Abstract
The characterization of RNA polymerase subunit genes has revealed that some subunits are shared by the three nuclear enzymes, some are homologous, and some are unique to RNA polymerases I, II, or III. We report here the isolation and characterization of the yeast RNA polymerase II subunit RPB11, which is encoded by a single copy RPB11 gene located directly upstream of the topoisomerase I gene, TOPI, on chromosome XV. The sequence of the gene predicts an RPB11 subunit of 120 amino acids (13,600 daltons), only two amino acids shorter than the RPB9 polypeptide, that co-migrates with RPB11 under most SDS-PAGE conditions. RPB11 was found to be an essential gene that encodes a protein closely related to an essential subunit shared by RNA polymerases I and III, AC19. RPB11 contains a 19 amino acid segment found in three other yeast RNA polymerase subunits and the bacterial RNA polymerase subunit α. Some mutations that affect RNA polymerase assembly map within this segment, suggesting that this region may play a role in subunit interactions. As the isolation of RPB11 completes the isolation of known yeast RNA polymerase II subunit genes, we briefly summarize the salient features of these twelve genes and the polypeptides that they encode.
RNA polymerases I, II, and III are at the core of an increasingly complex set of proteins that have roles in transcription and in its regulation. These multisubunit enzymes are highly conserved among eukaryotes (Young, 1991; Woychik and Young, 1990b; Sentenac, 1985). The Saccharomyces cerevisiae nuclear RNA polymerases are among the best studied eukaryotic RNA polymerases. Each of these enzymes comprises 10–13 polypeptides (Sentenac, 1985). To understand better the structure and function of eukaryotic RNA polymerases, the genes that encode S. cerevisiae RNA polymerase subunits are being isolated and used to examine the roles of the subunits in transcription.
The regulatory complexities of mRNA synthesis have led us to focus our attention on the structure and function of yeast RNA polymerase II. Thus far, the genes for the yeast RNA polymerase II subunit RPB1 (Young and Davis, 1983; Ingles et al., 1984; Allison et al., 1985); RPB2 (Sweetser et al., 1987); RPB3 (Kolodziej and Young, 1989); RPB4 (Woychik and Young, 1989); RPB5, RPB6, and RPB8 (Woychik et al., 1990); RPB7 (McKune et al., 1993); RPB9 (Woychik et al., 1991); RPB10 (Woychik and Young, 1990a); and RPB12 (also known as ABC10α; Carles et al., 1991; Treich et al., 1992) have been isolated and characterized. Structural and functional studies of yeast RNA polymerase II genes and their products have led to an improved picture of the enzyme. The three largest subunits — RPB1, RPB2, and RPB3 — are related to the prokaryotic core RNA polymerase subunits, are essential, and are probably largely responsible for RNA catalysis (Young, 1991). Five additional subunits — RPB5, RPB6, RPB8, RPB10, and RPB12 — are essential components shared by all three nuclear RNA polymerases (Woychik et al., 1990; Carles et al., 1991; Treich et al., 1992).
Upon SDS-polyacrylamide gel electrophoresis, purified yeast RNA polymerase II usually separates into 10 apparent polypeptides whose apparent molecular sizes are 220, 150, 44.5, 32, 27, 23, 16, 14.5, 12.6, and 10 kilodaltons (kDa; Sentenac, 1985). However, the enzyme actually comprises twelve subunits. The 10 kDa band consists of two proteins, the RPB10 and RPB12 (ABC10α) subunits (Carles et al., 1991; Treich et al., 1992). In addition, the 12.6 kDa band contains two polypeptides, the RPB9 subunit and an as yet uncharacterized subunit designated RPB11. This paper describes the isolation and characterization of this remaining RNA polymerase II subunit gene, RPB11.
Materials and methods
Yeast strains
Yeast strains used in this study were Z321 (MATa/MATα, ura3-52/ura3-52, his3Δ200/his3Δ200, leu2-3/leu2-3, leu2-112/leu2-112, lys2Δ201/lys2Δ201, ade2/ade2); WY-11 (MATa/MATα, ura3-52/ura3-52, his3Δ200/his3Δ200, leu2-3/leu2-3, leu2-112/leu2-112, lys2Δ201/lys2Δ201, ade2/ade2, RPB11/RPB11Δ1:: HIS3); and X2180-2, the diploid derivative of S288C (MATa mal gal2).
