All resistance” indicates any growth on ChromAgar™ supplemented with vancomycin only (to exclude Gram-positive species). *KESC, Klebsiella, Enterobacter, Serratia, Citrobacter spp. as identified on colorimetric media ChromAgar™ and confirmed by MALDI-TOF [16] as previously described [5]. Asterisks (*, **, ***) above bar charts indicate significant differences (p < 0.05)
Patients without resistant Enterobacteriaceae on admission were more likely to remain free of them after treatment with APP-β (or no drug) than cefepime (Fig. 1; *p = 0.004), and this association held when APP-β and cefepime treatment were directly compared (n = 45 vs. 22, p = 0.016). This was also true for APP-β (Fig. 1; **p = 0.002) and gentamicin resistance (Fig. 1; ***p = 0.014) when considered individually (Table 1). Backwards stepwise logistic regression analysis also linked cefepime exposure more strongly to later APP-β-resistance (OR 2.285 (CI 1.096–4.764); p = 0.027) than length of stay, age, or admission APACHE II score. Cefepime was also more strongly associated with APP-β-resistant Escherichia coli than APP-β itself (Pearson’s chi-square test, p = 0.027).
Table 1.
Effect of antibiotic on gain and loss of resistance in Enterobacteriaceae after 72 h in ICU
| Resistance | Treatmenta | Gainedb | Lostb | No changec | |
|---|---|---|---|---|---|
| Sensitive | Resistant | ||||
| Timentin and/or gentamicin | Cefepime | 15 | 7 | 22 | 17 |
| APP-β | 14 | 6 | 45c | 13 | |
| None | 13 | 3 | 43a | 8 | |
| p = 0.610 | p = 0.34 | p = 0.004a | p = 0.07 | ||
| Timentin | Cefepime | 15 | 8 | 22 | 16 |
| APP-β | 14 | 6 | 45a | 13 | |
| None | 12 | 3 | 44a | 8 | |
| p = 0.550 | p = 0.20 | p = 0.002a | p = 0.10 | ||
| Gentamicin | Cefepime | 8 | 7 | 39 | 7 |
| APP-β | 6 | 8 | 62a | 2 | |
| None | 5 | 2 | 57a | 3 | |
| p = 0.456 | p = 0.16 | p = 0.014a | p = 0.06 | ||
aAPP-β, antipseudomonal penicillin/β-lactamase; none, no cefepime or APP-β
bNumber of patients
cSignificant difference (p < 0.05)
Our analysis showed that high-level homogeneity of β-lactam antibiotics within cycles was not associated with overall increased resistance, in agreement with other studies on antibiotic cycling in which Gram-negative bacterial susceptibility was not significantly altered [2, 6, 7]. The apparent ecological effects that we describe are consistent with our own data regarding MRSA and P. aeruginosa [5], challenging antimicrobial homogeneity as a driver of resistance per se [8], an idea that was premised on a mathematical model which was recently disputed [9]. Antibiotic use is recognized as the single most powerful selective pressure for the emergence of resistance particularly in environments where usage is high (ICU). However, the different strategies implemented to curb the rise of resistance in hospitals, including cycling, have had variable outcomes due to the complex relationship between use of specific drugs and resistance patterns in bacterial populations [10]. In our study, despite stable overall resistance rates, treatment with cefepime was a significant independent predictor of acquisition of antibiotic-resistant Gram-negative organisms and was also strongly associated with increased resistance to APP-β, but not to cefepime or extended-spectrum β-lactams (Table 1), in agreement with other studies on cefepime use in hospitalized patients [6, 11].
Our data strongly point to in vivo ecological effects of antibiotics rather than specific selection pressure associated with use of a specific antimicrobial class. However, ecological perturbation does not readily explain gentamicin and β-lactam resistance after cefepime, as these phenotypes are typically plasmid-encoded in the Enterobacteriaceae. We therefore directly compared E. coli populations from each of 12 patients before and after cefepime treatment (Additional file 1: Methods) and found no increase in virulence-associated types nor dominance of any single resistant clone (Table 2). Cultured isolates were of limited diversity, almost all of the B2 and D phylogenetic subtypes. There were three or less clearly distinguishable restriction types, and antibiotic resistance phenotypes gave no hint of underlying processes. However, a general effect on mobile genetic elements was suggested by the increased complexity and abundance of self-transmissible resistance plasmids and by enrichment for mobile resistance genes not relevant to cefepime (e.g., strAB, blaTEM, blaSHV; Fig. 2).
