Abstract
Polianthes tuberosa is a popular ornamental plant. Its floral scent volatiles mainly consist of terpenes and benzenoids that emit a charming fragrance. However, our understanding of the molecular mechanism responsible for the floral scent of P. tuberosa is limited. Using transcriptome sequencing and de novo assembly, a total of 228,706,703 high-quality reads were obtained, which resulted in the identification of 96,906 unigenes (SRA Accession Number SRP126470, TSA Acc. No. GGEA00000000). Approximately 41.85% of the unigenes were functionally annotated using public databases. A total of 4,694 differentially expressed genes (DEGs)were discovered during flowering. Gas chromatography-mass spectrometry analysis revealed that the majority of the volatiles comprised benzenoids and small amounts of terpenes. Homology analysis identified 13 and 17 candidate genes associated with terpene and benzenoid biosynthesis, respectively. Among these, PtTPS1, PtDAHPSs, PtPAL1, and PtBCMT2 might play important roles in regulating the formation of floral volatiles. The data generated by transcriptome sequencing provide a critical resource for exploring concrete characteristics as well as for supporting functional genomics studies. The results of the present study also lay the foundation for the elucidation of the molecular mechanism underlying the regulation of floral scents in monocots.
Introduction
Floral scent is a crucial characteristic of ornamental plants, attracting pollinators for sexual reproduction and acting as a defense mechanism against pathogens, parasites, and herbivores [1–2]. Floral volatiles are mainly emitted by floral organs at specific flowering periods [3]. Approximately 1,700 floral volatiles from 100 plant species have been identified to date [1]. Floral volatiles consist of terpenoids, phenylpropanoids/benzenoids, and fattyacid derivatives, which are synthesized via different plant pathways [2].
Terpenes, the largest class of floral volatiles, are mainly composed of monoterpenes (C10), sesquiterpenes (C15), and diterpenes (C20)[3]. Volatile terpenoids are synthesized from two common isoprene precursors, namely, isopentenyl pyrophosphate (IPP) and dimethylallyl pyrophosphate (DMAPP). In the cytosol, IPP and DMAPP are derived from acetyl-CoA through the mevalonate (MVA) pathway and are catalyzed by farnesyl diphosphate synthase (FPPS) to form FPP. Then, cytosolic/mitochondrial terpene synthases (TPSs) convert FPP into various sesquiterpenes [4]. In plastids, IPP and DMAPP, which are synthesized from pyruvate and glyceraldehyde-3-phosphate by the 2-C-methyl-D-erythritol 4-phosphate (MEP) pathway, are converted into geranyl diphosphate (GPP) by GPP synthase (GPPS). Subsequently, plastidial TPSs catalyze the conversion of GPP into various monoterpenes and diterpenes [5]. The TPS gene family is further divided into seven subfamilies, namely, TPS-a to TPS-g [6–8]. Some TPSs involved in the biosynthesis of volatile terpenes have been investigated, including those in Arabidopsis thaliana (Arabidopsis) [9, 10], Rose hybrida (rose) [11], Antirrhinum majus (snapdragon) [12, 13], Nicotiana suaveolens (tobacco) [14], Lilium spp (lily) [15], and Cananga odorata (Ylang Ylang) [16].
Phenylpropanoids and benzenoids are the second largest class of floral volatiles that are generated in plastids via the shikimate pathway [17–18]. Phenylalanine ammonialyase (PAL) is the first committed step in the biosynthesis of benzenoids/phenylpropanoids and converts phenylalanine into trans-cinnamic acid [19]. The production of volatile benzenoids from cinnamic acid requires the shortening of the side chain by a C2 unit via the CoA-dependent β-oxidative pathway or CoA-independent non-β-oxidative pathway [19,20]. The β-oxidative pathway has recently been reported in the model plants Petunia×hybrida (petunia) [21–24] and A. thaliana (Arabidopsis)[25]. Current understanding of the non-β-oxidative pathway, including related enzymes, is limited.
Polianthes tuberosa is a member of the Agavoideae family that originated in Mexico and is mainly recognized for its aromatic oils and ornamental flowers [26]. Currently, it is cultivated in various tropical and subtropical regions. However, no genomic information on this species has been generated to date. When in bloom, the flowers of P. tuberosa emit an intense fragrance that consists of floral volatiles, mainly benzenoids such as methyl benzoate, methyl salicylate, methyl isoeugenol, and benzyl benzoate, and some terpenes, which include 1,8-cineol, farnesene, and germacrene D [27–30]. Previous studies have mainly focused on the composition analysis of floral scents, and the molecular mechanisms underlying the biosynthesis and regulation of these compounds remain unclear. To date, only two PALs and two DXRs have been identified in P. tuberosa[30]. Furthermore, the majority of research investigations that have focused on molecular mechanisms offloral scents involve dicots. In monocots, transcriptome analysis of floral scents has been reported for Hedychium coronarium (garland-flower)[31], Cymbopogon flexuosus (aromatic grasses)[32], Vanda Mimi Palmer (vandaceous orchid)[33], and Lilium ‘Siberia’ (lily)[34]. Thus, our understanding of the molecular mechanism underlying floral scent formation in P. tuberosa is limited.
De novo transcriptome analysis has become a useful tool in the discovery of genes involved in various metabolic pathways, including the determination of sequence and expression patterns without the need for a reference genome [35–37]. In the present study, we used RNA-seq technology to analyze the transcriptome of P. tuberosa flowers during four developmental stages. Four digital gene expression (DGE) libraries were obtained to analyze the gene expression patterns involved in floral scent formation during flowering. The results of this study can improve our understanding of the molecular mechanisms underlying floral scent formation in P. tuberosa as well as provide an important bioinformatics resource for investigating other biological mechanisms.
Methods
Plant material
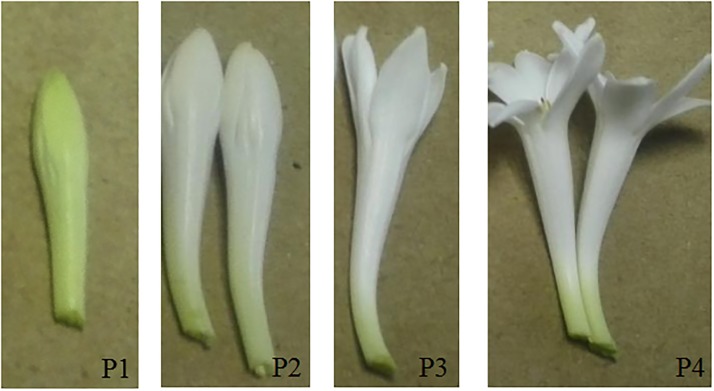
P. tuberosa, which was planted at the Fujian Academy of Agricultural Sciences (Fuzhou, China) in April, bloomed in August. The plant materials used in this study were collected on August 20, 2016 at 5:00 AM. The conditions of the sites included a daily high-low temperature of 35°C/29°C and a light-dark photoperiod cycle of 14 h/10 h. The materials were collected at four different flowering stages, immediately frozen in liquid nitrogen, and stored at -80°C. The four different flower stages were as follows: P1, early bud stage: completely closed petals, green; P2, mid-bud stage, completely closed petals, white; P3, anthesis stage: semi-open petals; and P4, fully bloomed stage: completely open petals (Fig 1).
Fig 1. Floral developmental stages of P. tuberosa.
P1, early bud stage; P2, mid-bud stage; P3, anthesis stage; P4, fully bloomed stage.
Headspace collection and gas chromatography-mass spectrometry (GC-MS) analysis
The volatile compounds were collected using a headspace-solid phase microextraction (HS-SPME)/GC-MS method. The whole flower was sealed in a 20-mL extraction bottle and equilibrated at 50°C for 10 min. The volatile compounds were extracted and adsorbed for 30 min using an SPME fiber [polydimethylsiloxane (PDMS), diameter 65 μm]. Then, the trapped floral scent compounds were analyzed using a Shimadzu GCMS-TQ8040 system. An Rxi-5Sil MS capillary column (30 m × 0.25 mm × 0.25 μm) was used for the separation of volatile compounds with helium as the carrier gas, at a flow rate of 1 mL/min. The column operating conditions were as follows: an initial 50°C hold step for 2 min, followed by an increase to 170°C at a rate of 4°C/min (held for 1 min), then to 270°C at a rate of 20°C/min, and a final hold step for 2 min. The volatiles were identified by comparing the mass spectra and retention times with authentic standards.
