Abstract
Cell–cell adhesions maintain the mechanical integrity of multicellular tissues and have recently been found to act as mechanotransducers, translating mechanical cues into biochemical signals. Mechanotransduction studies have primarily focused on focal adhesions, sites of cell-substrate attachment. These studies leverage technical advances in devices and systems interfacing with living cells through cell–extracellular matrix adhesions. As reports of aberrant signal transduction originating from mutations in cell–cell adhesion molecules are being increasingly associated with disease states, growing attention is being paid to this intercellular signaling hub. Along with this renewed focus, new requirements arise for the interrogation and stimulation of cell–cell adhesive junctions. This review covers established experimental techniques for stimulation and interrogation of cell–cell adhesion from cell pairs to monolayers.
Keywords: Mechanobiology, Cell–cell adhesion, Cell mechanics, BioMEMS, FRET
1. Introduction
Our bodies experience a wide variety of mechanical inputs on a continuous basis. Whether from opening a door, noticing a tap on the shoulder, or perceiving the acceleration of an elevator, our brains translate physical forces into information that we use to understand and interact with our environment. In a similar way, cells within our body’s tissues generate, sense, and respond to mechanical cues within their local environment to direct normal physiological processes, and when things go awry, aberrant mechanical signaling can lead to the development and progression of disease.
In the body, physical forces are generated by and transmitted through the musculoskeletal system comprising bones and muscles as well as the structures that hold them together: joints, ligaments and tendons. In cells, this function is performed by cytoskeletal filaments and their associated adhesive organelles [1]. Cell–cell and cell–extracellular matrix (ECM) adhesive organelles are macromolecular structures that integrate individual cells into complex tissues by anchoring cytoskeletal components [2]. They have been the subject of intensive scientific investigation due to their importance in cell and tissue mechanics, bridging the gap between interactions at the molecular level to forces at the cellular scale. In recent years, attention has shifted toward understanding the integral roles cell–cell adhesions play in transducing mechanical cues into biochemical signals, a process often referred to as mechanotransduction [3]. Under normal homeostatic conditions, the adhesion/cytoskeleton systems form a highly integrated network that regulates tissue morphogenesis, collective cell migration, cell proliferation, and differentiation [4]. However, a number of pathological conditions and developmental disorders, including skin disorders [5], arthritis [6], atherosclerosis [7], and cancer [8], may emerge from aberrant cell–cell adhesion or mechanical cues. Hence, the study of cell–cell adhesion has taken center stage aiming to understand its capacity in maintaining tissue- and cell-level mechanical integrity and more importantly, in converting forces and stresses at the cell–cell junction into regulatory pathways that dictate cell behavior.
Over the past two decades, we have witnessed a wide variety of techniques to stimulate and probe cells in vitro. With the advancement and wide adoption of these techniques, we have established a large body of literature about how cells move, differentiate, connect with other cells and probe their environment. The techniques were not invented for the sole purpose of studying cell–cell adhesion; rather they were designed to probe mechanical responses and/or to stimulate biochemical responses in cells interacting physically in their suspended or adherent states. The consequence was that investigators gained many insights about how adhesive organelles give rise to cell and tissue level architecture. The majority of the techniques were designed to stimulate and probe cell–ECM interactions, which serve at the forefront of the physical interaction between cells and their external environment. During the course of such studies, we have learned that cell–cell interactions work together with and even regulate cell–ECM adhesions. Some of the probing and stimulation techniques require the presence of robust cell–cell junctions. Here, we present techniques that have been widely used to explore cell mechanics, and then how they can be applied for use in cell–cell adhesion studies.
2. Cell–cell adhesion complexes
There are four main types of specialized cell–cell junctions in mammalian cells. These include tight junctions, gap junctions, adherens junctions, and desmosomes [9,10]. Tight junctions seal the paracellular space, limiting the passage of molecules and ions through the space between cells, and stopping the movement of membrane proteins between the upper and lower portions of the cell [11]. Gap junctions function as pores between adjoining cells, allowing molecules, ions, and electrical currents to pass directly between cells [9]. This review will focus on adherens junctions and desmosomes, which are cadherin-based intercellular junctions that link to the actin and intermediate filament (IF) cytoskeletons, respectively (Fig. 1a).
Fig. 1.
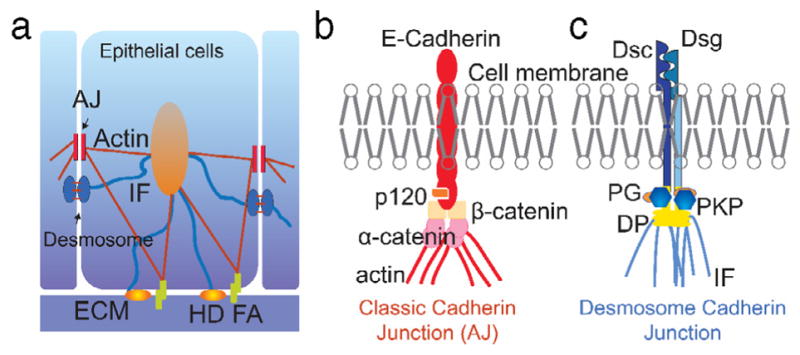
Cell–cell adhesion in epithelial cells. a. Adherens junctions (AJs) and desmosomes are cadherin-based intercellular junctions, which, along with adhesions at the cell–ECM (HD: hemidesmosome; FA: focal adhesion), are responsible for maintenance of the epithelial phenotype. b. The major components of the desmosome junction are desmocollin (Dsc), desmoglein (Dsg), plakoglobin (PG), plakophilin (PKP), and desmoplakin (DP), which connect to intermediate filaments (IFs). c. The major components of classical AJs are the transmembrane protein E-cadherin, p120, α-, and β-catenin.
2.1. Adherens junctions
Adherens junctions (AJs) are multiprotein complexes whose core components comprise transmembrane classical cadherins and intracellular armadillo family members (Fig. 1b). The extra-cellular domains of the classical cadherins, including E-cadherin, N-cadherin, VE-cadherin, and P-cadherin, form both hetero- and homophilic interactions and through calcium-dependent trans interactions link neighboring cells together [12]. On the intracellular side, they provide a platform for the recruitment of armadillo proteins p120 catenin and β-catenin. α-catenin interacts with β-catenin and provides the linkage to the actin cytoskeleton. Many of the classical cadherins have been shown to be mechanically sensitive [13], some capable of forming catch-bonds that strengthen and become longer lived in the presence of mechanical force [14]. Moreover, both extracellular and intracellular mechanical stimuli can promote force-mediated stabilization of the AJ/actin linkage through recruitment of the actin binding protein vinculin, which occurs through a conformational change in α-catenin that reveals a vinculin binding site [15].
2.2. Desmosomes
Desmosomes are also calcium-dependent adhesive junctions and have a molecular organization similar to that of the AJ (Fig. 1c) [16]. Desmosomal cadherins, which in humans include desmogleins (Dsg1–4) and desmocollins (Dsc1–3), are expressed in a tissue-type and differentiation-dependent manner [17,18]. They can form hetero- and/or homophilic trans interactions between their extracellular domains, though compared with classical cadherins these interactions are not as well understood [16,19]. The intracellular portions of desmosomal cadherins recruit the armadillo proteins plakoglobin and plakophilins. These interact with the plakin protein family member desmoplakin (DP), which in turn links the desmosomal core to the IF cytoskeleton [16]. The roles of the desmosome/IF network in mechanical signaling are not well understood. However, removing the keratin IF system was shown to affect the mechanical properties of cells as well as their ability to migrate [20]. Moreover, manipulating the strength of the desmosome/IF linkage using DP mutants regulates both intercellular forces in cell pairs and cell stiffness in cell pairs and larger groups of cells, through a process involving the actin cytoskeleton [21].
3. Techniques to study cell–cell adhesion
A wide variety of techniques have been used to study the mechanics of living cells. Early studies were done on groups of cells, using techniques such as substrate deformation, in which cells are cultured on a deformable substrate, and then subjected to a uniform monotonic or cyclic strain. To recapitulate the effects of fluid flow on cell layers, flow-induced shear experiments have also been performed, in which a fluid flowing over a cell layer imposes shear stress on the cells. To investigate cell-substrate and cell–cell interactions, traction force microscopy and micropost substrates were developed to enable the measurement of forces exerted by cells while stationary or in motion. In many instances, the mechanical properties of individual cells or cell pairs is of interest because they provide insight into how cells physically adapt to extracellular stimuli and their microenvironment. In such cases, methods with single cell resolution are employed. They include atomic force microscopy (AFM), optical traps/optical tweezers, magnetic beads, micropipette aspiration, and laser ablation. Another method to probe cell mechanical responses employs micro-electro mechanical systems (MEMS). These devices take advantage of actuators and sensors, with tunable displacement and force measurement resolution, by either directly attaching to the cell or moving a structure that the cell is attached to (resolutions for force and displacement are listed in Table 1). Some examples of microfabricated devices include uniaxial and biaxial pullers, micropillars, and cantilever beams.
Table 1.
Techniques to study cell–cell adhesion with resolutions in force and displacements.
| Technique | Force/Stress range (Resolution) | Displacement range (Resolution) |
|---|---|---|
| Deformable Substrates | 50–1000 Pa [22] | 0–100 μm [22] 0%–70% [23] (0.04%) |
| Fluid Flow | 0–22 Pa [24,25](0.05 Pa) | 0–50% [26] |
| Micropipette Aspiration | 0–700 Pa (0.1 Pa) [27,28] | 0–100 μm (25 nm) [29] |
| Optical Traps/stretcher | 0–300 pN (5 pN) [30,31] | 0–5 μm (1 nm) [32] |
| Magnetic Beads | 0–120 pN (1 pN) [33] | N/A |
| Atomic Force Microscopy | 0–20 nN (1 pN) [34] | 0–100 μm (1–5 nm) [34] |
| Uniaxial Puller | 0–1.5 μN (1 nN) [35] | 0–100 μm (40 nm) [35] |
| Biaxial Puller | 0–60 μN (1 nN) [36] | 0–3.4 μm (10 nm) [36] |
| Microposts | 0–100 nN (10 pN) [37] | 0–1000 nm (10 nm) [37] |
| Cantilever Beam Deflection | 0–1 μN (50 pN) [38] | 0–50 μm (10 nm) [38] |
3.1. Cell monolayer studies
3.1.1. Deformable substrates
By culturing cells on the surface of a deformable substrate, strains can easily be imposed on the cells by manipulating the substrate. Uniaxial and biaxial strains can be achieved, depending on how the substrate is deformed. There have been many studies on a variety of cell types, including bone tissue [22,39,40], lung cells [41,42], and neurons [43]. In addition, this method has been used to study cyclic loading on cell groups [23,44,45]. The mechanical properties of the substrate can be controlled by altering its thickness or its chemical composition [46], which in turn will affect the stiffness of the substrate, allowing for finer control over strain levels imposed on cells. Overall, this technique is simple to implement, but it does not offer high displacement resolution compared to other techniques.
Substrate deformation can be achieved by two modalities. The first is by stretching the substrate longitudinally. This method has been demonstrated to impose uniform longitudinal stresses on cells on the substrate uniaxially (Fig. 2a) or biaxially (Fig. 2b) and can also be used to impose oscillating stresses on cells [47,48]. The other method is substrate flection, in which the substrate is bent, usually using four-point bending [49] (Fig. 2c). This method offers the ability to achieve low strains, and has also been demonstrated to impose uniform longitudinal stresses on cells. Many studies have also used circular substrates with out-of-plane and in-plane deformation [50]. An issue with out-of-plane deformation is the heterogeneity of the strain imposed on the substrate. The radial component of strain is usually uniform, but the circumferential component varies from zero at the ends to some maximum in the center. In-plane deformation has been used to avoid problems of heterogeneous strain distributions that result from out-of-plane deformation. One in-plane deformation method is stretching the substrate around a circular ring (not shown, similar to Fig. 2e). This method is similar to the out-of-plane technique of stretching the substrate around a curved template (Fig. 2d), but with this method, the portion of the substrate inside the ring remains on a single plane, resulting in uniform strain distributions [51–53]. Similarly, a vacuum can be used to stretch a circular substrate around the perimeter of a raised central cylindrical platform (Fig. 2e). The portion of the substrate on the platform remains on a single plane, resulting in uniform strain distributions. This method has found wide success in commercialized systems such as Flexcell.
Fig. 2.
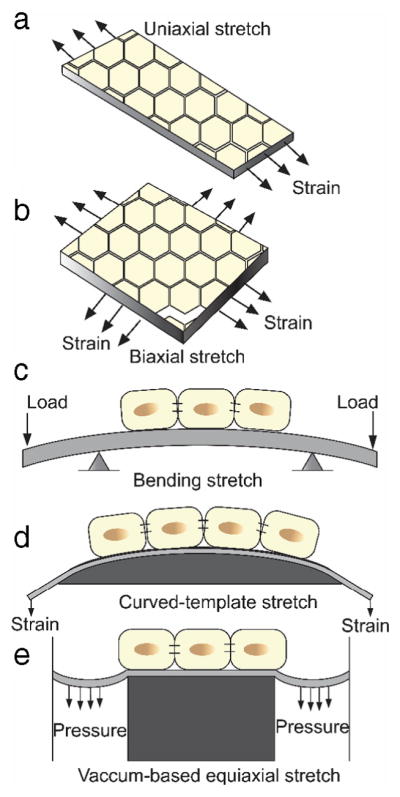
Mechanical stretching of a monolayer of cells. a. Uniaxial stretching; b: Biaxial stretching; c: Substrate flection; d: Stretching with curved template; e: Equiaxial stretching with vacuum suction.
Early attempts to stimulate a group of cells with mechanical strain aimed to understand their change in material properties in response to the stimulation [41,42], while underlying mechanisms of the biochemical and subsequently the biophysical responses are limited. This is mainly due to the nature of the technique but also the lack of tools to follow cellular responses at a small scale. In recent years, cyclic stretching with equiaxial strain generated most of the significant findings in how different cell types respond to oscillating strain. Commercialized instrumentation makes it a readily available technique in many biological labs; the capacity to work within the physiological cell culture condition propels its wide acceptance. The effects of cyclic stretching on the reorganization of cytoskeleton, the proliferation of different types of cells and the differentiation of stem cells have all been reported [54–57].
Stretch can be used to study mechanotransduction, as the process, in general, involves force-induced conformational changes in a mechanosensor protein that trigger additional protein–protein interactions. It is widely accepted that applied strains from stretching unfolds adhesion molecules in focal adhesion (FA) sites. For example, stretching induces conformational changes in the FA protein talin, which activates vinculin recruitment and ultimately leads to actin filament clustering and FA enhancement in a force-dependent manner [58]. Therefore, stretch can be used as a tool to examine force-induced alterations in mechanosensitive protein localization and recruitment, and when combined with molecular tension sensors (discussed below) can be used to observe force-induced protein conformational changes. These alterations in the FAs lead to other intracellular biochemical signals which result in a host of cellular behaviors.
