Abstract
Methicillin-resistant Staphylococcus aureus (MRSA) is a ubiquitous human inhabitant and one of the important pathogens of neonatal infections. MRSA is associated with significant mortality and morbidity, especially in very immature preterm neonates. Moreover, MRSA may be implicated in adverse long-term neonatal outcomes, posing a substantial disease burden. Recent advances in molecular microbiology have shed light on the evolution of MRSA population structure and virulence factors, which may contribute to MRSA epidemic waves worldwide. Equipped with remarkable genetic flexibility, MRSA has successfully developed resistance to an extensive range of antibiotics including vancomycin, as well as antiseptics. In the face of these new challenges from MRSA, our armamentarium of anti-infective strategies is very limited and largely dependent on prevention measures. Active surveillance cultures followed by decolonization may be a promising approach to control MRSA infections, with its efficacy and safety in the specific population of neonates yet to be addressed by large multicenter studies.
Keywords: Methicillin-resistant Staphylococcus aureus, Neonatal infections, Preterm infants, Molecular typing, Infection control
Introduction
Since the first reported case of methicillin-resistant Staphylococcus aureus (MRSA) infection in a neonate in the 1980s, MRSA has become a significant cause of neonatal morbidity and mortality [1]. Very immature preterm infants are particularly susceptible to MRSA infections, and neonates with a birth weight (BW) < 1,500 g may account for approximately 80% of all cases of neonatal MRSA infections [2, 3]. As demonstrated by a national cohort study in the USA, the incidence of infections due to MRSA remained stable over the last 2 decades at around 10 per 10,000 hospitalized infants, while that of infections caused by methicillin-susceptible Staphylococcus aureus remarkably decreased [3]. Moreover, recent epidemiologic studies have shed light on evolving MRSA clones and the increasing resistance of MRSA to antimicrobi al substances, indicating the remarkable adaptability of MRSA to anti-infective strategies and its significance as a successful global pathogen [4]. Until 2016, over 20 outbreaks of MRSA in neonatal intensive care units (NICU) have been reported worldwide, with the majority being located in Europe and the USA [5, 6, 7, 8, 9, 10, 11]. These challenges posed by MRSA prompt us to thoroughly and comprehensively investigate its epidemiology, clinical characteristics, and pathogenicity in order to reduce the burden of neonatal MRSA infections.
Literature Search
We searched PubMed for literature published from January 1, 1998, through February 1, 2018, using the following terms: “neonate” or “newborn” or “infant” or “preterm birth” or “preterm infant” or “premature birth,” and “methicillin-resistant Staphylococcus aureus” or “Staphylococcus aureus.” No language restriction was applied. The reference lists of the identified studies and review articles were searched for additional citations.
Neonatal MRSA Colonization and Infections
Current data on MRSA colonization in the general population is limited and mostly based on studies from developed regions. It has been demonstrated that approximately one third of the general population may be colonized by S. aureus [12, 13]. Neonates are likely to acquire S. aureus through the birth canal, breastfeeding, and contact with people and the surrounding environment [14, 15]. In one longitudinal study, colonization rates of S. aureus were reported to be 40–50% during the first 8 weeks of life, followed by a gradual decrease to around 20% at 6 months of age [14]. Similarly to adults, the prevalence of MRSA in the general neonatal population has been estimated to be 1.5% [13, 16], with anterior nares being the predominant site of MRSA colonization [16]. Other parts of the body, such as the umbilicus, the pharynx, the axilla, the groin, and the perineum, harbor MRSA to a lesser extent [14, 16]. Current data demonstrate a higher MRSA colonization rate in hospitalized neonates than in the general neonatal population, ranging from 3.9 to 32% among institutions [5, 6, 17, 18, 19, 20]. This might be explained by the enhanced prevalence of MRSA in healthcare settings compared to the community [12]. In Europe, 8 out of 30 countries reported percentages of MRSA in invasive S. aureus isolates to be above 25% [21]. Besides, hospitalized neonates are in close contact with healthcare workers, whose MRSA carriage rate seems to be 2–3 times higher than in the general population [11, 22]. The wide interinstitutional variance in MRSA colonization rate may be explained by patient demographical characteristics, MRSA screening policies, infection control strategies, and, more importantly, the presence of an MRSA outbreak. Notably, neonates who are not colonized on NICU admission may acquire MRSA within a median of 9 days (range: 1–91 days) [6], and the acquisition rate of MRSA during NICU hospitalization has been reported to be 6.1% (95% CI 2.8–10.6) [16].
Approximately 1 out of 5 neonates colonized with MRSA may develop an infection [5, 6, 18, 19], corresponding to a relative risk of 24.2 (95% CI 8.9–66.0) compared to noncolonized neonates [16]. The median time between MRSA colonization and the onset of infection may range from 4 to 9 days [18, 20]. Prematurity has been identified to be the principal risk factor for MRSA colonization and subsequent infections [2, 17, 19, 23]. The incidence of MRSA infections in neonates with a BW ≤1,000 g has been reported to be 53.4 per 10,000 infants, compared to 23.2, 7.9, and 5.0 per 10,000 infants in the BW categories of 1,001–1,500, 1,501–2,500, and > 2,500 g, respectively [2]. Other risk factors for MRSA infections include prolonged hospitalization, overcrowding and understaffing in neonatal wards, long-term use of respiratory support, intravascular catheters, antibiotics, and total parenteral nutrition, as well as surgical procedures [6, 18, 19]. The clinical manifestation of MRSA infections may range from mild focal infections, such as conjunctivitis and skin and soft tissue infections, to more severe forms like toxic shock syndrome [24] and even invasive infections such as sepsis, necrotizing pneumonia, meningitis, endocarditis, osteomyelitis, liver abscesses, and urinary tract infections [2, 18, 23, 25, 26, 27]. Sepsis, especially late-onset sepsis, is currently the most common manifestation of invasive MRSA infections, with one third of cases presenting as MRSA sepsis [2, 25]. However, though sporadically, early-onset sepsis caused by MRSA has been also reported in industrialized countries [23, 25]. Data from developing countries are lacking. It should be noted that clinical symptoms and signs at the onset of MRSA infections may be highly unspecific [23], underscoring the importance of close patient monitoring and timely access to laboratory and microbiological diagnostics. Currently, the mortality rate of MRSA infections varies widely among institutions, ranging from about 2.9 to 28% [3, 9, 19, 23]. Although there seems to be no difference between MRSA and methicillin-susceptible S. aureus in terms of their clinical presentation and mortality [3, 23, 28], neonates infected with MRSA may have a higher readmission rate and a longer infection course than MSSA cases [26, 29].
