Abstract
Background
Aberrant methylation of DNA is a key driver of hepatocellular carcinoma (HCC). In this study, we sought to integrate four cohorts profile datasets to identify such abnormally methylated genes and pathways associated with HCC.
Methods
To this end, we downloaded microarray datasets examining gene expression (GSE84402, GSE46408) and gene methylation (GSE73003, GSE57956) from the GEO database. Abnormally methylated differentially expressed genes (DEGs) were sorted and pathways were analyzed. The String database was then used to perform enrichment and functional analysis of identified pathways and genes. Cytoscape software was used to create a protein–protein interaction network, and MCODE was used for module analysis. Finally, overall survival analysis of hub genes was performed by the OncoLnc online tool.
Results
In total, we identified 19 hypomethylated highly expressed genes and 14 hypermethylated lowly expressed genes at the screening step, and finally found six mostly changed hub genes including MAD2L1, CDC20, CCNB1, CCND1, AR and ESR1. Pathway analysis showed that aberrantly methylated-DEGs mainly associated with the cell cycle process, p53 signaling, and MAPK signaling in HCC. After validation in TCGA database, the methylation and expression status of hub genes was significantly altered and same with our results. Patients with high expression of MAD2L1, CDC20 and CCNB1 and low expression of CCND1, AR, and ESR1 was associated with shorter overall survival.
Conclusions
Taken together, we have identified novel aberrantly methylated genes and pathways linked to HCC, potentially offering novel insights into the molecular mechanisms governing HCC progression and serving as novel biomarkers for precision diagnosis and disease treatment.
Electronic supplementary material
The online version of this article (10.1186/s12935-018-0629-5) contains supplementary material, which is available to authorized users.
Keywords: Hepatocellular carcinoma, Methylation, Hub genes
Background
Hepatocellular carcinoma (HCC), an inflammation-driven disease, is the third deadliest cancer worldwide, and HCC prevalence is predicted to continue to rise in coming years, serving as a major economic burden [1, 2]. Most cases of HCC occur in developing countries, such as China, and the leading cause of HCC is infection with hepatitis B virus (HBV); in contrast, the main cause in developed countries, such as the USA, is infection with hepatitis C virus (HCV) [3, 4]. Other risk factors for developing HCC include exposure to aflatoxin, alcohol intake, smoking, and diabetes [5]. The best curative treatments to date in early stage HCC patients involve surgical resection, tumor ablation, and potentially liver transplantation [6, 7]. However, the prognosis after curative therapy for HCC remains unsatisfactory because of a high postoperative recurrence rate. An improved understanding of the basic biology of HCC is needed to enhance prognostic predictions and to enhance therapeutic efficacy against this deadly disease.
The term epigenetics refers to heritable gene expression alterations no associated with DNA sequence changes [8]. The DNA methylation is closely related to embryonic development [9], regulation of gene expression [10], X-chromosome inactivation [11], genomic imprinting [12], and genomic stability [13]. Altered DNA methylation such as tumor suppressor gene hypermethylation or oncogene hypomethylation is thought to promote tumorigenesis. Genes including P15, P16, Ras association domain family 1 isoform A (RASSF1A), and Retinoblastoma 1 are inactivated in HCC due to promoter hypermethylation of these genes [14–17]. Given that methylation is potentially reversible, detection of such aberrant DNA methylation of tumor suppressors and oncogenes in HCC could be useful as a therapeutic target.
While altered methylation of many genes has been demonstrated to date in the context of HCC, a complete interaction network documenting the relationship between said genes remains to be produced. The comprehensive analysis of multiple datasets offers the power needed to properly identify and assess pertinent pathways and genes mediating the biological processes associated with HCC. To this end, we used datasets from microarrays examining gene expression (GSE84402, GSE46408) and gene methylation (GSE73003, GSE57956) to assess HCC gene and epigenetic signatures, allowing for identification of genes and pathways that were both abnormally methylated and differentially expressed. Using a protein–protein interaction network we were also able to identify key so-called “hub” genes central to these signaling events. Through this analysis, we believe it is possible to identify novel differentially methylated genes associated with HCC, offering key insights into the molecular mechanisms governing HCC development and progression.
Methods
Microarray data
Gene expression profiling datasets GSE84402 and GSE46408, and gene methylation profiling datasets GSE73003 and GSE57956, were downloaded from the gene expression omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo/) [18, 19]. 14 HCC and 14 normal specimens were obtained in GSE84402, while 6 HCC and 6 normal samples were obtained in GSE46408. GSE57956 included a total of 59 primary HCC tumor samples and 59 adjacent normal tissue samples, while GSE73003 included 40 paired normal and HCC samples from 20 patients.
Data acquisition and processing
Raw gene expression profiling datasets GSE84402 and GSE46408 were downloaded from GEO public repositories. Data processing was performed using robust multi-array average (RMA) in GeneSpring GX 11.5 (Agilent Technologies Pty Ltd), including background adjustment, normalization and log transformation of the values. The threshold set for up and down-regulated genes was a | log FC | ≥ 2 and P ≤ 0.05. For gene methylation profiling dataset (GSE73003 and GSE57956), the GEO2R software was used to analyze the raw data and to identify differentially methylated genes (DMGs). GEO2R is an interactive web tool which allows users to compare different groups of samples in a GEO series to screen genes that are differentially expressed in experimental conditions. The adjusted P-values (adj. P) and Benjamini and Hochberg false discovery rate were applied to provide a balance between discovery of statistically significant genes and limitations of false-positives. The DMGs cutoff criteria were P < 0.05 and |t| > 2. Additionally, one-sided tests were used to categorize the upregulated or downregulated DMGs. Finally, a Venn diagram was used to identify hypomethylated highly-expressed genes and hypermethylated lowly-expressed genes.
Functional and pathway enrichment analysis
Gene ontology (GO) analyses and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were conducted on identified genes with altered methylation/expression using the Search Tool for the Retrieval of Interacting Genes (STRING, https://string-db.org/). This allowed for protein–protein interaction (PPI) network generation and functional annotation of genes of interest. P < 0.05 was the threshold for statistical significance.
