Abstract
Huntington's disease (HD) is an inherited, progressive neurodegenerative disease caused by a CAG expansion in the Huntingtin (HTT) gene and various dysfunctions of biological processes in HD have been proposed. Although monogenic, the exact pathogenesis of HD currently remains unclear. To identify the synergistic microRNA (miRNA) pattern in HD, the miRNA expression profile dataset GSE64977 and the gene expression profile dataset GSE64810 were downloaded. Programming software R was used to identify differentially expressed genes (DEGs) and differentially expressed miRNAs (DEMs). Target genes of DEMs were predicted using the TargetScan database. Gene ontology (GO) function of DEGs was generated using the FunRich and a miRNA-mRNA interaction network was constructed using Cytoscape software. In total, 1,612 DEGs and 10 DEMs were identified. GO terms mainly included inflammatory response and immune response in DEGs. A total of 745 target genes were predicted from the DEMs and 33 overlaps were identified between these target genes and DEGs. The miRNA network demonstrated that hsa-miR-4488, hsa-miR-196a-5p, and hsa-miR-549a had a high degree and may be involved with the pathogenesis and potential therapeutic targets of HD.
Keywords: Huntington's disease, miRNAs, target genes, pathogenesis
Introduction
Huntington's disease (HD) is a frequent and incurable hereditary neurodegenerative disorder that impairs motor and cognitive functions (1). With autosomal dominant inheritance, typical mid-life onset, and unrelenting, progressive motor, cognitive and psychiatric symptoms over 15–20 years, the impact of HD on patients and their families is devastating (2). Although caused by a dominantly inherited CAG trinucleotide repeat expansion in the HD gene on chromosome 4 (3), the pathogenesis of HD has not yet been fully elucidated. Previous studies showed the number of CAG repeats was associated with the age of onset; however, only 50 to 70% of the variation can be attributed to repeat size (4). In addition, significant variation in clinical phenotypes is not well explained (5). Consequently, all these variations indicate other pathogenic factors such as heredity affect the disease progression. Recently, studies have focused on miRNAs, the small non-coding RNAs that participate in transcriptional regulation and translational repression of target genes (6). In HD, a neurodegenerative disease caused by a trinucleotide repeat expansion, miRNAs can interact with RelA/NFkB, p53 (7), Mitofusin2 (8), TBP (9), REST, or RE1 (10,11). Dysregulation of miRNA may also impact CAG length and affect the progression or severity of HD (12). Conversely, a certain degree of genetic heterogeneity of HD that may exhibit different miRNA expression in some cases has also been reported (13). In addition to research on the pathological mechanism, the role of miRNA in the treatment has been a subject of interest. Currently, miRNA-based Huntingtin (HTT)-lowering therapy is one of the most advanced gene strategies (14), silencing the HD gene by injecting artificial miRNA into the striatum of Q140/Q140 mice and transgenic sheep models and achieving the expected effect (15,16). However, bioinformatics studies focusing on miRNA and mRNA expression profiles in HD patients and healthy controls have not been published to date. Therefore, identifying the miRNA-mRNA interactions, understanding their synergistic effects on the pathogenesis, and exploring possible therapeutic approaches for HD are important.
To elucidate the miRNAs and associated target genes and pathways involved in HD, we downloaded miRNA and mRNA expression profiles of HD patients and healthy controls from the Gene Expression Omnibus (GEO) database. The differentially expressed genes (DEGs) and differentially expressed miRNAs (DEMs) target genes were identified and a miRNA-mRNA regulatory network established.
Materials and methods
miRNA and mRNA expression profiles
The mRNA and miRNA expression profiles of HD were downloaded from the GEO database (www.ncbi.nlm.nih.gov/geo) and were termed GSE64810 (https://www.ncbi.nlm.nih.gov/gds/?term=GSE64810; accessed August 8, 2017) and GSE64977 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE64977; accessed August 9, 2017), respectively (6,17). In the GSE64810 dataset, 20 HD and 49 neurologically normal control samples from post-mortem human subjects were included. In the GSE64977 dataset, expression profiles were obtained from 28 HD and 36 neurologically normal control prefrontal cortex samples. Both mRNA and miRNA profiling was performed using the Illumina HiSeq 2000 (Homo sapiens) platform (Illumina, Inc., San Diego, CA, USA).
Differential expression analysis for mRNA and miRNA expression profiling
Raw microarray data were first preprocessed including background correction and normalization. The probes corresponding to multiple genes were abandoned and multiple probes corresponding to one gene or miRNA were used to calculate the average expression level. Limma package (www.bioconductor.org/packages/release/bioc/html/limma.html) was used to identify DEGs and DEMs between HD and control samples. A two-tailed Student's t-test was used for statistical analysis. The DEGs and DEMs were considered significantly differentially expressed if the P-value was <0.05 and the log fold change (FC) was >1. The miRNAs were further classified using hierarchical clustering analysis.
Prediction of miRNAs associated with DEGs and construction of miRNA gene regulatory network
Target genes regulated by DEMs were predicted using TargetScan (http://www.targetscan.org/vert_71/), which is used to predict biological targets of miRNAs by searching for the presence of conserved 8mer, 7mer, and 6mer sites that match the seed region of each miRNA (18). In mammals, predictions are ranked based on the predicted efficacy of targeting as calculated using cumulative weighted context ++ scores of the sites (19). In the present study, the miRNA target gene set with context ++ scores <-0.4 were then used for further analysis. A Venn diagram of DEGs overlapping with DEM target genes was constructed using FunRich (20). Furthermore, Cytoscape was utilized to visualize the miRNA gene regulatory network (21).
Functional enrichment analyses
Functional enrichment analyses were performed using the open software FunRich (http://funrich.org/faq), which is a stand-alone software tool used mainly for functional enrichment and interaction network analysis of genes and proteins (20). Statistical cut-off of enrichment analyses in FunRich software was set to default with a P-value <0.05 after Bonferroni correction.
