Abstract
Background:
This study aimed to investigate the occurrence of aminoglycoside resistance and the prevalence of the resistance-modifying enzyme genes, ant(3”)-III, ant(6’)-Ia, aac(6′)-Ie-aph(2”)-Ia, and aph(2’)-Id, in Enterococcus strains isolated in Kermanshah Province, west of Iran.
Methods:
In this cross-sectional study, 108 enterococcal isolates from urine, wound, blood, and cerebrospinal fluid samples were collected. The Enterococcus species were recognized by standard phenotypic/biochemical tests. The antimicrobial resistance forms were detected using a disc diffusion method. Polymerase chain reaction was designed to identify aminoglycoside resistance genes, including ant(3”)-III, ant(6’)-Ia, aac(6′)-Ie-aph(2”)-Ia, and aph(2’)-Id.
Results:
Totally, 108 strains with a final diagnosis of Enterococcus were extracted from 84 (77.8%) urine, 14 (13%) wound, 6 (5.6%) blood, and 4 (3.7%) cerebrospinal fluid samples. Among the 108 Enterococcus specimens, 94 (87%) cases were Enterococcus faecalis and 14 (13%) were Enterococcus faecium. The highest frequency of resistance was observed for erythromycin (88.9%), while the lowest was found for streptomycin (44.4%). The frequency of high-level gentamicin resistance was 42.2%. Among the identified specimens, 42.6% contained the aac(6′)-Ie-aph(2”)-I gene, 20.4% contained the ant(6’)-Ia gene, and 15.7% contained the ant(3”)-III gene. A significant correlation was found between phenotypic gentamicin resistance and the presence of the aminoglycoside resistance genes (P<0.05).
Conclusion:
This study showed the high resistance of Enterococcus strains isolated from hospital samples. Compared with the previous studies, the strains isolated in our study showed a higher percentage of resistance to aminoglycosides.
Keywords: Aminoglycosides , Enterococcus , Aminoglycoside resistance genes , Prevalence
What’s Known
Enterococci are Gram-positive cocci capable of causing infections in humans and animals.
Evaluation of the prevalence of antibiotic resistance and the use of molecular typing techniques to identify genes responsible for resistance can help find ways to control these bacteria efficiently and reduce hospital-acquired infections caused by enterococci.
Results of studies in this field can significantly help prevent the spread of micro-organisms in hospital environments.
What’s New
Our study showed the high resistance of Enterococcus strains isolated from hospital samples in Kermanshah Province, west of Iran.
Introduction
Enterococci are Gram-positive cocci that are able to cause infection in humans and animals. These bacteria are the third common cause of infection in hospitalized patients in comparison with Escherichia coli and Staphylococcus aureus.1,2 Since 1980s, Enterococcus faecalis (E. faecalis) (80%-90%) and E. faecium (10%-15%) have had a higher prevalence than all the other strains.3 Enterococci are the main cause of 10% to 12% of hospital infections, 10% to 12% of urinary tract infections, and 5% to 10% of septicemia occurring in hospitals.4 The virulence of enterococci is due to not only the presence of virulence factors but also the resistance of the bacteria to various antibiotics.5 Common antibiotics such as glycopeptide antibiotics, beta-lactams, and aminoglycosides are utilized for the treatment of enterococcal infections.6 Enterococci can attain high-level aminoglycoside resistance. The resistance mechanism is attributed to the presence of aminoglycoside-modifying enzymes (AMEs).7 The most common enterococcal resistance gene to aminoglycoside is aac(6′)-Ie-aph(2”)-Ia, which is located on the Tn5281 transposon.8 Other enterococcal genes resistant to aminoglycoside include 2”-O phosphotransferase (APH(2’)), 3’-O phosphotransferase (APH(3’)), 3’-O adenyltransferase (ANT(3’)), 4’-O adenyltransferase (ANT(4’)), and 6’-O adenyltransferase (ANT(6’)).9,10 Clinical treatments for acute enterococcal infections need a mixture of a cell-wall active agent and an aminoglycoside, typically gentamicin.4,11 High-level gentamicin resistance (HLGR) in enterococci stops the synergism between gentamicin and antibiotics affecting bacterial walls such as vancomycin, ampicillin, and penicillin.12 The evaluation of the prevalence of antibiotic resistance and the use of molecular typing techniques for the identification of genes responsible for resistance can help find ways to control these bacteria efficiently and reduce hospital-acquired infections caused by enterococci. The results of studies in this field can significantly help thwart the spread of micro-organisms in hospital environments, prescribe and administer proper antibiotics for the treatment of resistant strains, prevent the increase in resistance to antibiotics, and reduce mortality in patients.13 Epidemiological studies have shown that in controlling the spread of bacterial resistance in a geographical region, it is necessary to obtain information about the status of bacterial resistance to the antibiotics in that particular area.