Generation and analysis of a protein–protein interaction (PPI) network
In order to interpret the drivers of carcinogenesis in a meaningful manner, a functional PPI analysis was necessary. The STRING database allowed us to generate PPI networks for both hypomethylated/and highly-expressed genes as well as for hypermethylated and lowly-expressed genes. A 0.4 interaction score served as the cutoff prior to visualization. Modules were then screened using Molecular Complex Detection (MCODE) in the Cytoscape software package, with MCODE score > 4 and number of nodes > 5. The top 3 Hub genes were selected by CytoHubba app in Cytoscape software. To determine the expression pattern of six hub genes in HCC, we used the datasets in the Oncomine (https://www.oncomine.org) and UALCAN (http://ualcan.path.uab.edu/) database. Oncomine and UALCAN are an online database consisting of previously published and open-access microarray data. The analysis enables multiple comparisons of gene expression between different studies; the significance of the gene expression across the available studies was also taken into account [20, 21]. The results were filtered by selecting hepatocellular carcinoma vs. normal tissue.
Hub gene validation
The Cancer Genome Atlas (TCGA) database has generated comprehensive, multi-dimensional maps of the key genomic changes in various types of cancers. MEXPRESS (http://mexpress.be/) is a data visualization tool designed for the easy visualization of TCGA expression, DNA methylation and clinical data, as well as the relationships between them. To confirm our results, we used the MEXPRESS to validate hypermethylation/low-expression hub genes and hypomethylation/high-expression hub genes in TCGA database. The probability of survival and significance were calculated using the OncoLnc database. OncoLnc (http://www.oncolnc.org/) is an online tool for interactively exploring survival correlations, containing survival data for 8647 patients from 21 cancer studies.
Results
Identification of abnormally methylated and differentially expressed genes in HCC
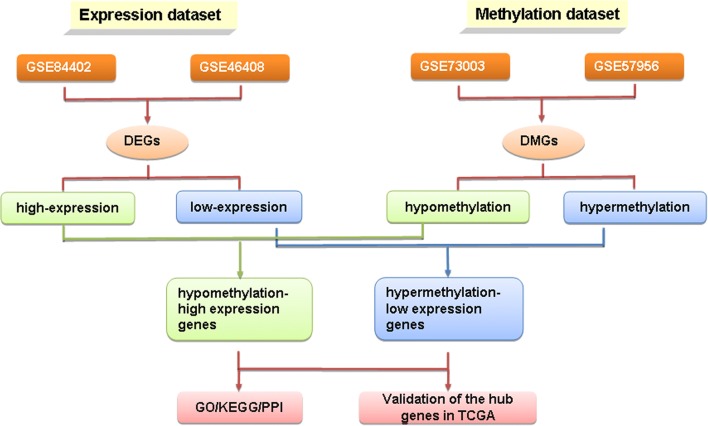
A study design flowchart is shown in Fig. 1. In gene expression microarrays, 127 genes were up-regulated in both datasets (207 in GSE84402, 851 in GSE46408) and 169 were down-regulated in both datasets (307 in GSE84402, 503 in GSE46408). For the gene methylation microarrays, 2139 overlapping hypermethylated genes (2468 in GSE73003, 3157 in GSE57956) and 4091 overlapping hypomethylated genes (4527 in GSE73003, 5334 in GSE57956) were identified. By comparing these two sets of genes, a total of 19 hypomethylated, highly-expressed genes and 14 hypermethylated, low-expression genes were identified (Fig. 2). A heat map of these genes in GSE84402 is shown in Fig. 3.
Fig. 1.
The flowchart of this study
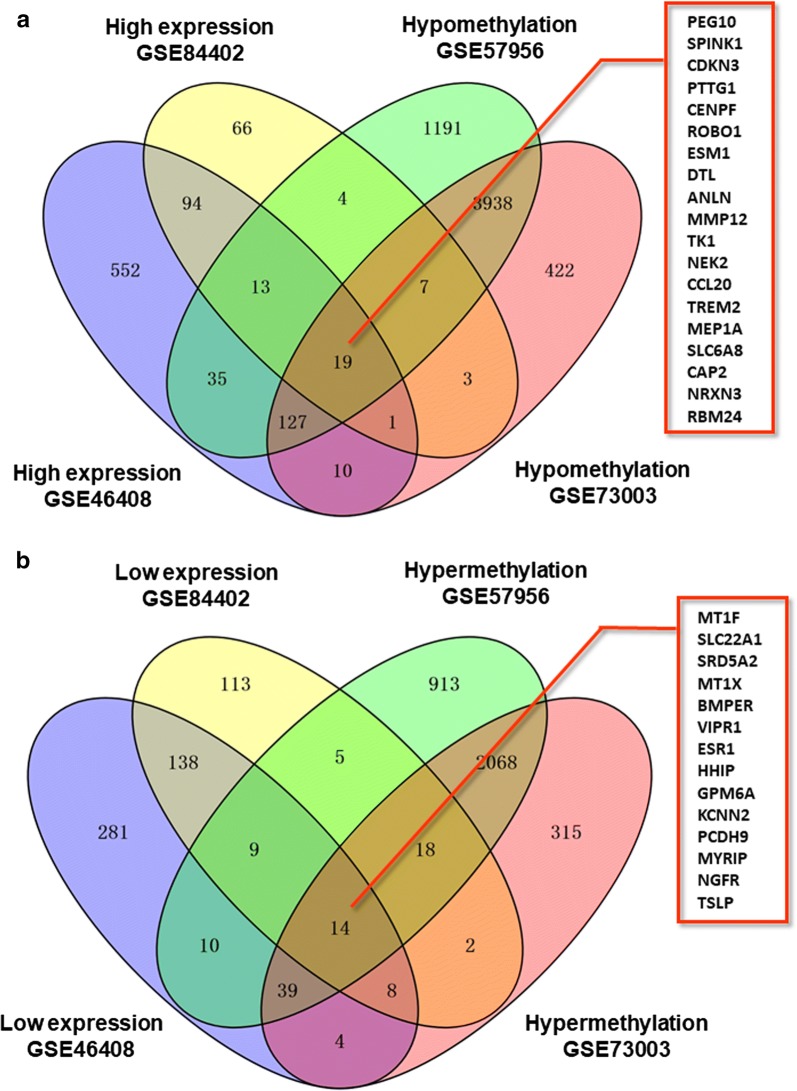
Fig. 2.
