Abstract
Vanadate, a protein tyrosine phosphatase inhibitor which elicits insulin-like effects, has previously been shown to inhibit expression of the insulin receptor gene at the transcriptional level in rat hepatoma cells. In an attempt to identify the DNA sequence and transcription factors potentially involved in this effect, a fragment of the proximal 5′flanking region of the IR gene (−1143/−252 upstream the ATG codon) has been cloned and functionally characterized. RNase protection allowed the identification of several transcription start sites in the conserved region of the gene, among which two major sites at −455 and −396. Upon fusion to the luciferase gene and transient transfection into hepatoma cells, the −1143/−252 fragment showed promoter activity. This was unaffected by deletion of the −1143/−761 sequence, but markedly decreased (90%) by additional deletion of the −760/−465 sequence. Treatment of hepatoma cells with vanadate led to a dose-dependent decrease in promoter activity of the 1143/−252, −760/−252 and −464/−252 constructs (change relative to untreated cells, 40, 55 and 23% at 125 μM, and 70, 85 and 62% at 250 μM, respectively). These data suggest that although the entire DNA sequence upstream the transcription start sites is probably involved in vanadate-induced inhibition, the short sequence downstream of position −464 and is sufficient for inhibition. Potential targets of vanadate are the transcription factors FoxO1 and HMGA1, two downstream targets of the insulin signaling pathway which have been shown to mediate the inhibitory effect of insulin on IR gene expression.
Keywords: Insulin receptor, Gene transcription, Vanadate, Rat, Liver, Hepatoma cells
Abbreviations: C/EBPβ, C/CAAT/enhancer binding protein β; FoxO1, Forkhead box protein O1; HMGA1, high mobility group A1 protein; HNF4, hepatocyte nuclear factor 4; IGFBP-1, insulin-like growth factor binding protein 1; IR, insulin receptor; PEPCK, phosphoenolpyruvate carboxykinase; PI3K, phosphatidyl inositol 3-kinase; SINE, short interspersed nuclear element; Sp1, specificity protein 1; STZ, streptozotocin; TCF7L2, T-cell specific transcription factor 7-like 2
Highlights
-
•
A fragment of the 5′flanking region of the rat insulin receptor gene (−1143/−252 upstream ATG) has been cloned.
-
•
Several transcription start sites have been mapped in the conserved −460/−360 sequence.
-
•
The −1143/−252 fragment shows promoter activity in hepatoma cells. Activity is inhibited by vanadate treatment.
-
•
The −760/−465 sequence is required for activity of the −1143/−252 fragment but not for vanadate-induced inhibition.
-
•
Potential targets of vanadate are FoxO1 and HMGA1, two nuclear effectors of the insulin signaling pathway.
1. Introduction
Vanadium is a transition metal found in relative abundance in nature and in trace amounts in living organisms. By inhibiting enzymes involved in phosphate transfer and release reactions, vanadium compounds affect a broad variety of cellular processes [1]. Since 1979, many reports have underlined their ability to exert insulin-like effects in isolated cells, to normalize glucose homeostasis and insulin sensitivity in rodent models of type I and II diabetes, and to improve insulin sensitivity in some diabetic human subjects [2]. The insulinomimetic action of vanadium compounds is linked to their ability to activate several components of the insulin signaling pathway, through inhibition of their dephosphorylation by protein tyrosine phosphatases [3], [4], [5]. Additionally, vanadium has been shown to correct the impaired expression of hepatic genes involved in carbohydrate and lipid metabolism in rodents with type 1 diabetes [6], [7], [8].
One major target of vanadium in cells is the insulin receptor, the most proximal component of the insulin signaling pathway. In addition to their ability to activate IR kinase and to inhibit the dephosphorylation of the tyrosine phosphorylated β subunit of the receptor [9], vanadium compounds have been shown to induce a decrease in IR protein [10], [11], [12] and mRNA [12] level in cultured cells and to prevent their increase in STZ-induced diabetic rats [13], [14], [15]. Although the in vivo effects of vanadium in diabetic rats may be mediated in part by reversal of hyperglycemia, studies with isolated cells clearly indicate a direct effect on IR gene expression. Furthermore, vanadate-induced inhibition of insulin receptor gene expression in Fao hepatoma cells has been shown to occur at a transcriptional level [12].
