Abstract
Genome-scale CRISPR-Cas9 Knockout Screening was applied to investigate novel targets in imatinib-resistant gastrointestinal stromal tumor (GIST). 20 genes and 2 miRNAs have been selected by total reads of sgRNA and sgRNA diversity, which has been further validated in imatinib-resistant GIST cells by CCK8 and qPCR analysis. Our study has finally revealed 9 genes (DBP, NR3C1, TCF12, TP53, ZNF12, SOCS6, ZFP36, ACYP1, and DRD1) involved in imatinib-resistant GIST-T1 cells. TP53 and SOCS6 may be the most promising candidate genes for imatinib-resistance due to the possible signaling pathway, such as apoptosis pathway and Wnt signaling pathway, JAK-STAT signaling pathway. It is necessary to perform more studies to discover novel targets in imatinib-resistant GIST, including DBP, NR3C1, TCF12, ZNF12, ZFP36, ACYP1 and DRD1.
Electronic supplementary material
The online version of this article (10.1186/s12943-018-0865-2) contains supplementary material, which is available to authorized users.
Keywords: Gastrointestinal stromal tumor, Imatinib resistance, Genome-scale CRISPR-Cas9 knockout screening
Gastrointestinal Stromal Tumor (GIST) is the most frequent mesenchymal tumor in the gastrointestinal tract [1]. The activating mutations in PDGFRA or KIT are observed in GIST, which are the key molecular drivers in tumor pathogenesis [2]. Imatinib mesylate, also known as Glivec®, is used as tyrosine kinase inhibitor for standard targeted therapy in GIST. However, secondary resistance to imatinib with disease progression is observed in about half of patients in 2 years of therapy. The mechanisms of imatinib-resistance in GIST have been validated in some extent, such as PI3K/AKT/mTOR pathway [3]. Due to the complexity of imatinib-resistant mechanisms, it is necessary to discover novel targets to imatinib-resistant in GIST. The RNA-guided CRISPR-associated nuclease Cas9 is an effective method to introduce targeted loss-of-function mutations at the specific sites in genome with low noise, consistent activity across reagents and minimal off-target effects [4]. This system has been previously reported to identify drug resistant genes with high efficiency in vitro [5]. Here, we sought to identify novel genes which are critically important to imatinib-resistance in GIST by genome-scale CRISPR-Cas9 knockout screening.
Genome-scale CRISPR-Cas9 knockout screening for imatinib resistance
To determine the minimum lethal dose (MLD) of imatinib in human GIST-derived cell line GIST-T1 cells, different concentrations of imatinib were added into GIST-T1 cells (results shown in Additional file 1: Figure S1). As shown in Fig. 1a, the cell number in control group was far more than that in imatinib groups (40 μg/mL) at day 4. 40 μg/mL was considered as the MLD of imatinib in GIST-T1 cells, which would be used in the following experiments.
Fig. 1.
Schematic and results of functional screening by sgRNA library and imatinib treatment. a Optical microscopic images of GIST-T1 cells treated with imatinib at day 4. b Structure of one vector lentiviralGeCKO system; c Schematic of imatinib-resistant GIST-T1 cells construction for high-throughput sequencing analysis; d Optical microscopic images of GIST-T1cells transfected with lentiviral sgRNA library and treated with imatinib. a-control, b-imatinib (40 μg/mL), c-transfection and imatinib (40 μg/mL)
To identify genes related to imatinib drug resistance in GIST, genome-scale CRISPR-Cas9 knockout screening was performed in the GIST-T1 cells. We applied a human genome-scale CRISPR knockout library A (hGeCKOa) and B (hGeCKOb) to transfect GIST-T1 cells separately. hGeCKOa and hGeCKOb contain 65,383 and 58,028 sgRNAs targeting 19,050 genes, respectively (see Additional file 2: Table S1 and Additional file 3: Table S2). A single vector lentiviralGeCKO system (lentiCRISPRv2, Fig. 1b) was used to deliver sgRNA, Cas9 and puromycin selection marker into GIST-T1 cells. Figure 1c displayed a schematic of imatinib-resistant GIST-T1 cells construction for high-throughput sequencing analysis. After imatinib treatment, the genomic DNA of surviving cells (imatinib-resistant GLIST-T1 cells) was extracted for PCR amplification of sgRNA-coding region, followed by high-throughput sequencing analysis. As shown in Fig. 1d, most of GIST-T1cells transfected with lentiviral sgRNA library were survived at day 6 after imatinib treatment, indicating that imatinib-resistant GIST-T1 cells were obtained. The successful transfection of lentiCRISPRv2 vector in GIST-T1 cells was confirmed by electrophoresis (results shown in Additional file 4: Figure S2A). After exposure to imatinib, a small group of GIST-T1 cells transfected with hGeCKOa or hGeCKOb was rendered drug resistance to imatinib, which was cultured and collected to extract DNA for PCR amplification (as shown in Additional file 1: Figure S2B-C).