Identification of aberrantly methylated and differentially expressed genes was analyzed by Funrich software. Different color areas represented different datasets. a Hypomethylation and high expression genes; b hypermethylation and low expression genes
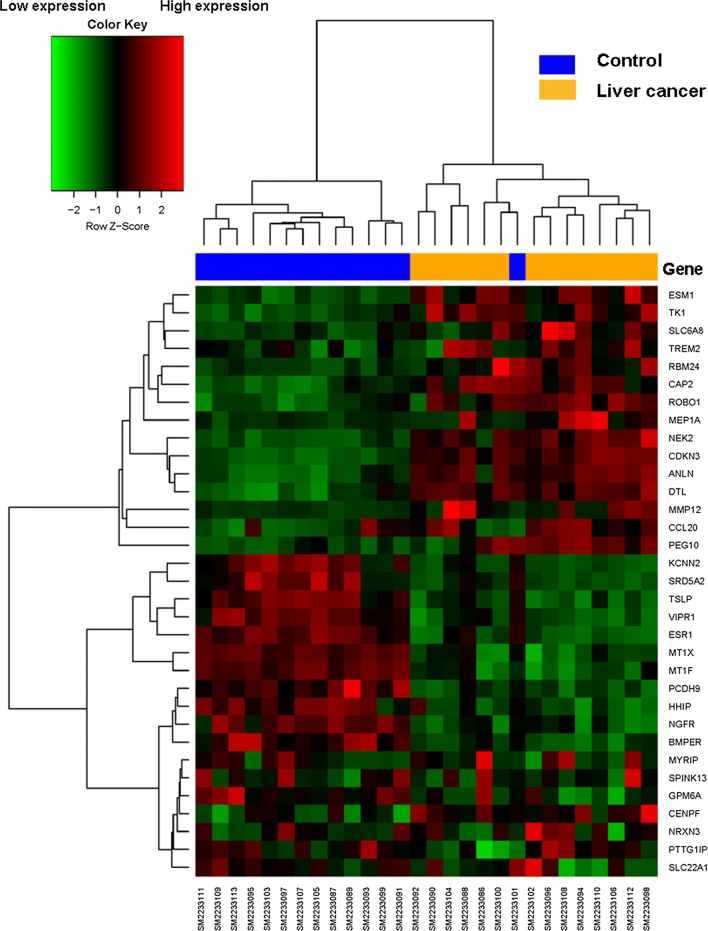
Fig. 3.
The heat map of 19 hypomethylation/high-expression genes and 14 hypermethylation/low-expression genes in GSE84402
Gene ontology and pathway functional enrichment analysis
The GO annotation and pathway enrichment analyses of identified aberrantly methylated and differentially expressed genes were implemented using the online tool STRING. Genes that were hypomethylated and highly expressed were enriched for cell cycle regulation, while the hypermethylated low-expression genes were primarily linked to cell proliferation, gene expression, and signal transduction. Cell component enrichment analysis indicated that hypomethylated highly-expressed genes were correlated with cytoskeletal part, whereas hypermethylation/low-expression genes were predominant at intracellular organelle lumen. As for molecular function, hypomethylation/high-expression genes were enriched mainly in roundabout binding, histone kinase activity, and protein binding, while hypermethylation/low-expression genes were mostly enriched in cyclin binding, receptor binding, and signal transducer activity (Table 1). The pathway analysis revealed that hypomethylated highly-expressed genes were linked to oocyte meiosis, cell cycle, and ubiquitin mediated proteolysis, while hypermethylation/low-expression genes significantly enriched in pathways in cancer, p53 signaling pathway, MAPK signaling pathway, and proteoglycans in cancer (Table 2).
Table 1.
Gene ontology analysis of aberrantly methylated-differentially expressed genes in hepatocellular carcinoma
| Category | ID | Term | Gene count | False discovery rate |
|---|---|---|---|---|
| Hypomethylation and high expression | ||||
| BP | GO.0051983 | Regulation of chromosome segregation | 13 | 3.94E−20 |
| BP | GO.0030071 | Regulation of mitotic metaphase/anaphase transition | 12 | 4.44E−20 |
| BP | GO.0007094 | Mitotic spindle assembly checkpoint | 11 | 1.30E−19 |
| BP | GO.0007088 | Regulation of mitotic nuclear division | 14 | 3.88E−19 |
| BP | GO.0010564 | Regulation of cell cycle process | 17 | 2.77E−15 |
| CC | GO.0000793 | Condensed chromosome | 10 | 3.89E−10 |
| CC | GO.0005819 | Spindle | 11 | 6.30E−10 |
| CC | GO.0031461 | Cullin-RING ubiquitin ligase complex | 7 | 1.34E−07 |
| CC | GO.0005813 | Centrosome | 8 | 0.00011 |
| CC | GO.0044430 | Cytoskeletal part | 12 | 0.000143 |
| MF | GO.0048495 | Roundabout binding | 3 | 0.000105 |
| MF | GO.0035173 | Histone kinase activity | 3 | 0.0059 |
| MF | GO.0005515 | protein binding | 21 | 0.0083 |
| Hypermethylation and low expression | ||||
| BP | GO.0042127 | Regulation of cell proliferation | 15 | 1.97E−06 |
| BP | GO.0010468 | Regulation of gene expression | 19 | 0.000139 |
| BP | GO.0009968 | Negative regulation of signal transduction | 10 | 0.000642 |
| BP | GO.0000082 | G1/S transition of mitotic cell cycle | 5 | 0.00149 |
| BP | GO.0051726 | Regulation of cell cycle | 9 | 0.00149 |
| CC | GO.0097458 | Neuron part | 10 | 0.00143 |
| CC | GO.0045202 | Synapse | 7 | 0.00802 |
| CC | GO.0070013 | Intracellular organelle lumen | 16 | 0.0137 |
| CC | GO.0005694 | Chromosome | 7 | 0.0138 |
| CC | GO.0005654 | Nucleoplasm | 13 | 0.0149 |
| MF | GO.0030332 | Cyclin binding | 4 | 4.14E−05 |
| MF | GO.0005515 | Protein binding | 21 | 0.000265 |
| MF | GO.0004871 | Signal transducer activity | 11 | 0.00462 |
| MF | GO.0005102 | Receptor binding | 10 | 0.00462 |
| MF | GO.0016538 | Cyclin-dependent protein serine/threonine kinase regulator activity | 3 | 0.00462 |
BP biological process, CC cell component, MF molecular function
Table 2.
