Abstract
Mice are used universally as model organisms for studying heart physiology, and a plethora of genetically modified mouse models exist to study cardiac disease. Transcriptomic data for whole-heart tissue are available, but not yet for isolated ventricular cardiomyocytes. Our lab therefore collected comprehensive RNA-seq data from wildtype murine ventricular cardiomyocytes as well as from knockout models of the ion channel regulators CASK, dystrophin, and SAP97. We also elucidate ion channel expression from wild-type cells to help forward the debate about which ion channels are expressed in cardiomyocytes. Researchers studying the heart, and especially cardiac arrhythmias, may benefit from these cardiomyocyte-specific transcriptomic data to assess expression of genes of interest.
Subject terms: Gene expression, Mechanisms of disease
Background & Summary
In this study, we present next-generation RNA sequencing (RNA-seq) data of murine ventricular cardiomyocytes (CMC). To date, only whole-heart RNA-seq data have been published1–3, in which a variety of cell types, such as fibroblasts, endothelial cells, and atrial and ventricular cardiomyocytes, are pooled. We endeavoured to provide RNA-seq data of isolated CMCs for several reasons. Firstly, since the pump function of the heart relies on proper CMC function, CMCs are the most thoroughly studied cardiac cell type. Researchers studying CMCs may benefit from CMC-specific RNA-seq data from which expression of genes of interest can be extracted. Secondly, because of the crucial role of ion channels in cardiac electrical excitability and arrhythmogenesis, researchers that study cardiac arrhythmias have debated the question of which ion channels are expressed in CMCs. However, existing ion channel expression data are low-throughput, often contradictory4–6, fragmented7, or expression is assessed in the whole heart. The present work reveals the expression of the more than 350 ion channel family members, including pore-forming and auxiliary subunits, in CMCs (see Fig. 1 and Tables 1, Tables 2 and Tables 3 (available online only)). We therefore believe that these data will be valuable for ion channel researchers attempting to resolve the ongoing debate.
Figure 1. Gene expression of ion channels in murine ventricular cardiomyocytes.
(a) Expression levels of voltage-gated ion channel genes: voltage-gated sodium channels (Na+; purple), voltage-gated calcium channels (Ca2+; blue), transient receptor potential cation channels (TRP; light blue), CS, CatSper channels (aqua), two-pore channels (2P; green), cyclic-nucleotide-regulated channels (cN; light green), calcium-activated potassium channels (KCa; ochre), voltage-gated potassium channels (K+; orange), inwardly rectifying potassium channels (Kir; red) and two-pore potassium channels (2PK; burgundy). (b) Expression levels of the ligand-gated purinergic receptor gene (PR; purple) and of ion channel genes from the “other” category: aquaporins (Aqp; blue), voltage-sensitive chloride channels (Cl-; light blue), calcium-activated chloride channels (CaCl-; green) and inositol triphosphate receptors (IP3; light green). (c) Expression levels of more ion channel genes from the “other” category: ryanodine receptors (Ryr; orange), gap junction proteins (GJ; red) and chloride intracellular channels (icCl-; burgundy). All expression levels are average TPM values of WT samples (n=5). Shown are genes with more than 75 reads per gene (normalized for gene length, prior to conversion to TPM) from Tables 1, Table 2 and Table 3 (available online only).
Table 1. Expression of voltage-gated ion channels.
| Gene | Protein | TPM |
|---|---|---|
| Voltage-gated ion channel genes, their respective proteins, and transcript per million (TPM) values averaged from five WT samples. | ||
|
Sodium channels, voltage-gated |
||
| Scn5a | Nav1.5 | 86.900 |
| Scn4a | Nav1.4 | 12.163 |
| Scn7a | Nav2.1 | 1.650 |
| Scn10a | Nav1.8 | 0.180 |
| Scn3a | Nav1.3 | 0.123 |
| Scn2a | Nav1.2 | 0.025 |
| Scn11a | Nav1.9 | 0.009 |
| Scn8a | Nav1.6 | 0.004 |
| Scn1a | Nav1.1 | 0.003 |
| Scn9a | Nav1.7 | 0.000 |
| Scn4b | β4 subunit | 17.765 |
| Scn1b | β1 subunit | 6.524 |
| Scn3b | β3 subunit | 0.074 |
| Scn2b | β2 subunit | 0.041 |
|
Calcium channels voltage-gated |
||
| Cacna1c | Cav1.2 | 32.050 |
| Cacna1s | Cav1.1 | 3.045 |
| Cacna1g | Cav3.1 | 3.006 |
| Cacna1h | Cav3.2 | 1.738 |
| Cacna1a | Cav2.1 | 0.360 |
| Cacna1d | Cav1.3 | 0.225 |
| Cacna1b | Cav2.2 | 0.028 |
| Cacna1i | Cav3.3 | 0.008 |
| Cacna1e | Cav2.3 | 0.003 |
| Cacna1f | Cav1.4 | 0.000 |
| Cacna2d1 | α2 and δ1 subunit | 30.799 |
| Cacna2d3 | α2 and δ3 subunit | 0.048 |
| Cacna2d4 | α2 and δ4 subunit | 0.000 |
| Cacnb2 | β2 subunit | 8.023 |