Recent studies have found that the stretching stimulation may not only strain mechanosensors at the FA but also at cell–cell adhesive junctions [59,60]. Beneath the complex mechanotransduction signaling network lies the coordinated crosstalk of integrins and cadherins that regulates cell signals and forces [61,62]. The force transmission can potentially be facilitated by the AJ protein α-catenin, which reinforces intercellular tension by tightening the linkage between AJs and F-actin when the cell–cell junction experiences strain that exposes the vinculin binding site in α-catenin [63]. It has also been reported that intercellular tension is increased when adherent cells are stretched, as measured by a Förster resonance energy transfer (FRET)-based tension sensor embedded in E-cadherin [59], pointing towards a direct impact of stretch stimulation at cell–cell junctions. Further, transcriptional activities of YAP and β-catenin upon mechanical strain function in a E-cadherin dependent manner [64]. Besides mechanosensor molecules at junctions, mechanical stretch also stimulates mechanosensitive ion channels, such as piezo1 [65].
3.1.2. Fluid flow
Fluid flow can be used to impose shear stress onto a group of cells, another type of mechanical stimulus. To this end, cells are cultured on the surface of a fluid flow channel. As fluid flows over the culture, shear stress is imposed on the cells from the boundary layer between the cells and the fluid. Fluid shear stress has been used in a variety of studies, including investigating the influence of fluid shear on proliferation of bovine aortic endothelial cells [66] and investigating rolling adhesion of white blood cells in shear flow [67]. The primary advantage of this method is the natural environment this study takes place in. Cells commonly interact with fluid flow in vivo, so this setup allows for a natural testing environment, thus fluid flow methods have been widely adopted to study the effect of blood flow on the physiology of endothelia [68–70].
Two types of fluid flow systems are commonly used. The first is a parallel plate system (Fig. 3a), in which fluid flow is driven through a small rectangular chamber using a pressure differential, normally a syringe pump. A variety of parallel plate systems have been developed [71–77], and technological advances allowed for smaller microscopic parallel plate systems [24,25,78]. In these studies, the dimensions of the channel can be varied to control the flow characteristics and thus shear stress, and are kept small to ensure a low Reynolds number and thus laminar flow. In one study, the width of the fluid flow channel was varied between 0.25 and 1 mm, which changed the shear stress imposed on a culture of fibroblasts [79]. To generate easily-controlled uniform shear stress, a cone and plate system filled with cell medium is employed (Fig. 3b), in which a cone is rotated along its axis above the surface of a circular plate, creating a fluid flow [80–84]. The tangential speed of the cone increases with increased distance from the axis of rotation, but the distance between the cone and flat plate also increases, resulting in uniform shear stress distributions on the cell monolayer along the plate and the cone surface. By varying the angle of the cone and the speed of rotation, shear stresses of different magnitudes can be achieved. Specialized fluid flow profiles and flow chamber geometries are often designed to generate different temporal and spatial shear stress gradients [26,85].
Fig. 3.
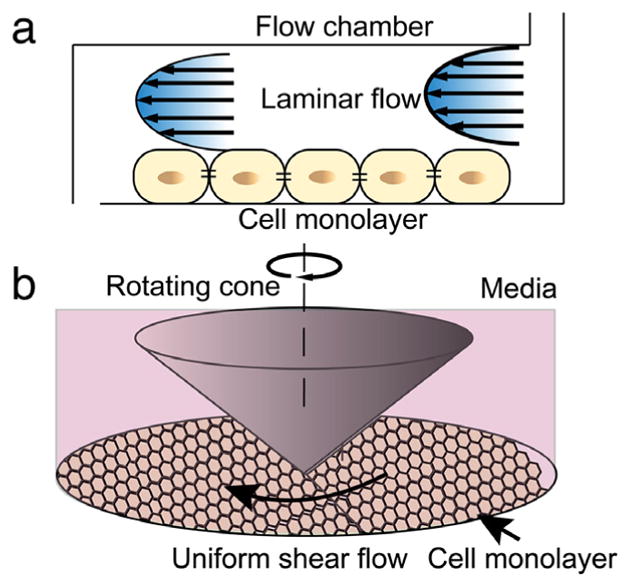
Fluid shear stimulation of a monolayer of cells. a. Fluid shear in a two-plate based flow chamber; b. Fluid shear with a plate and rotating cone system.
Fluid flow is best studied in the endothelial cell system to mimic the effect of blood flow on the vessel wall, particularly the endothelium, aiming to understand the process of atherosclerosis. In general, fluid flow produces two principal stress components: the stress perpendicular to the vessel wall and the stress parallel to the monolayer of endothelial cells. The former is the tensile stress exerting a dilating force on the vessel wall. The latter, fluid shear stress, represents the frictional force the fluid flow exerts on the endothelial cells, and it activates a cascade of mechanotransduction pathways that leads to endothelial cell proliferation [86], reorganization of the cytoskeleton [87] and even cell death [88]. Although the molecular basis for mechanosensing of fluid shear stress and for cells’ ability to discriminate flow patterns remains to be fully revealed, studies have shown a mechanotransduction cascade coordinates the redistribution of forces at cell–cell junctions and cell–ECM adhesions [89].
The endothelial AJ contains VE-cadherin and platelet endothelial cell adhesion molecule (PECAM)-1, both serving as anchors to the cell cytoskeleton and sustaining tension across the cell–cell junction. When a monolayer of endothelial cells is subject to fluid shear stress, though the cell layer tends to re-align itself in the direction of the flow to minimize drag resistance, the tension gradient at the apical surface of the cell membrane is eventually transmitted to junctional complexes. To maintain mechanical integrity, significant cytoskeletal remodeling takes place upon the onset of fluid flow [70]. Specifically, it has been reported that shear flow triggers the enhanced association of PECAM-1 with vimentin, which facilitates the transfer of actomyosin-generated tension within the actin network, thus relieving the stress at VE-cadherin adhesion complex [90,91]. This sequence of events leads to the rapid increase of tension at junctional PECAM-1 and a simultaneous drop of tension at junctional VE-cadherin, measured by FRET sensors in both adhesion molecules [90]. It is not surprising that PECAM-1 was identified as a mechanosensor that is associated with vimentin, an IF protein within endothelial cells. Similar roles for the IF-anchoring complex, the desmosome, are also reported in epithelial cells [92] with evidence of abundant crosstalk between desmosomes and AJs.
3.2. Single cell level stimulation and interrogation
Techniques used to quantify forces and stresses imposed on cells were developed to more fully understand biological responses to external mechanical stimuli. By studying isolated single cells or individual cells within a large group, complex responses to external mechanical stimuli can be recorded. However, these techniques offer new challenges to manipulate and position cells individually, with single cell control. Among the most common approaches are traction force microscopy (TFM), substrates with deformable microposts, micropipette aspiration, optical tweezers, magnetic twist cytometry (MTC), atomic force spectroscopy (AFM), laser ablation, and cantilever based spectroscopy. A description of each technique follows.
3.2.1. Traction force microscopy
TFM is the first mature technique to be employed to measure cell generated forces [93]. It consists of a flexible substrate containing embedded fluorescent beads over which adherent cells are cultured. In 2D TFM (Fig. 4a1), measurements of the substrate deformation together with the application of an inverse method using half-space elasticity solutions enables the construction of a map of the in-plane component of the traction vectors. As such the technique is capable of measuring forces at the single cell level. The displacements of beads due to substrate deformation are calculated by means of digital image correlation techniques [94]. Common substrates used for TFM include polyacrylamide or silicon based gels. They are selected due to their linear elasticity, optical transparency, and tunability of elastic moduli through polymer chain crosslinking, over several orders of magnitude [95]. Knowing the elastic moduli of the substrates, the displacement field can be converted to a traction force field using an inverse method [93, 96,97]. These forces are often on the order of nanonewton or tens of nanonewtons. To measure both the normal and in-plane components of traction vectors, 3D TFM has been developed in recent years [94,98–100]. Potential sources of inaccuracy in computing the displacement field include polymer degradation and nonlinearity of local deformation among others [101].
Fig. 4.
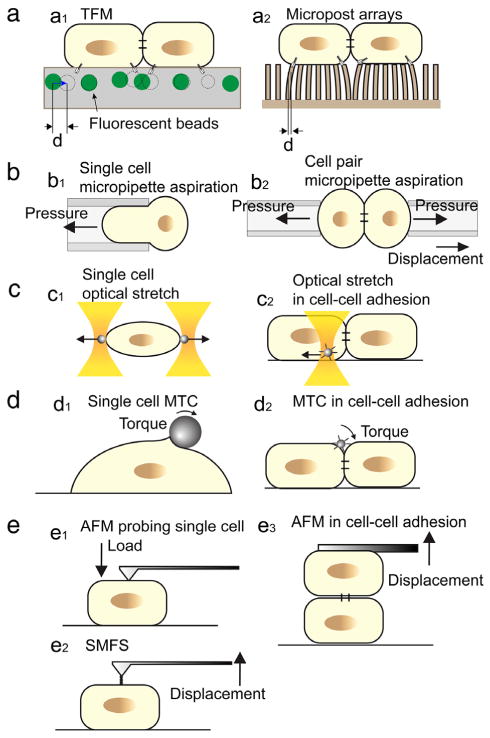
Cell force interrogation techniques and their use in cell–cell adhesion studies. a. TFM (a1) and micropost arrays (a2); b. Micropipette aspiration for single cells (b1) and cell–cell adhesion (b2); c. Optical trapping and stretcher for single cells (c1) and for cell–cell adhesion (c2); d. MTC for single cells (d1) and for cell–cell adhesion (d2); e. AFM based single mechanical interrogation (e1), single molecule force spectroscopy (SMFS) (e2) and single cell force spectroscopy (SCFS) for cell–cell adhesion (e3).
In the study of cell–cell adhesion, TFM has shown direct evidence of crosstalk between cell–cell and cell–ECM adhesions by indirectly quantifying cell–cell tugging forces through their force balance with cell–ECM traction forces [102–104]. These findings showed that integrin-mediated adhesions regulate tension and composition of cell–cell junctions [105–109]; conversely, cadherin-based cell–cell adhesions in epithelial cells modulate cell–ECM traction forces [102]. For example, inhibiting cell–cell adhesion in small colonies of keratinocytes by reducing calcium levels or genetically depleting cadherins (E-, and P-cadherin) results in remarkably different traction force patterns. Under control conditions, cooperative traction forces with maximum stress are localized to the periphery of cell colonies, and inhibiting cell–cell adhesion results in independent and significant traction forces evenly distributed throughout the colonies [102]. Similar cooperative activities of cell–cell and cell–ECM adhesion were also observed in heart muscle cells [110]. Studies using TFM also revealed that the force balance as well as molecular tension in E-cadherin are modulated by spatial distributions of FA sites [111]. TFM was also used advantageously to study mechanisms of cell migration [112].
3.2.2. Substrates with flexible microposts
A simple and elegant version of TFM was achieved by micro-fabrication of arrays of polydimethylsiloxane (PDMS) posts using micromolding [113] (Fig. 4a2). When cells are cultured over the substrate, in-plane components of adherent forces are revealed by the deflection of the microposts. Their cantilever geometry and chemical composition of the PDMS determine their stiffness. Hence, by using optical microscopy, the deflection amplitude and direction can be measured, from which the forces the cell is exerting on the microposts can be identified. The primary advantage of this technique is the large number of independent force measurements easily achieved as a function of position and time, which when combined, provide a vector map of traction forces. One embodiment of this technique used microposts with varying stiffness by changing the geometry of individual microposts. This allows for control of the sensitivity of force measurements on certain regions of the cell [113]. Further, by combining micropost fabrication with microcontact printing of ECM proteins, a cell pair can be confined to a predefined matrix pattern [114]. Relying on the zero net-force relationship in equilibrium condition, the forces being transmitted through the cell pair interface can be obtained from the vector sum of the forces acting on each set of microposts under the cell pair. Limitations of this technique are the lack of quantification of the component of the force vector normal to the substrate and the inability to use the microposts to apply forces on cells. By embedding a magnetic cobalt nanowire inside a subset of the microposts, application of forces to cells is possible using an externally applied magnetic field [37]. This approach was used to study the relationship between external mechanical forces and cytoskeleton induced forces. The applied forces were correlated to FAs and traction forces measured on non-magnetic posts near the magnetic posts [37]. Force stimulation has also been achieved by stretching the substrate containing micropost arrays [115].
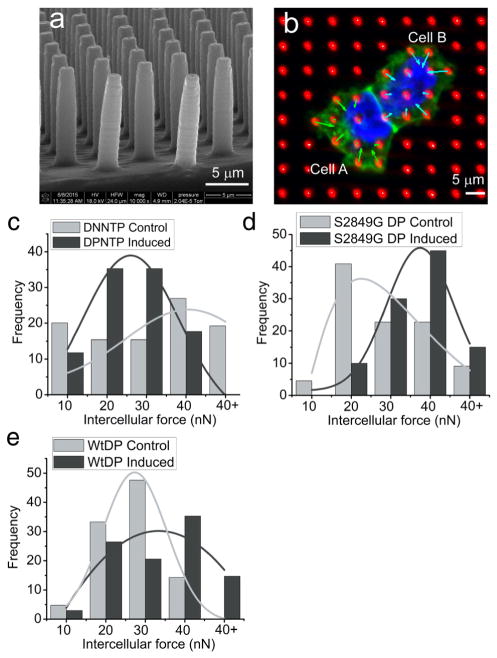
In the study of cell–cell adhesion, an early attempt using micropost arrays combined with microcontact printing showed that actomyosin generated contractile force regulates the size of AJs [114], indicating that cell contractility affects not only the composition at cell–ECM adhesions but also the composition of cell–cell junctions. In a recent study, the desmosome/IF linkage at cell–cell junctions has also been shown to regulate not only traction forces at cell–ECM adhesions, but also modulate forces at cell–cell junctions [21]. Using a combined approach of micropost arrays and AFM, epithelial cells with genetic mutation of a desmosome/IF linker molecule, DP, which lacks the IF-binding domain, exhibited significantly reduced traction forces and potentially increased tension or conformational changes in α-catenin of AJs [21,116]. This evidence lends support to the notion that abundant molecular communication exists between the two mechanically active junctional complexes [117]. The same study showed that the desmosome/IF linkage has a significant impact on global cell mechanics as well as tension within the cell–cell interface [21]. Different forms of DP were expressed in A431 cell lines to investigate the role of this linkage in regulating cell mechanics. These include: DPNTP, a truncated form of DP that lacks the IF binding domain, S2849G DP, which contains a serine to glycine mutation that enhances the desmosome/IF linkage, and wildtype DP (WtDP). Micropost arrays of different dimensions and elasticities were fabricated using micromolding (Fig. 5a, shown is micropost arrays with height of 10 μm and diameter of 2 μm and elastic modulus of 2.41 MPa). Cell–cell adhesion forces were derived by the zero-sum force balance between the two cells (Fig. 5b). Measurement results from the micropost arrays show that DPNTP expression results in a significant reduction of average intercellular forces while S2849G DP expression increases these forces significantly, and expression of WtDP only increases the intercellular forces slightly (Fig. 5c, d, e). These experimental data measured using micropost arrays suggest that the desmosome/IF linkage also modulates cell–ECM interactions, similar to AJs [102].