KEGG pathway analysis of aberrantly methylated-differentially expressed genes in hepatocellular carcinoma
| Pathway ID | Pathway name | Gene count | False discovery rate | Genes |
|---|---|---|---|---|
| Hypomethylation and high expression | ||||
| 4110 | Cell cycle | 12 | 1.03E−15 | ANAPC10, ANAPC4, BUB1, BUB1B, BUB3, CCNB1, CDC16, CDC20, CDC23, MAD1L1, MAD2L1, PTTG1 |
| 4114 | Oocyte meiosis | 10 | 6.60E−13 | ANAPC10, ANAPC4, AURKA, BUB1, CCNB1, CDC16, CDC20, CDC23, MAD2L1, PTTG1 |
| 4914 | Progesterone-mediated oocyte maturation | 8 | 1.84E−10 | ANAPC10, ANAPC4, BUB1, CCNB1, CDC16, CDC23, MAD1L1, MAD2L1 |
| 4120 | Ubiquitin mediated proteolysis | 6 | 1.11E−05 | ANAPC10, ANAPC4, CDC16, CDC20, CDC23, DDB1 |
| 4360 | Axon guidance | 4 | 0.00416 | ROBO1, SLIT1, SLIT2, SLIT3 |
| Hypermethylation and low expression | ||||
| 5200 | Pathways in cancer | 9 | 3.99E−07 | AR, CCND1, CDK4, CDK6, CDKN1A, HHIP, NTRK1, PTCH1, SHH |
| 4115 | p53 signaling pathway | 4 | 0.000145 | CCND1, CDK4, CDK6, CDKN1A |
| 4110 | Cell cycle | 4 | 0.00109 | CCND1, CDK4, CDK6, CDKN1A |
| 4010 | MAPK signaling pathway | 5 | 0.00115 | BDNF, NGF, NTF4, NTRK1, NTRK2 |
| 5205 | Proteoglycans in cancer | 4 | 0.00503 | CCND1, CDKN1A, ESR1, PTCH1 |
Construction and analysis of PPI networks
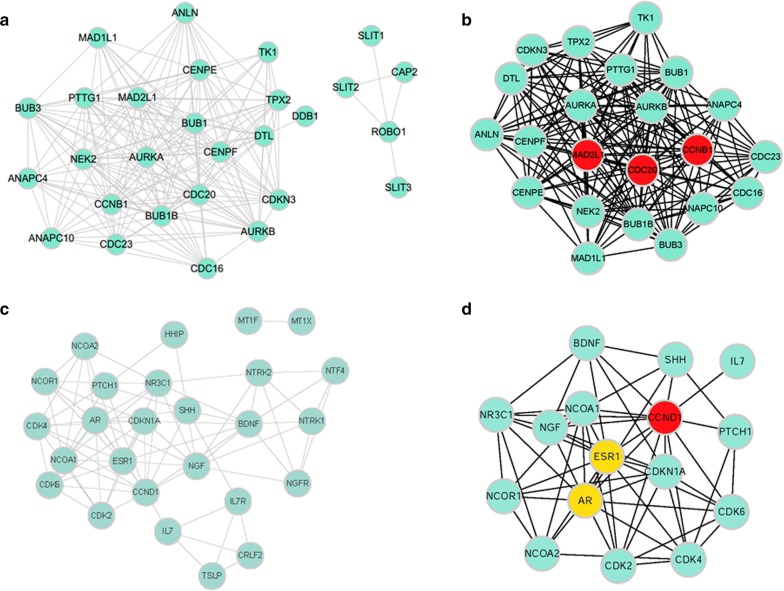
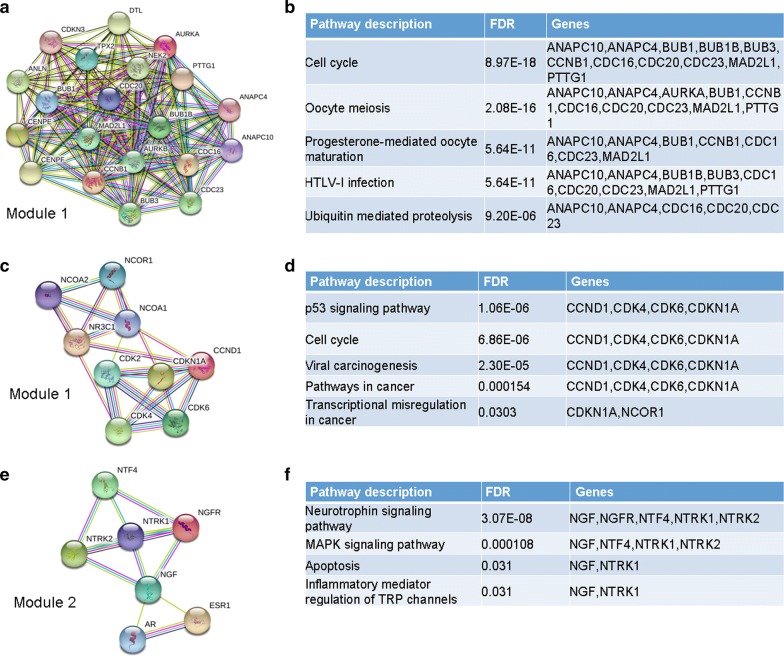
The STRING database was used for PPI network construction, with MCODE used for module analysis. Hub genes were selected using the cytoHubba Cytoscape package. The PPI network for genes that were hypomethylated and highly expressed is shown in Fig. 4a, with corresponding modules shown in Fig. 5a. The most significantly enriched functional modules were those linked to progesterone-mediated oocyte maturation, the cell cycle, HTLV-I infection, oocyte meiosis, and ubiquitin mediated proteolysis (Fig. 5b). Top three hub genes were MAD2L1, CDC20 and CCNB1 (Fig. 4b). The PPI network for genes that were hypermethylated and expressed at low levels is shown in Fig. 4c, with corresponding modules shown in Fig. 5c, e. Significant vital modules showed functions including p53 signaling pathway, cell cycle, viral carcinogenesis, transcriptional misregulation in cancer, neurotrophin signaling pathway, MAPK signaling pathway, apoptosis and inflammatory mediator regulation of TRP channels (Fig. 5d, f). Top three hub genes were CCND1, AR and ESR1 (Fig. 4d). Furthermore, we use Oncomine and UALCAN database to confirm the expression of hub genes in HCC (Fig. 6, Additional file 1: Figure S1). The data are in agreement with our results.
Fig. 4.
PPI network and hub genes of aberrantly methylated and differentially expressed genes. a PPI network and b Hub genes for hypomethylation/high-expression genes; c PPI network and d hub genes of hypermethylation/low-expression genes
Fig. 5.
