Abstract
Thermotolerant Campylobacter species C. jejuni and C. coli are actually recognized as the major bacterial agent responsible for food-transmitted gastroenteritis. The most effective antimicrobials against Campylobacter are macrolides and some, but not all aminoglycosides. Among these, susceptibility to streptomycin is reduced by mutations in the ribosomal RPSL protein or by expression of ANT(6)-I aminoglycoside O-nucleotidyltransferases. The presence of streptomycin resistance genes was evaluated among streptomycin-resistant Campylobacter isolated from humans and animals by using PCR with degenerated primers devised to distinguish ant(6)-Ia, ant(6)-Ib and other ant-like genes. Genes encoding ANT(6)-I enzymes were found in all possible combinations with a major fraction of the isolates carrying a previously described ant-like gene, distantly related and belonging to the new ant(6)-I sub-family ant(6)-Ie. Among Campylobacter isolates, ant(6)-Ie was uniquely found functional in C. coli, as shown by gene transfer and phenotype expression in Escherichia coli, unlike detected coding sequences in C. jejuni that were truncated by an internal frame shift associated to RPSL mutations in streptomycin resistant strains. The genetic relationships of C. coli isolates with ANT(6)-Ie revealed one cluster of strains presented in bovine and humans, suggesting a circulation pathway of Campylobacter strains by consuming contaminated calf meat by bacteria expressing this streptomycin resistance element.
Keywords: Campylobacter coli, Campylobacter jejuni, streptomycin-resistance, aminoglycoside adenylyl transferases, ANT(6)-I
Introduction
Campylobacteriosis is the main cause of foodborne diseases in the UE and in the United States [Collective Eurosurveillance Editorial Team, 2015; (Accessed March 2018) 1]. The drugs of choice for the treatment of campylobacteriosis were, mainly erythromycin (ERY) and ciprofloxacin (CIP), although quinolones are no longer effective after a fast rise in resistance mechanisms among Campylobacter species (Carreira et al., 2012; Hormeño et al., 2016). Aminoglycosides, the third class of antimicrobials used worldwide after sulfonamides and beta-lactams, are a recommended alternative for the treatment of difficult infections caused by thermotolerant Campylobacter spp. (Wieczorek and Osek, 2013). The advantages of using aminoglycosides compared to other antimicrobials are their concentration-dependent bactericidal activity and relatively predictable pharmacokinetics, and synergism with other antibiotics (Vakulenko and Mobashery, 2003). Among aminoglycosides, the first active molecule used was streptomycin (STR), produced by Streptomyces griseus. STR binds to the aminoacyl-tRNA site (A site) of the 16S rRNA in the 30S ribosomal subunit, inducing codon misreading and inhibiting of translocation (Moazed and Noller, 1987; Woodcock et al., 1991) which leads to inadequate protein production. When antibiotic resistance appears it is due to target modification of ribosomal components, antimicrobial modification, or lowering drug accumulation in the cell (Vakulenko and Mobashery, 2003). Like in other bacteria, mutation K43R of S12 protein, a component of the 30S ribosomal subunit encoded by the rpsL gene, confers high-level of STR resistance in Campylobacter (Olkkola et al., 2010). Besides that, two out of four ANT(6)-I subfamily members of aminoglycoside nucleotidyltransferases (also known as aminoglycosides adenyltransferases of the AADE family), ANT(6)-Ia and ANT(6)-Ib, are frequently involved in STR resistance in Campylobacter strains and probably evolved from Gram-positive bacteria (Pinto-Alphandary et al., 1990; Shaw et al., 1993; Gibreel et al., 2004; Nirdnoy et al., 2005; Abril et al., 2010; Qin et al., 2012; Zhao et al., 2016). An additional role in STR resistance of ANT-like protein has been suggested in C. coli (Olkkola et al., 2016).
The aim of this work was to characterize the STR resistance presented in Campylobacter isolates of human and animal origin, establishing the role of a new enzyme of the ANT(6)-I family, ANT(6)-Ie, detected in a significant fraction of STR resistant isolates which molecular typing evidenced spread between animal and human hosts.
Materials and Methods
Bacteria and Antimicrobial Resistance
Campylobacter spp. strains isolated from humans were previously described (Hormeño et al., 2016) and resulted from systematical screenings performed during 2010–2012 in fecal samples from gastroenteritis patients by the Microbiology services of three hospitals located in West-Center Spain: San Pedro de Alcántara, Cáceres; Campo Arañuelo, Cáceres; and Universitario de Salamanca, Salamanca. Campylobacter spp. isolated from bovine, fattening pigs and poultry were randomly selected in 2010–2012 from slaughterhouses located all around Spain by the Spanish Surveillance Network of Antimicrobial Resistance in Bacteria of Veterinary Origin (VAV Network; Moreno et al., 2000) and were partially described elsewhere (Florez-Cuadrado et al., 2016). From each farm, a single Campylobacter isolate was obtained by culturing pooled feces from animals (bovine and porcine) and cloacal or meat samples (poultry). Isolates were grown on blood agar, in a microaerophilic atmosphere (CampyGenTM, Thermo Scientific) at 42°C for 24–48 h and were identified by a Vitek-MS MALDI-TOF system (bioMérieux, Marcy-l’Etoile, France) to species level. The minimal inhibitory concentrations (MICs) for STR, ERY, gentamicin (GEN), CIP, and tetracycline (TET) were determined by agar dilution methods according to the guidelines of CLSI (Clinical and Laboratory Standards Institute [CLSI], 2010), including Campylobacter jejuni ATCC 33560 as the reference strain. Resistance was determined according to the EUCAST 2 (last accessed September of 2018), by using cut-off values [ecological cut-off value (ECOFF)] of 4 mg/L for STR, 4 mg/L (C. jejuni) or 8 mg/L (C. coli) for ERY, 2 mg/L for GEN, 0.5 mg/L for CIP, and 1 mg/L (C. jejuni) or 2 mg/L (C. coli) for TET. To test the presence of efflux pumps, MIC to STR were determined in the presence of the efflux pump inhibitor phenylalanine-arginine beta-naphthylamide (PaβN, Sigma) at a concentration of 20 mg/L.
