Significance
Membrane receptors sense and translate extracellular signals into specific intracellular functions and represent powerful molecules for biomedicine and synthetic biology. However, the computational design of receptors with novel signaling functions remains challenging because receptors often couple to multiple intracellular proteins with poorly characterized interactions. We developed a computational approach to model and rationally engineer orthogonal receptor-intracellular protein pairs that bind and signal with high selectivity. The orthogonal receptor-effector systems coupled with high efficiency and triggered the intended cellular functions without interfering with natural systems. The designed proteins displayed key, distinct sequence motifs when compared with native proteins, which expanded the alphabet of receptor-effector recognition. This design approach can be used to reprogram cellular functions in cell-engineering applications.
Keywords: protein design, cell signaling, G-protein–coupled receptor, protein binding, membrane protein
Abstract
Membrane receptors regulate numerous intracellular functions. However, the molecular underpinnings remain poorly understood because most receptors initiate multiple signaling pathways through distinct interaction interfaces that are structurally uncharacterized. We present an integrated computational and experimental approach to model and rationally engineer membrane receptor-intracellular protein systems signaling with novel pathway selectivity. We targeted the dopamine D2 receptor (D2), a G-protein–coupled receptor (GPCR), which primarily signals through Gi, but triggers also the Gq and beta-arrestin pathways. Using this approach, we designed orthogonal D2–Gi complexes, which coupled with high specificity and triggered exclusively the Gi-dependent signaling pathway. We also engineered an orthogonal chimeric D2–Gs/i complex that rewired D2 signaling from a Gi-mediated inhibitory into a Gs-dependent activating pathway. Reinterpreting the evolutionary history of GPCRs in light of the designed proteins, we uncovered an unforeseen hierarchical code of GPCR–G-protein coupling selectivity determinants. The results demonstrate that membrane receptor–cytosolic protein systems can be rationally engineered to regulate mammalian cellular functions. The method should prove useful for creating orthogonal molecular switches that redirect signals at the cell surface for cell-engineering applications.
Cells respond to their environment through networks of signaling molecules that act as sensors, transmitters, and activators (1). Upon receiving extracellular stimuli, membrane receptor sensors allosterically transmit signals across biological membranes by activating cytosolic transmitter proteins that trigger specific intracellular signaling cascades. The selective coupling of sensor to transmitter proteins is essential to guarantee appropriate physiological responses, but the molecular underpinnings are complex and remain poorly understood. For example, G-protein–coupled receptors (GPCRs), which constitute the largest family of signaling receptors (encoded by close to 800 genes in humans), can bind and activate different subsets of transmitter proteins (i.e., G proteins, beta-arrestins) upon sensing distinct extracellular stimuli. Severe drug side effects often result from the undesired activation of intracellular signaling pathways by drug-stimulated GPCRs (2). Due to the small number of G-protein classes and high evolutionary constraints, specific combinations of residues (i.e., selectivity barcode) on each G-protein class that govern their selective recognition by GPCRs have been readily identified (3). However, distinct subfamilies of GPCRs bind to identical G proteins using different sets of interacting hot-spot residues (3). Consequently, how GPCRs couple to selective G proteins remains very challenging to predict from sequence and necessitates 3D structural information. However, due to the inherent flexibility of GPCRs and G proteins and the intrinsic instability of the receptor active conformations, the agonist ligand–GPCR–G-protein active state complexes have proven very difficult to characterize. Their structure has been determined for only three receptors (B2AR, A2AR, and opsin) bound to two classes of G proteins [Gs and Gt (4–6)]. The lack of structural information hinders traditional structure–function relationships and protein-engineering studies targeting membrane receptor–cytosolic transmitter protein association and subsequent signaling pathways activation.
Engineered cells with novel properties show increasing therapeutic and biotechnological potential (7). Consequently, reprogramming of cellular functions is a major focus in synthetic biology and can be achieved through the design of novel signaling membrane receptors (8). Cytosolic transmitter proteins and a few families of membrane receptor sensors are built on a modular architecture consisting of multiple functionally distinct domains. Empiric combinations of domains from modular proteins enabled the engineering of protein variants with novel functions and spatiotemporal regulation (9–12). However, several important classes of membrane receptor sensors (e.g., GPCRs) are built on nonmodular architectures integrating multiple functionalities within the same protein domain (13, 14). Engineering receptors from these scaffolds require the modification of intrinsic protein properties, e.g., by mutagenesis (15) or protein loop insertion (16). Such approaches remain very challenging and experimentally time-consuming in the absence of a mechanistic understanding of signaling function and 3D structures for the large majority of receptors (17).
An optimal level of regulation and control of cellular functions requires orthogonal signaling components that do not interfere with the protein interaction networks of the host cell (18). While orthogonal extracellular ligand-receptor “input” interactions have been engineered using in vitro evolution techniques (15), orthogonal “output” signals remain far more challenging to design owing to the large number of receptor–cytosolic transmitter protein interactions, closely related transmitter proteins, and poorly characterized binding interfaces (19).