Protein sequence analysis
S. cerevisiae RNA polymerase II was purified from strain X2180-2 (Valenzuela et al., 1978), and the individual subunits were separated by SDS-PAGE. The RPB9/RPB11 doublet was electroblotted onto nitrocellulose and digested with trypsin, as described by Aebersold et al. (1987), omitting the NaOH wash to minimize any passive elution of protein. The resultant peptide mixture was chromatographed on a Hewlett-Packard 1090 HPLC column using a Vydac 2.1 mm × 150 mm C18 column. The gradient employed was essentially that previously described by Stone et al. (1989). The resulting samples were subjected to automated Edman degradation.
Plasmids
The plasmid pRSa3-9 (kindly provided by Rolf Sternglanz, State University of New York at Stony Brook) has a >6.5 kb RPB11- and TOP1- containing insert in YEp13; pRP11/1 has a Pst I fragment containing the RPB11 open reading frame removed and replaced by a BamH I site (the BamH I site is flanked by ∼600 bps of 5′ and ∼900 bps 3′ DNA with Pst I ends created by PCR) ligated to the Pst I site of pGem5Zf(+) (Promega); pRP11/2 is pRP11/1 cut with BamH I and ligated to a 1.75 kb HIS3 containing BamH I fragment.
Gene copy number: Southern analysis
Hybridization and wash conditions were as described previously (Woychik et al., 1990). The radiolabeled probe used for copy number analysis of RPB11 was made with a 270 bp Hind III/Nco I fragment containing the C-terminal end of the RPB11 coding sequence plus 200 bps of 3′ DNA.
Construction of the RPB11 deletion
The plasmid (pRP11/1) used to create RPB11Δ1::HIS3 yeast cells was constructed by first removing the RPB11 open reading frame by PCR of linearized pRSa3-9 DNA following the method of Ho et al. (1989) and replacing the RPB11 coding sequence with a unique BamH I restriction site. RPB11/RPB11Δ1::HIS3 yeast cells were selected on His− plates following the transformation of the yeast diploid Z321 with pRP11/2 cut with BstE II and Sac I. Genomic DNA was prepared from the His+ transformant WY-11 and subjected to Southern analysis to verify the substitution of the chromosomal copy of RPB11 with HIS3. WY-11 cells were then sporulated and subjected to tetrad analysis.
Results
Identification of a new RNA polymerase II subunit, RPB11
RNA polymerase II was purified by conventional methods (Valenzuela et al., 1978) and run on a 12.5% low-crosslinker SDS-polyacrylamide gel (Dreyfuss et al., 1984). Upon staining with Coomassie brilliant blue, we consistently resolved two distinct bands with estimated molecular weights equivalent to those predicted for the RPB9 subunit. The presence of this extra band was also noted by Edwards et al. (1990, 1991) after Coomassie brilliant blue staining of affinity purified S. cerevisiae RNA polymerase II run on 15% SDS-polyacrylamide gels. Peptide sequences were obtained from a trypsin-digested mixture of the RPB11/RPB9 proteins. The first peptide sequence obtained was used to isolate the RPB9 gene (Woychik et al., 1991). The peptide sequences obtained thereafter were compared to the predicted sequence of the RPB9 protein, and those not contained in RPB9 were used to isolate RPB11 gene.
Isolation of the RPB11 gene
The majority of RNA polymerase subunit genes were isolated by using oligonucleotides whose sequences were derived from the amino acid sequences of portions of the purified subunits. However, TFASTA database searches using the four RPB11-derived peptides revealed that one peptide, TNFETEWNLQTLAA, exactly matched a region within the predicted amino acid sequence of a gene directly upstream of the S. cerevisiae gene encoding topoisomerase I (TOP1; Thrash et al., 1985; Goto and Wang, 1985; Fig. 1A). A plasmid containing several kilobases of DNA upstream of the TOP1 gene was used to obtain the complete sequence of the RPB11 gene. The RPB11 gene encodes a protein containing all four RPB11 peptides obtained by microsequence analysis of purified protein (Fig. 1B). The RPB11 sequence predicts a protein 120 amino acids long with a molecular mass of 13,600 Da, which is close to its 13 kDa apparent molecular weight on SDS-PAGE (Sentenac, 1985).