Transcription of the human IR gene is critically regulated by the architectural transcription factor HMGA1. First described as a nuclear binding protein whose expression increases along with IR mRNA during myocyte and adipocyte differentiation [16], HMGA1 was later characterized [17] and shown to activate IR gene transcription as a multiprotein-DNA complex with the Sp1 and C/EBPβ transcription factors [18]. Genetic defects of HMGA1 in human subjects and disruption of the HMGA1 gene in mice adversely affect IR protein expression and impair insulin signaling, and conversely forced expression of HMGA1 in cells of affected subjects restores IR expression [19]. Recently, HMGA1 has been identified as a major downstream nuclear target of the PI3K/Akt insulin signaling pathway [20]. Serine phosphorylation of HMGA1 by Akt kinase, by reducing its DNA binding activity, mediates insulin-induced inhibition of IR gene transcription. Another important regulator of this process is FoxO1, a crucial effector of insulin action [21]. As a target of the PI3K/Akt signaling pathway, FoxO1 also mediates, via phosphorylation and nuclear exclusion, insulin-induced IR down regulation [21]. These observations raise the possibility that HMGA1 and/or FoxO1 are involved in the ability of vanadate to inhibit transcription of the IR gene.
To get some insight into the DNA sequence and transcription factor(s) potentially involved in this inhibitory effect of vanadate, a fragment of the proximal 5′flanking region of the rat IR gene (−1143/−252 upstream the ATG codon) has been cloned and characterized. Transcription start sites in this fragment have been mapped, and the full length and two 5′deleted fragments have been examined for their ability to direct expression of linked reporter luciferase in untreated and vanadate-treated rat hepatoma cells.
2. Materials and methods
2.1. Liver and hepatoma cells
Liver was obtained from male adult Sprague-Dawley rats. The Fao hepatoma cell line was derived from the Reuber H35 hepatoma [22]. Cells were cultured attached to Falcon Petri dishes in Coon's modified Ham's F12 medium supplemented with 5% fetal calf serum as previously described [12].
2.2. Extraction of nucleic acids
Genomic DNA was prepared from liver fragments (0.3–0.5 g) and Fao hepatoma cells (13 × 106 cells) by phenol extraction and ethanol precipitation after digestion by proteinase K. Total liver RNA was prepared by a single step procedure as described previously [12]. DNA and RNA integrity was assessed by electrophoresis on 0.8 or 1% agarose gels containing 0.1 mg/ml BET followed by UV visualization.
2.3. Cloning of fragments of the rat IR gene proximal 5′flanking region
The −1143/−252 fragment of the proximal 5′flanking region of the rat IR gene was generated by PCR amplification of liver genomic DNA using as primers the oligonucleotides 5′GCTCCAGACCTTTAAATTGGA-3’ (sense) and 5′ CGTCCTGTGTCCGCAATCGC-3’ (antisense). The former corresponds to the −1009/−989 sequence of the mouse IR gene (Genbank accession M28869.1), which exactly matches the −1143/−1123 sequence of the rat IR gene, and the latter to the −271/−252 sequence of the 347 bp 5′untranslated portion of rat IR cDNA [23] (Genbank accession M29014.1). The specificity of the −1143/−252 fragment was verified by Southern blotting using as a probe a 32P-labeled EcoR1-Eco47III fragment of rat IR gene cDNA (sequence −347/−15). Following cloning and sequencing of two additional fragments (−890/−514 and −1143/−563), the complete sequence of the −1143/−252 fragment was determined and the −1143/−1 sequence was deposited in Genbank (accession AJ006071). Two 5′ truncated DNA fragments, encompassing −760/−252 and −464/−252 sequences, were prepared by digestion of the parental −1143/−252 fragment cloned in the PCRTMII vector by digestion with the restriction enzymes Pvu2 and Dra1, respectively.
2.4. Sequence analysis of the rat IR gene proximal 5′flanking region
Database searches and alignments of the 5′flanking sequence of the rat IR gene with that of the mouse and human IR genes were performed using the blast program (ncbi.nlm.nih.gov/BLAST). The rat sequence was scanned for potential SINEs using SINE base search (http://sines.eimb.ru) and for potential transcription factor binding sites using the Jaspar software and Jaspar core database (http://jaspar.genereg.net).