Results
Differential expression analysis
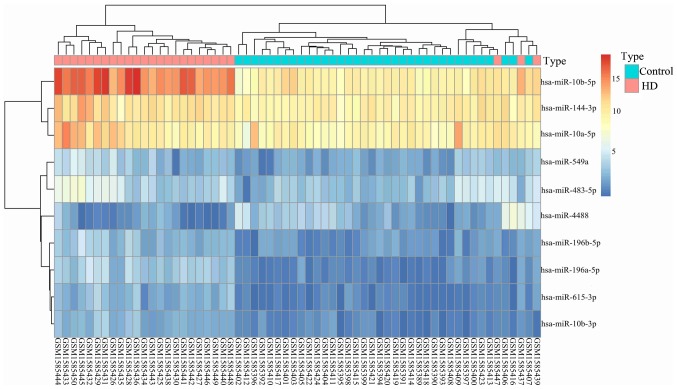
Based on the gene expression analysis, 8 upregulated and 2 downregulated DEMs were identified in HD compared with the normal controls (Table I). Based on the mRNA expression profile analysis, 1,612 DEGs were identified, including 945 upregulated and 667 downregulated DEGs. The top 10 DEGs are presented in Table II. The hierarchical clustering heat map of miRNA shows the differences between HD and normal controls (Fig. 1).
Table I.
Differentially expressed miRNAs between HD patients and healthy controls.
| miRNA | Log FC | P-value |
|---|---|---|
| hsa-miR-10b-5p | 4.175 | 9.10×10–23 |
| hsa-miR-196a-5p | 2.430 | 5.85×10–22 |
| hsa-miR-615-3p | 1.678 | 2.12×10–17 |
| hsa-miR-10b-3p | 1.480 | 4.79×10–14 |
| hsa-miR-196b-5p | 1.439 | 6.62×10–11 |
| hsa-miR-144-3p | 1.045 | 2.77×10-6 |
| hsa-miR-549a | 1.111 | 2.39×10-5 |
| hsa-miR-483-5p | 1.256 | 2.49×10-4 |
| hsa-miR-10a-5p | 1.042 | 6.0×10-4 |
| hsa-miR-4488 | −1.193 | 2.54×10-3 |
P<0.05 was considered to indicate a statistically significant difference. miRNA, microRNA; FC, fold-change; HD, Huntington's disease.
Table II.
Top 10 most differentially expressed mRNAs between HD patients and healthy controls.
| Gene | Log FC | P-value |
|---|---|---|
| PITX1 | 4.770 | 9.57×10–39 |
| POU4F2 | 3.962 | 3.42×10–23 |
| HAND1 | 3.703 | 1.46×10–17 |
| HOXD9 | 3.657 | 1.22×10–18 |
| SLC16A12 | 3.514 | 4.74×10–18 |
| PITX2 | 3.404 | 1.66×10–12 |
| BMP5 | 3.149 | 5.93×10–13 |
| OGN | 3.097 | 8.20×10–14 |
| SLC22A2 | 3.089 | 6.93×10–11 |
| IL1R2 | 3.038 | 3.35×10–12 |
P<0.05 was considered to indicate a statistically significant difference. FC, fold-change; HD, Huntington's disease.
Figure 1.
Hierarchical clustering heatmap of 10 DEMs in different datasets.
Functional enrichment analysis
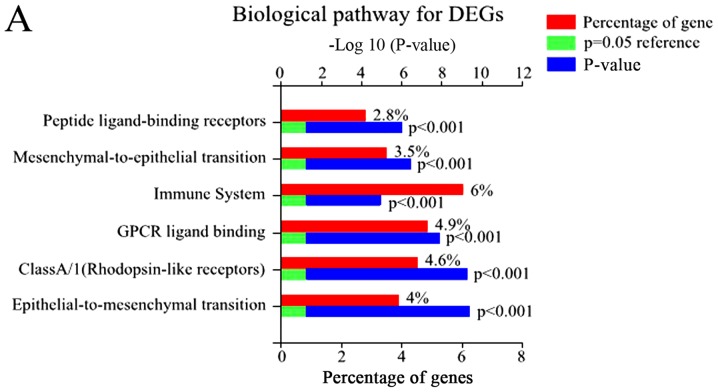
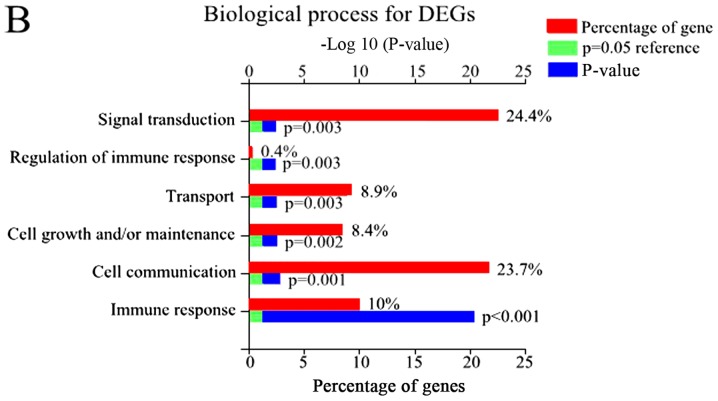
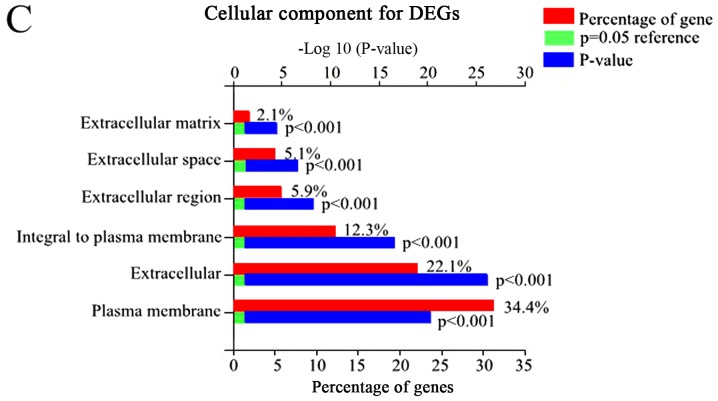
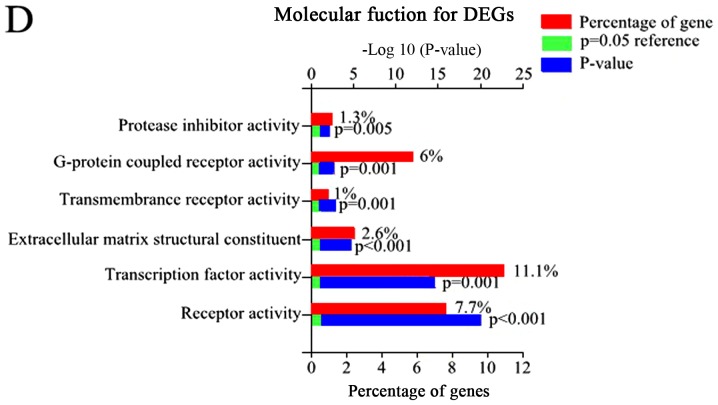
Using FunRich, 48 enriched GO terms were obtained for DEGs. The 10 most significantly enriched GO terms are listed in Table III, including integral components of the plasma membrane, inflammatory response, plasma membrane, and immune response. Furthermore, the most significantly enriched GO terms including biological processes (BP), molecular function (MF), and cellular component (CC) were analyzed (Fig. 2).