Neonatal infections confer an increased risk of short-term morbidity and mortality, as well as adverse long-term outcomes, especially in very immature preterm infants [30, 31, 32]. Few studies have provided pathogen-specific information on MRSA. However, there is a considerable amount of data on S. aureus being the second most common pathogen of neonatal late-onset sepsis in certain countries [33], closely following coagulase-negative staphylococci [34]. As shown by limited population-based studies, the incidence of S. aureus infections may be 0.6 per 1,000 live births [23], 4.0–5.3 per 1,000 neonatal admissions, and 22 per 1,000 hospitalized very low BW (VLBW) infants [3, 23], with MRSA accounting for approximately 28% of all S. aureus infections [3, 26]. In VLBW infants, those with gram-positive bacterial infections had a significantly higher risk of bronchopulmonary dysplasia and cerebral palsy than their counterparts without infections [30, 31, 32]. For infants with infections caused by non-coagulase-negative staphylococci gram-positive bacteria, the adjusted OR of bronchopulmonary dysplasia was shown to be 1.41 (95% CI 1.18–1.68) [30], and the adjusted OR of cerebral palsy appeared to range from 1.5 (95% CI 1.0–2.4) to 4.05 (95% CI 1.48–11.07) [31, 32]. Notably, intraventricular hemorrhage and infarction of the thalamus have been detected during or following S. aureus infection in VLBW infants [35]. The incidence of severe necrotizing enterocolitis and retinopathy of prematurity in very preterm infants with gram-positive bacterial infections was demonstrated to be nearly 3 times higher than in noninfected controls [30]. Additionally, infections caused by non-coagulase-negative staphylococci gram-positive bacteria may be associated with a 2-fold increased risk of vision and head growth impairment compared to uninfected infants [32]. In light of the above information, the burden of short-term and long-term neonatal morbidity caused by MRSA may be substantial, especially in VLBW infants. In the future, multicenter studies are needed to depict the clinical picture of MRSA infections.
Dynamic Shift in MRSA Population
MRSA was first reported in the 1960s, and the resistance of MRSA to methicillin and other β-lactam antibiotics is obtained through a mobile genetic element named staphylococcal chromosome cassette mec (SCCmec) [4]. Until the 1990s, MRSA was considered to be prevalent only within healthcare settings [4]. Since then, epidemic waves of MRSA infections have been reported in communities across continents [4, 36]. Community-associated MRSA (CA-MRSA) is conventionally recognized to be more virulent but less resistant to antibiotics than healthcare-associated MRSA (HA-MRSA) [37]. Genotypic characteristics underlying this phenotype difference include the exclusive carriage of Panton-Valentine leukocidin and a higher expression of other toxins in CA-MRSA than in HA-MRSA [37, 38]. Additionally, small-sized SCCmec types such as IV, V, and VII are overrepresented in CA-MRSA, whereas most HA-MRSA strains carry larger SCCmec types I, II, and III [37]. However, the evolution and transmission of MRSA has contributed to the recent emergence of CA-MRSA in healthcare settings including NICU [4, 17, 18, 26, 36, 37]. Accumulating evidence demonstrates that the genotypic and phenotypic distinction between CA-MRSA and HA-MRSA may have become less clear-cut [4, 36, 37, 38]. Notably, livestock-associated MRSA has emerged as another important source of MRSA infections in communities and healthcare settings, especially in Europe, indicating the significance of zoonotic reservoirs in MRSA transmission [39].
Pathogenic Factors of MRSA
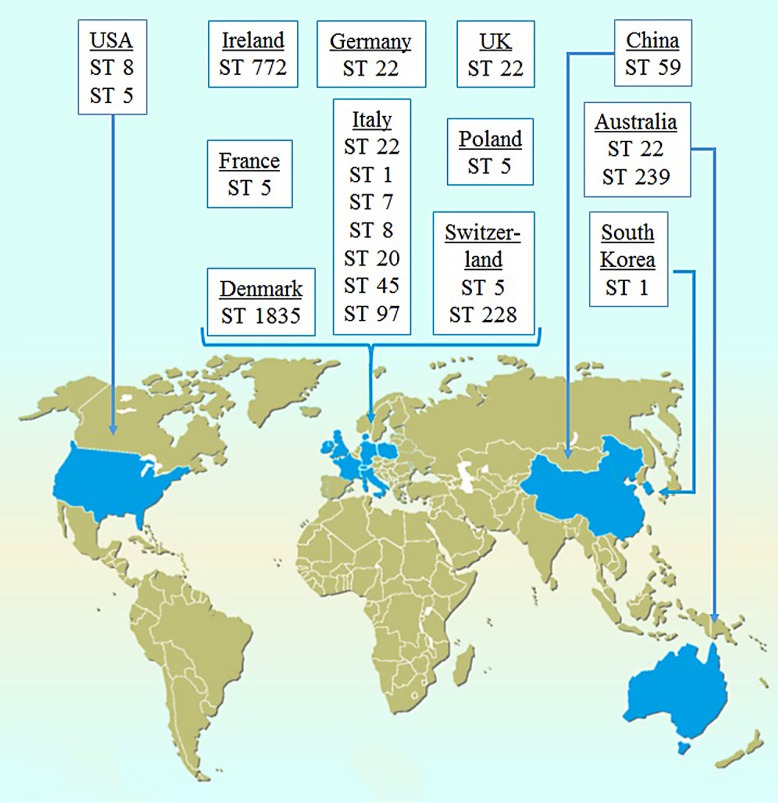
In the past decade, epidemiologic studies have shown that the majority of MRSA infections worldwide are attributable to a limited number of clonal lineages [4, 38, 39]. The most common MRSA strains isolated in neonatal wards belong to sequence types (ST) 1, 5, 8, and 22 (Fig. 1) [6, 7, 8, 9, 17, 18, 19, 22, 40, 41, 42, 43, 44]. Many of these clones are characterized by the ability to form biofilm, toxin production, and genetic flexibility, which will be briefly introduced in the following section. For more details on MRSA virulence, readers are referred to a review elsewhere [45].
