Abstract
Mitosis encompasses key molecular changes including chromatin condensation, nuclear envelope breakdown, and reduced transcription levels. Immediately after mitosis, the interphase chromatin structure is reestablished and transcription resumes. The reestablishment of the interphase chromatin is probably achieved by “bookmarking,” i.e., the retention of at least partial information during mitosis. To gain a deeper understanding of the contribution of histone modifications to the mitotic bookmarking process, we merged proteomics, immunofluorescence, and ChIP-seq approaches. We focused on key histone modifications and employed HeLa-S3 cells as a model system. Generally, in spite of the general hypoacetylation observed during mitosis, we observed a global concordance between the genomic organization of histone modifications in interphase and mitosis, suggesting that the epigenomic landscape may serve as a component of the mitotic bookmarking process. Next, we investigated the nucleosome that enters nucleosome depleted regions (NDRs) during mitosis. We observed that in ∼60% of the NDRs, the entering nucleosome is distinct from the surrounding highly acetylated nucleosomes and appears to have either low levels of acetylation or high levels of phosphorylation in adjacent residues (since adjacent phosphorylation may interfere with the ability to detect acetylation). Inhibition of histone deacetylases (HDACs) by the small molecule TSA reverts this pattern, suggesting that these nucleosomes are specifically deacetylated during mitosis. Altogether, by merging multiple approaches, our study provides evidence to support a model where histone modifications may play a role in mitotic bookmarking and uncovers new insights into the deposition of nucleosomes during mitosis.
The cell cycle is a dynamic orchestrated process that involves major changes to the chromatin structure (Raynaud et al. 2014) and the expression of hundreds of genes (Spellman et al. 1998; Cho et al. 2001; Whitfield et al. 2002; Bar-Joseph et al. 2008). During the cell cycle, DNA is replicated during S-phase and then condensed in mitosis prior to cell division.
During mitosis, the cell undergoes profound changes. The nuclear envelope breaks apart and the chromosomes change their shape into distinguishable thick structures. Multiple studies have examined the structure of mitotic chromosomes using microscopy (Vagnarelli 2013), but they lack fine-detail information on individual loci. Recently, genome-wide long-range interactions within mitotic chromosomes were investigated by Hi-C and 4C (Naumova et al. 2013; Dileep et al. 2015; Stevens et al. 2017). These studies demonstrate the disappearance of the consistent higher-order chromatin structure in mitosis, including both genome compartments and topological associated domains (TADs). Nevertheless, the unique features of mitotic chromatin at a finer genomic resolution remain unexplored. In addition to chromosome reshaping, mitotic chromosomes are characterized by the disassociation of numerous proteins including RNA polymerase II and many gene-specific as well as general transcription factors (Martinez-Balbas et al. 1995; Parsons and Spencer 1997; Kadauke and Blobel 2013). The disassociation of these transcription machinery components leads to a dramatic decrease in transcription (King and Barnhisel 1967; Gottesfeld and Forbes 1997; Liang et al. 2015; Palozola et al. 2017).
Immediately after mitosis, the chromatin structure is gradually reestablished—the chromosomes decondense throughout G1 (Belmont and Bruce 1994), the interphase TAD structures are reformed within 4 h after cytokinesis (Dileep et al. 2015), and transcription is reinitiated at the beginning of G1 (Prescott and Bender 1962; Gottesfeld and Forbes 1997; Prasanth et al. 2003). This complex orchestration of TAD reformation and controlled transcription reinitiation necessitates precise epigenomic markings, which are likely preserved during mitosis despite the radical changes to the chromatin. How is this epigenomic information stored during mitosis? The current view is that partial information is retained on mitotic chromosomes to serve as bookmarks for the reestablishment of the chromatin structure. In line with this view, even in their condensed state, mitotic chromosomes remain accessible, as is evident by both biophysical measurements (Chen et al. 2005; Caravaca et al. 2013) and DNase I sensitivity assays (Gazit et al. 1982; Kadauke et al. 2012). Furthermore, genomic chromatin accessibility maps by DNase I sensitivity and ATAC-seq comparing mitotic and interphase cells revealed that accessibility is generally preserved and is only locally modulated during mitosis (Hsiung et al. 2015; Teves et al. 2016). Yet, it remains unclear what serves as the bookmarks for the restoration of the global chromatin structure following the changes that occur during mitosis.
Much effort has been devoted to identifying components of the transcription machinery that may serve as cell cycle bookmarks. These studies focused mainly on general as well as specific transcription factors that are retained at a subset of their locations during mitosis (for review, see Kadauke and Blobel 2013; Zaidi et al. 2014). Additionally, preliminary evidence suggests that histone tail modifications themselves participate in carrying out the task of bookmarking. It has been reported that retaining H4K5ac levels during mitosis is crucial for the bookmarking of a reporter gene (Zhao et al. 2011). Further, the global levels of several histone modifications were shown to be preserved during mitosis (for review, see Wang and Higgins 2013). However, the mitotic histone modification levels were mostly assessed globally (e.g., by methods such as Western blots, mass spectrometry, or immuno-staining) and therefore lack spatial genomic information, with few exceptions where ChIP experiments demonstrated a mitotic preservation of histone acetylations at the promoters of several transcribed genes (Kouskouti and Talianidis 2005; Valls et al. 2005). An initial clue regarding the genome-wide organization of histone modifications was achieved by employing ChIP-seq to map H3K4me3, H3K27ac, and H2A.Z in interphase and mitotic cells. The genome-wide maps of H3K4me3 and H2A.Z revealed that the general genomic localization of these marks is conserved upon mitosis entry (Kelly et al. 2010; Liang et al. 2015). Conversely, most H3K27ac genome-wide studies describe a degree of locus specificity in the maintenance of H3K27ac during mitosis (Hsiung et al. 2016; Li et al. 2017; Liu et al. 2017a,b).
While the genomic localization of H3K4me3 and H2A.Z appears to be highly concordant between mitosis and interphase, the patterns of these histone marks were shown to dramatically differ between mitosis and interphase in specific genomic areas termed nucleosome depleted regions (NDRs) (Kelly et al. 2010; Liang et al. 2015). NDRs are loci highly depleted of nucleosomes and hence differ from the mostly uniform distribution of nucleosomes across the genome (Lee et al. 2004). NDRs are mainly located within promoters, enhancers, and insulators and thus are enriched with RNA polymerase and transcription factor binding sites (Fu et al. 2008; Henikoff 2008; Tsompana and Buck 2014). Promoter-associated NDRs were shown to disappear during mitosis and contain a nucleosome harboring H3K4me3 (Liang et al. 2015) and H2A.Z (Kelly et al. 2010). However, the histone composition of the nucleosome that occupies these NDRs, as well as the NDR occupation mechanism, remain unclear.
To better understand the role of mitotic histone modifications and nucleosome de-positioning, we investigated the global dynamics and genomic organization of histone modifications in mitosis and in interphase. Using HeLa-S3 cells as a model system, our study incorporated a synchronization approach that enables the collection of relatively pure preparations of mitotic cells; proteomics analysis focused on quantitative evaluation of the global levels of histone modifications; and finally, systematic genome localization mapping of RNA polymerase II (RNA PolII) and six key histone modifications (H3K4me1, H3K4me3, H3K9ac, H3K27ac, H3K27me3, and H3K36me3). Altogether, integration of these data provided two major insights. First, the epigenomic landscape does not change considerably during mitosis. Second, nucleosomes surrounding the NDR of promoters, enhancers, and insulators are repositioned and modified during mitosis.
Results
Efficient metaphase synchronization of HeLa-S3 cells using a kinesin 5 inhibitor (STC)
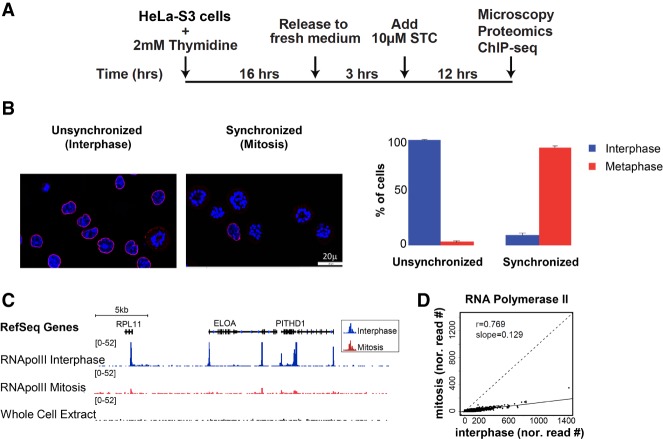
Mitotic cells constitute only about 2% of a nonsynchronized interphase population. Therefore, determining the chromatin landscape by proteomics and ChIP-seq of this cell cycle phase requires the generation of relatively pure preparations of mitotic cells. The kinesin 5 inhibitor STC (Skoufias et al. 2006) inhibits both separation of the duplicated centrosomes as well as bipolar spindle formation, resulting in cells arrested in a metaphase-like stage with monoastral spindles (Fig. 1A). Following treatment of HeLa-S3 cells with STC, we were able to achieve tight synchronization, with an average of ∼90% (SE = 1.6%; n = 8) of cells in a metaphase-like stage of mitosis (Fig. 1B). For simplicity, we refer below to the unsynchronized population as “interphase” cells and to the synchronized one as “mitosis.”