Enriched sgRNAs in GIST-T1 cells with imatinib resistance
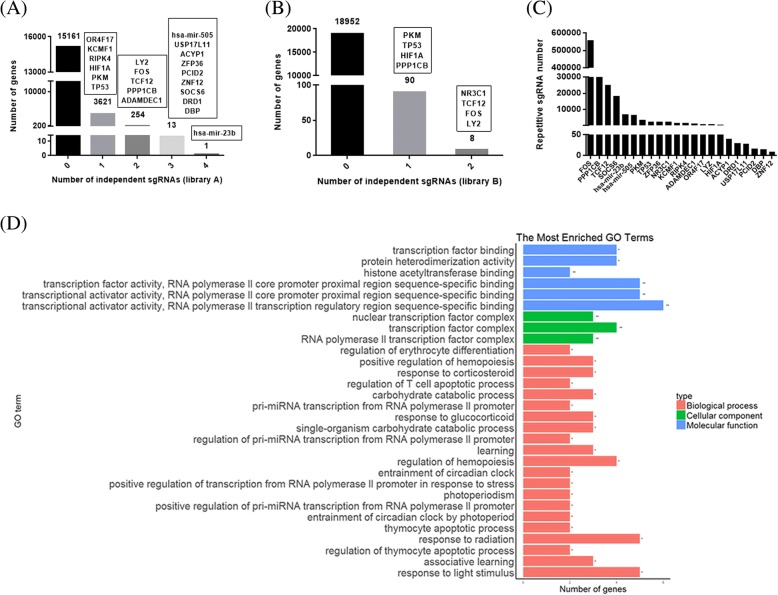
The results of high-throuoghput sequencing analysis have been shown in Additional file 5: Table S3. For a subset of genes/miRNA, we found enrichment of sgRNAs targeting each gene after imatinib treatment (Fig. 2a, b), suggesting that loss of these particular genes contributes to imatinib resistance. For further study, we have chosen 20 genes and 2 miRNAs by the total reads of sgRNAs and sgRNA diversity, as listed in Additional file 6: Table S4. The total reads of sgRNAs for selective genes/miRNAs were also shown in Fig. 2c. Gene ontology (GO) analysis was performed for the selected 20 genes to analyze their roles in biological process, molecular function and cellular component. Top 30 pathways from GO analysis of the 20 genes were shown in Fig. 2d. Detailed information of GO analysis for these genes was listed in Additional file 7: Table S5.
Fig. 2.
GeCKO screening in GIST-T1 cells reveals genes and miRNAs whose loss confers imatinib resistance. a & b Number of genes/miRNA with 0, 1, 2, 3 or 4 significantly enriched hGeCKOa (a) and hGeCKOb (b) sgRNAs targeting that genes/miRNA. Genes/miRNAs were selected by total reads of sgRNA and sgRNA diversity, as shown in black boxes. c Total reads of sgRNA for selected genes/miRNAs. d Gene ontology of candidate genes
GeCKO screening results of imatinib in GIST cells
To validate the function of candidate genes/miRNA identified from GeCKO screen, we generated individual gene or miRNA knockouts in GIST-T1 cells. We tested their susceptibility to imatinib in GIST-T1 cells by using CCK8 assays. As shown in Fig. 3a, there were 12 genes and 2 miRNAs being resistant to imatinib, especially SOCS6 and ZFP36. The majority of these resistant genes/miRNAs were targeted by two or more independent sgRNAs, indicating the tight relative between sgRNA diversity and resistant genes/miRNA. Optical microscopic images of GIST-T1cells with individual gene/miRNA knockouts have been shown in Additional file 8: Figure S3.
Fig. 3.