The present study aimed to investigate the occurrence of aminoglycoside resistance and the prevalence of the resistance-modifying enzyme genes ant(3”)-III, ant(6’)-Ia, aac(6′)-Ie-aph(2”)-Ia, and aph(2’)-Id in Enterococcus strains isolated from 2 hospitals in Kermanshah Province (Imam Khomeini and Imam Reza Kermanshah), west of Iran.
Materials and Methods
In this cross-sectional study, 108 enterococcal isolates from urine, wound, blood, and cerebrospinal fluid samples were collected from patients who referred to Imam Khomeini Hospital and Imam Reza Hospital, affiliated to Kermanshah University of Medical Sciences, between April and September 2016. The sample size was selected based on similar studies.14 The inclusion criterion was the diagnosis of Enterococcus, and the exclusion criterion was positive samples with other bacteria. The study was approved by our institutional review board, and written informed consent was obtained from all the patients.
Bacterial Isolates
Catalase tests, growth at 6.5% salt, bile esculin hydrolysis, and the pyrrolidonyl arylamidase (PYR) test were used to identify the genera and strains. Then, using the arabinose sugar fermentation process, E. faecalis (arabinose negative) was isolated from E. faecium (positive arabinose). The Enterococcus strains were investigated via biochemical reaction tests including fermentation of sugars (e.g., arabinose, sorbitol, mannitol, sorbose, and sucrose) and arginine dihydrolase.
Antibiotic Susceptibility Tests
Antimicrobial susceptibility was determined using the disc diffusion method (Kirby-Bauer) against gentamicin (10 µg), amikacin (30 µg), kanamycin (30 µg), and tobramycin (10 µg) (Mast, England) according to the guidelines of the Clinical and Laboratory Standards Institute (CLSI).15 In addition, a 120-µg gentamicin disc was used to identify HLGR. In the disc diffusion method, the discs were placed on the agar medium before they were incubated at 37 °C for 24 hours, and the diameter of the zone of inhibition was measured. Resistance was diagnosed8 by no zone and susceptibility by a zone of diameter ≥10mm. The results were confirmed through antibiotic susceptibility tests in triplicate for each sample.
Polymerase Chain Reaction Studies
Aminoglycoside resistance genes including ant(3”)-III, ant(6’)-Ia, aac(6′)-Ie-, and aph(2’)-Id were detected using polymerase chain reaction (PCR). DNA was also extracted through the boiling method as described previously.16 A fresh bacterial colony was suspended in 100 µL of sterile distilled water and boiled at 100 ºC for 10 minutes. After centrifugation, 3 mL of supernatant was used for the PCR assay with the primers described in table 1. The amplification of DNA was performed in a thermal cycler (Eppendorf, Germany). Subsequently, gene amplification was conducted on the AMEs genes. PCR was performed in a final volume of 25 μL with the following formula: 10X buffer=2.5 μL, dNTP mix=0.5 μL, forward primer=1 μL, reverse primer=1 μL, Taq polymerase=0.2 μL, H2O=12.5 μL, and MgCl2=0.75 μL. In addition, the thermocycler was programmed as follows: pre=denaturation temperature=94 °C for 5 minutes, denaturation temperature=94 °C for 30 seconds, annealing temperature=46 to 55 °C for 30 seconds, extension temperature=72 °C for 45 seconds, and post-extension temperature=72 °C for 5 minutes. The PCR products were electrophoresed in 1.5% agarose gels and visualized under ultraviolet light using a Gel Doc device (Bio-Rad, USA). Enterococcal strains carrying AME genes as positive controls were obtained from Kermanshah University of Medical Sciences.
Table 1.