Detection of Resistance Determinants
PCR was performed on DNA obtained by boiling, for 5 min, a suspension of one or two colonies from pure culture in 250 μL of milli-Q water, and recovering the supernatant after centrifugation at 10,000 ×g for 10 min. PCR was carried out with 1 μl of DNA, 0.2 mM of each dNTP (Biotools, Madrid, Spain), 0.5 μM of each primer [Stab Service (University of Extremadura, Badajoz, Spain)], 0.025 U/μl of Taq Polymerase (Biotools, Madrid, Spain) and 1X PCR buffer containing 1.5 mM MgCl2 (Biotools, Madrid, Spain), during 30 cycles of 94°C, 30 s; annealing temperature indicated in Table 1, 30 s; 72°C, 1 min. Amplicon purification was done with Speedtools PCR clean-up kit (Biotools, Madrid, Spain), following the manufacturer’s instructions. DNA sequencing were performed by STAB Service (DNA Sequencing facilities of the Universidad de Extremadura, Spain). In silico data analysis was carried out with bioinformatics tools available in NCBI3, SMS4, and EBI5.
Table 1.
Primers used in this work.
| Name | Sequence (5′-3′) | T1 | Bp2 | Reference |
|---|---|---|---|---|
| RPSLF | CCAGCGCTTAAAAAT TGTCC | 55 | 247 | Olkkola et al., 2010 |
| RPSLR | TATCAAGAGCACCA CGAACG | |||
| INT1F | GGCTCTCGGGTAAC ATCAAGG | 54 | 242 | Leverstein-van Hall et al., 2002 |
| INT1R | TCAGGAGATCGGAA GACCTC | |||
| CSF | GGCATCCAAGCAGCAAG | 56 | VAR3 | Lévesque et al., 1995 |
| CSR | AAAAGCAGACTTGA CCTGA | |||
| SAF | TGCAAAA(G/A)CC(G/C) GA(A/G)GATATGG | 56 | 305 | This work |
| SAR | TTCCTT(G/T)CG(G/A) CATA(G/T)CC(C/T)TT | |||
| SBF | GATTGT(T/C)CG(T/C)CAT GAGCTGCT | 57 | 327 | This work |
| SBR | GTGCTATCCAGGCAGC CGGTT | |||
| SCF | TGCCT(A/C)AAATTGG(G/A) T(G/A)AGTT | 52 | 368 | This work |
| SCR | ACCTAGCCA(A/G)ATTTCA AA(A/G)CCAAA | |||
| STREJF | TGCAAAGCGAAAA AAGAAT | 49 | 878 | This work |
| STREJR | TTATAATTTTCTTAAAAT TTTGCAAT | |||
| STRECF | TGCAAAATCAAGATAAAT TTTTAAAAC | 51 | 899 | This work |
| STRECR | TTACAATTTTCCTAAAAT TTTACAAT | |||
| STREFF | GTATGCGCAAAAATGAT TAAAG | 50 | 1110 | This work |
| STREFR | AAGGAAAAATTTAAATAT TGGTTTCA | |||
1Annealing temperatures for PCR. 2PCR-Product size in bp. 3Variable size depending on gene-cassette structure (Lévesque et al., 1995).
Mutations in the STR resistance region of the rpsL gene were screened by sequencing of the PCR amplicon produced by primers and conditions previously described (Table 1; Olkkola et al., 2010). Similarly, the possible presence of ant(3”)-Ia genes carried by Class-I integrons was evaluated by PCR with primers specific to intI and intI-associated gene cassettes (Table 1). Three sets of degenerated primers were designed to amplify internal fragments of genes ant(6)-I (Table 1): ant(6)-Ia (primers SAF and SAR), ant(6)-Ib (primers SBF and SBR), and ant(6)-Ie (primers SEF and SER). Further analysis was performed to amplify the (almost) full coding sequences of ant(6)-Ie genes (Table 1) from C. jejuni (primers STREJF and STREJR) and C. coli (primers STRECF and STRECR). Oligonucleotide design was performed with Oligo v.6 software.