Table III.
Top 10 GO functional annotation of differentially expressed genes.
| GO ID | P-value | Term |
|---|---|---|
| GO:0005887 | 6.55×10–21 | Integral component of plasma membrane |
| GO:0006954 | 1.54×10–19 | Inflammatory response |
| GO:0005886 | 4.77×10–19 | Plasma membrane |
| GO:0006955 | 3.86×10–19 | Immune response |
| GO:0005576 | 4.51×10–18 | Extracellular region |
| GO:0005615 | 1.43×10–17 | Extracellular space |
| GO:0045087 | 1.65×10–15 | Innate immune response |
| GO:0009952 | 1.61×10–14 | Anterior/posterior pattern specification |
| GO:0043565 | 1.22×10–11 | Sequence-specific DNA binding |
| GO:0042742 | 4.73×10–10 | Defense response to bacterium |
P<0.05 was considered to indicate a statistically significant difference. GO, Gene Ontology.
Figure 2.
The GO and biological pathway analysis of all DEGs. (A) Biological pathway analysis of DEGs, (B) BP of DEGs. (C) CC of DEGs. (D) MF of DEGs.
Synergistic miRNA network construction
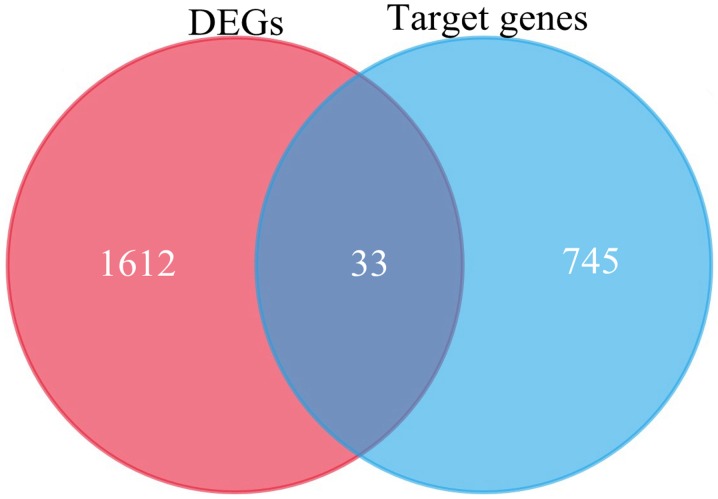
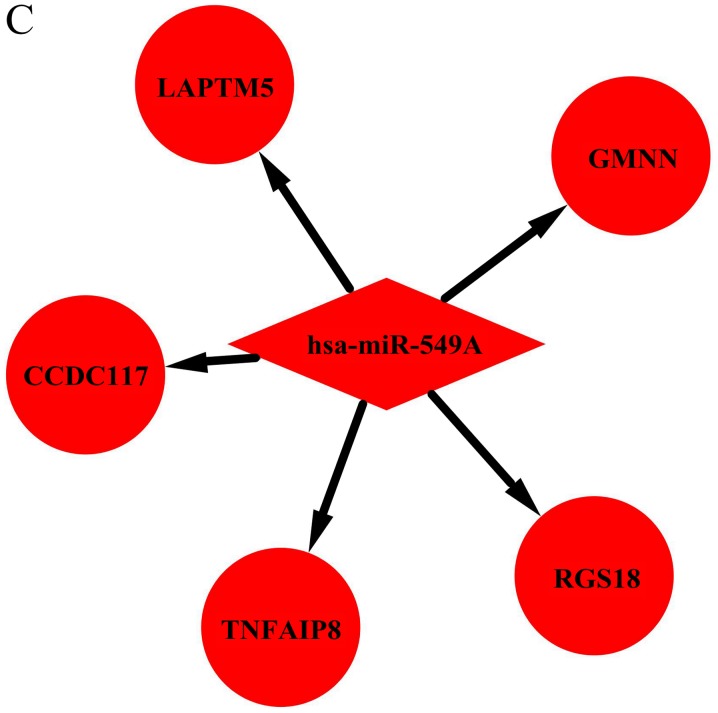
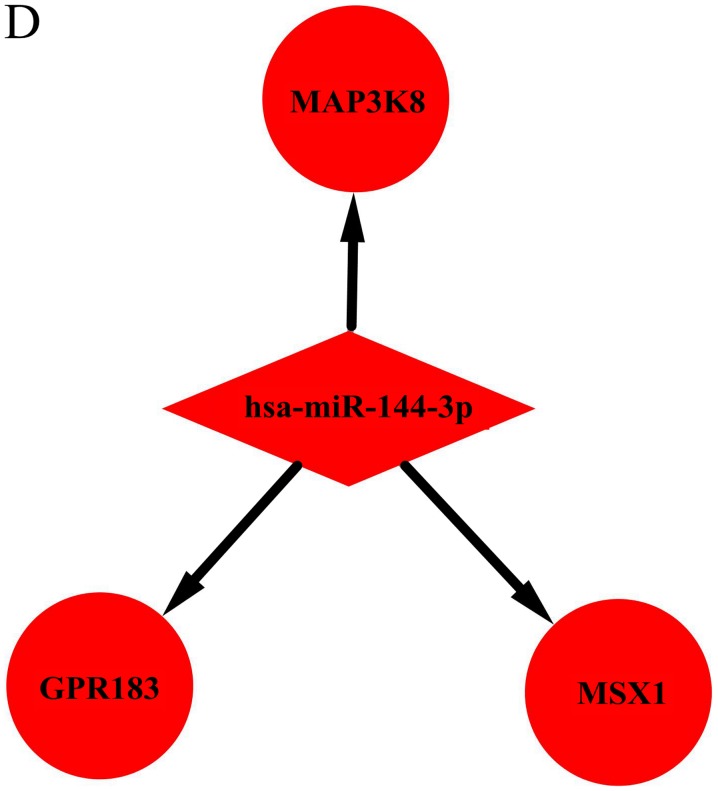
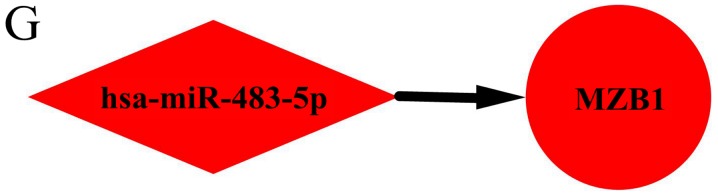
Utilizing TargetScan, 745 target genes of DEMs and 33 overlaps were identified between the target genes and DEGs (Fig. 3). We identified 33 possible miRNA-mRNA target pairs. In the network, hsa-miR-4488, hsa-miR-196a-5p, and hsa-miR-549a had a high degree and may be involved in the pathogenesis and used as potential therapeutic targets for HD (Fig. 4).