Table 2.
Antimicrobial resistance (AR) profiles of isolated E. coli representatives
| Patient | Isolate† | AR phenotype‡ | AR genotype§ |
|---|---|---|---|
| 1B | a B2 a D/E |
None None |
None |
| 1A | c B2 | None | None |
| 2B | d B2 e B2 |
None None |
None |
| 2A | e B2 d B2 |
None None |
None |
| 3B | f B2 f1 B2 h D/E |
None TET TET |
tet(B) aphA1 dfrA14 strA strB sul2 |
| 3A | f1 B2-D/E h D/E |
TET TET |
tet(B) |
| 4B | i B2 | AMP AMC CFZi TIMi | bla TEM sul2 |
| 4A | k B2 m D/E |
AMP CFZi CHLi AMP |
aadA blaSHV In |
| 5B | n B2 | AMP CFZi TIMi | aadA blaTEM In |
| 5A | n B2 | AMP CFZi TIMi | aadA blaTEM In |
| 6B | p B2-unk | AMP AMC TZP TIM CHL | aadA blaOXA-1 catA1 In |
| 6A | q B2 r B2 |
None AMP CFZi TIMi TET |
blaTEM tet(A) |
| 7B | s F | AMP CFZi TMP SXT | bla TEM dfrA14 sul2 strA strB |
| 7A | s1 D/E s2 F s2 D/E |
AMP AMCi CFZ TZP TIM TMP SXT AMP AMCi CFZ TZP TIM TOBi TMP SXT AMP AMCi CFZ TIM TMP SXT |
bla TEM dfrA14 strA strB sul2 |
| 8B | t B2 | AMP CFZi TIMi | aadA blaTEM In |
| 8A | t B2 | AMP AMCi CFZ TIM | aadA blaTEM In |
| 9B | u B2 v B2 |
AMP AMCi CFZi None |
bla TEM |
| 9A | u B2 | AMP AMCi CFZi TIMi | bla TEM |
| 10B | z B1 z1 B1 z2 B1 |
TET TET TET |
tet(B) |
| 10A | w B2 y unk |
None AMP TIMi TMP SXT |
bla TEM dfrA5 strA strB sul2 |
| 11B | w B2 | None | none |
| 11A | aa D/E | AMP AMC AZ CFZ FOX CAZ CRO LEX TIMi | blaCMY-2-like |
| 12B | bb B1 cc B1 |
AMP CFZi TIMi TMP SXT AMP TIMi CHLi TMP SXT |
blaTEM catA1 dfrA7 In |
| 12A | dd B2 ee B2 |
AMP AMCi CFZi TIM AMP AMCi CFZ TIMi |
bla TEM |
Underlined data not detected phenotypically by the BD PhoenixTMP system
B before antibiotic treatment (< 48 h ICU stay), A after antibiotic treatment (≥ 72 h ICU stay), i intermediate, In class 1 integron 5′- and/or 3′-conserved segments
†Defined by PFGE pattern (“a” to “ee”) and by phylogenetic grouping (A, B1, B2, D/E, F, unk (unknown) [19])
‡Not susceptible by BD PhoenixTMP screening of single E. coli colonies
§Genotype determined by NGS sequencing data analysis of pooled E. coli representatives for each patient, using BLAST comparisons [20] to the MARA database [17] and our in-house database of rep and mobilization genes (Additional file 1: Table S1)
Fig. 2.
