Abstract
Background
Dengue, an acute febrile illness caused by infection with dengue virus (DENV), is endemic in Bali, Indonesia. As one of the world’s most popular tourist destinations, Bali is regularly visited by domestic and international travellers, who are prone to infection by endemic pathogens, including DENV. Currently, limited data are available on the characteristics of dengue in travellers visiting Bali. Information on the epidemiology and virological aspects of dengue in these tourists is important to gain a better understanding of the dengue disease in international travellers.
Methods
We performed a prospective cross-sectional dengue study involving foreign travellers visiting Bali, Indonesia in the period of 2015–17. Patients presenting at Kasih Ibu Hospital with fever and clinical symptoms of dengue were asked to participate in the study. Clinical and laboratory assessments were performed and sera were collected for molecular analysis, which included DENV serotyping, genome sequencing and phylogenetic analysis.
Results
Among the 201 patients recruited, dengue was confirmed in 133 (66.2%) of them, based on detection of NS1 antigen and/or viral RNA. Of these, 115 (86.5%) manifested dengue fever (DF) and 18 (13.5%) dengue haemorrhagic fever (DHF). The temporal predominance of infecting DENV serotype was DENV-2 (48.7%), followed by DENV-3 (36.1%), DENV-1 (9.2%) and DENV-4 (3.4%). Phylogenetic analysis of DENV based on envelope gene sequences revealed that the source of DENVs was local endemic viruses.
Conclusion
Our study confirms that dengue is one of the causes of fever in travellers visiting Bali. Although it is a cause of significant morbidity, the majority of patients only experienced mild DF, with only a small proportion developing DHF. We revealed that DENVs isolated were autochthonous. Accurate diagnosis, preventive measures and continuous disease surveillance will be useful for better management of dengue infection in travellers.
Keywords: Dengue, serotype, genotype, travellers, Bali
Introduction
Dengue disease poses a significant public health challenge, with a global burden of 390 million infections occurring annually, with 70% of the world population at risk.1 Four DENV serotypes (DENV-1, -2, -3 and -4) circulate in tropical and subtropical regions of the world and are transmitted by the Aedes mosquito vectors.2 DENV shows very high genetic diversity, as demonstrated by the presence of four serotypes, within which there are several clusters of variants, termed genotypes.3 Dengue can be asymptomatic or clinically manifested as mild flu-like syndrome, known as classic Dengue Fever (DF), or in a more severe form known as Dengue Haemorrhagic Fever (DHF), to the potentially fatal Dengue Shock Syndrome (DSS).4
The evolution of global travel patterns characterised by dramatic increases in travel primarily those from emerging economies5 is found to translate into different geographic dispersion patterns of infectious diseases.6 These include DF, where more tourists visit dengue endemic countries7 and the consequence is an increase in the number of cases in returning travellers.8 While travel-related morbidity data are available, most of the data acquisition is still mainly driven by ‘Western countries’.6 It is increasingly important to conduct research not only in the receiving countries but also in the countries of acquisition where dengue is endemic7 including Indonesia where there is lack of data on exported dengue.
Indonesia is hyperendemic for dengue9 and classified as a country with frequent/continuous risks of dengue infection for visiting travellers in the dengue-risk map.10 The intensive labour flow and a large number of travellers in Bali have been reported to contribute to the spread of DENV infection.11 Previous reports indicate the hyperendemic transmission of all four DENV serotypes in Bali, where circulating DENVs include dominant local strains, suggesting that Bali is a melting pot of substantial DENV diversity and serves as a hub for dengue transmission and mixing of different strains.12–14
With the increase in a number of travellers in the coming decades as well as significant overall growth in the tourism sector worldwide,5 it is important to obtain data on exported dengue in travellers visiting Bali. As a very popular tourist destination, the endemicity of dengue in Bali can ultimately impact the health of foreign visitors. However, currently there are no data on dengue amongst travellers in Bali, including the clinical spectrum and infecting DENV serotypes. The information of whether travellers contract the disease locally or from other regions that were visited before Bali could not be ascertained. Molecular genotyping will be able to determine the genotype of the viruses and to some extent may also be used to track the origin of the viruses.15 Data on the clinical severity of the dengue and DENV serotypes and genotypes in travellers are important in order to gain comprehensive information on dengue disease in travellers. We conducted a fever survey, with the main focus on foreign tourists showing symptoms of dengue infection. The travellers were recruited from a local reference hospital, their demographic and clinical data assessed, and sera collected for the characterization of the infecting viruses, which included the serotype, genotype and the origin of the DENV.
Materials and Methods
Study design
The study was a prospective cross-sectional involving international travellers/tourists visiting Bali, Indonesia. The study protocol was approved by the Medical Research Ethic Committee of Udayana University/Sanglah General Hospital, Bali. Written informed consent was obtained from all subjects. Patients admitted to Kasih Ibu Hospital, Denpasar, Bali, Indonesia, were asked to participate in the study, and patient recruitment took place from April 2015 to December 2017. We recruited patients more than 1 year of age with a measured body temperature of ≥38°C and with less than 120 h/5 days of fever (acute phase), accompanied by at least one of the dengue symptoms, such as headache, rash, myalgia, arthralgia or retro-orbital pain.4 We excluded fever patients with clear symptoms of upper respiratory tract infections and/or those clearly diagnosed as non-dengue or who were unwilling to participate.