Fig. 1.
Predominant methicillin-resistant Staphylococcus aureus clones that have been reported in neonatal infections so far. The given MRSA strains were isolated at one or more neonatal wards in the particular country. ST, sequence type.
Biofilm Formation
Biofilms are a community of bacteria and bacteria-produced extracellular matrix [46]. They constitute a particular growth mode which protects bacteria from the attack of antimicrobial substances and the host immune system [46]. The predominant MRSA clones in the USA and Europe, i.e., ST8 and ST22, have been demonstrated to be strong biofilm producers [47]. The attachment and aggregation of MRSA on indwelling devices and host tissues is largely mediated via clumping factors and fibronectin-, elastin-, and collagen-binding proteins [45]. Apart from their roles in biofilm formation, these surface proteins may trigger an inflammatory response in the host, marked by the release of tumor necrosis factor-α, interleukin-1α (IL-1α), IL-1β, IL-6, and IL-10 [48, 49]. These inflammatory mediators may be implicated in the pathogenesis of S. aureus-induced endocarditis or arthritis, as demonstrated by animal experiments [48, 49].
Toxins
MRSA produces a vast repertoire of toxins, including phenol-soluble modulins, leukotoxins (Panton-Valentine leukocidin, LukED, and LukAB/DH), hemolysins, exfoliative toxins, staphylococcal enterotoxins, and toxic-shock syndrome toxin-1 [50]. Toxins may contribute to MRSA persistence in the host, relapse of infections, tissue damage, and the production of proinflammatory cytokines [47, 51, 52]. The neonatal immune response to MRSA has not been elucidated so far [53]. Some data suggest that neonates are prone to developing hyperinflammation during staphylococcal infections [54, 55]. Such imbalanced inflammation has been considered to be a key underlying mechanism of long-term neonatal morbidities, such as bronchopulmonary dysplasia, necrotizing enterocolitis, retinopathy of prematurity, and cerebral palsy [56].
Genetic Flexibility
The hypervariable genetic background of MRSA mainly lies in its acquisition of mobile genetic elements [37]. The uptake of SCCmec type IV, a Panton-Valentine leukocidin-encoding locus, and an arginine catabolic mobile element by an ancestor ST8 strain results in USA300, which is considered to be the most widespread MRSA clone in the USA and some European countries [37]. While SCCmec and Panton-Valentine leukocidin equip USA300 with antibiotic resistance and invasiveness, the arginine catabolic mobile element contributes to immune evasion and the enhanced survival capacity of MRSA within the host [37, 57]. Another example is the virulence factor phenol soluble modulin-mec, which combines antibiotic resistance and toxin production [51]. Recently, a newly identified gene locus named sasX, which may contribute to MRSA nasal colonization, lung infections, and abscess formation, has been observed to spread from ST239 to other MRSA clones like ST5 [58].
Treatment and Prevention of Neonatal MRSA Infections
Apart from β-lactam antibiotics, a wide range of other antibiotics have been demonstrated to exert a decreased efficacy against MRSA [2, 17], largely due to bacterial genetic mutations and acquisition of plasmids [4, 59]. Although vancomycin is considered to be the drug of last resort for staphylococcal infections, vancomycin-intermediate S. aureus and even vancomycin-resistant S. aureus have been reported [59, 60]. The reduced susceptibility of vancomycin to MRSA poses a great challenge for anti-infective treatment of MRSA in neonates, as vancomycin is the empiric antibiotic of choice in neonates presenting with sepsis and extensive skin infections, especially in regions where the prevalence of MRSA is high [61, 62]. Approaches targeting virulent determinants of MRSA may be promising [50], with their efficacy and safety yet to be validated in the neonatal population. So far, the best strategy to manage neonatal MRSA infections lies in prevention rather than treatment.