To address these limitations, we developed and applied a general computational experimental approach for rationally predicting and engineering membrane receptor–cytosolic transmitter protein pairs with novel binding specificities, which signal with fine-tuned pathway selectivity. The method integrates membrane protein homology modeling, ligand and protein docking, and design techniques to accurately model and design membrane receptor–transmitter protein coupling even in the absence of structural information on the components and their interactions. We demonstrate the power of the approach by engineering a highly orthogonal human dopamine D2 receptor (D2)–Gi protein pair signaling with higher pathway selectivity than the native system. We also engineered a D2-chimeric Gs–i pair redirecting the dopamine signal into the Gs-mediated pathway. In light of our designs, we have reinterpreted the sequence-structure relationships at GPCR–G-protein-binding interfaces and uncovered a hierarchical code of coupling selectivity determinants. The method should prove useful for creating orthogonal molecular switches redirecting signals at the cell surface for cell-engineering applications.
Results
Since native GPCRs often display substantial promiscuity and bind several transmitter proteins leading to complex intracellular signaling output (2, 20), we aimed, as a proof of concept, at modeling and designing orthogonal GPCR–G-protein pairs that exclusively signal through a selective pathway without interfering with native receptor and cytosolic transmitter protein components (Fig. 1A). Orthogonal signaling systems can be very useful for cell-engineering applications requiring a higher selectivity and level of control than native systems (18). However, computational design approaches require high-resolution structural information, and a large majority of GPCRs and interactions with cytosolic transmitter proteins remain structurally uncharacterized. To address this challenge, we developed a general integrated structure prediction, docking and design approach to accurately model and engineer receptor–transmitter protein structures and interactions (Fig. 1B and SI Appendix, Supplementary Methods).
Fig. 1.
Computational design of orthogonal GPCR–G-protein signaling pairs. (A) GPCR-mediated signaling rewiring through engineered orthogonal GPCR–G-protein pairs. A native promiscuous GPCR interacts and activates several G proteins (G1, G2), triggering distinct signaling pathways. The binding interface between the GPCR and G2 is designed to engineer an orthogonal highly selective protein pair that exclusively signals through the designed G2. (B) Integrated homology modeling, docking, and design for engineering orthogonal binding in the absence of experimental protein structures. The active state D2 structure bound to dopamine agonist ligand and the G-protein Gi is modeled by homology to the distant homologs beta2 adrenergic receptor (β2AR) and opsin, using an integrated homology modeling-docking technique (SI Appendix, Supplementary Methods and Fig. S1). The inactive-state D2 structure bound to the spiperone inverse agonist ligand is modeled starting from the close homolog dopamine D3 receptor (D3DR). Multistate design is applied to the active- and inactive-state D2 structures for engineering an orthogonal binding D2ortho–Giortho pair while ensuring that no selected mutations lead to an inactivated or constitutively active receptor (i.e., signaling without agonist stimulus).
Approach.
The main goal of our approach is to engineer GPCR–G-protein contacts so that the designed proteins couple only with each other without interfering with their native wild-type (WT) counterpart proteins. A secondary goal is to also achieve high selectivity against other endogenously expressed GPCR–G-protein systems. Additionally, to be useful in cell-engineering applications, the designed proteins should retain the properties of signaling switches and signal only upon receiving extracellular ligand agonist stimuli (21, 22). To achieve these goals, we further developed our homology modeling (23, 24) and design (25, 26) techniques (SI Appendix, Supplementary Methods, Supplementary Discussion, and Fig. S1).
Modeling and Design of Orthogonal D2–Gi Complexes.
We targeted D2, which primarily activates the G-protein Gi signaling pathway (27) (Fig. 1B). No structural information is available on D2, Gi, or the D2–Gi bound complex. The native D2 is promiscuous and, upon sensing dopamine, can also activate the Gq and beta-arrestin signaling pathways (28). D2 is central to several neuronal activities involved in sensing, behavioral, and motor functions, as well as severe diseases such as Parkinson’s and Alzheimer’s (29). Therefore, designed dopamine receptors with fine-tuned signaling properties would provide powerful tools for better studying signaling pathways and for future gene therapies against neurodegenerative diseases.
We modeled the ternary dopamine ligand–D2–Gi active state, starting from the B2AR-Gs and opsin-Gt distant homolog structures. The D2 inactive state was modeled from the close homolog dopamine D3 receptor structure. We then used the models as templates in design calculations that rapidly scan the binding interface [>3 × 1019 total possible amino acid combinations (SI Appendix, Supplementary Methods and Fig. S2)] for sets of mutations creating highly specific D2–Gi orthogonal interactions. Specifically, we selected mutations increasing the binding between the two designed proteins while simultaneously decreasing the interactions between the two pairs of WT and designed D2 and Gi proteins (i.e., D2WT–Gidesign and D2design–GiWT). Also, mutations that stabilize or destabilize the inactive resting state of the receptor, potentially leading to loss of activity or constitutive activity (i.e., lack of ligand-induced signaling control) were automatically discarded.