Serological and laboratory analyses
Serum samples were tested using dengue NS1 antigen and IgG/IgM serology rapid-tests (Standard Diagnostics, Korea) for dengue diagnosis. To determine the infection status of patients, samples were further tested using Dengue Duo IgG and IgM ELISA (Panbio, Alere, Australia) according to the manufacturer’s instructions. Common blood parameters such as haemoglobin, haematocrit, erythrocytes, platelets and leucocytes were documented. Determination of dengue severity was based on the WHO-SEARO dengue guidelines.4
Viral RNA detection and DENV genotype analysis
The presence of virus nucleic acid was detected using RT-PCR. Virus RNA was extracted from sera using QIAamp Viral RNA mini kit (Qiagen, Hilden, Germany) or the MagNA Pure LC Total Nucleic Acid kit and an automated MagNA Pure LC nucleic acid extraction system (Roche, Mannheim, Germany). The RT-PCR detection and serotyping were performed according to the method described elsewhere.13
The dengue NS1 and RT-PCR-negative samples were also tested for the presence of other arboviruses using pan-Flavivirus (such as Japanese Encephalitis, West Nile and Zika viruses) and pan-Alphavirus (such as the Chikungunya virus) RT-PCR panels, employing methods described elsewhere.16
For DENV genotype analysis, we sequenced the region of the envelope (E) protein, as described previously.17 Sequences were aligned using MUSCLE and phylogenetic trees and were inferred using the Maximum Likelihood method employing a general-time reversible (GTR) algorithm with four gamma parameters, invariant sites and 1000 boot-straps. The analysis was performed to determine the genotypes of the DENV according to classification by Goncalvez et al.,18 Twiddy et al.,19 Lanciotti et al.20 and Lanciotti et al.,21 for DENV-1, -2, -3 and -4, respectively. The E gene sequences of DENV obtained in this study have been deposited in the GenBank repository, with accession numbers MH173159–MH173175 and MH178411–MH178419.
Statistical analysis
Statistical analyses were performed using SPSS Version 18 (SPSS Inc., Chicago, IL) and R-statistics (www.r-project.org). A probability value of P ≤ 0.05 was considered statistically significant.
Results
Dengue cases and diagnosis in foreign travellers
Between April 2015 and December 2017, a total of 201 dengue-suspected foreign-traveller patients were enrolled. Dengue was confirmed in 133 (66.2%) patients, using NS1 antigen detection and/or viral nucleic acid detection by RT-PCR. The NS1 antigen was positive in 127 out of the 201 (63.2%). DENV serotype information was obtained for 119 out of the 133 (89.47%) dengue-confirmed patients. The age of the enrolled patients ranged from 15 to 78 (with a mean of 37 years old). In terms of gender, we observed a relatively equal proportion between males (43.6%) and females (56.4%). The majority (59.7%) of patients were of primary infection status.
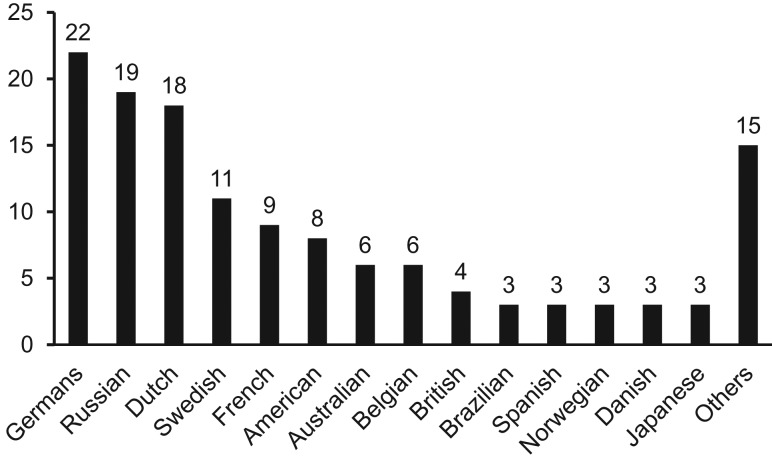
The top five nationalities of the patients with confirmed dengue cases were German (16.5%), Russian (14.3%), Dutch (13.5%), Swedish (8.3%) and French (6.8%) (Figure 1). Other arboviruses were not identified in DENV-negative samples (n = 68).
Figure 1.
Nationality of travellers involved in the study. Other nationalities include Austrian (2), Hungarian (2), Finnish (2), Estonian (2), Indian (1), Philippine (1), Lithuanian (1), Italian (1), New Zealander (1), Tunisian (1) and Cuban (1).