Figure 1.
Efficient metaphase synchronization of HeLa-S3 cells using a kinesin 5 inhibitor (STC) reveals low levels of RNA PolII binding during metaphase. (A) Schematic representation of the synchronization approach. (B) Synchronization was monitored by microscopy using both DAPI (blue) and Lamin B (red) staining. Only cells with the monopolar spindle appearance of the chromosomes and that lack a nuclear envelope were counted as cells in the metaphase stage. The percentage of cells in interphase (blue) and metaphase (red) are shown for both conditions, along with the standard error (SE). For simplicity, we refer to the unsynchronized population as “interphase” and to the synchronized one as “mitosis.” (C) Integrative Genomics Viewer (IGV) (Robinson et al. 2011; Thorvaldsdottir et al. 2013) tracks showing the alignment for RNA PolII ChIP-seq results on an ∼180-kb region on Chromosome 1. The image demonstrates reduced levels of RNA PolII binding in the metaphase sample (red) compared to the interphase sample (blue). (D) A scatter plot showing the normalized read counts at RNA PolII peaks (see Methods for more details) in mitosis versus interphase.
RNA PolII binding is maintained during mitosis at low levels
In accordance with previous studies (Parsons and Spencer 1997), we observed a dramatic reduction in RNA PolII occupancy in the mitotic samples (Fig. 1C,D). Nevertheless, in both interphase and mitotic samples, RNA PolII binding was almost exclusively at promoters (Supplemental Fig. S1). Recent findings suggest that a cell's transcription pattern is largely retained at a low level throughout mitosis (Palozola et al. 2017). However, we cannot rule out the possibility that the low peaks we observe in mitosis are due to the interphase cells that constitute ∼10% of our mitotic samples.
Identification of global changes in histone modifications via mass spectrometry-based proteomic analysis
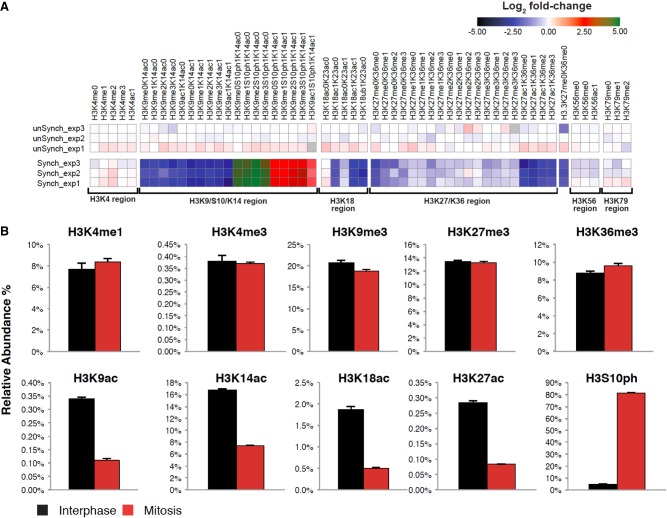
To gain insight into the global changes in histone modifications during mitosis, we employed a quantitative mass spectrometry (MS)-based approach (Jaffe et al. 2013). By implementing synthetic peptide standards, this MS approach provides high reproducibility and is ideal for comparing histone modification levels between mitosis and interphase. Using this method, we profiled the patterns of histone modifications in three biological replicates of mitotic and interphase samples (Fig. 2; Supplemental Fig. S2). In line with previous observations (Hendzel et al. 1997; Van Hooser et al. 1998; Johansen and Johansen 2006), the H3S10ph modification was highly enriched in the mitotic samples compared to interphase (∼17-fold; P-value < 10−3; two sided t-test) (Fig. 2), lending additional support to our synchronization approach. While we had no synthetic peptides in the assay that explicitly monitored the presence of H3S28ph (either alone or in combination with other marks), we could infer the presence of this modification by observing precursor m/z species consistent with the adduction of 79.966 Da on the corresponding H3 unphosphorylated peptides (e.g., H3K27me0K36me0 and H3K27ac1K36me0). H3S28ph was, like H3S10ph, enriched in the mitotic samples compared to interphase (see Supplemental Note for details). We further estimated the absolute occupancy percentage of individual histone marks by comparing mass spectral intensity vs. known concentrations of internal standards and summarizing the results. While the global levels of histone methylations were mostly similar between mitosis and interphase, histone acetylations as well as K18 ubiquitination were reduced in mitotic samples (approximately three- to sixfold; P-value < 0.015; two sided t-test) (Fig. 2).
Figure 2.
Identification of global changes in histone H3 modifications by quantitative targeted mass spectrometry analysis. (A) Heat map (log2 of fold change) of the different histone modifications as detailed on top. “Regions” (all peptides associated with specific modified/unmodified residue/s) are shown at the bottom. The peptides from these regions were then used to calculate the occupancy percentage of each specific histone modification presented in B. Data were normalized to the average signal of interphase samples. (B) Relative abundance of selected modifications on H3 tails from interphase or mitotic HeLa-S3 cells. The data were obtained by quantitative targeted mass spectrometry analysis. Each bar represents the percentage of the H3 peptides with the indicated modifications within the total H3 tail peptide population. Note the high levels of the mitosis unique modification, H3S10ph (∼17-fold increase, P-value < 10−3; two sided t-test), which provides evidence supporting the tight synchronization of the mitotic samples. Similar results were obtained for H4 modifications (Supplemental Fig. S2). Also note the general decrease in the level of all forms of the K27/K36 peptide in mitosis. This is most probably a result of the mitosis-associated phosphorylation of serine 28 (S28), which is located on the same peptide, reducing the abundance of the nonphosphorylated forms of this peptide in mitosis. We did not include the S28 phosphorylation in our analysis due to lack of an appropriate standard peptide. Yet, we can still infer the presence of the H3S28ph modification, and as detailed in the Supplemental Note, we do not expect the reduced confidence we have in detecting H3S28ph to impact our estimates of the total population measurements of H3.
We confirmed the distinction between histone methylation and acetylation by performing quantitative immunofluorescence and measuring the mitotic and interphase levels of H3K9ac and H3K4me3. While the levels of H3K4me3 were almost identical in mitosis and interphase as measured by immunofluorescence, H3K9ac levels were reduced in mitosis by 2.7-fold (P-value < 10−23; t-test) (Supplemental Fig. S3). Immunofluorescence measurements alone are not sufficient for accurate quantification of modification levels since antibody binding during mitosis may be influenced by chromatin condensation or H3S10 phosphorylation. Nevertheless, obtaining similar results from two independent techniques, i.e., immunofluorescence and mass spectrometry, strongly supports the conclusion that there is a key distinction between the protein level dynamics of histone acetylation versus methylation in the transition from interphase to mitosis. However, these results do not show how this difference is reflected spatially in the genomic organization of these histone modifications.
Analysis of the epigenomic landscape reveals high concordance between mitotic and interphase localization of histone marks
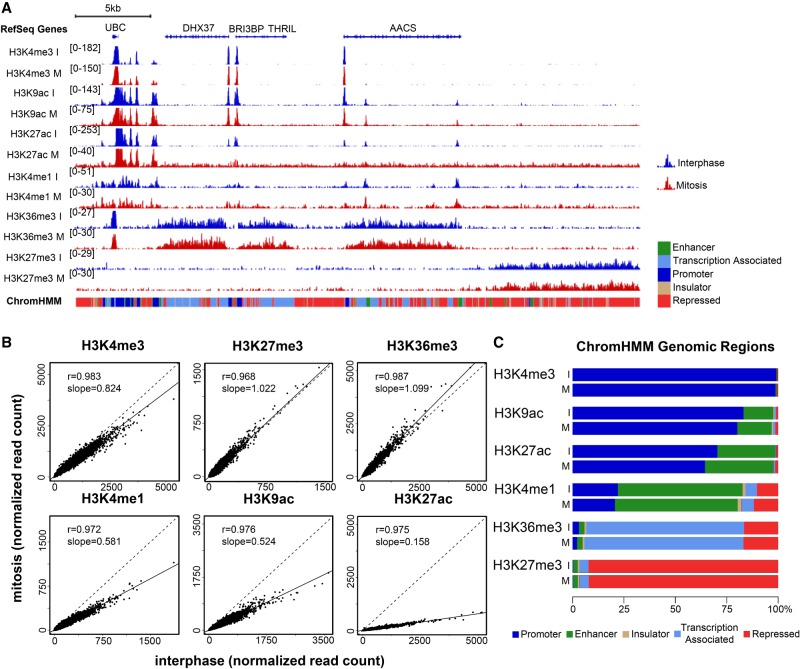
To follow up on the observations from the proteomics analysis and compare the genomic localization of histone marks between mitosis and interphase, we performed ChIP-seq focusing on key histone modifications. We chose six major histone modifications that together capture main chromatin features, including active promoters (H3K4me3, H3K9ac, and H3K27ac), putative enhancers (H3K4me1 and H3K27ac, which is also associated with active promoters), gene bodies (H3K36me3), and repressed regions (H3K27me3) (Fig. 3; Zhou et al. 2011). We found no major changes between the overall epigenomic landscape of the mitotic and interphase samples in two biological replicates (Fig. 3A; Supplemental Fig. S4). While this observation was true for all histone modifications examined, there is a clear distinction between methylation marks, which seem to remain intact (with one exception discussed below) and acetylation marks, which show a universal and uniform signal reduction in mitosis (Fig. 3B). This is in line with our proteomics and immunofluorescence observations regarding the global levels of the modifications in these cell cycle phases.