Fig. 5.
Micropost arrays reveal that desmosome/IF linkage regulates cell–cell adhesion forces. a. Micropost arrays 10 μm in height and 2 μm in diameter are fabricated using micromolding of PDMS. b. Force balance established between cell pairs are used to calculate cell–cell tugging force. c, d, e. Distribution of intercellular forces in nN as measured by micropost arrays for DPNTP, S2849G DP and WtDP, respectively. c, d and e are recreated from Fig. 2 in [21], reprinted with permission.).
3.2.3. Micropipette aspiration
Micropipette aspiration is a simple technique in which a single cell is partially or completely aspirated into the tip of a micropipette (Fig. 4b1). In an aspiration experiment, a micropipette with inner diameter smaller than that of a suspended cell, equipped with a micromanipulator positioning system, is brought into contact with a cell while a negative pressure is applied. The negative pressure forces the suspended cell to attach to the tip of the micropipette as a microscope imaging system records cell deformation during the process [118]. This simple procedure has been used to investigate membrane elasticity [119,120], to quantify mechanical and material properties of single cells and cell nuclei [121,122], to measure molecular adhesions between pairs of cells [123,124], and to study mechanotransduction within single cells [125,126]. The technique imposes large strains on cells, and as a result is incompatible with some cell types [27]. High deformation gradients along the edge of the pipette and friction between the pipette surface and the cell membrane often impact measurement results. Mathematical models have been used to convert resulting strains into stresses in erythrocytes [127,128] and chondrocytes [129], but they have limited accuracy due to the model assumptions. Finite element analyses with large deformations have been performed to determine mechanical properties of suspended cells, such as chondrocytes, with different viscoelastic models [28,130].
To study cell–cell adhesion, two cells are brought into contact with each other using two micropipettes in a process called dual micropipette aspiration (DPA) (Fig. 4b2). The micropipettes use a slight vacuum that can securely hold the cell on the micropipette without significant aspiration so as to avoid large strains. Cells form a cadherin-based cell–cell adhesion complex after a certain contact time. While one micropipette maintains a sufficient amount of pressure to hold the cells in place, the pressure in the other micropipette is increased while the pipette retracts until the two cells are separated. The forces required to break the cadherin adhesions can be calculated from the aspiration pressure required to separate the two cells [131,132], which is in the nanonewton range. When separating cell doublets with cadherin bonds, due to the short separation time of less than 1 s, the process can be regarded as separation of two elastic solids where the separation force depends on the density and dissociation rate of adhesion bonds [59]. We note that the micropipette aspiration technique can be easily integrated with fluorescence microscopy, which allows the imaging of the aggregation of adhesion molecules as well as the measurement of cell cortex deformation in real-time. Contact time can be controlled to ensure the maturity of the cell–cell adhesion complexes. In this context, the technique has been used to study cadherin binding forces between CHO cells and red blood cells [133] and to quantify cadherin-dependent cell–cell adhesion in E-cadherin expressing cell doublets [29].
Studies using micropipette aspiration have only been conducted on suspended cells until a recent report extended its utility to cells in adherent states. In the study, a single endothelial cell adherent on a substrate was aspirated from the apical surface using a micropipette tip, placed perpendicular to the substrate, while an inclined mirror was employed to measure the cell strains during aspiration [134]. The experiments revealed that the interaction between cytoskeletal actin and ECM, in adherent cells, increases the contractile forces within the cytoskeleton and thus the resistance to aspiration. This critical element was missing in studies on suspended cells. In addition, similar to micropipette aspiration in suspended cells, the nucleus was shown to be highly deformable under the considerably large strains resulting from aspiration. These experimental results were interpreted employing a viscoelastic finite element model.
3.2.4. Optical traps and stretchers
Optical traps use momentum conservation of diffracting photons to impart small forces on dielectric objects [135]. This method has been used to study many molecules [136], including the kinetics of RNA unfolding [137]. When used to study cells, dielectric microbeads are adhered to the cell and act as handles for the optical trap (Fig. 4c1). The adhesion strength limits the maximum force an optical trap can exert on a cell [30,138]. Once the microbeads are adhered to the cell, a laser is directed through one of the microbeads, refracting the laser. The refraction changes the momentum of the photons, and thus changes the momentum of the microbead, inducing a force. The microbead is attracted towards the focal point of the laser, and therefore the force can be controlled by altering how the laser is focused on the microbead [139]. For this to work, the refractive index of the bead must be larger than the refractive index of the medium surrounding the cell. Generally, this technique is used to apply static loading to cells, but can also be used for cyclic loading, using an acousto-optic modulator [140]. The two main advantages of these techniques are high force resolution and lack of physical contact between the actuation and the cell. Using this technique, sub pN forces were achieved, and cells were studied in their natural environments as they do not need to be physically attached to an instrument. However, the maximum achievable force is limited to a few hundred pN, as high laser power may impart radiation damage onto the cell.
A study that took full advantage of the resolution achieved with optical traps examined the impact malaria has on the mechanical properties of diseased red blood cells [141]. Healthy red blood cells [138] and infected blood cells [31,142] were stretched using an optical trap to quantify their elastic moduli. The diseased cells were tested in various stages of the infection, revealing the shear modulus steadily increased throughout the duration of the infection, increasing by about an order of magnitude at the final stage of the disease.
In the study of cell adhesion, optically trapped beads, a few micrometers in diameter, were tethered to matrix proteins such as functional peptides [143] or fibronectins [144] to measure adhesion forces (Fig. 4c2). The technique enabled real-time monitoring of traction forces, during the formation of cell adhesion sites, functioning as a high resolution (~pN) force sensor. However, due to the complexity of the optical setup, only a few beads could be tracked simultaneously, limiting its spatial resolution. The technique has also been deployed to investigate the interactions of AJs with filamentous actin (F-actin) [145]. To this end, two optical trapped beads attach to both ends of a single actin filament suspended above a purified cadherin–catenin complex. The objective was to mimic the parallel spatial arrangement of F-actin filaments with the cell membrane at the cell–cell junction. To apply tension, the cadherin–catenin complex was anchored onto a microsphere attached to a movable stage. During the cyclic motion of the stage, when the immobilized complex bound to the suspended filament, the interaction force can be obtained from the optically-displaced beads. The study revealed that α-catenin is required to form the AJ/actin filament tether and that the bond between the complex and the actin filament is tension-enhanced and catch-bond like.
A variation on optical traps, known as optical stretchers, uses divergent lasers that interact directly with the cell without the need for microbead handles [146–149]. This technique was most recently employed to probe the mechanics of cell–cell interfaces [149]. In this study, mature cell–cell junctions of a drosophila embryo were interrogated by optically imposing a deformation (<1 μm) on the cell–cell interface. Observation of the retraction of the cell–cell junction to the original shape and the restoration of force balance provides information about the mechanics of cell–cell junctions during early tissue morphogenesis. The study shows that tension at cell–cell junctions, on the order of 100 pN, can equilibrate over a few seconds and that the time-dependent properties of the junction can be reproduced by a simple viscoelastic model.
3.2.5. Magnetic beads
Magnetic beads can be used as handles to apply forces to a cell, in a process often referred to as magnetic twisting cytometry (MTC) (Fig. 4d1). The beads are coated with a group of molecules that allow them to bind to specific cell surface receptors [23]. Once they are attached, they can be manipulated through control of an external magnetic field. This method was developed in 1950 [150] and has been used in a variety of studies, including applying cyclic loadings to cells [33,151–154]. The technique has been mostly applied to investigate the mechanical properties, especially viscoelasticity, of cells [151], but was also applied to fundamental studies of the role of membrane forces in gene regulation [155]. Two main advantages of MTC are the ability to apply torque to cells and the ability to easily apply cyclic loadings through control of the magnetic field. However, due to unfavorable scaling of magnetic forces with size, applying large forces with this technique requires large beads relative to cell size [156].
Similar to the optical trapping method, MTC can apply and measure forces with pN resolution. To this end, magnetic beads are normally coated with RGD sequences that link them to integrin receptors, which led to the discovery of integrin-based mechanotransduction [153]. Recent studies on mechanotransduction at cell–cell junctions have revealed that applied forces at E-cadherin have profound impact on cell contractility and cell mechanics [157]. A combination of MTC for force loading at the cell–cell adhesion complex and traction force microscopy (TFM) to quantify traction forces at cell–ECM interactions was employed (Fig. 4d2). Beads coated with E-cadherin ligands were employed to induce an oscillating shear stress (~10 Pa) through modulation of the applied magnetic field. The study revealed that shear stress induced at cell–cell junctions can trigger a significant increase in traction forces. This also demonstrates a new mechanotransduction pathway from cell–cell junctions to FAs, potentially through EGFR-PI3K mediated pathways. It is yet to be determined the extent to which the α-catenin-actin filament linkages participate in the force sensing and transducing process, despite the many previous reports confirming their tension-maintaining and conformation-altering roles [145, 158].
3.2.6. Atomic force microscopy
Atomic force microscopy (AFM) is a technique that was originally developed to map surfaces of any material with nanometer resolution. A microfabricated silicon cantilever beam, having a sharp tip at its end, is brought into contact with a surface. As the tip interacts with the surface, the cantilever beam bends. By tracking its deflection, using a laser beam deflected off the back of the cantilever beam, the tip–surface interaction force can be measured when the cantilever stiffness is known. The technique has sub-pN force resolution and sub-nm displacement resolution, but has a limited maximum force and displacement. It has been adopted to the study of cell mechanics by relating the amount the cantilever beam deflects to the force the probe exerts. AFM has been successfully employed to study cell elastic [159–163] and viscoelastic properties [164], as well as nuclei stiffness [165] (Fig. 4e1). In addition to compressive forces, AFM can also be used to apply tensile forces to a cell. One way this can be done is by culturing cells directly on the AFM tip. This allows for easy manipulation of cells, and has been used to study cell–cell and cell-substrate interactions [166]. In addition, the AFM tip can be functionalized to bind to cell surface receptors in a technique widely known as single molecule force spectroscopy (SMFS) (Fig. 4e2). Once functionalized, the probe can be retracted to apply tensile forces and unwind or break molecular bonds [167]. SMFS has been used extensively to study binding affinities for a host of molecules, including the unbinding of DNA protein pairs [168,169].
In the study of cell–cell adhesion, AFM has been used in the context of single cell force spectroscopy (SCFS) [170,171] (Fig. 4e3). The technique is similar in principle with micropipette based cell–cell adhesion studies using DPA, where two cells are brought into contact and pulled apart to quantify their interaction. Compared with the DPA method, SCFS offers higher resolution (pN compared to nN for DPA) and less native strain to the cells before contact. In SCFS adhesion measurements, a living cell is first attached onto a tip-less AFM cantilever, normally by means of matrix protein coating, and the cell is brought into contact with another cell on the substrate by lowering the cantilever using a z-piezo stage. During the retraction phase, the interactions can be measured by recording the cantilever deflection. The sensitivity in force measurement and the fine position control, by the z-piezo stage, enables quantification of subtle cell–cell interactions during the initial phase of adhesion. These forces are often retraction rate-dependent and dwelling time-dependent [170,172]. A prolonged dwelling time often leads to the study of cooperative binding. SCFS studies have led to the conclusion that levels of E-cadherin determine the adhesion strength between different types of zebrafish progenitor cells during development [173]. Destructive methods are also used to probe cellular responses including the use of AFM probes [162,163] or laser pulses [174–176] to dissect cytoskeleton components.
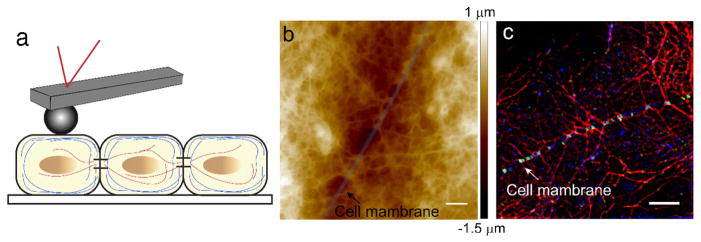
Performed on the center of individual cells, AFM nanomechanical studies can reveal subtle changes in global cell mechanics (illustrated Fig. 6a). AFM studies on A431 cells expressing various forms of DP have shown that removing the desmosome/IF linkage reduces cellular stiffness significantly compared with controls, conversely, enhancing the linkage increases cellular stiffness. These results suggest that changes at the cell–cell adhesion complex have a major impact on global cell mechanics. In addition, the stiffness changes observed on cells with DP mutations can be abrogated by depolymerizing actin filaments, indicating a potential crosstalk between the desmosome/IF and AJ/actin filament networks [21]. Furthermore, considering the significant difference in the mechanical properties of the cytoskeleton and the cytosol, AFM images obtained by applying a small force to the cell can reveal the cytoskeleton structures at the cell–cell adhesion junction as shown in Fig. 6b. The imaged structure can be compared to a correlative immunofluorescence image to identify the type of filaments (similar to the labeling of IF cytoskeleton in Fig. 6c). Mechanical measurements can be performed on individual filaments to reveal their tensional states when the junction complexes are modulated. These mechanical characterizations will inform our understanding from a global cell mechanics perspective, where the cell cytoskeleton is considered as a tensegrity structure [177].
Fig. 6.
AFM imaging reveals cytoskeletal bundles. a. AFM nanomechanical measurements reveal global cell mechanics. b. AFM image is scanned at the cell–cell junction. Scale bar: 2 μm. c. To identify the filaments in the AFM image, an immunofluorescence image is taken at the junction to visualize IF (Red: IF; Blue: plakoglobin; Green: DP). Scale bar: 5 μm. (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article.)
Cell–cell adhesion studies also take advantage of the binding force measurement capability of AFM. SMFS experiments on the desmosomal cadherin, Dsg3, have revealed the distinctive binding affinities of Dsg3 molecules at different regions of the cell surface [178]. Using a Dsg3 functionalized AFM probe, the study showed that homophilic Dsg3 binding forces are stronger at cell–cell contacts than at other cell surface areas. Further, this binding event can be effectively blocked by calcium depletion and Dsg3 antibodies [178]. In a separate study, the same group identified that inhibition of Dsg3 binding, which occurs in the autoimmune disease pemphigus vulgaris, does not lead to cell–cell adhesion loss, rather it alters downstream signaling events that may contribute to that effect, such as p38 mitogen-activated protein kinase (p38 MAPK) [179]. Studies using SMFS also revealed the mechanisms of bond formation between desmosomal cadherins: Dsc forms calcium-dependent homophilic bonds and Dsg forms calcium-independent heterophilic bonds with Dsc. [180]. Similar approaches have been taken in the study of E-cadherin based cell–cell adhesions, and the mechanism of catch bond formation between E-cadherin molecules has been demonstrated [181,182].