Figure 1.

The RPB11 gene. A. RPB11 is adjacent to the TOP1 gene. Arrows represent gene orientation; lengths of shaded and solid rectangles reflect relative gene sizes. B. Sequence of RPB11 DNA and the predicted amino acid sequence of the RPB11 subunit. The RPB11 peptides APNAVVITFEK, IQTTEGYDPK, LGALK, and TNFETEWNLQTLAA are underlined.
Copy number and chromosomal location of RPB11
Southern blots containing immobilized restriction digests of S. cerevisiae genomic DNA were probed with RPB11 DNA fragments at moderate stringency to determine the copy number of these genes. The pattern of hybridization, in which only a single band producing a strong signal was observed, indicated that RPB11 is a single copy gene in haploid yeast (Fig. 2). The pattern of hybridization obtained with the RPB11 DNA probes did not change over a range of hybridization and wash conditions and was consistent with that obtained with the oligonucleotide probes used for gene isolation and characterization.
Figure 2.
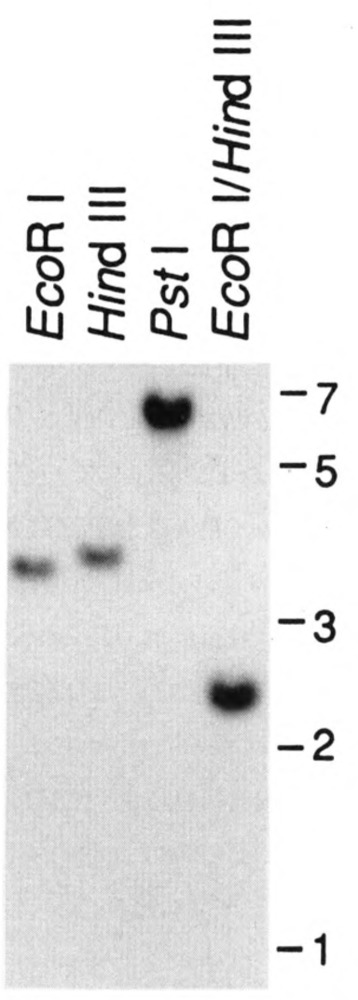
Copy number of the RPB11 gene. Genomic DNA prepared from haploid yeast cells was digested with the enzyme(s) indicated. DNA size markers (in kb) are shown to the right of each panel.
The gene encoding RPB11 is directly adjacent to the TOP1 gene, which has been genetically mapped to the right arm of chromosome XV, close to (<10 cM) the centromere (Thrash et al., 1985). Four of the twelve RNA polymerase II subunit genes — RPB2, RPB8, RPB10, and RPB11 — map to chromosome XV, while the other RNA polymerase II subunit genes are well dispersed among the chromosomes.
Cells lacking RPB11 are inviable
To determine whether the RPB11 subunit is required for cell viability, the entire RPB11 coding sequence was replaced with the nutritional marker HIS3 (RPB11Δ1::HIS3). One chromosomal copy of the RPB11 gene in yeast diploid cells was then replaced with RPB11Δ1::HIS3 by homologous recombination of RPB11-flanking DNA with the chromosomal DNA (Rothstein, 1983). The diploid cells obtained by this approach had one chromosome with a wild-type RNA polymerase subunit gene and one chromosome with a deletion allele. Tetrad analysis of the sporulation products of these diploid cells revealed that RPB11 is an essential gene, since there were two viable and two inviable spores per tetrad (data not shown). None of the spores containing the deletion allele produced viable cells (i.e., all viable spores were His−).
The deletion results are consistent with those of Goto and Wang (1985), that insertion of a selectable marker at the Hind III site upstream of TOPI suggested the existence of a single-copy, essential gene in this region. Disruption of the RPB11 gene at the Hind III site allows for the synthesis of a truncated RPB11 polypeptide lacking 20% of the carboxyl terminal residues, suggesting that expression of more than 80% of the RPB11 protein is required for normal cell growth and viability.