Figure 3.
High concordance between mitotic and interphase histone modification patterns. (A) Integrative Genomics Viewer (Robinson et al. 2011; Thorvaldsdottir et al. 2013) tracks showing the alignments for the indicated histone modification ChIP-seq results for interphase (blue) and mitosis (red) samples on an ∼315-kb region on Chromosome 12. The scale of each track was adjusted to the total number of reads using the Normalize Coverage Data option in IGV (see Methods). The ChromHMM annotation for each genomic region is shown below the plot using the same color code as in C. (B) Scatter plots showing the normalized read counts for all regions enriched in each modification (see Methods for more details) in mitosis versus interphase. It should be noted that ChIP-seq-based quantification may be less accurate than quantification based on mass spectrometry data due to inherent noise in the ChIP-seq method. Thus, in cases of a discrepancy, e.g., H3K4me1, we rely on the mass spectrometry results for quantification, and on the ChIP-seq results for localization. (C) Bar plots showing the percentage of reads in peak regions per ChromHMM category.
The only exception to our above observation is the ChIP-seq data for H3K4me1. The signal for this histone modification is moderately reduced in mitosis (Fig. 3B). However, since the proteomics analysis displays a high concordance between the levels of H3K4me1 in interphase and mitosis (even showing a slight increase in mitosis), the discrepancy might be related to the challenges in quantifying ChIP-seq signal due to the inherent noise in this method. We thus relied in this case on the MS data for assessing the levels of H3K4me1 and on the ChIP-seq data regarding the localization of this histone modification.
To quantify the genomic changes in the levels of histone modifications between mitosis and interphase, we used the ENCODE mappings of chromatin environments derived from the combined ChromHMM annotations (Ernst and Kellis 2012). We compared the organization of histone modifications within the chromatin environments and found that the distribution of the methylation signals in interphase and mitosis was almost identical. Conversely, the genomic distribution of the H3K27 and H3K9 acetylations differed slightly between these cell cycle phases. Both acetylations presented an increase in signal over enhancer regions (5% and 2%, respectively) and concurrent reduction in promoter regions (6% and 3%, respectively) in mitosis (Fig. 3C).
The nucleosome depleted regions become occupied during mitosis by a nucleosome distinguishable from its neighbors
Previous reports demonstrated that nucleosome depleted regions associated with the transcription start site (TSS) of promoters are lost during mitosis, and the gap is occupied by a nucleosome containing H3K4me3 and H2A.Z (Kelly et al. 2010; Liang et al. 2015). To better understand this mitotic change in the chromatin structure, we leveraged the variety of histone modifications we mapped in mitosis and interphase to study the histone modifications of the nucleosome that occupies the NDR during mitosis. Nucleosome depleted regions were also observed within enhancers and insulators (Henikoff 2008; Tsompana and Buck 2014). Together, the range of marks we profiled enabled the deep investigation of NDRs associated with promoters, putative enhancers, and insulators for the first time.
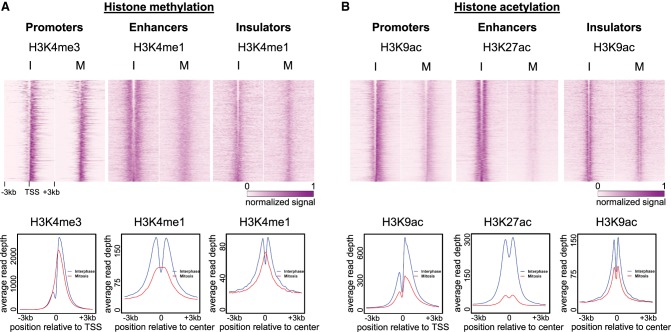
First, we focused on H3K4me3, which was previously shown to occupy promoter NDRs during mitosis (Liang et al. 2015). In line with Liang et al.’s report, the data sets we generated were mostly devoid of the promoter NDR structure in mitosis (Fig. 4A), even though our samples were synchronized using a different approach (STC vs. nocodazole). Additionally, extension of this analysis to include putative enhancers and insulators revealed that, in mitosis, a nucleosome containing methylated histones also occupies NDRs associated with these regulatory regions (Fig. 4A; Supplemental Figs. S5–S7).
Figure 4.
The nucleosome that enters the NDR during mitosis is distinguishable from the surrounding nucleosomes. Top: Heat maps showing the normalized and standardized read depth at 6-kb regions centered at either TSSs, enhancers, or insulators, for H3K4 methylation (n = 6488, 7062, 4367, respectively) (A), and H3K9 or H3K27 acetylation (n = 6808, 5565, 3033, respectively) (B). Bottom: Metagene plots showing the average read depth for all regions in each category are under the relevant heat map. Similar plots for additional modifications as well as a biological repeat are shown in Supplemental Figures S5–S7.
Next, we analyzed the organization of the histone acetylations H3K9ac and H3K27ac over NDRs. We observed that the binding structure was mostly preserved during mitosis as opposed to the histone methylation pattern, suggesting that the nucleosome that occupies the NDR is deacetylated (Fig. 4B; Supplemental Figs. S5–S7). To corroborate our observation, we analyzed data from G1E murine erythroblast cells (Hsiung et al. 2016). Consistently, in G1E murine cells the nucleosome that occupies the NDR is mostly deacetylated (Supplemental Fig. S8), indicating that this phenomenon is conserved between species and cell types. Altogether, these data suggest that the nucleosome that occupies NDRs during mitosis is methylated but not acetylated; hence, it is distinguishable from the neighboring nucleosomes which are dually marked by methylation and acetylation.
Classification of the NDRs according to nucleosome modifications during mitosis
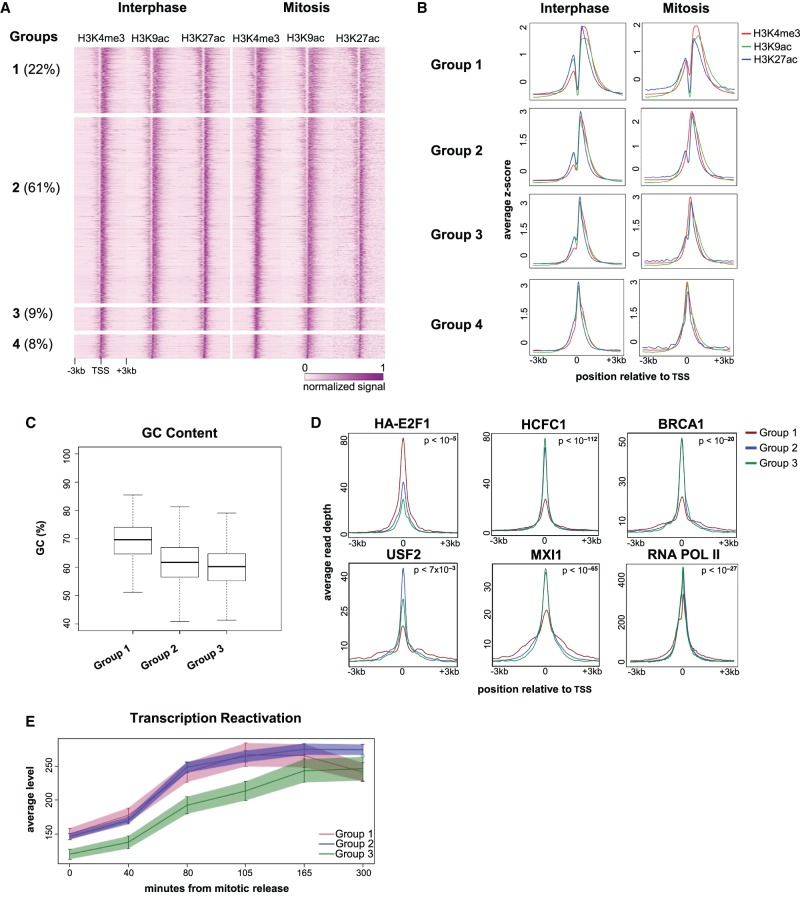
To further characterize the nucleosome that occupies TSS-associated NDRs during mitosis, we looked at the coverage profiles of genes marked with H3K4me3, H3K9ac, and H3K27ac at their TSSs and sorted the profiles by occupancy of mitotic H3K9ac at the TSS. The sorted profiles revealed four gene groups based on the combination of modifications occupying the NDR (Fig. 5A,B; Supplemental Fig. S9). The groups were classified as follows: Group 1, retention of the NDR during mitosis; Group 2, loss of the NDR during mitosis due to the entry of a nucleosome with solely methylated histones; Group 3, loss of the NDR during mitosis due to the entry of a nucleosome with both methylated and acetylated histones; Group 4, absence of the NDR in both interphase and mitosis. We disregarded Group 4 in further analyses since it already lacked the NDR structure in interphase. Of the promoters analyzed, we found that ∼22% belong to Group 1, 61% to Group 2, and 9% to Group 3.