RNA extraction, library construction, and RNA-seq
For transcriptome assembly, whole flowers of P. tuberosa (two biological replicates) were collected at the four flower developmental stages (S1 Fig). Total RNA was isolated using a universal RNA extraction kit (Bioteke Corporation, Beijing, China), and quality and quantity were examined with a NanoDrop 2000 UV-vis spectrophotometer (Thermo Scientific, MA, USA) and an Agilent 2100 Bioanalyzer (Agilent Technologies, CA, USA). Library construction was performed by the Biomarker Biotechnology Corporation (Beijing, China) using an mRNA-Seq sample prep kit (Illumina, CA, USA). Finally, the eight libraries were sequenced using an Illumina HiSeq™ 4000 platform.
De novo assembly and functional annotation
Raw reads from the four samples were collected, and low-quality reads were removed. The remaining high-quality clean reads were then extended into longer contigs via their overlapping regions using Trinity software [38], further assembled into transcripts through pair-end joining, and then clustered to unigenes. Based on sequence similarity, all assembled unigenes were compared to public databases such as the NCBI non-redundant protein (Nr) and non-redundant nucleotide (Nt) databases, GO, eukaryotic orthologs groups (KOG), KEGG, clusters of orthologous groups of proteins (COG), Swiss-Prot protein database, protein family (Pfam), and orthologous groups of genes (eggNOG) using an E-value cut-off of 10−5.
Expression annotation
The trimmed reads were aligned to the assembled transcriptome using Bowtie2 [39]. The abundance of each gene was estimated using RSEM (RNA-Seq by Expectation Maximization) [40]. Read counts per gene were calculated using fragments per kilobase of transcript per million mapped reads (FPKM) [41]. DEGs among eight different libraries were identified using DESeq [42]. An absolute false discovery rate < 0.01 and a fold-change value ≥ 2 were used as thresholds to confirm significant differences in expression levels.
Q-PCR analysis
RT-qPCR was performed using the fluorescent dye SYBR Green (TaKaRa,China) on an ABI 7500 real-time PCR System (Applied Biosystems, USA). Primer sequences were designed using PrimerPremier 5.0 software. Actin-1 was employed as an internal control. The PCR conditions were as follows: 95°C for 30 s, followed by 40 cycles of 95°C for 5 s, 60°C for 34 s, 72°C for 30 s, and then 95°C for 15 s, 60°C for 1 min, and 95°C for 15 s. Each reaction was performed in triplicate. Relative expression levels were calculated using the 2-ΔΔCt method [43].
Results
Identification of major floral compounds
To examine the composition of floral scents emitted by flowers, total volatile compounds from four flower developmental stages were analyzed by GC-MS. Low levels of benzenoids and terpenes were detected in green flower buds (P1), whereas these gradually increased at flower anthesis. At the anthesis stage (P3), the majority of the volatiles comprised benzenoids, coupled with lower amounts of terpenes such as methyl benzoate (17.07%), 1,8-cineole (10.00%), methyl salicylate (9.83%), germacrene D (7.68%), indole (6.65%), α-farnesene (4.92%), methyl anthranilate (4.29%), and methyl isoeugenol (3.56%) (Table 1).
Table 1. Relative abundance of volatile compounds in P. tuberosa flowers.
| Compound | Retention time (min) | Relative amount (%) at specific flower developmental stages | |||
|---|---|---|---|---|---|
| P1 | P2 | P3 | P4 | ||
| Benzenoids | |||||
| Methyl benzoate | 12.514 | 4.13 ± 0.20 | 26.75 ± 0.08 | 17.07 ± 0.21 | 7.49 ± 1.63 |
| Methyl salicylate | 16.292 | 3.28 ± 0.43 | 12.99 ± 0.23 | 9.83 ± 0.86 | 7.32 ± 0.32 |
| Indole | 19.959 | — | 1.03 ± 0.14 | 6.65 ± 0.02 | 9.78 ± 0.10 |
| Methyl anthranilate | 21.557 | — | 0.42 ± 0.01 | 4.29 ± 0.88 | 6.43 ± 0.42 |
| Eugenol | 21.949 | — | 0.28 ± 0.04 | 0.95 ± 0.01 | 1.13 ± 0.14 |
| Trans-iseugenol | 25.073 | — | 0.52 ± 0.04 | 2.91 ± 0.11 | 2.32 ± 0.36 |
| Methyl isoeugenol | 26.576 | — | 1.68 ± 0.06 | 3.56 ± 0.14 | 3.23 ± 0.08 |
| Benzyl benzoate | 34.181 | 0.35 ± 0.15 | 1.12 ± 0.11 | 1.09 ± 0.94 | 2.55 ± 0.10 |
| Terpenes | |||||
| 1,8-Cineol | 9.948 | — | 4.46 ± 0.45 | 10.00 ± 0.39 | 9.42 ± 0.47 |
| α-Terpineol | 16.405 | — | 1.35 ± 0.28 | 1.91 ± 0.01 | 2.55 ± 1.17 |
| Geraniol | 18.457 | — | 0.28 ± 0.55 | 1.01 ± 0.08 | 1.15 ± 0.01 |
| β-Bourbonene | 23.032 | 0.47 ± 0.02 | 0.7 ± 0.08 | 1.38 ± 0.41 | 3.29 ± 0.80 |
| Germacrene D | 26.176 | 3.86 ± 0.26 | 5.85 ± 0.16 | 7.68 ± 1.41 | 5.75 ± 0.26 |
| α-Farnesene | 26.884 | — | 2.58 ± 0.11 | 4.92 ± 0.71 | 6.94 ± 0.10 |
| Fatty acid derivatives | |||||
| (E)-2-Hexenal | 2.81 | 7.53 ± 0.04 | 4.95 ± 0.11 | 3.51 ± 0.88 | 5.12 ± 0.13 |
| 2-Pentylfuran | 8.273 | 6.24 ± 0.10 | 2.27 ± 0.23 | 1.34 ±0.21 | 0.99 ± 0.34 |
| 1,3-Hexadiene | 9.869 | 13.03 ± 0.12 | 3.89 ± 0.30 | — | — |
| (E)-2-Octenal | 11.050 | 12.52 ± 0.40 | 4.87 ± 0.23 | 1.49 ± 0.48 | 2.44 ± 0.02 |
| Nonanal | 12.928 | 3.77 ± 0.45 | 1.99 ± 0.01 | 1.10 ± 0.12 | 1.54 ± 0.16 |
| (E, Z)-2,6-Nonadiena | 14.773 | 4.03 ± 0.49 | 2.76 ± 0.11 | 0.40 ± 0.03 | 1.81 ± 0.08 |
| 2-Nonenal | 15.076 | 8.95 ± 0.41 | 3.12 ± 0.07 | 0.32 ± 0.04 | 1.73 ± 0.11 |
| 2,4-Decadienal | 19.993 | 5.41 ± 0.32 | 4.03 ± 0.47 | 0.56 ± 0.02 | 0.35 ± 0.12 |
de novo assembly, annotation of transcriptome, and identification of DEGs
Because no genomic information on P. tuberosa is currently available, transcriptome sequencing and de novo assembly were conducted in this study. A total of 228.71 million clean reads were generated from eight libraries (S4 Table). De novo assembly of these reads resulted in 96,906 unigenes with a mean length of 1,022 bp. The unigene details are shown in Table 2. Of the unigenes, 31,615 were longer than 1000 bp, 32.6% of the total.