DENV serotype distribution
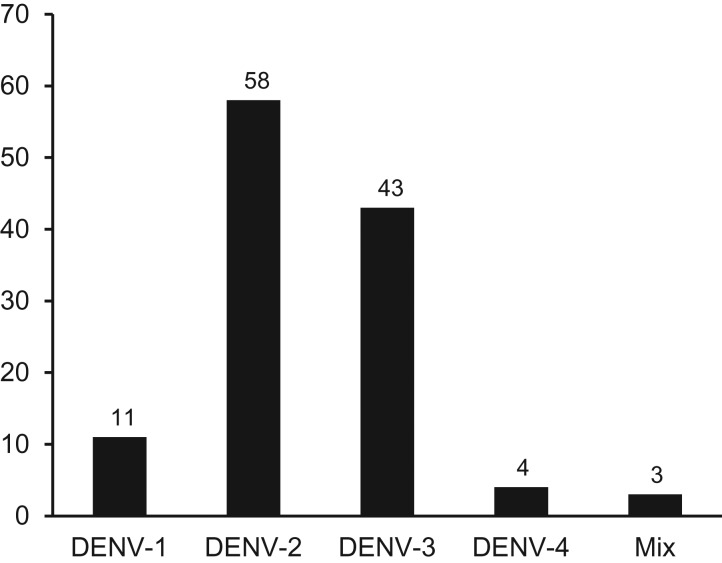
DENV serotyping using RT-PCR revealed the presence of all four DENV serotypes in the foreign patients. The predominant serotype was DENV-2 (48.7%), followed by DENV-3 (36.1%), DENV-1 (9.2%) and DENV-4 (3.4%) (Figure 2). Three samples were detected to have concurrent/mixed infection of DENV serotypes, of which two patients were infected by DENV-1 & DENV-2, and one by DENV-1 & DENV-4.
Figure 2.
DENV serotype distribution among travellers visiting Bali 2015–17.
Patient clinical manifestations
Based on WHO-SEARO dengue clinical manifestation, we classified 115 (86.5%) patients as DF, while 18 (13.5%) patients were DHF. Within the DHF group, one patient was classified as DHF Grade III (DSS). No fatalities were reported. We observed that the most common clinical manifestation were fever, myalgia and headache (Table 1). Other common symptoms of dengue such as malaise, loss of appetite, retro-orbital and joint pain, were also observed in smaller proportion. Rash was prominent in only 9.8% of the patients.
Table 1.
Clinical manifestations of 133 dengue-confirmed travellers
| Clinical features | N | % |
|---|---|---|
| Fever | 130 | 97.7 |
| Myalgia | 118 | 88.7 |
| Headache | 109 | 82.0 |
| Malaise | 93 | 69.9 |
| Loss of appetite | 86 | 64.7 |
| Retro-orbital pain | 86 | 64.7 |
| Joint pain | 55 | 41.4 |
| Vomiting | 27 | 20.3 |
| Sore throat | 25 | 18.8 |
| Bleeding | 19 | 14.3 |
| Rash | 13 | 9.8 |
DENV genotype analysis
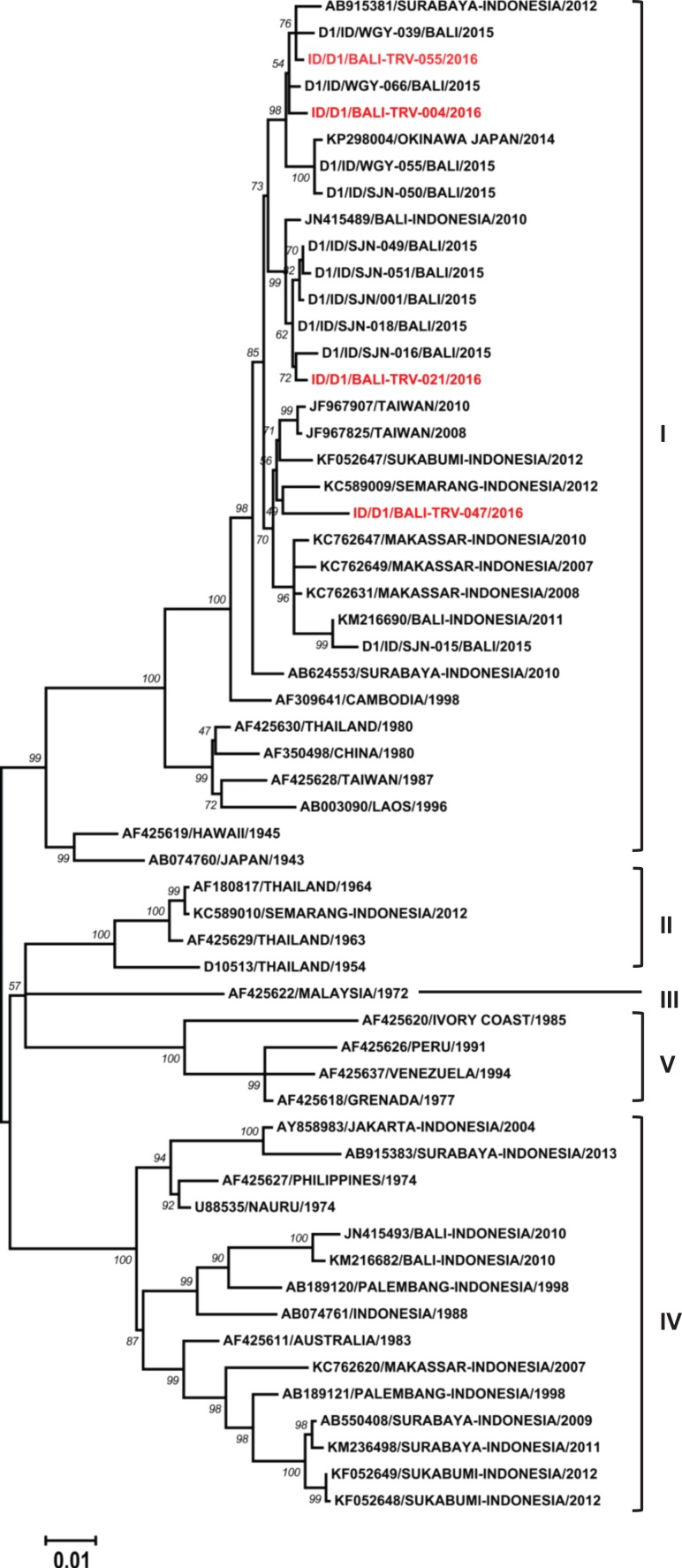
To determine the origin and genetic profile of the DENV infecting Bali travellers, we performed a phylogenetic analysis. A total of 26 representative DENV E gene sequences from all four serotypes were successfully obtained and phylogenetic trees were constructed. Four DENV-1 isolates were classified as Genotype I according to classification by Goncalvez et al.18 (Figure 3). The isolates were closely related to recent DENV strains isolated from local Balinese patients13 and other cities in Indonesia.