GeCKO screening results of imatinib in GIST cells. a CCK8 assay for GIST-T1 cells with individual gene or miRNA knockouts; b qPCR analysis for 12 genes and 2 miRNAs in GIST-T1 cells (imatinib-sensitive), imatinib-resistant GIST-48 and imatinib-resistant GIST-430 cells
Furthermore, qPCR was applied to determine the expression levels of 12 genes and 2 miRNAs in GIST-T1 cells (imatinib-sensitive), imatinib-resistant GIST-48 and imatinib-resistant GIST-430 cells with the aim of confirming their functions. As shown in Fig. 3(b), the expression levels of 9 genes were significantly lower in imatinib-resistant GIST-48 and imatinib-resistant GIST-430 cells than that in GIST-T1 cells, including SOCS6, ZFP36, DBP, NR3C1, TCF12, TP53, ZNF12, ACYP1, and DRD1. These 9 genes (DBP, NR3C1, TCF12, TP53, ZNF12, SOCS6, ZFP36, ACYP1, and DRD1) might be the potential genes for imatinib-resistance, while HIF1A, LYZ, mir-23b-3p and mir-505-3p showed uncertain relationship with imatinib-resistance.
It is worth mentioning that these resistant genes included previously reported genes of TP53 in imatinib resistance [6]. In addition, reported genes/miRNAs involved in other drug resistance were also detected, such as SOCS6 [7], ZFP36 [8], and NR3C1 [9]. Except for these reported genes in drug resistance, we have revealed novel genes involved in imatinib intoxication mechanisms, including TCF12, DRD1, ZNF12, DBP, and ACYP1.
Fas-mediated apoptosis pathway and the loss of TP53 were associated with imatinib resistance in chronic myeloid leukemia, indicating that the mechanism of action of imatinib is possibly responsible for imatinib resistance in cancers [6, 10]. Similarly, the loss of TP53 in GIST-T1 cells showed obvious imatinib resistance in our study, which was consistent with the reported study. The induction of SOCS6 [7] and ZFP36 [8] may contribute to drug resistance in breast cancer and omental adipose tissue, respectively. In contrast, the loss of SOCS6 and ZFP36 in GIST-T1 cells resulted in imatinib resistance. Further investigation should be performed to validate the function and mechanisms of SOCS6 and ZFP36 in imatinib resistance. The KEGG pathway analysis of candidate genes has been listed in Additional file 9: Table S6. Additional file 10: Figure S4A has shown the potential signaling pathway contributed to imatinib resistance in GIST, including apoptosis pathway and Wnt signaling pathway, JAK-STAT signaling pathway, which may be related with the loss of TP53 and SOCS6 respectively. More targets to imatinib-resistance would be identified from these candidate genes in the following study. Protein-protein interaction (PPI) network of these candidate genes has been shown in Additional file 10: Figure S4B, which would provide a reference for the discovery of potential targets in imatinib-resistant GIST.
Conclusions
Our study has validated 9 genes involved in imatinib-resistant GIST-T1 cells by genome-scale CRISPR-Cas9 knockout screening. TP53 and SOCS6 may be the most promising candidate gene for imatinib-resistance due to the possible signaling pathway, such as apoptosis pathway and Wnt signaling pathway, JAK-STAT signaling pathway. It is necessary to perform more studies to discover novel targets in imatinib-resistant GIST, including DBP, NR3C1, TCF12, ZNF12, ZFP36, ACYP1 and DRD1.
Additional files
Figure S1. Optical microscopic images of GIST-T1 cells treated with imatinib (40, 60 and 80 μg/mL) with observation at day 2, 4 and 5. (TIF 314 kb)
Table S1. human geckov2 library a. (XLSX 2104 kb)
Table S2. human geckov2 library b. (XLSX 1864 kb)
Figure S2. PCR amplification of sgRNA region for deep-sequencing analysis. (A) Successful transfection of lentiCRISPRv2 vector in GIST-T1 cells, as indicated by electrophoresis of the PCR amplification region (192 bp) from lentiCRISPRv2 vector. M: 2000 bp DNA marker; 1: PCR amplification products of lentiCRISPRv2 vector transfected with GIST-T1 cells; 2: PCR amplification products of GIST-T1 cells without lentiCRISPRv2 vector transfection. (B) PCR amplification of sgRNA region from the sgRNA library for deep-sequencing analysis, as indicated by electrophoresis. M: 2000 bp DNA marker. (C) Sequence of sgRNA region (642 bp) for PCR amplification. The black part: linker adaptor; The red part: variable sequence (24 bp) for sequencing analysis. (TIF 81 kb)
Table S3. The results of high-throughput sequencing analysis. (XLSX 263 kb)
Table S4. Candidate genesmiRNAs with sgRNA sequence, total reads and diversity. (DOCX 13 kb)
Table S5. Detailed information of GO analysis for the selected 20 genes. (DOCX 13 kb)
Figure S3. Optical microscopic images of GIST-T1 cells with individual gene/miRNA knockouts and imatinib treatment. (TIF 606 kb)
Table S6. KEGG pathway analysis of candidate genes. (DOCX 14 kb)
Figure S4. (A)The potential signaling pathway contributed to imatinib resistance in GIST. The green boxes and the solid arrows represented the previously reported signaling pathways related with imatinib resistance; the orange boxes and the dotted arrows represented the potential signaling pathway contributed to imatinib resistance in GIST. (B) Validated genes (9 genes) in protein-protein interaction network. (TIF 319 kb)
Acknowledgments
Funding
The study was supported by grants from the National Natural Science Foundation of China (grant no. 81272556), Guangdong Medical Science and Research Foundation (A2018304), the Science and Technology Project of Guangdong Province (grant no. 2017A030311035) and the Science and Technology Program of Guangzhou, China (grant no. 2014Y2–00137).