Primers used to identify genes for resistance to aminoglycosides in Enterococcus
| Primer name | Sequence | PCR product (bp) |
|---|---|---|
| ant (3”)-III | F-CACGCTATTACGAACTATGA | 284 |
| R-TAAGAAAGAACATCACCACGA | ||
| ant (6’)-Ia | F-ACTCGGGGATTGATAGGC | 597 |
| R-GCTGCTAAAGCTGCGCTT | ||
| aac (6′)-Ie- aph (2”)-Ia | F-GAGCAATAAGGGCATACCAAAAATC | 505 |
| R-CCGTGCATTTGTCTTAAAAAACTGG | ||
| aph (2’)-Id | F-GTGGTTTTTACAGGAATGCCATC | 641 |
| R-CCCTCTTCATACCAATCCATATAACC |
Statistical Analysis
The nominal variables were identified using frequencies. The significance of the obtained results was detected via the χ2 test at a significance level of P≤0.05 using SPSS, version 16.
Results
In this study, 108 strains with a final diagnosis of Enterococcus were extracted from 84 (77.8%) urine, 14 (13%) wound, 6 (5.6%) blood, and 4 (3.7%) cerebrospinal fluid samples. From this total, 94 (87%) isolates were E. faecalis and 14 (13%) were E. faecium. Among the 108 Enterococcus specimens, 94 (87%) cases were E. faecalis, and 14 (13%) were E. faecium. The highest frequency of resistance was observed for erythromycin (88.9%), while the lowest was found for streptomycin (44.4%). The frequency of HLGR was 42.2% (table 2). (Table 2 presents the antibiogram pattern in the disc diffusion method.) Among the identified specimens, 42.6% contained the aac(6′)-Ie-aph(2”)-Ia gene, 20.4% contained the ant(6’)-Ia gene, and 15.7% contained the ant(3”)-III gene (table 3). (Table 3 presents the prevalence of the genes responsible for resistance to aminoglycosides in the E. faecalis and E. faecium strains.) The aminoglycoside resistance genes ant(3”)-III, ant(6’)-Ia, aac(6′)-Ie-aph(2”)-Ia, and aph(2’)-Id were detected by PCR (figure 1- figure 2). Data analysis revealed a significant correlation between the resistance genes and phenotype resistance (P<0.05). Tables 4 and 5 depict the frequencies of the AME genes resistant to aminoglycosides in the enterococci and the pertinent correlations.
Table 2.
Antibiogram pattern in the disc diffusion method
| Bacteria | Percentage (%) of Enterococcus isolates based on the antibiotic resistance | ||||||
|---|---|---|---|---|---|---|---|
| TOB | HLGR | GM | ERM | STR | KA | AK | |
| Enterococcus faecalis | 62 (57.4) | 33 (30.6) | 76 (70.4) | 85 (78.7) | 24 (22.2) | 67 (62) | 60 (55.6) |
| Enterococcus faecium | 8 (7.4) | 13 (12) | 13 (12) | 11 (10.2) | 3 (2.8) | 12 (11.1) | 8 (7.4) |
| Total number (%) | 70 (64.8) | 46 (42.6) | 46 (42.6) | 96 (88.9) | 27 (44.4) | 79 (73.1) | 68 (63) |
TOB: Tobramycin; GM: Gentamicin; ERM: Erythromycin; STR: Streptomycin; KA: Kanamycin; AK: Amikacin
Table 3.
Incidence of the aminoglycoside-modifying enzyme (AME) genes in each of the enterococcal species
| AME Gene | E. faecalis (94 isolates) | E. faecium (14 isolates) | Total |
|---|---|---|---|
| aac (6′)-Ie-aph (2”)-Ia | 33 (30.6%) | 13 (12%) | 46 (42.6%) |
| aph (2’)-Id | 0 | 0 | 0 |
| ant (6’)-Ia | 19 (17.6%) | 3 (2.8%) | 22 (20.4%) |
| ant (3”)-III | 15 (13.9%) | 2 (1.9%) | 17 (15.7%) |
Figure1.