2.5. Identification of transcription start sites in rat IR gene 5′flanking region
Transcription start sites in the −1143/−252 fragment of the rat IR gene were mapped using a RNase protection assay. The −464/−252 and −760/−252 fragments generated as indicated above were used as templates to synthesize two [P32] labeled antisense RNA probes using T7 RNA polymerase. Each probe (500,000 cpm) was hybridized with 40 μg of total liver RNA in 20 μl of 40 mM Pipes buffer, pH 6.5, containing 80% (v/v) formamide, 0.4 M NaCl and 1 mM EDTA. After incubation for 15 h at 55 °C, the nonhybridized probes were digested by RNase T1 (20 U/ml) and RNase A (40 μg/ml) in RNase buffer for 30 min at 37 °C, and hybrids were analyzed on sequencing polyacrylamide gels (6% polyacrylamide/7M urea). The size of the protected RNA fragments was determined using a dideoxy sequencing ladder run in parallel on the same gels.
2.6. Construction of the IR-luciferase gene fusion plasmids, transfection of hepatoma cells, and determination of luciferase activity
The −1143/−252, −760/−252 and −464/−252 fragments of the rat IR gene cloned in the PCRTMII vector were subcloned in pGL3, a low background promoter-less luciferase plasmid. IR inserts were excised from the IR-PCRTMII constructs by digestion with EcoR1 (−1143/−252 insert), Hind3 + EcoR1 (−760/−252 insert) and Apa1 + EcoR1 (−464/−252 insert), respectively. Owing to the lack of compatible cloning sites in the pGL3 vector, inserts were first cloned in the Bluescript vector and then excised by digestion with Xho1 + Sma1 (−1143/−252 fragment) and Kpn1 + Sma1 (−760/−252 and −464/−252 fragments) prior to subcloning in the pGL3 vector. The correct orientation of the inserts in the PGL3 constructs was verified by sequencing.
Fao hepatoma cells were transfected with empty pGL3 and IR-pGL3 constructs using fugene and then treated with vanadate or left untreated over three days. On day 1, cells were seeded in 6 well tissue culture dishes (300,000 cells per well) in 3 ml of Ham F12 medium supplemented with 10% calf serum, 1% penicillin-streptomycin (10,000 U/ml) and 2% fungizone (250 μg/ml). On day 2, 2 μg of plasmid DNA in 10 μl of water was mixed with 6 μl of fugene in 100 μl of medium, and after incubation for 15 min at 23 °C, this mixture was added dropwise to the cells. After 3–8 h, the medium was replaced by fresh medium. On day 3, sodium orthovanadate was added at the indicated concentration. On day 4, cells were washed in Mg++ and Ca++ free PBS and lysed in 0.2 ml of 100 mM KPO4, pH 7.8. Lysates were clarified by centrifugation and assayed in triplicate for luciferase activity using a luminometer. After normalization for protein concentration, activity of IR-pGL3 constructs was expressed as fold increase relative to activity of empty pGL3. All data shown were the mean ± SEM of 3–4 determinations in separate transfection experiments. Comparison between experimental groups were performed by analysis of variance followed by the Fisher, Scheffe and Bonferroni/Dunn tests. Differences were considered significant at the level of P < 0.05.
3. Results and discussion
3.1. Structural characterization of the proximal 5′flanking region of the rat IR gene
The proximal 5′flanking region of the IR gene was first cloned in human [24] and mouse [25] over 1.8 and 1.2 kb upstream the ATG, respectively. Alignment of the −1143/−1 sequence of the rat IR gene (this study, Genbank accession AJ006071) with the 1009/−1 sequence of the mouse IR gene (Genbank accession M28869) reveals a 86% identity at −766/−1 and −734/−1, and a 83% identity at −1143/−993 and −1009/−858, respectively. In contrast, identity of the rat sequence with the −1823/−1 sequence of the human IR gene (Genbank accession M23100) is only 68% at −558/−1 and −536/−1, respectively, with a maximum (83%) at −523/−407 (rat) and −545/−429 (human). Alignments of rat (NC_005111.4), mouse (NC_000074.6) and human (NC_018930.2) genomic clones over 10 kb upstream the IR gene coding sequence confirm these observations and show that a high identity (>78%) between the rat and mouse sequences is maintained up to −4.2 kb, whereas there is little identity between the rat and human sequences upstream of −0.6 kb.