Core modules for aberrantly methylated and differentially expressed genes. Hypomethylation/high-expression genes a Module 1 and b the enrichment and pathways analysis of module 1; Hypermethylation/low-expression genes: c Module 1 and d the enrichment and pathways analysis of module 1; e Module 2 and f the enrichment and pathways analysis of module 2
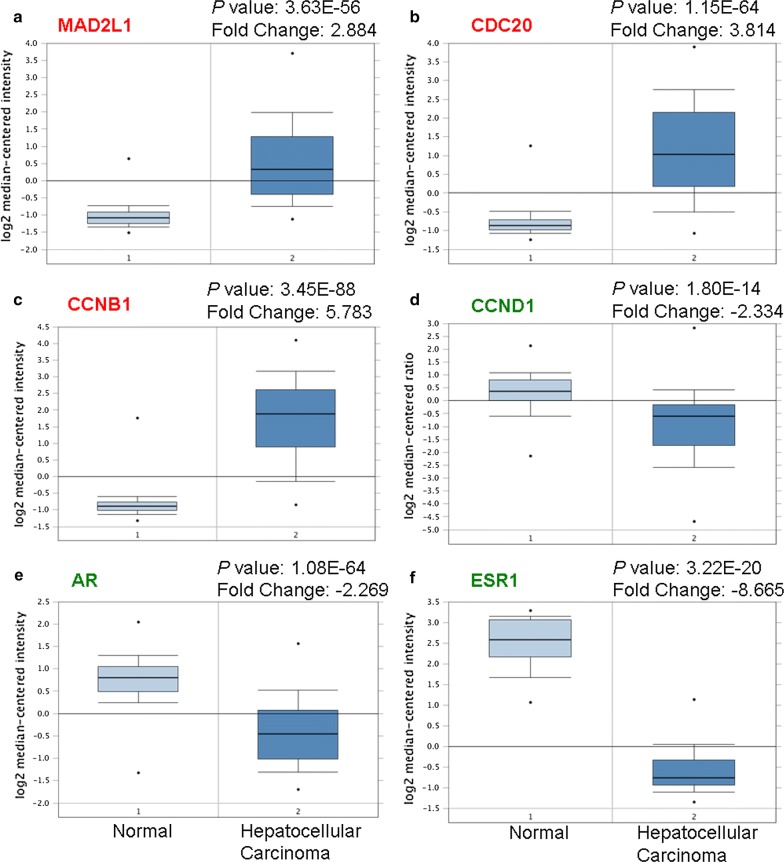
Fig. 6.
Validation of the expression of hub genes in Oncomine database. The expression level of a MAD2L1, b CDC20, c CCNB1, d CCND1, e AR, and f ESR1 were detected in Oncomine database. Red: Hypomethylation/high-expression genes; Green: Hypermethylation/low-expression genes
Hub gene validation
Hub genes for the Hypermethylated/low-expression and Hypomethylation/high-expression datasets were then validated using the TCGA database in order to affirm the validity of these findings. The MEXPRESS web tool allowed for easy and rapid assessment of the relationships between methylation and gene expression within TCGA at a single-gene level. By ordering samples based on gene expression levels, MEXPRESS makes it clear that hub gene expression was negatively correlated with methylation, and a Pearson correlation confirmed this finding. For the hypermethylation/low-expression hub genes, gene expression tended to be higher in normal samples than in tumor samples (Additional file 2: Figure S2). However, tumor samples tended to have higher expression than normal samples for hypomethylation/high-expression hub genes (Additional file 3: Figure S3). The outcomes are summarized in Table 3. Gene expression and methylation were altered and consistent with our findings, indicating that these results are reliable and reproducible.
Table 3.
Validation of the hub genes in TCGA database
| Hub gene | Methylation status | P value | Expression status | P value |
|---|---|---|---|---|
| Hypomethylation/high-expression | ||||
| MAD2L1 | Hypomethylation | < 2.2e−16 | High expression | < 2.2e−16 |
| CDC20 | Hypomethylation | 6.12e−13 | High expression | < 2.2e−16 |
| CCNB1 | Hypomethylation | 2.87e−5 | High expression | < 2.2e−16 |
| Hypermethylation/low-expression | ||||
| CCND1 | Hypermethylation | 1.27e−5 | Low expression | 1.27e−5 |
| AR | Hypermethylation | 2.96e−4 | Low expression | 1.47e−9 |
| ESR1 | Hypermethylation | 0.0258 | Low expression | < 2.2e−16 |
The relationship between hub genes and the survival in hepatocellular carcinoma
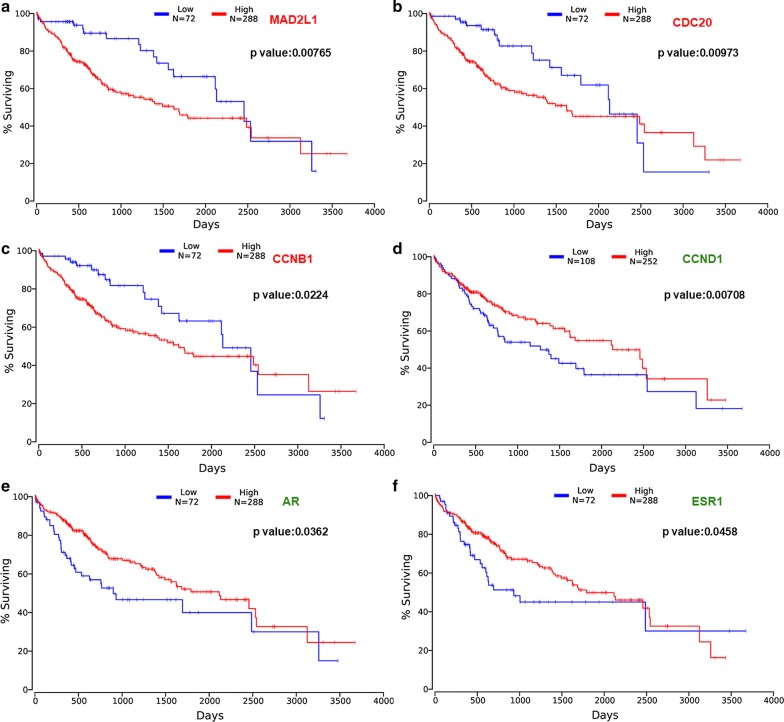
The prognostic value of six hub genes was assessed by OncoLnc database. The threshold was adjusted to cox P value < 0.05. Patients with high expression of MAD2L1, CDC20 and CCNB1 and low expression of CCND1, AR, and ESR1 was associated with shorter overall survival (Fig. 7). The survival data showed that MAD2L1, CDC20 and CCNB1 played an oncogenic role, while CCND1, AR, and ESR1 genes were associated with better survival in HCC.
Fig. 7.