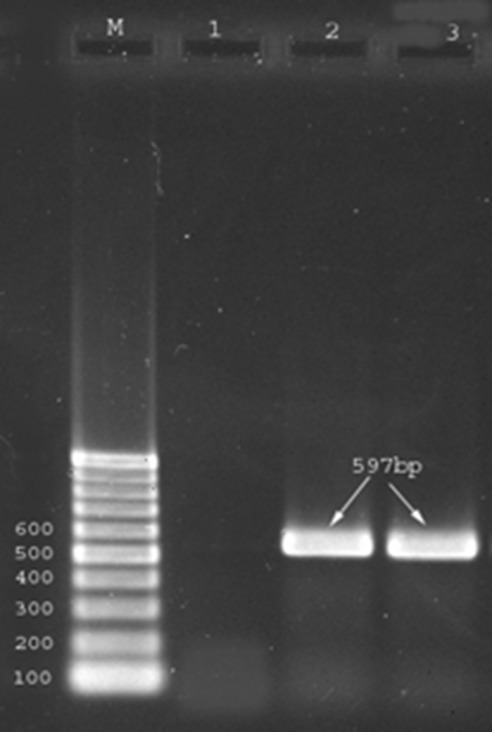

Amplified products generated by polymerase chain reaction (PCR). Lane M is 100-bp marker. Lane 1 shows negative PCR control, Lane 2 is positive control ant (6’)-Ia (597bp) gene; and Lane 3 is sample positive.
Figure2.
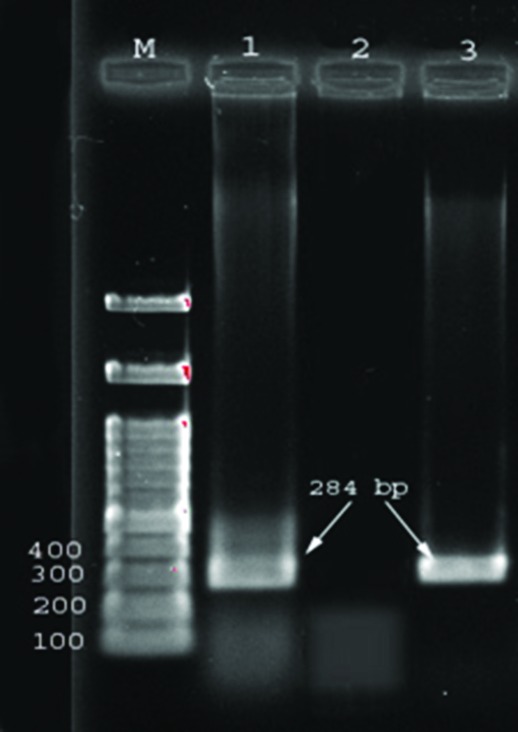
Amplified products generated by polymerase chain reaction (PCR). Lane M is 100-bp markers. Lane 1 is positive control ant(3”)-III (284bp) gene, Lane 2 shows negative PCR control, and Lane 3 is sample positive.
Table 4.
Frequency of the aminoglycoside-modifying enzyme genes resistant to aminoglycosides in the Enterococcus isolates
| Antibiotics | E. faecalis | E. faecium | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ant (3”)-III | ant (6’)-Ia | aac (6′)-Ie | aph (2’)-Id | ant (3”)-III | ant (6’)-Ia | aac (6′)-Ie | aph (2’)-Id | |||||||||
| Pos | neg | pos | neg | pos | neg | pos | neg | Pos | neg | pos | neg | pos | neg | pos | neg | |
| Gentamicin | 13 | 63 | 19 | 57 | 33 | 43 | 0 | 76 | 2 | 11 | 3 | 10 | 12 | 1 | 0 | 13 |
| Tobramycin | 1 | 7 | 19 | 43 | 16 | 46 | 0 | 62 | 14 | 48 | 3 | 5 | 7 | 1 | 0 | 8 |
| Kanamycin | 13 | 54 | 19 | 48 | 23 | 44 | 0 | 67 | 1 | 11 | 3 | 9 | 11 | 1 | 0 | 12 |
| Amikacin | 14 | 46 | 19 | 41 | 19 | 41 | 0 | 60 | 1 | 7 | 3 | 5 | 7 | 1 | 0 | 8 |
| Streptomycin | 14 | 10 | 16 | 8 | 4 | 20 | 0 | 24 | 2 | 1 | 2 | 1 | 3 | 0 | 0 | 3 |
| Erythromycin | 15 | 70 | 19 | 66 | 28 | 57 | 0 | 85 | 2 | 9 | 3 | 8 | 10 | 1 | 0 | 11 |
| HLGR | 3 | 30 | 3 | 30 | 3 | 30 | 0 | 33 | 2 | 11 | 3 | 10 | 3 | 10 | 0 | 13 |
Pos: Positive; neg: Negative
Table 5.