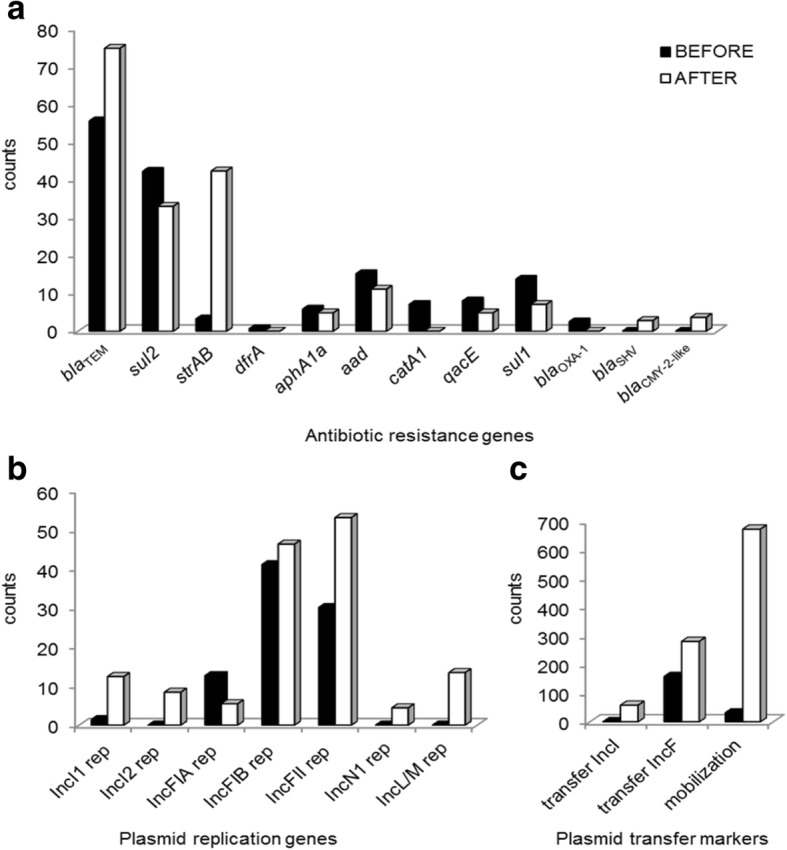
Antibiotic resistance markers detected before and after cefepime exposure in E. coli isolates from ICU patients. Resistance genes (a), plasmid replication genes (b), and plasmid transfer markers (c) were identified using the sequencing output from MiSeq sequencing (250 bp; paired-end) of pooled representatives (2–6 colonies per patient) of the E. coli population before (black) and after (white) cefepime treatment. Markers were detected by alignment with resistance genes and mobile genetic elements in the MARA database [17] and plasmids markers in our in-house database (Additional file 1 Methods). aad includes aadA1, aadA6, and aadA10; “transfer” comprises marker genes for conjugation/self-transmission (tra, nik, etc.); “mobilization” indicates relaxase genes of mobilizable plasmid types (Additional file 1: Table S1) [18]
In animal models, a proteobacterial bloom that accompanies colitis was associated with accelerated plasmid transfer between species [12], and a similar proteobacterial bloom is relatively prolonged after third-generation cephalosporins compared to penicillins [13, 14], providing a potential biological explanation for our findings (Fig. 2). Antibiotic treatment modifies the microbial community structure in the gut by shifting the competitive balance between sensitive bacteria and resistant/pathogenic subpopulations [15]. These subpopulations carry different resistant determinants that may come to predominate both by amplification of the original carriers and/or spread to other species. In Gram-negative enterobacteria, antibiotic resistance develops mainly via horizontal transfer of resistance genes that often cluster together in the same genetic locus, either on the chromosome or on plasmids, giving rise to multiple resistant types. Use of one antibiotic may drive selection of resistance to an entirely different class of drugs due to both cross-resistance mechanisms and co-localization of genetic elements. Perhaps more importantly resistance determinants are also associated with diverse mobile genetic elements (transposons, insertion sequences, plasmids) that allow for the movement of multidrug resistance loci between bacterial cells [15].
Even though selection and spread of specific resistance might be constrained by fitness requirements, antibiotic activity itself is known to promote horizontal gene transfer by triggering recombination and conjugation events, which will affect population-level resistance patterns [15], and by acceleration of gene transfer during population expansion events [13]. Together, these data indicate that cefepime exposure differentially drives antibiotic resistance in the microflora other than by direct phenotypic selection and are consistent with descriptions of enhanced plasmid transfer in other gut dysbioses [13]. This provides a potential explanation for resistance (e.g., to extended-spectrum β-lactam antibiotics) in Enterobacteriaceae that has been linked to exposure to late-generation cephalosporins, such as cefepime [14], and seems likely generalizable to third-generation cephalosporins, which have similar activities, gut penetration and associations with antibiotic resistance. It appears unlikely from (narrower-spectrum) first-generation cephalosporins, but reminds us that unmeasured impacts on the microbiome are key outcome determinants that have yet to be fully explored.