MRSA infections are largely preceded by MRSA colonization, and each additional day of exposure to an untreated colonized infant may increase the acquisition risk of MRSA by 6% [63]. Therefore, decolonization may prevent MRSA infections by eliminating the bacterial reservoir [64, 65]. A recent survey showed that 86% of NICU from 58 countries have adopted active surveillance cultures plus MRSA decolonization strategies using nasal mupirocin with or without antiseptic bath [66]. Active surveillance cultures and decolonization have contributed to a dramatic decrease in MRSA colonization and infections in some centers and seem to be cost-effective [67, 68]. However, in other centers, up to 50% of neonates may be persistently colonized or recolonized by MRSA, and a fraction of patients may still develop a subsequent infection despite decolonization [11, 63, 69]. Notably, the current MRSA decolonization protocol largely stems from guidelines for adults [65], and robust evidence regarding its extrapolation to neonates is scarce [70]. Currently, the following questions on MRSA surveillance and decolonization in neonates are yet to be addressed: (1) the implementation of universal versus targeted active surveillance cultures in neonates at high risk for MRSA colonization and infection [10, 64], (2) the definition of optimum surveillance time points to better identify colonized neonates [10, 64], and (3) the benefit of screening extranasal sites for early identification of colonized infants [10, 16, 64]. Moreover, the importance of new screening methods, such as polymerase chain reaction and whole-genome sequencing, needs to be defined [6, 7, 10, 16, 64]. Since MRSA resistant to mupirocin and antiseptics have been reported [18, 44, 71], the efficacy of nasal mupirocin application in primary and persistent MRSA decolonization warrants verification [10, 70, 72], Finally, potential adverse effects of antiseptic baths, such as skin irritation and toxicity [73], as well as nonantimicrobial strategies like maternal skin-to-skin contact for MRSA decolonization remain subjects of discussion [74].
Apart from surveillance and decolonization, strict hand hygiene is universally recognized to be of primary importance in interrupting MRSA transmission [64, 65]. Other measures that may contribute to the prevention of MRSA infections include cohorting and isolation of MRSA-positive patients, barrier precautions, education of healthcare workers, avoidance of ward crowding, and, in some units, surveillance and decolonization of healthcare workers and visiting parents [6, 11, 22, 64, 65, 69].
Conclusion
The enhanced survival rate of very immature preterm infants may contribute to a growing population of infants at risk for MRSA colonization and subsequent infections. Despite abundant data reported from neonatal wards around the world, there is a lack of multicenter or population-based studies to unravel the epidemiology and clinical characteristics of neonatal MRSA colonization and infections. Such data are imperative to accurately assess the disease burden of MRSA and to assist in decision-making processes regarding surveillance and prevention. Furthermore, new methods of MRSA detection and eradication are mandated to confront the rapid evolution of MRSA population structure and pathogenic factors. Due to the rising antibiotic resistance of MRSA and uncertainties in the efficacy and safety of decolonization strategies in neonates, the key to managing neonatal MRSA infections still lies in basic prevention measures.
Disclosure Statement
The authors declare no potential conflict of interests.
Funding Sources
The authors declare no funding sources.
References
- 1.Weeks JL, Garcia-Prats JA, Baker CJ. Methicillin-resistant Staphylococcus aureus osteomyelitis in a neonate. JAMA. 1981;245:1662–1664. [PubMed] [Google Scholar]
- 2.Lessa FC, Edwards JR, Fridkin SK, Tenover FC, Horan TC, Gorwitz RJ. Trends in incidence of late-onset methicillin-resistant Staphylococcus aureus infection in neonatal intensive care units: data from the National Nosocomial Infections Surveillance System, 1995–2004. Pediatr Infect Dis J. 2009;28:577–581. doi: 10.1097/INF.0b013e31819988bf. [DOI] [PubMed] [Google Scholar]
- 3.Ericson JE, Popoola VO, Smith PB, Benjamin DK, Fowler VG, Benjamin DK, Clark RH, Milstone AM. Burden of invasive Staphylococcus aureus infections in hospitalized infants. JAMA Pediatr. 2015;169:1105–1111. doi: 10.1001/jamapediatrics.2015.2380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Uhlemann AC, Otto M, Lowy FD, DeLeo FR. Evolution of community- and healthcare-associated methicillin-resistant Staphylococcus aureus. Infect Genet Evol. 2014;21:563–574. doi: 10.1016/j.meegid.2013.04.