Figure 3.
Venn diagram of DEGs overlapping with DEM target genes using FunRich.
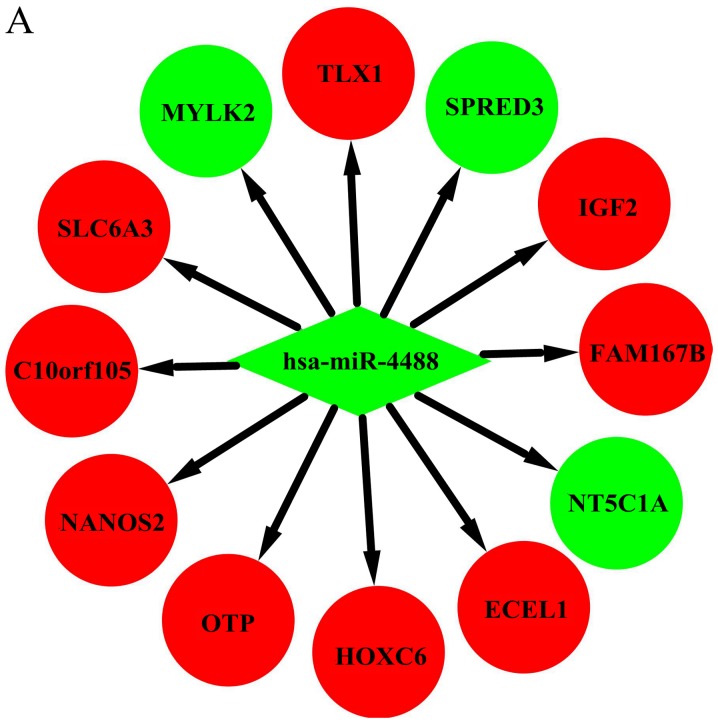
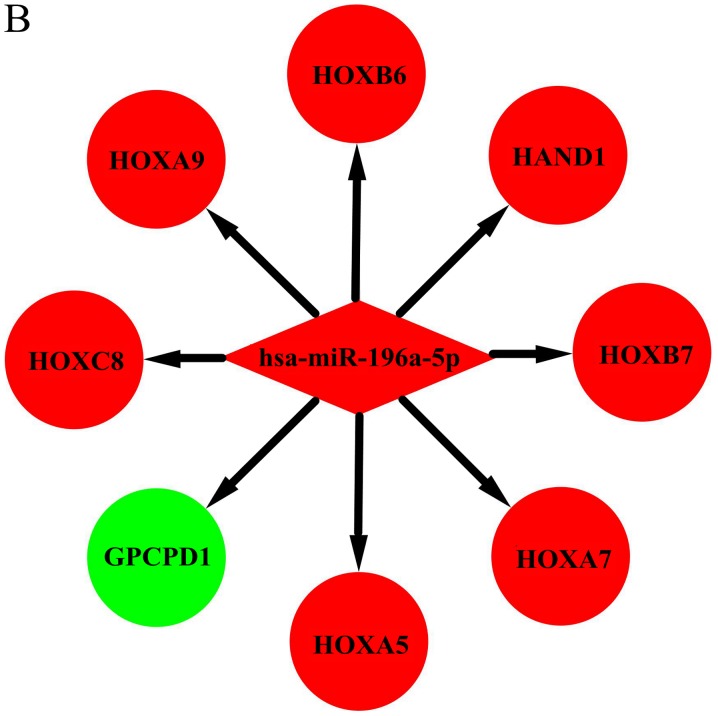
Figure 4.
Interaction network of DEMs and target genes in HD. Interaction networks for (A) hsa-miR-4488, (B) hsa-miR-196a-5p, (C) hsa-miR-459A, The red and green colors represent upregulation and downregulation, respectively. Interaction network of DEMs and target genes in HD. (D) hsa-miR-144-3p, (E) hsa-miR-10b-3p, (F) hsa-miR-10a-5p and (G) hsa-miR-483-5p. The red and green colors represent upregulation and downregulation, respectively.