Figure 5.
Characterization of the TSS-associated NDRs. (A) A heat map showing the normalized and standardized z-scores at 6-kb regions centered at the TSS of 5731 genes that are occupied by H3K4me3, H3K9ac, and H3K27ac. The regions were manually divided into four groups (Groups 1–4) after visual inspection; the percentage of regions in each group is shown in parentheses. (B) Metagene plots showing the average occupancy of each modification for the four groups. (C) Boxplot showing the difference in the GC content at the TSS for Groups 1–3. The difference between Group 1 and each of the other groups (Group 2 and Group 3) is highly significant (P-value = 1.67 × 10−133 and 3.97 × 10−84; t-test, and effect size = 0.88 for Group 2 and 1.12 for Group 3). (D) Metagene plots showing ENCODE ChIP-seq data and our RNA PolII data for Groups 1–3; P-values were obtained either by comparing Group 1 to the combination of Groups 2 and 3 (for HCFC1, BRCA1, MXI1, and RNA PolII), or by determining the maximal P-value from all pairwise comparisons (for HA-E2F1 and USF1). For a comprehensive list of all P-values, see Supplemental Table S2. The same analysis for all available ENCODE HeLa-S3 data is shown in Supplemental Figures S11, S12. (E) Profiles showing transcription reactivation kinetics following release from mitosis. The difference between Groups 1 and 2 versus Group 3 is significant (P-value = 0.00235; ANOVA. Note: This tests the change over time of the transcription levels and is therefore not sensitive to the difference in initial level [time = 0] between the groups).
To further assess these promoters, we first characterized the associated genes. We found that, of the genes in each of Groups 1–3, 88%–92% were shown to be transcribed in HeLa cells (Hart et al. 2013), 5% are cell cycle genes (as defined by Whitfield et al. 2002), 59%–61% are housekeeping genes, and 13%–15% are tissue-specific genes (as defined by Uhlen et al. 2015; Supplemental Table S1). Second, we compared the percentage of GC in each group and found that Group 1 has a higher GC content than Groups 2 and 3 (∼70% vs. 62% and 60% in Groups 2 and 3, respectively) (Fig. 5C). Thus, the GC content appears to distinguish between regions that retain the NDR during mitosis and regions that lose it, regardless of the histone modifications on the NDR-occupying nucleosome. As GC content may be associated with the binding of specific transcription factors (TFs), we next employed DREME, a discriminative motif discovery tool (Bailey 2011), to identify motifs enriched in Group 1 relative to the other groups. Indeed, along with the high GC content observed in Group 1, many of the motifs were enriched for GC (Supplemental Fig. S10).
To further investigate the characteristics of these three groups, we studied the binding patterns of TFs mapped by ENCODE in HeLa-S3 cells as well as our RNA PolII occupancy data from interphase cells. Similar to the GC content distinction, RNA PolII and most of the TFs (e.g., HCFC1, BRCA1, and MXI1) (Fig. 5D; Supplemental Fig. S11) demonstrated a binding pattern that differed between Group 1 and the two other groups (2 and 3), whereas only a limited number of factors (e.g., E2F1 and USF2) (Fig. 5D; Supplemental Fig. S11) show distinct binding patterns for each group. This observation, together with the GC content results, suggests that there is an intrinsic division between genes that have a nucleosome in the NDR during mitosis versus genes that do not.
Next, to better understand what distinguishes Group 2 from Group 3, we evaluated the association of histone deacetylase (HDAC) binding to each of these groups. Analysis of our and public HDAC2 and HDAC6 ChIP-seq data from unsynchronized cells (Wang et al. 2009; Ram et al. 2011) revealed a statistically significant difference between these groups: 43% of the TSSs in Group 2, whose NDR-occupying nucleosome is deacetylated, overlapped with an HDAC peak, while only 35% of the TSSs in Group 3, whose NDR-occupying nucleosome is acetylated, were occupied by an HDAC (P-value ∼ 0.001; χ2 test). Thus, it appears that the presence of HDAC may contribute to the observed difference in the acetylation patterns.
Finally, to identify the potential function of the differential NDR patterns, we analyzed gene reactivation kinetics following mitotic exit using a recently published data set (Palozola et al. 2017). We found that genes in Groups 1 and 2 are reactivated significantly earlier than the genes in Group 3 (P-value = 0.00235; ANOVA) (Fig. 5E). This finding suggests that the NDR patterns may differentiate between genes that are important promptly following mitotic exit versus genes that are only necessary later. In line with this notion, Gene Ontology analysis of the genes associated with the three groups revealed that Groups 1 and 2 are enriched for genes that may be relevant immediately upon mitotic exit, such as genes involved in cell signaling, localization, and RNA processing (Supplemental Fig. S13). These results suggest that the NDR pattern seen at the promoters of genes in Groups 1 and 2 may be important for the rapid reactivation of transcription following mitosis.
Mitosis-associated nucleosome deposition into the NDR
Nucleosome occupancy at NDRs during mitosis was previously explained as a consequence of the penetration of the +1 nucleosome into the NDR (Kelly et al. 2010). However, the absence of acetylation of this nucleosome challenges this model. We envisioned two possible models for the integration of the nucleosome into the NDR during mitosis, the first model being the incorporation into the NDR of a new nucleosome whose histones are methylated but not acetylated. The second model proposes, as postulated previously, that the surrounding nucleosomes shift into the NDR in mitosis, and the histones are actively deacetylated by HDACs.
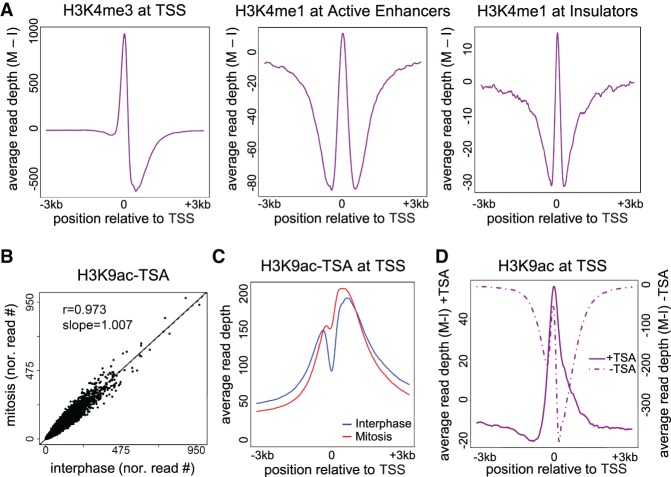
To discriminate between the two models, we first evaluated the dynamics of H3K4me3 in interphase and mitotic samples. H3K4me3 was shown both by a previous study and our analysis to occupy the NDR during mitosis. To evaluate nucleosome movement from our data, we measured the difference in the H3K4me3 signal in mitosis compared to interphase for promoter NDRs (delta metagene plots) (see Methods). We observed that the increase in nucleosome occupancy at the TSS is accompanied by a decrease in nucleosome occupancy in the downstream region, suggesting a shift of the +1 nucleosome into the NDR. This observation is in line with the findings of Kelly et al. (2010). Furthermore, we identified similar patterns of surrounding nucleosomes entering NDRs during mitosis at putative enhancers and insulators (Fig. 6A; Supplemental Fig. S14).
Figure 6.
Nucleosome shifting and deacetylation may explain NDR disappearance. (A) Delta metagene plots (see Methods for details) showing the average read depth of the indicated modifications after subtracting the interphase depths from the mitotic depths. (B) A scatter plot showing the normalized read counts at H3K9ac peaks on cells treated with TSA in mitosis versus interphase. (C) Metagene plot for H3K9ac data after TSA treatment. (D) Delta metagene plot for H3K9ac data before (dotted line) and after (solid line) TSA treatment. Similar results were obtained in a biological replicate (Supplemental Fig. S14).