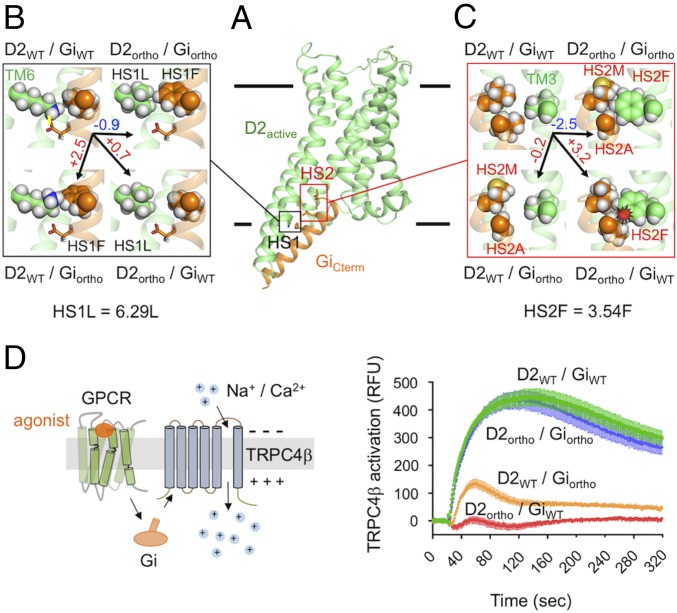
From more than 3 × 1011 combinations of mutations designed at the predicted D2–Gi binding interface, we identified two hot spot motifs (HS1, HS2) predicted to rewire the interactions between D2 and Gi into a highly selective D2ortho and Giortho pair (SI Appendix, Supplementary Methods and Table S1). The hot spots HS1 and HS2 are deeply buried at the binding interface, involving, respectively, two and three complementary mutations on D2 and Gi (Fig. 2 A–C). In our predictions for HS1 (Fig. 2B), substitutions of Lys204 to Leu on transmembrane helix 6 (TM6) of D2 [HS1L; position 6.29 in Ballesteros-Weinstein (BW) notation] and of Leu293 to Phe on Gi (HS1F) provided stronger binding [−0.9 Rosetta Energy Unit (REU)] through better shape complementarity and removal of a partially buried charged residue. The mutations promoted selectivity by decreasing the interactions between D2WT and Giortho through Phe293 (HS1F) (+2.5 REU), predicted to prevent Lys204 from interacting with the stabilizing Asp290 on Gi. In our predictions for HS2 (Fig. 2C), substitutions of Val100 to Phe on TM3 of D2 (HS2F; position 3.54 in BW notation), of Leu297 to Ala (HS2A), and of Leu302 to Met (HS2M) on Gi provided stronger binding (−2.5 REU) through better shape complementarity. The mutations promoted selectivity by decreasing the interactions between D2ortho and GiWT (+3.2 REU) through steric clashes between Phe100 (HS2F) and Leu297.
Fig. 2.
Designed D2–Gi orthogonal pair achieves high activity and coupling selectivity. (A) Representative active-state model of D2 (D2active) bound to the C-terminal helix of Gi (GiCterm). The designed binding interface includes two hot-spot motifs (HS1 and HS2). (B) Zoomed view of the D2–Gi interface at HS1. Predicted energetic contribution of designed residues at HS1 to the binding affinity of the designed D2ortho–Giortho and the noncognate ortho/WT pairs. D2 and Gi residues are colored in green and gold, respectively. The highlighted designed D2 residue HS1L (mutation K204L) is located on TM6 at the Ballesteros Weinstein (BW) position 6.29. Stabilization and destabilization effects correspond to negative and positive energy differences from WT, respectively. Energy differences are in Rosetta Energy Units and shown on top of the black arrows. (C) Similar to B but for HS2. The highlighted designed D2 residue HS2F (mutation V100F) is located on TM3 at the BW position 3.54. (D) TrpC4β channel activation upon D2 activation by dopamine through the Gi signaling pathway. Fluorescence changes are measured for cells stably expressing TrpC4β channels cotransfected with plasmids coding the D2 and Gi variants indicated on each curve (SI Appendix, Supplementary Methods). Under the experimental conditions, the D2 variants were expressed at the same level on the cell surface as measured by ELISA (SI Appendix, Fig. S3A).
Orthogonal D2 Activates Orthogonal Gi with High Efficiency.
We validated the designed signaling switch using a cell-based assay reporting Gi-dependent activation of the TrpC4β channel (30) (Fig. 2D and SI Appendix, Supplementary Methods). All assays were performed under conditions where D2 variants were expressed at the same level on the cell surface (SI Appendix, Fig. S3A). Corroborating our predictions, the designed D2ortho and Giortho pair responded strongly to dopamine with similar amplitude to the native D2WT–GiWT pair, while the competitor D2ortho–GiWT and D2WT–Giortho pairs displayed minimal responses to dopamine. No signal was detected for the D2ortho–GiWT pair, while a small and transient response was observed for the D2WT–Giortho pair (<20% of that measured for the cognate D2WT–GiWT and D2ortho–Giortho pairs). Overall, these experimental measurements strongly validate our predictions and suggest that the designed proteins function as intended.
The Designed Residue Hot Spots Promote Efficient Coupling of the Orthogonal Proteins.