| Cacnb1 | β1 subunit | 1.714 |
| Cacnb3 | β3 subunit | 0.398 |
| Cacnb4 | β4 subunit | 0.108 |
| Cacng6 | γ6 subunit | 1.513 |
| Cacna2d2 | γ1 subunit | 0.455 |
| Cacng7 | γ7 subunit | 0.261 |
| Cacng1 | γ7 subunit | 0.024 |
| Cacng4 | γ4 subunit | 0.010 |
| Cacng5 | γ5 subunit | 0.008 |
| Cacng2 | γ2 subunit | 0.002 |
| Cacng3 | γ3 subunit | 0.000 |
| Cacng8 | γ8 subunit | 0.000 |
|
Transient receptor potential cation channels |
||
| Trpm7 | TRPM7 | 5.530 |
| Pkd1 | TRPP1 | 5.248 |
| Pkd2l2 | TRPP5 | 4.380 |
| Mcoln1 | TRPML1 | 2.836 |
| Trpm4 | TRPM4 | 2.431 |
| Pkd2 | TRPP2 | 2.424 |
| Trpc1 | TRPC1 | 1.836 |
| Trpc3 | TRPC3 | 0.875 |
| Trpv2 | TRPV2 | 0.391 |
| Trpv4 | TRPV4 | 0.366 |
| Trpm3 | TRPM3 | 0.287 |
| Trpc6 | TRPC6 | 0.051 |
| Trpm6 | TRPM6 | 0.037 |
| Trpc4 | TRPC4 | 0.033 |
| Trpv1 | TRPV1 | 0.026 |
| Trpa1 | TRPA1 | 0.022 |
| Mcoln3 | TRPML3 | 0.018 |
| Trpv3 | TRPV3 | 0.016 |
| Mcoln2 | TRPML2 | 0.011 |
| Trpm2 | TRPM2 | 0.011 |
| Trpm5 | TRPM5 | 0.005 |
| Trpv5 | TRPV5 | 0.005 |
| Trpv6 | TRPV6 | 0.005 |
| Trpc7 | TRPC7 | 0.002 |
| Pkd2l1 | TRPP3 | 0.000 |
| Trpc2 | TRPC2 | 0.000 |
| Trpc5 | TRPC5 | 0.000 |
| Trpm1 | TRPM1 | 0.000 |
| Trpm8 | TRPM8 | 0.000 |
|
CatSper channels |
||
| Catsper2 | CATSPER2 | 1.744 |
| Catsper3 | CATSPER3, CACRC | 0.127 |
| Catsper4 | CATSPER4 | 0.060 |
| Catsperd | δ subunit | 0.058 |
| Catsperg1 | γ1 subunit | 0.031 |
| Catsperb | β subunit | 0.003 |
| Catsper1 | CATSPER1 | 0.000 |
| Catsperg2 | γ2 subunit | 0.000 |
|
Two-pore channels |
||
| Tpcn1 | TPCN1 | 4.183 |
| Tpcn2 | TPCN2 | 0.446 |
|
Cyclic nucleotide-regulated channels |
||
| Hcn2 | HCN2 | 9.736 |
| Hcn4 | HCN4 | 3.661 |
| Cnga3 | CNGA3 | 1.880 |
| Cngb3 | CNGB3 | 1.051 |
| Hcn1 | HCN1 | 0.229 |
| Cngb1 | CNGB1 | 0.067 |
| Hcn3 | HCN3 | 0.056 |
| Cnga1 | CNGA1 | 0.000 |
| Cnga2 | CNGA2 | 0.000 |
| Cnga4 | CNGA4 | 0.000 |
|
Potassium channels, calcium-activated |
||
| Kcnn1 | KCa2.1 | 5.677 |
| Kcnn2 | KCa2.2 | 3.107 |
| Kcnt2 | Kna | 0.287 |
| Kcnmb1 | β1 subunit | 0.127 |
| Kcnn3 | KCa2.3 | 0.043 |
| Kcnt1 | KCa4.1 | 0.019 |
| Kcnu1 | KCa5.1 | 0.014 |
| Kcnn4 | KCa3.1 | 0.008 |
| Kcnma1 | KCa1.1 | 0.007 |
| Kcnmb2 | β2 subunit | 0.000 |
| Kcnmb3 | β3 subunit | 0.000 |
| Kcnmb4 | β4 subunit | 0.000 |
|
Potassium channels, voltage-gated |
||
| Kcng2 | Kv6.2 | 44.470 |
| Kcnh2 | Kv11.1 | 17.558 |
| Kcnd2 | Kv4.2 | 12.960 |
| Kcnq1 | Kv7.1 | 12.079 |
| Kcnb1 | Kv2.1 | 8.310 |
| Kcna5 | Kv1.5 | 4.250 |
| Kcnv2 | Kv8.2 | 3.502 |
| Kcnd3 | Kv4.3 | 3.270 |
| Kcna7 | Kv1.7 | 2.800 |
| Kcne1 | KCNE1 | 1.917 |
| Kcnq4 | Kv7.4 | 1.326 |
| Kcna4 | Kv1.4 | 0.907 |
| Kcne4 | KCNE4 | 0.833 |
| Kcnf1 | Kv5.1 | 0.000 |
| Kcnc3 | Kv3.3 | 0.566 |
| Kcnc1 | Kv3.1 | 0.237 |
| Kcna1 | Kv1.1 | 0.175 |
| Kcne2 | KCNE2 | 0.005 |
| Kcna6 | Kv1.6 | 0.119 |
| Kcnab3 | KCAB3 | 0.113 |
| Kcna2 | Kv1.2 | 0.105 |
| Kcnab2 | KCAB2 | 0.088 |
| Kcnab1 | KCAB1 | 0.056 |
| Kcns1 | Kv9.1 | 0.051 |
| Kcnq5 | Kv7.5 | 0.048 |
| Kcnc4 | Kv3.4 | 0.041 |
| Kcnd1 | Kv4.1 | 0.031 |
| Kcnc2 | Kv3.2 | 0.029 |
| Kcnh1 | Kv10.1 | 0.027 |
| Kcng4 | Kv6.4 | 0.020 |
| Kcna3 | Kv1.3 | 0.019 |
| Kcns3 | Kv9.3 | 0.012 |
| Kcnh3 | Kv12.2 | 0.008 |
| Kcnh6 | Kv11.2 | 0.007 |
| Kcng3 | Kv6.3 | 0.005 |
| Kcne3 | KCNE3 | 0.060 |
| Kcnq3 | Kv7.3 | 0.004 |
| Kcnb2 | Kv2.2 | 0.003 |
| Kcnq2 | Kv7.2 | 0.002 |
| Kcnh8 | Kv12.1 | 0.002 |
| Kcna10 | Kv1.8 | 0.000 |
| Kcng1 | Kv6.1 | 0.000 |
| Kcnh4 | Kv12.3 | 0.000 |
| Kcnh5 | Kv10.2 | 0.000 |
| Kcnh7 | Kv11.3 | 0.000 |
| Kcns2 | Kv9.2 | 0.000 |
| Kcnv1 | Kv8.1 | 0.000 |
|
Potassium channels, inwardly rectifying |
||
| Abcc9 | SUR2A,SUR2B | 85.126 |
| Kcnj11 | Kir6.2 | 51.960 |
| Kcnj3 | Kir3.1 | 26.920 |
| Kcnj5 | Kir3.4 | 24.795 |
| Kcnj2 | Kir2.1 | 20.573 |
| Kcnj12 | Kir2.2 | 12.050 |
| Kcnj8 | Kir6.1 | 8.852 |
| Abcc8 | SUR1 | 8.225 |
| Kcnj14 | Kir2.4, Kir1.3 | 0.620 |
| Kcnj4 | Kir2.3 | 0.322 |
| Kcnj15 | Kir4.2 | 0.105 |
| Kcnj9 | Kir3.3 | 0.016 |
| Kcnj1 | Kir1.1 | 0.003 |
| Kcnj10 | Kir4.1, Kir1.2 | 0.002 |
| Kcnj13 | Kir7.1, Kir1.4 | 0.000 |
| Kcnj16 | Kir5.1 | 0.000 |
| Kcnj6 | Kir3.2 | 0.000 |
|
Potassium channels, two-P |
||
| Kcnk3 | K2P3.1, TASK-1 | 76.710 |
| Kcnk6 | K2P6.1, TWIK-2 | 0.580 |
| Kcnk1 | K2P1.1, TWIK-1 | 0.280 |
| Kcnk5 | K2P5.1, TASK-2 | 0.104 |
| Kcnk2 | K2P2.2, TREK-1 | 0.078 |
| zKcnk13 | K2P13.1, THIK-1 | 0.066 |
| Kcnk7 | K2P7.1 | 0.007 |
| Kcnk10 | K2P10.1, TREK-2 | 0.006 |
| Kcnk12 | K2P12.1, THIK-2 | 0.000 |
| Kcnk15 | K2P15.1, TASK-5 | 0.000 |
| Kcnk16 | K2P16.1, TASLK-1 | 0.000 |
| Kcnk18 | K2P18.1, TRESK-2 | 0.000 |
| Kcnk4 | K2P4.1, TRAAK | 0.000 |
| Kcnk9 | K2P9.1, TASK-3 | 0.000 |
|
Hydrogen voltage-gated ion channels |
||
| Hvcn1 | Hv1 | 0.138 |
Table 2. Expression of ligand-gated ion channels.