Next, to further provide support for the second model, we treated interphase and mitotic cells with the global HDAC inhibitor TSA. Perturbation of HDAC activity eliminates the general mitotic hypoacetylation, and ChIP-seq performed on perturbed samples demonstrates highly similar H3K9ac signal intensities in interphase and mitosis (Fig. 6B; Supplemental Fig. S14). Additionally, inhibition of HDAC activity altered the histone acetylation pattern we observed previously at the NDRs, and in TSA-treated mitotic cells, the NDR-occupying nucleosome has acetylated histones (Fig. 6C; Supplemental Fig. S14). Delta metagene plots demonstrated that the increase in the level of acetylated histones at the NDR is accompanied by a decrease in the signal at the surrounding regions, which was indiscernible in untreated cells due to global mitotic deacetylation (Fig. 6D; Supplemental Fig. S14). This further supports the notion that the +1 nucleosome shifts into the NDR from a surrounding position. We observed a similar nucleosomal shift in enhancer and insulator regions (Supplemental Fig. S15). Finally, H3K4me3 patterns were not affected by TSA treatment, thus minimizing the possibility of a global effect (Supplemental Fig. S16).
Discussion
In this study, we aimed to gain a better understanding of the mitotic bookmarking process and the organization of chromatin during this phase. We designed an interdisciplinary approach that merges proteomics, immunofluorescence, automated ChIP-seq, and computational analysis that allowed the robust identification of changes between the interphase and mitotic chromatin organization. We incorporated the results of biological replicates for all measurements to obtain quantitative information regarding the global amounts of modified histones in each cell cycle phase. This was achieved by employing MS and immunofluorescence and by generating genome-wide chromatin maps for RNA PolII and six key histone modifications. These epigenomic landscapes provide insight into the interphase and mitosis spatial chromatin organization. To the best of our knowledge, we are the first to conduct a systematic molecular survey that combines global (proteomics and immunofluorescence) and spatial (epigenomics) measurements in interphase and mitosis.
We analyzed mass spectrometry as well as ChIP-seq data to study global changes in histone modifications during mitosis. MS data yield reliable quantitative results and therefore can be used to measure global changes in histone modification levels. The incorporation of MS data is particularly valuable when assessing histone acetylation levels, since mitosis-specific H3S10 and H3S28 phosphorylations may limit the functionality of H3K9ac and H3K27ac antibodies, thereby affecting the ChIP-seq results (Supplemental Fig. S17; Rothbart et al. 2015; Cornett et al. 2017). For this reason, we rely on the MS results to evaluate the mitotic levels of these acetylations, rather than on the ChIP-seq data. Additionally, ChIP-seq data contribute valuable spatial information, even though it may be influenced by the genomic distribution of mitotic phosphorylation. The integration of these varying approaches has allowed us to gain insight into both the quantification and localization of each tested histone modification.
All of our results (excluding the immunofluorescence experiments) are based on a prolonged mitotic arrest. Although this approach is widely used in the field (Kadauke et al. 2012; Hsiung et al. 2015, 2016; Palozola et al. 2017), we cannot rule out the possibility that some of our observations may be a consequence of this abnormal mitosis phase. It should be noted that despite the prolonged mitotic arrest, the majority of cells retained the ability to successfully re-enter the cell cycle (Supplemental Fig. S18). Further method development is needed in order to perform similar experiments on unperturbed mitotic cells.
Analyzing the proteomic and epigenomic data sets provided several major insights. We observed that although mitotic chromatin is highly condensed, global levels of histone methylation is mostly maintained. In contrast, histone acetylation abundance was reduced in mitotic samples as seen by measuring both global protein levels (via proteomics and immunofluorescence) and DNA association (via ChIP-seq). Yet, the change in acetylation levels between mitosis and interphase was moderate (about two- to sixfold) compared, for instance, to the variation observed in H3S10ph levels (about 17-fold) between these two phases (Fig. 2). Furthermore, comparison of the genomic localization between interphase and mitotic samples demonstrated that the organization of these modifications was tightly maintained. Altogether, apart from the patterns we observed in NDRs and discuss below, we did not identify local differences in the organization of the histone modification patterns between mitosis and interphase.
While previous studies assumed a global change in chromatin structure and mainly focused on the bookmarking effect of a limited number of transcription factors, our study suggests that the overall epigenomic structure in interphase appears to be vastly maintained during mitosis. This observation is in line with several recent studies, which indicate various levels of similarity between the mitotic and interphase chromatin structure. First, it was shown both in Drosophila and mice that a large portion of the transcription machinery, including many transcription factors and chromatin regulators, remain on the chromosomes during mitosis (Black et al. 2016; Teves et al. 2016; Liu et al. 2017b). Second, a genome-wide DNase-seq experiment revealed that genome accessibility is widely preserved and only locally modulated during mitosis (Hsiung et al. 2015). Finally, a recent study utilized 5-EU labeling to demonstrate that many genes exhibit transcription during mitosis (Palozola et al. 2017). Taken together, these, as well as our observations, suggest that despite their condensation, mitotic chromosomes retain a base level of their original chromatin structure and are partially involved in active transcription. Thus, the transcription that occurs during the G1 phase may be a mere continuation of the transcription that took place in the previous cell cycle and therefore may not require a specific bookmarking mechanism.
Recently, several studies employed ChIP-seq to assess histone modification levels during mitosis. There are, however, conflicting observations regarding the direction and uniformity of the changes in mitotic H3K27ac. While we observed a uniform decrease in H3K27ac levels in mitosis relative to interphase (Fig. 2; Supplemental Fig. S3B), a recent study in breast cancer MCF-7 cells saw a uniform increase in the levels of this modification in mitotic cells (Liu et al. 2017a). Other studies have demonstrated that the change in H3K27ac levels in mitosis is heterogeneous (Hsiung et al. 2016; Liu et al. 2017b). We cannot rule out the possibility that these variations are a consequence of technical differences (e.g., synchronization techniques) between the studies. However, the variations may also reflect a biological distinction between cancer or fully differentiated cells and cells that carry the potential for further differentiation. In line with this notion, mitosis constitutes a window of opportunity for altering cellular fates via mitotic chromatin reprogramming (Halley-Stott et al. 2014). Thus, H3K27ac, a modification linked to cell fate decisions (Long et al. 2016), may exhibit a heterogeneous change in profile in cells possessing the potency to differentiate (ESC and G1E erythroblasts) and a uniformly changing profile in cancer cells (HeLa-S3 and MCF-7).
Global deacetylation of histones during mitosis has been previously observed, and blocking it by HDAC inhibition causes severe mitotic defects (Cimini et al. 2003). Our results, which show a uniform genome-wide deacetylation of H3K9 and H3K27, strongly support a structural role of the deacetylation (Li et al. 2006) over a transcription-dependent mechanism (Noh et al. 2009). Indeed, blocking histone deacetylation by TSA strongly reduces the cells’ ability to exit mitosis following STC release (Supplemental Fig. S18).
Previous studies demonstrated that during mitosis there is a shift of the +1 nucleosome into the NDR that is associated with transcription start sites (Kelly et al. 2010; Liang et al. 2015). By leveraging the comprehensive data sets we generated, we were able to expand upon these initial observations and study a broad set of histone modifications, as well as the NDRs associated with enhancers and insulators. We observed that most of the nucleosomes that enter TSS-associated NDRs (∼61%) are devoid of histone acetylations (H3K9ac and H3K27ac). This observation may, in fact, reflect a specific phosphorylation of the nucleosome that enters the NDR, which can hinder the ability to detect these acetylations (Supplemental Fig. S17), rather than a deacetylation. Ruling out this alternative explanation requires further work, such as systematic ChIP-seq-based mapping of H3S10ph and H3S28ph during mitosis. Nevertheless, the significantly higher association of HDACs with Group 3 genes, as well as the effect of TSA on the acetylation levels of the NDR-associated nucleosome, support a deacetylation-based mechanism.
The transcription reactivation kinetics of genes with an acetylated NDR-associated nucleosome (Group 3) is slower than that of other genes (Groups 1 and 2) (Fig. 5E). A possible explanation for this finding is that the deacetylation serves as a means for bookmarking the region as an NDR in interphase and enables the removal of the nucleosome that enters this region immediately following mitosis. In line with this hypothesis, it has been shown that some chromatin remodeling complexes are affected by histone modifications (Swygert and Peterson 2014). Thus, the deacetylation may facilitate the removal of the nucleosome that enters the NDR by recruiting specific remodeling complexes after the mitosis phase. Further studies are needed to evaluate this hypothesis and investigate whether nucleosome repositioning is indeed important for the bookmarking process.
Not all NDRs undergo the same mitosis-associated changes. We found that in most cases there is a penetrating nucleosome that is deacetylated; however, there is also a group of promoters which retain their NDRs during mitosis (Group 1) (Fig. 5). Compared to the rest of the promoters, Group 1's promoters are already distinguishable during interphase in several ways: (1) These regions have a higher GC content; (2) the binding of many TFs is less localized to the TSS of these promoters (Fig. 5); and (3) the NDR seems to be wider (see, for example, the H3K4me3 plot in Supplemental Fig. S12). These findings suggest that basic differences in the promoter architecture reflect the mechanism that preserves the NDR during interphase. Only some of the mechanisms associated with these different promoter architectures are active during mitosis, and this may explain the differential loss of some NDRs. Thus, for instance, sequence-based mechanisms should be preserved during mitosis, whereas interphase NDRs that are kept via the binding of transcription factors (Struhl and Segal 2013) may be lost as cells undergo mitosis. Further work is needed to understand these mechanisms and their importance in mitotic function.