To stringently assess the accuracy of the D2ortho–Giortho-binding predictions and the specific role of each designed motif, we deconstructed the designed interface by backmutating the engineered hot-spot residues predicted to confer high affinity between D2ortho and Giortho to the corresponding native residues. This led us to model and validate four D2–Gi pairs incorporating distinct combinations of WT and designed residues (SI Appendix, Supplementary Discussion and Fig. S4). The measured activations of designed Gi by designed D2 were in good agreement with the calculated effects of individual designed motifs in all four D2–Gi pairs, supporting the accuracy of the modeled D2ortho–Giortho-binding interface.
The Designed Residue Hot Spots Confer High Coupling Specificity.
We next validated the accuracy of the designed interactions conferring binding selectivity. We backmutated the engineered hot-spot residues predicted to destabilize the binding interface between the WT (D2WT and GiWT) and designed proteins (D2ortho and Giortho) to the corresponding native residues. To that end, we modeled and characterized three pairs of designed and WT protein pairs. Except for the D2WT–GiHSF1- pair, the experimentally measured activations were in good agreement with the calculated effects of the designed residues on the coupling selectivity (SI Appendix, Supplementary Discussion and Figs. S5 and S6).
Overall, the strong correlation between predicted and measured responses for 9 of the 10 combinations of designed variants support the high coupling efficiency and specificity provided by the designed hot-spot residues and the accuracy of the rational structure-based design approach.
Orthogonal D2 Signals Exclusively Through the Gi Signaling Pathway.
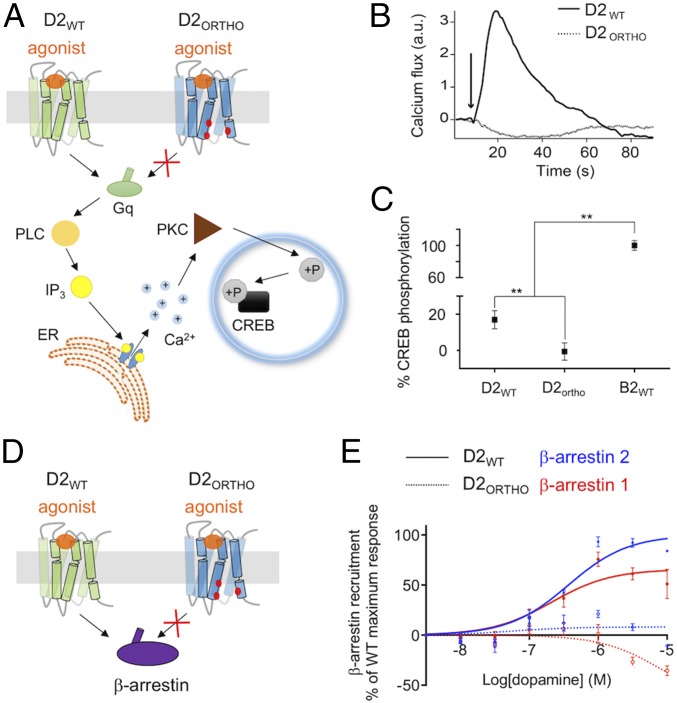
Upon dopamine sensing, the WT D2 receptor is known to signal through both Gi- and Gq-dependent pathways and to couple to beta-arrestins. One major goal in synthetic biology is to engineer molecular systems with a high level of functional precision and selectivity. Therefore, in addition to being orthogonal, we designed the D2 receptor to exclusively trigger the Gi-dependent signaling pathway. To validate the selectivity against alternative G-protein signaling pathways, we measured the Gq-pathway–mediated increase in intracellular calcium flux as well as the Gq- and Gs-pathway–mediated phosphorylation of the transcription factor cAMP response element binding protein (CREB) (Fig. 3 and SI Appendix, Supplementary Methods and Fig. S7). D2 signaling through Gi was prevented using pertussis toxin treatment to selectively measure signaling through the Gq pathway. Consistent with previous findings, we observed that D2WT promotes small but significant calcium release and phosphorylation of CREB (28). By contrast, D2ortho did not produce any detectable amount of released calcium or phosphorylated CREB (pCREB) upon stimulation by dopamine. Interestingly, none of the backmutated D2 variants retaining only one of the two designed hot-spot residues (i.e., HS1L-, HS2F-) promoted any measurable pCREB production upon stimulation by dopamine (SI Appendix, Fig. S7). These results indicate that each designed hot spot is sufficient to disrupt the weak coupling of D2 to Gq.
To evaluate the selectivity of the designed D2 on beta-arrestin recruitment, we used enhanced bystander bioluminescence resonance energy transfer (ebBRET) (31). Upon stimulation of the WT receptor with dopamine, beta-arrestin–RlucII is recruited to the plasma membrane labeled with the plasma membrane marker rGFP-CAAX, leading to a time- and concentration-dependent increase of the BRET signal (Fig. 3C and SI Appendix, Fig. S8). Consistent with the predictions, no significant recruitment by D2ortho was observed for beta-arrestin 1 or 2. Coexpression of GRK2 led to phosphorylation of the receptor C-terminal tail and enhanced beta-arrestin recruitment for both D2WT and D2ortho. However, while detectable, the dopamine-mediated beta-arrestin recruitment measured for D2ortho was only around 25% of that of D2WT.