Table 2. Quality parameters of the P. tuberosa transcriptome.
| Length (bp) | Unigenes | |
|---|---|---|
| Number | Percentage | |
| 200–300 | 9,519 | 9.82% |
| 300–500 | 23,480 | 24.23% |
| 500–1,000 | 32,292 | 33.32% |
| 1,000–2,000 | 19,453 | 20.07% |
| 2,000+ | 12,162 | 12.55% |
| Total number | 96,906 | |
| Total length | 99,108,304 | |
| N50 length | 1,519 | |
| Mean length | 1,022.726 | |
Of the 96,906 unigenes, 40,563 assembled sequences were annotated using BLAST analysis against eight public databases, accounting for approximately 41.86% of all unigenes, with an E-value ≤1e-5. The annotation details are shown in S5 Table. In total, 23,396 unigenes were grouped into three major categories, molecular function, cellular components, and biological processes by GO annotation (S2 Fig). The number of annotated DEGs in P1 vs. P2 was higher than that in P2 vs. P3 and P3 vs. P4, suggesting that a large number of genes may be involved in flowering, particularly at the initial developmental stage. However, a large number of assembled sequences remained unannotated, which may be due to non-coding RNAs, untranslated regions, or specifically expressed in P. tuberosa. The special expression genes can be a great resource for the identification and study of novel genes [44, 45].
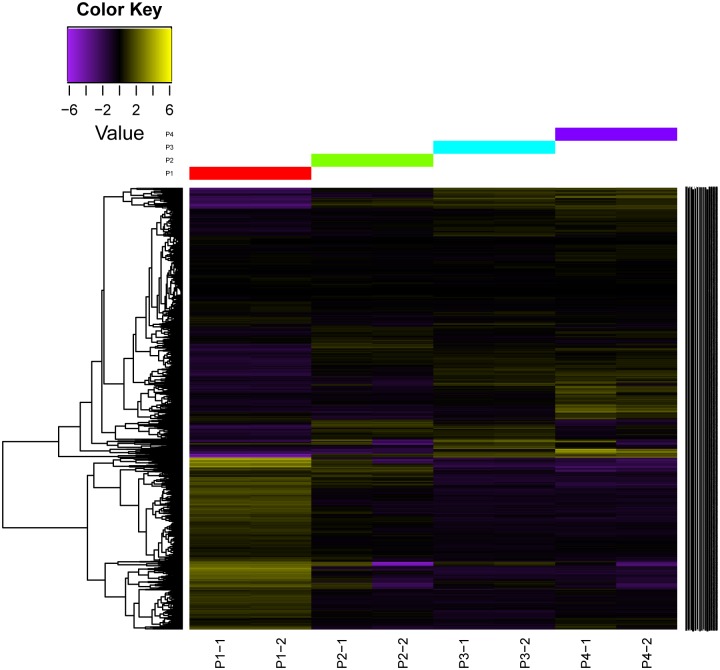
Upon flowering, vast quantities of benzenoid volatiles accumulated in P. tuberosa from P1-P3, then decreased from P3-P4. To identify and select DEGs during flowering, the expression level of each gene in the eight libraries was compared in pairs of consecutive stages. The DEGs details are presented in S3 Fig. The number of DEGs in P1 vs. P2 was higher than that in P2 vs. P3 and P3 vs. P4, suggesting that more complex biological events occur during the initial stage of development. The transcript abundances were clustered by hierarchical cluster analysis (Fig 2). Expression analysis revealed that P1 had a pronounced difference from P2, P3 and P4, while a similar pattern was observed between P2, P3 and P4.
Fig 2. Expression profiles and cluster analysis of unigenes during four stages of flowering development.
Secondary metabolic pathways identified in flowering
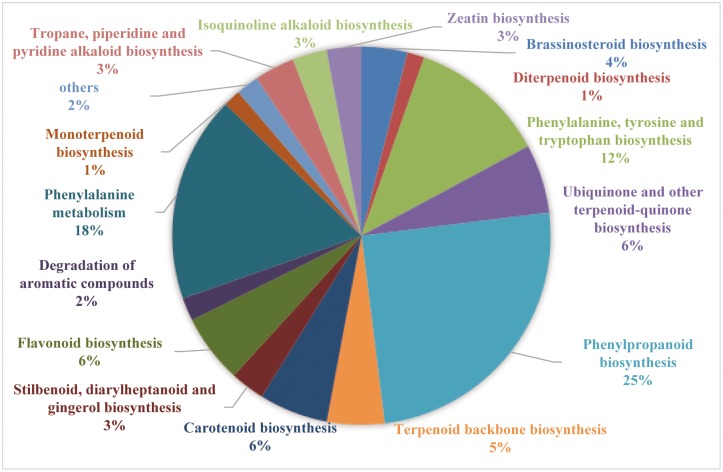
To identify unigenes related to secondary metabolic pathways, KEGG annotation was conducted by mapping the reference canonical pathways. About 14,771 (18.52%) unigenes were assigned to 113 KEGG pathways. Of these, 21 pathways of secondary metabolism were identified (184 DEGs). Phenylpropanoid biosynthesis and phenylalanine metabolism was the largest group (Fig 3), which was consistent with large amounts of benzenoid volatiles. Also, large numbers of DEGs involved in secondary metabolites, like terpenoid backbone biosynthesis, ubiquinone, and other types of terpenoid-quinone biosynthesis, monoterpenoid biosynthesis, and diterpenoid biosynthesis were represented, which was consistent with large amounts of some terpene volatiles. These unigene analyses are a valuable resource for gene mining and functional analysis of P. tuberosa.
Fig 3. KEGG analysis of DEGs involved in secondary metabolism during flowering.
Identification of putative genes related to benzenoid biosynthesis
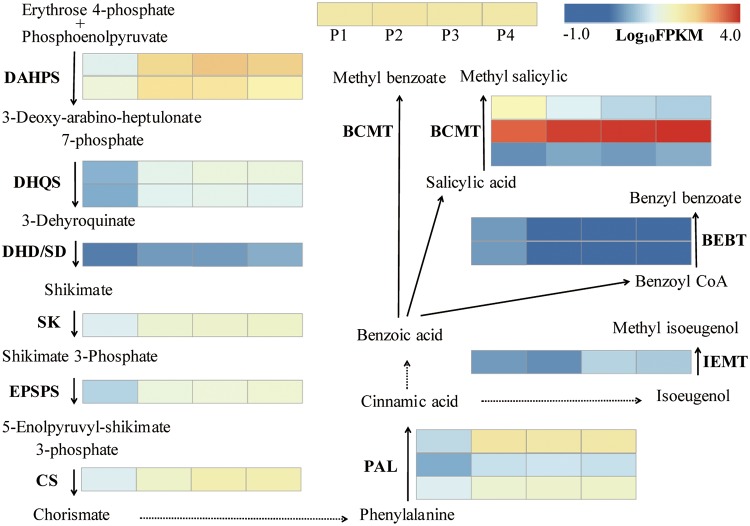
Eight genes of the shikimate pathway, which supplies the carbon flux for benzenoid biosynthesis in plants, were here identified. These are two 3-deoxy-7-phosphoheptulonate synthases (DAHPSs) and 3-dehydroquinate synthases (DHQSs), one each of dehydroquinate dehydratase/shikimate dehydrogenase (DHD/SHD), shikimate kinase (SK), 3-phospho shikimate 1-carboxyvinyltransferase (EPSPS), and the chorismate synthase (CS) gene. The expression levels of the eight genes during flowering development are depicted in Fig 4. DAHPS represents the first reaction step, which converts erythrose 4-phosphate and phosphoenolpyruvate into 3-deoxy-arabino-heptulonate 7-phosphate [46]. Both PtDAHP1 and PtDAHP2 contained an N-terminal chloroplasttransit peptide (S4 Fig) and had a typical class-II DAHP synthase family domain (PLN02291). Homology analysis showed the predicted amino acid sequence of PtDAHP1 and PtDAHP2 to be very highly conserved in core sequences, and both of them were upregulated during flowering, suggesting that they play key roles in regulating the shikimate pathway. The following six genes were upregulated from the P1–P4 stages, indicating the predominant role of benzenoid volatiles in P. tuberosa (Fig 4, S5 Fig).