3.2.7. Laser ablation
The contractile forces present within cell–cell contacts of living cells can be directly measured using laser ablation [183]. The cell cortex at sites of cell–cell contact is physically cut using a high-power laser (e.g. two-photon laser). A fiduciary fluorescently tagged membrane marker is used to track the displacement of the vertices at either side of an ablated cell–cell junction. The rate of vertex recoil can then be used to extract force-related parameters including the contractile force that was present within the junction as well as the ratio between junction elasticity and the viscosity of the cytoplasm [183]. Tension within cell–cell junctions can then be compared across experimental conditions by comparing initial recoil velocities. This approach has been used to determine the contributions of Src kinase [184], actin regulating proteins like N-WASP [185] and the Rho guanosine nucleotide exchange factor Ect2 [186], and myosin II [187] to the generation of tension at sites of cell–cell contact. Finally, this method is capable of measuring cell–cell forces both in traditional cell culture models as well as in vivo models like zebrafish [188].
4. MEMS and beyond for parallel stimulation and interrogation
The aforementioned techniques, while effective, present some limitations in terms of force and displacement resolutions, and imaging modalities. To overcome such limitations, researchers resorted to the design flexibility offered by MEMS through creation of specialized platforms for cell–cell adhesion studies. Parallel stimulation and measurement of forces were achieved by employing compliant mechanisms embodied in various configurations [189].
4.1. Moveable structures
In a moveable platform MEMS device, a cell is adhered to a platform that is split into two or more parts. The cell is adhered to the platform while the parts are together, and then the parts of the platform are separated using an external actuator, e.g., piezoelectric actuator, and mechanical linkages. As the parts of the platform separate, the cell is stretched, and the degree to which the cell is deformed can be controlled by the separation distance between the parts of the platform. Two variations of this technique have been implemented, a uniaxial puller and a biaxial puller.
An example uniaxial puller consists of two platforms, one of which is fixed while the other is moveable (Fig. 7a). The moveable platform is attached to an external piezoelectric actuator, which can control the displacement of the platform. In one study, a uniaxial puller was used to study mechanical properties of hydrated collagen fibrils [190]. An electrostatic comb drive actuator was employed to actuate one of the platforms, while the other was held rigidly in place. The main advantages of using an electrostatic comb-drive actuator include low power consumption using moderate driving voltages, and high speed and accuracy. Also, use of an electrostatic comb drive actuator allowed for cyclic loading of the cell. A biaxial puller was developed that used an electrostatic comb-drive actuator and a cleverly designed kinematic linkage that allowed for controlled actuation of four segments of a platform at the same time [36] (Fig. 7b). In this setup, one part of the stage was fixed, while the other three were connected to a kinematic linkage and an electrostatic actuator. When the actuator moves, the linkage causes the three mobile portions of the platform to move in mutually orthogonal directions from each other. If small displacements are assumed, the relative motion away from each other is at the same speed. This results in uniform biaxial strain on the cell. It is worth mentioning that electrostatic actuation is not shown in the diagrams of Fig. 7. Electrostatic actuation in a liquid environment for biological applications has been developed using a differential actuation design [191]. The system is operated at high frequencies to compensate for the attenuation due to impedance loss in conductive media [192]. In sum, the MEMS-based stretching technique has not been reliably deployed to study cell–cell adhesion mainly because of the difficulty in aligning the cell doublets in a spatial arrangement where the strain direction is perpendicular to the cell–cell junction. In addition, the difficulty also lies in the compatibility of MEMS materials for cell attachment and growth.
Fig. 7.
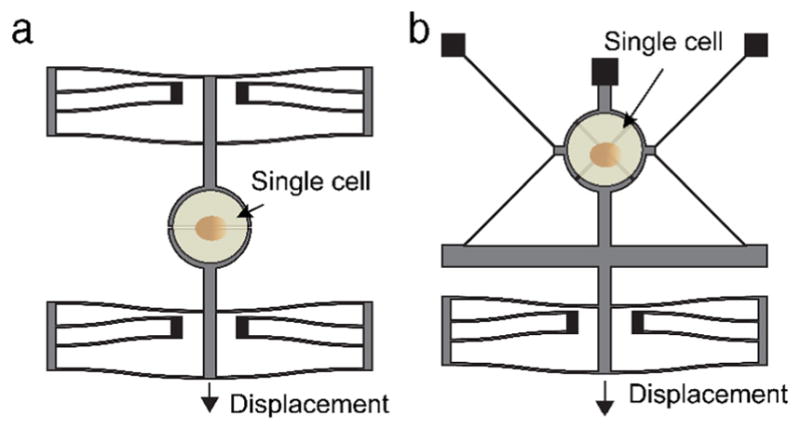
MEMS based single cell interrogation and stimulation systems. a. Uniaxial single cell stretching; b: Biaxial single cell stretching. (Reprinted with permission from Fig. 4 of [189].).
To investigate cell–cell junctions and meet the challenges mentioned, we fabricated and integrated a microdevice capable of parallel stretching and sensing of forces (Fig. 8). The device is floating above a window through a silicon wafer and consists of folded beams (load sensor) and shuttles to support rafts on which cell pairs are cultured. It is designed to apply tension to a pair of cells placed between the two shuttles. The stretching is achieved by means of an external manipulator and a metallic needle that interfaces with the device through an aperture made on the right shuttle (Fig. 8a). Considering the large deformability of cells and the high load resolution needed to capture breaking of desmosome cell–cell adhesions, we use a set of folded beams over a length of 500 μm possessing a stiffness of 12 pN/nm, which allows measurement of forces as low as 250 pN using digital image correlation (DIC). This force is equivalent to the strength of ~3–6 cadherin bonds [193]. DIC enables correlation of features in a pair of images using software analysis. A mathematical correlation algorithm is applied to track the position of features (here we track features included in the surface of the shuttles shown in Fig. 8b) from one image to the next relative to an initial reference image. The accuracy of this method is approximately ± 0.1 pixels (resolutions of 20 nm have been achieved [194] when a 100X objective, 1.40 N.A., is used).
Fig. 8.
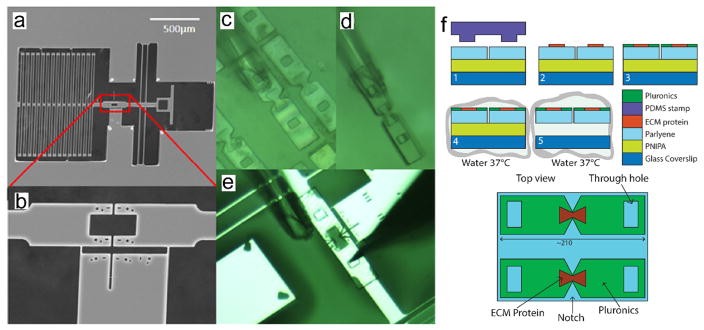
Parallel stimulation and interrogation of cell–cell adhesion with bioMEMS system. a. SEM image of the microfabricated device, composed of folded beams (load sensor), and shuttles to mount the rafts in which cells are cultured (b). c–e. The parylene C rafts are successfully released from the substrate by dissolving a sacrificial layer (PNIPA) in 37 °C water, manipulated using a micropipette and positioned on a device. f. Raft fabrication and release. Top view of the rafts with stamped ECM proteins.
To contain the live cells, a microfabricated parylene C raft in the shape of a bowtie, which is patterned with extracellular matrix (ECM) proteins, is placed on the device in a gap between the shuttles (Fig. 8c–e). The raft is fabricated by patterning the deposited parylene C layer on top of a sacrificial layer, PNIPA. The patterned raft can be released by increasing the media temperature to above 37 degrees, which dissolves the PNIPA (Fig. 8f). Cells are then plated over the microdevice platform, and a pair of cells is placed across the two half rafts using micropipette aspiration. The cell pair is confined in position by the physical well in the parylene layer, which is approximately 8 μm in depth. The shuttles bear an opening that corresponds to the location of the cell pair on the rafts such that the cells can be observed by an inverted fluorescence microscope. Simultaneous imaging by confocal fluorescence microscopy would enable tracking of the evolution of intercellular adhesions and cytoskeletal networks.
4.2. 3D Nanofabrication
Advances in nanofabrication techniques promise to enable unique cell adhesion studies. For example, two-photon polymerization (TPP) has enabled the fabrication of devices at nanoscale and more importantly using biocompatible fabrication materials [195]. A new class of micro-scaffold with nanometer scale features has been developed for cell attachment and growth [196–199], for force measurement from cell adhesion induced interactions [200], and for stimulation of FAs [201]. This method is still in its early stages, but it is very promising because it offers biocompatibility and the potential to probe cells within a 3D environment.
5. Sensing intra -cellular forces with FRET imaging
To gain insights into the conformational changes of mechanosensor molecules and to reveal the mechanisms of mechanosensing, genetically encoded tension sensors were developed to introduce fluorophores with compatible emission/absorption spectra [202–205]. FRET is based on the nonradiative energy transfer between one fluorophore, the donor, and another fluorophore, the acceptor. The efficiency of this transfer depends highly on the relative distance between the fluorophores. The Förster distance, a characteristic length at which efficiency of energy transfer equals 50%, is a few nanometers [206]. Thus, FRET works with a length scale that is comparable with conformation changes of molecules within cells. Tremendous progress has been made in using FRET for the study of cell generated forces and mechanosensing of external forces. There have been reports of FRET measurements for detecting force-activated RhoA [207,208], Rac [209], Focal adhesion kinase (FAK) [210] and Src kinase [211].
Cell–cell junction molecules, E-cadherin [59,111,212], VE-cadherin [213], PECAM-1 [90], as well as linker molecules α-catenin [214] have been reported as effective mechanosensors. FRET sensors are most effective when used in parallel with other mechanical stimulation techniques, such as MTC [157,215]. Mechanical stress or strain applied to cell–cell junctions pose a conformational change to mechanosensors at the junction, and this change and the resulting tension within the sensor can be revealed qualitatively, possibly quantitatively with calibration [216], by monitoring the FRET ratio. Surprisingly, use of an E-cadherin tension sensor demonstrated that membrane-associated E-cadherin is under constitutive actomyosin generated tension, irrespective of whether or not it is present in cell–cell contacts; however, E-cadherin tension is increased at cell–cell contacts when adhering cells are stretched [59]. Moreover, introducing an α-catenin FRET sensor into AJs revealed a rapid and reversible conformational change when activated by mechanical strain. Further, it confirms that force dependent conformational change is followed by recruitment of vinculin to its α-catenin binding site [215]. An α-actinin FRET sensor was used to demonstrate that fluid shear stresses are transmitted to AJs through force redistribution within cytoskeletal binding proteins, and changes in cytoskeletal tension and reorganization are upstream in the response of cells to flow [217]. Finally, other types of cell–cell junctions, including tight junctions, can affect the forces within AJs. Depletion of the tight junction component ZO1 in endothelial cells resulted in a redistribution of actomyosin and a decrease in AJ tension as assessed using a VE-cadherin tension sensor [218].
6. Outlook
This review covers the most widely used techniques in stimulating and probing cell generated forces, focusing on cell–cell adhesion studies. While effective, these techniques are mostly restricted to in vitro studies, and it is still rather difficult to probe physical interactions in vivo. These techniques generally require specialized instrumentation and detailed calibration to realize desirable signal-to-noise ratio and resolution. Thus, improvements should be made to facilitate the adoption of these techniques in routine laboratory protocols. Researchers need also to consider a variety of issues in deciding which method to adopt. Besides displacement resolution (micrometer or nanometer) and force resolution (piconewton or nanonewton), the in vivo or in vitro observation conditions, the dimensionality of measurements (2D or 3D), the temporal resolution of the mechanotransduction pathways are all factors that influence inter- and intra-cellular processes. Specifically, for cell–cell adhesion studies, it is not yet possible to develop novel platforms for applying precise external load (force or strain) to a single cell–cell junction while simultaneously using confocal microscopy to measure its response. This capability would enhance our understanding of how mechanosensing modules in intercellular junctions drive cytoskeletal remodeling and potentially transcriptional responses affecting tissue differentiation and function. BioMEMS holds great potential to tackle these issues through innovative designs and advances in materials and fabrication processes. In this respect, it would be ideal to combine bioMEMS systems with well characterized FRET sensors to enable the application and measurement of external forces while unraveling the intracellular processes.
Acknowledgments
The authors acknowledge contributions made by O. Loh, M. Naghari, A. Beese, and K. Nandy towards the design and acquisition of images discussed in relation to cell–cell junction testing with microsystems. This work would have not been possible without the support from the McCormick School of Engineering through a catalyst award. This work was in part supported by NIH grants from NIAMS ( AR041836 and AR043380) and NCI (CA122151). R. Yang gratefully acknowledges financial support from the Nebraska Center for Integrated Biomolecular Communication (NCIBC) (NIH National Institutes of General Medical Sciences P20 GM113126). A. Monemian and J. Rosenbohm helped with literature search and their contributions are acknowledged.