Overall, together with several recent studies, our work provides an important cornerstone necessary for shifting the paradigm that has lingered for many years. We can reason that, while structurally the chromatin goes through major changes during mitosis, molecular characteristics such as the organization of histone modifications are maintained, even if their levels may be reduced. This provides a simpler explanation for the restoration of chromatin structure following mitosis.
Methods
Cell culture and synchronization
HeLa-S3 cells were grown in Spinner flasks in Dulbecco's Modified Eagle's Medium (DMEM) supplemented with 10% fetal bovine serum (FBS), penicillin-streptomycin, L-glutamine, sodium pyruvate, and 0.1% Pluronic F-68. Cells were incubated at 37°C and 5% CO2. Cells were presynchronized in G1/S by addition of 2 mM thymidine for 16 h, washed with PBS, released for 3 h in fresh medium, and arrested with S-trityl-L-cysteine (STC) (164739, Sigma) for 12 h. HDAC blocking was done with 150 nM Trichostatin A (TSA, T1952, SIGMA) which was added either to the unsynchronized culture or 3 h after adding the STC, for 9 h.
Immunofluorescence and microscopy
Cells were fixed with either 70% ethanol or 4% paraformaldehyde, blocked with 5% BSA, and stained either with mouse anti-LaminA (1:1000; L1293, Sigma) followed by donkey anti-mouse DyLight 550 (1:400; ab98795, Abcam), or with rabbit anti-H3K9ac (1:1000; C5B11, Cell Signaling Technology) or H3K4me3 (1:1000; C42D8, Cell Signaling Technology) followed by Alexa Fluor 488 goat anti-rabbit IgG(H+L) (1:400; A11011, Invitrogen). Cells were counterstained with DAPI.
Proteomics
Three replicates of HeLa-S3 cells (either unsynchronized or STC synchronized) were harvested. Exactly 2.1 million cells from each sample were snap-frozen in liquid nitrogen and stored at −80°C. Global chromatin profiling (GCP) of cell pellets to quantify the levels of histone modification was performed as described previously (Creech et al. 2015). In brief, cells were lysed, nuclei were isolated, and histones were extracted by sulfuric acid and were then precipitated by trichloroacetic acid. Next, the samples were propionylated, desalted, and digested with trypsin overnight. Samples were subjected to a second propionylation, and then desalted using a C18 Sep-Pak Cartridge (Waters). Prior to MS analysis, a mix of isotopically labeled synthetic peptides was added to each sample. Next, the peptides were separated on a C18 column (15 cm × 75 µm ID, ReproSil-Pur 120 C18-AQ, 1.9 µm) using an EASY-nLC 1000 (Thermo Fisher Scientific) and analyzed using the GCP assay (a parallel reaction monitoring targeted LCMS method) using a Q Exactive-plus mass spectrometer (Thermo Fisher Scientific) as described previously (Creech et al. 2015).
The GCP employs high mass accuracy (∼3 parts per million)/high resolution (R = 17,500 at m/z 400) mass spectrometry; thus, it enables distinguishing between an adduct of trimethylation (42.0470 Da) from acetylation (42.0106 Da).
Standard operating procedures (SOPs) for the GCP assay, including synthetic peptide master mixture formulation, can be found at: https://panoramaweb.org/labkey/wiki/LINCS/Overview%20Information/page.view?name=sops.
Chromatin immunoprecipitation followed by sequencing (ChIP-seq)
ChIP-seq was performed either manually on individual samples or automatically using the Bravo liquid handling platform (Agilent model 16050-102, “Bravo”) as described previously (Busby et al. 2016). In accordance with our efforts to promote reproducibility, all antibodies used were monoclonal. The following antibodies were used: anti-RNA polymerase II 8WG16 (ab817, Abcam), anti-H3K4me1 (D1A9, Cell Signaling Technology), anti-H3K4me3 (C42D8, Cell Signaling Technology), anti-H3K9ac (C5B11, Cell Signaling Technology), anti-H3K27ac (D5E4, Cell Signaling Technology), anti-H3K27me3 (C36B11, Cell Signaling Technology), anti-H3K36me3 (D5A7, Cell Signaling Technology), and anti-CTCF G.758.4 (MA5-11187, Invitrogen). The specificity of all antibodies targeting histone modifications were assessed by us (Busby et al. 2016) and others (Rothbart et al. 2015), and the array data sets are publicly available at www.histoneantibodies.com.
Bioinformatic analysis
ChIP-seq reads were aligned to the human genome (hg19) using either BWA (Li and Durbin 2009) or Bowtie (Langmead and Salzberg 2012). Aligning to GRCh38 is expected to provide similar results, as only a small number of bases change genome-wide. The major difference between the releases is in centromere assembly (Guo et al. 2017), which were excluded from our analyses due to their repetitive nature. All regions listed in the ENCODE hg19 blacklist, as well as regions in genomic bins (2 kb) that had high coverage in our control file (top 0.2%), were excluded from all analyses. Duplicate alignments were removed with Picard Tools MarkDuplicates (http://broadinstitute.github.io/picard/). Only QC-passed reads were used for the analyses (see Supplemental Table S3 for read counts of all sequenced samples).
Peak detection on merged replicates (with a whole-cell extract data set as a control) was done using HOMER (Heinz et al. 2010) with default histone style parameters for all histone modifications, and default factor style parameters for RNA PolII and CTCF.
Scatter plots were made from read counts (calculated using BEDTools multicov [Quinlan and Hall 2010]) at merged interphase and mitosis peaks and normalized to 10 million total reads. ChromHMM (Chromatin State Discovery and Characterization) genomic annotations were downloaded from the UCSC Genome Browser for HeLa-S3 cells (Ernst and Kellis 2013) and combined into five main categories. The ChromHMM distribution plot (Fig. 3C) shows the percentage of reads in peak regions per ChromHMM annotation.
IGV snapshots show TDF files created from the aligned sequence data (using the count command from IGV tools [Robinson et al. 2011; Thorvaldsdottir et al. 2013]). The scale of each track was adjusted to the total number of reads using the Normalize Coverage Data option in IGV which multiplies each value by [1,000,000/(totalReadCount)] (Fig. 1C; Supplemental Fig. S3A).
For generating coverage profiles, metagene plots, and delta metagene plots, we first fixed the length of each read to the average of the mitosis and interphase fragment length modes. We analyzed the following NDRs: TSSs (using gene coordinates from UCSC [Karolchik et al. 2004]); enhancers (using regions with H3K4me1, H3K27ac, and UCSC EP300 HeLa-S3 peaks [wgEncodeEH001820]); and insulators (using merged interphase and mitosis CTCF ChIP-seq peaks), each containing a unique set of loci. In each set, only regions located >6 kb from each other and overlapping modification peaks within 500 bp on either side of the NDR, were included in the analyses.
Coverage profiles (heat maps) were created from the read depth (calculated using SAMtools bedcov [Li et al. 2009]) of each modification in a window of 6 kb around the NDR in 25-bp bins. Read depth was normalized to 10 million total reads, and the depths of each row were then standardized to the range [0, 1]. Heat maps are sorted by NDR occupancy in interphase.
Combined coverage profiles (Fig. 5A; Supplemental Fig. S9) only include regions with peaks for all of the modifications of interest overlapping the NDR. The normalized read depth was converted to a z-score for mitosis and interphase of each modification separately, and then each row was standardized (to [0, 1]). Heat maps are sorted by TSS occupancy of H3K9ac in mitosis. Visual inspection of the sorted profiles revealed gene groups based on the combination of modifications at the TSS. Metagene plots were produced by averaging the coverage profiles. Delta metagene plots were made by averaging the result of subtracting the interphase coverage profiles from the mitosis coverage profiles.
For the boxplots (Fig. 5C), GC content was measured around the TSS (±500 bp) of genes using BEDTools nuc, and corresponding P-values (t-test) and effect sizes were calculated.
HDAC2 and HDAC6 peaks for CD4 (Wang et al. 2009) and K562 cells (Ram et al. 2011) were downloaded and merged. The percentage of genes in each group that overlapped an HDAC peak at the TSS (1 bp) was determined and P-values were calculated using a χ2 test.
ChIP-seq data sets were downloaded from ENCODE (see Supplemental Table S2 for ENCODE file names) and metagene profiles of gene groups were created using deepTools plotProfile (Ramirez et al. 2016). Counts at the TSS (20 bp on either side) were totaled and corresponding P-values (t-test) were calculated.
Lists of HeLa-S3 transcribed genes (only genes with a zFPKM > −3) (Hart et al. 2013), cell cycle genes (Whitfield et al. 2002), and housekeeping/tissue-specific genes (www.proteinatlas.org) were downloaded and the percentage of genes in each group that overlapped each of these data sets was determined.