Fig 4. Expression profiles of genes in the shikimate and benzenoid biosynthesis.
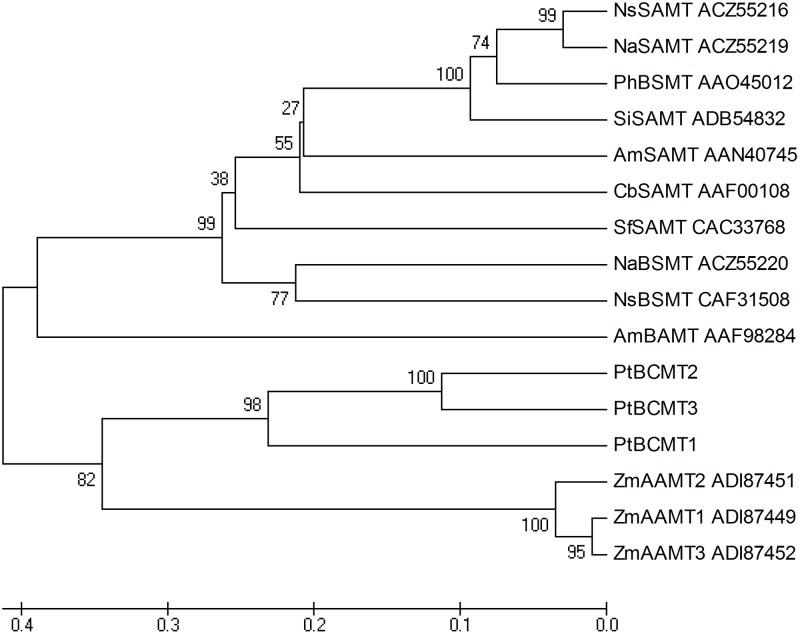
To further investigate benzenoid biosynthesis in P. tuberosa and to confirm the identities of related genes in the pathway, the putative genes of benzenoid biosynthesis were investigated using sequence homology analysis. PAL is the first rate-limiting enzyme in benzenoid biosynthesis. It converts phenylalanine into cinnamic acid [19]. Three PtPAL genes were detected in the assembled transcriptome. Phylogenetic analysis of the corresponding amino acid sequences showed three PtPALs clustered into one phylogenetic group (S6 Fig), and their expression levels increased during flowering, particularly that of PtPAL1, which significantly increased from P1 to P2 and remained high through P3 and P4 (Fig 4). From the tree, PtPAL2 closely matched the PtPAL3. This may be due to gene duplication events, which may have mutated or diverged later from the common ancestor. In the final step, floral volatiles such as methyl isoeugenol, benzyl benzoate, methyl benzoate, methyl salicylate, and methyl anthranilate are synthesized. This process is catalyzed by S-adenosyl-L-methionine:(iso)eugenol O-methyltransferase(IEMT), benzyl alcohol benzoyl transferase (BEBT), and benzenoid carboxyl methyltransferase (BCMT). BCMT include salicylic acid methyltransferase (SAMT), benzoic acid carboxyl methyltransferase (BAMT), anthranilic acid methyltransferase (AAMT), and benzoic acid/salicylic acid carboxyl methyltransferases (BSMT) [47, 48]. Phylogenetic analysis of the deduced amino acid sequence showed that three PtBCMTs are very similar to maize AAMT but are different from other dicot BAMTs, BSMTs, and SAMTs (Fig 5). PtBCMT2 was upregulated during the four stages of flower development, whereas the expression of PtBCMT1 decreased and that of PtBCMT3 remained low. The expression of two PtBEBTs decreased sharply during flowering development (Fig 4). PtIEMT was upregulated during flowering development, which coincides with the emission of methyl isoeugenol volatiles.
Fig 5. Phylogenetic reconstruction of the PtBCMTs and other BCMTs.
Ns, Nicotiana suaveolens; Na, N.alata; Ph, Petunia hybrida; Sl, Solanum lycopersium; Am, Antirrhinum majus; Sb, Sorghum bicolor; Sf, Stephanotis floribunda; Zm, Zea mays.
Identification of putative genes related to terpene biosynthesis
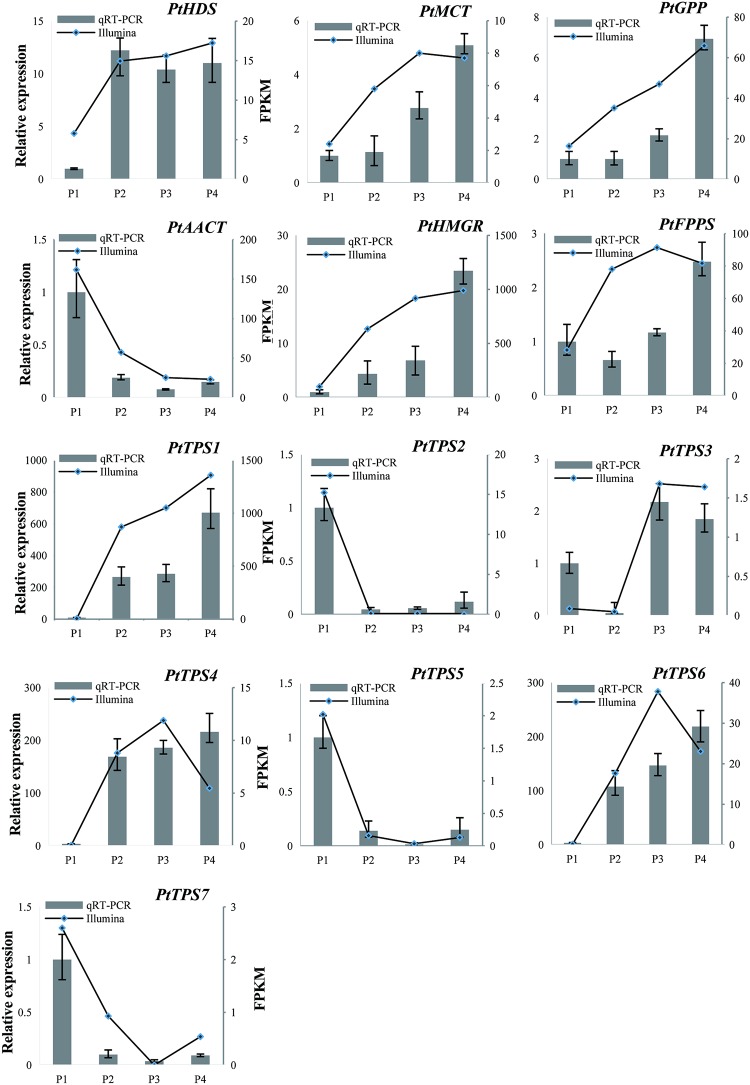
The biosynthesis and emission of terpenes have been investigated in various plants. To establish further details on terpene biosynthesis in P. tuberosa, genes related to terpene biosynthesis were mined. The identified terpenoid biosynthesis genes involved in the MEP and MVA pathways were as follows: 4-hydroxy-3-methylbut-2-en-1-yl diphosphate synthase (HDS), GPPS, 2-C-methyl-D-erythritol 4-phosphate cytidylyltransferase (MCT), FPPS, acetyl-CoA acetyltransferase (AACT), and hydroxymethylglutaryl-CoA reductase (HMGR). DXS and AACT are the first enzymes of the MEP and MVA pathways, respectively. Six PtDXS were observed, all of which had very low expression levels, and PtAACT was significantly downregulated from P1–P4, indicating the minor role of terpene volatiles in P. tuberosa. PtGPPS and PtFPPS catalyze two common isoprene precursors (IPP and DMAPP) to produce GPP and FPP, which are the precursors of monoterpenes and sesquiterpenes [49]. Only one PtGPPS and PtFPPS was identified in the P. tuberosa transcriptome, which showed a 4.01- and 2.90-fold increase in expression from the P1 to P4 stages, respectively (Fig 6), indicating the crucial role in terpene biosynthesis.
Fig 6. qRT-PCR validation of selected unigenes in the MEP, MVA, and terpene biosynthetic pathways.