| Gene | Protein | TPM |
|---|---|---|
| Ligand-gated ion channel genes, their respective proteins, and transcript per million (TPM) values averaged from five WT samples. | ||
|
5-HT (serotonin) receptors, ionotropic |
||
| Htr3a | 5-HT3A | 0.023 |
| Htr3b | 5-HT3B | 0.000 |
|
Acetylcholine receptors, nicotinic |
||
| Chrna2 | ACHA2 | 1.073 |
| Chrnb1 | ACHB | 0.668 |
| Chrnb2 | ACHB2 | 0.066 |
| Chrng | ACHG | 0.028 |
| Chrna10 | ACH10 | 0.019 |
| Chrna1 | ACHA | 0.014 |
| Chrne | ACHE | 0.007 |
| Chrna5 | ACHA5 | 0.003 |
| Chrna3 | ACHA3 | 0.000 |
| Chrna4 | ACHA4 | 0.000 |
| Chrna6 | ACHA6 | 0.000 |
| Chrna7 | ACHA7 | 0.000 |
| Chrna9 | ACHA9 | 0.000 |
| Chrnb3 | ACHB3 | 0.000 |
| Chrnb4 | ACHB4 | 0.000 |
| Chrnd | ACHD | 0.000 |
|
GABA(A) receptors |
||
| Gabrr2 | GBRR2 | 0.855 |
| Gabra3 | GBRA3 | 0.155 |
| Gabre | GBRE | 0.102 |
| Gabrb3 | GBRB3 | 0.058 |
| Gabrq | GBRT | 0.022 |
| Gabrg3 | GBRG3 | 0.021 |
| Gabrb2 | GBRB2 | 0.019 |
| Gabra2 | GBRA2 | 0.006 |
| Gabrd | GBRD | 0.004 |
| Gabra5 | GBRA5 | 0.004 |
| Gabrg1 | GBRG1 | 0.002 |
| Gabra4 | GBRA4 | 0.002 |
| Gabra1 | GBRA1 | 0.002 |
| Gabra6 | GBRA6 | 0.000 |
| Gabrb1 | GBRB1 | 0.000 |
| Gabrg2 | GBRB2 | 0.000 |
| Gabrp | GBRP | 0.000 |
| Gabrr1 | GBRR1 | 0.000 |
| Gabrr3 | GBRR3 | 0.000 |
|
Glutamate receptors, ionotropic |
||
| Grik5 | GRIK5 | 0.830 |
| Grin2c | NMDE3 | 0.407 |
| Grin3b | NMD3B | 0.151 |
| Gria3 | GRIA3 | 0.148 |
| Grin2d | NMDE4 | 0.106 |
| Gria1 | GRIA1 | 0.022 |
| Grik4 | GRIK4 | 0.018 |
| Grik3 | GRIK3 | 0.016 |
| Grik2 | GRIK2 | 0.010 |
| Grin3a | NMD3A | 0.008 |
| Grin2a | NMDE1 | 0.006 |
| Gria4 | GRIA4 | 0.005 |
| Grid2 | GRID2 | 0.003 |
| Gria2 | GRIA2 | 0.002 |
| Grid1 | GRID1 | 0.001 |
| Grik1 | GRIK1 | 0.001 |
| Grin1 | NMDZ1 | 0.001 |
| Grin2b | NMDE2 | 0.000 |
|
Glycine receptors |
||
| Glra4 | GLRA4 | 0.045 |
| Glra1 | GLRA1 | 0.000 |
| Glra2 | GLRA2 | 0.000 |
| Glra3 | GLRA3 | 0.000 |
|
Purinergic receptors, ionotropic |
||
| P2rx5 | P2X5 | 8.390 |
| P2rx4 | P2X4 | 2.570 |
| P2rx6 | P2X6 | 1.069 |
| P2rx7 | P2X7 | 0.349 |
| P2rx3 | P2X3 | 0.191 |
| P2rx1 | P2X1 | 0.065 |
| P2rx2 | P2X2 | 0.010 |
|
Zinc-activated channels |
||
| not expressed in mice |
Table 3. Expression of other ion channels.