Availability of data and materials
All data generated or analysed during this study are included in this published article and its supplementary information files.
Abbreviations
- ACYP1
Acylphosphatase
- AKT
Protein kinase B
- CRISPR
Clustered regularly interspaced short palindrome repeats
- DBP
D-box binding PAR bZIP transcription factor
- DRD1
Dopamine receptor D1
- GIST
Gastrointestinal stromal tumor
- HIF1A
Hypoxia inducible factor 1 alpha subunit
- KEGG
Kyoto encyclopedia of genes and genomes
- MLD
Minimum lethal dose
- mTOR
Mammalian target of the rapamycin
- NR3C1
Nuclear receptor subfamily 3 group C member 1
- PCR
Polymerase chain reaction
- PDGFRA
Platelet-derived growth factor receptor A kinase
- PI3K
Phosphatidylinositol-3-kinase
- qPCR
Real-time quantitative qPCR
- RIPK4
Receptor interacting serine/threonine kinase 4
- SOCS6
Suppressor of cytokine signaling 6
- TCF12
Transcription factor 12
- TP53
Tumor protein p53
- ZFP36
ZFP36 ring finger protein
- ZNF12
Zinc finger protein 12
Authors’ contributions
QW, WL, JC, JW, PY and TZ designed the study and wrote the manuscript; ZC, FH, FW, HC, HH performed the experiments. JZ, ZY and WC collect and analyzed the data. All authors read and approved the final manuscript.
Not applicable
Not applicable
The authors declare that they have no competing interests.
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Jie Cao, Email: doctor_caojie@126.com.
Jianchang Wei, Email: wei12600@126.com.
Ping Yang, Email: northpark@163.com.
Tong Zhang, Email: stong0707@126.com.
Zhuanpeng Chen, Email: 382547177@qq.com.
Feng He, Email: hefengmm@163.com.
Fang Wei, Email: 15521287698@126.com.
Huacui Chen, Email: 791205503@qq.com.
He Hu, Email: 446021441@qq.com.
Junbin Zhong, Email: abinabc@163.com.
Zhi Yang, Email: 35429564@qq.com.
Wensong Cai, Email: caiwensong@163.com.
Wanglin Li, Email: lwl31312008@sina.com.
Qiang Wang, Email: wangqiang973@126.com.