Additional file
Methods. This file describes the methods used to obtain and analyze the data presented in this manuscript and includes Table S1. (entitled “Markers for transmissible antibiotic resistance included in our in-house screening”) and additional references pertaining to the methodology. (DOCX 24 kb)
Acknowledgements
The authors acknowledge Agnieszka M Wiklendt for her technical support.
Funding
This work was supported by grants from the Australian National Health and Medical Research Council (NHMRC) to JI (1001020; 1046889).
Availability of data and materials
The datasets supporting the conclusions of this article are included within the article (and its additional files).
Authors’ contributions
CV designed and performed all analyses for the E. coli characterization study, participated in the analysis of clinical data, and wrote the manuscript. ANG performed the initial culture work for resistance data from the clinical specimens and analysis of clinical data and participated in clinical study design and manuscript preparation. BEW performed the analysis of clinical data. GT supported the bioinformatic screening of sequencing data for resistance, mobile element, and plasmid markers. IP participated in the study design. SRP participated in the study design and analysis of sequence data and created the plasmid marker database. JRI designed the study, supervised all analysis, and wrote the manuscript. All authors approved the final version of the manuscript.
Ethics approval and consent to participate
The previously published clinical component in the parent study (Ginn et al. 2012 [5]) was conducted under a waiver of consent, under the auspices of the relevant Human Research Ethics Committees of the Sydney West Area Health Service and the Royal Brisbane and Women’s Hospital.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Footnotes
Electronic supplementary material
The online version of this article (10.1186/s40635-018-0185-2) contains supplementary material, which is available to authorized users.
Contributor Information
Carola Venturini, Email: carola.venturini@sydney.edu.au.
Andrew N. Ginn, Email: andrew.ginn@sydney.edu.au
Brooke E. Wilson, Email: Brooke.Wilson3@health.nsw.gov.au
Guy Tsafnat, Email: guy.tsafnat@mq.edu.au.
Ian Paulsen, Email: ian.paulsen@mq.edu.au.
Sally R. Partridge, Email: sally.partridge@health.nsw.gov.au
Jonathan R. Iredell, Phone: 0061 2 86273411, Email: jonathan.iredell@sydney.edu.au
References
- 1.Lankelma JM, van Vught LA, Belzer C, Schultz MJ, van der Poll T, de Vos WM, Wiersinga WJ. Critically ill patients demonstrate large interpersonal variation in intestinal microbiota dysregulation: a pilot study. Intensive Care Med. 2016;43(1):59–68. doi: 10.1007/s00134-016-4613-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Macfarlane S. Antibiotic treatments and microbes in the gut. Environ Microbiol. 2014;16(4):919–924. doi: 10.1111/1462-2920.12399. [DOI] [PubMed] [Google Scholar]
- 3.Kalil AC. Is cefepime safe for clinical use? A Bayesian viewpoint. J Antimicrob Chemother. 2011;66(6):1207–1209. doi: 10.1093/jac/dkr138. [DOI] [PubMed] [Google Scholar]
- 4.Arizpe A, Reveles KR, Patel SD, Aitken SL. Updates in the management of cephalosporin-resistant Gram-negative bacteria. Curr Infect Dis Rep. 2016;18(12):39. doi: 10.1007/s11908-016-0552-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ginn AN, Wiklendt AM, Gidding HF, George N, O’Driscoll JS, Partridge SR, O’Toole BI, Perri RA, Faoagali J, Gallagher JE, Lipman J, Iredell JR. The ecology of antibiotic use in the ICU: homogeneous prescribing of cefepime but not Tazocin selects for antibiotic resistant infection. PLoS One. 2012;7(6):e38719. doi: 10.1371/journal.pone.0038719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Sarraf-Yazdi S, Sharpe M, Bennet KM, Dotson TL, Anderson DJ, Vaslef SN. A 9-year retrospective review of antibiotic cycling in a surgical intensive care unit. J Surg Res. 2012;176(2):e73–e78. doi: 10.1016/j.jss.2011.12.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Cadena J, Taboada CA, Burgess DS, Ma JZ, Lewis IIJS, Freytes CO, Patterson JE. Antibiotic cycling to decrease bacterial antibiotic resistance: a 5-year experience on a bone marrow transplant unit. Bone Marrow Transplant. 2007;40:151–155. doi: 10.1038/sj.bmt.1705704. [DOI] [PubMed] [Google Scholar]