| Gene | Protein | TPM |
|---|---|---|
| Other ion channel genes, their respective proteins, and transcript per million (TPM) values averaged from five WT samples. | ||
|
Acid-sensing (proton-gated) ion channels |
||
| Asic3 | ASIC3 | 0.068 |
| Asic1 | ASIC1 | 0.053 |
| Asic4 | ASIC4 | 0.009 |
| Asic2 | ASIC2 | 0.000 |
|
Aquaporins |
||
| Aqp1 | AQP1 | 82.978 |
| Aqp7 | AQP7 | 5.290 |
| Aqp8 | AQP8 | 4.274 |
| Aqp4 | AQP4 | 2.601 |
| Aqp11 | AQP11 | 0.154 |
| Aqp6 | AQP6 | 0.047 |
| Aqp2 | AQP2 | 0.043 |
| Aqp5 | AQP5 | 0.018 |
| Aqp9 | AQP9 | 0.007 |
| Aqp12 | AQP12 | 0.000 |
| Aqp3 | AQP3 | 0.000 |
| Mip | MIP | 0.000 |
|
Chloride channels, voltage-sensitive |
||
| Clcn4 | CLCN4 | 15.442 |
| Clcn7 | CLCN7 | 8.289 |
| Clcn3 | CLCN3 | 5.871 |
| Clcn6 | CLCN6 | 2.146 |
| Clcn1 | CLCN1 | 1.737 |
| Clcn2 | CLCN2 | 0.667 |
| Clcnkb | CLCKB | 0.550 |
| Clcn5 | CLCN5 | 0.216 |
| Clcnka | CLCKA | 0.003 |
|
Cystic fibrosis transmembrane conductance regulators |
||
| Cftr | CFTR | 0.006 |
|
Calcium-activated chloride channels |
||
| Ano10 | ANO10 | 12.374 |
| Ano8 | ANO8 | 4.659 |
| Best3 | BEST3 | 4.218 |
| Ano4 | ANO4 | 1.961 |
| Ano1 | ANO1 | 1.565 |
| Ano5 | ANO5 | 1.432 |
| Ano6 | ANO6 | 1.228 |
| Ano3 | ANO3 | 0.033 |
| Best1 | BEST1 | 0.016 |
| Ano9 | ANO9 | 0.011 |
| Best2 | BEST2 | 0.006 |
| Ano2 | ANO2 | 0.000 |
| Ano7 | ANO7 | 0.000 |
|
Chloride intracellular channels |
||
| Clic4 | CLIC4 | 72.409 |
| Clic5 | CLIC5 | 65.201 |
| Clic1 | CLIC1 | 10.695 |
| Clic3 | CLIC3 | 0.398 |
| Clic6 | CLIC6 | 0.039 |
|
Gap junction proteins |
||
| Gja1 | CXA1 | 82.934 |
| Gja3 | CXA3 | 7.013 |
| Gjc1 | CXG1 | 2.952 |
| Gja4 | CXA4 | 1.946 |
| Gja5 | CXA5 | 0.613 |
| Gja6 | CXA6 | 0.147 |
| Gjc2 | CXG2 | 0.095 |
| Gjd3 | CXD3 | 0.069 |
| Gjb5 | CXB5 | 0.019 |
| Gjc3 | CXG3 | 0.015 |
| Gjb2 | CXB2 | 0.005 |
| Gja10 | CXA10 | 0.000 |
| Gja8 | CXA8 | 0.000 |
| Gjb1 | CXB1 | 0.000 |
| Gjb3 | CXB3 | 0.000 |
| Gjb4 | CXB4 | 0.000 |
| Gjb6 | CXB6 | 0.000 |
| Gjd2 | CXD2 | 0.000 |
| Gjd4 | CXD4 | 0.000 |
| Gje1 | GJE1 | 0.000 |
|
IP3 receptors |
||
| Itpr1 | ITPR1 | 4.382 |
| Itpr2 | ITPR2 | 1.929 |
| Itpr3 | ITPR3 | 0.798 |
|
Pannexins |
||
| Panx2 | PANX2 | 0.207 |
| Panx1 | PANX1 | 0.137 |
| Panx3 | PANX3 | 0.000 |
|
Ryanodine receptors |
||
| Ryr2 | RYR2 | 225.160 |
| Ryr3 | RYR3 | 0.220 |
| Ryr1 | RYR1 | 0.047 |
|
Sodium leak channels, non-selective |
||
| Nalcn | NALCN | 0.049 |
|
Sodium channels, non-voltage-gated |
||
| Scnn1a | SCNNA | 0.077 |
| Scnn1b | SCNNB | 0.000 |
| Scnn1g | SCNNG | 0.000 |
We have also included cardiac-specific knockout models of the ion channel regulators dystrophin, synapse-associated protein-97 (SAP97), and calmodulin-activated serine kinase (CASK). They interact with ion channels and modify their cell biological properties, such as membrane localization3,8–11. Notably, CASK provides a direct link between ion channel function and gene expression. It regulates transcription factors (TFs) in the nucleus, such as Tbr-1, and induces transcription of T-element-containing genes12. CASK also regulates TFs of the basic helix-loop-helix family, which bind E-box elements in promoter regions, by modulating the inhibitor of the DNA-binding-1 TF13. Additionally, CASK and SAP97 directly interact with each other11. For these reasons, we include CASK, SAP97, and dystrophin knockout mice to investigate whether these three proteins have a similar effect on gene expression, which may suggest their involvement in similar pathways. However, research beyond the scope of this paper would be needed to determine whether CASK-dependent TF regulation caused the differential expression that we observed.
To date, mutations in approximately 27 ion channel genes have been associated with cardiac arrhythmias, such as congenital short- and long-QT syndrome (SQTS and LQTS), Brugada syndrome (BrS), and conduction disorders (see http://omim.org)14–16. Notably, our ion channel expression data, as presented in Fig. 1 and Tables 1, Table 2 and Table 3 (available online only), reveal that several arrhythmia-associated ion channel genes are not or are scarcely expressed in murine ventricular CMCs (including Kcne2, Kcne3, Scn2b, and Scn3b). Although murine and human ion channel expression may differ, we are presently unaware of any available transcriptome of human CMCs17,18. We are also unable to either exclude or assess the effect of enzymatic isolation on the transcriptome. Finally, other cardiac cell types such as (myo)fibroblasts may express these ion channels and therefore may be important for arrhythmogenesis. Indeed, many ion channel genes that are not expressed in cardiomyocytes have been reported in murine whole-heart tissue2. These include Scn1a, Scn3b, 10 voltage-gated Ca2+ channels, 10 Kv channels, and four two-pore K+ channels. Conversely, all ion channel genes expressed in CMCs are also reported in whole-heart expression data.