References
- 1.Lai S, Wang G, Cao X, Luo X, Wang G, Xia X, Hu J, Wang J. KIT over-expression by p55PIK-PI3K leads to Imatinib-resistance in patients with gastrointestinal stromal tumors. Oncotarget. 2016;7:1367–1379. doi: 10.18632/oncotarget.6011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Lasota J, Miettinen M. KIT and PDGFRA mutations in gastrointestinal stromal tumors (GISTs) Semin Diagn Pathol. 2006;23:91–102. doi: 10.1053/j.semdp.2006.08.006. [DOI] [PubMed] [Google Scholar]
- 3.Li J, Dang Y, Gao J, Li Y, Zou J, Shen L. PI3K/AKT/mTOR pathway is activated after imatinib secondary resistance in gastrointestinal stromal tumors (GISTs) Med Oncol. 2015;32:111. doi: 10.1007/s12032-015-0554-6. [DOI] [PubMed] [Google Scholar]
- 4.Evers B, Jastrzebski K, Heijmans JP, Grernrum W, Beijersbergen RL, Bernards R. CRISPR knockout screening outperforms shRNA and CRISPRi in identifying essential genes. Nat Biotechnol. 2016;34:631–633. doi: 10.1038/nbt.3536. [DOI] [PubMed] [Google Scholar]
- 5.Kurata M, Rathe SK, Bailey NJ, Aumann NK, Jones JM, Veldhuijzen GW, Moriarity BS, Largaespada DA. Using genome-wide CRISPR library screening with library resistant DCK to find new sources of Ara-C drug resistance in AML. Sci Rep. 2016;6:36199. doi: 10.1038/srep36199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Al-Achkar W, Wafa A, Moassass F, Othman MA. A novel dic (17,18) (p13.1;q11.2) with loss of TP53 and BCR/ABL rearrangement in an Imatinib resistant chronic myeloid leukemia. Mol Cytogenet. 2012;5:36. doi: 10.1186/1755-8166-5-36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Shen R, Wang Y, Wang CX, Yin M, Liu HL, Chen JP, Han JQ, Wang WB. MiRNA-155 mediates TAM resistance by modulating SOCS6-STAT3 signalling pathway in breast cancer. Am J Transl Res. 2015;7:2115–2126. [PMC free article] [PubMed] [Google Scholar]
- 8.Bouchard L, Vohl MC, Deshaies Y, Rheaume C, Daris M, Tchernof A. Visceral adipose tissue zinc finger protein 36 mRNA levels are correlated with insulin, insulin resistance index, and adiponectinemia in women. Eur J Endocrinol. 2007;157:451–457. doi: 10.1530/EJE-07-0073. [DOI] [PubMed] [Google Scholar]
- 9.Liang YN, Tang YL, Ke ZY, Chen YQ, Luo XQ, Zhang H, Huang LB. MiR-124 contributes to glucocorticoid resistance in acute lymphoblastic leukemia by promoting proliferation, inhibiting apoptosis and targeting the glucocorticoid receptor. J Steroid Biochem Mol Biol. 2017;172:62–68. doi: 10.1016/j.jsbmb.2017.05.014. [DOI] [PubMed] [Google Scholar]
- 10.Zheng Q, Cao J, Hamad N, Kim HJ, Moon JH, Sohn SK, Jung CW, Lipton JH, Kim DD. Single nucleotide polymorphisms in apoptosis pathway are associated with response to imatinib therapy in chronic myeloid leukemia. J Transl Med. 2016;14:82. doi: 10.1186/s12967-016-0837-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figure S1. Optical microscopic images of GIST-T1 cells treated with imatinib (40, 60 and 80 μg/mL) with observation at day 2, 4 and 5. (TIF 314 kb)
Table S1. human geckov2 library a. (XLSX 2104 kb)
Table S2. human geckov2 library b. (XLSX 1864 kb)
Figure S2. PCR amplification of sgRNA region for deep-sequencing analysis. (A) Successful transfection of lentiCRISPRv2 vector in GIST-T1 cells, as indicated by electrophoresis of the PCR amplification region (192 bp) from lentiCRISPRv2 vector. M: 2000 bp DNA marker; 1: PCR amplification products of lentiCRISPRv2 vector transfected with GIST-T1 cells; 2: PCR amplification products of GIST-T1 cells without lentiCRISPRv2 vector transfection. (B) PCR amplification of sgRNA region from the sgRNA library for deep-sequencing analysis, as indicated by electrophoresis. M: 2000 bp DNA marker. (C) Sequence of sgRNA region (642 bp) for PCR amplification. The black part: linker adaptor; The red part: variable sequence (24 bp) for sequencing analysis. (TIF 81 kb)
Table S3. The results of high-throughput sequencing analysis. (XLSX 263 kb)
Table S4. Candidate genesmiRNAs with sgRNA sequence, total reads and diversity. (DOCX 13 kb)
Table S5. Detailed information of GO analysis for the selected 20 genes. (DOCX 13 kb)
Figure S3. Optical microscopic images of GIST-T1 cells with individual gene/miRNA knockouts and imatinib treatment. (TIF 606 kb)
Table S6. KEGG pathway analysis of candidate genes. (DOCX 14 kb)
Figure S4. (A)The potential signaling pathway contributed to imatinib resistance in GIST. The green boxes and the solid arrows represented the previously reported signaling pathways related with imatinib resistance; the orange boxes and the dotted arrows represented the potential signaling pathway contributed to imatinib resistance in GIST. (B) Validated genes (9 genes) in protein-protein interaction network. (TIF 319 kb)
Data Availability Statement
All data generated or analysed during this study are included in this published article and its supplementary information files.