Fig. 3.
Designed D2–Gi orthogonal pair achieves high signaling pathway selectivity. (A) Calcium release and CREB phosphorylation upon D2 activation by dopamine through the Gq signaling pathway. (B) After addition of 20 μM of dopamine (arrow), change in intracellular concentration of calcium ion was measured using a Ca2+-sensitive fluorescent probe on cells transiently transfected with either pcDNA3.1-D2WT (solid line) or pcDNA3.1-D2ORTHO (dashed line). (C) pCREB levels are measured by ELISA on cells transiently transfected with the above-mentioned constructs (SI Appendix, Supplementary Methods). pCREB levels are normalized to those obtained upon β2AR activation by isoproterenol through the Gs signaling pathway. Signaling through GiWT is abrogated with pertussis toxin (SI Appendix, Supplementary Methods). **P < 0.05. (D and E) β-Arrestin recruitment by D2 receptors upon stimulation by dopamine using ebBRET on cells transiently transfected with the above-mentioned constructs. Signals are normalized to that obtained with D2WT and β-arrestin 2. Under the experimental conditions, the D2 variants were expressed at the same level on the cell surface as measured by ELISA (SI Appendix, Figs. S3B, S7B, and S8D).
Altogether, these results demonstrate the high signaling pathway selectivity of the orthogonal signaling D2–Gi switch and suggest that the designed mutations prevent the coupling of D2 to alternative Gq-, Gs-, and beta-arrestin–mediated signaling pathways.
Engineering a D2–Gs-Signaling Switch.
We next attempted to design a GPCR–G-protein complex that rewires the receptor signaling into an alternative nonnative pathway. Since D2 primarily signals through Gi, which inhibits the production of cAMP, we decided to engineer a D2–Gs complex that would revert D2 signaling into increasing cAMP levels instead. Because D2 does not display any measurable coupling to Gs, we decided to harness the binding potency and selectivity of Giortho onto the signaling function of Gs by creating a chimeric Gs-iortho protein. Using the structural models of D2–Gi- and B2–Gs-binding interfaces, we engineered key D2-binding hot-spot residues from the Giortho C-terminal domain onto that of Gs. Consistent with our predictions, the chimeric Gs-iortho protein redirected D2ortho signaling into substantial Gs-dependent Epac activation and phosphorylation of CREB (SI Appendix, Fig. S9). Additionally, Gs-iortho was very selective for D2ortho and displayed minimal coupling to D2WT. Chimeric WT G proteins had been previously engineered by swapping a few residues from the C-terminal helix of distinct G-protein classes (32). While functional, these chimeric proteins lacked the binding selectivity provided by our designed orthogonal interface. Our results suggest that orthogonal pairs of GPCRs and chimeric G proteins may prove useful for rewiring signaling pathways in cell-engineering applications.
Altogether, these results indicate that the structure-based modeling and engineering approach provides a general versatile platform for reprogramming the intracellular signaling selectivity of GPCRs, even those without solved 3D structures.
Uncovering Two Classes of GPCR–G-Protein Recognition Determinants.
The high functional precision of the novel engineered orthogonal protein complexes prompted us to analyze the designed binding selectivity determinants within the context of the evolutionary history of naturally evolved GPCR–G-protein interfaces (Fig. 4A).
Fig. 4.
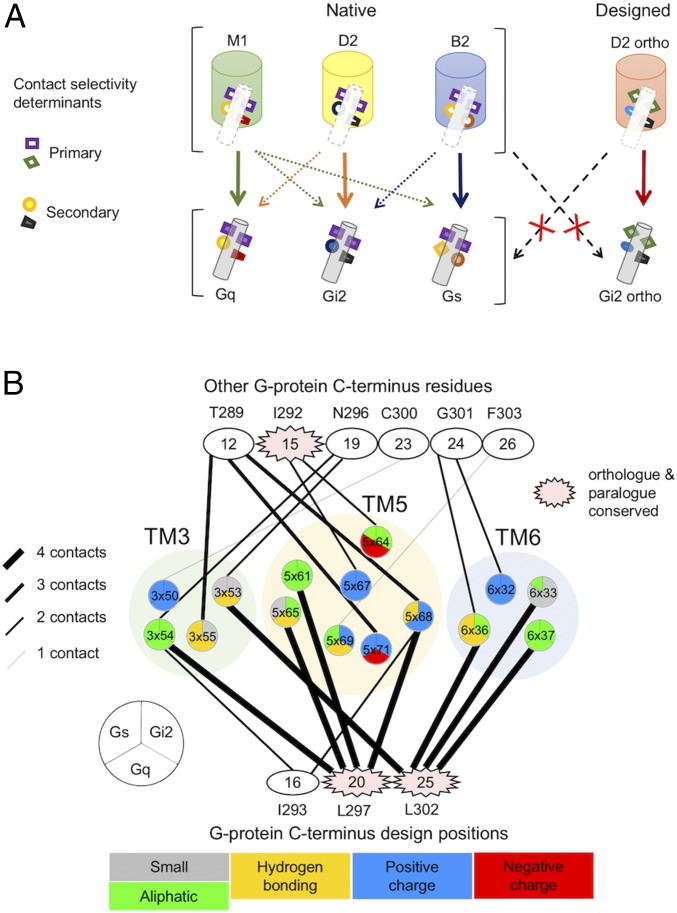
Two classes of contact selectivity determinants of GPCR–G-protein coupling. (A) Schematic representation of the contact selectivity determinants at the GPCR–G-protein–binding interface. Primary determinants are located at highly conserved positions on naturally evolved G proteins. Modifying the primary contacts results in major selectivity switches. Native GPCR–G-protein systems differ in the secondary contact determinants that fine-tune their binding specificity profile, enabling GPCRs to recognize multiple G proteins with distinct affinities. Muscarinic M1, dopamine D2, and beta2 adrenergic receptors are provided as examples with their corresponding G-protein–coupling profiles. (B) Sequence-structure relationships at the binding interface of three major classes of GPCR–G-protein systems. C-terminal residues of the G protein are separated in two sets of positions: positions selected for design based on D2–Gi models (Bottom) and nondesigned positions (Top). G-protein position numbering is given based on the consensus G-protein aligned sequences (within ovals) and based on the exact residue type and number in Gi2 (above or below ovals). Each G-protein position is connected to a GPCR position based on the number of interaction contacts observed at the D2–Gi interface. Each position on the GPCR is represented as a pie divided into three regions that describe the chemical property of the consensus amino acid sequence coupling to each family of G proteins (i.e., Gs, Gi, Gq) (SI Appendix, Supplementary Methods). Sixty percent of the GPCR residues contacting G proteins do not share the same chemical properties in different GPCR–G-protein classes. Compared with previous studies (3), the amino acid type of residues contacting G proteins differs even between GPCR subfamilies coupling to the same G-protein class.