In sum, this study presents RNA-seq data from wildtype murine ventricular CMCs, as well as from SAP97, CASK, and dystrophin knockouts and controls (see Fig. 2 for a schematic overview of study design). We performed differential gene expression analysis to compare the knockouts to their controls, and we extracted wildtype ion channel gene expression data (Tables 1, Table 2 and Table 3 (available online only), Fig. 1). We believe that these data will be valuable for researchers studying cardiomyocytes and ion channels to assess expression of genes of interest.
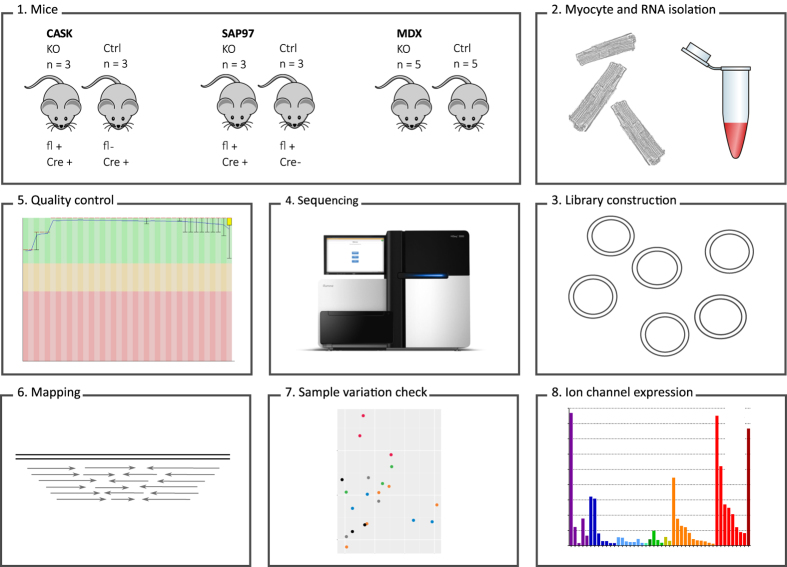
Figure 2. Experimental design and workflow.
(1) 22 mice with six different genetic backgrounds (CASK KO and control, SAP97 KO and control, and MDX and control) were used. fl+, first exon of gene is floxed; Cre+, Cre recombinase is expressed. (2) Cardiomyocytes were isolated on a Langendorff system and RNA was isolated with a FFPE Clear RNAready kit. (3) Libraries were constructed with 1 μg RNA per sample using a TrueSeq Stranded Total RNA protocol and (4) sequenced on an Illumina HiSeq3000 machine. (5) Quality of the reads was assessed with FastQC, and (6) reads were mapped to the Mus musculus reference genome (GRCm38.83) with Tophat. (7) To assess sample variation within each group, we performed principle component analyses (PCA) (see Fig. 3). (8) Lastly, ion channel expression was determined.
Methods
Mouse models
All animal experiments conformed to the Guide to the Care and Use of Laboratory Animals (US National Institutes of Health, publication No. 85-23, revised 1996); have been approved by the Cantonal Veterinary Administration, Bern, Switzerland; and have complied with the Swiss Federal Animal Protection Law. Mice were kept on a 12-hour light/dark cycle. Lights were on from 6:30 AM to 6:30 PM. To avoid the influence of circadian rhythm, mice were sacrificed between 10:00 AM and 1:00 PM. Mice were all male and were between the ages of 8 and 15 weeks.
MHC-Cre
The cardiac-specific murine alpha-myosin heavy chain (μMHC) promoter drives the expression of Cre recombinase, which, in turn, can recombine LoxP sequences. The μMHC-Cre strain was generated as previously described19 and acquired from the Jackson Laboratory (stock #011038).
CASK and SAP97 knockout mice
CASK KO and SAP97/Dlg1 KO mice were generated as previously described9,20. Both the CASK and SAP97 mouse lines were on mixed backgrounds. The appropriate control mice were selected in accordance with the publications that characterized both mouse lines9,20. CASK control mice express Cre while the first CASK exon is not floxed. SAP97 control mice are Cre-negative and the first SAP97 gene was floxed.
Dystrophin knockout (MDX-5CV) mice
The MDX-5CV strain demonstrates total deletion of the dystrophin protein. It was created as previously described21, and acquired from the Jackson laboratory (stock #002379). MDX mice were on pure Bl6/Ros backgrounds. Control mice were on pure Bl6/J background, except for MDX_Ct5 and MDX_5, which were Bl6/Ros mice backcrossed three times on Bl6/J.
Cardiomyocyte isolation
Mice (n=3–5 per genotype, male, age 10–15 weeks) were heparinized (intraperitoneal injection of 100 μL heparin (5000 U/mL; Biochrom AG)) and killed by cervical dislocation. Hearts were excised, and the aortas were cannulated in ice-cold phosphate-buffered saline (PBS). Subsequently, hearts were perfused on a Langendorff system in a retrograde manner at 37 °C with 5 mL perfusion buffer (1.5 mL/min; in mM: 135 NaCl, 4 KCl, 1.2 NaH2PO4, 1.2 MgCl2, 10 HEPES, 11 glucose), followed by the application of type II collagenase (Worthington CLS2; 25 mL of 1 mg/mL in perfusion buffer with 50 μM CaCl2). Left and right ventricles were triturated in PBS to dissociate individual ventricular cardiomyocytes and then filtered through a 100 μm filter.
RNA extraction and sequencing
RNA-seq was performed by the Next Generation Sequencing Platform at the University of Bern. Total RNA was isolated from freshly dissociated cardiomyocytes with an FFPE Clear RNAready kit (AmpTec, Germany), which included a DNase treatment step. RNA quality was assessed with Qubit and Bioanalyzer, and RNA quantity was checked with Qubit.
To allow sequencing of long non-coding RNA (lncRNA), libraries were constructed with 1 μg RNA using the TruSeq Stranded Total RNA kit after Ribo-Zero Gold (Illumina) treatment for rRNA depletion. Library molecules with inserts <300 base pairs (bp) were removed. Paired-end libraries (2x150 bp) were sequenced on an Illumina HiSeq3000 machine.