As with the human [18] and mouse [19] IR genes, the 5′flanking sequence of the rat IR gene displays a high GC content (68% of the −575/−1 sequence). In addition, it harbours two 14 bp repeats (5′TCCCTAGGCTTTTA 3′) at −1073−1060 and −941/−928; a 77 bp sequence with 65% identity to the B4 member of the SINE family at −1018/−942; three 12–16 bp AT-rich (>80%) sequences at −997/−986, −765/−750 and −747/−734, potentially involved in the binding of HMGA1; a polyT sequence at −820/−791; and a polyC sequence at −551/−540. Potential binding sites for a number of transcription factors shown by chromatin immunoprecipitation to bind to DNA sequences in the proximal 5′flanking region of the human IR gene (http://www.genome-euro.ucsc.edu) are detectable in that of the rat gene. In addition to binding sites for Sp1, C/EBPβ, FoxO1, TCF7L2 and the glucocorticoid receptors, which are known to activate transcription of the human IR gene, there are binding sites for CREB, E2F1 and EGR1, which activate transcription of the IGF1 receptor gene, and for HNF4, a factor involved in hepatocyte differentiation. Alignment of the 5′flanking region of the rat, mouse and human IR genes over 2 kb upstream the ATG codon shows that number and distribution of potential transcription factor binding sites between the conserved and nonconserved portion depend on the particular factor considered, but are generally comparable for the three species.
3.2. Mapping of transcription start sites
As in many housekeeping genes, multiple transcription start sites have been identified in the 5′flanking sequence of the human and mouse IR genes. Primer extension and nuclease S1 protection assays have allowed the identification of seven sites at −550/−400 in the human gene [26], [27], [28] and of two sites at −469 and −424 in the mouse gene [25]. To map the transcription start sites in the 5′flanking sequence of the rat IR gene, a RNase protection assay was used. Liver RNA was hybridized with 32P-labeled antisense RNA probes complementary to the −464/−252 and −760/−252 DNA sequences, following which the size of RNase-protected hybrids was assessed by gel electrophoresis (Fig. 1). With the −464/−252 probe (Pvu2 generated), one major protected fragment of 144 bp, and four minor fragments of 194, 131, 117 and 111 bp were identified, indicating a major transcription start site at −396, and four minor sites at −446, −383 −369 and −363, respectively. With the −760/−252 probe (Dra1 generated), one major protected fragment of 203 bp and three minor protected fragments of 180, 144 and 117 bp were detected, indicating a major site at −455 and three minor sites at −432, −396 and −369. It is of interest that, as in the human and mouse IR genes, the transcription start sites identified in the rat IR gene are mainly localized in the conserved region of the gene.
Fig. 1.

Localization of transcriptional start sites in the 5′flanking region of the rat insulin receptor gene by RNase protection analysis. The −464/−252 and −760/−252 IR gene fragments generated from the −1143/−252 parental fragment by Pvu2 and Dra1, respectively, were used as templates to synthesize two [P32] labeled antisense RNA probes. Following incubation of these probes with 40 μg of total liver RNA, nonhybridized probes were digested by a mixture of RNase T1 and RNase A, and hybrids were analyzed by electrophoresis on 6% polyacrylamide gels containing 6M urea and radioautography. The size of the protected RNA fragments (arrows on the left) was determined using a dideoxy sequencing ladder run in parallel on the same gels (shown on the right).
3.3. Basal and vanadate-sensitive promoter activity of IR gene fragments in transfected hepatoma cells
Based on the expression of 5′deleted fragments of the human [26], [27], [28], [29], [30], [31] and mouse [25] IR genes linked to chloramphenicol acetyltransferase in transiently transfected cells, the majority of promoter activity has been mapped to sequences −646/−290 and −447/−412, respectively. In an attempt to identify the rat IR gene sequence required for basal and vanadate-sensitive promoter activity, three fragments (−1143/−252, 760/−252 and −464/−252) of IR genomic DNA linked to luciferase were examined for promoter activity in transiently transfected rat Fao hepatoma cells (Fig. 2).
Fig. 2.
Promoter activity of rat insulin receptor gene 5′flanking sequences in untreated and vanadate-treated Fao hepatoma cells. Fao hepatoma cells were transfected with the indicated IR gene fragment fused to the pGL3 plasmid as described in Materials and Methods. Transfected cells were left untreated or treated with vanadate at the indicated concentration for 24 h. Following cell lysis, lysates were clarified and assayed for luciferase activity with results normalized to protein and expressed relatively to empty pGL3 activity. For each of the three IR-luciferase constructs, asterisks indicate a statistically significant difference between untreated and vanadate-treated cells.
In untreated cells, promoter activity of the −760/−252 fragment was similar to that of the −1143/−252 fragment, but relative to these fragments activity of the −464/−252 fragment was decreased by 90%. These findings indicate that, consistent with deletional analysis of the 5′flanking region of the human IR gene, the −760/−465 sequence of the rat IR gene is important for activation of transcription. Interestingly, this sequence harbours a 16 bp AT-rich sequence at −765/−750, potentially involved in the binding of the transcription factor HMGA1, a conserved CEBPβ binding motif at −744/−734, and multiple potential Sp1 binding sites at −600/−500 (Fig. 3). Association of these factors within a multiprotein-DNA complex has been shown to be essential for activation of human IR gene transcription [19].