Functional Expression in E. coli
The expression of ant(6)-Ie from C. coli was tested through cloning the complete gene in the vector pGem-T Easy (Promega®), according to the manufacturer’s instructions. The full length of the gene including its promoter sequence was amplified by using primers STREFF and STREFR (Table 1), designed from the genome sequence of C. coli Z163 (ZP_14079546.1) and assuming that σ70 Campylobacter promoters have a well-conserved -10 box and lack the -35 box presented in other bacteria (Petersen et al., 2003). The ligation mixture was electroporated in Escherichia coli XL1-Blue MRF’ and transformants were selected in Luria-Bertani medium supplemented with 100 mg/L ampicillin.
Multilocus Sequence Typing of Campylobacter Isolates From Human and Animal Origin
A group of Campylobacter isolates was genotyped for flaA-SVR (short variable region of flaA gene) and multilocus sequence typing (MLST). PCR fragments of the housekeeping genes aspA (aspartase A), glnA (glutamine synthetase), gltA (citrate synthase), glyA (serine hydroxymethyltransferase), pgm (phosphoglucomutase), tkt (transketolase), and uncA (ATP synthase a subunit), as well as flaA gene (flagellin), were amplified and sequenced as described elsewhere (Ugarte-Ruiz et al., 2013). Allele numbers were assigned by sequence comparisons against the existing sequences deposited in the Campylobacter MLST database6.
Results
Streptomycin Resistance Phenotypes in Isolates From Human Origin
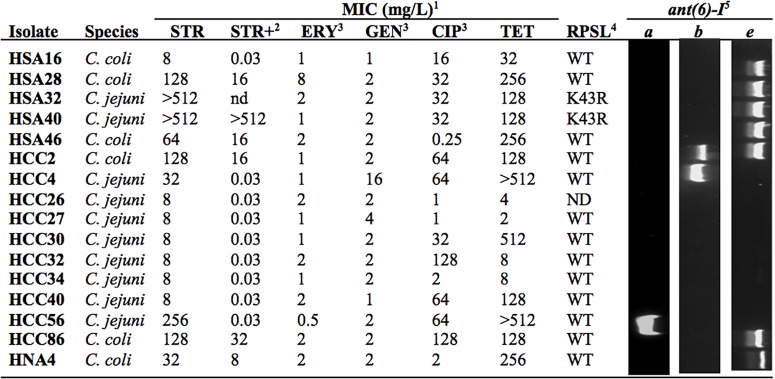
Based on the ECOFF defined by EUCAST for STR resistance of Campylobacter (MIC > 4 mg/L), 16 out of 141 human isolates are above the threshold (Figure 1). Among these it was possible to identify three different phenotypes: high-level resistance, shown by two C. jejuni strains (MIC > 512 mg/L), medium-level resistance, in two C. jejuni and five C. coli isolates (32 ≤ MIC ≤ 256 mg/L), and low-level resistance, with inhibition of growth immediately above ECOFF, detected in six C. jejuni and one C. coli (MIC = 8 mg/L). Treatment with the efflux pump inhibitor PAβN reduced MICs in all the isolates, with the exception of the highly resistant HSA40, with maximal susceptibility attained in two isolates from the medium-level resistance group plus in the seven isolates with the lowest resistance level (Figure 1). Among analyzed isolates, low susceptibility against clinically relevant antimicrobials was generally found to CIP and/or TET but not to ERY or GEN, although three low-level resistant strains to STR were also found near the cut-off for CIP and TET (HCC26, HCC27, and HCC34; Figure 1).
FIGURE 1.
Phenotypic and genotypic analysis of streptomycin (STR) resistant isolates. 1Minimal inhibitory concentrations for STR, erythromycin (ERY), gentamicin (GEN), ciprofloxacin (CIP), and tetracycline (TET). 2MIC were determined in the presence of PaβN (mg/L). 3Data previously reported (Hormeño et al., 2016). 4Mutations in the RPSL coding sequence were detected by sequencing (WT, no mutation). 5Genes ant(6)-I were amplified with PCR with specific primers. ND, not determined.
rpsL Polymorphism Among Streptomycin Resistant Isolates
The rpsL gene region determining resistance to aminoglycosides (Olkkola et al., 2010) was amplified and sequenced in 15 Campylobacter isolates with MICs above STR ECOFF value (Accession Nos. LT605180, LT605181, LT605182, LT605184, LT605185, LT605186, LT605187, LT605190, LT605191, LT605192, LT605193, LT605194, LT605195, LT605196, and LT605197). Among 11 polymorphic positions detected, only one was expressed at protein level corresponding to mutation K43R (not shown). This occurred in two C. jejuni isolates, HSA32 and HSA40 (Accession Nos. LT605194 and LT605195), having both the high-level resistant phenotype (Figure 1).