RNA-seq data analysis
Between 17.5 and 56.4 million read pairs were obtained per sample and the quality of the reads was assessed using FastQC v.0.11.2 (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/). Ribosomal RNA (rRNA) was removed by mapping the reads with Bowtie2 v.2.2.1 (ref. 22) to a collection of rRNA sequences (NR_003279.1, NR_003278.3 and NR_003280.2) downloaded from NCBI (www.ncbi.nlm.nih.gov). No quality trimming was required.The remaining reads were mapped to the Mus musculus reference genome (GRCm38.83) with Tophat v.2.0.13 (ref. 23). We used htseq-count v.0.6.1 (ref. 24) to count the number of reads overlapping with each gene, as specified in the Ensembl annotation (GRCm38.83). Detailed information about the genes including the Entrez Gene ID, the MGI symbol and the description of the gene was obtained using the Bioconductor package BioMart v.2.26.1 (ref. 25).
Raw reads were corrected for gene length and TPM (transcripts per million) values were calculated to compare the expression levels among samples. Gene lengths for the latter step were retrieved from the Ensembl annotation (GRCm38.83) as the total sum of all exons.
Principal component analysis (PCA) plots were done in DESeq2 v.1.10.1 (ref. 26) (https://bioconductor.org/packages/release/bioc/html/DESeq2.html) using the 500 genes with the most variable expression across samples. A regularized log transformation was applied to the counts before performing the PCA.
Statistics
To assess differential gene expression between genotypes, a Wald test was performed with the Bioconductor package DESeq2 v.1.10.1 (ref. 26). We considered p values of up to 0.01, accounting for a Benjamini-Hochberg false discovery rate adjustment, to indicate significant difference. Statistical tools used included DESeq2, R-3.2.5 (https://cran.r-project.org), and Biomart_2.26.1 (www.biomart.org).
Data Records
The data were submitted to NCBI Gene Expression Omnibus (GEO) (Data Citation 1). This GEO project contains raw data and TPM values from all samples, and differential gene expression analysis between knockout and control samples.
Technical Validation
RNA metrics
RNA-seq yielded 1.0 billion read pairs in total, with an average of 44.5 million read pairs per sample (standard deviation 8.4 million). The number of read pairs (in millions) was 306 for CASK KO and Ctrl, 268 for SAP97 KO and Ctrl, and 404 for MDX and Ctrl (see Table 4 for an overview of RNA-seq metrics, including mapping rates). One sample (MDX_1) yielded few reads and was therefore excluded from further analyses. The proportion of reads mapping to annotated exons ranged from 65 to 77%. Mapping, no-feature (2–13%), and ambiguous (11–23%) read pairs together accounted for 89–97% of the total number of RNA reads (Table 4). Read pairs covered 49,671 genes of the Mus musculus reference genome (GRCm83.38).
Table 4. RNA-seq raw data and mapping metrics.
| Sample ID | Genotype | # read pairs total | # non-rRNA read pairs | % of total | Insert size | # read pairs mapping to a gene | % of total | # no-feature read pairs | % of total | # ambiguous read pairs | % of total |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Total and non-ribosomal RNA read pairs, average RNA fragment size (bp), and mapping metrics, including absolute number and percentages of read pairs mapping to all annotated exons of the mouse reference genome, and no-feature and ambiguous reads, per sample. Note the low number of read pairs in MDX_1, which is therefore excluded from further analysis. CASK KO and Ctrl, SAP97 KO and Ctrl ns=3, MDX KO n=4, MDX Ctrl n=5. | |||||||||||
| CASK_Ct1 | WT+Cre | 47,543,799 | 47,343,548 | 99.58 | 492 | 34,076,980 | 71.67 | 2,721,942 | 5.73 | 8,287,647 | 17.43 |
| CASK_Ct2 | WT+Cre | 45,437,500 | 45,287,356 | 99.67 | 476 | 33,988,592 | 74.8 | 1,440,229 | 3.17 | 7,578,641 | 16.68 |
| CASK_Ct3 | WT+Cre | 55,117,414 | 54,944,721 | 99.69 | 479 | 40,469,641 | 73.42 | 1,790,829 | 3.25 | 11,381,005 | 20.65 |
| CASK_KO1 | CASK_fl+Cre | 45,685,573 | 45,565,815 | 99.74 | 472 | 32,765,612 | 71.72 | 5,738,568 | 12.56 | 5,504,670 | 12.05 |
| CASK_KO2 | CASK_fl+Cre | 55,895,769 | 55,607,105 | 99.48 | 511 | 39,344,558 | 70.39 | 2,372,238 | 4.24 | 12,476,403 | 22.32 |
| CASK_KO3 | CASK_fl+Cre | 56,437,329 | 56,008,449 | 99.24 | 499 | 42,159,185 | 74.7 | 2,804,256 | 4.97 | 9,655,232 | 17.11 |
| MDX_1 | MDX | 17,485,935 | 17,320,513 | 99.05 | 380 | (sample exluded) | |||||
| MDX_2 | MDX | 39,536,744 | 39,037,330 | 98.74 | 447 | 27,475,113 | 69.49 | 3,258,092 | 8.24 | 6,543,268 | 16.55 |
| MDX_3 | MDX | 39,626,959 | 39,432,254 | 99.51 | 455 | 27,841,146 | 70.26 | 2,169,924 | 5.48 | 7,327,584 | 18.49 |