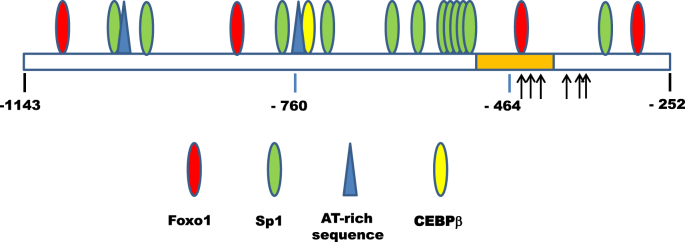
Fig. 3.
Schematic representation of the 5′flanking −1143/−252 sequence of the rat IR gene. This figure shows the conserved sequence (−523/−407) (orange box), the transcription start sites (arrows), and the binding sites of transcription factors potentially involved in IR gene transcription in untreated and vanadate-treated hepatoma cells (colored ellipses and triangles).
Treatment of Fao cells with vanadate led to a significant, dose-dependent decrease of promoter activity of each of the three IR gene fragments (Fig. 2). Relative to untreated cells, the changes in activity of the −1143/−252, −760/−252 and −464/−252 fragments induced by vanadate were respectively, 40%, 55% and 23% at 125 μM, and 70%, 85% and 62% at 250 μM, with no statistically significant differences between the three fragments. This suggests that although the entire −1143/−252 sequence may be involved in the inhibitory effect of vanadate, the short DNA sequence downstream of position −464 and upstream of the transcription start sites is sufficient for inhibition. Examination of the −1143/−252 sequence reveals, in addition to the potential binding sites for HMGA1, CEBPβ and Sp1 mentioned above, three potential binding sites for FoxO1 at −1109/−1099, −861/−851 and −450/−440 (Fig. 3). Since deletion of the −1143/−761 and −760/−465 sequences does not suppress the inhibitory effect of vanadate on promoter activity, the most proximal FoxO1 binding site at −450/−440, also the best conserved, can be considered as at least one potential site of action of vanadate.
The involvement of FoxO1 in the inhibitory effect of vanadate on IR gene transcription shown here is supported by several observations. First, as insulin, organic vanadium compounds stimulate, via Akt activation, serine phosphorylation of FoxO1 [32], and by resulting in FoxO1 nuclear exclusion this process has been shown to mediate the inhibitory effect of insulin on transcription of the IR gene [21] and other insulin-sensitive genes [33]. Second, some genes whose expression is inhibited by insulin, such as the PEPCK, glucose-6-phosphatase and IGFBP-1 genes, are also down-regulated by vanadate, and for at least two of them, glucose 6-phosphatase [34] and δ-aminolevulinate synthase [35], increased phosphorylation and/or nuclear exclusion of FoxO1 in response to vanadate have been demonstrated. Third, the increased activity of the PEPCK, glucose-6 phosphatase and IGFBP-1 gene promoters induced by overexpression of the FoxO1 related factor FoxO4 in HepG2 cells has been shown to be repressed by bisperxovanadate [36].
Since potential binding sites for HMGA1 and C/EBPβ are present in the rat IR gene upstream of position −730 in the IR gene, these transcription factors may also be involved in the ability of vanadate to inhibit the expression of the −1143/−252 and −760/−252 IR-luciferase constructs. As a downstream target of the PI3K/Akt insulin signaling pathway [20], HMGA1 is expected to undergo serine phosphorylation and a reduced ability to bind to IR DNA in vanadate-treated hepatoma cells. Interestingly by modulating FoxO1 binding to DNA and FoxO1 transactivation, HMGA1 has recently been shown to be essential for FoxO1-induced expression of the IGFBP-1 gene in liver cells, and thus identified as an important factor in modulating FoxO1 activity in the context of insulin signaling [37], [38]. Also supporting a potential involvement of C/EBPβ, vanadate treatment has been shown to decrease, in vitro, C/EBPβ mRNA in Fao hepatoma cells (Bortoli S, unpublished observations) and to reverse, in vivo, the overexpression of C/EBPβ mRNA in liver of STZ-induced diabetic rats [39].