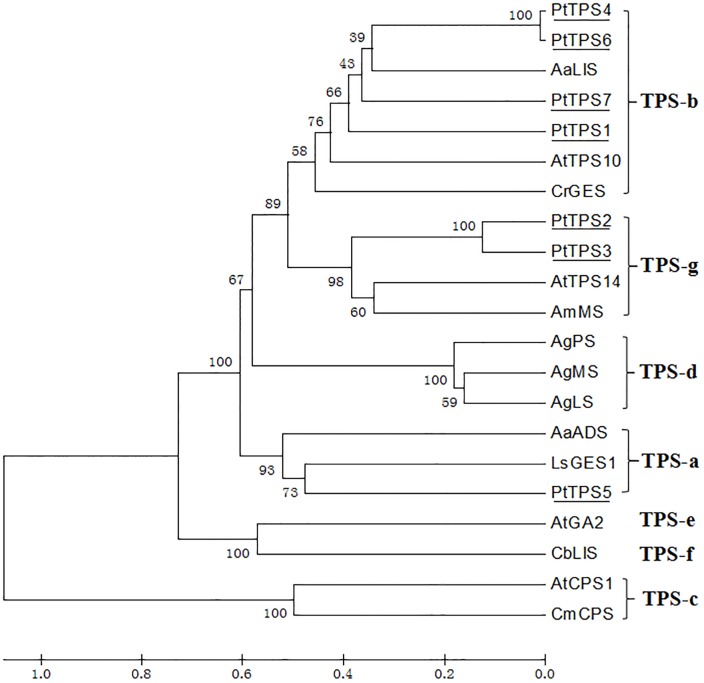
TPSs exclusively catalyze GPP and FPP to monoterpenes and sesquiterpenes, respectively. P. tuberosa RNA-seq analysis identified four complete and three partial TPSs. Phylogenetic analysis of seven PtTPSs revealed that PtTPS1, PtTPS4, PtTPS6, and PtTPS7 could be clustered into the TPS-b subfamily, which consisted mainly of mono-TPSs, whereas PtTPS5 belonged to the TPS-a subfamily, which was composed of sesqui-TPSs, and PtTPS2 and PtTPS3 were assigned to the TPS-g subfamily (Fig 7). The deduced amino acids of PtTPS2, PtTPS4, PtTPS5, and PtTPS6 contained the motif DDXXD, which is a conserved domain in TPSs (S7 Fig). The expression of PtTPS1 was significantly higher than that of the other six PtTPSs, suggesting that it played an important role in terpene biosynthesis.
Fig 7. Phylogenetic analysis TPSs of P. tuberosa.
A: Phylogenetic reconstruction of P. tuberosaTPSswith other plant TPSs using the MEGA 5 program; B: Alignment of deduced amino acid sequences of PtTPS2, PtTPS4, PtTPS5, and PtTPS6. AaLIS, linalool synthase (AAF13356); AtTPS10, Arabidopsis thaliana myrcene/ocimene synthase (AAG09310); CrGES, Catharanthus roseus geraniol synthase (AFD64744); AtTPS14, A. thaliana terpene synthase 14 (NP176361); AmMS, Antirrhium majus myrcene synthase (AAO41727); AgPS, Abies grandis pinene synthase (AAB71085); AgMS, Abies grandis myrcene synthase (AAB71084); AgLS, Abies grandis 4S-limonene synthase (AAB70907); AaADS, amorpha-4,11-dienesynthase (AFA34434); LsGES1, Lactuca sativus germacrene A synthase (AAM11626); AtGA2, A. thaliana ent-kaurene synthase (AAC39443); CbLIS, Clarkia breweri S-linalool synthase (AAC49395); AtCPS1, A. thaliana copalyl diphosphate synthase (NP_192187); and CmCPS, Cucurbita maxima copalyl diphosphate synthase (AAD04292).
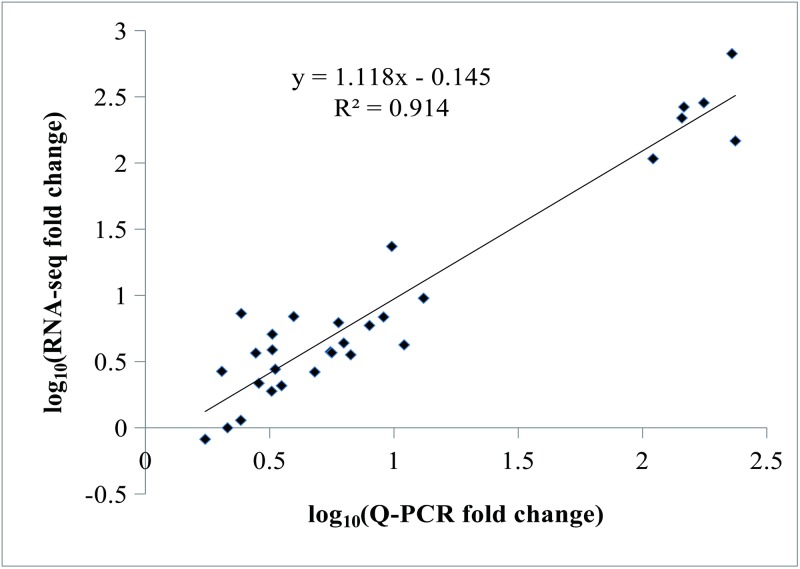
Based on the log10 fold-changes in expression observable in the scatter plot (Fig 8), a positive correlation was observed between the results of qRT-PCR and RNA-Seq analyses (R2 = 0.914).
Fig 8. Scatter plot of 10 selected genes using RNA-seq and qRT-PCR analysis.
The RNA-Seq fold-change represents the FPKM ratios of P2 (P3, P4) to P1; Q-PCR fold change refers to the relative expression from P2 (P3, P4) to P1.
Discussion
P. tuberosa is a popular flower with a distinct floral scent. However, our understanding of the molecular mechanisms responsible for its floral scent is limited because genomic information on this species is currently unavailable. The present study utilized extensive cDNA sequence data to identify genes that control floral scent compounds and to further analyze floral scent biosynthesis in P. tuberosa. Here, eight cDNA libraries during flowering of P. tuberosa were obtained through transcriptome sequencing. A total of 96,906 unigenes was obtained. Approximately 41.85% of the assembled unigene sequences were annotated by public databases. The availability of data for P. tuberosa provides an important resource for exploring the characteristics of this species as well as for mining key genes and their functions in floral scent development.
Floral scents mainly consist of terpenoids and benzenoids/phenylpropanoids, which are made up of a complex mixture of low-molecular-mass volatiles. The study of both volatile compounds and gene expression profiling have proven to be very powerful as a means of identifying candidate genes involved in the formation of floral scent. In Arabidopsis, Chen first detected floral volatile compounds from flowers. Then, candidate AtTPS genes were explored using the genomic information and RT-PCR [9]. In the absence of genome sequences, a combination of expressed sequence tags (EST) and metabolic profiling could be used to investigate scent-related genes, such as those in petunias [50], roses [11], and snapdragons [51]. In the present study, transcriptome sequencing combined with volatile compounds was used to explore floral scent formation. The results of GC-MS showed that the majority of floral volatile compounds in P. tuberosa were benzenoid compounds, which included methyl benzoate (17.07%), methyl salicylate (9.83%), indole (6.65%), methyl anthranilate (4.29%), and methyl isoeugenol (3.56%) (Table 1). High levels of some terpenes, including 1,8-cineole (10.00%), germacrene D (7.68%), and α-farnesene (4.92%), were also detected. This was consistent with the high volatile content previously reported in P. tuberosa flowers, which was attributed to benzenoid compounds and terpenes [29–30]. We also identified 13 and 17 candidate genes in P. tuberosa that were related to terpene and benzenoid biosynthesis, respectively. These genes and their expression patterns are an important resource for exploring floral scent in P. tuberosa. Several candidate genes, such as PtTPS1, PtDAHPSs, PtPAL1, and PtBCMT2, which are up-regulated and related to high contents of corresponding floral volatiles, might play crucial roles in floral scent.
