The CRISPR/Cas system has revolutionized the field of genome-editing as it is easier, faster, less expensive, and more efficient than traditional methods. In a very short time, its applications have become very diverse and widely used. Recently, this technology has been applied in human embryos to correct pathogenic mutations and has re-ignited the ethical debate of germ-line editing1,2. Besides the ethical considerations, the possibility of creating off-target mutations with unknown consequences is a concern.
There are many reports of high-frequency off-target mutations in human and mouse cell-lines3,4, but unwanted errors in mammalian embryo editing have been deemed as a rarity5–9. In a study with human embryo editing, one embryo was profiled for off-target mutations by whole-genome sequencing2. Even though they report no off-target mutations, the number is too small (n = 1) for statistical significance and the on-target locus has 1 mismatch with the sgRNA used. Previous studies with mouse embryos have reported rare and low-frequency mutations at sites with >1 mismatches8,10. Here we report high-frequency mutations at an off-target site with three mismatches using this gene editing technology in the mouse embryo.
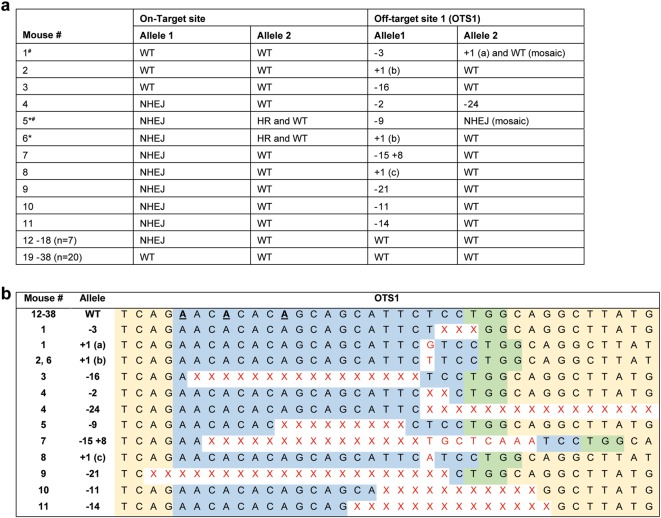
We targeted Dicer1 to introduce a mutation at a Serine residue by pronuclear injection of sgRNA (7.5 ng/µl), spCas9 mRNA (10 ng/µl), and a 123 base single-stranded donor oligo (20 ng/µl) into mouse embryos using previously described methodology11. The sgRNA has a quality score of 76 (out of 100) with 124 potential off-target sites according to the crispr.mit.edu site. In all 38 mice from 200 injected zygotes, we used the polymerase chain reaction to amplify and Sanger sequence a 1 Kbp region of the on-target and the top two candidate off-target sites that contain a proto-spacer adjacent motif (PAM) site. We observed the on-target efficiency of 40% (15/38 animals) with non-homologous end joining (NHEJ) being the predominant repair mechanism (Fig. 1a). Only 5% of the mice (2/38) had the intended mutation introduced by homologous recombination (both were mosaic based on sequencing and validated by genotyping of F1 progeny). Surprisingly, at the top predicted off-target site (intergenic) with 3 mismatches at positions 1, 4, and 8 (where 1 is farthest from the PAM site), we observed mutations in 29% (11/38) of mice (Fig. 1b). Off-target mutations at this site were observed in 3 mice (#1, #2, and #3) that did not have mutations at the on-target site. Additionally, bi-allelic off-target mutations were observed in 2 mice (#4 and #5). At the second candidate off-target site with 4 mismatches at positions 1, 2, 5, and 8 followed by a PAM, sequencing results did not reveal mutations in any of the 38 mice. We further evaluated 3 additional off-target sites in six animals (with on-target mutations) and identified no additional mutations. Since Dicer1 is an essential gene, bi-allelic mutations may result in embryonic lethality. As a result, we may have missed additional on- and off-target mutations.