Figure 3.
Phylogenetic tree and genotype classification of DENV-1 isolates from travellers to Bali (red font) with other reference strains. Bootstrap values of >50 are shown. The scale bar indicates nucleotide substitutions per site.
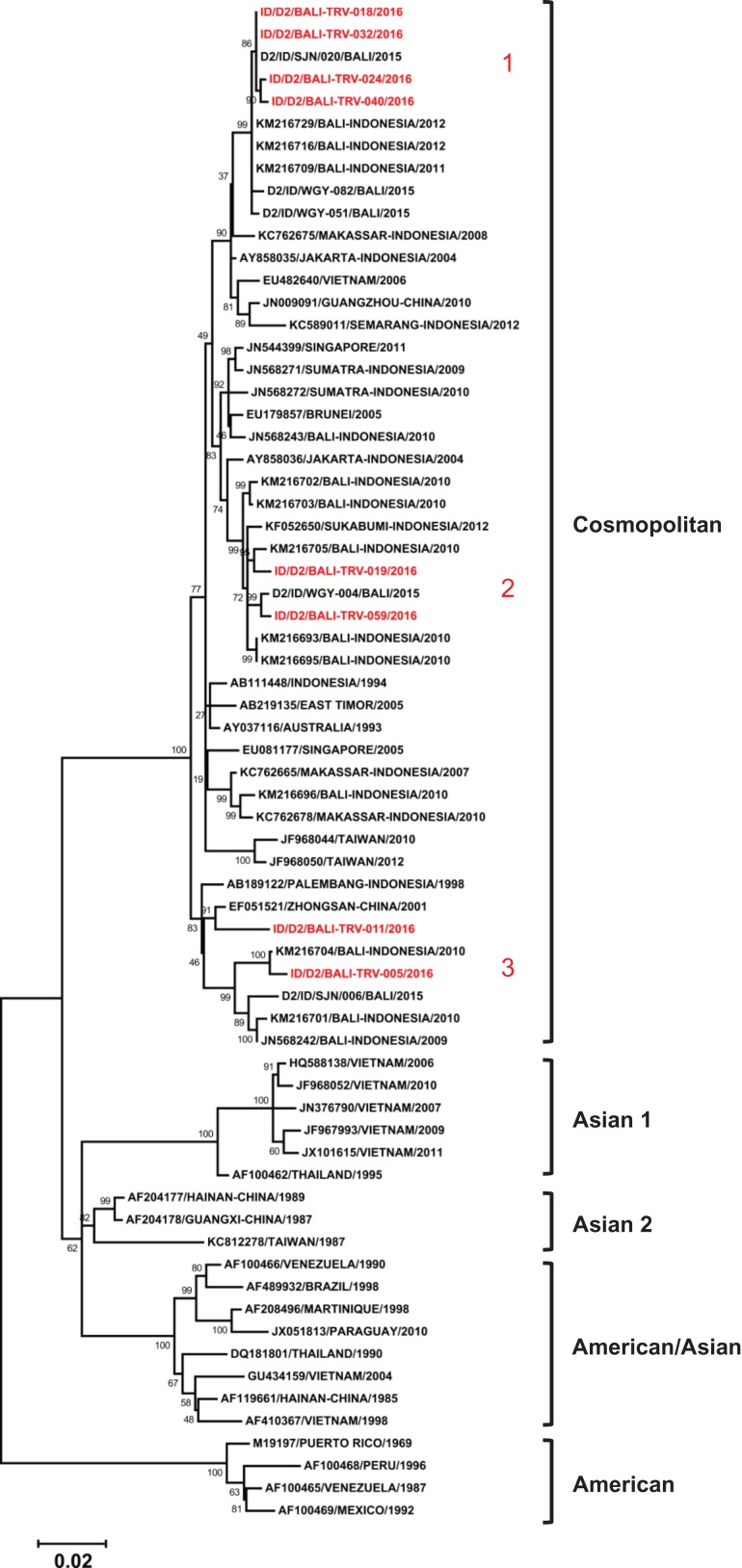
Eight DENV-2 isolates obtained belonged to the Cosmopolitan genotype, based on Twiddy et al.’s classification19 (Figure 4). A close relationship between the isolates from local Bali strains and other cities in Indonesia was also observed. Three lineages furtherly generated within the Cosmopolitan genotype were observed to form monophyletic clades with DENV strains from Bali.