The ANT(6)-I Family in Campylobacter
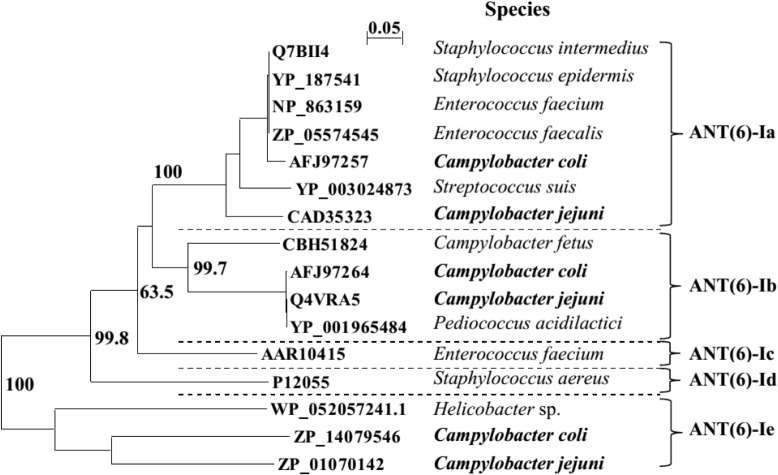
The NCBI database includes sequences for three members of the ANT(6) protein family previously described in Campylobacter: ANT(6)-Ia, ANT(6)-Ib, and ANT-like sequence cluster (Abril et al., 2010; Olkkola et al., 2016). The phylogenetic relationships previously defined within the ANT(6)-I family (Abril et al., 2010) were re-analyzed (Figure 2), including C. jejuni and C. coli for clusters ANT(6)-Ia and ANT(6)-Ib, plus the new and distantly related family member previously identified as ANT-like (Olkkola et al., 2016). Supported by bootstrapping with a threshold near 70%, ANT-like sequences cluster is a new member of the protein family that will be named hereafter ANT(6)-Ie (Figure 2), the fifth described ANT(6) (aminoglycoside 6-adenyltransferase) enzyme.
FIGURE 2.
The ANT(6)-I phylogenetic tree. Multiple sequence alignment was performed by Clustal X 2.1. The phylogenetic tree was deduced by neighbor joining algorithm, excluding positions with gaps and emulated by NJPlot 2.3. Bootstrap values (N, 100; seeds, 111) are indicated for branches supporting sequence clustering and assuming previous data (Abril et al., 2010).
ANT(6)-I Detection in Streptomycin Resistant Isolates
The role of ANT(6)-I enzymes on STR resistance of Campylobacter was addressed by using specific primers designed to detect the coding sequences for ANT(6)-Ia, ANT(6)-Ib, and ANT(6)-Ie, including degenerated positions for efficient amplification of homologs of either C. jejuni or C. coli for every subfamily (Table 1). Among the 16 Campylobacter isolates resistant to STR detected in this work from human infections, nine were positive for the presence of ant(6)-I genes with two isolates positive for the subfamilies ant(6)-Ia, one for ant(6)-Ib and seven for ant(6)-Ie (Figure 1). The unique two C. jejuni isolates presenting ant(6)-Ie also have the RSPL polymorphism K43R and the high-resistance phenotype, whereas the six isolates with low-level of resistance did not carry any of the screened genes.
The nucleotide sequences of the seven ant(6)-Ie genes detected among human isolates, including the six Campylobacter strains presenting this gene as the unique aminoglycoside 6-adenyltransferase enzyme, revealed different functional roles on STR resistance depending on Campylobacter species. The ant(6)-Ie genes from the two C. jejuni isolates were found non-functional when compared with the reference used to define the protein subfamily (ZP_01070142, Figure 2), sharing both the unique polymorphism C-394-Δ (Accession No. LT605198, isolate HSA32), an out of frame deletion that produces the premature arrest of translation and the loss of 55% of protein sequence from its C-terminal end. In contrast, the four ant(6)-Ie genes from C. coli strains HCC2, HSA28, HSA86, and HCC46 presented identical sequences to ZP_14079546.1, whereas the polymorphism C466T originating variant P156S in the encoded protein was detected in the gene from HNA4 isolate (Accession No. LT605200).
Functional Expression in E. coli of ANT(6)-Ie
The coding sequence for ANT(6)-Ie from HNA4 was amplified and cloned in pGEM-T vector and E. coli XL1 Blue (MRF’) cells. Cells carrying the recombinant vector expressed resistance to STR with a MIC of 64 mg/L, significantly higher than the control cells transformed with a non-recombinant vector (MIC = 8 mg/L). Besides, both recipient and transformants cells remained sensitive to other antimicrobials tested showing aminoglycoside specificity of the ant(6)-Ie gene: spectinomycin (MIC ≤ 32 mg/L), GEN (MIC ≤ 1 mg/L), apramycin (MIC ≤ 4 mg/L), and neomycin (MIC ≤ 4 mg/L).
Genetic and Phenotype Relationships Among Human and Animal Streptomycin Resistant Isolates Carrying ANT(6)-Ie
We screened for the three ANT(6)-I encoding genes in Campylobacter among 65 STR resistant isolates from the three most common food-producing livestock: poultry, pigs, and cattle (Table 2). All ant(6)-I genotypes were detected, with C. coli being largely the most prevalent species among streptomycin resistant isolates. Interestingly, the presence of the single-gene ant(6)-Ie genotype represents a major fraction of STR resistant C. coli, with one fourth of isolates.