The floral volatile compounds of P. tuberosa mainly include benzenoids/phenylpropanoids. In plants, the shikimate pathway, which provides metabolic flux to secondary metabolites, is the precursor in benzenoid biosynthesis. It is mostly regulated at the gene expression level [52]. DAHPS is the first enzyme in the shikimate pathway (Fig 4) [47]. It has been found that many plants contained two DAHPS genes, and the gene was upregulated in response to demands for increased emission of benzenoid floral volatiles [52–54]. In petunias, PhDAHP1 was upregulated during flowering and RNA interference (RNAi) suppression of PhDAHP1 revealed that emission of floral benzenoids decreased significantly, while RNAi of PhDAHP2 in which transcript remained constitutive showed no change in emission of floral benzenoids, suggesting that PhDAHP1 was responsible for the benzenoid/phenylpropanoid biosynthesis [54]. In P. tuberosa, there were a total of two DAHPS genes, PtDAHPS1 and PtDAHPS2, which were significantly upregulated during flowering and remained at high levels in P3, appeared to play crucial roles in the shikimate pathway. The expression level of most genes in the shikimate pathway were upregulated during flowering. These were related to the high levels of benzenoid emissions, suggesting that the enzymes of shikimate pathway are important to the formation of benzenoid volatiles.
PAL catalyzed the transition of phenylalanine to trans-cinnamate and directed the carbon flow from the shikimate pathway to phenypropanoid metabolism. Several copies of PAL genes were found in some plant species, such as four genes in Arabidopsis, five in poplars and nine in rice [55]. The PAL genes had different functions in poplars and Arabidopsis, [56]. A total of three PtPALs were found in P. tuberosa, all of which had different expression patterns, consistent with their different functions. Of these, PtPAL1 significantly increased from P1–P2 and remained high in P3 and P4, indicating the important role in the formation of benzenoid volatiles. BCMT, which catalyzes the biosynthesis of methyl benzoate, methyl salicylate, and methyl anthranilate, has been extensively investigated in petunia and snapdragon [31, 48, 49]. Phylogenetic analysis showed three PtBCMTs to be closely related to maize AAMT but distantly related to other BAMTs, BSMTs, and SAMTs (Fig 5). This may be because the enzymes in this group have broad substrate specificity and are more similar to each other in terms of catalyzing different substrates in this species than enzymes having the same functions in other species [57]. Only PtBCMT2 showed sustained upregulation of expression. It also showed a positive correlation with the emission of methyl benzoate (Table 1, Fig 4). PtBCMT2 seems to be primarily responsible for the biosynthesis of methyl benzoate. BEBT is the final enzyme in the biosynthesis of benzyl benzoate (Fig 4). The expression level of two PtBEBTs was low and decreased sharply during flowering, which coincides with the low benzyl benzoate content during flowering. Previous studies have shown the volatile emission in P. tuberosa to be rhythmic. Peak levels of benzyl benzoate were observed from evening until midnight, but these were relatively low during the day [30], and the expression level of BEBT was closely coordinated with benzyl benzoate emissions [30]. In our study, the activity of BEBT was low, which may reflect the time of our sample collection (5:00 AM).
DXS catalyzes the first step of the MEP pathway, which is the pathway upstream of monoterpene production. Transgenic plants accumulate different levels of monoterpene compounds than wild-type, so DXS acts as a limiting enzyme to control the biosynthesis of monoterpene [58]. In the present study, a total of six PtDXSs were identified, but all showed low expression levels. This may be attributable to the relatively low monoterpenecontents because benzenoid compounds are the main volatiles. TPSs convert GPP or FPP to various terpenes via a one-step method, and plants have a wide range of TPSs [59]. In Hedychium coronarium, most of the 17 TPSs were upregulated during flowering development [31]. We found approximately 18 TPSs from the transcriptome data of P. tuberosa. However, only 7 of the 18 PtTPS transcripts had relatively high expression levels, whereas the rest of the 11 exhibited very low expression levels. This may be because terpene volatiles are not the main volatile compounds of this species, and these genes have functions other than those involving volatile synthesis, and some have even completely lost their functions [59]. Some plant models possess 24–69 TPS genes [59]. Some TPS genes are expressed in specific tissues or need to be induced [6], indicating that there may be more TPS genes in P. tuberosa that have yet to be identified. TPSs are generally classified into seven clades, and TPSs mainly belong to only two or three clades in some plants, revealing the expansion of specific types of genes [59]. Among the 7 TPSs, four belong to TPS-b, one to TPS-a, and two to TPS-g. In higher plants, terpene emissions are regulated by the pattern of expression of TPS genes [4]. In snapdragons, the emission of volatile terpenes, such as (E)-β-ocimene and myrcene, are closely associated with expression of their related TPS genes during flowering [12]. PtTPS1 which belonged to TPS-b (consisted mainly of mono-TPSs) showed the highest expression levels from P2 to P4, suggesting that this gene plays a crucial role in the biosynthesis of terpenes. The transcription level of PtTPS1 showed positive correlation with the emission of 1,8-cineol, which is present in large quantities in terpene compounds (Table 1, Fig 6). It may be that PtTPS1s are cineol synthase, which are involved in1,8-cineol formation.
Supporting information
(DOCX)
(XLS)
(XLS)
(DOCX)
(DOCX)
(DOCX)
(DOCX)
(DOCX)
(DOCX)
(DOCX)
(DOCX)
(DOCX)
Abbreviations
- AACT
acetyl-CoA acetyltransferase
- AAMT
anthranilic acid methyltransferase
- BAMT
benzoic acid carboxyl methyltransferase
- BCMT
benzenoid carboxyl methyltransferase
- BEBT
benzyl alcohol benzoyl transferase
- BSMT
benzoic acid/salicylic acid carboxyl methyltransferases
- CS
chorismate synthase
- DAHPS
3-deoxy-7-phosphoheptulonate synthase
- DHD/SHD
dehydroquinate dehydratase/shikimate dehydrogenase
- DHQS
3-dehydroquinate synthase
- DMAPP
dimethylallyl pyrophosphate
- EPSPS
3-phospho shikimate 1-carboxyvinyltransferase
- FPKM
fragments per kilobase of transcript per million mapped reads
- FPPS
arnesyl pyrophosphate synthase
- GPPS
geranyl pyrophosphate synthase
- HDS
4-hydroxy-3-methylbut-2-en-1-yl diphosphate synthase
- HMGR
hydroxymethylglutaryl-CoA reductase
- IEMT
S-adenosyl-L-methionine: (iso)eugenol O-methyltransferase
- IPP
isopentenyl pyrophosphate
- MCT
2-C-methyl-D-erythritol 4-phosphate cytidylyltransferase
- MEP
methyl-erythritol-phosphate
- MVA
mevalonic acid
- PAL
phenylalanine ammonialyase
- SAMT
salicylic acid methyltransferase
- SK
shikimate kinase
- TPSs
terpene synthase
Data Availability
The raw sequences were found in NCBI SRA (accession SRP126470); the assembled sequences were found in NCBI TSA (accession GGEA00000000).