4. Conclusion
Consistent with its ability to inhibit transcription of the endogenous IR gene in rat Fao hepatoma cells, vanadate treatment inhibits the promoter activity of the proximal 5′flanking region of the IR gene in transfected cells. It affects to a comparable extent the activity of the 1143/−252, −760/−252 and −464/−252 gene fragments, suggesting that the short sequence downstream of position −464 and upstream of transcription start sites is sufficient for inhibition. Potential targets of vanadate are the transcription factors FoxO1 and HMGA1, both of which have been shown to mediate, via Akt-dependent serine phosphorylation and reduced DNA binding activity, insulin-induced inhibition of transcription of the IR gene. Furthermore, FoxO1 has been implicated in vanadate-induced down regulation of several other insulin-sensitive genes. Analysis of Foxo1 and HMGA1 serine phosphorylation and association of these factors with IR genomic DNA in control and vanadate-treated hepatoma cells should help to prove their involvement in vanadate-induced inhibition of IR gene transcription.
Author contributions
The three authors conceived and coordinated the study. MC and SB designed and performed the experiments and analyzed the results. BD wrote the paper and SB contributed to the writing. All authors approved the final version of the manuscript.
Conflict of interest
The authors have no conflict of interest to declare.
Funding source
This work was supported by Institut National de la Santé et de la Recherche Médicale and Université Paris Descartes.
Acknowledgments
We gratefully acknowledge the assistance of Soledad Lopez in the cloning of the 5′flanking region of the rat IR gene and thank Claude Forest for his advice in the preparation of the manuscript.
This work is dedicated to the memory of our colleague Jean-François Savouret, with whom we had stimulating discussions.
References
- 1.Rehder D. The role of vanadium in biology. Metallomics. 2015;7:730–742. doi: 10.1039/c4mt00304g. [DOI] [PubMed] [Google Scholar]
- 2.Srivastava A.K., Mehdi M.Z. Insulino-mimetic and anti-diabetic effects of vanadium compounds. Diabet. Med. 2005;22:2–13. doi: 10.1111/j.1464-5491.2004.01381.x. [DOI] [PubMed] [Google Scholar]
- 3.Mehdi M.Z., Srivastava A.K. Organo-vanadium compounds are potent activators of the protein kinase B signaling pathway and protein tyrosine phosphorylation: mechanism of insulinomimesis. Arch. Biochem. Biophys. 2005;440:158–164. doi: 10.1016/j.abb.2005.06.008. [DOI] [PubMed] [Google Scholar]
- 4.Mehdi M.Z., Pandey S.K., Théberge J.F., Srivastava A.K. Insulin signal mimicry as a mechanism for the insulin-like effects of vanadium. Cell Biochem. Biophys. 2006;44:73–81. doi: 10.1385/CBB:44:1:073. [DOI] [PubMed] [Google Scholar]
- 5.Irving E., Stoker A.W. Vanadium compounds as PTP inhibitors. Molecules. 2017;22:2269. doi: 10.3390/molecules22122269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Valera A., Rodriguez-Gil J.E., Bosch F. Vanadate treatment restores the expression of key enzymes in the glucose and ketone bodies metabolism in the liver of diabetic rats. J. Clin. Invest. 1993;92:4–11. doi: 10.1172/JCI116580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Brichard S.M., Desbuquois B., Girard F. Vanadate treatment of diabetic rats reverses the impaired expression of genes involved in hepatic glucose metabolism: effects on glycolytic and gluconeogenic enzymes and on glucose transporter Glut2. Mol. Cell. Endocrinol. 1993;91:91–97. doi: 10.1016/0303-7207(93)90259-m. [DOI] [PubMed] [Google Scholar]
- 8.Brichard S.M., Ongemba L.N., Girard F., Henquin J.C. Tissue specific correction of lipogenic gene expression in diabetic rats given vanadate. Diabetologia. 1994;37(1994):1065–1072. doi: 10.1007/BF00418369. [DOI] [PubMed] [Google Scholar]