Table 2.
ant(6)-I genotypes of streptomycin resistant Campylobacter isolates.
| Host |
ant(6)-I profile1 |
||||||||
|---|---|---|---|---|---|---|---|---|---|
| a | b | e | a/b | a/e | b/e | a/b/e | Ø | Σ | |
| Human2 | 1 | 1 | 6(4) | - | - | 1(1) | - | 7(1) | 16 (6) |
| Poultry | 13(8) | 1(1) | - | 10(9) | 1(1) | - | 2(1) | 5(1) | 32 (21) |
| Porcine | 4(4) | - | 10(10) | 2(2) | 7(7) | 4(4) | 1(1) | 1(1) | 29 (29) |
| Bovine | 2(1) | - | 1(1) | - | 1(1) | - | - | - | 4 (3) |
| Σ | 20(13) | 2(1) | 17(15) | 12(11) | 9(9) | 5(5) | 3(2) | 6(3) | 81 (59) |
Multilocus sequence typing plus flaA typing was performed in 14 C. coli isolates carrying ant(6)-Ie as the only determinant expressing STR resistance (Table 3). The multilocus analysis allowed the detection of a cluster of strains (ST-827, clonal complex 828) including two isolates from human origin plus one from bovine. Moreover, one of the human and the bovine origin isolates shared the same flaA allele 236 and the same resistance profile against the five clinically relevant antimicrobials tested, which is considered an indication of a probable common clonal origin.
Table 3.
Molecular and antimicrobial resistance typing of Campylobacter isolates carrying1 ant(6)-Ie.
| MIC (mg/L)2 |
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Strain | Year | Origin | STR | ERY | GEN | CIP | TET | CC3 | ST4 | flaA |
| ZTA10/00526CPD | 2010 | Porcine | ≥32 | 1 | 4 | ≥8 | ≥32 | ST-828 | 7337 | ND |
| ZTA10/00602CPD | 2010 | Porcine | ≥32 | ≥64 | 4 | ≥8 | ≥32 | ND | 7340 | ND |
| ZTA10/00794CPD | 2010 | Porcine | ≥32 | 1 | 4 | ≥8 | ≥32 | ST-828 | 829 | ND |
| ZTA10/01257CPD | 2010 | Bovine | ≥32 | 1 | 2 | ≥8 | ≥32 | ST-828 | 827 | 0236 |
| ZTA10/01418CPD | 2010 | Porcine | ≥32 | ≥64 | 2 | ≥8 | ≥32 | ST-828 | 1413 | ND |
| ZTA10/02049CPD | 2010 | Porcine | ≥32 | 2 | 2 | ≥8 | ≥32 | ST-828 | 4950 | ND |
| ZTA11/00514CP | 2011 | Porcine | ≥32 | ≥64 | 2 | ≥8 | ≥32 | ND | 7341 | 0662 |
| ZTA11/00726CP | 2011 | Porcine | ≥32 | 1 | 4 | 0.13 | ≥32 | ST-828 | 7338 | ND |
| ZTA11/01342CP | 2011 | Porcine | ≥32 | ≥64 | 4 | 0.25 | ≥32 | ST-828 | 1413 | ND |
| ZTA11/03282CP | 2011 | Porcine | ≥32 | 0.5 | 1 | ≥8 | ≥32 | ST-828 | 1096 | 0319 |
| ZTA11/03389CP | 2011 | Porcine | ≥32 | ≥64 | 2 | ≥8 | ≥32 | ST-828 | 2733 | ND |
| HSA028 | 2010 | Human | 128 | 8 | 2 | 32 | 256 | ST-828 | 827 | 0236 |
| HSA046 | 2010 | Human | 64 | 2 | 2 | 0.25 | 256 | ST-828 | 827 | 0255 |
| HNA4 | 2010 | Human | 32 | 2 | 2 | 2 | 256 | ND | 7339 | 0633 |
1The fourteen C. coli isolates presenting ant(6)-Ie as the unique streptomycin (STR) resistance determinant (Table 2). 2Minimal inhibitory concentrations for STR, erythromycin (ERY), gentamicin (GEN), ciprofloxacin (CIP), and tetracycline (TET). 3Clonal Complex. 4Sequence Types and flaA alleles were assigned by MLST database (see footnote 6). ND, not determined.
Discussion
This work shows the main role of adenylyl transferases belonging to the ANT(6)-I family on STR resistance in Campylobacter. Previous reports had described the phenotypic expression of ANT(6)-I enzymes (Nirdnoy et al., 2005; Abril et al., 2010; Qin et al., 2012; Olkkola et al., 2016), and now strong evidence is provided supporting the role of ANT(6)-Ie on STR resistance. Although ANT(6)-Ie coding sequences were detected in the two most frequent Campylobacter species, C. jejuni and C. coli, the association with STR resistance was only proved in C. coli since no C. jejuni isolate carried this coding sequence as the unique candidate to express the phenotype (Figure 1 and Table 2).