Funding Statement
Special Project of Public Welfare Research Institutes in Fujian (2018R1026-6), and Innovation Team Project of the Fujian Academy of Agricultural Science (STIT2017-2-9) supported this study. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Knudsen JT, Eriksson R, Gershenzon J, Stahl B. Diversity and distribution of floral scent. Botanical Review. 2006; 72: 1–120. [Google Scholar]
- 2.Muhlemann JK, Klempien A, Dudareva N. Floral volatiles: from biosynthesis to function. Plant, Cell and Environment. 2014; 37, 1936–1949. 10.1111/pce.12314 [DOI] [PubMed] [Google Scholar]
- 3.Dudareva N, Klempien A, Muhlemann JK, Kaplan I. Biosynthesis, function and metabolic engineering of plant volatile organic compounds. New Phytologist.2013; 198: 16–32. 10.1111/nph.12145 [DOI] [PubMed] [Google Scholar]
- 4.Tholl D. Terpene synthases and the regulation, diversity and biological roles of terpene metabolism. Curr Opin Plant Biol. 2006;9(3):297–304. 10.1016/j.pbi.2006.03.014 [DOI] [PubMed] [Google Scholar]
- 5.Vranova E, Coman D, Gruissem W. Network analysis of the MVA and MEP pathways for isoprenoid synthesis. Annu Rev Plant Biol. 2013;64:665–700. 10.1146/annurev-arplant-050312-120116 [DOI] [PubMed] [Google Scholar]
- 6.Bohlmann J, Meyer-Gauen G, Croteau R. Plant terpenoid synthases: molecular biology and phylogenetic analysis. Proceedings of the National Academy of Sciences, US.1998; 95:4126–4133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lee S, Chappell J. Biochemical and genomic characterization of terpene synthases in Magnolia grandiflora. Plant Physiology. 2008; 147: 1017–1033. 10.1104/pp.108.115824 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Martin DM, Aubourg S, Schouwey MB, Daviet L, Schalk M, Toub O, et al. Functional annotation, genomeorganization and phylogeny of the grapevine (Vitis vinifera) terpene synthase gene family based on genome assembly, FLcDNA cloning, and enzyme assays. BMC Plant Biology.2010; 10:226 10.1186/1471-2229-10-226 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Chen F, Tholl D, D’Auria JC, Farooq A, Pichersky E, Gershenzon J.Biosynthesis and emission of terpenoid volatiles from Arabidopsis flowers. Plant Cell. 2003;15(2):481–494. 10.1105/tpc.007989 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Tholl D, Chen F, Petri J, Gershenzon J, Pichersky E. Two sesquiterpene synthases are responsible for the complex mixture of sesquiterpenes emitted from Arabidopsis flowers. Plant J. 2005;42(5):757–771. 10.1111/j.1365-313X.2005.02417.x [DOI] [PubMed] [Google Scholar]
- 11.Guterman I, Shalit M, Menda N, Piestun D, Dafny-Yelin M, Shalev G, et al. Rose scent: Genomics approach to discovering novel floral fragrance-related genes. Plant Cell. 2002;14(10):2325–2338. 10.1105/tpc.005207 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Dudareva N, Martin D, Kish CM, Kolosova N, Gorenstein N, Faldt J, et al. (E)-beta-ocimene and myrcene synthase genes of floral scent biosynthesis in snapdragon: function and expression of three terpene synthase genes of a new terpene synthase subfamily. Plant Cell.2003;15(5):1227–1241. 10.1105/tpc.011015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Nagegowda DA, Gutensohn M, Wilkerson CG, Dudareva N. Two nearly identical terpene synthases catalyze the formation of nerolidol and linalool in snapdragon flowers. Plant J. 2008;55(2):224–239. 10.1111/j.1365-313X.2008.03496.x [DOI] [PubMed] [Google Scholar]
- 14.Roeder S, Hartmann A, Effmert U, Piechulla B. Regulation of simultaneous synthesis of floral scent terpenoids by the 1,8-cineole synthase of Nicotiana suaveolens. Plant Mol Biol. 2007;65(1–2):107–124. 10.1007/s11103-007-9202-7 [DOI] [PubMed] [Google Scholar]
- 15.Zhang HX, Hu ZH, Leng PS, Wang WH, Xu F, Zhao J. Qualitative and quantitative analysis of floral volatile components from different varieties of Lilium spp. Scientia Agri Sin. 2013; 46(4): 790–799. [Google Scholar]
- 16.Jin J, Kim MJ, Dhandapani S, Tjhang JG, Yin JL,Wong L, et al. The floral transcriptome of ylang ylang (Cananga odorata var. fruticosa) uncovers biosynthetic pathways for volatileorganic compounds and a multifunctional and novel sesquiterpene synthase. Journal of Experimental Botany. 2015; 66(13): 3959–3975. 10.1093/jxb/erv196 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Orlova I, Marshall-Colón A, Schnepp J, Wood B, Varbanova M, Fridman E. Reduction of benzenoid synthesis in petunia flowers reveals multiple pathways to benzoic acid and enhancement in auxin transport. The Plant Cell.2006; 18: 3458–3475. 10.1105/tpc.106.046227 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Vogt T. Phenylpropanoid biosynthesis. Molecular Plant.2010; 3:2–20. 10.1093/mp/ssp106 [DOI] [PubMed] [Google Scholar]
- 19.Wildermuth MC. Variations on a theme: synthesis and modification of plant benzoic acids. Curr Opin Plant Biol. 2006;9(3):288–296. 10.1016/j.pbi.2006.03.006 [DOI] [PubMed] [Google Scholar]
- 20.Boatright J, Negre F, Chen XL, Kish CM, Wood B, Peel G, et al. Understanding in vivo benzenoid metabolism in petunia petal tissue. Plant Physiol. 2004;135(4):1993–2011. 10.1104/pp.104.045468 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Klempien A, Kaminaga Y, Qualley A, Nagegowda DA, Widhalm JR, Orlova I,et al. Contribution of CoA ligases to benzenoid biosynthesis in petunia flowers. Plant Cell. 2012;24(5):2015–2030. 10.1105/tpc.112.097519 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Qualley AV, Widhalm JR, Adebesin F, Kish CM, Dudareva N. Completion of the core beta-oxidative pathway of benzoic acid biosynthesis in plants. P Natl Acad Sci Usa. 2012;109(40):16383–16388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Van Moerkercke A, Schauvinhold I, Pichersky E, Haring MA, Schuurink RC. A plant thiolase involved in benzoic acid biosynthesis and volatile benzenoid production. Plant J. 2009;60(2):292–302. 10.1111/j.1365-313X.2009.03953.x [DOI] [PubMed] [Google Scholar]
- 24.Colquhoun TA, Marciniak DM, Wedde AE, Kim JY, Schwieterman ML, Levin LA, et al. A peroxisomally localized acyl-activating enzyme is required for volatile benzenoid formation in a Petunia×hybrida cv. ‘Mitchell Diploid’flower. J Exp Bot. 2012;63(13):4821–4833. 10.1093/jxb/ers153 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Bussell JD, Reichelt M, Wiszniewski AAG, Gershenzon J, Smith SM.Peroxisomal ATP-binding cassette transporter COMATOSE and the multifunctional protein ABNORMAL INFLORESCENCE MERISTEM are required for the production of benzoylated metabolites in Arabidopsis seeds. Plant Physiol. 2014;164(1):48–54. 10.1104/pp.113.229807 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Waithaka K, Reid MS, Dodge LL. Cold storage and flower keeping quality of cut tuberose (Polianthes tuberosa L.). J Hortic Sci Biotechnol. 2001; 76:271–275 [Google Scholar]
- 27.Rakthaworn P, Dilokkunanant U, Sukkatta U, Vajrodaya S, Haruethaitanasan V, Pitpiangchan P, et al. Extraction methods for tuberose oil and their chemical components. Kasetsart J (Nat. Sci.). 2009; 43(5): 204–211. [Google Scholar]
- 28.Kanani M, Nazarideljou MJ. Methyl Jasmonate and α-aminooxi-β-phenyl Propionic Acid Alter Phenylalanine Ammonia-Lyase Enzymatic Activity to Affect the Longevity and Floral Scent of Cut Tuberose. Hortic Environ Biotechnol. 2017; 58(2): 136–143. [Google Scholar]
- 29.Maiti S, Mitra A. Morphological, physiological and ultrastructural changes in flowers explain the spatio-temporal emission of scent volatiles in Polianthes tuberosa L. Plant Cell Physiol. 2017; 58: 2095–2111. 10.1093/pcp/pcx143 [DOI] [PubMed] [Google Scholar]