Figure 4.
Phylogenetic tree and genotype classification of DENV-2 isolates from travellers to Bali (red font) with other reference strains. Bootstrap values of >50 are shown. The scale bar indicates nucleotide substitutions per site. Arabic numbers in red font denote lineage clustering.
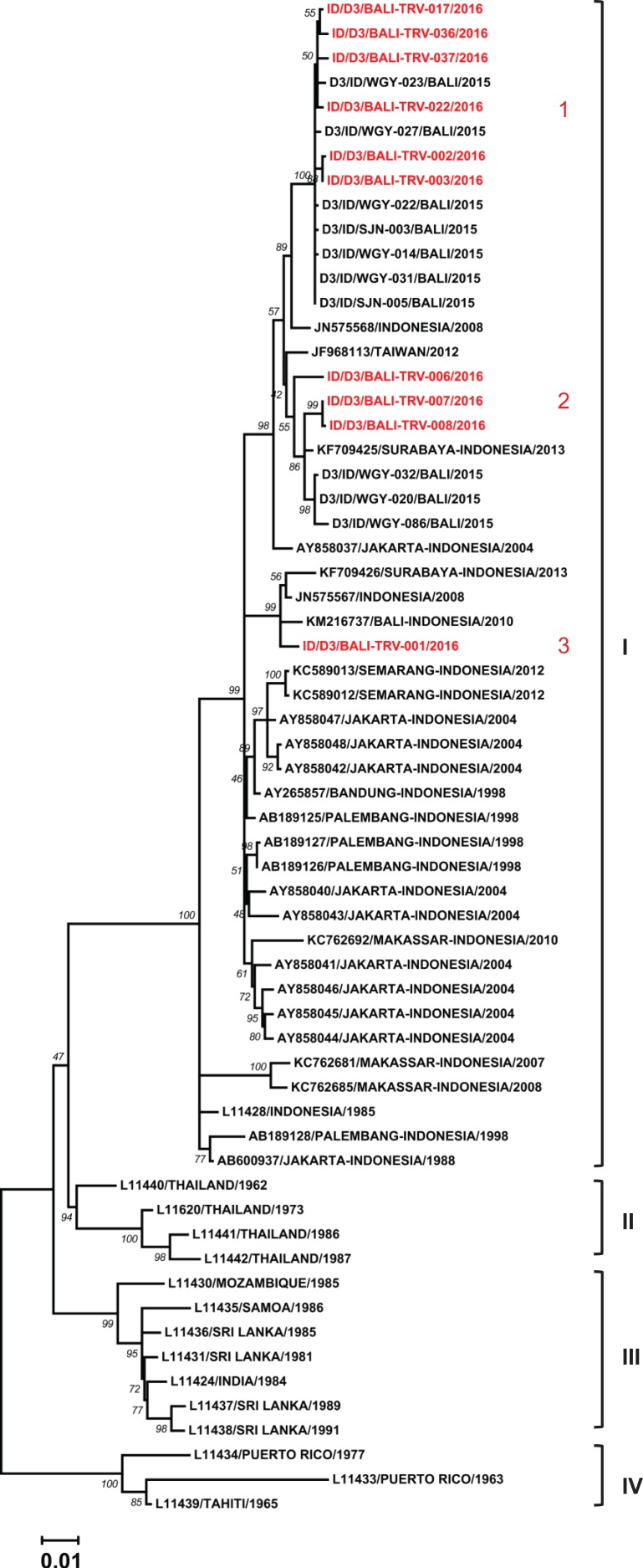
We successfully sequenced the E genes of 10 DENV-3 isolates. All the isolates were grouped together into Genotype I following Lanciotti et al.’s classification20 (Figure 5). The isolates were closely related to local strains from Bali and other cities in Indonesia. In addition, further sub-genotype grouping was formed, reflecting a total of three distinct lineages.
Figure 5.
Phylogenetic tree and genotype classification of DENV-3 isolates from travellers to Bali (red font) with other reference strains. Bootstrap values of >50 are shown. The scale bar indicates nucleotide substitutions per site. Arabic numbers in red font denote lineage clustering.