RPB11 is related to a subunit common to S. cerevisiae RNA polymerase I and III
Computer searches revealed that the RPB11 amino acid sequence is 30% identical and an additional 24% of the residues are related (scoring +1 or greater on the Dayhoff mutation data matrix [Dayhoff et al., 1983]) to S. cerevisiae RNA polymerase subunit AC19 (Dequard-Chablat et al., 1991; Fig. 3). AC19 is one of two subunits common only to RNA polymerases I and III. The other subunit common to yeast RNA polymerase I and III, AC40, is related to the RNA polymerase II subunit RPB3. RPB3 and AC40 are 31% identical, and an additional 20% of the residues are conserved substitutions (Kolodziej and Young, 1989). The same level of sequence similarity exists between RPB3 and AC40 as does that between RPB11 and AC19 (50% of the residues are identical or conserved). These results suggest that the AC40 and AC19 subunits are functionally related to RPB3 and RPB11.
Figure 3.

S. cerevisiae RPB11 is related to yeast RNA polymerase I and III subunit AC19 and contains a conserved 19 amino acid region present in the bacterial a subunit and other eukaryotic proteins related to α. A. Identical amino acid residues are boxed; conserved residues are in bold. B. RPB11 and RPB3, yeast RNA polymerase II subunits; AC19 and AC40, shared subunits of yeast RNA polymerase I and III; α, E. coli RNA polymerase subunit; RNLVA, α homologue in Marchantia polymorpha chloroplast DNA (Dequard-Chablat et al., 1991); cnjC, Tetrahymena conjugation-specific protein related to α, RPB3, and AC40 (Martindale, 1990). The shaded regions represent identical and conserved residues scoring +1 or greater on the Dayhoff mutation data matrix. The residues which contribute to a defect in RNA polymerase subunit assembly are marked with an asterisk. The RPB3 mutation A30 → D30 is represented in the mutant rpb3-1 (Kolodziej and Young, 1989). The α mutation R45 → C45 is represented in the mutant rpoA112 (Igarashi et al., 1990). Some of the sequence similarities were demonstrated in previous reports (Dequard-Chablat et al., 1991; Martindale, 1990).
Interestingly, RPB3, AC40, AC19, and RPB11 all contain a highly conserved region 19 amino acids in length that is present in the bacterial RNA polymerase II α subunit. This region is also represented in the human RPB3 counterpart (Pati and Weissman, 1990), the α homologue in chloroplasts (Dequard-Chablat et al., 1991), and a gene related to α (cnjC) in Tetrahymena (Martindale, 1990). Database searches using the peptide representing this region in RPB11 did not reveal other proteins with known functions that also contain this conserved region.
Discussion
The isolation and characterization of RPB11 completes the isolation of the genes that encode the twelve subunits of yeast RNA polymerase II. Sequence analysis, mutagenesis, and epitope-tagging experiments made possible by the cloning of these genes have provided a clearer picture of RNA polymerase II subunit composition, stoichiometry, and function (Table 1). The three largest subunits, RPB1, RPB2, and RPB3, have counterparts in the bacterial core enzyme. However, yeast RNA polymerase II does not appear to have a subunit that is closely related to the bacterial o subunit family. Deletion analysis revealed that ten of the twelve subunit genes are essential for viability. The conditional growth phenotypes that are observed for RPB4 and RPB9 deletion mutants suggest that these subunits have a role enzyme stability or in fine tuning the efficiency of the transcription apparatus.
Table 1.
Yeast RNA polymerase II subunit genes.