Interestingly, while the only starting information of our approach was the predicted structure of the D2–Gi complex, the hot-spot designed motifs included positions on Gi that are universally conserved among naturally evolved G proteins (positions H5.20 and H5.25, Fig. 4B and SI Appendix, Fig. S10). As such, these positions are currently thought to be essential for proper GPCR–G-protein coupling and G-protein activation (3). Within this model, any substitutions at these binding sites is predicted to disrupt signaling function. By contrast, our findings indicate that alternative binding motifs that perfectly maintain optimal G-protein activation function can be engineered at these sites. The results imply that, instead of the residue types on G proteins (3), the chemical contacts between GPCRs and G proteins at the conserved positions are critical for both function and selectivity. Since optimal contacts can be created from multiple combinations of residue types with high chemical and structural complementarity, this expands the range of binding and signaling selectivity that GPCR–G-protein complexes can achieve beyond that explored by nature. To achieve high specificity while maintaining high affinity, the computational design approach introduced “large key–small lock” residue motifs at hot spots HS1 and HS2. A large phenylalanine at each hot spot motif provided the “key” while small hydrophobic “lock” residues maintained optimal surface complementarity and interactions on the other side of the binding interface. The asymmetric design of the engineered hot-spot motifs contrasted with that of native GPCR–G-proteins composed of residues of similar sizes (Lys, Leu at HS1 and Val, Leu, Leu at HS2, Fig. 2 B and C) and provided the desired specificity.
In light of the designs, we propose a reinterpretation of recent bioinformatics findings (3), leading to a paradigm shift based on the existence of two classes of contact selectivity determinants at GPCR–G-protein interfaces (Fig. 4B). The primary class is the major binding affinity and specificity determinant enabling optimal docking of G proteins in the TM6–TM7-binding groove of GPCRs. Chemical complementarity is key at the primary contact sites, and small structural changes on one side of the interface can disrupt binding, enabling the design of complete selectivity switches. The primary class of contacts is under strong evolutionary constraints and characterized by a high level of sequence conservation in naturally evolved G proteins. Optimal contact with the primary sites on G proteins is maintained through diverse combinations of residue types in GPCRs (Fig. 4B and ref. 3), highlighting the multiple possible solutions for satisfying chemical and structural complementarity.
The secondary class fine-tunes the binding selectivity achieved by the primary sites and has been used by nature to evolve GPCR–G-protein systems with a wide range of binding specificity and signaling profiles. Since most naturally evolved G proteins share the same primary sites recognized by all GPCRs, modulation of the secondary sites often results in GPCRs coupling to several G proteins and displaying a significant level of signaling promiscuity. The secondary contacts are highly specific to each G-protein class and GPCR subfamily and encode complex networks of interactions (Fig. 4B). This level of binding selectivity fine-tuning remains very challenging to predict from sequence alone and requires high-resolution structural information.
Discussion
Here, we describe a general approach to accurately model and engineer membrane receptor–cytosolic transmitter protein systems with diverse binding and signaling pathway selectivity even without protein structure information. We successfully engineered orthogonal dopamine D2 receptor–Gi protein pairs that couple with high selectivity and, unlike the native D2 receptor, exclusively trigger the Gi-dependent signaling pathway. We also engineered an orthogonal D2–Gs–i chimeric protein pair that redirected D2 signaling from the inhibitory Gi- to the activating Gs-dependent signaling pathway. Remarkably, 10 among 11 engineered receptor–transmitter protein pairs displayed the relative levels of molecular recognition and function predicted by the calculations. The method is very efficient and requires only distant homolog structural information without restriction on the protein topology, size, and nature of protein-binding interfaces. Therefore, it can be applied to a wide diversity of membrane receptor–transmitter protein-signaling classes beyond the family of GPCR–G-protein systems.