Besides ANT(6)-I, an additional STR resistance determinant is ANT(3”)-Ia or AADA which also confers resistance to spectinomycin. This enzyme is highly prevalent among enterobacteria (Shaw et al., 1993) and has been detected associated to class I integrons and their gene cassettes in Campylobacter, although only anecdotally (Ouellette et al., 1987; O’Halloran et al., 2004). Indeed, several reports have described the unsuccessful search of ant(3”) in Campylobacter (van Essen-Zandbergen et al., 2007; Piccirillo et al., 2013). Similarly, all STR resistant isolates from humans analyzed in the present work have been screened for int1 or associated gene cassettes, unsuccessfully (data not shown). Thus, ANT(6)-I enzymes might be the unique adenylyl transferases with significant relevance in STR resistance in Campylobacter.
To the best of our knowledge, this is the first report showing a RPSL mutation in C. jejuni isolates conferring STR resistance. In a previous study, with C. coli, it was found that isolates presenting high-level resistance to STR shared the mutation K43R in RPSL (Olkkola et al., 2010), similarly to the two C. jejuni isolates from humans, detected in this work, with MIC > 512 mg/L (Figure 1). Although both isolates also carry ant(6)-Ie genes, resistance to STR might be determined by RPSL mutation since the adenylyl transferase coding sequence is truncated and most probably not functional. In addition, there was no contribution to this phenotype from efflux pump activity, as deduced by the lack of any effect on MIC by PAβN treatment (Figure 1).
A group of six C. jejuni and one C. coli isolates from humans that expressed low-level STR-resistance, did not contain any of the screened determinants and presented a strong decreased MIC to STR in the presence of PAβN (Figure 1). Thus, efflux pump activity must be responsible for low-level STR resistance of these strains, similarly to Mycobacterium tuberculosis where the effect of outward transporters is known to increase modestly the MIC for STR (Spies et al., 2008). At least three different efflux pump systems have been shown to be up-regulated in Campylobacter strains resistant to a broad range of antimicrobials (Lin et al., 2005; Akiba et al., 2006; Jeon et al., 2011), so they could be candidates for determinants to the low level STR resistance. In addition, treatment with PAβN produced a strong effect on MIC of Campylobacter isolates carrying ant(6)-I genes, mostly for those with ant(6)-Ia or ant(6)-Ib as unique resistance determinants (Figure 1). This observation might indicate that, among human isolates analyzed in this work, the only functional adenylyl transferase gene is ant(6)-Ie and that even these isolates require efflux pump activity to support the medium-level of resistance. Treatment of ant(6)-Ie carrying strains with PAβN reduces their STR MIC to low-level resistance, which might correspond to their in vivo expression level. Synergic effects of efflux pumps have been evidenced in Campylobacter with resistance determinants for quinolones and macrolides, gyrA and 23S rRNA gene mutations, respectively (Luo et al., 2003; Cagliero et al., 2006; Corcoran et al., 2006). Indeed, three Campylobacter isolates showing low-level resistance to STR were also found to have low-level resistant to CIP and TET (Figure 1), lacked the gyrA C-257-T mutation conferring low susceptibility to fluoroquinolones (Hormeño et al., 2016) and also tetO, the major TET resistant determinant in this species (not shown, authors’ personal communication). A weak overexpression of efflux pump activity might be involved in the antimicrobial resistance phenotype of these strains.
The set of primers described in this work allows specific detection of the three ant(6)-I genes described in Campylobacter, including those belonging to ant(6)-Ie and encoding a new subfamily of aminoglycoside O-nucleotidyltransferases (Figure 2) that provides functional information for hundreds of orthologs annotated as hypothetical proteins, mainly from Campylobacter and related organisms like Helicobacter. In addition, the molecular and antimicrobial resistance typing of Campylobacter isolates expressing ANT(6)-Ie has revealed a spread pathway for this zoonotic pathogen between cattle and humans.
Author Contributions
SP and AQ conceived and designed the study. LH, MU-R, GP, CB, and DF-C acquired the samples and data. LH, MU-R, GP, DF-C, and MC performed the laboratory analysis. SV, SP, LD, MC, and AQ analyzed and interpreted the data. MC and AQ wrote the manuscript. All authors have approved the final article.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Funding. Authors wish to thank for their support to the Ministry of Innovation, Science and Technology of Spain (AGL2012-39028-C03 and AGL2016-74882-C3), the Department of Economy and Infrastructure of the regional government of Extremadura, Spain (Group CTS001 and project IB16073), the University of Extremadura (Group MIVET), the Spanish Ministry of Agriculture, and the Autonomous Community of Madrid (S2009/AGR-1489 and S2013/ABI-2747), the FPI program (BES-2013–065003) from the Spanish Ministry of Economy and Competitiveness, Fundação para a Ciência e Tecnologia (FCT), through the strategic project UID/MAR/04292/2013 granted to MARE and the Integrated Programme of SR&TD “Smart Valorization of Endogenous Marine Biological Resources Under a Changing Climate” (reference Centro-01-0145-FEDER- 000018), co-funded by Centro 2020 program, Portugal 2020, European Union, through the European Regional Development Fund.