Correlation between the frequency of aminoglycoside-modifying enzyme genes resistant to aminoglycosides in the Enterococcus isolates
| Antibiotics | ant (3”)-III (positive) | ant (6’)-Ia (positive) | aac (6′) Ie (positive) | aph (2’) Id (positive) |
|---|---|---|---|---|
| Gentamicin | 15 | 22* | 35* | 89 |
| Tobramycin | 15* | 22* | 23 | 70 |
| Kanamycin | 14 | 22* | 34 | 79 |
| Amikacin | 15* | 22* | 26 | 68 |
| Streptomycin | 16* | 18* | 7* | 27 |
| Erythromycin | 17 | 22 | 38 | 96 |
P<0.05
Figure3.
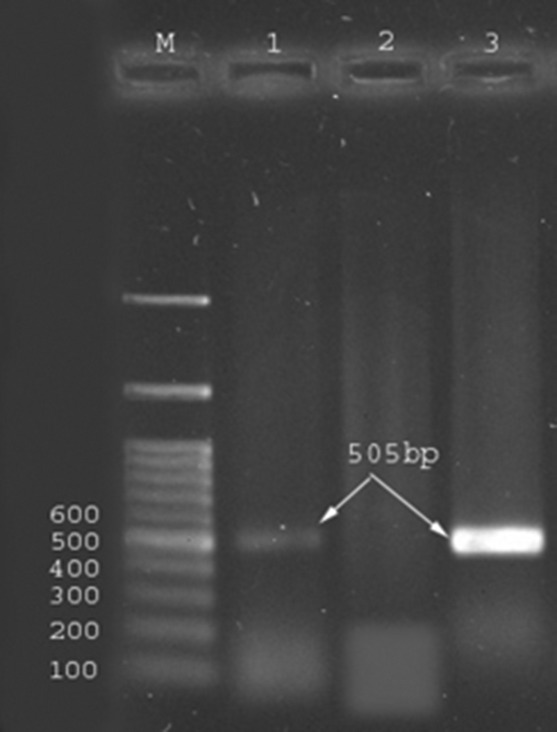
Amplified products generated by polymerase chain reaction (PCR). Lane M is 100-bp markers. Lane 1 is positive control aac(6′)-Ie-aph(2”)-Ia (505bp) gene, Lane 2 shows negative PCR control, and Lane 3 is sample positive.
Discussion
In recent years, multi-drug resistant Gram-positive microorganisms have been recognized as one of the main causes of death in hospitals.17 Among enterococci, E. faecalis and E. faecium are the 2 dominant strains commonly isolated from human infections. E. faecalis has a strong capability to bind and proliferate in the intestine and consequently plays a greater role in enterococcal infections. On the other hand, E. faecium has a high potential to become resistant to multiple antibiotics and, thus, accounts for a high percentage of resistance to different antibiotics. 18,19 Studies that have investigated the prevalence of resistance to gentamicin in the United States, Canada, Latin America, Europe, and Asia have shown that 14% to 40% of enterococci are resistant to gentamicin.20 The aim of the current study was to determine the prevalence of AMEs in hospitals in Kermanshah Province, Iran.
In the present study, among all the strains isolated from enterococci, 87% were E. faecalis and 13% were E. faecium. The difference in the incidence of these species among the 108 Enterococcus-containing samples was significant. Our finding is consistent with the results of other studies such as those conducted by Li et al.21 and Mohammadi et al., 22 who reported that the prevalence of E. faecalis was higher than that of E. faecium in their clinical specimens. This finding may be due to the capability of E. faecalis to adapt to the body’s condition and its higher level of presence in the body, particularly in the gastrointestinal tract.23 Several studies have indicated that the prevalence of E. faecium is higher than that of E. faecalis in clinical specimens, which is not consistent with our findings.24-26 It is possible that the sampling methods create the diversity in the frequency patterns of these 2 species.
The majority of the bacteria were isolated from urine samples (77.8%) and the lowest number of the strains was isolated from the cerebrospinal fluid (3.7%). In line with our results, the studies by El-Ghazawy et al.27 and Mittal et al.28 demonstrated that most bacteria were isolated from urine samples.