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ramsing BG, Arpi M, Andersen EA, Knabe N, Mogensen D, Buhl D, Westh H, Ostergaard C. First outbreak with MRSA in a Danish neonatal intensive care unit: risk factors and control procedures. PLoS One. 2013;8:e66904. doi: 10.1371/journal.pone.0066904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Nübel U, Nachtnebel M, Falkenhorst G, Benz ler J, Hecht J, Kube M, Bröcker F, Moelling K, Bührer C, Gastmeier P, Piening B, Behnke M, Dehnert M, Layer F, Witte W, Eckmanns T. MRSA transmission on a neonatal intensive care unit: epidemiological and genome-based phylogenetic analyses. PLoS One. 2013;8:e54898. doi: 10.1371/journal.pone.0054898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Köser CU, Holden MT, Ellington MJ, Cartwright EJ, Brown NM, Ogilvy-Stuart AL, Hsu LY, Chewapreecha C, Croucher NJ, Harris , SR, Sanders M, Enright MC, Dougan G, Bentley SD, Parkhill J, Fraser LJ, Betley JR, Schulz-Trieglaff OB, Smith GP, Peacock SJ. Rapid whole-genome sequencing for investigation of a neonatal MRSA outbreak. N Engl J Med. 2012;366:2267–2275. doi: 10.1056/NEJMoa1109910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Leroyer C, Lehours P, Tristan A, Boyer F, Marie V, Elleau C, Nolent P, Venier AG, Brissaud O, de Barbeyrac B, Megraud F, Rogues AM. Outbreak in newborns of methicillin-resistant Staphylococcus aureus related to the sequence type 5 Geraldine clone. Am J Infect Control. 2016;44:e9–e11. doi: 10.1016/j.ajic.2015.09.020. [DOI] [PubMed] [Google Scholar]
- 9.Ilczyszyn WM, Sabat AJ, Akkerboom V, Szkarlat A, Klepacka J, Sowa-Sierant I, Wasik B, Kosecka-Strojek M, Buda A, Miedzobrodzki J, Friedrich AW. Clonal structure and characterization of Staphylococcus aureus strains from invasive infections in paediatric patients from south Poland: association between age, spa types, clonal complexes, and genetic markers. PLoS One. 2016;11:e0151937. doi: 10.1371/journal.pone.0151937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Giuffrè M, Bonura C, Cipolla D, Mammina C. MRSA infection in the neonatal intensive care unit. Expert Rev Anti Infect Ther. 2013;11:499–509. doi: 10.1586/eri.13.28. [DOI] [PubMed] [Google Scholar]
- 11.Scheithauer S, Trepels-Kottek S, Häfner H, Keller D, Ittel T, Wagner N, Heimann K, Schoberer M, Schwarz R, Haase G, Waitschies B, Orlikowsky T, Lemmen S. Healthcare worker-related MRSA cluster in a German neonatology level III ICU: a true European story. Int J Hyg Environ Health. 2014;217:307–311. doi: 10.1016/j.ijheh.2013.07.006. [DOI] [PubMed] [Google Scholar]
- 12.Salgado CD, Farr BM, Calfee DP. Community-acquired methicillin-resistant Staphylococcus aureus: a meta-analysis of prevalence and risk factors. Clin Infect Dis. 2003;36:131–139. doi: 10.1086/345436. [DOI] [PubMed] [Google Scholar]
- 13.Gamblin J, Jefferies JM, Harris S, Ahmad N, Marsh P, Faust SN, Fraser S, Moore M, Roderick P, Blair I, Clarke SC. Nasal self-swabbing for estimating the prevalence of Staphylococcus aureus in the community. J Med Microbiol. 2013;62:437–440. doi: 10.1099/jmm.0.051854-0. [DOI] [PubMed] [Google Scholar]
- 14.Peacock SJ, Justice A, Griffiths D, de Silva GD, Kantzanou MN, Crook D, Sleeman K, Day NP. Determinants of acquisition and carriage of Staphylococcus aureus in infancy. J Clin Microbiol. 2003;41:5718–5725. doi: 10.1128/JCM.41.12.5718-5725.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Roca A, Bojang A, Camara B, Oluwalana C, Lette K, West P, D'Alessandro U, Bottomley C. Maternal colonization with Staphylococcus aureus and group B streptococcus is associated with colonization in newborns. Clin Microbiol Infect. 2017;23:974–979. doi: 10.1016/j.cmi.2017.04.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zervou FN, Zacharioudakis IM, Ziakas PD, Mylonakis E. MRSA colonization and risk of infection in the neonatal and pediatric ICU: a meta-analysis. Pediatrics. 2014;133:e1015–e1023. doi: 10.1542/peds.2013-3413. [DOI] [PubMed] [Google Scholar]
- 17.Kuo CY, Huang YC, Huang DT, Chi H, Lu CY, Chang LY, Chi CY, Ho YH, Huang YC, Liu CC, Huang LM. Prevalence and molecular characterization of Staphylococcus aureus colonization among neonatal intensive care units in Taiwan. Neonatology. 2014;105:142–148. doi: 10.1159/000356733. [DOI] [PubMed] [Google Scholar]
- 18.Reich PJ, Boyle MG, Hogan PG, Johnson AJ, Wallace MA, Elward AM, Warner BB, Burnham CA, Fritz SA. Emergence of community-associated methicillin-resistant Staphylococcus aureus strains in the neonatal intensive care unit: an infection prevention and patient safety challenge. Clin Microbiol Infect. 2016;22:645. doi: 10.1016/j.cmi.2016.04.013. e1–e8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Giuffrè M, Amodio E, Bonura C, Geraci DM, Saporito L, Ortolano R, Corsello G, Mammina C. Methicillin-resistant Staphylococcus aureus nasal colonization in a level III neonatal intensive care unit: Incidence and risk factors. Am J Infect Control. 2015;43:476–481. doi: 10.1016/j.ajic.2014.12.027. [DOI] [PubMed] [Google Scholar]
- 20.Kim YH, Chang SS, Kim YS, Kim EA, Yun SC, Kim KS, Pi SY. Clinical outcomes in methicillin-resistant Staphylococcus aureus-colonized neonates in the neonatal intensive care unit. Neonatology. 2007;91:241–247. doi: 10.1159/000098171. [DOI] [PubMed] [Google Scholar]
- 21.European Centre for Disease Prevention and Control Antimicrobial resistance surveillance in Europe in 2014: annual report of the European Antimicrobial Resistance Surveillance Network (EARS-Net) 2015 http://ecdc.europa.eu/en/publications/Publications/antimicrobial-resistance-europe-2014.pdf (accessed December 20, 2017) [Google Scholar]