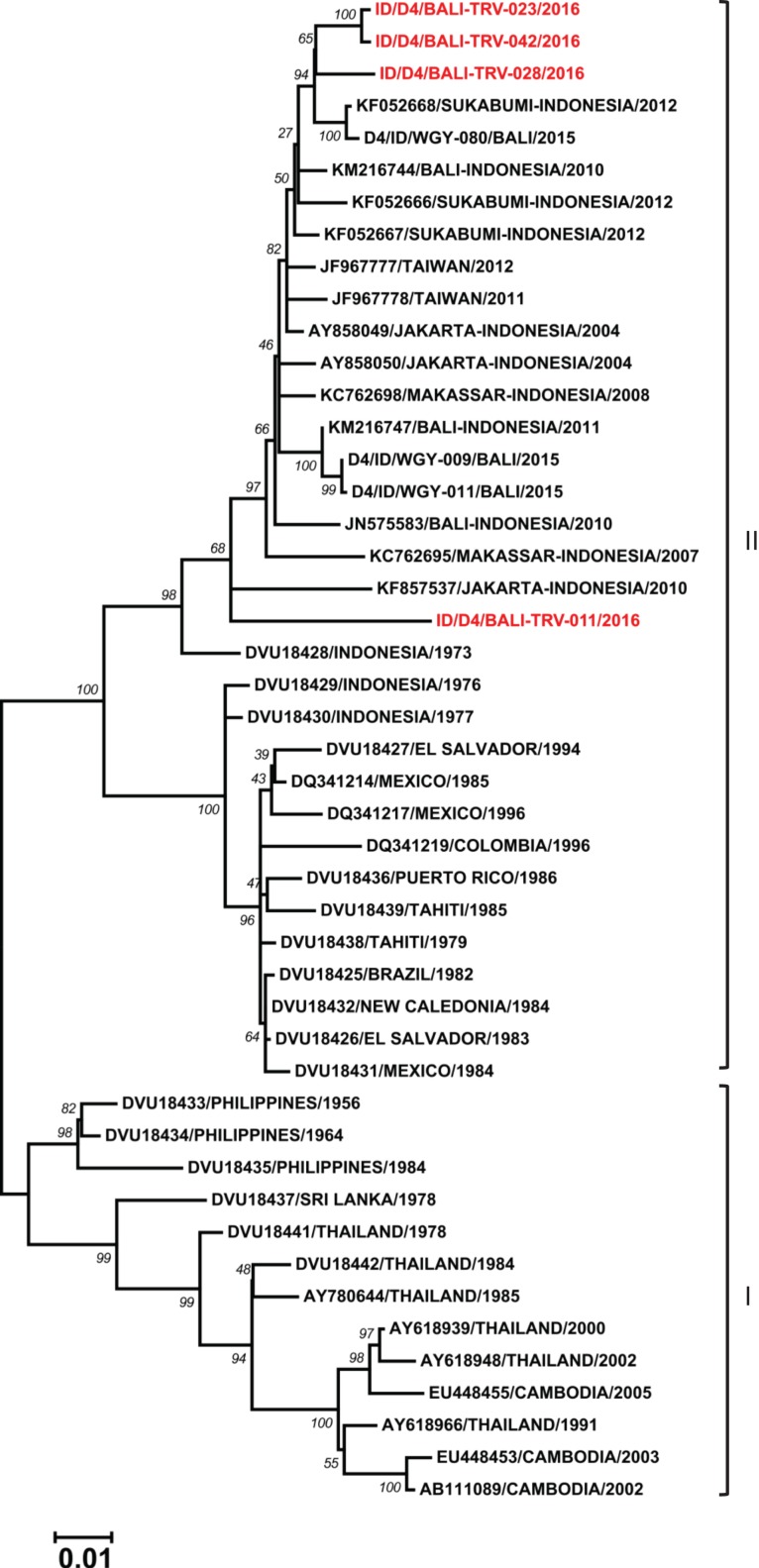
Four isolates of DENV-4 obtained from travellers to Bali were classified as members of Genotype II, following Lanciotti et al.21 (Figure 6). Among them, a relationship with local Bali strains was observed for three isolates, while one showed a close relationship with an older strain of DENV-4 from Indonesia and a strain from Jakarta from 2010 (unpublished).
Figure 6.
Phylogenetic tree and genotype classification of DENV-4 isolates from travellers to Bali (red font) with other reference strains. Bootstrap values of >50 are shown. The scale bar indicates nucleotide substitutions per site.
Discussion
We have previously reported the hyperendemicity of DENV in local Balinese people.13 In this study, we performed a surveillance of DENV in foreign travellers, and confirmed that dengue is one of the causes of febrile illnesses in travellers. We also screened the dengue-negative samples for other arboviral diseases that are endemic or previously detected in Bali, such as Japanese Encephalitis, Zika and Chikungunya virus infections,22–24 using pan-flavivirus and pan-alphavirus RT-PCR and virus isolation; however, we did not detect any positive results. The data demonstrate the important contribution of DENV infection in travellers to Bali presenting with febrile illness.
During our study, all four DENV serotypes were detected which further supports the hyperendemicity of dengue in Bali. However, most patients in this study were infected with DENV-2, in contrary to the dengue infection in local Balinese people in 2015, where DENV-3 was predominant.13 However, our results are in line with data reported by Ernst et al.,12 which showed the same predominance of DENV-2 in Australian travellers visiting Bali in 2010. The predominance of DENV-2 was also reported in other cities in Indonesia in 2012.25 Considering that our samples were collected from 2015 to 2017, the results may indicate a change in serotype predominance over time and reflect the dynamics of dengue in Bali.