References
- 1.Yusko EC, Asbury CL. Force is a signal that cells cannot ignore. Mol Biol Cell. 2014;25:3717–3725. doi: 10.1091/mbc.E13-12-0707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Gumbiner BM. Cell adhesion: the molecular basis of tissue architecture and morphogenesis. Cell. 1996;84:345–357. doi: 10.1016/s0092-8674(00)81279-9. [DOI] [PubMed] [Google Scholar]
- 3.Ingber DE. Cellular mechanotransduction: putting all the pieces together again. FASEB J. 2006;20:811–827. doi: 10.1096/fj.05-5424rev. [DOI] [PubMed] [Google Scholar]
- 4.DuFort CC, Paszek MJ, Weaver VM. Balancing forces: architectural control of mechanotransduction. Nat Rev Mol Cell Biol. 2011;12:308–319. doi: 10.1038/nrm3112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Winograd-Katz SE, Fassler R, Geiger B, Legate KR. The integrin adhesome: from genes and proteins to human disease. Nat Rev Mol Cell Biol. 2014;15:273–288. doi: 10.1038/nrm3769. [DOI] [PubMed] [Google Scholar]
- 6.Veale DJ, Maple C. Cell adhesion molecules in rheumatoid arthritis. Drugs Aging. 1996;9:87–92. doi: 10.2165/00002512-199609020-00003. [DOI] [PubMed] [Google Scholar]
- 7.Hahn C, Schwartz MA. Mechanotransduction in vascular physiology and atherogenesis. Nat Rev Mol Cell Biol. 2009;10:53–62. doi: 10.1038/nrm2596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Jaalouk DE, Lammerding J. Mechanotransduction gone awry. Nat Rev Mol Cell Biol. 2009;10:63–73. doi: 10.1038/nrm2597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.El-Sabban ME, Abi-Mosleh LF, Talhouk RS. Developmental regulation of gap junctions and their role in mammary epithelial cell differentiation. J Mammary Gland Biol Neoplasia. 2003;8:463–473. doi: 10.1023/B:JOMG.0000017432.04930.76. [DOI] [PubMed] [Google Scholar]
- 10.Rodriguez-Boulan E, Macara IG. Organization and execution of the epithelial polarity programme. Nature Rev Mol Cell Biol. 2014;15:225–242. doi: 10.1038/nrm3775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Suzuki T. Regulation of intestinal epithelial permeability by tight junctions. Cell Mol Life Sci. 2013;70:631–659. doi: 10.1007/s00018-012-1070-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Miller PW, Clarke DN, Weis WI, Lowe CJ, Nelson WJ. The evolutionary origin of epithelial cell-cell adhesion mechanisms. Curr Top Membranes. 2013;72:267–311. doi: 10.1016/B978-0-12-417027-8.00008-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Muhamed I, Wu J, Sehgal P, Kong X, Tajik A, Wang N, Leckband DE. E-cadherin-mediated force transduction signals regulate global cell mechanics. J Cell Sci. 2016;129:1843–1854. doi: 10.1242/jcs.185447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Sivasankar S. Tuning the kinetics of cadherin adhesion. J Invest Dermatol. 2013;133:2318–2323. doi: 10.1038/jid.2013.229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Yao M, Qiu W, Liu R, Efremov AK, Cong P, Seddiki R, Payre M, Lim CT, Ladoux B, Mège R-M, Yan J. Force-dependent conformational switch of α-catenin controls vinculin binding. Nature Commun. 2014;5 doi: 10.1038/ncomms5525. [DOI] [PubMed] [Google Scholar]
- 16.Saito M, Tucker DK, Kohlhorst D, Niessen CM, Kowalczyk AP. Classical and desmosomal cadherins at a glance. J Cell Sci. 2012;125:2547–2552. doi: 10.1242/jcs.066654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Rübsam M, Broussard JA, Wickström SA, Nekrasova O, Green KJ, Niessen CM. Adherens junctions and desmosomes coordinate mechanics and signaling to orchestrate tissue morphogenesis and function: An evolutionary Perspective. Cold Spring Harb Perspect Biol. 2017:a029207. doi: 10.1101/cshperspect.a029207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Johnson JL, Najor NA, Green KJ. Desmosomes: regulators of cellular signaling and adhesion in epidermal health and disease. Cold Spring Harb Perspect Med. 2014;4:a015297. doi: 10.1101/cshperspect.a015297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Harrison OJ, Brasch J, Lasso G, Katsamba PS, Ahlsen G, Honig B, Shapiro L. Structural basis of adhesive binding by desmocollins and desmogleins. Proc Natl Acad Sci. 2016;113:7160–7165. doi: 10.1073/pnas.1606272113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Seltmann K, Fritsch AW, Käs JA, Magin TM. Keratins significantly contribute to cell stiffness and impact invasive behavior. Proc Natl Acad Sci. 2013;110:18507–18512. doi: 10.1073/pnas.1310493110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Broussard JA, Yang R, Huang C, Nathamgari SSP, Beese AM, Godsel LM, Hegazy MH, Lee S, Zhou F, Sniadecki NJ, Green KJ, Espinosa HD. The desmoplakin/intermediate filament linkage regulates cell mechanics. Mol Biol Cell. 2017 doi: 10.1091/mbc.E16-07-0520. mbc. E16-07-0520. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Pioletti DP, Müller J, Rakotomanana LR, Corbeil J, Wild E. Effect of micromechanical stimulations on osteoblasts: development of a device simulating the mechanical situation at the bone–implant interface. J Biomech. 2003;36:131–135. doi: 10.1016/s0021-9290(02)00301-9. [DOI] [PubMed] [Google Scholar]
- 23.Wang H, Ip W, Boissy R, Grood ES. Cell orientation response to cyclically deformed substrates: experimental validation of a cell model. J Biomech. 1995;28:1543–1552. doi: 10.1016/0021-9290(95)00101-8. [DOI] [PubMed] [Google Scholar]
- 24.Tkachenko E, Gutierrez E, Ginsberg MH, Groisman A. An easy to assemble microfluidic perfusion device with a magnetic clamp. Lab Chip. 2009;9:1085–1095. doi: 10.1039/b812184b. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Young EW, Wheeler AR, Simmons CA. Matrix-dependent adhesion of vascular and valvular endothelial cells in microfluidic channels. Lab Chip. 2007;7:1759–1766. doi: 10.1039/b712486d. [DOI] [PubMed] [Google Scholar]
- 26.Bagi Z, Frangos JA, Yeh JC, White CR, Kaley G, Koller A. PECAM-1 mediates NO-dependent dilation of arterioles to high temporal gradients of shear stress. Arterioscler Thromb Vasc Biol. 2005;25:1590–1595. doi: 10.1161/01.ATV.0000170136.71970.5f. [DOI] [PubMed] [Google Scholar]
- 27.Hochmuth RM. Micropipette aspiration of living cells. J Biomech. 2000;33:15–22. doi: 10.1016/s0021-9290(99)00175-x. [DOI] [PubMed] [Google Scholar]
- 28.Baaijens FPT, Trickey WR, Laursen TA, Guilak F. Large deformation finite element analysis of micropipette aspiration to determine the mechanical properties of the chondrocyte. Ann Biomed Eng. 2005;33:494–501. doi: 10.1007/s10439-005-2506-3. [DOI] [PubMed] [Google Scholar]
- 29.Chu YS, Thomas WA, Eder O, Pincet F, Perez E, Thiery JP, Dufour S. Force measurements in E-cadherin-mediated cell doublets reveal rapid adhesion strengthened by actin cytoskeleton remodeling through Rac and Cdc42. J Cell Biol. 2004;167:1183–1194. doi: 10.1083/jcb.200403043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Dao M, Lim CT, Suresh S. Mechanics of the human red blood cell deformed by optical tweezers. J Mech Phys Solids. 2003;51:2259–2280. [Google Scholar]
- 31.Mills J, Diez-Silva M, Quinn D, Dao M, Lang M, Tan K, Lim C, Milon G, David P, Mercereau-Puijalon O. Eeffect of plasmodial RESA protein on deformability of human red blood cells harboring Plasmodium falciparum. Proc Natl Acad Sci. 2007;104:9213–9217. doi: 10.1073/pnas.0703433104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Suresh S, Spatz J, Mills JP, Micoulet A, Dao M, Lim CT, Beil M, Seufferlein T. Connections between single-cell biomechanics and human disease states: gastrointestinal cancer and malaria. Acta Biomater. 2005;1:15–30. doi: 10.1016/j.actbio.2004.09.001. [DOI] [PubMed] [Google Scholar]
- 33.Maksym GN, Fabry B, Butler JP, Navajas D, Tschumperlin DJ, Laporte JD, Fredberg JJ. Mechanical properties of cultured human airway smooth muscle cells from 0.05 to 0.4 Hz. J Appl Physiol. 2000;89:1619–1632. doi: 10.1152/jappl.2000.89.4.1619. [DOI] [PubMed] [Google Scholar]
- 34.Giessibl FJ. AFM’s path to atomic resolution. Mater Today. 2005;8:32–41. [Google Scholar]
- 35.Serrell DB, Oreskovic TL, Slifka AJ, Mahajan RL, Finch DS. A uniaxial bioMEMS device for quantitative force–displacement measurements. Biomed Microdevices. 2007;9:267–275. doi: 10.1007/s10544-006-9032-4. [DOI] [PubMed] [Google Scholar]
- 36.Scuor N, Gallina P, Panchawagh H, Mahajan R, Sbaizero O, Sergo V. Design of a novel MEMS platform for the biaxial stimulation of living cells. Biomed Microdevices. 2006;8:239–246. doi: 10.1007/s10544-006-8268-3. [DOI] [PubMed] [Google Scholar]
- 37.Sniadecki NJ, Anguelouch A, Yang MT, Lamb CM, Liu Z, Kirschner SB, Liu Y, Reich DH, Chen CS. Magnetic microposts as an approach to apply forces to living cells. Proc Natl Acad Sci. 2007;104:14553–14558. doi: 10.1073/pnas.0611613104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Rajagopalan J, Tofangchi A, Saif MTA. Linear high-resolution bioMEMS force sensors with large measurement range. J Microelectromech Syst. 2010;19:1380–1389. [Google Scholar]
- 39.Basso N, Heersche JN. Characteristics of in vitro osteoblastic cell loading models. Bone. 2002;30:347–351. doi: 10.1016/s8756-3282(01)00678-0. [DOI] [PubMed] [Google Scholar]
- 40.Jones D, Nolte H, Scholübbers J, Turner E, Veltel D. Biochemical signal transduction of mechanical strain in osteoblast-like cells. Biomaterials. 1991;12:101–110. doi: 10.1016/0142-9612(91)90186-e. [DOI] [PubMed] [Google Scholar]
- 41.Trepat X, Grabulosa M, Puig F, Maksym GN, Navajas D, Farré R. Viscoelasticity of human alveolar epithelial cells subjected to stretch. Am J Physiol-Lung Cell Mol Physiol. 2004;287:L1025–L1034. doi: 10.1152/ajplung.00077.2004. [DOI] [PubMed] [Google Scholar]
- 42.Trepat X, Puig F, Gavara N, Fredberg JJ, Farre R, Navajas D. Effect of stretch on structural integrity and micromechanics of human alveolar epithelial cell monolayers exposed to thrombin. Am J Physiol-Lung Cell Mol Physiol. 2006;290:L1104–L1110. doi: 10.1152/ajplung.00436.2005. [DOI] [PubMed] [Google Scholar]
- 43.Pfister BJ, Weihs TP, Betenbaugh M, Bao G. An in vitro uniaxial stretch model for axonal injury. Ann Biomed Eng. 2003;31:589–598. doi: 10.1114/1.1566445. [DOI] [PubMed] [Google Scholar]
- 44.Leung D, Glagov S, Mathews M. A new in vitro system for studying cell response to mechanical stimulation: different effects of cyclic stretching and agitation on smooth muscle cell biosynthesis. Exp Cell Res. 1977;109:285–298. doi: 10.1016/0014-4827(77)90008-8. [DOI] [PubMed] [Google Scholar]
- 45.Shukla A, Dunn A, Moses M, Van Vliet K. Endothelial cells as mechanical transducers: enzymatic activity and network formation under cyclic strain. Mech Chem Biosyst. 2004;1:279–290. [PubMed] [Google Scholar]
- 46.Pelham RJ, Wang Y-l. Cell locomotion and focal adhesions are regulated by substrate flexibility. Proc Natl Acad Sci. 1997;94:13661–13665. doi: 10.1073/pnas.94.25.13661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Somjen D, Binderman I, Berger E, Harell A. Bone remodelling induced by physical stress is prostaglandin E2 mediated. Biochim Biophys Acta. 1980;627:91–100. doi: 10.1016/0304-4165(80)90126-9. [DOI] [PubMed] [Google Scholar]
- 48.Meikle MC, Reynolds JJ, Sellers A, Dingle JT. Rabbit cranial sutures in vitro: a new experimental model for studying the response of fibrous joints to mechanical stress. Calcif Tissue Int. 1979;28:137–144. doi: 10.1007/BF02441232. [DOI] [PubMed] [Google Scholar]
- 49.Bottlang M, Simnacher M, Schmitt H, Brand R, Claes L. A cell strain system for small homogeneous strain applications-ein zellstimulations-system zur applikation kleiner homogener dehnungen. Biomed Tech/Biomed Eng. 1997;42:305–309. doi: 10.1515/bmte.1997.42.11.305. [DOI] [PubMed] [Google Scholar]
- 50.Hasegawa S, Sato S, Saito S, Suzuki Y, Brunette D. Mechanical stretching increases the number of cultured bone cells synthesizing DNA and alters their pattern of protein synthesis. Calcif Tissue Int. 1985;37:431–436. doi: 10.1007/BF02553714. [DOI] [PubMed] [Google Scholar]
- 51.Hung C, Williams J. A method for inducing equi-biaxial and uniform strains in elastomeric membranes used as cell substrates. J Biomech. 1994;27:227–232. doi: 10.1016/0021-9290(94)90212-7. [DOI] [PubMed] [Google Scholar]
- 52.Schaffer JL, Rizen M, L’Italien GJ, Benbrahim A, Megerman J, Gerstenfeld LC, Gray ML. Device for the application of a dynamic biaxially uniform and isotropic strain to a flexible cell culture membrane. J Orthop Res. 1994;12:709–719. doi: 10.1002/jor.1100120514. [DOI] [PubMed] [Google Scholar]
- 53.Williams J, Chen J, Belloli D. Strain fields on cell stressing devices employing clamped circular elastic diaphragms as substrates. J Biomech Eng. 1992;114:377–384. doi: 10.1115/1.2891398. [DOI] [PubMed] [Google Scholar]
- 54.Fu JP, Wang YK, Yang MT, Desai RA, Yu XA, Liu ZJ, Chen CS. Mechanical regulation of cell function with geometrically modulated elastomeric substrates (vol 7, pg 733, 2010) Nature Methods. 2011;8:184. doi: 10.1038/nmeth.1487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Faust U, Hampe N, Rubner W, Kirchgessner N, Safran S, Hoffmann B, Merkel R. Cyclic stress at mHz frequencies aligns fibroblasts in direction of zero strain. Plos One. 2011;6 doi: 10.1371/journal.pone.0028963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.De R, Safran SA. Dynamical theory of active cellular response to external stress. Phys Rev E. 2008;78 doi: 10.1103/PhysRevE.78.031923. [DOI] [PubMed] [Google Scholar]