- 9.Posner B.I., Faure R., Burgess J.W., Bevan A.P., Lachance D., Zhang-Sun G., Fantus I.G., Ng J.B., Hall D.A., Lum B.S., Shaver A. Peroxovanadium compounds. A new class of potent phosphotyrosine phosphatase inhibitors which are insulin mimetic. J. Biol. Chem. 1994;269:4596–4604. [PubMed] [Google Scholar]
- 10.Marshall S., Monzon K. Down regulation of cell surface insulin receptors in primary cultured rat adipocytes by sodium vanadate. Endocrinology. 1987;121:1116–1122. doi: 10.1210/endo-121-3-1116. [DOI] [PubMed] [Google Scholar]
- 11.Torossian K., Freedman D., Fantus I.G. Vanadate downregulates cell surface insulin and growth hormone receptors and inhibits insulin receptor degradation. J. Biol. Chem. 1988;263:9353–9359. [PubMed] [Google Scholar]
- 12.Bortoli S., Amessou M., Collinet M., Desbuquois B., Lopez S. Vanadate, but not insulin, inhibits insulin receptor gene expression in rat hepatoma cells. Endocrinology. 1997;138:4821–4829. doi: 10.1210/endo.138.11.5521. [DOI] [PubMed] [Google Scholar]
- 13.Meyerovitch J., Farfel Z., Sack J., Shechter Y. Oral administration of vanadate normalizes blood glucose levels in streptozotocin-treated rats. Characterization and mode of action. J. Biol. Chem. 1987;262:6658–6662. [PubMed] [Google Scholar]
- 14.Venkatesan N., Avidan A., Davidson M.B. Antidiabetic action of vanadyl in rats independent of in vivo insulin-receptor kinase activity. Diabetes. 1991;40:492–498. doi: 10.2337/diab.40.4.492. [DOI] [PubMed] [Google Scholar]
- 15.Amessou M., Bortoli S., Liemans V., Collinet M., Desbuquois B., Brichard S., Girard D. Treatment of streptozotocin-induced diabetic rats with vanadate and phlorizin prevents the over-expression of the liver insulin receptor gene. Eur. J. Endocrinol. 1999;140:79–86. doi: 10.1530/eje.0.1400079. [DOI] [PubMed] [Google Scholar]
- 16.Brunetti A., Foti D., Goldfine I.D. Identification of unique nuclear regulatory proteins for the insulin receptor gene, which appear during myocyte and adipocyte differentiation. J. Clin. Invest. 1993;92:1288–1295. doi: 10.1172/JCI116702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Brunetti A., Manfioletti G., Chiefari E., Goldfine I.D., Foti D. Transcriptional regulation of human insulin receptor gene by the high-mobility group protein HMGI(Y) FASEB J. 2001;15:492–500. doi: 10.1096/fj.00-0190com. [DOI] [PubMed] [Google Scholar]
- 18.Foti D., Iuliano R., Chiefari E., Brunetti A. A nucleoprotein complex containing Sp1, C/EBPβ, and HMGI-Y controls human insulin receptor gene transcription. Mol. Cell Biol. 2003;23:2720–2732. doi: 10.1128/MCB.23.8.2720-2732.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Foti D., Chiefari E., Fedele M., Iuliano R., Brunetti L., Paonessa F., Manfioletti G., Barbetti F., Brunetti A., Croce C.M., Fusco A., Brunetti A. Lack of the architectural factor HMGA1 causes insulin resistance and diabetes in humans and mice. Nat. Med. 2005;11:765–773. doi: 10.1038/nm1254. [DOI] [PubMed] [Google Scholar]
- 20.Chiefari E., Nevolo M.T., Arcidiacono B., Maurizio E., Nocera A., Iiritano S., Sgarra R., Possidente K., Palmieri C., Paonessa F., Brunetti G., Manfioletti G., Foti D., Brunetti A. HMGA1 is a novel downstream nuclear target of the insulin receptor signaling pathway. Sci. Rep. 2012;2:251. doi: 10.1038/srep00251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Puig O., Tjian R. Transcriptional feedback control of insulin receptor by dFOXO/FOXO1. Genes Dev. 2005;19:2435–2444. doi: 10.1101/gad.1340505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Deschatrette J., Weiss M.C. Characterization of differentiated and dedifferentiated clones from a rat hepatoma. Biochimie. 1974;56:1603–1611. doi: 10.1016/s0300-9084(75)80286-0. [DOI] [PubMed] [Google Scholar]
- 23.Goldstein B.J., Dudley A.L. The rat insulin receptor: primary structure and conservation of tissue-specific alternative messenger mRNA splicing. Mol. Endocrinol. 1990;4:235–244. doi: 10.1210/mend-4-2-235. [DOI] [PubMed] [Google Scholar]