| Gene | SDS-PACE mobility (kDa) | Protein mass (kDa) | Stoichiometry | Chromosomal location | Deletion phenotype | Sequence similarity |
|---|---|---|---|---|---|---|
| RPB1 a | 220 | 190 | 1.1 | IV | Inviable | β′c |
| RPB2 | 150 | 140 | 1.0 | XV | Inviable | βd |
| RPB3 | 45 | 35 | 2.1 | IX | Inviable | Human hRPB44e, AC40 f, αf |
| RPB4 | 32 | 25 | 0.5 | X | Conditional | Acidic region of σ70 g |
| RPB5 | 27 | 25 | 2.0 | II | Inviable | Human hRPB33h, rpoHi |
| RPB6 | 23 | 18 | 0.9 | XVI | Inviable | |
| RPB7 | 17 | 19 | 0.5 | IV or XII | Inviable | |
| RPB8 | 14 | 17 | 0.8 | XV | Inviable | |
| RPB9 | 13 | 14 | 2.0 | VII | Conditional | Drosophila 14 kDa proteinj |
| RPB10 | 10 | 8.3l | 0.9 | XV | Inviable | |
| RPB11 | 13 | 14 | – | XV | Inviable | AC19k |
| RPB12 b | 10 | 7.7 | – | VIII | Inviable |
a Also known as RP021, RPB220. b Also known as RPC10. c Allison et al., 1985. d Sweetser et al., 1987. e Pati and Weissman, 1990. f Kolodziej and Young, 1989. g Woychik and Young, 1989. h Pati and Weissman, 1989. i Klenk et al., 1992. j Woychik and Young, 1991. k Dequard-Chablat et al., 1991. l The published DNA sequence contained an extra A at position 95 of the open reading frame. The corrected RPB10 sequence extends the open reading frame to encode a protein 70 amino acids in length.
The common functions of the three RNA polymerases are reflected in features that are shared by these enzymes. The three largest subunits of RNA polymerase II are related in size and sequence to the three largest subunits of RNA polymerases I and III (Young, 1991; Kolodziej and Young, 1989). In addition, five identical subunits (RPB5, RPB6, RPB8, RPB10, and RPB12) are found in all three nuclear RNA polymerases (Woychik et al., 1990; Carles et al., 1991; Triech et al., 1992). We have found a new sequence relationship among RNA polymerase subunits, as RPB11 is very similar to a subunit shared by RNA polymerases I and III, AC19. Thus, at least nine of the twelve known RNA polymerase II subunits are related or identical to the subunits of RNA polymerases I and III. These results suggest that the three eukaryotic nuclear RNA polymerases have significant structural and functional relationships.
There is a conserved 19 amino acid region present in RPB3, RPB11, AC40, AC19, and the bacterial RNA polymerase subunit α. The E. coli α subunit plays an important role in subunit assembly, since its dimerization is the first step in the sequential assembly of subunits to form the bacterial holoenzyme α → α2 → α2β → α2ββ′ (Yura and Ishihama, 1979; Ishihama, 1981; Chamberlin, 1982). A temperature-sensitive mutation that falls within the conserved 19 amino acid segment of the E. coli α subunit blocks subunit assembly (Igarashi et al., 1990; Kawakami and Ishihama, 1980). In addition, one of two RPB3 mutations that produce an assembly defect also maps within this conserved region (Kolodziej and Young, 1989). These results suggest that this 19 amino acid region may be involved in RNA polymerase subunit interactions. Future genetic and mutational studies will likely provide more precise clues to the functional role of this conserved region.
Acknowledgments
We thank Rolf Sternglanz for providing the RPB11-containing plasmids and Parag Sadhale for comments on the manuscript.
The majority of this work was supported by Public Health Service grant GM-34365 and by Public Health Service postdoctoral fellowship GM-11605 to N. A. Woychik from the National Institutes of Health. R. A. Young is Burroughs Wellcome Sholar.
The costs of publishing this article were defrayed in part by the payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 USC Section 1734 solely to indicate this fact.