- 30.Maiti S, Moon UR, Bera P, Samanta T, Mitra A. The in vitro antioxidant capacities of Polianthes tuberosa L. flower extracts. Acta Physiol. Plant. 2014; 36: 2597–2605. [Google Scholar]
- 31.Yue Y, Yu R, Fan Y. Transcriptome profiling provides new insights into the formation of floral scent in Hedychium coronarium. BMC Genomics. 2015; 16:470 10.1186/s12864-015-1653-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Meena S, Kumar SR, Venkata Rao DK, Dwivedi V, Shilpashree HB, Rastogi S, et al. De novo sequencing and analysis of lemongrass transcriptome provide first insights into the essential oil biosynthesis of Aromatic Grasses. Front. Plant Sci. 2016; 7:1129 10.3389/fpls.2016.01129 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Toh C, Mohd Hairul AR, Ain NM, Namasivayam P, Go R, Abdullah NAP, Abdullah MO, Abdullah JO. Floral micromorphology and transcriptome analyses of a fragrant Vandaceous Orchid, Vanda Mimi Palmer, for its fragrance production sites. BMC Res Notes. 2017; 10:554 10.1186/s13104-017-2872-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Hu Z, Tang B, Wu Q, Zheng J, Leng P, Zhang K.Transcriptome Sequencing Analysis Reveals a Difference in Monoterpene Biosynthesis between Scented Lilium ‘Siberia’ and Unscented Lilium ‘Novano’. Front. Plant Sci. 2017; 8:1351 10.3389/fpls.2017.01351 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Martin JA, Wang Z. Next-generation transcriptome assembly. Nat Rev Genet. 2011; 12: 671–682. 10.1038/nrg3068 [DOI] [PubMed] [Google Scholar]
- 36.Ward JA, Ponnala L, Weber CA. Strategies for transcriptome analysis in nonmodel plants. Am J Bot.2012; 99: 267–276. 10.3732/ajb.1100334 [DOI] [PubMed] [Google Scholar]
- 37.Hamilton JP, Buell CR. Advances in plant genome sequencing. Plant J. 2012; 70: 177–190. 10.1111/j.1365-313X.2012.04894.x [DOI] [PubMed] [Google Scholar]
- 38.Grabherr GM, Haas BJ, Yassour M, Levin ZJ, Thompson AD, Amit I, et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. NatBiotechnol. 2011; 29: 644–652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Langmead B, Trapnell C, Pop M, Salzberg SL. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biology Italic. 2009; 10(3): R25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Li B, Colin ND. RSEM: accurate transcript quantification from RNA Seq data with or without a reference genome. BMC Bioinformatics Italic. 2011; (12):323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol. 2010;28:511–515. 10.1038/nbt.1621 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Anders S, Huber W. Differential expression analysis for sequence count data. Genome Biol. 2010;11:R106 10.1186/gb-2010-11-10-r106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2-DELTADELTACT method. Methods.2001;25(4):402–408. 10.1006/meth.2001.1262 [DOI] [PubMed] [Google Scholar]
- 44.Chung PJ, Jung H, Jeong DH, Ha SH, Choi YD, Kim JK. Transcriptome profiling of drought responsive noncoding RNAs and their target genes in rice. BMC Genomics. 2016;17:563 10.1186/s12864-016-2997-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Kwenda S, Birch PR, Moleleki LN. Genome-wide identification of potato long intergenic noncoding RNAs responsive to Pectobacterium carotovorum subspecies brasiliense infection. BMC Genomics. 2016;17:614 10.1186/s12864-016-2967-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Tzin V, Malitsky S, Ben Zvi MM, Bedair M, Sumner L, Aharoni A, et al. Expression of a bacterial feedback-insensitive 3-deoxy-D-arabino-heptulosonate7-phosphate synthase of the shikimate pathway in Arabidopsis elucidates potential metabolic bottlenecks between primary and secondary metabolism. New Phytol. 2012;194(2):430–439. 10.1111/j.1469-8137.2012.04052.x [DOI] [PubMed] [Google Scholar]
- 47.Murfitt LM, Kolosova N, Mann CJ, Dudareva N. Purification and characterization of S-adenosyl-L-methionine: benzoic acid carboxyl methyltransferase, the enzyme responsible for biosynthesis of the volatile ester methyl benzoate in flowers of Antirrhinum majus. Arch Biochem Biophys. 2000;382(1):145–151. 10.1006/abbi.2000.2008 [DOI] [PubMed] [Google Scholar]
- 48.Negre F, Kish CM, Boatright J, Underwood B, Shibuya K, Wagner C, et al. Regulation of methylbenzoate emission after pollination in snapdragon and petunia flowers. Plant Cell. 2003;15(12):2992–3006. 10.1105/tpc.016766 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Vandermoten S, Haubruge E, Cusson M. New insights into short-chain prenyltransferases: structural features, evolutionary history and potential for selective inhibition. Cell Mol Life Sci. 2009;66(23):3685–3695. 10.1007/s00018-009-0100-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Verdonk JC, Haring MA, van Tunen AJ,Schuurink RC.ODORANT1 regulates fragrance biosynthesis in petunia flowers. Plant Cell. 2005;17:1612–1624 10.1105/tpc.104.028837 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Muhlemann JK, Maeda H, Chang CY, Miguel PS, Baxter I, Cooper B, et al. Developmental changes in the metabolic network of snapdragon flowers. 2012; Plos One 7, e40381 10.1371/journal.pone.0040381 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Tzin V and Galili G. The biosynthetic pathways for shikimate and aromatic amino acids in Arabidopsis thaliana In The Arabidopsis Book Torii K., ed. (Rockville MD: The American Society of Plant Biologists; ), 2010; p. e0132 10.1199/tab.0132 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Maeda H and Dudareva N. The shikimate pathway and aromatic amino acid biosynthesis in plants. Annu. Rev. Plant Biol. 2012; 63:73–105. 10.1146/annurev-arplant-042811-105439 [DOI] [PubMed] [Google Scholar]
- 54.Langer KM, Jones CR, Jaworski EA, Rushing GV, Kim JY, Clark DG, et al. PhDAHP1 is required for floral volatile benzenoid/phenylpropanoid biosynthesis in Petunia x hybrida cv ʽMitchell Diploid.’ Phytochemistry. 2014;103:22–31. 10.1016/j.phytochem.2014.04.004 [DOI] [PubMed] [Google Scholar]
- 55.Hamberger B, Ellis M, Friedmann M, de Azevedo Sousa C, Barbazuk B, Douglas C.Genome-wide analyses of phenylpropanoid-related genes in Populus trichocarpa, Arabidopsis thaliana, and Oryza sativa: the Populus lignin toolbox and conservation and diversification of angiosperm gene families. Can J. Bot. 2007; 85: 1182–1201. [Google Scholar]
- 56.Kao YY, Harding SA, Tsai CJ. Differential expression of two distinct phenylalanine ammonia-lyase genes in condensed tannin-accumulating and lignifying cells of quaking aspen. Plant Physiol. 2002; 130: 796–807. 10.1104/pp.006262 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Pott MB, Hippauf F, Saschenbrecker S, Chen F, Ross J, Kiefer I, et al. Biochemical and structural characterization of benzenoid carboxyl methyltransferases involved in floral scent production in Stephanotis floribunda and Nicotiana suaveolens. Plant Physiol. 2004;135(4):1946–1955. 10.1104/pp.104.041806 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Estévez JM, Cantero A, Reindl A, Reichler S,León P. 1-Deoxy-d-xylulose-5-phosphate synthase, a limiting enzyme for plastidicisoprenoid biosynthesis in plants. J. Biol. Chem. 2001; 276, 22901–22909. 10.1074/jbc.M100854200 [DOI] [PubMed] [Google Scholar]
- 59.Chen F, Tholl D, Bohlmann J, Pichersky E. The family of terpene synthases in plants: a mid-size family of genes for specialized metabolism that is highly diversified throughout the kingdom. Plant J. 2011;66(1):212–229. 10.1111/j.1365-313X.2011.04520.x [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
(DOCX)
(XLS)
(XLS)
(DOCX)
(DOCX)
(DOCX)
(DOCX)
(DOCX)
(DOCX)
(DOCX)
(DOCX)
(DOCX)
Data Availability Statement
The raw sequences were found in NCBI SRA (accession SRP126470); the assembled sequences were found in NCBI TSA (accession GGEA00000000).