The accurate structural models of the uncharacterized GPCR–Gi family substantially expanded our knowledge of the GPCR–G-protein recognition molecular code. Previous sequence-based bioinformatics studies identified mainly class-specific binding determinants on the G proteins (3). The lack of structural information on GPCR–G-protein systems and their complex evolutionary history made similar findings very difficult on GPCRs. Using the structural modeling and design framework, we evolved and identified key 3D contacts between GPCR and G proteins, which inherently define protein–protein recognition. Through engineered functional receptor–transmitter protein pairs not yet identified in nature, we discovered a hierarchical code of contacts determining GPCR–G-protein functional coupling. Interestingly, the identification of tight binding but nonsignaling GPCR–G-protein pairs that correlate with nonnative binding modes suggests that conformational constraints are important determinants of G-protein activation in addition to protein contacts defining binding affinity and specificity. Adding a conformational dimension to the GPCR–G-protein–coupling alphabet will be important to establish a unified code integrating recognition and signaling for predicting function. Overall, our findings and approach provide an integrated sequence–structure framework for predicting how distinct GPCR families manage to recognize a small number of G proteins with diverse and precise selectivity.
A hierarchy of contacts where a minority of hot-spot motifs guarantees primary recognition while the others encode second-order fine-tuning of binding selectivity has been widely used by nature for evolving cognate proteins with a large spectrum of binding affinities and specificities. A hierarchy of recognition determinants also provides an effective framework for creating orthogonal functionalities through minimal perturbations of the native proteins. Our findings suggest that a large diversity of orthogonal and functional receptor–transmitter protein complexes that do not couple with native systems may be designed. As such, they may be used as plug-in molecular devices for precise manipulation of cellular functions with minimal interference. Since many synthetic biology approaches involve the engineering of molecular systems with a high level of functional precision and selectivity, the method paves the road for creating cellular signaling pathways.
Rationally engineering orthogonal protein systems with fined-tuned signaling properties at the cell surface will greatly aid in the design of custom signaling networks for better studying of complex disease mechanisms and for advancing cell and gene therapeutics.
Methods
The cDNA clones for the human dopamine D2DR (long isoform) receptor and Gi protein in pcDNA3.1(+) vectors were a generous gift from T. Wensel (Baylor College of Medicine, Houston). All functional assays were performed as described in SI Appendix, Supplementary Methods. RosettaMembrane’s homology modeling protocol was used to generate models of D2DR WT providing structural templates for the design calculations (23). (SI Appendix, Supplementary Methods).
Supplementary Material
Acknowledgments
This work was supported by National Institutes of Health (NIH) Grant 1R01GM097207; by a supercomputer allocation from the Extreme Science and Engineering Discovery Environment (XSEDE) (MCB120101) (to P.B.); and by NIH National Institute of General Medical Sciences Training Fellowship T32GM008280 (to M.Y. and K.M.C.). C.A. is supported by a Leukemia and Lymphoma Society Translational Research Program grant and a Cancer Prevention and Research Institute of Texas Individual Investigator Research Award (RP160345).
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1718489115/-/DCSupplemental.
References
- 1.Gordley RM, Bugaj LJ, Lim WA. Modular engineering of cellular signaling proteins and networks. Curr Opin Struct Biol. 2016;39:106–114. doi: 10.1016/j.sbi.2016.06.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Manglik A, et al. Structure-based discovery of opioid analgesics with reduced side effects. Nature. 2016;537:185–190. doi: 10.1038/nature19112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Flock T, et al. Selectivity determinants of GPCR-G-protein binding. Nature. 2017;545:317–322. doi: 10.1038/nature22070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Rasmussen SG, et al. Crystal structure of the β2 adrenergic receptor-Gs protein complex. Nature. 2011;477:549–555. doi: 10.1038/nature10361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Carpenter B, Nehmé R, Warne T, Leslie AG, Tate CG. Structure of the adenosine A(2A) receptor bound to an engineered G protein. Nature. 2016;536:104–107. doi: 10.1038/nature18966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Scheerer P, et al. Crystal structure of opsin in its G-protein-interacting conformation. Nature. 2008;455:497–502. doi: 10.1038/nature07330. [DOI] [PubMed] [Google Scholar]