| MDX_4 | MDX | 42,406,246 | 40,919,896 | 96.49 | 488 | 29,805,158 | 70.28 | 2,905,415 | 6.85 | 6,497,990 | 15.32 |
| MDX_5 | MDX | 50,934,677 | 47,518,076 | 93.29 | 484 | 34,233,480 | 67.21 | 1,864,210 | 3.66 | 10,028,518 | 19.69 |
| MDX_Ct1 | WT | 48,311,563 | 46,288,106 | 95.81 | 380 | 32,827,800 | 67.95 | 4,353,181 | 9.01 | 7,779,264 | 16.1 |
| MDX_Ct2 | WT | 47,283,192 | 46,988,962 | 99.38 | 446 | 32,237,279 | 68.18 | 2,304,142 | 4.87 | 10,939,883 | 23.14 |
| MDX_Ct3 | WT | 35,275,617 | 34,938,284 | 99.04 | 427 | 24,235,276 | 68.7 | 3,631,537 | 10.29 | 4,922,208 | 13.95 |
| MDX_Ct4 | WT | 33,977,175 | 32,900,815 | 96.83 | 515 | 25,298,933 | 74.46 | 1,713,065 | 5.04 | 4,619,558 | 13.6 |
| MDX_Ct5 | WT | 49,379,536 | 45,499,492 | 92.14 | 485 | 32,227,570 | 65.27 | 1,210,708 | 2.45 | 10,698,976 | 21.67 |
| SAP_Ct1 | WT+Cre | 47,930,112 | 47,715,719 | 99.55 | 461 | 34,192,649 | 71.34 | 1,965,652 | 4.1 | 9,896,590 | 20.65 |
| SAP_Ct2 | WT+Cre | 44,934,245 | 44,566,395 | 99.18 | 444 | 30,350,071 | 67.54 | 4,879,732 | 10.86 | 7,491,483 | 16.67 |
| SAP_Ct3 | WT+Cre | 43,586,968 | 43,382,766 | 99.53 | 451 | 29,836,839 | 68.45 | 1,332,267 | 3.06 | 8,751,881 | 20.08 |
| SAP_KO1 | SAP_fl+Cre | 44,319,566 | 44,146,526 | 99.61 | 452 | 34,155,235 | 77.07 | 2,606,959 | 5.88 | 5,090,692 | 11.49 |
| SAP_KO2 | SAP_fl+Cre | 41,547,517 | 41,397,099 | 99.64 | 469 | 28,697,320 | 69.07 | 3,765,768 | 9.06 | 7,431,842 | 17.89 |
| SAP_KO3 | SAP_fl+Cre | 46,143,349 | 45,812,985 | 99.28 | 443 | 30,710,635 | 66.55 | 5,476,238 | 11.87 | 8,174,538 | 17.72 |
Quality assessment
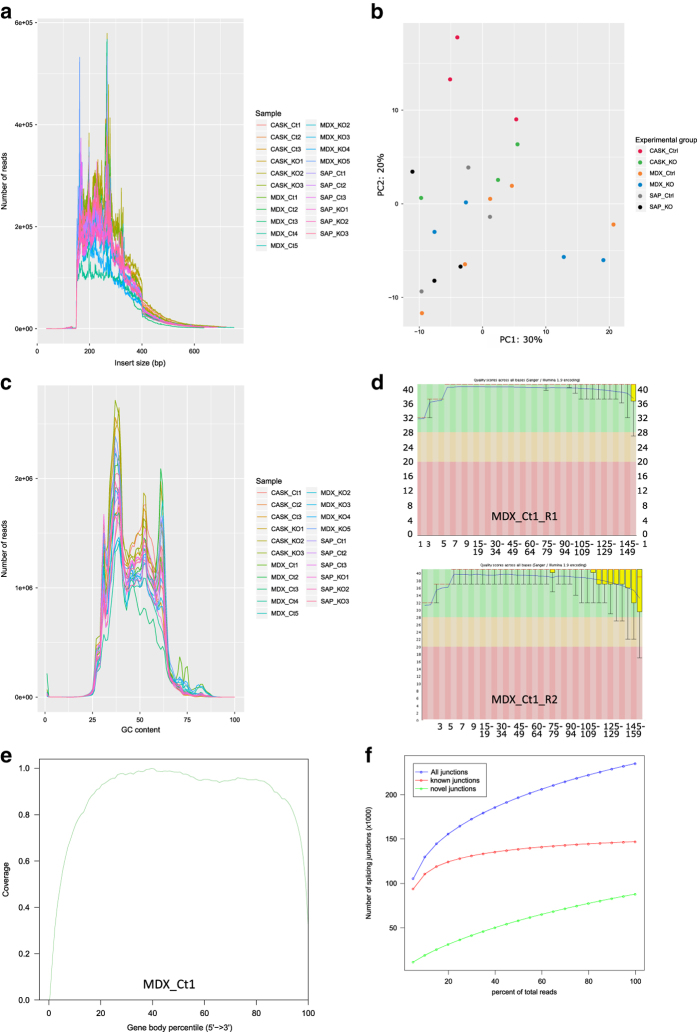
The quality of all samples was assessed with FastQC. Except for MDX_1, all samples were of high quality. Where applicable, a representative example (MDX_Ct1) is shown. Firstly, the insert size histogram (Fig. 3a) shows that the inferred insert size of each sample exceeded 150 base pairs, demonstating that the sequencing was not contaminated by adapter sequences. Secondly, the GC content plot (Fig. 3c) ideally shows a roughly normal distribution centred around the average GC content of the genome, which varies between species. The peaks observed in Fig. 3c are likely caused by sequences that are detected at high copy numbers, and should not pose problems for downstream analyses. Furthermore, Phred scores (Fig. 3d) are well within the green area of the graph indicating good base quality along the length of reads. As well, the gene coverage graph (Fig. 3e) of sample MDX_Ct1 shows that reads are distributed evenly along the length of the gene body. Because the gene coverage for all other samples is highly comparable to that of MDX_Ct1, only one example is shown. Lastly, the saturation report (Fig. 3f) represents the number of splice junctions detected using different subsets of the data from 5 to 100% of all reads. At sequencing depths sufficient to perform alternative splicing analysis, at least the red line, representing known junctions, should reach a plateau where adding more data does not much increase the number of detected junctions. Only MDX_1 does not reach this plateau.
Figure 3. Quality control.
(a) Histogram of inferred insert size for each sample, which represents distance between the two reads of one RNA fragment. (b) Principle component analyses (PCA) plots were performed to assess variability of samples within and between groups. Plot of the first two axes from a PCA based on the 500 genes with the most variable expression across all samples except MDX_1. CASK control (red, n=3) and KO (green, n=3); MDX control (orange, n=5) and KO (blue, n=4); SAP97 control (grey, n=3) and KO (black, n=3). (c) Distribution of GC content of the reads for each sample. (d) Base quality (Phred scores) along the length of the reads in each FastQC file of MDX_Ct1 as representative sample. The box plots are drawn as follows: red line, median; yellow box, range between upper and lower quartiles; whiskers, range between 10 and 90% quantiles. The blue line shows the mean quality. Y-axis represents quality scores across all bases. X-axis represents position in read (bp). (e) Gene body coverage. Distribution of reads along the length of the genes (5’-end on the left, 3’-end on the right). Shown image of sample MDX_Ct1 is representative for all samples. (f) Saturation report, depicting the number of splice junctions detected using different subsets of the data from 5 to 100% of all reads. Red, known junction based on the provided genome annotation; green, novel junctions; blue, all junctions. The red line reaches a plateau where adding more data does not increase the number of detected junctions, indicating that the sequencing depth suffices for performing alternative splicing analysis.