Discussion
HD is an inherited progressive neurodegenerative disorder that usually affects people in midlife (22). Previous studies have implicated that gene expression may be altered at more than one stage of RNA processing, translation, protein post-transcriptional trafficking or modifications (23). miRNAs are small non-coding RNAs that can influence diverse ranges of cellular processes (24). Recently, dysregulation of miRNAs has been reportedly associated with neurological and neurodegenerative disorders (25) and several studies have explored the functions of miRNAs in HD patients (10,26). Although altered expression of miRNAs has been reported in cellular models (10) and mouse models of HD (27) using quantified microarray technology, a bioinformatics study of miRNA and mRNA expression profiles in HD patients and healthy controls has not yet been performed. In the present study, we elucidated the synergistic effects of miRNAs on the pathogenesis of HD by constructing a miRNA-mRNA interaction network.
Using a bioinformatics approach to analyze differential expression profiles of mRNA and miRNA, HD patients showed significant differences in mRNA and miRNA expression when compared with neurologically normal controls. In total, 1,612 DEGs and 10 DEMs were identified. GO function (BP, CC, and MF) and biological pathway analysis of DEGs demonstrated the majority of genes were involved in some processes and pathways, such as signal transduction in BP (28), plasma membrane in CC (29), transcription factor activity in MF (30), and immune system in biological pathway (28). Changes in Ca2+ signaling and/or transduction systems and misfolded protein-plasma membrane interactions affected HD initiation and progression as reported by Fan and Shrivastava (29,31). Many previous studies showed that aberrant transcriptional regulations play important roles in the molecular pathogenesis of HD. The HTT gene interferes with some important transcription factors (32); among these, Sp1 and TAFII130 were shown disrupted in the early stage of HD progression (33). Furthermore, both innate and adaptive immune systems have been previously suggested to be activated during the progression of HD (34). Reportedly, HD is characterized by cellular and molecular features of inflammation (cytokine expression and microglia activation) and HTT mRNA expression in immune cells is on average higher than that observed in most organs (35). miRNAs may also influence the dysregulated production of both type 1 and type 2 cytokines observed at different stages of HD. This result is attracting attention as pathogenetic mechanism and as possible therapeutic approach with immunomodulation (36). Combined with the previous research conclusions, the present functional analysis may provide novel therapeutic targets or possible pathogenesis to be further studied.
In the context of the miRNA-mRNA interaction network, hsa-miR-4488, hsa-miR-196a-5p, and hsa-miR-549a had a high degree in this study (Table IV). Hsa-miR-4488 was downregulated and had 12 target genes in our study. Lee et al also found that miR-4488 was differentially downregulated in their HD patients (27). Among the target genes of hsa-miR-4488 in the present study, SPRED3 (37), IGF2 (38), HOXC6 (39), NANOS2 (40), SLC6A3 (41), TLX1 (42), and NT5C1A (43) have been reported to play various roles in the pathogenesis or development of HD. Although we could not determine the association of FAM167B, ECEL1, OTP, C10orf105, or MYLK2 with HD, further experimental studies are necessary to elucidate their possible relationships with HD. Previous studies also showed hsa-miR-196a plays an important role in the pathogenesis and progression of HD (44), Fu et al suggested that hsa-miR-196a altered the RIG-I-like receptor signaling pathway and the immune system, as well as changed the expression of several well-defined pathways of HD, such as apoptosis and cell adhesion (45). A previous study showed that HOX is indirectly involved in the neuroprotective response in HD and increased expression of HOX genes can enhance H3K27me3 or impair PcG repression (39). In our present study, hsa-miR-196a-5p was found to mainly interact with HOX genes, indicating hsa-miR-196a-5p may be involved in the neuroprotective response in HD. Reportedly, miR-196a dominantly altered the immune system or adaptive immune system (45). During the last decade, a hyperreactive immune system has been recognized as an important feature of HD pathogenesis. Macrophages in individuals with manifested HD and even pre-manifested HD have been reported to release more proinflammatory cytokines such as TNF-alpha (46) and prototypical anti-inflammatory cytokines such as IL-10 (47) and IL-13 (48). Similarly, Dobson et al showed that at baseline, monocytes from HD subjects released more cytokines than monocytes isolated from healthy volunteers, and this abnormality could be modulated by laquinimod, which exerts an immunomodulatory effect on isolated HD monocytes (49).
Table IV.
Some of the miRNAs associated with HD.
| Author, year | miRNA | Change in expression | Role in HD | Target genes | (Refs.) |
|---|---|---|---|---|---|
| Burgunder J-M et al, 2014 | miR-144-3p | Upregulated | Not found | GPR183, MAP3K8, MSX1 | (4) |
| miR-10b-3p | Upregulated | Influence CAG length in HD | HOXD1, ZG16B | ||
| miR-10a-5p | Upregulated | Not found | HOXB3, TFAP2C | ||
| miR-483-5p | Upregulated | Not found | MZB1 | ||
| Traeger U et al, 2014; Shrivastava AN et al, 2017; van Hagen M et al, 2017; Fan MMY et al, 2007; Moumne L et al, 2013; Dunah AW et al, 2002; Squitieri F et al, 2006 | miR-4488 | Downregulated | Post-translational protein modification; post-transcriptional regulation by preventing translational initiation; | SPRED3, IGF2, HOXC6, NANOS2, SLC6A3, TLX1, NT5C1A, FAM167B, ECEL1, OTP, C10orf105, MYLK2 | (28–34) |
| Butland SL et al, 2014; Salem L et al, 2016; Hoss AG et al, 2014; Rokavec M et al, 2014 | miR-549a | Upregulated | Regulate transcription factors, affect autophagy and endoplasmic reticulum stress pathway | CCDC117, GMNN, LAPTM5, RGS18, TNFAIP8 | (37–40) |
| Cheng P-H et al, 2013; Fu M-H et al, 2015; Zou J et al, 2015 | miR-196a | Upregulated | Neuroprotective effect by interacting with HOX genes; affects the entire immune system leading to disorder of cytokine secretion | GPCPD1, HAND1, HOXA5, HOXA7, HOXA9, HOXB6, HOXB7, HOXC8 | (44–46) |
P<0.05 was considered to indicate a statistically significant difference. miR, microRNA; HD, Huntington's disease.