Prognostic value of six hub genes in hepatocellular carcinoma. Prognostic value of a MAD2L1, b CDC20, c CCNB1, d CCND1, e AR, and f ESR1 were obtained in OncoLnc database. The survival curve comparing the patients with high (red) and low (blue) expression in HCC
Discussion
Exploring the mechanisms underlying HCC development and progression not only has prognostic implications, but may also be helpful in monitoring treatment response, surveillance for tumor recurrence, and guidance of clinical decisions. Modern advances in sequencing technologies and microarray development have provided ample high-throughput opportunities to study disease-related biology, allowing for simultaneous assessment of gene methylation and expression for thousands of genes in the human genome. In the present study, we identified 19 hypomethylated, highly-expressed genes and 14 hypermethylated, low-expression genes, using bioinformatic analysis. Functional enrichment of these genes revealed that aberrant methylation indeed affects certain pathways and hub genes. These results can provide novel insight into the explanation of HCC pathogenesis.
The GO enrichment analysis revealed that the primary biological processes of the hypomethylated/highly-expressed genes were the regulation of cell cycle processes, chromosome segregation, and mitotic nuclear division while the hypermethylated/low-expression genes were involved mainly in controlling cell proliferation, gene expression, and the mitotic G1/S transition. This is expected given that the chromosome segregation process occurs during mitosis, which is a part of the cell cycle. The G1/S cell cycle transition is tightly controlled. Deregulation of this key checkpoint can allow cells to undergo transformation, thereby permitting tumorigenesis. This finding is consistent with the fundamental role played by cell cycle regulators in cell proliferation, invasion, and metastasis in HCC. KEGG pathway analysis of hypomethylation/high-expression genes revealed that they were linked to the cell cycle, oocyte meiosis, and ubiquitin-mediated proteolysis. The cell cycle and oocyte meiosis are vital for cell proliferation in tumor cells, and the ubiquitin proteasome pathway functions to regulate cell cycle control and the DNA damage response in tumor genesis [22]. The KEGG pathway analysis of hypermethylation/low-expression genes suggested that methylation may be linked to HCC development through the p53 and MAPK signaling pathways. p53 is a tumor suppressor to conserve genome stability by preventing mutations caused by cellular stress or DNA damage. Together, these results suggest that hypermethylation and hypomethylation are key mediators of cancer development and progression.
The PPI network of hypomethylated/highly-expressed genes provides insight into the functional associations between them, and from this, the top three hub genes were selected: MAD2L1, CDC20, and CCNB1. Mitotic Arrest Deficient 2 Like 1 (MAD2L1) and Cell Division Cycle 20 (CDC20) are two key mitotic checkpoint genes. Both MAD2L1 and CDC20 were more highly expressed in higher grade tumors than in low-grade tumors. High MAD2L1 or CDC20 levels may allow for the development of multi-organ aggressive tumors, including those affecting the breasts, lungs, liver, and stomach [23–25]. Collectively, these findings suggest that MAD2L1 and CDC20 may be key regulators of tumor severity, ultimately dictating patient survival. Cyclin B1 (CCNB1), complexing with CDC2, is a G2/M-phase checkpoint regulator that is vital for regulation of proliferation and DNA synthesis. CCNB1 overexpression has been found to occur in HCC [26] and many other cancer, often being linked to progression, recurrence, and to poor prognoses [27]. Therefore, MAD2L1, CDC20, and CCNB1 may all be abnormally methylated genes that modulate the cell cycle and proliferation in HCC. With regard to the hypermethylated/low-expression genes, the most prominent hub genes were CCND1, AR, and ESR1. Cyclin D1 (CCND1) is a proto-oncogene regulating G1 to S phase progression; it participates in the Wnt/β-catenin pathway [28, 29]. Androgen receptor (AR) is a steroid hormone receptor superfamily member that is involved in human hepatocarcinogenesis [30]. It alters the AR-dependent transcriptome and stimulates oncogenic growth. Estrogen receptor 1 (ESR1) functions as a transcription factor, regulating cell cycle, cell proliferation, apoptosis, and inflammation-associated gene expression [31]. Research has shown that estrogen-depleted postmenopausal women undergo more rapid progression from HCV-infection to HCC development [32]. Aberrant expression of ESR subtypes may contribute to the progression of HCC. These three genes are related to prognosis, tumorigenesis, and metastasis of HCC. Furthermore, survival analysis of hub genes revealed that MAD2L1, CDC20 and CCNB1 play an oncogenic role, while CCND1, AR, and ESR1 genes were associated with favorable patient survival in HCC.
The core PPI network module for hypomethylated/highly-expressed genes was linked to the cell cycle, oocyte meiosis, and ubiquitin-mediated proteolysis, indicating that these pathways are key targets of hypermethylation. The top two modules of the hypermethylated/low-expression gene PPI network were those linked to the p53 signaling pathways, viral carcinogenesis and neurotrophin signaling pathways. p53 signaling conserves the stability of the genome. The leading cause of HCC is infection with HBV or HCV. It is reasonable that viral carcinogenesis is involved in the development of HCC. Neurotrophins and neurotrophin receptors are found on tumor and stromal cells, and are linked to many kinds of tumor development. Brain-derived neurotrophic factor and nerve growth factor are both neurotrophins linked to tumor development, promoting proliferation, angiogenesis, and invasion.
While previously groups have assessed arrays cataloging gene expression or methylation, the two have not been examined simultaneously. Furthermore, single studies generally lack the power needed to identify critical regulatory genes and signaling pathways. Our research used a bioinformatics workflow to jointly analyze extant gene expression and gene methylation profiling microarrays, allowing for more powerful and precise insights into these screening results. However, we only validated candidate abnormally methylated genes that were differentially expressed using the TCGA database. Further experiments will be necessary in order to confirm that these genes and pathways are linked to HCC.
Conclusion
In summary, this study provides a comprehensive bioinformatics analysis of aberrantly methylated DEGs that may be involved in the progression and development of HCC. In addition, six mostly changed hub genes were identified, including MAD2L1, CDC20, CCNB1, CCND1, AR, and ESR1. These novel findings may contribute to the unraveling of the pathogenesis of HCC, and these candidate genes may be optimal abnormal methylation-based biomarkers that can be used to accurately diagnose and treat HCC.
Additional files
Additional file 1: Figure S1. Validation of the expression of hub genes in UALCAN database. Red: Hypomethylation/high-expression genes; Green: Hypermethylation/low-expression genes.
Additional file 2: Figure S2. Validation of the hypermethylation/low-expression hub genes in TCGA database. For the hypermethylation/low-expression hub genes, normal samples tended to have higher expression than tumor samples.
Additional file 3: Figure S3. Validation of the hypomethylation/high-expression hub genes in TCGA database. Tumor samples tended to have higher expression than normal samples for hypomethylation/high-expression hub genes.