Gene expression variation of biological replicates
We performed Principle Component Analyses (PCA) to assess whether samples from the same experimental group have similar gene expression profiles (Fig. 3b). Of note, samples within each sample group still show considerable variation. The mixed genetic background of most sample groups may explain this variation; only the MDX control mice are on a pure Bl6/J background. The variation seen in MDX control mice is likely due to a batch effect, as two rounds of samples were sequenced. However, considering that PCA plots are based on the 500 genes with the highest variability in one sample, our genes of interest, including all ion channel genes, show similar expression levels throughout all samples.
Ion channel expression
Based on the list of ion channel genes from HUGO Gene Nomenclature Committee (https://www.genenames.org/cgi-bin/genefamilies/set/177), we distilled ion channel expression from WT mice expressed as TPM (Tables 1, Table 2 and Table 3 (available online only), Fig. 1).
Additional information
How to cite this article: Chevalier, M. et al, Transcriptomic analyses of murine ventricular cardiomyocytes. Sci. Data 5:180170 doi: 10.1038/sdata.2018.170 (2018).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Material
Acknowledgments
The research was financially supported by Swiss National Science Foundation Grant 310030_165741 (HA). We thank the Next Generation Sequencing Platform of the University of Bern, in particular Muriel Fragnière, for performing the high-throughput sequencing experiments; prof. Rolf Jaggi for excellent technical advice on RNA isolation from cardiomyocytes; and Joseph Allan for language editing.
Footnotes
The authors declare no competing interests.
Data Citations
- 2017. Gene Expression Omnibus. GSE102772
References
- Cerrone M. et al. Plakophilin-2 is required for transcription of genes that control calcium cycling and cardiac rhythm. Nat Commun 8, 106 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harrell M. D., Harbi S., Hoffman J. F., Zavadil J. & Coetzee W. A. Large-scale analysis of ion channel gene expression in the mouse heart during perinatal development. Physiol. Genomics 28, 273–283 (2007). [DOI] [PubMed] [Google Scholar]
- Petitprez S. et al. SAP97 and dystrophin macromolecular complexes determine two pools of cardiac sodium channels Nav1.5 in cardiomyocytes. Circ. Res. 108, 294–304 (2011). [DOI] [PubMed] [Google Scholar]
- Zimmer T., Haufe V. & Blechschmidt S. Voltage-gated sodium channels in the mammalian heart. Glob. Cardiol. Sci. Pract 4, 449–463 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Balse E. et al. Dynamic of ion channel expression at the plasma membrane of cardiomyocytes. Physiol. Rev. 92, 1317–1358 (2012). [DOI] [PubMed] [Google Scholar]
- Westenbroek R. E. et al. Localization of sodium channel subtypes in mouse ventricular myocytes using quantitative immunocytochemistry. J. Mol. Cell. Cardiol. 64, 69–78 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stroud D. M. et al. Contrasting Nav1.8 Activity in Scn10a-/- Ventricular Myocytes and the Intact Heart. J. Am. Heart Assoc 5, 1–10 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leonoudakis D., Conti L. R., Radeke C. M., McGuire L. M. & Vandenberg C. A. A multiprotein trafficking complex composed of SAP97, CASK, Veli, and Mint1 is associated with inward rectifier Kir2 potassium channels. J. Biol. Chem. 279, 19051–19063 (2004). [DOI] [PubMed] [Google Scholar]
- Eichel C. A. et al. Lateral membrane-specific MAGUK CASK down-regulates NaV1.5 channel in cardiac myocytes. Circ. Res. 119, 544–556 (2016). [DOI] [PubMed] [Google Scholar]
- Abriel H., Rougier J. S. & Jalife J. Ion channel macromolecular complexes in cardiomyocytes: roles in sudden cardiac death. Circ. Res. 116, 1971–1988 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nafzger S. & Rougier J. S. Calcium/calmodulin-dependent serine protein kinase CASK modulates the L-type calcium current. Cell Calcium 61, 10–21 (2017). [DOI] [PubMed] [Google Scholar]
- Hsueh Y. P., Wang T. F., Yang F. C. & Sheng M. Nuclear translocation and transcription regulation by the membrane-associated guanylate kinase CASK/LIN-2. Nature 404, 298–302 (2000). [DOI] [PubMed] [Google Scholar]
- Sun R. et al. Human calcium/calmodulin-dependent serine protein kinase regulates the expression of p21 via the E2A transcription factor. Biochem. J. 419, 457–466 (2009). [DOI] [PubMed] [Google Scholar]
- Ashcroft F. M. From molecule to malady. Nature 440, 440–447 (2006). [DOI] [PubMed] [Google Scholar]
- Rickert-Sperling S., Kelly R. G. & Driscoll D. J. Congenital Heart Diseases: The Broken Heart 721–736 (Springer-Verlag, 2016). [Google Scholar]
- Lahrouchi N., Behr E. R. & Bezzina C. R. Next-generation sequencing in post-mortem genetic testing of young sudden cardiac death cases. Front. Cardiovasc. Med 3, 13 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heinig M. A. et al. Natural genetic variation of the cardiac transcriptome in non-diseased donors and patients with dilated cardiomyopathy. Genome Biol. 18, 170 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- van den Berg C. W. et al. Transcriptome of human foetal heart compared with cardiomyocytes from pluripotent stem cells. Development 142, 3231–3238 (2015). [DOI] [PubMed] [Google Scholar]
- Agah R. et al. Gene recombination in postmitotic cells. Targeted expression of Cre recombinase provokes cardiac-restricted, site-specific rearrangement in adult ventricular muscle in vivo. J. Clin. Invest. 100, 169–179 (1997). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gillet L. et al. Cardiac-specific ablation of synapse-associated protein SAP97 in mice decreases potassium currents but not sodium current. Heart Rhythm 12, 181–192 (2015). [DOI] [PubMed] [Google Scholar]
- Chapman V. M., Miller D. R., Armstrong D. & Caskey C. T. Recovery of induced mutations for X chromosome-linked muscular dystrophy in mice. Proc. Natl. Acad. Sci. USA 86, 1292–1296 (1989). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langmead B. & Salzberg S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anders S., Pyl P. T. & Huber W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Durinck S. et al. BioMart and Bioconductor: a powerful link between biological databases and microarray data analysis. Bioinformatics 21, 3439–3440 (2005). [DOI] [PubMed] [Google Scholar]
- Love M. I., Huber W. & Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Citations
- 2017. Gene Expression Omnibus. GSE102772