- 57.Kaunas R, Nguyen P, Usami S, Chien S. Cooperative effects of rho and mechanical stretch on stress fiber organization. Proc Natl Acad Sci USA. 2005;102:15895–15900. doi: 10.1073/pnas.0506041102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.del Rio A, Perez-Jimenez R, Liu R, Roca-Cusachs P, Fernandez JM, Sheetz MP. Stretching single talin rod molecules activates vinculin binding. Science. 2009;323:638–641. doi: 10.1126/science.1162912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Borghi N, Sorokina M, Shcherbakova OG, Weis WI, Pruitt BL, Nelson WJ, Dunn AR. E-cadherin is under constitutive actomyosin-generated tension that is increased at cell–cell contacts upon externally applied stretch. Proc Natl Acad Sci. 2012;109:12568–12573. doi: 10.1073/pnas.1204390109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Mertz Aaron F, Banerjee Shiladitya YC, Goldstein Jill M, Rosowski Kathryn A, Revilla Stephen F, Niessen Carien M, Cristina Marchetti M, Dufresne Eric R. Valerie Horsley, Cadherin-based intercellular adhesions organize epithelial cell–matrix traction forces. Proc Natl Acad Sci USA. 2013 doi: 10.1073/pnas.1217279110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Mui KL, Chen CS, Assoian RK. The mechanical regulation of integrin-cadherin crosstalk organizes cells, signaling and forces. J Cell Sci. 2016;129:1093–1100. doi: 10.1242/jcs.183699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Bays JL, Peng X, Tolbert CE, Guilluy C, Angell AE, Pan Y, Superfine R, Burridge K, DeMali KA. Vinculin phosphorylation differentially regulates mechanotransduction at cell–cell and cell–matrix adhesions. J Cell Biol. 2014;205:251–263. doi: 10.1083/jcb.201309092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Yonemura S, Wada Y, Watanabe T, Nagafuchi A, Shibata M. alpha-Catenin as a tension transducer that induces adherens junction development. Nature Cell Biol. 2010;12:533–U535. doi: 10.1038/ncb2055. [DOI] [PubMed] [Google Scholar]
- 64.Benham-Pyle BW, Pruitt BL, Nelson WJ. Mechanical strain induces E-cadherin–dependent Yap1 and β-catenin activation to drive cell cycle entry. Science. 2015;348:1024–1027. doi: 10.1126/science.aaa4559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Gudipaty SA, Lindblom J, Loftus PD, Redd MJ, Edes K, Davey CF, Krishnegowda V, Rosenblatt J. Mechanical stretch triggers rapid epithelial cell division through Piezo1. Nature. 2017;543:118. doi: 10.1038/nature21407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Levesque M, Nerem R, Sprague E. Vascular endottielial cell proliferation in culture and the influence of flow. Biomaterials. 1990;11:702–707. doi: 10.1016/0142-9612(90)90031-k. [DOI] [PubMed] [Google Scholar]
- 67.Dong C, Lei XX. Biomechanics of cell rolling: shear flow, cell–surface adhesion, and cell deformability. J Biomech. 2000;33:35–43. doi: 10.1016/s0021-9290(99)00174-8. [DOI] [PubMed] [Google Scholar]
- 68.White CR, Frangos JA. The shear stress of it all: the cell membrane and mechanochemical transduction. Phil Trans R Soc B. 2007;362:1459–1467. doi: 10.1098/rstb.2007.2128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Fisher AB, Chien S, Barakat AI, Nerem RM. Endothelial cellular response to altered shear stress. Am J Physiol Lung Cell Mol Physiol. 2001;281:L529–533. doi: 10.1152/ajplung.2001.281.3.L529. [DOI] [PubMed] [Google Scholar]
- 70.Malek AM, Izumo S. Mechanism of endothelial cell shape change and cytoskeletal remodeling in response to fluid shear stress. J Cell Sci. 1996;109(pt 4):713–726. doi: 10.1242/jcs.109.4.713. [DOI] [PubMed] [Google Scholar]
- 71.Chun TH, Itoh H, Ogawa Y, Tamura N, Takaya K, Igaki T, Yamashita J, Doi K, Inoue M, Masatsugu K. Shear stress augments expression of C-type natriuretic peptide and adrenomedullin. Hypertension. 1997;29:1296–1302. doi: 10.1161/01.hyp.29.6.1296. [DOI] [PubMed] [Google Scholar]
- 72.Diacovo TG, Roth SJ, Morita CT, Rosat JP, Brenner MB, Springer TA. Interactions of human alpha/beta and gamma/delta T lymphocyte subsets in shear flow with E-selectin and P-selectin. J Exp Med. 1996;183:1193–1203. doi: 10.1084/jem.183.3.1193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Frangos J, McIntire L, Eskin S. Shear stress induced stimulation of mammalian cell metabolism. Biotechnol Bioeng. 1988;32:1053–1060. doi: 10.1002/bit.260320812. [DOI] [PubMed] [Google Scholar]
- 74.Hung CT, Pollack SR, Reilly TM, Brighton CT. Real-time calcium response of cultured bone cells to fluid flow. Clin Orthop Relat Res. 1995;313:256–269. [PubMed] [Google Scholar]
- 75.Kuijper P, Torres HG, Van Der Linden J, Lammers J, Sixma J, Koenderman L, Zwaginga J. Platelet-dependent primary hemostasis promotes selectin-and integrin-mediated neutrophil adhesion to damaged endothelium under flow conditions. Blood. 1996;87:3271–3281. [PubMed] [Google Scholar]
- 76.Levesque M, Nerem R. The elongation and orientation of cultured endothelial cells in response to shear stress. J Biomech Eng. 1985;107:341–347. doi: 10.1115/1.3138567. [DOI] [PubMed] [Google Scholar]
- 77.Tseng H, Peterson TE, Berk BC. Fluid shear stress stimulates mitogen-activated protein kinase in endothelial cells. Circ Res. 1995;77:869–878. doi: 10.1161/01.res.77.5.869. [DOI] [PubMed] [Google Scholar]
- 78.Song JW, Gu W, Futai N, Warner KA, Nor JE, Takayama S. Computer-controlled microcirculatory support system for endothelial cell culture and shearing. Anal Chemistry. 2005;77:3993–3999. doi: 10.1021/ac050131o. [DOI] [PubMed] [Google Scholar]
- 79.Lu H, Koo LY, Wang WM, Lauffenburger DA, Griffith LG, Jensen KF. Microfluidic shear devices for quantitative analysis of cell adhesion. Anal Chemistry. 2004;76:5257–5264. doi: 10.1021/ac049837t. [DOI] [PubMed] [Google Scholar]
- 80.Dewey C., Jr Effects of fluid flow on living vascular cells. J Biomech Eng. 1984;106:31–35. doi: 10.1115/1.3138453. [DOI] [PubMed] [Google Scholar]
- 81.Furukawa KS, Ushida T, Nagase T, Nakamigawa H, Noguchi T, Tamaki T, Tanaka J, Tateishi T. Quantitative analysis of cell detachment by shear stress. Mater Sci Eng C. 2001;17:55–58. [Google Scholar]
- 82.Hermann C, Zeiher AM, Dimmeler S. Shear stress inhibits H2O2-induced apoptosis of human endothelial cells by modulation of the glutathione redox cycle and nitric oxide synthase. Arterioscler Thromb Vasc Biol. 1997;17:3588–3592. doi: 10.1161/01.atv.17.12.3588. [DOI] [PubMed] [Google Scholar]
- 83.Mohtai M, Gupta M, Donlon B, Ellison B, Cooke J, Gibbons G, Schurman D, Smith RL. Expression of interleukin-6 in osteoarthritic chondrocytes and effects of fluid-induced shear on this expression in normal human chondrocytes in vitro. J Orthop Res. 1996;14:67–73. doi: 10.1002/jor.1100140112. [DOI] [PubMed] [Google Scholar]
- 84.van Grondelle Av, Worthen GS, Ellis D, Mathias MM, Murphy RC, Strife RJ, Reeves JT, Voelkel NF. Altering hydrodynamic variables influences PGI2 production by isolated lungs and endothelial cells. J Appl Physiol. 1984;57:388–395. doi: 10.1152/jappl.1984.57.2.388. [DOI] [PubMed] [Google Scholar]
- 85.Haidekker MA, White CR, Frangos JA. Analysis of temporal shear stress gradients during the onset phase of flow over a backward-facing step. J Biomech Eng. 2001;123:455–463. doi: 10.1115/1.1389460. [DOI] [PubMed] [Google Scholar]
- 86.White CR, Stevens HY, Haidekker M, Frangos JA. Temporal gradients in shear but not spatial gradients stimulate erk1/2 activation in human endothelial cells. Am J Physiol-Heart C. 2005;289:H2350–H2355. doi: 10.1152/ajpheart.01229.2004. [DOI] [PubMed] [Google Scholar]
- 87.Osborn EA, Rabodzey A, Dewey CF, Jr, Hartwig JH. Endothelial actin cytoskeleton remodeling during mechanostimulation with fluid shear stress. Am J Physiol Cell Physiol. 2006;290:C444–452. doi: 10.1152/ajpcell.00218.2005. [DOI] [PubMed] [Google Scholar]
- 88.Kadohama T, Nishimura K, Hoshino Y, Sasajima T, Sumpio BE. Effects of different types of fluid shear stress on endothelial cell proliferation and survival. J Cell Physiol. 2007;212:244–251. doi: 10.1002/jcp.21024. [DOI] [PubMed] [Google Scholar]
- 89.Tzima E, Irani-Tehrani M, Kiosses WB, Dejana E, Schultz DA, Engelhardt B, Cao GY, DeLisser H, Schwartz MA. A mechanosensory complex that mediates the endothelial cell response to fluid shear stress. Nature. 2005;437:426–431. doi: 10.1038/nature03952. [DOI] [PubMed] [Google Scholar]
- 90.Conway DE, Breckenridge MT, Hinde E, Gratton E, Chen CS, Schwartz MA. Fluid shear stress on endothelial cells modulates mechanical tension across VE-cadherin and PECAM-1. Curr Biol. 2013;23:1024–1030. doi: 10.1016/j.cub.2013.04.049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Conway DE, Schwartz MA. Mechanotransduction of shear stress occurs through changes in VE-cadherin and PECAM-1 tension: implications for cell migration. Cell Adhesion Migr. 2015;9:335–339. doi: 10.4161/19336918.2014.968498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Bazellieres E, Conte V, Elosegui-Artola A, Serra-Picamal X, Bintanel-Morcillo M, Roca-Cusachs P, Munoz JJ, Sales-Pardo M, Guimera R, Trepat X. Control of cell–cell forces and collective cell dynamics by the intercellular adhesome. Nature Cell Biol. 2015;17:409–420. doi: 10.1038/ncb3135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Dembo M, Wang YL. Stresses at the cell-to-substrate interface during locomotion of fibroblasts. Biophys J. 1999;76:2307–2316. doi: 10.1016/S0006-3495(99)77386-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Franck C, Hong S, Maskarinec SA, Tirrell DA, Ravichandran G. Three-dimensional full-field measurements of large deformations in soft materials using confocal microscopy and digital volume correlation. Exp Mech. 2007;47:427–438. [Google Scholar]
- 95.Han SJ, Oak Y, Groisman A, Danuser G. Traction microscopy to identify force modulation in subresolution adhesions. Nature Methods. 2015;12:653–656. doi: 10.1038/nmeth.3430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Schwarz US, Balaban NQ, Riveline D, Bershadsky A, Geiger B, Safran SA. Calculation of forces at focal adhesions from elastic substrate data: The effect of localized force and the need for regularization. Biophys J. 2002;83:1380–1394. doi: 10.1016/S0006-3495(02)73909-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Schwarz US, Soine JRD. Traction force microscopy on soft elastic substrates: A guide to recent computational advances. Bba-Mol Cell Res. 2015;1853:3095–3104. doi: 10.1016/j.bbamcr.2015.05.028. [DOI] [PubMed] [Google Scholar]
- 98.Steinwachs J, Metzner C, Skodzek K, Lang N, Thievessen I, Mark C, Munster S, Aifantis KE, Fabry B. Three-dimensional force microscopy of cells in biopolymer networks. Nature Methods. 2016;13:171–176. doi: 10.1038/nmeth.3685. [DOI] [PubMed] [Google Scholar]
- 99.Franck C, Maskarinec SA, Tirrell DA, Ravichandran G. Three-dimensional traction force microscopy: a new tool for quantifying cell–matrix interactions. PLoS One. 2011;6:e17833. doi: 10.1371/journal.pone.0017833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Maskarinec SA, Franck C, Tirrell DA, Ravichandran G. Quantifying cellular traction forces in three dimensions. Proc Natl Acad Sci USA. 2009;106:22108–22113. doi: 10.1073/pnas.0904565106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Legant WR, Miller JS, Blakely BL, Cohen DM, Genin GM, Chen CS. Measurement of mechanical tractions exerted by cells in three-dimensional matrices. Nature Methods. 2010;7:969–U113. doi: 10.1038/nmeth.1531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Mertz AF, Che Y, Banerjee S, Goldstein JM, Rosowski KA, Revilla SF, Niessen CM, Marchetti MC, Dufresne ER, Horsley V. Cadherin-based intercellular adhesions organize epithelial cell–matrix traction forces. Proc Natl Acad Sci USA. 2013;110:842–847. doi: 10.1073/pnas.1217279110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Reinhart-King CA, Dembo M, Hammer DA. Cell-cell mechanical communication through compliant substrates. Biophys J. 2008;95:6044–6051. doi: 10.1529/biophysj.107.127662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Sim JY, Moeller J, Hart KC, Ramallo D, Vogel V, Dunn AR, Nelson WJ, Pruitt BL. Spatial distribution of cell–cell and cell–ECM adhesions regulates force balance while maintaining E-cadherin molecular tension in cell pairs. Mol Biol Cell. 2015;26:2456–2465. doi: 10.1091/mbc.E14-12-1618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Yamada S, Nelson WJ. Localized zones of Rho and Rac activities drive initiation and expansion of epithelial cell–cell adhesion. J Cell Biol. 2007;178:517–527. doi: 10.1083/jcb.200701058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Tseng Q, Duchemin-Pelletier E, Deshiere A, Balland M, Guillou H, Filhol O, Thery M. Spatial organization of the extracellular matrix regulates cell–cell junction positioning. Proc Natl Acad Sci USA. 2012;109:1506–1511. doi: 10.1073/pnas.1106377109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Martinez-Rico C, Pincet F, Thiery JP, Dufour S. Integrins stimulate e-cadherin-mediated intercellular adhesion by regulating src-kinase activation and actomyosin contractility. J Cell Sci. 2010;123:712–722. doi: 10.1242/jcs.047878. [DOI] [PubMed] [Google Scholar]