We assessed the infection status of the patients and found that the majority (59.7%) of DENV infections were primary. This was not unexpected, considering that the patients came from countries where dengue is not commonly present (Figure 1). We presume that they have contracted dengue for the first time during their visit to Bali. The presence of remarkably high dengue-seropositive travellers (40.3%) in this study may have implications for dengue vaccination programs in international travellers. With travel-acquired dengue increasing, a safe and efficacious vaccine might be beneficial to prevent dengue in travellers as well as to reduce the risk of introduction to non-endemic countries.
In general, a relatively milder disease severity was observed, with most of the patients categorized as having DF. Mild dengue is more frequently reported in travellers compared to people from endemic areas.7 Moreover, a higher proportion of primary infection might be contributing to the mild disease manifestation. However, in this study, a small proportion of patients experiencing DHF was still observed.
To understand the origin of the DENV infecting travellers visiting Bali, we employed phylogenetic analysis using E gene sequences. Phylogenetic analysis based on partial and/or complete genomic sequences has been successfully used to elucidate the origins, genetic diversity, transmission dynamics and epidemic potential of DENV.15 Using this method, we identified the DENV infecting travellers as Genotype I, Cosmopolitan genotype, Genotype I and Genotype II, for DENV-1, DENV-2, DENV-3 and DENV-4, respectively. Overall, the genetic profiles of DENV in travellers to Bali were similar to the reported strains of local Balinese.13 Similar to previous findings,13 we did not find Genotype IV of DENV-1, which may indicate that this older lineage of DENV-1 has been replaced by the more recent Genotype I.26 Moreover, we observed a close relationship between the isolates and local Bali strains, strains from other cities in Indonesia, as well as strains from imported cases from Australia and the region. No DENV isolates were found to originate from or related to countries outside Indonesia. Overall, our study confirmed that only local endemic strains were responsible for dengue infection in international travellers to Bali, and supports the potential of Bali and Indonesia in general as the source of dengue case importation to other countries.13,27
The study has several limitations. Our patient recruitment was performed only in one hospital. However, the Kasih Ibu Hospital is the reference hospital for expatriates and international travellers, therefore the study population should be quite representative. In addition, only symptomatic dengue patients were recruited; therefore, we may have underestimated the true burden of dengue on international travellers. With regard to the DENV genetic analysis, we only used E gene sequences and a more in-depth evolutionary analysis using whole DENV genome sequences may be needed to better characterize the genetic make-up of DENV in travellers to Bali. We also did not have access to data on travellers’ personal measures on protection against mosquito bite (e.g. the use of mosquito-repellent). Nevertheless, our study is the first to report the incidence of dengue in travellers visiting Bali and to provide data on the clinical spectrum, epidemiology and virological aspects of the disease.
In conclusion, our study demonstrates that dengue infection is one of the causes of febrile illness in travellers visiting the island. Although affecting travellers’ health, the majority of patients only experienced mild DF. All four DENV serotypes were found to infect the travellers reflecting the hyperendemicity of dengue in the region, which may favour the spread of DENV to other countries through visiting travellers. Monitoring and continuous dengue surveillance, supported by an accurate diagnosis of the disease and preventive measures, including vaccination, improving healthcare access for travellers, will benefit travellers visiting endemic areas including Bali.
Acknowledgements
We would like to thank the patients and medical staff involved in this study as well as the United States Centers for Disease Control and Prevention for their support on non-dengue arboviral molecular testing used in this study. The assistance of Rama Dhenni from Eijkman Institute for the detection of non-dengue arboviruses and Made Satya Dharmayanti from Warmadewa University was greatly appreciated.
Conflict of interest: The authors have declared that there are no conflicts of interest.
Funding
This work was supported by grant from the International Society for Travel Medicine Research Awards to S.M. and R.T.S.
Author Contributions
S.M. and R.T.S. conceived and designed the study. S.M. and I.K.A.S. collected the samples and performed clinical assessments. S.M. and B.Y. performed the experiments and data analysis. K.S.A.M. provided resources and reviewed the manuscript. S.M., B.Y. and R.T.S. wrote the manuscript. All authors read and approved the final manuscript.