- 22.Brennan GI, Shore AC, Corcoran S, Tecklenborg S, Coleman DC, O'Connell B. Emergence of hospital- and community-associated Panton-Valentine leukocidin-positive methicillin-resistant Staphylococcus aureus genotype ST772-MRSA-V in Ireland and detailed investigation of an ST772-MRSA-V cluster in a neonatal intensive care unit. J Clin Microbiol. 2012;50:841–847. doi: 10.1128/JCM.06354-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Vergnano S, Menson E, Smith Z, Kennea N, Embleton N, Clarke P, Watts T, Heath PT. Characteristics of invasive Staphylococcus aureus in United Kingdom neonatal units. Pediatr Infect Dis J. 2011;30:850–885. doi: 10.1097/INF.0b013e318224546d. [DOI] [PubMed] [Google Scholar]
- 24.Takahashi N, Uehara R, Nishida H, Sakuma I, Yamasaki C, Takahashi K, Honma Y, Momoi MY, Uchiyama T. Clinical features of neonatal toxic shock syndrome-like exanthematous disease emerging in Japan. J Infect. 2009;59:194–200. doi: 10.1016/j.jinf.2009.06.010. [DOI] [PubMed] [Google Scholar]
- 25.Isaacs D, Fraser S, Hogg G, Li HY. Staphylococcus aureus infections in Australasian neonatal nurseries. Arch Dis Child Fetal Neonatal Ed. 2004;89:F331–F335. doi: 10.1136/adc.2002.009480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Engelman D, Hofer A, Davis JS, Carapetis JR, Baird RW, Giffard PM, Holt DC, Tong SY. Invasive Staphylococcus aureus infections in children in tropical northern Australia. J Pediatric Infect Dis Soc. 2014;3:304–311. doi: 10.1093/jpids/piu013. [DOI] [PubMed] [Google Scholar]
- 27.Filleron A, Lotthé A, Jourdes E, Jeziorski E, Prodhomme O, Didelot MN, Parer S, Marchandin H, Cambonie G. Atypical pneumonia linked to community-acquired Staphylococcus aureus cross-transmission in the nursery. Neonatology. 2013;104:156–160. doi: 10.1159/000351019. [DOI] [PubMed] [Google Scholar]
- 28.Wisgrill L, Zizka J, Unterasinger L, Rittenschober-Böhm J, Waldhör T, Makristathis A, Berger A. Active surveillance cultures and targeted decolonization are associated with reduced methicillin-susceptible Staphylococcus aureus infections in VLBW infants. Neonatology. 2017;112:267–273. doi: 10.1159/000477295. [DOI] [PubMed] [Google Scholar]
- 29.Rana D, Abughali N, Kumar D, Super DM, Jacobs MR, Kumar ML. Staphylococcus aureus, including community-acquired methicillin-resistant S. aureus, in a level III NICU: 2001 to 2008. Am J Perinatol. 2012;29:401–408. doi: 10.1055/s-0032-1304819. [DOI] [PubMed] [Google Scholar]
- 30.Shah J, Jefferies AL, Yoon EW, Lee SK, Shah PS, Canadian Neonatal Network Risk factors and outcomes of late-onset bacterial sepsis in preterm neonates born at <c; 32 weeks' gestation. Am J Perinatol. 2015;32:675–682. doi: 10.1055/s-0034-1393936. [DOI] [PubMed] [Google Scholar]
- 31.Schlapbach LJ, Aebischer M, Adams M, Natalucci G, Bonhoeffer J, Latzin P, Nelle M, Bucher HU, Latal B, Swiss Neonatal Network and Follow-Up Group Impact of sepsis on neurodevelopmental outcome in a Swiss National Cohort of extremely premature infants. Pediatrics. 2011;128:e348–e357. doi: 10.1542/peds.2010-3338. [DOI] [PubMed] [Google Scholar]
- 32.Stoll BJ, Hansen NI, Adams-Chapman I, Fanaroff AA, Hintz SR, Vohr B, Higgins RD, National Institute of Child Health and Human Development Neonatal Research Network Neurodevelopmental and growth impairment among extremely low-birth-weight infants with neonatal infection. JAMA. 2004;292:2357–2365. doi: 10.1001/jama.292.19.2357. [DOI] [PubMed] [Google Scholar]
- 33.Dong Y, Speer CP. Late-onset neonatal sepsis: recent developments. Arch Dis Child Fetal Neonatal Ed. 2015;100:F257–F263. doi: 10.1136/archdischild-2014-306213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Dong Y, Speer CP. The role of Staphylococcus epidermidis in neonatal sepsis: guarding angel or pathogenic devil? Int J Med Microbiol. 2014;304:513–520. doi: 10.1016/j.ijmm.2014.04.013. [DOI] [PubMed] [Google Scholar]
- 35.Claessens LC, Zonnenberg IA, van den Dungen FA, Vermeulen RJ, van Weissenbruch MM. Cerebral ultrasound abnormalities in preterm infants caused by late-onset sepsis. PLoS One. 2017;12:e0173227. doi: 10.1371/journal.pone.0173227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.DeLeo FR, Otto M, Kreiswirth BN, Chambers HF. Community-associated meticillin-resistant Staphylococcus aureus. Lancet. 2010;375:1557–1568. doi: 10.1016/S0140-6736(09)61999-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Otto M. Community-associated MRSA: what makes them special? Int J Med Microbiol. 2013;303:324–330. doi: 10.1016/j.ijmm.2013.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Vandenesch F, Naimi T, Enright MC, Lina G, Nimmo GR, Heffernan H, Liassine N, Bes M, Greenland T, Reverdy ME, Etienne J. Community-acquired methicillin-resistant Staphylococcus aureus carrying Panton-Valentine leukocidin genes: worldwide emergence. Emerg Infect Dis. 2003;9:978–984. doi: 10.3201/eid0908.030089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Grundmann H, Aanensen DM, van den Wijngaard CC, Spratt BG, Harmsen D, Friedrich AW, European Staphylococcal Reference Laboratory Working Group Geographic distribution of Staphylococcus aureus causing invasive infections in Europe: a molecular-epidemiological analysis. PLoS Med. 2010;7:e1000215. doi: 10.1371/journal.pmed.1000215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Schlebusch S, Price GR, Hinds S, Nourse C, Schooneveldt JM, Tilse MH, Liley HG, Wallis T, Bowling F, Venter D, Nimmo GR. First outbreak of PVL-positive nonmultiresistant MRSA in a neonatal ICU in Australia: comparison of MALDI-TOF and SNP-plus-binary gene typing. Eur J Clin Microbiol Infect Dis. 2010;29:1311–1314. doi: 10.1007/s10096-010-0995-y. [DOI] [PubMed] [Google Scholar]