References
- Aebersold R. H., Leavitt J., Saavedra R. A., and Hood L. E. (1987), Proc Natl Acad Sci USA 84, 6970–6974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Allison L. A., Moyle M., Shales M., and Ingles C. J. (1985), Cell 42, 599–610. [DOI] [PubMed] [Google Scholar]
- Carles C., Treich I., Bouet F., Riva M., and Sentenac A. (1991), J Biol Chem 266, 24092–24096. [PubMed] [Google Scholar]
- Chamberlin M. J. (1982), in The Enzymes (Boyer P., ed.), Academic Press, New York, pp. 61–86. [Google Scholar]
- Dayhoff M. O., Barker W. C., and Hunt L. T. (1983), Methods Enzymol 91, 524–545. [DOI] [PubMed] [Google Scholar]
- Dequard-Chablat M., Riva M., Carles C., and Sentenac A. (1991), J Biol Chem 266, 15300–15307. [PubMed] [Google Scholar]
- Dreyfuss G., Adam S. A., and Choi Y. D. (1984), Mol Cell Biol 4, 415–423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edwards A. M., Darst S. A., Feaver W. J., Thompson N. E., Burgess R. R., and Kornberg R. D. (1990), Proc Natl Acad Sci USA 87, 2122–2126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edwards A. M., Kane C. M., Young R. A., and Kornberg R. D. (1991), J Biol Chem 266, 71–75. [PubMed] [Google Scholar]
- Goto T. and Wang J. C. (1985), Proc Natl Acad Sci USA 82, 7178–7182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ho S. N., Hunt H. D., Horton R. M., Pullen J. K., and Pease L. R. (1989), Gene 77, 51–59. [DOI] [PubMed] [Google Scholar]
- Igarashi K., Fujita N., and Ishihama A. (1990), Nucleic Acids Res 18, 5945–5948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ingles C. J., Himmelfarb H. J., Shales M., Greenleaf A. L., and Friesen J. D. (1984), Proc Natl Acad Sci USA 81, 2157–2161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishihama A. (1981), Adv Biophys 14, 1–35. [PubMed] [Google Scholar]
- Klenk H., Palm P., Lottspeich F., and Zillig W. (1992), Proc Natl Acad Sci USA 89, 407–410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawakami K. and Ishihama A. (1980), Biochemistry 19, 3491–3495. [DOI] [PubMed] [Google Scholar]
- Kolodziej P. A. and Young R. A. (1989), Mol Cell Biol 9, 5387–5394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martindale D. W. (1990), Nucleic Acids Res 18, 2953–2960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKune K., Richards K. L., Edwards A. M., Young R. A., and Woychik N. A. (1993), Yeast, in press. [DOI] [PubMed]
- Pati U. K. and Weissman S. M. (1989), J Biol Chem 264, 13114–13121. [PubMed] [Google Scholar]
- Pati U. K. and Weissman S. M. (1990), J Biol Chem 265, 8400–8403. [PubMed] [Google Scholar]
- Rothstein R. J. (1983), Methods Enzymol 101, 202–211. [DOI] [PubMed] [Google Scholar]
- Sentenac A. (1985), CRC Crit Rev Biochem 18, 31–90. [DOI] [PubMed] [Google Scholar]
- Stone K. L., LoPresti M. B., Williams N. D., Crawford J. M., DeAngelis R., and Williams K. R. (1989), in Techniques in Protein Chemistry (Hugli T., ed.), Academic Press, San Diego, pp. 377–391. [Google Scholar]
- Sweetser D., Nonet M., and Young R. A. (1987), Proc Natl Acad Sci USA 84, 1192–1196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thrash C., Bankier A. T., Barrell B. G., and Sternglanz R. (1985), Proc Natl Acad Sci USA 82, 4374–4378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Treich I., Carres C., Riva M., and Sentenac A. (1992), Gene Expr 2, 31–37. [PMC free article] [PubMed] [Google Scholar]
- Valenzuela P., Bell G. I., Weinburg F., and Rutter W. J. (1978), Methods Cell Biol 19, 1–26. [DOI] [PubMed] [Google Scholar]
- Woychik N. A., Lane W. S., and Young R. A. (1991), J Biol Chem 266, 19053–19055. [PubMed] [Google Scholar]
- Woychik N. A., Liao S.-M., Kolodziej P. A., and Young R. A. (1990), Genes Dev 4, 313–323. [DOI] [PubMed] [Google Scholar]
- Woychik N. A. and Young R. A. (1989), Mol Cell Biol 9, 2854–2859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woychik N. A. and Young R. A. (1990a), J Biol Chem 265, 17816–17819. [PubMed] [Google Scholar]
- Woychik N. A. and Young R. A. (1990b), Trends Biochem Sci 15, 347–351. [DOI] [PubMed] [Google Scholar]
- Young R. A. (1991), Annu Rev Biochem 60, 689–715. [DOI] [PubMed] [Google Scholar]
- Young R. A. and Davis R. W. (1983), Science 222, 778–782. [DOI] [PubMed] [Google Scholar]
- Yura T. and Ishihama A. (1979) Annu Rev Genet 13, 59–97. [DOI] [PubMed] [Google Scholar]