In addition, we found another miRNA, hsa-miR-549a, was upregulated in HD patients and could regulate five target genes. When reviewing the literature, we found that GMNN (50), TNFAIP18 (51), LAPTM5 (52), and RGS18 (53) played roles in the pathogenesis of neurodegenerative diseases through various mechanisms, including regulation of transcription factors and affecting autophagy and endoplasmic reticulum stress pathways and might be involved in the pathogenesis of HD. Other affected miRNAs found in our study including hsa-miR-144-3p, hsa-miR-10b-3p, hsa-miR-10a-5p, and hsa-miR-483-5p have been reported as differentially expressed in HD samples; hsa-miR-10b-3p showed a significant association with CAG length in HD (6). Regarding the other three miRNAs, more research is be needed to identify their exact relationships with HD.
Although extensive research has identified aberrantly expressed miRNAs in HD, the molecular mechanisms underlying the pathological implications remain largely unknown. Using expression profile datasets, we compared the genomic expression status of HD and revealed differentially expressed mRNAs and DEMs. We identified DEGs and constructed a miRNA-mRNA regulatory network. We found hsa-miR-4488, hsa-miR-196a-5p, and hsa-miR-549a had a high degree and may be involved in the pathogenesis of HD. Studies in this field could help improve the understanding of how miRNAs mediate the etiopathological mechanisms of HD. Since the neuroprotective effects of certain miRNAs have been demonstrated in animal studies, the therapeutic potential of miRNAs should be further investigated and followed by molecular validation.
Acknowledgements
Not applicable.
Glossary
Abbreviations
- HD
huntington disease
- miRNA
microRNA
- DEMs
differentially expressed miRNAs
- DEGs
differentially expressed genes
- PPI
protein-protein interaction
- GEO
Gene Expression Omnibus
- GO
gene ontology
- KEGG
Kyoto Encyclopedia of Genes and Genomes
- FC
fold change
- MCODE
molecular complex detection
- BP
biological processes
- MF
molecular function
- CC
cellular component
Funding
The present study was supported by the National Natural Science Foundation of China (grant no. 81371271) and was sponsored by the Liaoning Bai Qian Wan Talents Program [grant no. (2015)41].
Availability of data and materials
The datasets analyzed in the present study are available in the GEO repository, https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE64977 and https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE64810.
Authors' contributions
XD designed the research, performed data and statistical analysis, and drafted the manuscript. SC conceived and designed the research, and revised the manuscript for important intellectual content. All authors read and approved the final manuscript.
Ethics approval and consent to participate
The present study was approved by the Ethics Committee of the Shengjing Hospital of China Medical University, Liaoning, China.
Patient consent for publication
Written informed consent was obtained from all volunteers for the publication of the associated data.
Competing interests
The authors declare that they have no competing interests.
References
- 1.Huang WJ, Chen WW, Zhang X. Huntington's disease: Molecular basis of pathology and status of current therapeutic approaches. Exp Ther Med. 2016;12:1951–1956. doi: 10.3892/etm.2016.3566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ghosh R, Tabrizi SJ. Clinical features of Huntington's disease. Adv Exp Med Biol. 2018;1049:1–28. doi: 10.1007/978-3-319-71779-1_1. [DOI] [PubMed] [Google Scholar]
- 3.McColgan P, Tabrizi SJ. Huntington's disease: A clinical review. Eur J Neurol. 2018;25:24–34. doi: 10.1111/ene.13413. [DOI] [PubMed] [Google Scholar]
- 4.Burgunder JM. Genetics of Huntington's disease and related disorders. Drug Discov Today. 2014;19:985–989. doi: 10.1016/j.drudis.2014.03.005. [DOI] [PubMed] [Google Scholar]
- 5.Djoussé L, Knowlton B, Hayden M, Almqvist EW, Brinkman R, Ross C, Margolis R, Rosenblatt A, Durr A, Dode C, et al. Interaction of normal and expanded CAG repeat sizes influences age at onset of Huntington disease. Am J Med Genet A. 2003;119A:279–282. doi: 10.1002/ajmg.a.20190. [DOI] [PubMed] [Google Scholar]
- 6.Hoss AG, Labadorf A, Latourelle JC, Kartha VK, Hadzi TC, Gusella JF, MacDonald ME, Chen JF, Akbarian S, Weng Z, et al. miR-10b-5p expression in Huntington's disease brain relates to age of onset and the extent of striatal involvement. BMC Med Genomics. 2015;8:10. doi: 10.1186/s12920-015-0083-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ghose J, Sinha M, Das E, Jana NR, Bhattacharyya NP. Regulation of miR-146a by RelA/NFkB and p53 in STHdh(Q111)/Hdh(Q111) cells, a cell model of Huntington's disease. PLos One. 2011;6:e23837. doi: 10.1371/journal.pone.0023837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bucha S, Mukhopadhyay D, Bhattacharyya NP. Regulation of mitochondrial morphology and cell cycle by microRNA-214 targeting Mitofusin2. Biochem Biophys Res Commun. 2015;465:797–802. doi: 10.1016/j.bbrc.2015.08.090. [DOI] [PubMed] [Google Scholar]
- 9.Sinha M, Ghose J, Das E, Bhattarcharyya NP. Altered microRNAs in STHdh(Q111)/Hdh(Q111) cells: miR-146a targets TBP. Biochem Biophys Res Commun. 2010;396:742–747. doi: 10.1016/j.bbrc.2010.05.007. [DOI] [PubMed] [Google Scholar]
- 10.Johnson R, Zuccato C, Belyaev ND, Guest DJ, Cattaneo E, Buckley NJ. A microRNA-based gene dysregulation pathway in Huntington's disease. Neurobiol Dis. 2008;29:438–445. doi: 10.1016/j.nbd.2007.11.001. [DOI] [PubMed] [Google Scholar]