In the present research, apropos antibiotic susceptibility as assessed via the disc diffusion method, the strains showed the highest resistance to erythromycin (88.9%) and gentamicin (82.4%) and the lowest resistance to streptomycin (44.4%). In addition, 42.6% of the strains were resistant to a high level of gentamicin, which is consistent with results of an investigation by Mirnejad et al.29
In our study, E. faecalis strains resistant to HLGR accounted for 30.6% all the strains. Further, the resistance rate to gentamicin was 70.4%. Chiming in with our results, in the study by Li et al.,21 resistance to gentamicin was 58.8%. However, the prevalence rates of the E. faecium strains resistant to HLGR and gentamicin were 12% and 12%, respectively. In other words, the total resistance of both strains to HLGR was 42.6%. This finding is consistent with the results of a study conducted in 2009 in Iran by Behnoud et al. ,30 who reported that 32.43% of their enterococci cases were resistant to HLGR. Moreover, this finding is concordant with the results of an investigation carried out in 2006 by Feizabadi et al.,19 who reported that 52% of their cases were resistant to HLGR. According to Ben Saeid et al.,31 all the E. faecalis and E. faecium strains (5.2% and 6.1%, correspondingly) in their investigation had an HLGR phenotype.
During the last 30 years in Iran, aminoglycosides, particularly gentamicin, have been widely used for the treatment of most infections. This could be the principal reason for the high prevalence of HLGR strains in hospitals in Iran.32 In our study, all the HLGR strains had the aac(6’)-Ie-aph(2’’) gene. Consistent with our results, Faizabadi et al.19 showed that HLGR strains had the aac(6’)-Ie-aph(2’’) gene.
E. faecalis and E. faecium contained the aac(6’)-Ie-aph(2’’) gene with a prevalence rate of 30.6% and 92%, respectively. In the study by Li et al.,21 the prevalence rate of this gene was 49.4%. Elsewhere Jackson33 reported that 23% of the E. faecalis strains and 8.5% of the E. faecium strains in their study featured the aac(6’)-Ie-aph(2’’) gene. Additionally, 24% of all the strains in that investigation had an HLGR phenotype. According to Padmasini et al.,26 17.9% of the E. faecalis strains and 21.9% of the E. faecium strains had the aac(6’)-Ie-aph(2’’) gene.
In our study, the aph(2’)-Id gene was not found in the E. faecalis and E. faecium strains. This finding is in line with the results reported by Padmasini et al.26 In contrast, Li et al.21 reported a prevalence rate of 1.3% for this gene. The prevalence rate of ant(6’)-Ia in the E. faecalis and E. faecium was 17.6% and 2.8, respectively, which was lower than the prevalence rate of 31.3% reported by Li et al. 21 The prevalence rate of the ant (6’)-Ia gene was 7.8% in a research conducted by Said LB.31
Conclusion
This study showed the high resistance of Enterococcus strains isolated from hospital samples. Compared with the previous studies, the strains isolated in our study exhibited a higher percentage of resistance to aminoglycosides. The excessive use of these antibiotics can be the main reason for the high incidence of antibiotic resistance.
Acknowledgement
The authors wish to thank all the patients and health staff who participated in this study. This work received no financial support.
Conflict of Interest:None declared.
References
- 1.Willems RJ, Hanage WP, Bessen DE, Feil EJ. Population biology of Gram-positive pathogens: high-risk clones for dissemination of antibiotic resistance. FEMS Microbiol Rev. 2011;35:872–900. doi: 10.1111/j.1574-6976.2011.00284.x. [ PMC Free Article] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Cattoir V, Leclercq R. Twenty-five years of shared life with vancomycin-resistant enterococci: is it time to divorce? J Antimicrob Chemother. 2013;68:731–42. doi: 10.1093/jac/dks469. [DOI] [PubMed] [Google Scholar]
- 3.Nasaj M, Mousavi SM, Hosseini SM, Arabestani MR. Prevalence of Virulence Factors and Vancomycin-resistant Genes among Enterococcus faecalis and E. faecium Isolated from Clinical Specimens. Iran J Public Health 2016;45:806–13. [ PMC Free Article] [PMC free article] [PubMed] [Google Scholar]