For the transcription reactivation kinetics analysis (Fig. 5E), data were downloaded (Palozola et al. 2017) and profiles of the mean for each gene group were plotted. Error bars and shading depict the standard error at each time point. The P-value was calculated using a Repeated Measures ANOVA performed on Groups 1 and 2 combined versus Group 3.
GO analysis was done with the GOrilla (Gene Ontology enRIchment anaLysis and visuaLizAtion) online tool (Eden et al. 2009).
Data access
The genomic data generated as part of this study have been submitted to the NCBI Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo) under accession number GSE108173. The original mass spectra have been deposited in the public proteomics repository MassIVE (Mass Spectrometry Interactive Virtual Environment; https://massive.ucsd.edu/) under accession number MSV000082818.
Supplementary Material
Acknowledgments
We thank Dr. Michele Busby and Dr. Melissa Gymrek for fruitful discussions; Yutong Qiu, Miguel Alcantar, and Michal Schlesinger for technical assistance; and Dr. Jason Ernst for assistance with HeLa-S3 ChromHMM data. This research was supported by the Israel Science Foundation (grant no. 184/16 to I.S.), the Israel Science Foundation – Broad Institute Joint Program (grant no. 1972/15 to I.S. and A.G.), the United States–Israel Binational Science Foundation (BSF), Jerusalem, Israel (grant no. 2015272 to I.S. and A.G.), and National Institutes of Health grants (U54 HG008097 to J.D.J. and MIRA R35GM124736 to S.B.R.).
Footnotes
[Supplemental material is available for this article.]
Article published online before print. Article, supplemental material, and publication date are at http://www.genome.org/cgi/doi/10.1101/gr.230300.117.
References
- Bailey TL. 2011. DREME: motif discovery in transcription factor ChIP-seq data. Bioinformatics 27: 1653–1659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bar-Joseph Z, Siegfried Z, Brandeis M, Brors B, Lu Y, Eils R, Dynlacht BD, Simon I. 2008. Genome-wide transcriptional analysis of the human cell cycle identifies genes differentially regulated in normal and cancer cells. Proc Natl Acad Sci 105: 955–960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belmont AS, Bruce K. 1994. Visualization of G1 chromosomes: a folded, twisted, supercoiled chromonema model of interphase chromatid structure. J Cell Biol 127: 287–302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Black LK, Petruk S, Fenstermaker TK, Hodgson JW, Caplan JL, Brock HW, Mazo A. 2016. Chromatin proteins and RNA are associated with DNA during all phases of mitosis. Cell Discov 2: 16038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Busby M, Xue C, Li C, Farjoun Y, Gienger E, Yofe I, Gladden A, Epstein CB, Cornett EM, Rothbart SB, et al. 2016. Systematic comparison of monoclonal versus polyclonal antibodies for mapping histone modifications by ChIP-seq. Epigenetics Chromatin 9: 49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caravaca JM, Donahue G, Becker JS, He X, Vinson C, Zaret KS. 2013. Bookmarking by specific and nonspecific binding of FoxA1 pioneer factor to mitotic chromosomes. Genes Dev 27: 251–260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen D, Dundr M, Wang C, Leung A, Lamond A, Misteli T, Huang S. 2005. Condensed mitotic chromatin is accessible to transcription factors and chromatin structural proteins. J Cell Biol 168: 41–54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cho RJ, Huang M, Campbell MJ, Dong H, Steinmetz L, Sapinoso L, Hampton G, Elledge SJ, Davis RW, Lockhart DJ. 2001. Transcriptional regulation and function during the human cell cycle. Nat Genet 27: 48–54. [DOI] [PubMed] [Google Scholar]
- Cimini D, Mattiuzzo M, Torosantucci L, Degrassi F. 2003. Histone hyperacetylation in mitosis prevents sister chromatid separation and produces chromosome segregation defects. Mol Biol Cell 14: 3821–3833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cornett EM, Dickson BM, Rothbart SB. 2017. Analysis of histone antibody specificity with peptide microarrays. J Vis Exp 10.3791/55912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Creech AL, Taylor JE, Maier VK, Wu X, Feeney CM, Udeshi ND, Peach SE, Boehm JS, Lee JT, Carr SA, et al. 2015. Building the Connectivity Map of epigenetics: chromatin profiling by quantitative targeted mass spectrometry. Methods 72: 57–64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dileep V, Ay F, Sima J, Vera DL, Noble WS, Gilbert DM. 2015. Topologically associating domains and their long-range contacts are established during early G1 coincident with the establishment of the replication-timing program. Genome Res 25: 1104–1113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eden E, Navon R, Steinfeld I, Lipson D, Yakhini Z. 2009. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics 10: 48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ernst J, Kellis M. 2012. ChromHMM: automating chromatin-state discovery and characterization. Nat Methods 9: 215–216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ernst J, Kellis M. 2013. Interplay between chromatin state, regulator binding, and regulatory motifs in six human cell types. Genome Res 23: 1142–1154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fu Y, Sinha M, Peterson CL, Weng Z. 2008. The insulator binding protein CTCF positions 20 nucleosomes around its binding sites across the human genome. PLoS Genet 4: e1000138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gazit B, Cedar H, Lerer I, Voss R. 1982. Active genes are sensitive to deoxyribonuclease I during metaphase. Science 217: 648–650. [DOI] [PubMed] [Google Scholar]
- Gottesfeld JM, Forbes DJ. 1997. Mitotic repression of the transcriptional machinery. Trends Biochem Sci 22: 197–202. [DOI] [PubMed] [Google Scholar]
- Guo Y, Dai Y, Yu H, Zhao S, Samuels DC, Shyr Y. 2017. Improvements and impacts of GRCh38 human reference on high throughput sequencing data analysis. Genomics 109: 83–90. [DOI] [PubMed] [Google Scholar]
- Halley-Stott RP, Jullien J, Pasque V, Gurdon J. 2014. Mitosis gives a brief window of opportunity for a change in gene transcription. PLoS Biol 12: e1001914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hart T, Komori HK, LaMere S, Podshivalova K, Salomon DR. 2013. Finding the active genes in deep RNA-seq gene expression studies. BMC Genomics 14: 778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, Cheng JX, Murre C, Singh H, Glass CK. 2010. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell 38: 576–589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hendzel MJ, Wei Y, Mancini MA, Van Hooser A, Ranalli T, Brinkley BR, Bazett-Jones DP, Allis CD. 1997. Mitosis-specific phosphorylation of histone H3 initiates primarily within pericentromeric heterochromatin during G2 and spreads in an ordered fashion coincident with mitotic chromosome condensation. Chromosoma 106: 348–360. [DOI] [PubMed] [Google Scholar]
- Henikoff S. 2008. Nucleosome destabilization in the epigenetic regulation of gene expression. Nat Rev Genet 9: 15–26. [DOI] [PubMed] [Google Scholar]
- Hsiung CC, Morrissey CS, Udugama M, Frank CL, Keller CA, Baek S, Giardine B, Crawford GE, Sung MH, Hardison RC, et al. 2015. Genome accessibility is widely preserved and locally modulated during mitosis. Genome Res 25: 213–225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsiung CC, Bartman CR, Huang P, Ginart P, Stonestrom AJ, Keller CA, Face C, Jahn KS, Evans P, Sankaranarayanan L, et al. 2016. A hyperactive transcriptional state marks genome reactivation at the mitosis–G1 transition. Genes Dev 30: 1423–1439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jaffe JD, Wang Y, Chan HM, Zhang J, Huether R, Kryukov GV, Bhang HE, Taylor JE, Hu M, Englund NP, et al. 2013. Global chromatin profiling reveals NSD2 mutations in pediatric acute lymphoblastic leukemia. Nat Genet 45: 1386–1391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johansen KM, Johansen J. 2006. Regulation of chromatin structure by histone H3S10 phosphorylation. Chromosome Res 14: 393–404. [DOI] [PubMed] [Google Scholar]
- Kadauke S, Blobel GA. 2013. Mitotic bookmarking by transcription factors. Epigenetics Chromatin 6: 6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kadauke S, Udugama MI, Pawlicki JM, Achtman JC, Jain DP, Cheng Y, Hardison RC, Blobel GA. 2012. Tissue-specific mitotic bookmarking by hematopoietic transcription factor GATA1. Cell 150: 725–737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karolchik D, Hinrichs AS, Furey TS, Roskin KM, Sugnet CW, Haussler D, Kent WJ. 2004. The UCSC Table Browser data retrieval tool. Nucleic Acids Res 32: D493–D496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kelly TK, Miranda TB, Liang G, Berman BP, Lin JC, Tanay A, Jones PA. 2010. H2A.Z maintenance during mitosis reveals nucleosome shifting on mitotically silenced genes. Mol Cell 39: 901–911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- King DW, Barnhisel ML. 1967. Synthesis of RNA in mammalian cells during mitosis and interphase. J Cell Biol 33: 265–272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kouskouti A, Talianidis I. 2005. Histone modifications defining active genes persist after transcriptional and mitotic inactivation. EMBO J 24: 347–357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langmead B, Salzberg SL. 2012. Fast gapped-read alignment with Bowtie 2. Nat Methods 9: 357–359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee CK, Shibata Y, Rao B, Strahl BD, Lieb JD. 2004. Evidence for nucleosome depletion at active regulatory regions genome-wide. Nat Genet 36: 900–905. [DOI] [PubMed] [Google Scholar]