- 108.Weber GF, Bjerke MA, DeSimone DW. Integrins and cadherins join forces to form adhesive networks. J Cell Sci. 2011;124:1183–1193. doi: 10.1242/jcs.064618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Maruthamuthu V, Sabass B, Schwarz US, Gardel ML. Cell-ECM traction force modulates endogenous tension at cell–cell contacts. Proc Natl Acad Sci USA. 2011;108:4708–4713. doi: 10.1073/pnas.1011123108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.McCain ML, Lee H, Aratyn-Schaus Y, Kleber AG, Parker KK. Cooperative coupling of cell–matrix and cell–cell adhesions in cardiac muscle. Proc Natl Acad Sci USA. 2012;109:9881–9886. doi: 10.1073/pnas.1203007109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Sim JY, Moeller J, Hart KC, Ramallo D, Vogel V, Dunn AR, Nelson WJ, Pruitt BL. Spatial distribution of cell–cell and cell–ECM adhesions regulates force balance while maintaining E-cadherin molecular tension in cell pairs. Mol Biol Cell. 2015;26:2456–2465. doi: 10.1091/mbc.E14-12-1618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Meili R, Alonso-Latorre B, del Alamo JC, Firtel RA, Lasheras JC. Myosin II is essential for the spatiotemporal organization of traction forces during cell motility. Mol Biol Cell. 2010;21:405–417. doi: 10.1091/mbc.E09-08-0703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Tan JL, Tien J, Pirone DM, Gray DS, Bhadriraju K, Chen CS. Cells lying on a bed of microneedles: an approach to isolate mechanical force. Proc Natl Acad Sci. 2003;100:1484–1489. doi: 10.1073/pnas.0235407100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Liu Z, Tan JL, Cohen DM, Yang MT, Sniadecki NJ, Ruiz SA, Nelson CM, Chen CS. Mechanical tugging force regulates the size of cell–cell junctions. Proc Natl Acad Sci USA. 2010;107:9944–9949. doi: 10.1073/pnas.0914547107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Shao Y, Mann JM, Chen W, Fu J. Global architecture of the F-actin cytoskeleton regulates cell shape-dependent endothelial mechanotransduction. Integr Biol (Camb) 2014;6:300–311. doi: 10.1039/c3ib40223a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Biswas KH, Hartman KL, Zaidel-Bar R, Groves JT. Sustained alpha-catenin activation at e-cadherin junctions in the absence of mechanical force. Biophys J. 2016;111:1044–1052. doi: 10.1016/j.bpj.2016.06.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Borghi N, Lowndes M, Maruthamuthu V, Gardel ML, Nelson WJ. Regulation of cell motile behavior by crosstalk between cadherin-and integrin-mediated adhesions. Proc Natl Acad Sci. 2010;107:13324–13329. doi: 10.1073/pnas.1002662107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Rand R, Burton A. Mechanical properties of the red cell membrane: I. Membrane stiffness and intracellular pressure. Biophys J. 1964;4:115–135. doi: 10.1016/s0006-3495(64)86773-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Henriksen JR, Ipsen JH. Measurement of membrane elasticity by micro-pipette aspiration. Eur Phys J E. 2004;14:149–167. doi: 10.1140/epje/i2003-10146-y. [DOI] [PubMed] [Google Scholar]
- 120.Bo L, Waugh RE. Determination of bilayer-membrane bending stiffness by tether formation from giant. Thin-Walled Vesicles Biophys J. 1989;55:509–517. doi: 10.1016/S0006-3495(89)82844-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Hochmuth R, Ting-Beall H, Beaty B, Needham D, Tran-Son-Tay R. Viscosity of passive human neutrophils undergoing small deformations. Biophys J. 1993;64:1596–1601. doi: 10.1016/S0006-3495(93)81530-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Sato M, Levesque MJ, Nerem RM. Micropipette aspiration of cultured bovine aortic endothelial cells exposed to shear stress. Arterioscl Thromb Vasc Biol. 1987;7:276–286. doi: 10.1161/01.atv.7.3.276. [DOI] [PubMed] [Google Scholar]
- 123.Evans E, Yeung A. Apparent viscosity and cortical tension of blood granulocytes determined by micropipet aspiration. Biophys J. 1989;56:151–160. doi: 10.1016/S0006-3495(89)82660-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Vaziri A, Mofrad MR. Mechanics and deformation of the nucleus in micropipette aspiration experiment. J Biomech. 2007;40:2053–2062. doi: 10.1016/j.jbiomech.2006.09.023. [DOI] [PubMed] [Google Scholar]
- 125.Ricca BL, Venugopalan G, Fletcher DA. To pull or be pulled: parsing the multiple modes of mechanotransduction. Curr Opin Cell Biol. 2013;25:558–564. doi: 10.1016/j.ceb.2013.06.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Mitrossilis D, Fouchard J, Guiroy A, Desprat N, Rodriguez N, Fabry B, Asnacios A. Single-cell response to stiffness exhibits muscle-like behavior. Proc Natl Acad Sci. 2009;106:18243–18248. doi: 10.1073/pnas.0903994106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Secomb T. Red blood cell mechanics and capillary blood rheology. Cell Biochem Biophys. 1991;18:231–251. doi: 10.1007/BF02989816. [DOI] [PubMed] [Google Scholar]
- 128.Secomb T, Hsu R. Red blood cell mechanics and functional capillary density. Int J Microcirc. 1995;15:250–254. doi: 10.1159/000179026. [DOI] [PubMed] [Google Scholar]
- 129.Wu J, Herzog W, Epstein M. Modelling of location-and time-dependent deformation of chondrocytes during cartilage loading. J Biomech. 1999;32:563–572. doi: 10.1016/s0021-9290(99)00034-2. [DOI] [PubMed] [Google Scholar]
- 130.Zhou EH, Lim CT, Quek ST. Finite element simulation of the micropipette aspiration of a living cell undergoing large viscoelastic deformation. Mech Adv Mater Struct. 2005;12:501–512. [Google Scholar]
- 131.Sung KLP, Sung LA, Crimmins M, Burakoff SJ, Chien S. Determination of junction avidity of cytolytic T cell and target cell. Science. 1986;234:1405–1409. doi: 10.1126/science.3491426. [DOI] [PubMed] [Google Scholar]
- 132.Sung KP, Saldivar E, Phillips L. Interleukin-1β induces differential adhesiveness on human endothelial cell surfaces. Biochem Biophys Res Commun. 1994;202:866–872. doi: 10.1006/bbrc.1994.2010. [DOI] [PubMed] [Google Scholar]
- 133.Tabdili H, Langer M, Shi QM, Poh YC, Wang N, Leckband D. Cadherin-dependent mechanotransduction depends on ligand identity but not affinity. J Cell Sci. 2012;125:4362–4371. doi: 10.1242/jcs.105775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Reynolds NH, Ronan W, Dowling EP, Owens P, McMeeking RM, McGarry JP. On the role of the actin cytoskeleton and nucleus in the biomechanical response of spread cells. Biomaterials. 2014;35:4015–4025. doi: 10.1016/j.biomaterials.2014.01.056. [DOI] [PubMed] [Google Scholar]
- 135.Ashkin A, Dziedzic JM. Optical trapping and manipulation of viruses and bacteria. Science. 1987;235:1517–1521. doi: 10.1126/science.3547653. [DOI] [PubMed] [Google Scholar]
- 136.Mehta AD, Rief M, Spudich JA, Smith DA, Simmons RM. Single-molecule biomechanics with optical methods. Science. 1999;283:1689–1695. doi: 10.1126/science.283.5408.1689. [DOI] [PubMed] [Google Scholar]
- 137.Wen JD, Manosas M, Li PT, Smith SB, Bustamante C, Ritort F, Tinoco I. Force unfolding kinetics of RNA using optical tweezers. I. Effects of experimental variables on measured results. Biophys J. 2007;92:2996–3009. doi: 10.1529/biophysj.106.094052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Mills J, Qie L, Dao M, Lim C, Suresh S. Nonlinear elastic and viscoelastic deformation of the human red blood cell with optical tweezers. MCB-Tech Sci Press. 2004;1:169–180. [PubMed] [Google Scholar]
- 139.Svoboda K, Block SM. Biological applications of optical forces. Annu Rev Biophys Biomol Struct. 1994;23:247–285. doi: 10.1146/annurev.bb.23.060194.001335. [DOI] [PubMed] [Google Scholar]
- 140.Sleep J, Wilson D, Simmons R, Gratzer W. Elasticity of the red cell membrane and its relation to hemolytic disorders: an optical tweezers study. Biophys J. 1999;77:3085–3095. doi: 10.1016/S0006-3495(99)77139-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Dao M, Lim CT, Suresh S. Mechanics of the human red blood cell deformed by optical tweezers. J Mech Phys Solids. 2003;51:2259–2280. [Google Scholar]
- 142.Suresh S, Spatz J, Mills J, Micoulet A, Dao M, Lim C, Beil M, Seufferlein T. Reprint of: connections between single-cell biomechanics and human disease states: gastrointestinal cancer and malaria. Acta Biomaterialia. 2015;23:S3–S15. doi: 10.1016/j.actbio.2015.07.015. [DOI] [PubMed] [Google Scholar]
- 143.Foillard S, Dumy P, Boturyn D. Highly efficient cell adhesion on beads functionalized with clustered peptide ligands. Org Biomol Chem. 2009;7:4159–4162. doi: 10.1039/b911440h. [DOI] [PubMed] [Google Scholar]
- 144.Schwingel M, Bastmeyer M. Force mapping during the formation and maturation of cell adhesion sites with multiple optical tweezers. PLoS One. 2013;8:e54850. doi: 10.1371/journal.pone.0054850. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Buckley CD, Tan J, Anderson KL, Hanein D, Volkmann N, Weis WI, Nelson WJ, Dunn AR. Cell adhesion. The minimal cadherin-catenin complex binds to actin filaments under force. Science. 2014;346:1254211. doi: 10.1126/science.1254211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Guck J, Ananthakrishnan R, Cunningham CC, Käs J. Stretching biological cells with light. J Phys: Condens Matter. 2002;14:4843. [Google Scholar]
- 147.Guck J, Ananthakrishnan R, Mahmood H, Moon TJ, Cunningham CC, Käs J. The optical stretcher: a novel laser tool to micromanipulate cells. Biophys J. 2001;81:767–784. doi: 10.1016/S0006-3495(01)75740-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Guck J, Chiang J, Kas J. Proc Mol Biol Cell. Amer Soc Cell Biology Publ Office; 9650 Rockville Pike, Bethesda, MD 20814 USA: The optical stretcher ua novel tool to characterize the cytoskeleton; pp. 105A–105A. [Google Scholar]
- 149.Bambardekar K, Clement R, Blanc O, Chardes C, Lenne PF. Direct laser manipulation reveals the mechanics of cell contacts in vivo. Proc Natl Acad Sci USA. 2015;112:1416–1421. doi: 10.1073/pnas.1418732112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Crick F, Hughes A. The physical properties of cytoplasm: A study by means of the magnetic particle method Part I. Experimental. Exp Cell Res. 1950;1:37–80. [Google Scholar]
- 151.Puig-De-Morales M, Grabulosa M, Alcaraz J, Mullol J, Maksym GN, Fredberg JJ, Navajas D. Measurement of cell microrheology by magnetic twisting cytometry with frequency domain demodulation. J Appl Physiol. 2001;91:1152–1159. doi: 10.1152/jappl.2001.91.3.1152. [DOI] [PubMed] [Google Scholar]
- 152.Fabry B, Maksym GN, Hubmayr RD, Butler JP, Fredberg JJ. Implications of heterogeneous bead behavior on cell mechanical properties measured with magnetic twisting cytometry. J Magn Magn Mater. 1999;194:120–125. [Google Scholar]
- 153.Wang N, Butler JP, Ingber DE. Mechanotransduction across the cell-surface and through the cytoskeleton. Science. 1993;260:1124–1127. doi: 10.1126/science.7684161. [DOI] [PubMed] [Google Scholar]
- 154.Wang N, Ingber DE. Probing transmembrane mechanical coupling and cytomechanics using magnetic twisting cytometry. Biochem Cell Biol. 1995;73:327–335. doi: 10.1139/o95-041. [DOI] [PubMed] [Google Scholar]
- 155.Tajik A, Zhang Y, Wei F, Sun J, Jia Q, Zhou W, Singh R, Khanna N, Belmont AS, Wang N. Transcription upregulation via force-induced direct stretching of chromatin. Nature Mater. 2016;15:1287–1296. doi: 10.1038/nmat4729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156.Tseng Y, Kole TP, Wirtz D. Micromechanical mapping of live cells by multiple-particle-tracking microrheology. Biophys J. 2002;83:3162–3176. doi: 10.1016/S0006-3495(02)75319-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157.Muhamed I, Wu J, Sehgal P, Kong X, Tajik A, Wang N, Leckband DE. E-cadherin-mediated force transduction signals regulate global cell mechanics. J Cell Sci. 2016;129:1843–1854. doi: 10.1242/jcs.185447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Barry AK, Tabdili H, Muhamed I, Wu J, Shashikanth N, Gomez GA, Yap AS, Gottardi CJ, de Rooij J, Wang N, Leckband DE. alpha-catenin cytomechanics–role in cadherin-dependent adhesion and mechanotransduction. J Cell Sci. 2014;127:1779–1791. doi: 10.1242/jcs.139014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 159.Charras G, Lehenkari PP, Horton M. Atomic force microscopy can be used to mechanically stimulate osteoblasts and evaluate cellular strain distributions. Ultramicroscopy. 2001;86:85–95. doi: 10.1016/s0304-3991(00)00076-0. [DOI] [PubMed] [Google Scholar]
- 160.Radmacher M. 4.-Measuring the elastic properties of living cells by the atomic force microscope. Methods Cell Biol. 2002;68:67–90. doi: 10.1016/s0091-679x(02)68005-7. [DOI] [PubMed] [Google Scholar]
- 161.Fung CKM, Seiffert-Sinha K, Lai KWC, Yang R, Panyard D, Zhang J, Xi N, Sinha AA. Investigation of human keratinocyte cell adhesion using atomic force microscopy. Nanomedicine: Nanotechnol Biol Med. 2010;6:191–200. doi: 10.1016/j.nano.2009.05.008. [DOI] [PubMed] [Google Scholar]
- 162.Yang R, Song B, Sun Z, Lai KWC, Fung CKM, Patterson KC, Seiffert-Sinha K, Sinha AA, Xi N. Cellular level robotic surgery: Nanodissection of intermediate filaments in live keratinocytes. Nanomedicine: Nanotechnol Biol Med. 2015;11:137–145. doi: 10.1016/j.nano.2014.08.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163.Seiffert-Sinha K, Yang R, Fung CK, Lai KW, Patterson KC, Payne AS, Xi N, Sinha AA. Nanorobotic investigation identifies novel visual, structural and functional correlates of autoimmune pathology in a blistering skin disease model. PLoS One. 2014;9:e106895. doi: 10.1371/journal.pone.0106895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164.Mahaffy R, Park S, Gerde E, Käs J, Shih C. Quantitative analysis of the viscoelastic properties of thin regions of fibroblasts using atomic force microscopy. Biophys J. 2004;86:1777–1793. doi: 10.1016/S0006-3495(04)74245-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 165.Vaziri A, Lee H, Mofrad MK. Deformation of the cell nucleus under indentation: mechanics and mechanisms. J Mater Res. 2006;21:2126–2135. [Google Scholar]
- 166.Benoit M. 5.-Cell adhesion measured by force spectroscopy on living cells. Methods Cell Biol. 2002;68:91–114. doi: 10.1016/s0091-679x(02)68006-9. [DOI] [PubMed] [Google Scholar]