- 4.Gikas A, Christidou A, Scoulica E, Nikolaidis P, Skoutelis A, Levidiotou S, et al. Epidemiology and molecular analysis of intestinal colonization by vancomycin-resistant enterococci in greek hospitals. J Clin Microbiol. 2005;43:5796–9. doi: 10.1128/JCM.43.11.5796-5799.2005. [ PMC Free Article] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Yamane K, Wachino J, Doi Y, Kurokawa H, Arakawa Y. Global spread of multiple aminoglycoside resistance genes. Emerg Infect Dis. 2005;11:951–3. doi: 10.3201/eid1106.040924. [ PMC Free Article] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Sood S, Malhotra M, Das BK, Kapil A. Enterococcal infections & antimicrobial resistance. Indian J Med Res. 2008;128:111–21. [PubMed] [Google Scholar]
- 7.Ramirez MS, Tolmasky ME. Aminoglycoside modifying enzymes. Drug Resist Updat. 2010;13:151–71. doi: 10.1016/j.drup.2010.08.003. [ PMC Free Article] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lebreton F, van Schaik W, McGuire AM, Godfrey P, Griggs A, Mazumdar V, et al. Emergence of epidemic multidrug-resistant Enterococcus faecium from animal and commensal strains. MBio. 2013;4 doi: 10.1128/mBio.00534-13. [ PMC Free Article] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Strateva T, Atanasova D, Savov E, Petrova G, Mitov I. Incidence of virulence determinants in clinical Enterococcus faecalis and Enterococcus faecium isolates collected in Bulgaria. Braz J Infect Dis. 2016;20:127–33. doi: 10.1016/j.bjid.2015.11.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Zou LK, Wang HN, Zeng B, Li JN, Li XT, Zhang AY, et al. Erythromycin resistance and virulence genes in Enterococcus faecalis from swine in China. New Microbiol. 2011;34:73–80. [PubMed] [Google Scholar]
- 11.Fisher K, Phillips C. The ecology, epidemiology and virulence of Enterococcus. Microbiology. 2009;155:1749–57. doi: 10.1099/mic.0.026385-0. [DOI] [PubMed] [Google Scholar]
- 12.Marothi YA, Agnihotri H, Dubey D. Enterococcal resistance--an overview. Indian J Med Microbiol. 2005;23:214–9. [PubMed] [Google Scholar]
- 13.Singh A, Goering RV, Simjee S, Foley SL, Zervos MJ. Application of molecular techniques to the study of hospital infection. Clin Microbiol Rev. 2006;19:512–30. doi: 10.1128/CMR.00025-05. [ PMC Free Article] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Tilea B, Teches S, Voidazan S, Brinzaniuc K, Tilea I. Infective Endocarditis–An Observational Study. ARS Medica Tomitana. 2015;21:87–94. doi: 10.1515/arsm-2015-0026. [DOI] [Google Scholar]
- 15.Jorgensen JH, Hindler JF. New consensus guidelines from the Clinical and Laboratory Standards Institute for antimicrobial susceptibility testing of infrequently isolated or fastidious bacteria. Clin Infect Dis. 2007;44:280–6. doi: 10.1086/510431. [DOI] [PubMed] [Google Scholar]
- 16.Colak D, Naas T, Gunseren F, Fortineau N, Ogunc D, Gultekin M, et al. First outbreak of vancomycin-resistant enterococci in a tertiary hospital in Turkey. J Antimicrob Chemother. 2002;50:397–401. doi: 10.1093/jac/dkf134. [DOI] [PubMed] [Google Scholar]
- 17.Hayakawa K, Martin ET, Gudur UM, Marchaim D, Dalle D, Alshabani K, et al. Impact of different antimicrobial therapies on clinical and fiscal outcomes of patients with bacteremia due to vancomycin-resistant enterococci. Antimicrob Agents Chemother. 2014;58:3968–75. doi: 10.1128/AAC.0294314. [ PMC Free Article] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Zarrilli R, Tripodi MF, Di Popolo A, Fortunato R, Bagattini M, Crispino M, et al. Molecular epidemiology of high-level aminoglycoside-resistant enterococci isolated from patients in a university hospital in southern Italy. J Antimicrob Chemother. 2005;56:827–35. doi: 10.1093/jac/dki347. [DOI] [PubMed] [Google Scholar]
- 19.Feizabadi MM, Maleknejad P, Asgharzadeh A, Asadi S, Shokrzadeh L, Sayadi S. Prevalence of aminoglycoside-modifying enzymes genes among isolates of Enterococcus faecalis and Enterococcus faecium in Iran. Microb Drug Resist. 2006;12:265–8. doi: 10.1089/mdr.2006.12.265. [DOI] [PubMed] [Google Scholar]