Fig. 1. On- and off-target changes observed for all 38 mice born from CRISPR/Cas9 targeted embryos.
a Allelic description for all mice at both on- and off-target sites. *, mosaic at on-target site; #, mosaic at off-target site; WT, wild-type; NHEJ, non-homologous end-joining; HR, homologous recombination. b List of all mutations observed at an off-target site with three mismatches. Off-target site is in blue, mismatches are bolded and underlined, PAM is in green, deletions are highlighted as X in red, and insertions are in red letters. Note: For mouse # 4 in (b), the whole deletion is not depicted (to fit the figure)
We have generated six other alleles in the laboratory using six additional sgRNAs by injection of zygotes, and found no other mutations upon screening the top 5 off-target candidates for each sgRNA in all mice with the intended on-target mutation. In contrast to the off-target site with mutations in this study (where all mismatches are outside of the seed region), candidate off-target sites with a PAM for all other sgRNAs –contained ≥3 mismatches with ≥1 mismatches in the seed region. Our results, while rare, provide important and compelling evidence that high-efficiency off-target mutations can occur in mammalian embryos. Our data are consistent with previous observations that this system is more tolerant when mismatches are away from the seed region. Combined, these data underscore the need to identify and screen potential off-target events when engineering genetic models, with an emphasis on sites that have ≤3 mismatches outside of the seed region. Importantly, unlike cell-lines, mice can be backcrossed to remove inter-chromosomal off-target mutations. Finally, this observation needs to be considered as the technology advances for therapeutic purposes.
Electronic supplementary material
List of candidate off-target sites for the selected sgRNA
Acknowledgements
This work was supported by the MD Anderson Cancer Center Support Grant CA16672.
Conflict of interest
The authors declare that they have no conflict of interest.
Footnotes
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Information accompanies this paper at (10.1038/s41419-018-1146-0).
References
- 1.Ma, H. et al. Correction of a pathogenic gene mutation in human embryos. Nature, 548, 413–419 (2017). [DOI] [PubMed]
- 2.Tang L, et al. CRISPR/Cas9-mediated gene editing in human zygotes using Cas9 protein. Mol. Genet. Genom. 2017;292:525–533. doi: 10.1007/s00438-017-1299-z. [DOI] [PubMed] [Google Scholar]
- 3.Hsu PD, et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 2013;31:827–832. doi: 10.1038/nbt.2647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Fu Y, et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat. Biotechnol. 2013;31:822–826. doi: 10.1038/nbt.2623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Iyer V, et al. Off-target mutations are rare in Cas9-modified mice. Nat. Methods. 2015;12:479. doi: 10.1038/nmeth.3408. [DOI] [PubMed] [Google Scholar]
- 6.Nakajima K, et al. Exome sequencing in the knockin mice generated using the CRISPR/Cas system. Sci. Rep. 2016;6:34703. doi: 10.1038/srep34703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Iyer V, et al. No unexpected CRISPR-Cas9 off-target activity revealed by trio sequencing of gene-edited mice. PLoS. Genet. 2018;14:e1007503. doi: 10.1371/journal.pgen.1007503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Yang H, et al. One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell. 2013;154:1370–1379. doi: 10.1016/j.cell.2013.08.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Hay EA, et al. An analysis of possible off target effects following CAS9/CRISPR targeted deletions of neuropeptide gene enhancers from the mouse genome. Neuropeptides. 2017;64:101–107. doi: 10.1016/j.npep.2016.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Shen B, et al. Efficient genome modification by CRISPR-Cas9 nickase with minimal off-target effects. Nat. Methods. 2014;11:399–402. doi: 10.1038/nmeth.2857. [DOI] [PubMed] [Google Scholar]
- 11.Aryal NK, Wasylishen AR, Pant V, Riley-Croce M, Lozano G. Loss of digestive organ expansion factor (Diexf) reveals an essential role during murine embryonic development that is independent of p53. Oncotarget. 2017;8:103996–104006. doi: 10.18632/oncotarget.22087. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
List of candidate off-target sites for the selected sgRNA