References
- Abril C., Brodard I., Perreten V. (2010). Two novel antibiotic resistance genes, tet(44) and ant(6)-Ib, are located within a transferable athogenicity island in Campylobacter fetus subsp fetus. Antimicrob. Agents Chemother. 54 3052–3055. 10.1128/AAC.00304-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Akiba M., Lin J., Barton Y. W., Zhang Q. J. (2006). Interaction of CmeABC and CmeDEF in conferring antimicrobial resistance and maintaining cell viability in Campylobacter jejuni. J. Antimicrob. Chemother. 57 52–60. 10.1093/jac/dki419 [DOI] [PubMed] [Google Scholar]
- Cagliero C., Mouline C., Cloeckaert A., Payot S. (2006). Synergy between efflux pump CmeABC and modifications in ribosomal proteins L4 and L22 in conferring macrolide resistance in Campylobacter jejuni and Campylobacter coli. Antimicrob. Agents Chemother. 50 3893–3896. 10.1128/AAC.00616-06 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carreira A. C., Clemente L., Rocha T., Tavares A., Geraldes M., Barahona M. J., et al. (2012). Comparative genotypic and antimicrobial susceptibility analysis of zoonotic Campylobacter species isolated from broilers in a nationwide survey Portugal. J. Food Prot. 75 2100–2109. 10.4315/0362-028X.JFP-12-183 [DOI] [PubMed] [Google Scholar]
- Clinical and Laboratory Standards Institute [CLSI] (2010). Methods for Antimicrobial Dilution and Disk Susceptibility Testing for Infrequently-Isolated or Fastidious Bacteria: Approved Guidelines Approved Guidelines (M45-A). Wayne, PA: CLSI. [Google Scholar]
- Collective Eurosurveillance Editorial Team (2015). The 2013 joint ECDC/EFSA report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks published. Euro Surveill. 20:21021. 10.2807/ese.20.04.21021-en [DOI] [PubMed] [Google Scholar]
- Corcoran D., Quinn T., Cotter L., Fanning S. (2006). An investigation of the molecular mechanisms contributing to high-level erythromycin resistance in Campylobacter. Int. J. Antimicrob. Agents 27 40–45. 10.1016/j.ijantimicag.2005.08.019 [DOI] [PubMed] [Google Scholar]
- Florez-Cuadrado D., Ugarte-Ruiz M., Quesada A., Palomo G., Domínguez L., Porrero M. C. (2016). Description of an erm(B)-carrying Campylobacter coli isolate in Europe. J. Antimicrob. Chemother. 71 841–843. 10.1093/jac/dkv383 [DOI] [PubMed] [Google Scholar]
- Gibreel A., Skold O., Taylor D. E. (2004). Characterization of plasmid-mediated aphA-3 kanamycin resistance in Campylobacter jejuni. Microb. Drug Resist. 10 98–105. 10.1089/1076629041310127 [DOI] [PubMed] [Google Scholar]
- Hormeño L., Palomo G., Ugarte-Ruiz M., Porrero M. C., Borge C., Vadillo S., et al. (2016). Identification of the main quinolone resistance determinant in Campylobacter jejuni and Campylobacter coli by MAMA-DEG PCR. Diagn. Microbiol. Infect. Dis. 84 236–239. 10.1016/j.diagmicrobio.2015.11.002 [DOI] [PubMed] [Google Scholar]
- Jeon B., Wang Y., Hao H., Barton Y.-W., Zhang Q. (2011). Contribution of CmeG to antibiotic and oxidative stress resistance in Campylobacter jejuni. J. Antimicrob. Chemother. 66 79–85. 10.1093/jac/dkq418 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leverstein-van Hall M. A., Paauw A., Box A. T. A., Blok H. E. M., Verhoef J., Fluit A. C. (2002). Presence of integron-associated resistance in the community is widespread and contributes to multidrug resistance in the hospital. J. Clin. Microbiol. 40 3038–3040. 10.1128/JCM.40.8.3038-3040.2002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lévesque C., Piché L., Larose C., Roy P. H. (1995). PCR mapping of integrons reveals several novel combinations of resistance genes. Antimicrob. Agents Chemother. 39 185–191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin J., Akiba M., Sahin O., Zhang Q. J. (2005). CmeR functions as a transcriptional repressor for the multidrug efflux pump CmeABC in Campylobacter jejuni. Antimicrob. Agents Chemother. 49 1067–1075. 10.1128/AAC.49.3.1067-1075.2005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo N., Sahin O., Lin J., Michel L. O., Zhang Q. J. (2003). In vivo selection of Campylobacter isolates with high levels of fluoroquinolone resistance associated with gyrA mutations and the function of the CmeABC efflux pump. Antimicrob. Agents Chemother. 47 390–394. 10.1028/AAC.47.1.390-394.2003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moazed D., Noller H. F. (1987). Interaction of antibiotics with functional sites in 16S ribosomal RNA. Nature 327 389–394. 10.1038/327389a0 [DOI] [PubMed] [Google Scholar]
- Moreno M. A., Domínguez L., Teshager T., Herrero I. A., Porrero M. C. (2000). Antibiotic resistance monitoring: the Spanish programme. The VAV Network. Red de Vigilancia de Resistencias Antibióticas en Bacterias de Origen Veterinario. Int. J. Antimicrob. Agents 14 285–290. 10.1016/S0924-8579(00)00138-2 [DOI] [PubMed] [Google Scholar]