- 20.Thaver D, Ali SA, Zaidi AK. Antimicrobial resistance among neonatal pathogens in developing countries. Pediatr Infect Dis J. 2009;28:S19–21. doi: 10.1097/INF.0b013e3181958780. [DOI] [PubMed] [Google Scholar]
- 21.Li W, Li J, Wei Q, Hu Q, Lin X, Chen M, et al. Characterization of aminoglycoside resistance and virulence genes among Enterococcus spp. isolated from a hospital in China. Int J Environ Res Public Health 2015;12:3014–25. doi: 10.3390/ijerph120303014. [ PMC Free Article] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Mohammadi F, Tabaraie B, Sadeghifard N, Ghafoorian S, Maleki A, Davoodian E, et al. Evaluation of drug resistance frequency among Entrococci faecium and E. faecalis strains and detection of VanA/B genes in vancomycin resistance isolated by PCR method in ilam and kermanshah hospitals. Journal of Ilam University of Medical Sciences 2011;19:1–8. [Google Scholar]
- 23.Brinster S, Posteraro B, Bierne H, Alberti A, Makhzami S, Sanguinetti M, et al. Enterococcal leucine-rich repeat-containing protein involved in virulence and host inflammatory response. Infect Immun. 2007;75:4463–71. doi: 10.1128/IAI.0027907. [ PMC Free Article] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Dadfarma N, Imani Fooladi AA, Oskoui M, Mahmoodzadeh Hosseini H. High level of gentamicin resistance (HLGR) among enterococcus strains isolated from clinical specimens. J Infect Public Health. 2013;6:202–8. doi: 10.1016/j.jiph.2013.01.001. [DOI] [PubMed] [Google Scholar]
- 25.Rahimi F, Talebi M, Saifi M, Pourshafie MR. Distribution of enterococcal species and detection of vancomycin resistance genes by multiplex PCR in Tehran sewage. Iran Biomed J. 2007;11:161–7. [PubMed] [Google Scholar]
- 26.Padmasini E, Padmaraj R, Ramesh SS. High level aminoglycoside resistance and distribution of aminoglycoside resistant genes among clinical isolates of Enterococcus species in Chennai, India. ScientificWorldJournal. 2014;2014 doi: 10.1155/2014/329157. [ PMC Free Article] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.El-Ghazawy IF, Okasha HA, Mazloum SM. A study of high level aminoglycoside resistant enterococci. Afr J Microbiol Res. 2016;10:572–7. doi: 10.5897/AJMR2015.7943. [DOI] [Google Scholar]
- 28.Mittal S, Singla P, Deep A, Bala K, Sikka R, Garg M, et al. Vancomycin and High Level Aminoglycoside Resistance in Enterococcus spp. in a Tertiary Health Care Centre: A Therapeutic Concern. 2016;2016:8262561. doi: 10.1155/2016/8262561. [ PMC Free Article] [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Mirnejad R, Sajjadi N, Masoumi Zavaryani S, Piranfar V, Hajihosseini M, Roshanfekr M. Identification of aminoglycoside resistance genes by Triplex PCR in Enterococcus spp. isolated from ICUs. 2016;24:222–9. [PubMed] [Google Scholar]
- 30.Behnood A, Farajnia S, Moaddab SR, Ahdi-Khosroshahi S, Katayounzadeh A. Prevalence of aac(6’)-Ie-aph(2’’)-Ia resistance gene and its linkage to Tn5281 in Enterococcus faecalis and Enterococcus faecium isolates from Tabriz hospitals. Iran J Microbiol. 2013;5:203–8. [ PMC Free Article] [PMC free article] [PubMed] [Google Scholar]
- 31.Ben Said L, Klibi N, Lozano C, Dziri R, Ben Slama K, Boudabous A, et al. Diversity of enterococcal species and characterization of high-level aminoglycoside resistant enterococci of samples of wastewater and surface water in Tunisia. Sci Total Environ. 2015;530-531:11–7. doi: 10.1016/j.scitotenv.2015.05.091. [DOI] [PubMed] [Google Scholar]
- 32.Shokoohi Zade L, Mohabbati Mobarez A, Alebooye M, Ranjbar R, Zali M. Antibiotic Resistance Patterns of E. Faecium and E. Faecalis Strains Isolated from ICUs. Medical Laboratory Journal. 2015;8:34–41. [Google Scholar]
- 33.Jackson CR, Fedorka-Cray PJ, Barrett JB, Ladely SR. High-level aminoglycoside resistant enterococci isolated from swine. Epidemiol Infect. 2005;133:367–71. doi: 10.1017/s0950268804003395. [ PMC Free Article] [DOI] [PMC free article] [PubMed] [Google Scholar]