References
- 1. Bhatt S, Gething PW, Brady OJ et al. . The global distribution and burden of dengue. Nature 2013; 496:504–7. 10.1038/nature12060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Lambrechts L, Scott TW, Gubler DJ. Consequences of the expanding global distribution of Aedes albopictus for dengue virus transmission. PLoS Negl Trop Dis 2010; 4:e646 10.1371/journal.pntd.0000646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Holmes EC. RNA virus genomics: a world of possibilities. J Clin Invest 2009; 119:2488–95. 10.1172/JCI38050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. WHO-SEARO Comprehensive guidelines for prevention and control of dengue and dengue haemorrhagic fever In: Revised and expanded. New Delhi, India: World Health Organization, 2011. [Google Scholar]
- 5. Glaesser D, Kester J, Paulose H, Alizadeh A, Valentin B. Global travel patterns: an overview. J Travel Med 2017; 24(4). 10.1093/jtm/tax007. [DOI] [PubMed] [Google Scholar]
- 6. Wilder-Smith A. Closing the gap in travel medicine. J Travel Med 2017; 24(4). 10.1093/jtm/tax027. [DOI] [PubMed] [Google Scholar]
- 7. Wilder-Smith A. Dengue in international travelers: quo vadis? J Travel Med 2013; 20:341–3. 10.1111/jtm.12063. [DOI] [PubMed] [Google Scholar]
- 8. Ratnam I, Leder K, Black J, Torresi J. Dengue fever and international travel. J Travel Med 2013; 20:384–93. 10.1111/jtm.12052. [DOI] [PubMed] [Google Scholar]
- 9. Setiati TE, Wagenaar JF, de Kruif MD, Mairuhu AT, van Gorp EC, Soemantri A. Changing epidemiology of dengue haemorrhagic fever in Indonesia. Bull WHO 2006; 30:1–14. [Google Scholar]
- 10. Jentes ES, Lash RR, Johansson MA et al. . Evidence-based risk assessment and communication: a new global dengue-risk map for travellers and clinicians. J Travel Med 2016; 23(6). 10.1093/jtm/taw062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Yoshikawa MJ, Kusriastuti R. Surge of dengue virus infection and chikungunya Fever in Bali in 2010: the burden of mosquito-borne infectious diseases in a tourist destination. Trop Med Health 2013; 41:67–78. 10.2149/tmh.2011-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Ernst T, McCarthy S, Chidlow G et al. . Emergence of a new lineage of dengue virus type 2 identified in travelers entering Western Australia from Indonesia, 2010–2012. PLoS Negl Trop Dis 2015; 9:e0003442 10.1371/journal.pntd.0003442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Megawati D, Masyeni S, Yohan B et al. . Dengue in Bali: clinical characteristics and genetic diversity of circulating dengue viruses. PLoS Negl Trop Dis 2017; 11:e0005483 10.1371/journal.pntd.0005483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Warrilow D, Northill JA, Pyke AT. Sources of dengue viruses imported into Queensland, Australia, 2002–2010. Emerg Infect Dis 2012; 18:1850–1857. 10.3201/eid1811.120014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Weaver SC, Vasilakis N. Molecular evolution of dengue viruses: contributions of phylogenetics to understanding the history and epidemiology of the preeminent arboviral disease. Infect Genet Evol 2009; 9:523–40. 10.1016/j.meegid.2009.02.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Perkasa A, Yudhaputri F, Haryanto S et al. . Isolation of Zika Virus from Febrile Patient, Indonesia. Emerg Infect Dis 2016; 22:924–5. 10.3201/eid2205.151915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Lestari CSW, Yohan B, Yunita A et al. . Phylogenetic and evolutionary analyses of dengue viruses isolated in Jakarta, Indonesia. Virus Genes 2017. 10.1007/s11262-017-1474-7. [DOI] [PubMed] [Google Scholar]
- 18. Goncalvez AP, Escalante AA, Pujol FH et al. . Diversity and evolution of the envelope gene of dengue virus type 1. Virology 2002; 303:110–9. [DOI] [PubMed] [Google Scholar]
- 19. Twiddy SS, Farrar JJ, Vinh Chau N et al. . Phylogenetic relationships and differential selection pressures among genotypes of dengue-2 virus. Virology 2002; 298:63–72. [DOI] [PubMed] [Google Scholar]
- 20. Lanciotti RS, Lewis JG, Gubler DJ, Trent DW. Molecular evolution and epidemiology of dengue-3 viruses. J Gen Virol 1994; 75:65–75. [DOI] [PubMed] [Google Scholar]
- 21. Lanciotti RS, Gubler DJ, Trent DW. Molecular evolution and phylogeny of dengue-4 viruses. J Gen Virol 1997; 78:2279–84. [DOI] [PubMed] [Google Scholar]
- 22. Leung GHY, Baird RW, Druce J, Anstey NM. Zika virus infection in Australia following a monkey bite in Indonesia. Southeast Asian J Trop Med Public Health 2015; 46:460–4. [PubMed] [Google Scholar]
- 23. Sari K, Myint KSA, Andayani AR et al. . Chikungunya fever outbreak identified in North Bali, Indonesia. Trans R Soc Trop Med Hyg 2017; 111:325–7. 10.1093/trstmh/trx054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. de Jong W, Rusli M, Bhoelan S et al. . Endemic and emerging acute virus infections in Indonesia: an overview of the past decade and implications for the future. Crit Rev Microbiol 2018:1–17. 10.1080/1040841X.2018.1438986. [DOI] [PubMed] [Google Scholar]
- 25. Nusa R, Prasetyowati H, Meutiawati F et al. . Molecular surveillance of Dengue in Sukabumi, West Java province, Indonesia. J Infect Dev Ctries 2014; 8:733–41. [DOI] [PubMed] [Google Scholar]
- 26. Sasmono RT, Wahid I, Trimarsanto H et al. . Genomic analysis and growth characteristic of dengue viruses from Makassar, Indonesia. Infect Genet Evol 2015; 32:165–77. 10.1016/j.meegid.2015.03.006. [DOI] [PubMed] [Google Scholar]
- 27. Lestari CSW, Yohan B, Yunita A et al. . Phylogenetic and evolutionary analyses of dengue viruses isolated in Jakarta, Indonesia. Virus Genes 2017; 53:778–88. 10.1007/s11262-017-1474-7. [DOI] [PubMed] [Google Scholar]