Authors’ contributions
Conceived and designed the study strategy: XC, CDW; Acquisition of data: statistical analysis and interpretation of data GRF, YQT; Drafting or revision of the manuscript: YQT, GRF; Reference collection and data management: CC, HYS; Wrote the manuscript: YQT; Prepared the tables and figures: GRF; Study supervision: XC, CDW; All authors read and approved the final manuscript.
Acknowledgements
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Availability of data and materials
The datasets generated and/or analysed during the current study are available in the GEO repository, https://www.ncbi.nlm.nih.gov/geo/.
Consent for publication
Not applicable.
Ethics approval and consent to participate
Not applicable.
Funding
This work was supported by the National Natural Science Foundation of China (No. 81500796) and Research fund of Wuhan Union hospital (No. 02.03.2017-321).
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Abbreviations
- HCC
hepatocellular carcinoma
- DEGs
differentially expressed genes
- DMGs
differentially methylated genes
- PPI
protein-protein interaction
- RASSF1A
Ras association domain family 1 isoform A
- GEO
gene expression omnibus
- KEGG
Kyoto Encyclopedia of Genes and Genomes
- STRING
Search Tool for the Retrieval of Interacting Genes
- MCODE
Molecular Complex Detection
- TCGA
The Cancer Genome Atlas
- GO
gene ontology
- MAD2L1
Mitotic Arrest Deficient 2 Like 1
- CDC20
Cell Division Cycle 20
- CCNB1
cyclin B1
- CCND1
cyclin D1
- AR
androgen receptor
- ESR1
estrogen receptor 1
- NGF
nerve growth factor
- BDNF
brain-derived neurotrophic factor
Contributor Information
Guorun Fan, Email: guorun2006@126.com.
Yaqin Tu, Email: tuyaqin79@163.com.
Cai Chen, Email: 495542137@qq.com.
Haiying Sun, Email: 540007181@qq.com.
Chidan Wan, Phone: 86-27-85726798, Email: chidanwan@163.com.
Xiong Cai, Phone: 86-27-85726798, Email: caixiong@live.com.
References
- 1.Addeo R, Caraglia M, Del Prete S. Highlights of regional meeting of Italian Southern Oncological Group (GOIM): focus on hepatocellular carcinoma: biological and clinical background, therapeutic guide-lines and perspectives. 7 November 2008, Naples, Italy. Expert Opin Investig Drugs. 2009;18(3):373–378. doi: 10.1517/14712590802680158. [DOI] [PubMed] [Google Scholar]
- 2.Ringelhan M, Pfister D, O’Connor T, Pikarsky E, Heikenwalder M. The immunology of hepatocellular carcinoma. Nat Immunol. 2018;19(3):222–232. doi: 10.1038/s41590-018-0044-z. [DOI] [PubMed] [Google Scholar]
- 3.Jaquet A, Tchounga B, Tanon A, Bagny A, Ekouevi DK, Traore HA, Sasco AJ, Maiga M, Dabis F. Etiology of hepatocellular carcinoma in West Africa, a case-control study. Int J Cancer. 2018;143(4):869–877. doi: 10.1002/ijc.31393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ligat G, Schuster C, Baumert TF. HBV core variants, liver fibrosis and hepatocellular carcinoma. Hepatology. 2018 doi: 10.1002/hep.30231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011;61(2):69–90. doi: 10.3322/caac.20107. [DOI] [PubMed] [Google Scholar]
- 6.Venook AP, Papandreou C, Furuse J, de Guevara LL. The incidence and epidemiology of hepatocellular carcinoma: a global and regional perspective. Oncologist. 2010;15(Suppl 4):5–13. doi: 10.1634/theoncologist.2010-S4-05. [DOI] [PubMed] [Google Scholar]
- 7.Younossi Z, Anstee QM, Marietti M, Hardy T, Henry L, Eslam M, George J, Bugianesi E. Global burden of NAFLD and NASH: trends, predictions, risk factors and prevention. Nat Rev Gastroenterol Hepatol. 2018;15(1):11–20. doi: 10.1038/nrgastro.2017.109. [DOI] [PubMed] [Google Scholar]
- 8.Wong CC, Qian Y, Yu J. Interplay between epigenetics and metabolism in oncogenesis: mechanisms and therapeutic approaches. Oncogene. 2017;36(24):3359–3374. doi: 10.1038/onc.2016.485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Yin LJ, Zhang Y, Lv PP, He WH, Wu YT, Liu AX, Ding GL, Dong MY, Qu F, Xu CM, et al. Insufficient maintenance DNA methylation is associated with abnormal embryonic development. BMC Med. 2012;10:26. doi: 10.1186/1741-7015-10-26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Shi B, Thomas AJ, Benninghoff AD, Sessions BR, Meng Q, Parasar P, Rutigliano HM, White KL, Davies CJ. Genetic and epigenetic regulation of major histocompatibility complex class I gene expression in bovine trophoblast cells. Am J Reprod Immunol. 2018;79(1):e12779. doi: 10.1111/aji.12779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Pasque V, Karnik R, Chronis C, Petrella P, Langerman J, Bonora G, Song J, Vanheer L, Sadhu Dimashkie A, Meissner A, et al. X chromosome dosage influences DNA methylation dynamics during reprogramming to mouse iPSCs. Stem Cell Rep. 2018;10(5):1537–1550. doi: 10.1016/j.stemcr.2018.03.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lee CC, Peng SH, Shen L, Lee CF, Du TH, Kang ML, Xu GL, Upadhyay AK, Cheng X, Yan YT, et al. The role of N-alpha-acetyltransferase 10 protein in DNA methylation and genomic imprinting. Mol Cell. 2017;68(1):89–103. doi: 10.1016/j.molcel.2017.08.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hernandez-Saavedra D, Strakovsky RS, Ostrosky-Wegman P, Pan YX. Epigenetic regulation of centromere chromatin stability by dietary and environmental factors. Adv Nutr. 2017;8(6):889–904. doi: 10.3945/an.117.016402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Xu T, Cui T, Peng L, Kong S, Zou J, Tian X. The anti-hepatocellular carcinoma activity of Mel-P15 is mediated by natural killer cells. Oncol Lett. 2017;14(6):6901–6906. doi: 10.3892/ol.2017.7018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Sun G, Zhang C, Feng M, Liu W, Xie H, Qin Q, Zhao E, Wan L. Methylation analysis of p16, SLIT2, SCARA5, and Runx3 genes in hepatocellular carcinoma. Medicine. 