- 41.Song X, Cheung S, Klontz K, Short B, Campos J, Singh N. A stepwise approach to control an outbreak and ongoing transmission of methicillin-resistant Staphylococcus aureus in a neonatal intensive care unit. Am J Infect Control. 2010;38:607–611. doi: 10.1016/j.ajic.2010.02.017. [DOI] [PubMed] [Google Scholar]
- 42.Sax H, Posfay-Barbe K, Harbarth S, Francois P, Touveneau S, Pessoa-Silva CL, Schrenzel J, Dharan S, Gervaix A, Pittet D. Control of a cluster of community-associated, methicillin-resistant Staphylococcus aureus in neonatology. J Hosp Infect. 2006;63:93–100. doi: 10.1016/j.jhin.2005.11.016. [DOI] [PubMed] [Google Scholar]
- 43.Li S, Ning X, Song W, Dong F, Zheng Y, Chen Q, Qiao Y, Li J, Wang L, Wang Q, Yue L, Wang A, Yu S, Yao K, Yang Y, Shen X. Clinical and molecular characteristics of community-acquired methicillin-resistant Staphylococcus aureus infections in Chinese neonates. APMIS. 2015;123:28–36. doi: 10.1111/apm.12304. [DOI] [PubMed] [Google Scholar]
- 44.Park SH, Kim SY, Lee JH, Park C, Lee DG. Community-genotype strains of methicillin-resistant Staphylococcus aureus with high-level mupirocin resistance in a neonatal intensive care unit. Early Hum Dev. 2013;89:661–665. doi: 10.1016/j.earlhumdev.2013.04.014. [DOI] [PubMed] [Google Scholar]
- 45.Otto M. MRSA virulence and spread. Cell Microbiol. 2012;14:1513–1521. doi: 10.1111/j.1462-5822.2012.01832.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Costerton JW, Stewart PS, Greenberg EP. Bacterial biofilms: a common cause of persistent infections. Science. 1999;284:1318–1322. doi: 10.1126/science.284.5418.1318. [DOI] [PubMed] [Google Scholar]
- 47.Baldan R, Testa F, Lorè NI, Bragonzi A, Cichero P, Ossi C, Biancardi A, Nizzero P, Moro M, Cirillo DM. Factors contributing to epidemic MRSA clones replacement in a hospital setting. PLoS One. 2012;7:e43153. doi: 10.1371/journal.pone.0043153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Veloso TR, Chaouch A, Roger T, Giddey M, Vouillamoz J, Majcherczyk P, Que YA, Rousson V, Moreillon P, Entenza JM. Use of a human-like low-grade bacteremia model of experimental endocarditis to study the role of Staphylococcus aureus adhesins and platelet aggregation in early endocarditis. Infect Immun. 2013;81:697–703. doi: 10.1128/IAI.01030-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Josefsson E, Juuti K, Bokarewa M, Kuusela P. The surface protein Pls of methicillin-resistant Staphylococcus aureus is a virulence factor in septic arthritis. Infect Immun. 2005;73:2812–2817. doi: 10.1128/IAI.73.5.2812-2817.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Kong C, Neoh HM, Nathan S. Targeting Staphylococcus aureus toxins: a potential form of anti-virulence therapy. Toxins (Basel) 2016;8:E72. doi: 10.3390/toxins8030072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Queck SY, Khan BA, Wang R, Bach TH, Kretschmer D, Chen L, Kreiswirth BN, Peschel A, Deleo FR, Otto M. Mobile genetic element-encoded cytolysin connects virulence to methicillin resistance in MRSA. PLoS Pathog. 2009;5:e1000533. doi: 10.1371/journal.ppat.1000533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Kretschmer D, Nikola N, Dürr M, Otto M, Peschel A. The virulence regulator Agr controls the staphylococcal capacity to activate human neutrophils via the formyl peptide receptor 2. J Innate Immun. 2012;4:201–212. doi: 10.1159/000332142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Power Coombs MR, Kronforst K, Levy O. Neonatal host defense against staphylococcal infections. Clin Dev Immunol. 2013;2013:826303. doi: 10.1155/2013/826303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Glaser K, Speer CP. Toll-like receptor signaling in neonatal sepsis and inflammation: a matter of orchestration and conditioning. Expert Rev Clin Immunol. 2013;9:1239–1252. doi: 10.1586/1744666X.2013.857275. [DOI] [PubMed] [Google Scholar]
- 55.Zhang JP, Yang Y, Levy O, Chen C. Human neonatal peripheral blood leukocytes demonstrate pathogen-specific coordinate expression of TLR2, TLR4/MD2, and MyD88 during bacterial infection in vivo. Pediatr Res. 2010;68:479–483. doi: 10.1203/PDR.0b013e3181f90810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Dong Y, Speer CP, Glaser K. Beyond sepsis: Staphylococcus epidermidis is an underestimated but significant contributor to neonatal morbidity. Virulence. 2018;9:621–633. doi: 10.1080/21505594.2017.1419117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Planet PJ, LaRussa SJ, Dana A, Smith H, Xu A, Ryan C, Uhlemann AC, Boundy S, Goldberg J, Narechania A, Kulkarni R, Ratner AJ, Geoghegan JA, Kolokotronis SO, Prince A. Emergence of the epidemic methicillin-resistant Staphylococcus aureus strain USA300 coincides with horizontal transfer of the arginine catabolic mobile element and speG-mediated adaptations for survival on skin. MBio. 2013;4:00889–13. doi: 10.1128/mBio.00889-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Li M, Du X, Villaruz AE, Diep BA, Wang D, Song Y, Tian Y, Hu J, Yu F, Lu Y, Otto M. MRSA epidemic linked to a quickly spreading colonization and virulence determinant. Nat Med. 2012;18:816–819. doi: 10.1038/nm.2692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Hafer C, Lin Y, Kornblum J, Lowy FD, Uhlemann AC. Contribution of selected gene mutations to resistance in clinical isolates of vancomycin-intermediate Staphylococcus aure us. Antimicrob Agents Chemother. 