- Li H, Durbin R. 2009. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25: 1754–1760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y, Kao GD, Garcia BA, Shabanowitz J, Hunt DF, Qin J, Phelan C, Lazar MA. 2006. A novel histone deacetylase pathway regulates mitosis by modulating Aurora B kinase activity. Genes Dev 20: 2566–2579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R, Genome Project Data Processing Subgroup. 2009. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25: 2078–2079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li M, Sun Q, Wang X. 2017. Transcriptional landscape of human cancers. Oncotarget 8: 34534–34551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liang K, Woodfin AR, Slaughter BD, Unruh JR, Box AC, Rickels RA, Gao X, Haug JS, Jaspersen SL, Shilatifard A. 2015. Mitotic transcriptional activation: clearance of actively engaged Pol II via transcriptional elongation control in mitosis. Mol Cell 60: 435–445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y, Chen S, Wang S, Soares F, Fischer M, Meng F, Du Z, Lin C, Meyer C, DeCaprio JA, et al. 2017a. Transcriptional landscape of the human cell cycle. Proc Natl Acad Sci 114: 3473–3478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y, Pelham-Webb B, Di Giammartino DC, Li J, Kim D, Kita K, Saiz N, Garg V, Doane A, Giannakakou P, et al. 2017b. Widespread mitotic bookmarking by histone marks and transcription factors in pluripotent stem cells. Cell Rep 19: 1283–1293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Long HK, Prescott SL, Wysocka J. 2016. Ever-changing landscapes: transcriptional enhancers in development and evolution. Cell 167: 1170–1187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martinez-Balbas MA, Dey A, Rabindran SK, Ozato K, Wu C. 1995. Displacement of sequence-specific transcription factors from mitotic chromatin. Cell 83: 29–38. [DOI] [PubMed] [Google Scholar]
- Naumova N, Imakaev M, Fudenberg G, Zhan Y, Lajoie BR, Mirny LA, Dekker J. 2013. Organization of the mitotic chromosome. Science 342: 948–953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Noh EJ, Lim DS, Jeong G, Lee JS. 2009. An HDAC inhibitor, trichostatin A, induces a delay at G2/M transition, slippage of spindle checkpoint, and cell death in a transcription-dependent manner. Biochem Biophys Res Commun 378: 326–331. [DOI] [PubMed] [Google Scholar]
- Palozola KC, Donahue G, Liu H, Grant GR, Becker JS, Cote A, Yu H, Raj A, Zaret KS. 2017. Mitotic transcription and waves of gene reactivation during mitotic exit. Science 358: 119–122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parsons GG, Spencer CA. 1997. Mitotic repression of RNA polymerase II transcription is accompanied by release of transcription elongation complexes. Mol Cell Biol 17: 5791–5802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prasanth KV, Sacco-Bubulya PA, Prasanth SG, Spector DL. 2003. Sequential entry of components of the gene expression machinery into daughter nuclei. Mol Biol Cell 14: 1043–1057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prescott DM, Bender MA. 1962. Synthesis of RNA and protein during mitosis in mammalian tissue culture cells. Exp Cell Res 26: 260–268. [DOI] [PubMed] [Google Scholar]
- Quinlan AR, Hall IM. 2010. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26: 841–842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ram O, Goren A, Amit I, Shoresh N, Yosef N, Ernst J, Kellis M, Gymrek M, Issner R, Coyne M, et al. 2011. Combinatorial patterning of chromatin regulators uncovered by genome-wide location analysis in human cells. Cell 147: 1628–1639. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramirez F, Ryan DP, Gruning B, Bhardwaj V, Kilpert F, Richter AS, Heyne S, Dundar F, Manke T. 2016. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res 44: W160–W165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raynaud C, Mallory AC, Latrasse D, Jegu T, Bruggeman Q, Delarue M, Bergounioux C, Benhamed M. 2014. Chromatin meets the cell cycle. J Exp Bot 65: 2677–2689. [DOI] [PubMed] [Google Scholar]
- Robinson JT, Thorvaldsdottir H, Winckler W, Guttman M, Lander ES, Getz G, Mesirov JP. 2011. Integrative genomics viewer. Nat Biotechnol 29: 24–26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothbart SB, Dickson BM, Raab JR, Grzybowski AT, Krajewski K, Guo AH, Shanle EK, Josefowicz SZ, Fuchs SM, Allis CD, et al. 2015. An interactive database for the assessment of histone antibody specificity. Mol Cell 59: 502–511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skoufias DA, DeBonis S, Saoudi Y, Lebeau L, Crevel I, Cross R, Wade RH, Hackney D, Kozielski F. 2006. S-trityl-L-cysteine is a reversible, tight binding inhibitor of the human kinesin Eg5 that specifically blocks mitotic progression. J Biol Chem 281: 17559–17569. [DOI] [PubMed] [Google Scholar]
- Spellman PT, Sherlock G, Zhang MQ, Iyer VR, Anders K, Eisen MB, Brown PO, Botstein D, Futcher B. 1998. Comprehensive identification of cell cycle–regulated genes of the yeast Saccharomyces cerevisiae by microarray hybridization. Mol Biol Cell 9: 3273–3297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stevens TJ, Lando D, Basu S, Atkinson LP, Cao Y, Lee SF, Leeb M, Wohlfahrt KJ, Boucher W, O'Shaughnessy-Kirwan A, et al. 2017. 3D structures of individual mammalian genomes studied by single-cell Hi-C. Nature 544: 59–64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Struhl K, Segal E. 2013. Determinants of nucleosome positioning. Nat Struct Mol Biol 20: 267–273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swygert SG, Peterson CL. 2014. Chromatin dynamics: interplay between remodeling enzymes and histone modifications. Biochim Biophys Acta 1839: 728–736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Teves SS, An L, Hansen AS, Xie L, Darzacq X, Tjian R. 2016. A dynamic mode of mitotic bookmarking by transcription factors. eLife 5: e22280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thorvaldsdottir H, Robinson JT, Mesirov JP. 2013. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 14: 178–192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsompana M, Buck MJ. 2014. Chromatin accessibility: a window into the genome. Epigenetics Chromatin 7: 33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uhlen M, Fagerberg L, Hallstrom BM, Lindskog C, Oksvold P, Mardinoglu A, Sivertsson A, Kampf C, Sjostedt E, Asplund A, et al. 2015. Proteomics. Tissue-based map of the human proteome. Science 347: 1260419. [DOI] [PubMed] [Google Scholar]
- Vagnarelli P. 2013. Chromatin reorganization through mitosis. Adv Protein Chem Struct Biol 90: 179–224. [DOI] [PubMed] [Google Scholar]
- Valls E, Sanchez-Molina S, Martinez-Balbas MA. 2005. Role of histone modifications in marking and activating genes through mitosis. J Biol Chem 280: 42592–42600. [DOI] [PubMed] [Google Scholar]
- Van Hooser A, Goodrich DW, Allis CD, Brinkley BR, Mancini MA. 1998. Histone H3 phosphorylation is required for the initiation, but not maintenance, of mammalian chromosome condensation. J Cell Sci 111: 3497–3506. [DOI] [PubMed] [Google Scholar]
- Wang F, Higgins JM. 2013. Histone modifications and mitosis: countermarks, landmarks, and bookmarks. Trends Cell Biol 23: 175–184. [DOI] [PubMed] [Google Scholar]
- Wang Z, Zang C, Cui K, Schones DE, Barski A, Peng W, Zhao K. 2009. Genome-wide mapping of HATs and HDACs reveals distinct functions in active and inactive genes. Cell 138: 1019–1031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whitfield ML, Sherlock G, Saldanha AJ, Murray JI, Ball CA, Alexander KE, Matese JC, Perou CM, Hurt MM, Brown PO, et al. 2002. Identification of genes periodically expressed in the human cell cycle and their expression in tumors. Mol Biol Cell 13: 1977–2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaidi SK, Grandy RA, Lopez-Camacho C, Montecino M, van Wijnen AJ, Lian JB, Stein JL, Stein GS. 2014. Bookmarking target genes in mitosis: a shared epigenetic trait of phenotypic transcription factors and oncogenes? Cancer Res 74: 420–425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao R, Nakamura T, Fu Y, Lazar Z, Spector DL. 2011. Gene bookmarking accelerates the kinetics of post-mitotic transcriptional re-activation. Nat Cell Biol 13: 1295–1304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou VW, Goren A, Bernstein BE. 2011. Charting histone modifications and the functional organization of mammalian genomes. Nat Rev Genet 12: 7–18. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.