2017;96(41):e8279. doi: 10.1097/MD.0000000000008279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zhang X, Guo C, Wu X, Li AX, Liu L, Tsark W, Dammann R, Shen H, Vonderfecht SL, Pfeifer GP. Analysis of liver tumor-prone mouse models of the hippo kinase scaffold proteins RASSF1A and SAV1. Cancer Res. 2016;76(9):2824–2835. doi: 10.1158/0008-5472.CAN-15-3010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Ishak CA, Marshall AE, Passos DT, White CR, Kim SJ, Cecchini MJ, Ferwati S, MacDonald WA, Howlett CJ, Welch ID, et al. An RB-EZH2 complex mediates silencing of repetitive DNA sequences. Mol Cell. 2016;64(6):1074–1087. doi: 10.1016/j.molcel.2016.10.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Wang H, Huo X, Yang XR, He J, Cheng L, Wang N, Deng X, Jin H, Wang C, Zhao F, et al. STAT3-mediated upregulation of lncRNA HOXD-AS1 as a ceRNA facilitates liver cancer metastasis by regulating SOX4. Mol Cancer. 2017;16(1):136. doi: 10.1186/s12943-017-0680-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Mah WC, Thurnherr T, Chow PK, Chung AY, Ooi LL, Toh HC, Teh BT, Saunthararajah Y, Lee CG. Methylation profiles reveal distinct subgroup of hepatocellular carcinoma patients with poor prognosis. PLoS ONE. 2014;9(8):e104158. doi: 10.1371/journal.pone.0104158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Roessler S, Jia HL, Budhu A, Forgues M, Ye QH, Lee JS, Thorgeirsson SS, Sun Z, Tang ZY, Qin LX, et al. A unique metastasis gene signature enables prediction of tumor relapse in early-stage hepatocellular carcinoma patients. Cancer Res. 2010;70(24):10202–10212. doi: 10.1158/0008-5472.CAN-10-2607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Chen X, Cheung ST, So S, Fan ST, Barry C, Higgins J, Lai KM, Ji J, Dudoit S, Ng IO, et al. Gene expression patterns in human liver cancers. Mol Biol Cell. 2002;13(6):1929–1939. doi: 10.1091/mbc.02-02-0023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Tu Y, Chen C, Pan J, Xu J, Zhou ZG, Wang CY. The Ubiquitin Proteasome Pathway (UPP) in the regulation of cell cycle control and DNA damage repair and its implication in tumorigenesis. Int J Clin Exp Pathol. 2012;5(8):726–738. [PMC free article] [PubMed] [Google Scholar]
- 23.Liu X, Zhou W, Zhang X, Ding Y, Du Q, Hu R. 1-L-MT, an IDO inhibitor, prevented colitis-associated cancer by inducing CDC20 inhibition-mediated mitotic death of colon cancer cells. Int J Cancer. 2018;143(6):1516–1529. doi: 10.1002/ijc.31417. [DOI] [PubMed] [Google Scholar]
- 24.Wang Z, Katsaros D, Shen Y, Fu Y, Canuto EM, Benedetto C, Lu L, Chu WM, Risch HA, Yu H. Biological and clinical significance of MAD2L1 and BUB1, genes frequently appearing in expression signatures for breast cancer prognosis. PLoS ONE. 2015;10(8):e0136246. doi: 10.1371/journal.pone.0136246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Schvartzman JM, Sotillo R, Benezra R. Mitotic chromosomal instability and cancer: mouse modelling of the human disease. Nat Rev Cancer. 2010;10(2):102–115. doi: 10.1038/nrc2781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Covini G, Chan EK, Nishioka M, Morshed SA, Reed SI, Tan EM. Immune response to cyclin B1 in hepatocellular carcinoma. Hepatology. 1997;25(1):75–80. doi: 10.1002/hep.510250114. [DOI] [PubMed] [Google Scholar]
- 27.Egloff AM, Vella LA, Finn OJ. Cyclin B1 and other cyclins as tumor antigens in immunosurveillance and immunotherapy of cancer. Cancer Res. 2006;66(1):6–9. doi: 10.1158/0008-5472.CAN-05-3389. [DOI] [PubMed] [Google Scholar]
- 28.Karami F, Mohammadi-Yeganeh S, Abedi N, Koochaki A, Kia V, Paryan M. Bioinformatics prediction and in vitro analysis revealed that miR-17 targets cyclin D1 mRNA in triple negative breast cancer cells. Chem Biol Drug Des. 2016;87(3):317–320. doi: 10.1111/cbdd.12671. [DOI] [PubMed] [Google Scholar]
- 29.Karimkhanloo H, Mohammadi-Yeganeh S, Ahsani Z, Paryan M. Bioinformatics prediction and experimental validation of microRNA-20a targeting Cyclin D1 in hepatocellular carcinoma. Tumour Biol. 2017;39(4):1010428317698361. doi: 10.1177/1010428317698361. [DOI] [PubMed] [Google Scholar]
- 30.Zhang H, Li XX, Yang Y, Zhang Y, Wang HY, Zheng XFS. Significance and mechanism of androgen receptor overexpression and androgen receptor/mechanistic target of rapamycin cross-talk in hepatocellular carcinoma. Hepatology. 2018;67(6):2271–2286. doi: 10.1002/hep.29715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Heldring N, Pike A, Andersson S, Matthews J, Cheng G, Hartman J, Tujague M, Strom A, Treuter E, Warner M, et al. Estrogen receptors: how do they signal and what are their targets. Physiol Rev. 2007;87(3):905–931. doi: 10.1152/physrev.00026.2006. [DOI] [PubMed] [Google Scholar]
- 32.Iyer JK, Kalra M, Kaul A, Payton ME, Kaul R. Estrogen receptor expression in chronic hepatitis C and hepatocellular carcinoma pathogenesis. World J Gastroenterol. 2017;23(37):6802–6816. doi: 10.3748/wjg.v23.i37.6802. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1: Figure S1. Validation of the expression of hub genes in UALCAN database. Red: Hypomethylation/high-expression genes; Green: Hypermethylation/low-expression genes.
Additional file 2: Figure S2. Validation of the hypermethylation/low-expression hub genes in TCGA database. For the hypermethylation/low-expression hub genes, normal samples tended to have higher expression than tumor samples.
Additional file 3: Figure S3. Validation of the hypomethylation/high-expression hub genes in TCGA database. Tumor samples tended to have higher expression than normal samples for hypomethylation/high-expression hub genes.
Data Availability Statement
The datasets generated and/or analysed during the current study are available in the GEO repository, https://www.ncbi.nlm.nih.gov/geo/.