2012;56:5845–5851. doi: 10.1128/AAC.01139-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Pogorzelska-Maziarz M, Carter EJ, Manning ML, Larson EL. State health department requirements for reporting of antibiotic-resistant infections by providers, United States, 2013 and 2015. Public Health Rep. 2017;132:32–36. doi: 10.1177/0033354916681507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Sivanandan S, Soraisham AS, Swarnam K. Choice and duration of antimicrobial therapy for neonatal sepsis and meningitis. Int J Pediatr. 2011;2011:712150. doi: 10.1155/2011/712150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Liu C, Bayer A, Cosgrove SE, Daum RS, Fridkin SK, Gorwitz RJ, Kaplan SL, Karchmer AW, Levine DP, Murray BE, Rybak M, Talan DA, Chambers HF, Infectious Diseases Society of America Clinical practice guidelines by the Infectious Diseases Society of America for the treatment of methicillin-resistant Staphylococcus aureus infections in adults and children. Clin Infect Dis. 2011;52:e18–e55. doi: 10.1093/cid/ciq146. [DOI] [PubMed] [Google Scholar]
- 63.Pierce R, Lessler J, Popoola VO, Milstone AM. Meticillin-resistant Staphylococcus aureus (MRSA) acquisition risk in an endemic neonatal intensive care unit with an active surveillance culture and decolonization programme. J Hosp Infect. 2017;95:91–97. doi: 10.1016/j.jhin.2016.10.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Newnam KM. Surveillance and isolation of methicillin-resistant Staphylococcus aureus colonization in the neonatal intensive care unit. Adv Neonatal Care. 2016;16:298–307. doi: 10.1097/ANC.0000000000000312. [DOI] [PubMed] [Google Scholar]
- 65.Gerber SI, Jones RC, Scott MV, Price JS, Dworkin MS, Filippell MB, Rearick T, Pur SL, McAuley JB, Lavin MA, Welbel SF, Garcia-Houchins S, Bova JL, Weber SG, Arnow PM, Englund JA, Gavin PJ, Fisher AG, Thomson RB, Vescio T, Chou T, Johnson DC, Fry MB, Molloy AH, Bardowski L, Noskin GA. Management of outbreaks of methicillin-resistant Staphylococcus aureus infection in the neonatal intensive care unit: a consensus statement. Infect Control Hosp Epidemiol. 2006;27:139–145. doi: 10.1086/501216. [DOI] [PubMed] [Google Scholar]
- 66.Milstone AM, Song X, Coffin S, Elward A, Society for Healthcare Epidemiology of America's Pediatric Special Interest Group Identification and eradication of methicillin-resistant Staphylococcus aureus colonization in the neonatal intensive care unit: results of a national survey. Infect Control Hosp Epidemiol. 2010;31:766–768. doi: 10.1086/653615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.You JH, Chan CY, Wong MY, Ip M. Active surveillance and decolonization of methicillin-resistant Staphylococcus aureus on admission to neonatal intensive care units in Hong Kong: a cost-effectiveness analysis. Infect Control Hosp Epidemiol. 2012;33:1024–1030. doi: 10.1086/667735. [DOI] [PubMed] [Google Scholar]
- 68.Morioka I, Takahashi N, Kitajima H. Prevalence of MRSA-colonization patients in Japanese neonatal care units in 2011. Pediatr Int. 2014;56:211–214. doi: 10.1111/ped.12232. [DOI] [PubMed] [Google Scholar]
- 69.Popoola VO, Budd A, Wittig SM, Ross T, Aucott SW, Perl TM, Carroll KC, Milstone AM. Methicillin-resistant Staphylococcus aureus transmission and infections in a neonatal intensive care unit despite active surveillance cultures and decolonization: challenges for infection prevention. Infect Control Hosp Epidemiol. 2014;35:412–418. doi: 10.1086/675594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Nelson MU, Bizzarro MJ, Dembry LM, Baltimore RS, Gallagher PG. One size does not fit all: why universal decolonization strategies to prevent methicillin-resistant Staphylococcus aureus colonization and infection in adult intensive care units may be inappropriate for neonatal intensive care units. J Perinatol. 2014;34:653–655. doi: 10.1038/jp.2014.125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Fritz SA, Hogan PG, Camins BC, Ainsworth AJ, Patrick C, Martin MS, Krauss MJ, Rodriguez M, Burnham CA. Mupirocin and chlorhexidine resistance in Staphylococcus aureus in patients with community-onset skin and soft tissue infections. Antimicrob Agents Chemother. 2013;57:559–568. doi: 10.1128/AAC.01633-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Pierce R, Bryant K, Elward A, Lessler J, Milstone AM. Bacterial infections in neonates following mupirocin-based MRSA decolonization: a multicenter cohort study. Infect Control Hosp Epidemiol. 2017;38:930–936. doi: 10.1017/ice.2017.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Chapman AK, Aucott SW, Milstone AM. Safety of chlorhexidine gluconate used for skin antisepsis in the preterm infant. J Perinatol. 2012;32:4–9. doi: 10.1038/jp.2011.148. [DOI] [PubMed] [Google Scholar]
- 74.Lamy Filho F, de Sousa SH, Freitas IJ, Lamy ZC, Simões VM, da Silva AA, Barbieri MA. Effect of maternal skin-to-skin contact on decolonization of methicillin-oxacillin-resistant Staphylococcus in neonatal intensive care units: a randomized controlled trial. BMC Pregnancy Childbirth. 2015;15:63. doi: 10.1186/s12884-015-0496-1. [DOI] [PMC free article] [PubMed] [Google Scholar]