- Nirdnoy W., Mason C. J., Guerry P. (2005). Mosaic structure of a multiple-drug-resistant, conjugative plasmid from Campylobacter jejuni. Antimicrob. Agents Chemother. 49 2454–2459. 10.1128/AAC.49.6.2454-2459.2005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- O’Halloran F., Lucey B., Cryan B., Buckley T., Fanning S. (2004). Molecular characterization of class 1 integrons from Irish thermophilic Campylobacter spp. J. Antimicrob. Chemother. 53 952–957. 10.1093/jac/dkh193 [DOI] [PubMed] [Google Scholar]
- Olkkola S., Culebro A., Juntunen P., Hanninen M.-L., Rossi M. (2016). Functional genomics in Campylobacter coli identified a novel streptomycin resistance gene located in a hypervariable genomic region. Microbiology 162 1157–1166. 10.1099/mic.0.000304 [DOI] [PubMed] [Google Scholar]
- Olkkola S., Juntunen P., Heiska H., Hyytiainen H., Hanninen M. L. (2010). Mutations in the rpsL gene are involved in streptomycin resistance in Campylobacter coli. Microb. Drug Resist. 16 105–110. 10.1089/mdr.2009.0128 [DOI] [PubMed] [Google Scholar]
- Ouellette M., Gerbaud G., Lambert T., Courvalin P. (1987). Acquisition by a Campylobacter-like strain of aphA-1, a kanamycin resistance determinant from members of the family Enterobacteriaceae. Antimicrob. Agents Chemother. 31 1021–1026. 10.1128/AAC.31.7.1021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petersen L., Larsen T. S., Ussery D. W., On S. L. W., Krogh A. (2003). RpoD promoters in Campylobacter jejuni exhibit a strong periodic signal instead of a-35 box. J. Mol. Biol. 326 1361–1372. 10.1016/S0022-2836(03)00034-2 [DOI] [PubMed] [Google Scholar]
- Piccirillo A., Dotto G., Salata C., Giacomelli M. (2013). Absence of class 1 and class 2 integrons among Campylobacter jejuni and Campylobacter coli isolated from poultry in Italy. J. Antimicrob. Chemother. 68 2683–2685. 10.1093/jac/dkt242 [DOI] [PubMed] [Google Scholar]
- Pinto-Alphandary H., Mabilat C., Courvalin P. (1990). Emergence of aminoglycoside resistance genes aadA and aadE in the genus Campylobacter. Antimicrob. Agents Chemother. 34 1294–1296. 10.1128/AAC.34.6.1294 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qin S., Wang Y., Zhang Q., Chen X., Shen Z., Deng F., et al. (2012). Identification of a novel genomic island conferring resistance to multiple aminoglycoside antibiotics in Campylobacter coli. Antimicrob. Agents Chemother. 56 5332–5339. 10.1128/AAC.00809-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaw K. J., Rather P. N., Hare R. S., Miller G. H. (1993). Molecular genetics of aminoglycoside resistance genes and familial relationships of the aminoglycoside-modifying enzymes. Microbiol. Rev. 57 138–163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spies F. S., da Silva P. E. A., Ribeiro M. O., Rossetti M. L., Zaha A. (2008). Identification of mutations related to streptomycin resistance in clinical isolates of Mycobacterium tuberculosis and possible involvement of efflux mechanism. Antimicrob. Agents Chemother. 52 2947–2949. 10.1128/AAC.01570-07 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ugarte-Ruiz M., Wassenaar T. M., Gomez-Barrero S., Porrero M. C., Navarro-Gonzalez N., Dominguez L. (2013). The effect of different isolation protocols on detection and molecular characterization of Campylobacter from poultry. Lett. Appl. Microbiol. 57 427–435. 10.1111/lam.12130 [DOI] [PubMed] [Google Scholar]
- Vakulenko S. B., Mobashery S. (2003). Versatility of aminoglycosides and prospects for their future. Clin. Microbiol. Rev. 16 430–450. 10.1128/CMR.16.3.430-450.2003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Essen-Zandbergen A., Smith H., Veldman K., Mevius D. (2007). Occurrence and characteristics of class 1, 2 and 3 integrons in Escherichia coli, Salmonella and Campylobacter spp. in the Netherlands. J. Antimicrob. Chemother. 59 746–750. 10.1093/jac/dkl549 [DOI] [PubMed] [Google Scholar]
- Wieczorek K., Osek J. (2013). Antimicrobial resistance mechanisms among Campylobacter. Biomed. Res. Int. 2013:340605 10.1155/2013/340605 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woodcock J., Moazed D., Cannon M., Davies J., Noller H. F. (1991). Interaction of antibiotics with A- and P-site-specific bases in 16S ribosomal RNA. EMBO J. 10 3099–3103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao S., Tyson G. H., Chen Y., Li C., Mukherjee S., Young S., et al. (2016). Whole-genome sequencing analysis accurately predicts antimicrobial resistance phenotypes in Campylobacter spp. Appl. Environ. Microbiol. 82 459–466. 10.1128/AEM.02873-15 [DOI] [PMC free article] [PubMed] [Google Scholar]