- 167.Hinterdorfer P. 6.-Molecular recognition studies using the atomic force microscope. Methods Cell Biol. 2002;68:115–141. doi: 10.1016/s0091-679x(02)68007-0. [DOI] [PubMed] [Google Scholar]
- 168.Willemsen OH, Snel MM, Cambi A, Greve J, De Grooth BG, Figdor CG. Biomolecular interactions measured by atomic force microscopy. Biophys J. 2000;79:3267–3281. doi: 10.1016/S0006-3495(00)76559-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 169.Marshall BT, Long M, Piper JW, Yago T. Direct observation of catch bonds involving cell-adhesion molecules. Nature. 2003;423:190. doi: 10.1038/nature01605. [DOI] [PubMed] [Google Scholar]
- 170.Benoit M, Gabriel D, Gerisch G, Gaub HE. Discrete interactions in cell adhesion measured by single-molecule force spectroscopy. Nature Cell Biol. 2000;2:313–317. doi: 10.1038/35014000. [DOI] [PubMed] [Google Scholar]
- 171.Friedrichs J, Legate KR, Schubert R, Bharadwaj M, Werner C, Mullner DJ, Benoit M. A practical guide to quantify cell adhesion using single-cell force spectroscopy. Methods. 2013;60:169–178. doi: 10.1016/j.ymeth.2013.01.006. [DOI] [PubMed] [Google Scholar]
- 172.Puech PH, Poole K, Knebel D, Muller DJ. A new technical approach to quantify cell–cell adhesion forces by AFM. Ultramicroscopy. 2006;106:637–644. doi: 10.1016/j.ultramic.2005.08.003. [DOI] [PubMed] [Google Scholar]
- 173.Krieg M, Arboleda-Estudillo Y, Puech PH, Kafer J, Graner F, Muller DJ, Heisenberg CP. Tensile forces govern germ-layer organization in zebrafish. Nature Cell Biol. 2008;10:429–U122. doi: 10.1038/ncb1705. [DOI] [PubMed] [Google Scholar]
- 174.Heisterkamp A, Maxwell IZ, Mazur E, Underwood JM, Nickerson JA, Kumar S, Ingber DE. Pulse energy dependence of subcellular dissection by femtosecond laser pulses. Opt Express. 2005;13:3690–3696. doi: 10.1364/opex.13.003690. [DOI] [PubMed] [Google Scholar]
- 175.Kumar S, Maxwell IZ, Heisterkamp A, Polte TR, Lele TP, Salanga M, Mazur E, Ingber DE. Viscoelastic retraction of single living stress fibers and its impact on cell shape, cytoskeletal organization, and extracellular matrix mechanics. Biophys J. 2006;90:3762–3773. doi: 10.1529/biophysj.105.071506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 176.Brugues A, Anon E, Conte V, Veldhuis JH, Gupta M, Colombelli J, Munoz JJ, Brodland GW, Ladoux B, Trepat X. Forces driving epithelial wound healing. Nature Phys. 2014;10:683–690. doi: 10.1038/nphys3040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177.Ingber DE, Tensegrity I. Cell structure and hierarchical systems biology. J Cell Sci. 2003;116:1157–1173. doi: 10.1242/jcs.00359. [DOI] [PubMed] [Google Scholar]
- 178.Vielmuth F, Hartlieb E, Kugelmann D, Waschke J, Spindler V. Atomic force microscopy identifies regions of distinct desmoglein 3 adhesive properties on living keratinocytes. Nanomed-Nanotechnol. 2015;11:511–520. doi: 10.1016/j.nano.2014.10.006. [DOI] [PubMed] [Google Scholar]
- 179.Vielmuth F, Waschke J, Spindler V. Loss of desmoglein binding is not sufficient for keratinocyte dissociation in pemphigus. J Invest Dermatol. 2015;135:3068–3077. doi: 10.1038/jid.2015.324. [DOI] [PubMed] [Google Scholar]
- 180.Lowndes M, Rakshit S, Shafraz O, Borghi N, Harmon RM, Green KJ, Sivasankar S, Nelson WJ. Different roles of cadherins in the assembly and structural integrity of the desmosome complex. J Cell Sci. 2014;127:2339–2350. doi: 10.1242/jcs.146316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181.Rakshit S, Zhang YX, Manibog K, Shafraz O, Sivasankar S. Ideal, catch, and slip bonds in cadherin adhesion. Proc Natl Acad Sci USA. 2012;109:18815–18820. doi: 10.1073/pnas.1208349109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 182.Manibog K, Li H, Rakshit S, Sivasankar S. Resolving the molecular mechanism of cadherin catch bond formation. Nature Commun. 2014;5 doi: 10.1038/ncomms4941. [DOI] [PubMed] [Google Scholar]
- 183.Liang X, Michael M, Gomez GA. Measurement of mechanical tension at cell–cell junctions using two-photon laser ablation. Bio-Protocol. 2016;6:e2068. doi: 10.21769/BioProtoc.2068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 184.Gomez GA, McLachlan RW, Wu SK, Caldwell BJ, Moussa E, Verma S, Bastiani M, Priya R, Parton RG, Gaus K, Sap J, Yap AS. An RPTPα/Src family kinase/Rap1 signaling module recruits myosin IIB to support contractile tension at apical E-cadherin junctions. Mol Biol Cell. 2015;26:1249–1262. doi: 10.1091/mbc.E14-07-1223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 185.Wu SK, Gomez GA, Michael M, Verma S, Cox HL, Lefevre JG, Parton RG, Hamilton NA, Neufeld Z, Yap AS. Cortical F-actin stabilization generates apical–lateral patterns of junctional contractility that integrate cells into epithelia. Nature Cell Biol. 2014;16:167–178. doi: 10.1038/ncb2900. [DOI] [PubMed] [Google Scholar]
- 186.Ratheesh A, Gomez GA, Priya R, Verma S, Kovacs EM, Jiang K, Brown NH, Akhmanova A, Stehbens SJ, Yap AS. Centralspindlin and [alpha]-catenin regulate rho signalling at the epithelial zonula adherens. Nature Cell Biol. 2012;14:818–828. doi: 10.1038/ncb2532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 187.Priya R, Gomez GA, Budnar S, Verma S, Cox HL, Hamilton NA, Yap AS. Feedback regulation through myosin II confers robustness on RhoA signalling at E-cadherin junctions. Nature Cell Biol. 2015;17:1282–1293. doi: 10.1038/ncb3239. [DOI] [PubMed] [Google Scholar]
- 188.Smutny M, Behrndt M, Campinho P, Ruprecht V, Heisenberg C-P. Uv laser ablation to measure cell and tissue-generated forces in the zebrafish embryo in vivo and ex vivo. In: Nelson CM, editor. Tissue Morphogenesis: Methods and Protocols. Springer; New York, New York, NY: 2015. pp. 219–235. [DOI] [PubMed] [Google Scholar]
- 189.Loh O, Vaziri A, Espinosa HDSM. The potential of MEMS for advancing experiments and modeling in cell mechanics. Exp Mech. 2009;49:105–124. [Google Scholar]
- 190.Eppell S, Smith B, Kahn H, Ballarini R. Nano measurements with microdevices: mechanical properties of hydrated collagen fibrils. J R Soc Interface. 2006;3:117–121. doi: 10.1098/rsif.2005.0100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191.Mukundan V, Pruitt BL. Mems electrostatic actuation in conducting biological media. J Microelectromech Syst. 2009;18:405–413. doi: 10.1109/JMEMS.2009.2013398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 192.Mukundan V, Ponce P, Butterfield HE, Pruitt BL. Modeling and characterization of electrostatic comb-drive actuators in conducting liquid media. J Micromech Microeng. 2009;19 doi: 10.1088/0960-1317/19/6/065008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 193.Panorchan P, Thompson MS, Davis KJ, Tseng Y, Konstantopoulos K, Wirtz D. Single-molecule analysis of cadherin-mediated cell–cell adhesion. J Cell Sci. 2006;119:66–74. doi: 10.1242/jcs.02719. [DOI] [PubMed] [Google Scholar]
- 194.Berfield TA, Patel HK, Shimmin RG, Braun PV, Lambros J, Sottos NR. Fluorescent image correlation for nanoscale deformation measurements. Small. 2006;2:631–635. doi: 10.1002/smll.200500289. [DOI] [PubMed] [Google Scholar]
- 195.Greiner AM, Richter B, Bastmeyer M. Micro-engineered 3D scaffolds for cell culture studies. Macromol Biosci. 2012;12:1301–1314. doi: 10.1002/mabi.201200132. [DOI] [PubMed] [Google Scholar]
- 196.Greiner AM, Jackel M, Scheiwe AC, Stamow DR, Autenrieth TJ, Lahann J, Franz CM, Bastmeyer M. Multifunctional polymer scaffolds with adjustable pore size and chemoattractant gradients for studying cell matrix invasion. Biomaterials. 2014;35:611–619. doi: 10.1016/j.biomaterials.2013.09.095. [DOI] [PubMed] [Google Scholar]
- 197.Klein F, Richter B, Striebel T, Franz CM, von Freymann G, Wegener M, Bastmeyer M. Two-component polymer scaffolds for controlled three-dimensional cell culture. Adv Mater. 2011;23:1341–1345. doi: 10.1002/adma.201004060. [DOI] [PubMed] [Google Scholar]
- 198.Richter B, Hahn V, Bertels S, Claus TK, Wegener M, Delaittre G, Barner-Kowollik C, Bastmeyer M. Guiding Cell Attachment in 3D Microscaffolds Selectively Functionalized with Two Distinct Adhesion Proteins. Adv Mater. 2017;29 doi: 10.1002/adma.201604342. [DOI] [PubMed] [Google Scholar]
- 199.Richter B, Pauloehrl T, Kaschke J, Fichtner D, Fischer J, Greiner AM, Wedlich D, Wegener M, Delaittre G, Barner-Kowollik C, Bastmeyer M. Three-dimensional microscaffolds exhibiting spatially resolved surface chemistry. Adv Mater. 2013;25:6117–6122. doi: 10.1002/adma.201302678. [DOI] [PubMed] [Google Scholar]
- 200.Klein F, Striebel T, Fischer J, Jiang Z, Franz CM, von Freymann G, Wegener M, Bastmeyer M. Elastic fully three-dimensional microstructure scaffolds for cell force measurements. Adv Mater. 2010;22:868–871. doi: 10.1002/adma.200902515. [DOI] [PubMed] [Google Scholar]
- 201.Scheiwe AC, Frank SC, Autenrieth TJ, Bastmeyer M, Wegener M. Subcellular stretch-induced cytoskeletal response of single fibroblasts within 3D designer scaffolds. Biomaterials. 2015;44:186–194. doi: 10.1016/j.biomaterials.2014.12.018. [DOI] [PubMed] [Google Scholar]
- 202.Austen K, Ringer P, Mehlich A, Chrostek-Grashoff A, Kluger C, Klingner C, Sabass B, Zent R, Rief M, Grashoff C. Extracellular rigidity sensing by talin isoform-specific mechanical linkages. Nature Cell Biol. 2015;17:1597–1606. doi: 10.1038/ncb3268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 203.Cost AL, Ringer P, Chrostek-Grashoff A, Grashoff C. How to measure molecular forces in cells: A guide to evaluating genetically-encoded FRET-based tension sensors. Cell Mol Bioeng. 2015;8:96–105. doi: 10.1007/s12195-014-0368-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 204.Jurchenko C, Salaita KS. Lighting up the force: Investigating mechanisms of mechanotransduction using fluorescent tension probes. Mol Cell Biol. 2015;35:2570–2582. doi: 10.1128/MCB.00195-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 205.Kong HJ, Polte TR, Alsberg E, Mooney DJ. FRET measurements of cell-traction forces and nano-scale clustering of adhesion ligands varied by substrate stiffness. Proc Natl Acad Sci USA. 2005;102:4300–4305. doi: 10.1073/pnas.0405873102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 206.Berney C, Danuser G. FRET or no FRET: a quantitative comparison. Biophys J. 2003;84:3992–4010. doi: 10.1016/S0006-3495(03)75126-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 207.Pertz O, Hodgson L, Klemke RL, Hahn KM. Spatiotemporal dynamics of RhoA activity in migrating cells. Nature. 2006;440:1069–1072. doi: 10.1038/nature04665. [DOI] [PubMed] [Google Scholar]
- 208.Machacek M, Hodgson L, Welch C, Elliott H, Pertz O, Nalbant P, Abell A, Johnson GL, Hahn KM, Danuser G. Coordination of Rho GTPase activities during cell protrusion. Nature. 2009;461:99–103. doi: 10.1038/nature08242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 209.Ouyang M, Sun J, Chien S, Wang Y. Determination of hierarchical relationship of Src and Rac at subcellular locations with FRET biosensors. Proc Natl Acad Sci USA. 2008;105:14353–14358. doi: 10.1073/pnas.0807537105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 210.Seong J, Ouyang M, Kim T, Sun J, Wen PC, Lu S, Zhuo Y, Llewellyn NM, Schlaepfer DD, Guan JL, Chien S, Wang Y. Detection of focal adhesion kinase activation at membrane microdomains by fluorescence resonance energy transfer. Nature Commun. 2011;2:406. doi: 10.1038/ncomms1414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 211.Wang Y, Botvinick EL, Zhao Y, Berns MW, Usami S, Tsien RY, Chien S. Visualizing the mechanical activation of Src. Nature. 2005;434:1040–1045. doi: 10.1038/nature03469. [DOI] [PubMed] [Google Scholar]
- 212.Cai D, Chen SC, Prasad M, He L, Wang X, Choesmel-Cadamuro V, Sawyer JK, Danuser G, Montell DJ. Mechanical feedback through E-cadherin promotes direction sensing during collective cell migration. Cell. 2014;157:1146–1159. doi: 10.1016/j.cell.2014.03.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 213.Daneshjou N, Sieracki N, van Nieuw Amerongen GP, Conway DE, Schwartz MA, Komarova YA, Malik AB. Rac1 functions as a reversible tension modulator to stabilize VE-cadherin trans-interaction. J Cell Biol. 2015;208:23–32. doi: 10.1083/jcb.201409108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 214.Kim TJ, Zheng S, Sun J, Muhamed I, Wu J, Lei L, Kong X, Leckband DE, Wang Y. Dynamic visualization of α-catenin reveals rapid, reversible conformation switching between tension states. Curr Biol. 2015;25:218–224. doi: 10.1016/j.cub.2014.11.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 215.Kim TJ, Zheng S, Sun J, Muhamed I, Wu J, Lei L, Kong X, Leckband DE, Wang Y. Dynamic visualization of alpha-catenin reveals rapid, reversible conformation switching between tension states. Curr Biol. 2015;25:218–224. doi: 10.1016/j.cub.2014.11.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 216.Brenner MD, Zhou R, Conway DE, Lanzano L, Gratton E, Schwartz MA, Ha T. Spider silk peptide is a compact, linear nanospring ideal for intracellular tension sensing. Nano Lett. 2016;16:2096–2102. doi: 10.1021/acs.nanolett.6b00305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 217.Verma D, Bajpai VK, Ye N, Maneshi MM, Jetta D, Andreadis ST, Sachs F, Hua SZ. Flow induced adherens junction remodeling driven by cytoskeletal forces. Exp Cell Res. 2017;359:327–336. doi: 10.1016/j.yexcr.2017.08.009. [DOI] [PubMed] [Google Scholar]
- 218.Tornavaca O, Chia M, Dufton N, Almagro LO, Conway DE, Randi AM, Schwartz MA, Matter K, Balda MS. ZO-1 controls endothelial adherens junctions, cell–cell tension, angiogenesis, and barrier formation. J Cell Biol. 2015;208:821–838. doi: 10.1083/jcb.201404140. [DOI] [PMC free article] [PubMed] [Google Scholar]