- 24.Seino S., Seino M., Nishi S., Bell G.I. Structure of the human insulin receptor gene and characterization of its promoter. Proc. Natl. Acad. Sci. U.S.A. 1989;86:114–118. doi: 10.1073/pnas.86.1.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sibley E., Kastelic T., Kelly T.J., Lane D. Characterization of the mouse insulin receptor gene promoter. Proc. Natl. Acad. Sci. U.S.A. 1989;86:9732–9736. doi: 10.1073/pnas.86.24.9732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Mamula P.W., Wong K.-Y., Maddux B.A., McDonald A.R., Goldfine I.D. Sequence and analysis of promoter region of human insulin-receptor gene. Diabetes. 1988;37:1241–1246. doi: 10.2337/diab.37.9.1241. [DOI] [PubMed] [Google Scholar]
- 27.Tewari D.S., Cook D.M., Taub R. Characterization of the promoter region and 3' end of human insulin receptor gene. J. Biol. Chem. 1989;264:15238–15245. [PubMed] [Google Scholar]
- 28.McKeon C., Moncada V., Pham T., Salvatore P., Kadowaki T., Accili D., Taylor S.I. Structural and functional analysis of the insulin receptor promoter. Mol. Endocrinol. 1990;4:647–656. doi: 10.1210/mend-4-4-647. [DOI] [PubMed] [Google Scholar]
- 29.Araki E., Murakami T., Shirotani T., Kanai F., Shinohara Y., Shimada F., Mori M., Shichiri M., Ebina Y. A cluster of four Sp1 binding sites required for efficient expression of the human insulin receptor gene. J. Biol. Chem. 1991;266:3944–3948. [PubMed] [Google Scholar]
- 30.Cameron K.E., Resnik J., Webster N.J.G. Transcriptional regulation of the human insulin receptor promoter. J. Biol. Chem. 1992;267:17375–17383. [PubMed] [Google Scholar]
- 31.Lee J.-K., Tam J.W.O., Tsai M.-J., Tsai S.Y. Identification of cis- and trans-acting factors regulating the expression of the human insulin receptor gene. J. Biol. Chem. 1992;267:4638–4645. [PubMed] [Google Scholar]
- 32.Vardatsikos G., Mehdi M.Z., Srivastava A.K. Bis(maltolato)-oxovanadium (IV)-induced phosphorylation of PKB, GSK-3 and Foxo1 contributes to its glucoregulatory responses. Int. J. Mol. Med. 2009;24:303–309. doi: 10.3892/ijmm_00000233. [DOI] [PubMed] [Google Scholar]
- 33.Mounier C., Posner B.I. Transcriptional regulation by insulin: from the receptor to the gene. Can. J. Physiol. Pharmacol. 2006;84:713–724. doi: 10.1139/y05-152. [DOI] [PubMed] [Google Scholar]
- 34.Hiromura M., Nakayama A., Adachi Y., Doi M., Sakurai H. Action mechanism of bis(allixinato)oxovanadium(IV) as a novel potent insulin-mimetic complex: regulation of GLUT4 translocation and FoxO1 transcription factor. J. Biol. Inorg. Chem. 2007;12:1275–1287. doi: 10.1007/s00775-007-0295-x. [DOI] [PubMed] [Google Scholar]
- 35.Oliveri L.M., Davio C., Battle A.M., Gerez E.N. ALAS1 gene expression is down-regulated by Akt-mediated phosphorylation and nuclear exclusion of FOXO1 by vanadate in diabetic mice. Biochem. J. 2012;442:303–310. doi: 10.1042/BJ20111005. [DOI] [PubMed] [Google Scholar]
- 36.Yang Z., Whelan J., Babb R., Bowen B.R. An mRNA splice variant of the AFX gene with altered transcriptional activity. J. Biol. Chem. 2002;277:8068–8075. doi: 10.1074/jbc.M106091200. [DOI] [PubMed] [Google Scholar]
- 37.Arcidiacono B., Chiefari E., Messineo S., Bilotta F.L., Pastore I., Corigliano D.M., Foti D.P., Brunetti A. HMGA1 is a novel transcriptional regulator of the FoxO1 gene. Endocrine. 2018;60:56–64. doi: 10.1007/s12020-017-1445-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Chiefari E., Arcidiacono B., Palmieri C., Corigliano D.M., Morittu V.M., Britti D., Armoni M., Foti D.P., Brunetti A. Cross-talk among HMGA1 and FoxO1 in control of nuclear insulin signaling. Sci. Rep. 2018;8:8540. doi: 10.1038/s41598-018-26968-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Bosch F., Sabater J., Valera A. Insulin inhibits liver expression of the CCAAT/enhancer-binding protein beta. Diabetes. 1995;44:267–271. doi: 10.2337/diab.44.3.267. [DOI] [PubMed] [Google Scholar]