- 11.Packer AN, Xing Y, Harper SQ, Jones L, Davidson BL. The bifunctional microRNA miR-9/miR-9* regulates REST and CoREST and is downregulated in Huntington's disease. J Neurosci. 2008;28:14341–14346. doi: 10.1523/JNEUROSCI.2390-08.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Langfelder P, Gao F, Wang N, Howland D, Kwak S, Vogt TF, Aaronson JS, Rosinski J, Coppola G, Horvath S, Yang XW. MicroRNA signatures of endogenous Huntingtin CAG repeat expansion in mice. PLoS One. 2018;13:e0190550. doi: 10.1371/journal.pone.0190550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kartsaki E, Spanaki C, Tzagournissakis M, Petsakou A, Moschonas N, Macdonald M, Plaitakis A. Late-onset and typical Huntington disease families from Crete have distinct genetic origins. Int J Mol Med. 2006;17:335–346. [PubMed] [Google Scholar]
- 14.Miniarikova J, Evers MM, Konstantinova P. Translation of MicroRNA-based huntingtin-lowering therapies from preclinical studies to the Clinic. Mol Ther. 2018;26:947–962. doi: 10.1016/j.ymthe.2018.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Keeler AM, Sapp E, Chase K, Sottosanti E, Danielson E, Pfister E, Stoica L, DiFiglia M, Aronin N, Sena-Esteves M. Cellular analysis of silencing the Huntington's disease gene using AAV9 mediated delivery of artificial micro rna into the striatum of Q140/Q140 mice. J Huntingtons Dis. 2016;5:239–248. doi: 10.3233/JHD-160215. [DOI] [PubMed] [Google Scholar]
- 16.Pfister EL, DiNardo N, Mondo E, Borel F, Conroy F, Fraser C, Gernoux G, Han X, Hu D, Johnson E, et al. Artificial miRNAs reduce human mutant huntingtin throughout the striatum in a transgenic sheep model of Huntington's disease. Hum Gene Ther. 2018 Feb 23; doi: 10.1089/hum.2017.199. (Epub ahead of print) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Labadorf A, Hoss AG, Lagomarsino V, Latourelle JC, Hadzi TC, Bregu J, MacDonald ME, Gusella JF, Chen JF, Akbarian S, et al. Correction: RNA sequence analysis of human huntington disease brain reveals an extensive increase in inflammatory and developmental gene expression. PLoS One. 2016;11:e0160295. doi: 10.1371/journal.pone.0160295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005;120:15–20. doi: 10.1016/j.cell.2004.12.035. [DOI] [PubMed] [Google Scholar]
- 19.Agarwal V, Bell GW, Nam JW, Bartel DP. Predicting effective microRNA target sites in mammalian mRNAs. Elife. 2015;4:e05005. doi: 10.7554/eLife.05005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Pathan M, Keerthikumar S, Ang CS, Gangoda L, Quek CY, Williamson NA, Mouradov D, Sieber OM, Simpson RJ, Salim A, et al. FunRich: An open access standalone functional enrichment and interaction network analysis tool. Proteomics. 2015;15:2597–2601. doi: 10.1002/pmic.201400515. [DOI] [PubMed] [Google Scholar]
- 21.Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–2504. doi: 10.1101/gr.1239303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lund E, Guttinger S, Calado A, Dahlberg JE, Kutay U. Nuclear export of microRNA precursors. Science. 2004;303:95–98. doi: 10.1126/science.1090599. [DOI] [PubMed] [Google Scholar]
- 23.Cha JH. Transcriptional signatures in Huntington's disease. Prog Neurobiol. 2007;83:228–248. doi: 10.1016/j.pneurobio.2007.03.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Bartel DP. MicroRNAs: Target recognition and regulatory functions. Cell. 2009;136:215–233. doi: 10.1016/j.cell.2009.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Junn E, Mouradian MM. MicroRNAs in neurodegenerative diseases and their therapeutic potential. Pharmacol Ther. 2012;133:142–150. doi: 10.1016/j.pharmthera.2011.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Jin J, Cheng Y, Zhang Y, Wood W, Peng Q, Hutchison E, Mattson MP, Becker KG, Duan W. Interrogation of brain miRNA and mRNA expression profiles reveals a molecular regulatory network that is perturbed by mutant huntingtin. J Neurochem. 2012;123:477–490. doi: 10.1111/j.1471-4159.2012.07925.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Lee ST, Chu K, Im WS, Yoon HJ, Im JY, Park JE, Park KH, Jung KH, Lee SK, Kim M, Roh JK. Altered microRNA regulation in Huntington's disease models. Exp Neurol. 2011;227:172–179. doi: 10.1016/j.expneurol.2010.10.012. [DOI] [PubMed] [Google Scholar]
- 28.Träger U, Andre R, Lahiri N, Magnusson-Lind A, Weiss A, Grueninger S, McKinnon C, Sirinathsinghji E, Kahlon S, Pfister EL, et al. HTT-lowering reverses Huntington's disease immune dysfunction caused by NFκB pathway dysregulation. Brain. 2014;137:819–833. doi: 10.1093/brain/awt355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Shrivastava AN, Aperia A, Melki R, Triller A. Physico-pathologic mechanisms involved in neurodegeneration: Misfolded protein-plasma membrane interactions. Neuron. 2017;95:33–50. doi: 10.1016/j.neuron.2017.05.026. [DOI] [PubMed] [Google Scholar]
- 30.van Hagen M, Piebes DGE, de Leeuw WC, Vuist IM, van Roon-Mom WMC, Moerland PD, Verschure PJ. The dynamics of early-state transcriptional changes and aggregate formation in a Huntington's disease cell model. BMC Genomics. 2017;18:373. doi: 10.1186/s12864-017-3745-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Fan MM, Raymond LA. N-methyl-D-aspartate (NMDA) receptor function and excitotoxicity in Huntington's disease. Prog Neurobiol. 2007;81:272–293. doi: 10.1016/j.pneurobio.2006.11.003. [DOI] [PubMed] [Google Scholar]
- 32.Moumne L, Betuing S, Caboche J. Multiple aspects of gene dysregulation Huntington's disease. Front Neurol. 2013;4:127. doi: 10.3389/fneur.2013.00127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Dunah AW, Jeong H, Griffin A, Kim YM, Standaert DG, Hersch SM, Mouradian MM, Young AB, Tanese N, Krainc D. Sp1 and TAFII130 transcriptional activity disrupted in early Huntington's disease. Science. 2002;296:2238–2243. doi: 10.1126/science.1072613. [DOI] [PubMed] [Google Scholar]