- 8.Bergstrom CT, Lo M, Lipsitch M. Ecological theory suggests that antimicrobial cycling will not reduce antimicrobial resistance in hospitals. PNAS. 2004;101:13285–13290. doi: 10.1073/pnas.0402298101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Beardmore RE, Peña-Miller R, Gori F, Iredell J. Antibiotic cycling and antibiotic mixing: which one best mitigates antibiotic resistance? Mol Biol Evol. 2017;34(4):802–817. doi: 10.1093/molbev/msw292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Lipsitch M, Samore MH. Antimicrobial use and antimicrobial resistance: a population perspective. Emerg Infect Dis. 2002;8(4):347–354. doi: 10.3201/eid0804.010312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.De Araujo OR, Cardoso Bourguignon da Silva D, Diegues AR, Arkader R, Aparecida E, Cabral F, Rodriguez Afonso M, Louzada ME, de Cássia A, Albertoni S Cefepime restriction improves gram-negative overall resistance patterns in neonatal intensive care unit. Braz J Infect Dis 11(2):277–280 [DOI] [PubMed]
- 12.Stecher B, Denzler R, Maier L, Bernet F, Sanders MJ, Pickard DJ, Barthel M, Westendorf AM, Krogfelt KA, Walker AW, Ackermann M, Dobrindt U, Thomson NR, Hardt W-D. Gut inflammation can boost horizontal gene transfer between pathogenic and commensal Enterobacteriaceae. Proc Natl Acad Sci U S A. 2012;109(4):1269–1274. doi: 10.1073/pnas.1113246109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Antonopoulos DA, Huse SM, Morrison HG, Schmidt TM, Sogin ML, Young VB. Reproducible community dynamics of the gastrointestinal microbiota following antibiotic perturbation. Infect Immun. 2009;77:2367–2375. doi: 10.1128/IAI.01520-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Meletiadis J, Turlej-Rogacka A, Lerner A, Adler A, Taconelli E, Mouton JW, the SATURN Diagnostic Study Group Amplification of antimicrobial resistance in gut flora of patients treated with ceftriaxone. Antimicrob Agents Chemother. 2017;61(11):e00473–e00417. doi: 10.1128/AAC.00473-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Modi SR, Collins J, Relman DA. Antibiotics and the gut microbiota. J Clin Invest. 2014;124(10):4212–4218. doi: 10.1172/JCI72333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kok J, Thomas LC, Olma T, Chen SCA, Iredell JR. Identification of bacteria in blood culture broths using matrix-assisted laser desorption-ionization Sepsityper™ and time of flight mass spectrometry. PLoS One. 2011;6(8):e23285. doi: 10.1371/journal.pone.0023285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Partridge SR, Tsafnat G. Automated annotation of mobile antibiotic resistance in Gram-negative bacteria: the Multiple Antibiotic Resistance Annotator (MARA) and database. J Antimicrob Chemother. 2018;73(4):883–890. doi: 10.1093/jac/dkx513. [DOI] [PubMed] [Google Scholar]
- 18.Garcillán-Barcia MP, Alvarado A, de la Cruz F. Identification of bacterial plasmids based on mobility and plasmid population biology. FEMS Microbiol Rev. 2011;35(5):936–956. doi: 10.1111/j.1574-6976.2011.00291.x. [DOI] [PubMed] [Google Scholar]
- 19.Clermont O, Christenson JK, Denamur E, Gordon DM. The Clermont Escherichia coli phylo-typing method revisited: improvement of specificity and detection of new phylo-groups. Environ Microbiol Rep. 2013;5(1):58–65. doi: 10.1111/1758-2229.12019. [DOI] [PubMed] [Google Scholar]
- 20.Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Methods. This file describes the methods used to obtain and analyze the data presented in this manuscript and includes Table S1. (entitled “Markers for transmissible antibiotic resistance included in our in-house screening”) and additional references pertaining to the methodology. (DOCX 24 kb)
Data Availability Statement
The datasets supporting the conclusions of this article are included within the article (and its additional files).