- 7.Lim WA, June CH. The principles of engineering immune cells to treat cancer. Cell. 2017;168:724–740. doi: 10.1016/j.cell.2017.01.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Arber C, Young M, Barth P. Reprogramming cellular functions with engineered membrane proteins. Curr Opin Biotechnol. 2017;47:92–101. doi: 10.1016/j.copbio.2017.06.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Roybal KT, et al. Engineering T cells with customized therapeutic response programs using synthetic notch receptors. Cell. 2016;167:419–432.e16. doi: 10.1016/j.cell.2016.09.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.St-Pierre F, Chavarha M, Lin MZ. Designs and sensing mechanisms of genetically encoded fluorescent voltage indicators. Curr Opin Chem Biol. 2015;27:31–38. doi: 10.1016/j.cbpa.2015.05.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Dotti G, Gottschalk S, Savoldo B, Brenner MK. Design and development of therapies using chimeric antigen receptor-expressing T cells. Immunol Rev. 2014;257:107–126. doi: 10.1111/imr.12131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Schwarz KA, Daringer NM, Dolberg TB, Leonard JN. Rewiring human cellular input-output using modular extracellular sensors. Nat Chem Biol. 2017;13:202–209. doi: 10.1038/nchembio.2253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Katritch V, Cherezov V, Stevens RC. Structure-function of the G protein-coupled receptor superfamily. Annu Rev Pharmacol Toxicol. 2013;53:531–556. doi: 10.1146/annurev-pharmtox-032112-135923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.von Heijne G. Membrane-protein topology. Nat Rev Mol Cell Biol. 2006;7:909–918. doi: 10.1038/nrm2063. [DOI] [PubMed] [Google Scholar]
- 15.Dong S, Rogan SC, Roth BL. Directed molecular evolution of DREADDs: A generic approach to creating next-generation RASSLs. Nat Protoc. 2010;5:561–573. doi: 10.1038/nprot.2009.239. [DOI] [PubMed] [Google Scholar]
- 16.Siuda ER, et al. Spatiotemporal control of opioid signaling and behavior. Neuron. 2015;86:923–935. doi: 10.1016/j.neuron.2015.03.066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Pieper U, et al. Coordinating the impact of structural genomics on the human α-helical transmembrane proteome. Nat Struct Mol Biol. 2013;20:135–138. doi: 10.1038/nsmb.2508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kapp GT, et al. Control of protein signaling using a computationally designed GTPase/GEF orthogonal pair. Proc Natl Acad Sci USA. 2012;109:5277–5282. doi: 10.1073/pnas.1114487109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Moreira IS. Structural features of the G-protein/GPCR interactions. Biochim Biophys Acta. 2014;1840:16–33. doi: 10.1016/j.bbagen.2013.08.027. [DOI] [PubMed] [Google Scholar]
- 20.Wacker D, et al. Structural features for functional selectivity at serotonin receptors. Science. 2013;340:615–619. doi: 10.1126/science.1232808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Deupi X, Kobilka BK. Energy landscapes as a tool to integrate GPCR structure, dynamics, and function. Physiology (Bethesda) 2010;25:293–303. doi: 10.1152/physiol.00002.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Han M, Smith SO, Sakmar TP. Constitutive activation of opsin by mutation of methionine 257 on transmembrane helix 6. Biochemistry. 1998;37:8253–8261. doi: 10.1021/bi980147r. [DOI] [PubMed] [Google Scholar]
- 23.Chen KY, Sun J, Salvo JS, Baker D, Barth P. High-resolution modeling of transmembrane helical protein structures from distant homologues. PLOS Comput Biol. 2014;10:e1003636. doi: 10.1371/journal.pcbi.1003636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Feng X, Ambia J, Chen KM, Young M, Barth P. Computational design of ligand-binding membrane receptors with high selectivity. Nat Chem Biol. 2017;13:715–723. doi: 10.1038/nchembio.2371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Havranek JJ, Harbury PB. Automated design of specificity in molecular recognition. Nat Struct Biol. 2003;10:45–52. doi: 10.1038/nsb877. [DOI] [PubMed] [Google Scholar]
- 26.Humphris EL, Kortemme T. Design of multi-specificity in protein interfaces. PLoS Comput Biol. 2007;3:e164. doi: 10.1371/journal.pcbi.0030164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Sung YM, Wilkins AD, Rodriguez GJ, Wensel TG, Lichtarge O. Intramolecular allosteric communication in dopamine D2 receptor revealed by evolutionary amino acid covariation. Proc Natl Acad Sci USA. 2016;113:3539–3544. doi: 10.1073/pnas.1516579113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Yan Z, Feng J, Fienberg AA, Greengard P. D(2) dopamine receptors induce mitogen-activated protein kinase and cAMP response element-binding protein phosphorylation in neurons. Proc Natl Acad Sci USA. 1999;96:11607–11612. doi: 10.1073/pnas.96.20.11607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Girault JA, Greengard P. The neurobiology of dopamine signaling. Arch Neurol. 2004;61:641–644. doi: 10.1001/archneur.61.5.641. [DOI] [PubMed] [Google Scholar]
- 30.Luo J, Zhu Y, Zhu MX, Hu H. Cell-based calcium assay for medium to high throughput screening of TRP channel functions using FlexStation 3. J Vis Exp. 2011:3149. doi: 10.3791/3149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Namkung Y, et al. Monitoring G protein-coupled receptor and β-arrestin trafficking in live cells using enhanced bystander BRET. Nat Commun. 2016;7:12178. doi: 10.1038/ncomms12178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Conklin BR, et al. Carboxyl-terminal mutations of Gq alpha and Gs alpha that alter the fidelity of receptor activation. Mol Pharmacol. 1996;50:885–890. [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.