- 34.Squitieri F, Cannella M, Sgarbi G, Maglione V, Falleni A, Lenzi P, Baracca A, Cislaghi G, Saft C, Ragona G, et al. Severe ultrastructural mitochondrial changes in lymphoblasts homozygous for Huntington disease mutation. Mech Ageing Dev. 2006;127:217–220. doi: 10.1016/j.mad.2005.09.010. [DOI] [PubMed] [Google Scholar]
- 35.Crotti A, Glass CK. The choreography of neuroinflammation in Huntington's disease. Trends Immunol. 2015;36:364–373. doi: 10.1016/j.it.2015.04.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Björkqvist M. lmmunomodulation-a disease-modifying avenue for treatment of Huntington's disease? J Neurochem. 2016;137:670–672. doi: 10.1111/jnc.13539. [DOI] [PubMed] [Google Scholar]
- 37.Butland SL, Sanders SS, Schmidt ME, Riechers SP, Lin DT, Martin DD, Vaid K, Graham RK, Singaraja RR, Wanker EE, et al. The palmitoyl acyltransferase HIP14 shares a high proportion of interactors with huntingtin: implications for a role in the pathogenesis of Huntington's disease. Hum Mol Genet. 2014;23:4142–4160. doi: 10.1093/hmg/ddu137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Salem L, Saleh N, Désaméricq G, Youssov K, Dolbeau G, Cleret L, Bourhis ML, Azulay JP, Krystkowiak P, Verny C, et al. Insulin-like growth factor-1 but not insulin predicts cognitive decline in Huntington's disease. PLoS One. 2016;11:e0162890. doi: 10.1371/journal.pone.0162890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Hoss AG, Kartha VK, Dong X, Latourelle JC, Dumitriu A, Hadzi TC, Macdonald ME, Gusella JF, Akbarian S, Chen JF, et al. MicroRNAs located in the Hox gene clusters are implicated in Huntington's disease pathogenesis. PLoS Genet. 2014;10:e1004188. doi: 10.1371/journal.pgen.1004188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Rokavec M, Li H, Jiang L, Hermeking H. The p53/miR-34 axis in development and disease. J Mol Cell Biol. 2014;6:214–230. doi: 10.1093/jmcb/mju003. [DOI] [PubMed] [Google Scholar]
- 41.Shapshak P. Molecule of the month: miRNA and proteins DARPP-32, DRD1, SLC6A3, and CK2. Bioinformation. 2013;9:274–275. doi: 10.6026/97320630009274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ooi L, Wood IC. Regulation of gene expression in the nervous system. Biochem J. 2008;414:327–341. doi: 10.1042/BJ20080963. [DOI] [PubMed] [Google Scholar]
- 43.Lloyd TE, Christopher-Stine L, Pinal-Fernandez I, Tiniakou E, Petri M, Baer A, Danoff SK, Pak K, Casciola-Rosen LA, Mammen AL. Cytosolic 5′-nucleotidase 1A As a target of circulating autoantibodies in autoimmune diseases. Arthritis Care Res (Hoboken) 2016;68:66–71. doi: 10.1002/acr.22600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Cheng PH, Li CL, Chang YF, Tsai SJ, Lai YY, Chan AW, Chen CM, Yang SH. miR-196a ameliorates phenotypes of Huntington disease in cell, transgenic mouse, and induced pluripotent stem cell models. Am J Hum Genet. 2013;93:306–312. doi: 10.1016/j.ajhg.2013.05.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Fu MH, Li CL, Lin HL, Tsai SJ, Lai YY, Chang YF, Cheng PH, Chen CM, Yang SH. The potential regulatory mechanisms of miR-196a in Huntington's disease through bioinformatic analyses. PLoS One. 2015;10:e0137637. doi: 10.1371/journal.pone.0137637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Zou J, Guo P, Lv N, Huang D. Lipopolysaccharide-induced tumor necrosis factor-α factor enhances inflammation and is associated with cancer (Review) Mol Med Rep. 2015;12:6399–6404. doi: 10.3892/mmr.2015.4243. [DOI] [PubMed] [Google Scholar]
- 47.Gerard C, Bruyns C, Marchant A, Abramowicz D, Vandenabeele P, Delvaux A, Fiers W, Goldman M, Velu T. Interleukin 10 reduces the release of tumor necrosis factor and prevents lethality in experimental endotoxemia. J Exp Med. 1993;177:547–550. doi: 10.1084/jem.177.2.547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Nicoletti F, Mancuso G, Cusumano V, Di Marco R, Zaccone P, Bendtzen K, Teti G. Prevention of endotoxin-induced lethality in neonatal mice by interleukin-13. Eur J Immunol. 1997;27:1580–1583. doi: 10.1002/eji.1830270639. [DOI] [PubMed] [Google Scholar]
- 49.Dobson L, Träger U, Farmer R, Hayardeny L, Loupe P, Hayden MR, Tabrizi SJ. Laquinimod dampens hyperactive cytokine production in Huntington's disease patient myeloid cells. J Neurochem. 2016;137:782–794. doi: 10.1111/jnc.13553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Lee HK, Lee HS, Moody SA. Neural transcription factors: From embryos to neural stem cells. Mol Cells. 2014;37:705–712. doi: 10.14348/molcells.2014.0227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Stefani IC, Wright D, Polizzi KM, Kontoravdi C. The role of ER stress-induced apoptosis in neurodegeneration. Curr Alzheimer Res. 2012;9:373–387. doi: 10.2174/156720512800107618. [DOI] [PubMed] [Google Scholar]
- 52.Glowacka WK, Alberts P, Ouchida R, Wang JY, Rotin D. LAPTM5 protein is a positive regulator of proinflammatory signaling pathways in macrophages. J Biol Chem. 2012;287:27691–27702. doi: 10.1074/jbc.M112.355917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Raju HB, Tsinoremas NF, Capobianco E. Emerging putative associations between non-coding RNAs and protein-coding genes in neuropathic pain: Added value from reusing microarray data. Front Neurol. 2016;7:168. doi: 10.3389/fneur.2016.00168. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets analyzed in the present study are available in the GEO repository, https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE64977 and https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE64810.